Figure S1. MUT-16 localization in L4 hermaphrodite, adult hermaphrodite, and adult male germlines. MUT-16 DAPI. Phillips et al. S-1. male.
|
|
|
- Sara Price
- 6 years ago
- Views:
Transcription
1 Supplementary Material for Phillips et al. Figure S1. MUT-16 localization in L4 hermaphrodite, adult hermaphrodite, and adult male germlines. Figure S2. Mutator foci and P granules localize independently of one another. Figure S3. Localization of MUT-16 and MUT-7 in CSR-1/WAGO class 22G mutants. Figure S4. ClustalW alignment of mut-16 in Caenorhabditis species. Figure S5. Characterization of HA::ego-1 FLAG::rrf-1 transgene. Table S1. List of genes tested by RNAi for effect on Mutator foci. Table S2. Homology and amino acid composition of mut-16 orthologs. Table S3. Classification of low and high sirna yielding genes (separate file). Table S4. sirna reads from high sirna yielding transposons. Table S5. Strains used. Table S6. Primers used. L4 3 day adult Figure S1. MUT-16 localization in L4 hermaphrodite, adult hermaphrodite, and adult male germlines. male MUT-16 DAPI Phillips et al. S-1
2 A B C MUT- 16 association with nuclear pores MUT- 16 nuclear pores DAPI merge D MUT- 7 association with P granules MUT- 7 PGL- 1 DAPI merge P granule components localizes independently of mutator proteins wild- type mut- 16 (pk710) rde- 2 (pk1657) mut- 2 (r459) mut- 7 (pk720) mut- 15 (tm1358) MUT- 16 localizes independently of P granule components control RNAi MUT- 16 PGL- 1 mut- 14 (pk738) glh- 1 RNAi MUT- 16 PGL- 1 pgl- 1 RNAi MUT- 16 PGL- 1 Figure S2. Mutator foci and P granules localize independently of one another. Phillips et al. S-2
3 MUT-16::GFP MUT-7::GFP rrf-1(nec1) ego-1(om97) drh-3 (tm1217) ekl-1 (ok1197) Figure S3. Localization of MUT-16 and MUT-7 in CSR-1/WAGO class 22G mutants. Phillips et al. S-3
4 Figure S4. ClustalW alignment of mut-16 in Caenorhabditis species. briggsae_mut-16 MSGLCENSFEQRTVLIGSHLGISATGLVMTSQEDDYPDDFDITTSEQ------GDADDIQ 54 remanei_mut MTEVNDEDYPEELDLSHTTEENEGIVGSVPSDE 33 brenneri_mut MSHSDDDYPELDVSTDGNEN----ALTAADGY 28 elegans_mut MSESDDDYPELDISDQYIDPLGIVVGPPPASY 32 japonica_mut MSDESYADLDLDSSYNEE EQPVQ 23.::.*.: : briggsae_mut-16 NHCLGAPPISDFDSNEEQ DSENELEDSYDSGSEPFDVDGYDYTT-EPL 101 remanei_mut-16 MHFIGAPPGSDSEVDEEYR QYQRDPYSDYSSSDNEFDCDDYLKNVPREL 82 brenneri_mut-16 GEVVGAPPSSDVESDESAYHTPSSEENVIRQYEYNECDLYDSGSEEFDVEGYENEQLADL 88 elegans_mut-16 TETDREETPMQNRTEDDTN------SYGNSSGEHDYDSYLDSGDDDFDVDAYYANDNMDE 86 japonica_mut-16 NHVIGPAPTSSEDDEMDTR SVDSYMSGESDYDLENATPYIEPSVEELAKLD : : briggsae_mut-16 PAMPDD---LVGNGAQN--YADMNDFGGDPRELLPIFFTSTLAAHPLRLKEGYDPEELDK 156 remanei_mut-16 PELTND---LIVSGSDNTRLQELQDLG-DPVEFLKVFLTASMMGQPLKVKDGFDPEDLDK 138 brenneri_mut-16 HIHDMSSYVDVGNEQESNAAHNFQEFEDSNKEIFQVLLPSAMIASIEHMTIGFEPSVIND 148 elegans_mut-16 PPPETIPDNLIQNVIGRNDNADYDFSDASNPEIMKLFLSSSLLSNPLKFRSGYSSDELDD 146 japonica_mut-16 ENLEPELPEHLIAAENYSDDIYASNSNEYVETVIKYFLPAAICSTPLLLRNGFSPDEVDK 134 :..: ::.:::.. *:... ::. briggsae_mut-16 CLRENMGFTLPVVATILLGSHITDGLPNDSKTEYAIALANKNWLTRNGDKFLPIFPESEK 216 remanei_mut-16 CCSELMGITLTGVASVLLGAELQN-IPNRSKTEMAFALANRGWLRAVNSKFYPVIPETEN 197 brenneri_mut-16 CLKD-VGYCLEVVAMMLLGAEKFATIPRAP-VDWCHALKDANHLLFENGKFYPRVPESLR 206 elegans_mut-16 CLKSCMGYSLQVTAVLLLPPEIISQLPNDSKTEHAHALVRGGWLQSKEGAFFPVISDSER 206 japonica_mut-16 CLTDEGQISLEATAYILLGARIVENLPNKSKTEFCHALVKAGWLTEKQGKFFPVLPESEK 194 *. *.* :**.. :*...:. **. *. * *..::. briggsae_mut-16 EFITSLIEGSELLASKEETKKKEAQVFPSMEKEIRAHQTFNMIVELLQIVRDELRQRKVP 276 remanei_mut-16 DFVLSLIEGAERLQHLDENKKREAEHYPSAEKEINALISFNMIVELMDAVREEYRIMLLP 257 brenneri_mut-16 AHCTSLIEGASRLSQFEEKKKKEQETYKSEEFELRVLYTFNMIAELLNAVRDEQRQLKIG 266 elegans_mut-16 ETVVSLMNGSEEQHKRQERKKKEADTFESEEKEIRTLLTFNMIAELLMAVRNEYSIRSVK 266 japonica_mut-16 EFILELIDGAENTKYYEERKKKEAEQYPSEEAEIRAHITFNFICELLSAVQRELQQNKIK 254.*::*:. :* **:* : : * * *:.. :**:* **: *: * : briggsae_mut-16 YQNIMFAFNEMAKGKKHAHVFEKYSKQLELDS-TMEWNTKWFNQYTNRSTLKKFVQMSKF 335 remanei_mut-16 YQTLSNGYLDMVTGKKYPEIHKKYFNKLNLDE-KKVWNSEWFNAFTGRSTLKKFVQMARF 316 brenneri_mut-16 YSHLMNAYQAMVEGLKYHDIFKRYKGTLSLNDPKKLWNSQWFNEYTGRSSLMKFIHMARF 326 elegans_mut-16 YQILSTAYTNMVTGAAHANIFRKYKDILQLDP-EKLWNNDWFKEYTNRGTLKKFLTTARF 325 japonica_mut-16 YQTLSTEFEKLVSGQKNANLFSKYKHLLDLES-DAKWDSEWFKKYTNRSSLKKFVTMPKF 313 *. : : :. *.:. :* *.*: *:..**: :*.*.:* **:.:* briggsae_mut-16 SEIVVSNMK--DSPFYFRADDEG-DRPVKMFTDDDIKAVKDKWMSGNERNKNFGS--NYE 390 remanei_mut-16 AEIVVVENP--TSQFCFRADSE--NRPVRMFTANDIKDVEEKWSSGNEKNKNFGR--NYG 370 brenneri_mut-16 SEIAVTNTN--PRTYYFRADDDVPQRDVKLFIKKDFDEVREKWRSGNQANRNFGR--NYG 382 elegans_mut-16 SEIVVSQANGKTVELYFRADDEG-NRPVVLFTDEHIADVRNKWKTGNQRNQNYGSQGNYR 384 japonica_mut-16 AAIVADQTS----EFRFRTDNVH------LFTNEDLDEVRQRWR : *.. : **:*. :*..: *.::* briggsae_mut-16 RSGQRGDRGGPSRYQQQPKKVIEQDPNYRSSTFNGGISGA-ANDDGSLEPSSSSRTYDNQ 449 remanei_mut-16 GQGQRGLPRGAS---QQPKRKIEQDPNYRSSTFAGGIN-D-DDDDGSLMPSTSARENQAG 425 brenneri_mut-16 AGGQRGEGRGPS---QQKKKTIEQDPNYKSARFNGLMD---GVDDGSLEPSTSTRYNRE- 435 elegans_mut-16 AGGQRSDDRRGP---QQRRNVIVPDPNYQPSTFAGGISNN-ADDDGSLQPTTSSHFNRNT 440 japonica_mut-16 RPAARADRRWQSGSQNAKSRENVPDPDYRPAAVSGGISALDEDDDGSLRPTTSSNHRNRH 407. *.. :. **:*:.:. * :. ***** *::*:. briggsae_mut-16 PSSRTR CHRSPTPSPERRRENPVAERSTIRDESTQRRSESPPPVTR 495 remanei_mut-16 PSSSSQPV YGRRSPSPTVRNQRARSPSQESRNSNTVPQSSYRPIDPFAG 474 brenneri_mut RSPPRPIRNRSLSPRENRSPIPARSPSPFDEPTSQAQE 473 elegans_mut-16 DRSTSRP PRAPTSPVNRVMETDPLMGQGTSSGAPQRSAIP-NPFGG 485 japonica_mut-16 HSPTATIRDSSRTRPTTSSHPARSRSPSITTKTTRTRRDSSENPPPEPLNMEPAANPFGG 467 *:..... briggsae_mut-16 EVFDPFGGGPPLNQKTEDNRGNNRGRGFGNGNFSS remanei_mut-16 AMVHPISAGVNGNRNGQGGYETRRLQTFGASSSESDEVSEGCYSDDDPEDQKRKTIERNE 534 brenneri_mut-16 QAARPSSPE elegans_mut-16 APALSRSTITNGNRGPSYGDRGERVQDVGDTTSDSEITSEGSYSD japonica_mut-16 PLPPPARAEHRPAHQPPPPNFPRTADYAGYSSTEDDITSDGSYTD Phillips et al. S-4
5 briggsae_mut LAKPVARIDHIGDTSSETSSRFNT AYSPPRRRTPSPKPRRNV 572 remanei_mut-16 KRLRKEAREMERRKKHKAEVHVPPVTRFQTNPFYKHKSARAPEPANPRASTAVEQEEREE 594 brenneri_mut SQSDPPQRVALSDPFGGR 500 elegans_mut EDPEQKEIKRQRRKDKLKKKQERELRSREKHTKSKQQPPSKIETRFNTYKKKSES 585 japonica_mut EKSEEKERRRDDRQKKLAKKQKKLME RQNAPPKPPPDNSRNRFRVSS 559. briggsae_mut-16 SAEPEIPSPQPQPTQNQTPSPFLGQESPPS---IDNDRMDDYMPPPPAAPVVRQLPLAVS 629 remanei_mut-16 LVVPPRQSPFDAPSALAPAPAAPAPVSQPSNSDAQVNAHEDVLPPPPAAVAANTLPLFVS 654 brenneri_mut-16 PVQRREETNFGGSSDNQLTRDNYGR NRAGEVRRIDVHLI 539 elegans_mut-16 SATDTSNTPPVDTVNVALPTPVVESSSTTAAPSIPVSTRPEVVVPPENPAPLREVGNFYS 645 japonica_mut-16 FAPVPESPTPPPPAAPAAPPPLQETRNDPD PPPSLSGQIPLTIS : briggsae_mut-16 SSRHDA-EHMTAQVPYRPAEGVLERNKAEEEKKKRRQEIDDVRIESENRQPPTQHPFLTS 688 remanei_mut-16 SSTHDQ-DRCSAQMPYKPAEGVLERNKQEEAERLRREQLSEMRMRSGFNQVPASHPWMSA 713 brenneri_mut-16 GETSSE-GEITSDGSYSNED-----EDTKQAKRERRLAVKKKKDEREMR elegans_mut-16 KSNHDE-DRRNVQLPFTPADTHKPIKVAPKEPVRNPLLKERPSANGFINRRLPSHPAPPP 704 japonica_mut-16 ESRFDEEDQARGSIPFRPMPTSRPVEDERAAPVPPVPAPQPRTRVVHPDIARRQRDQNES : :. briggsae_mut-16 S------RAKPTTSRNPEQGYSDPVTTNGFARRNVAQPTYAATMAAQAPPAPPAPAPQPS 742 remanei_mut-16 ASKPGPSQPQAYEEPAPTKGFGTTLPPTVQRSVPQPPPQQAPQMQQQQEPIYSTPFVETV 773 brenneri_mut RRKQHEENRNKPIEQTRFSVSQFKKKTPQNAPRSPERRRQESPLPFEDEPTTS 635 elegans_mut-16 VN-----QSQPANQPMQTAVYQNSHPGAPYIPQQPTYQPQLPVQQPQPHQYAPQPIHHQQ 759 japonica_mut-16 VSRPMEPGVPGGFRGARKVEPAQAPPPPPPQTAPPPQPPRQPHQQPPPPLHNYQQFSDNR briggsae_mut-16 PVP PPVFPLHAPMQAPLQVPVSRTNSVIKIILKPSILE 780 remanei_mut-16 SLSSLIVLNNSLGFQYQGRNRQGMAPQNYAQPAPQNYAQPAPQNYAQPAPSQQQYPQHQN 833 brenneri_mut-16 TEP LPPVELPVEEPTPVQTTTADSDDEPLPVRPQ 669 elegans_mut-16 PIHQP MHGQQYPPVNQQQPIYQQPAPQYPPYNSIQN 795 japonica_mut-16 RGSQDYG GMNGRPESPPPIPEQQHHYQAPEPQYPHYQQQPN : briggsae_mut-16 EIDKEGGHLNRGYPLPPQQYPPQNQYQPPLQQQIQ----QPNYQYPQQNQYQQGPPPPQY 836 remanei_mut-16 LVQQNPPLFPDHSASQPQYQPMQQQYQQPLPQNPYPIGSQSSSTLPQQQPYHHQNQDTYR 893 brenneri_mut-16 ATNKAPAVVTDTMTN ALSHQAQVPYR 695 elegans_mut-16 NPQHGPSPFNYSQVPQPAYNHVGQQPSHMSNQPHINQNGYQNSYNPNQGPTSSDPNYGCN 855 japonica_mut-16 PYHERNNQPNRQFAPQPNNQPYSMGEPEQPREYNMPNYYNPNYQPPPPPPQHQQQPVYGQ briggsae_mut-16 PMETPQYNPTPPPPPRAEYSSNYPPPQNQMNR-----PPQQSYQDQQYSRNQYSSNGNWS 891 remanei_mut-16 PTSRNHANPQHMDNQYSSMRDAMRPPVPRNDMNYDSIPSNSLHQIPPMNQPQYSSQQRQY 953 brenneri_mut-16 PSALDVQKKKMEQEEKAKKASMN AVKKYGADAVLAYHPALFSQNSGPG 743 elegans_mut-16 PQFNHYGSRSVYHEDHSSQRRRSPDQFPPNPP---EYDPHGNFKLADYERDRMTVGYSQN 912 japonica_mut-16 NYHDPPRESRSSHFNNFQQPPVQSAPPPPHTSRNSHHQPFDHYFPSGNGNAHYNSNNGYN briggsae_mut-16 NPAYPQQPRSPPLPNGPPQDYNRWRDRDPPPRQPPPATYGRRDVGRPS remanei_mut-16 NDVYP--PVNPYNNRVPSTDFNDRRIRNEQWHQVQPGSYDPYGRSQNPGPGSSYNPRGEE 1011 brenneri_mut-16 PSSSPLPPPQAYIEPEPPSGFGNSQ elegans_mut-16 PHQFDHHGSHMPHQSQP-QGYDNFNGNSAPYFNKNGGQSNHQPEAQRS japonica_mut LPFANRQQQYHDQPPPPHNNDRWNPYATARRPPPATTGRGVSLLS *... briggsae_mut-16 --LSTFSLLAEEDDRRAAPQRPDVGRMRDIIMSIAYNCRTKGCQLDKERLKYEVCQSRFQ 997 remanei_mut-16 NCASFFGRLRKAAAASGGGESEEVKQIKIEIQRIVFDYASENRELTLSELKANLVRKMRH 1071 brenneri_mut elegans_mut FSVLSSNRQPSNRELIFQDGIEKELRDIILRYRSMNLTVLTVQELRTEVSRRPAI 1014 japonica_mut SHPRDSTQLSAEEDRRIQRGRMSVMRYMEDCERSRSQVTGYDLRSAQHNGEVH 982 briggsae_mut-16 QHFPGGPEWFDFTSFIRNELRGTMEVRGNENGAVWYELLRN remanei_mut-16 LQF------FDVHEFIQTYLRNQVVIVNNGPYGPVVRPT brenneri_mut RFVDYIKRFFV elegans_mut-16 PRY------IDIVQYIRDSSSVAIVERGDIEPYVVLKDDIRN japonica_mut-16 IGGN-----ENIIAFIRRYMSAIVGIGFGRDANGQEIDIFYVIQD :*: Phillips et al. S-5
6 A Somatic RNAi Fertility lin-29 nhr-23 wild-type ego-1 rrf ego-1 rrf-1;; HA::ego-1 FLAG::rrf B EGO-1 PGL-1 merge RRF-1 PGL-1 merge Figure S5. Characterization of HA::ego-1 FLAG::rrf-1 transgene. Phillips et al. S-6
7 Supplemental Figure Legends Figure S1. MUT-16 localization in L4 hermaphrodite, adult hermaphrodite, and adult male germlines. MUT-16 foci are present in the transition zone region of L4 hermaphrodite, three day old adult hermaphrodite, and adult male dissected germlines stained with DAPI (blue) and anti-gfp (recognizing MUT-16::GFP). All images are projections of 3D images following deconvolution. Scale bars represent 5 µm. Figure S2. Mutator foci and P granules localize independently of one another. (A) MUT-16 (red) associates with nuclear pores (immunostaining with mab414 - green) in C. elegans germlines. Images are single focal planes following taken from a 3D image following deconvolution. (B) MUT-7 (red) costained with P granules (immunostaining with PGL-1 antibody K76 green) at the diplotene stage of meiosis. Note that some P granules and Mutator foci have detached from the nuclear periphery and are now visible in the cytoplasm. Images are projections of 3D images following deconvolution. (C) MUT-16 (red) and PGL-1 (green) expression in adult C. elegans germlines feeding on E. coli expressing control (L4440), glh-1 and pgl-1 dsrna. PGL-1 and MUT-16 were visualized a PGL-1::RFP and MUT-16::GFP strain. Images are single focal planes from dissected (L4440 and glh-1) and intact (pgl-1) adult C. elegans. (D) Wild-type and mutator mutants were dissected and stained with DAPI (blue) and antibodies against PGL-1 (green). All images are projections of 3D images following deconvolution. Scale bars represent 5 µm. Figure S3. Localization of MUT-16 and MUT-7 in CSR-1/WAGO class 22G mutants. Balanced rrf-1(nec1) ego-1(om97), drh-3(tm1217); or ekl-1(ok1197) were introduced into the MUT-16::GFP or MUT-7::GFP strains. Homozygous rrf-1(nec1) ego-1(om97), drh-3(tm1217); or ekl-1(ok1197) animals were selected from heterozygous parents, and were subsequently dissected and stained with DAPI (blue) and anti- GFP (red). All images are projections of 3D images following deconvolution. Scale bars represent 5 µm. Figure S4. ClustalW alignment of mut-16 in Caenorhabditis species. Glutamines are highlighted in yellow, asparagines are highlighted in green, and prolines are highlighted in blue. Figure S5. Characterization of HA::EGO-1 FLAG::RRF-1 transgene. (A) Phenotypes of ego-1 rrf-1 mutant worms in the presence and absence of a rescuing transgene. Fertility scored as presence of viable progeny (+++) or complete sterility (-). lin-29 RNAi scored as 100% vulval bursting (+++) or 100% viable adults (-). nhr-23 RNAi scored as 100% larval arrest (+++) or 100% viable adults (-). (B) HA::EGO-1 (red, anti-ha immunostaining top panels) or FLAG::RRF-1 (red, anti-flag immunostaining bottom panels) costained with PGL-1 (green, anti-pgl-1(k76) immunostaining). DNA is counterstained with DAPI (blue, in merge). All images are projections of 3D images following deconvolution. Scale bars represent 5 µm. Phillips et al. S-7
8 Gene Description sirna Pathway +/- mutator foci L4440 negative control + mut-16 Mutator - Q/N rich domain WAGO-class 22G, ERGO-1-class 26G - mut-7 Mutator - 3'-5' exonuclease WAGO-class 22G, ERGO-1-class 26G - mut-2 Mutator - nucleotidyl transferase WAGO-class 22G, ERGO-1-class 26G + mut-15 Mutator - novel probable WAGO-class 22G + ergo-1 Argonaute ERGO-1-class 26G + dcr-1 DICER RNase III exo-sirna, ERGO-1 and ALG-3/4-class 26G + rde-4 dsrna binding protein exo-sirna, ERGO-1 and ALG-3/4-class 26G + ekl-1 Tudor domain WAGO-class 22G,CSR-class 22G + csr-1 Argonaute CSR-class 22G + cde-1 nucleotidyl transferase CSR-class 22G + rde-1 Argonaute exo-sirna + drh-1 DEAD box helicase exo-sirna + rsd-2 Required for sirna spreading + ppw-2 Argonaute + sago-1 Argonaute + Table S1. RNAi of each of the indicated genes was performed using the MUT-7::GFP strain, and scored for the presence or absence of Mutator foci. Negative (L4440 empty vector) and positive (mut-16 and mut-7) controls are included. Phillips et al. S-8
9 Table S2. Homology and amino acid composition of mut-16 orthologs BLAST results using N-terminal (1-530) or C-terminal ( ) regions of C. elegans mut-16 mut-16 (N-term.) % Coverage % Identity % Positive C. briggsae (29-557) 100% 42% 56% C. remanei (6-519) 99% 40% 56% C. brenneri (1-556) 99% 38% 54% mut-16 (C-term.) % Coverage % Identity % Positive C. briggsae none N/A N/A C. remanei none N/A N/A C. brenneri none N/A N/A Percentage glutamine, asparagine and proline residues in full length and C-terminal regions of mut-16 orthologs mut-16 (full length) Q/N Q N P C. elegans (1-1050) 14.8% 7.4% 7.4% 10.1% C. briggsae (1-1038) 13.1% 7.0% 6.1% 12.0% C. remanei (1-1104) 15.0% 8.3% 6.7% 10.0% C. brenneri (1-779) 10.4% 5.0% 5.4% 7.7% C. japonica (1-1022) 12.5% 6.4% 6.1% 13.4% mut-16 (C-term.) Q/N Q N P C. elegans ( ) 28.5% 18.2% 10.3% 17.0% C. briggsae ( ) 21.6% 14.4% 7.2% 24.4% C. remanei ( ) 27.6% 19.1% 8.5% 17.0% C. brenneri ( ) 11.8% 7.5% 4.3% 15.6% C. japonica ( ) 23.6% 13.3% 10.3% 24.3% Phillips et al. S-9
10 Table S4. sirna reads from high sirna yielding transposons*. Sequence Name wt RPM mut-16 RPM mut-16/wt Chromosome Genomic Cooridinates Transposon Classification Y102A5D V Retrotransposon-like T27C V Retrotransposon-like T05C V Unknown T05H V DDE Transposase family T23B V Unknown Y20F I Retrotransposon-like K03D IV Unknown F14D II Tc8/TURMOIL1/Tourist K03D IV Retrotransposon-like T23B V Retrotransposon-like F58H IV Retrotransposon-like R06A II DDE Transposase family Y73B3A X DDE Transposase family R07H IV Tc5 C03A V Retrotransposon-like Y43F4A III Retrotransposon-like R13D V Retrotransposon-like Y48G1BM I Retrotransposon-like Y37H2A V Retrotransposon-like F30F I Tc4 H25K IV Retrotransposon-like F07B V Retrotransposon-like K10E I Tc4 Y38H6C V Tc4 C44B IV Retrotransposon-like F21E X LINE2D_CE K06C V Retrotransposon-like Y54C5B II Tc5 M IV Tc5 Y38F2AR IV Tc5 F57G V TURMOIL2/Tourist none II Tc5 none IV Tc5 F26H I DDE Transposase family T07G IV Tc5 T25G X Unknown F38E X LINE2A_CE ZK II Retrotransposon-like C40A II Retrotransposon-like C27H IV DDE Transposase family H28G X DDE Transposase family Y51A2D V DDE Transposase family W06G V DDE Transposase family T14G X DDE Transposase family K03H IV DDE Transposase family C52D IV DDE Transposase family ZK V Retrotransposon-like W04G I DDE Transposase family T16A II Unknown Multiple* Multiple ~32 loci Tc1 family Multiple* Multiple ~14 loci rte-1 family Multiple* Multiple ~22 loci Tc3 family * Tc1, rte-1 and Tc3 family transposons produced too few sirna reads to be classified as high sirna yielding transpsosons after accounting for copy number. Phillips et al. S-10
11 Table S5. Strains used in this study. Strain name Genotype Description N2 wild-type NL1810 mut-16(pk710) I - outcrossed 4x NL3531 rde-2(pk1657) I - outcrossed 4x TW332 mut-2(r459) I - outcrossed 4x WM30 mut-2(ne298) I - outcrossed 4x NL1820 mut-7(pk720) III - outcrossed 4x GR1747 mut-15(tm1358) V - outcrossed 4x NL1838 mut-14(pk738) V - outcrossed 4x WM187 avr-14(ad1302) rrf-1(nec1) ego-1(om97)/ht2[qis48] I; +/ht2 III; avr-15(ad1051) glc-1(pk54) V GR1748 unc-119(ed3) III; mgsi2[mut-16p::mut-16::gfp::mut-16-3'utr] IV mut-16::gfp GR1749 mut-7(pk720) unc-119(ed3) III; mgsi13[mut-7p::mut-7::gfp::mut-7-3'utr] IV mut-7::gfp GR1750 mgsi14[mut-7p::mut-7::mcherry::mut-7-3'utr] II; mut-7(pk720) unc-119(ed3/ed9) III mut-7::mcherry GR1751 unc-119(ed3/ed9) III; mgsi15[mut-14p::mut-14::mcherry::mut-14-3'utr] II; mut-14(pk738) V mut-14::mcherry GR1752 mut-2(ne298) I; unc-119(ed3) III; mgsi16[mut-2p::mut-2::gfp::mut-2-3'utr] IV mut-2::gfp GR1753 mut-2(ne298) I; unc-119(ed3/ed9) III; mgsi17[mut-2p::mut-2::mcherry::mut-2-3'utr] II mut-2::mcherry GR1754 unc-119(ed3/ed9) III; mgsi18[mut-15p::mut-15::mcherry::mut-15-3'utr] II; mut-15(tm1358) V mut-15::mcherry GR1755 rde-2(pk1657) I; unc-119(ed3) III; mgsi19[rde-2p::rde-2::gfp::rde-2-3'utr] IV rde-2::gfp GR1756 rde-2(pk1657) I; unc-119(ed3/ed9) III; mgsi20[rde-2p::rde-2::mcherry::rde-2-3'utr] II rde-2::mcherry GR1757 unc-119(ed3) III; mgsi2[mut-16p::mut-16::gfp::mut-16-3'utr] IV; [nmy-2::pgl-1::mrfp; unc-119] mut-16::gfp; pgl-1::rfp GR1759 GR1760 GR1761 GR1762 GR1763 GR1764 GR1765 GR1766 GR1767 GR1768 GR1769 GR1770 GR1771 GR1772 GR1773 GR1774 GR1775 GR1776 GR1777 GR1778 GR1779 GR1780 GR1781 GR1782 GR1783 mut-2(ne298) I; unc-119(ed3) III; mgsi2[mut-16p::mut-16::gfp::mut-16-3'utr] IV mut-7(pk720) unc-119(ed3) III; mgsi2[mut-16p::mut-16::gfp::mut-16-3'utr] IV rde-2(pk1657) I; unc-119(ed3) III; mgsi2[mut-16p::mut-16::gfp::mut-16-3'utr] IV unc-119(ed3) III; mgsi2[mut-16p::mut-16::gfp::mut-16-3'utr] IV; mut-14(pk738) V unc-119(ed3) III; mgsi2[mut-16p::mut-16::gfp::mut-16-3'utr] IV; mut-15(tm1358) V mut-2(ne298) I; mut-7(pk720) unc-119(ed3) III; mgsi13[mut-7p::mut-7::gfp::mut-7-3'utr] IV rde-2(pk1657) I; mut-7(pk720) unc-119(ed3) III; mgsi13[mut-7p::mut-7::gfp::mut-7-3'utr] IV mut-7(pk720) unc-119(ed3) III; mgsi13[mut-7p::mut-7::gfp::mut-7-3'utr] IV; mut-14(pk738) V mut-7(pk720) unc-119(ed3) III; mgsi13[mut-7p::mut-7::gfp::mut-7-3'utr] IV; mut-15(tm1358) V mut-16(pk710) I; mut-7(pk720) unc-119(ed3) III; mgsi13[mut-7p::mut-7::gfp::mut-7-3'utr] IV mut-2(ne298) I; mut-7(pk720) unc-119(ed3) III; mgsi16[mut-2p::mut-2::gfp::mut-2-3'utr] IV mut-2(ne298) rde-2(pk1657) I; unc-119(ed3) III; mgsi16[mut-2p::mut-2::gfp::mut-2-3'utr] IV mut-2(ne298) I; unc-119(ed3) III; mgsi16[mut-2p::mut-2::gfp::mut-2-3'utr] IV; mut-14(pk738) V mut-2(ne298) I; unc-119(ed3) III; mgsi16[mut-2p::mut-2::gfp::mut-2-3'utr] IV; mut-15(tm1358) V mut-2(ne298) mut-16(pk710) I; unc-119(ed3) III; mgsi16[mut-2p::mut-2::gfp::mut-2-3'utr] IV mut-2(ne298) rde-2(pk1657) I; unc-119(ed3) III; mgsi19[rde-2p::rde-2::gfp::rde-2-3'utr] IV rde-2(pk1657) I; mut-7(pk720) unc-119(ed3) III; mgsi19[rde-2p::rde-2::gfp::rde-2-3'utr] IV rde-2(pk1657) I; unc-119(ed3) III; mgsi19[rde-2p::rde-2::gfp::rde-2-3'utr] IV; mut-14(pk738) V rde-2(pk1657) I; unc-119(ed3) III; mgsi19[rde-2p::rde-2::gfp::rde-2-3'utr] IV; mut-15(tm1358) V rde-2(pk1657) mut-16(pk710) I; unc-119(ed3) III; mgsi19[rde-2p::rde-2::gfp::rde-2-3'utr] IV mut-2(ne298) I; mgsi18[mut-15p::mut-15::mcherry::mut-15-3'utr] II; unc-119(ed3/ed9) III; mut-15(tm1358) V mgsi18[mut-15p::mut-15::mcherry::mut-15-3'utr] II; mut-7(pk720) unc-119(ed3/ed9) III; mut-15(tm1358) V rde-2(pk1657) I; mgsi18[mut-15p::mut-15::mcherry::mut-15-3'utr] II; unc-119(ed3/ed9) III; mut-15(tm1358) V mgsi18[mut-15p::mut-15::mcherry::mut-15-3'utr] II; unc-119(ed3/ed9) III; mut-14(pk738) mut-15(tm1358) V mut-16(pk710) I; mgsi18[mut-15p::mut-15::mcherry::mut-15-3'utr] II; unc-119(ed3/ed9) III; mut-15(tm1358) V GR1784 mut-2(ne298) I; mgsi15[mut-14p::mut-14::mcherry::mut-14-3'utr] II; unc-119(ed3/ed9) III; mut-14(pk738) V GR1785 mgsi15[mut-14p::mut-14::mcherry::mut-14-3'utr] II; mut-7(pk720) unc-119(ed3/ed9) III; mut-14(pk738) V GR1786 rde-2(pk1657) I; mgsi15[mut-14p::mut-14::mcherry::mut-14-3'utr] II; unc-119(ed3/ed9) III; mut-14(pk738) V GR1787 mgsi15[mut-14p::mut-14::mcherry::mut-14-3'utr] II; unc-119(ed3/ed9) III; mut-14(pk738) mut-15(tm1358) V GR1788 mut-16(pk710) I; mgsi15[mut-14p::mut-14::mcherry::mut-14-3'utr] II; unc-119(ed3/ed9) III; mut-14(pk738) V GR1789 GR1790 GR1791 GR1792 avr-14(ad1302) ego-1(om97) rrf-1(nec1)/ht2[qis48] I; mut-7(pk720) unc-119(ed3)/ht2 III; mgsi13[mut-7p::mut-7::gfp::mut-7-3'utr] IV avr-14(ad1302) ego-1(om97) rrf-1(nec1)/ht2[qis48] I; unc-119(ed3)/ht2 III; mgsi2[mut-16p::mut-16::gfp::mut-16-3'utr] IV drh-3(tm1217) I/hT2[qIs48] I; unc-119(ed3)/ht2 III; mgsi2[mut-16p::mut-16::gfp::mut-16-3'utr] IV ekl-1(ok1197) I/hT2[qIs48] I; unc-119(ed3)/ht2 III; mgsi2[mut-16p::mut-16::gfp::mut-16-3'utr] IV GR1793 mut-2(ne298) I; mgsi17[mut-2p::mut-2::mcherry::mut-2-3'utr] II; unc-119(ed3/ed9) III; mgsi2[mut-16p::mut-16::gfp::mut-16-3'utr] IV mut-2::mcherry; mut-16:gfp GR1794 mgsi18[mut-15p::mut-15::mcherry::mut-15-3'utr] II; unc-119(ed3/ed9) III; mgsi2[mut-16p::mut-16::gfp::mut-16-3'utr] IV; mut-15(tm1358) V mut-15::mcherry; mut-16::gfp GR1795 rde-2(pk1657) I; mgsi20[rde-2p::rde-2::mcherry::rde-2-3'utr] II; unc-119(ed3/ed9) III; mgsi2[mut-16p::mut-16::gfp::mut-16-3'utr] IV rde-2::mcherry; mut-16::gfp GR1796 mgsi15[mut-14p::mut-14::mcherry::mut-14-3'utr] II; unc-119(ed3/ed9) III; mgsi2[mut-16p::mut-16::gfp::mut-16-3'utr] IV; mut-14(pk738) V mut-14::mcherry; mut-16::gfp GR1797 mgsi14[mut-7p::mut-7::mcherry::mut-7-3'utr] II; mut-7(pk720) unc-119(ed3) III; mgsi2[mut-16p::mut-16::gfp::mut-16-3'utr] IV mut-7::mcherry; mut-16::gfp GR1798 mut-2(ne298) I; mgsi14[mut-7p::mut-7::mcherry::mut-7-3'utr] II; mut-7(pk720) unc-119(ed3) III; mgsi16[mut-2p::mut-2::gfp::mut-2-3'utr] IV mut-7::mcherry; mut-2::gfp GR1912 ego-1(om97) rrf-1(nec1) I; mgsi24[ego-1p::3xha::ego-1::3xflag::rrf-1::rrf-1 3 UTR] II; unc-119(ed9) III HA::ego-1 FLAG:rrf-1 GR1913 ego-1(om97) rrf-1(nec1) I; mgsi24[ego-1p::3xha::ego-1::3xflag::rrf-1::rrf-1 3 UTR] II; unc-119(ed9) III; mgsi2[mut-16p::mut-16::gfp::mut-16-3 UTR] IV HA::ego-1 FLAG:rrf-1; mut- 16::gfp Phillips et al. S-11
12 Table S6. Primers used in this work. Cloning primers mut-16 promoter mut-16 gene mut-16 3'UTR mut-7 promoter mut-7 gene mut-7 3'UTR mut-2 promoter mut-2 gene mut-2 3'UTR rde-2 promoter rde-2 gene rde-2 3'UTR mut-15 promoter mut-15 gene mut-15 3'UTR mut-14 promoter mut-14 gene mut-14 3'UTR ego-1 promoter ego-1 gene/region between ego-1 and rrf-1 region between ego-1 and rrf-1 rrf-1 gene/3'utr ctgtgtttttctgccgtcgac tttctgcaaatatcgagtatttgag atgtccgaaagtgatgatgattatc gtttcggatatcatctttcaaaacg gaatttacttgttcctattactgttc aatacgtcatatcataaattcacaaag aacaataatggagaacgtttgtcg tttcgaaacggacaagcttgattc atggaagaagaaccgtacaaaag acattcctggctggtgctc attcgaaatcccgttccccttc gtagagccttgcgaaatgaac aaacactaatacatttcacttacaatc tgttgttactacaaaaaatattcaattattattc atgtctcaaccaaataaagatcag tacaaatgagttccgacagtag tcttaatatgtcgatgttttgtc ggattgtagttagtttgattgc gcaatgtttaaatatacggtaac ctgaatagcaagaaaatgaatatag atgcataatgggtaccattcttattttc ctttcgaactgaaataataatttgaatag aaactgcacaatcgacaaatcac gtttctctttgtaaacctgaaaaatg caaaaatcgattatttaacaaaaacg ttctttgatacttatctaaatacttc atggataactccaatggaatcg aacaggaacagctgatgttgc atatgtatttttcttccaacttctc acttcccgtctattaatctgtaaaac cgtcgattttgcagagttttgc ctgaaaaatatgattttctcgatc atgtcatatccctggagtaac agcccatgagtcgatattgttg aataattgatttactgtcagtagtatttag tgattcccattcctttgtttttg atggctgaagctgtttgctg tgttgcgaggattcgggata ggggacgaaggttatcgtgg ccgacatgacgaggagtctg agaggacacatcaacatctag gacgaggagtctgaaaaattttat tcggaacgaagtcatggattc aatgttcttggatcgaagacc Phillips et al. S-12
Two classes of silencing RNAs move between Caenorhabditis elegans tissues.
 Two classes of silencing RNAs move between Caenorhabditis elegans tissues. Antony M Jose, Giancarlo A Garcia, and Craig P Hunter. Supplementary Figures, Figure Legends, and Tables. Supplementary Figure
Two classes of silencing RNAs move between Caenorhabditis elegans tissues. Antony M Jose, Giancarlo A Garcia, and Craig P Hunter. Supplementary Figures, Figure Legends, and Tables. Supplementary Figure
b number of motif occurrences
 supplementary information DOI: 1.138/ncb194 a b number of motif occurrences number of motif occurrences number of motif occurrences I II III IV V X TTGGTCAGTGCA 3 8 23 4 13 239 TTGGTCAGTGCT 7 5 8 1 3 83
supplementary information DOI: 1.138/ncb194 a b number of motif occurrences number of motif occurrences number of motif occurrences I II III IV V X TTGGTCAGTGCA 3 8 23 4 13 239 TTGGTCAGTGCT 7 5 8 1 3 83
MUT-16 promotes formation of perinuclear Mutator foci required for RNA silencing in the C. elegans germline
 MUT-16 promotes formation of perinuclear Mutator foci required for RNA silencing in the C. elegans germline Carolyn M. Phillips, Taiowa A. Montgomery, Peter C. Breen, and Gary Ruvkun 1 Department of Molecular
MUT-16 promotes formation of perinuclear Mutator foci required for RNA silencing in the C. elegans germline Carolyn M. Phillips, Taiowa A. Montgomery, Peter C. Breen, and Gary Ruvkun 1 Department of Molecular
MUT-14 and SMUT-1 DEAD Box RNA Helicases Have Overlapping Roles in Germline RNAi and Endogenous sirna Formation
 Current Biology 24, 839 844, April 14, 2014 ª2014 Elsevier Ltd All rights reserved http://dx.doi.org/10.1016/j.cub.2014.02.060 MUT-14 and DEAD Box RNA Helicases Have Overlapping Roles in Germline RNAi
Current Biology 24, 839 844, April 14, 2014 ª2014 Elsevier Ltd All rights reserved http://dx.doi.org/10.1016/j.cub.2014.02.060 MUT-14 and DEAD Box RNA Helicases Have Overlapping Roles in Germline RNAi
mut-16 and other mutator-class genes modulate 22G and 26G sirna pathways in
 Supplemental Information mut-16 and other mutator-class genes modulate 22G and 26G sirna pathways in Caenorhabditis elegans. Chi Zhang, Taiowa A. Montgomery, Harrison W. Gabel, Sylvia E. J. Fischer, Carolyn
Supplemental Information mut-16 and other mutator-class genes modulate 22G and 26G sirna pathways in Caenorhabditis elegans. Chi Zhang, Taiowa A. Montgomery, Harrison W. Gabel, Sylvia E. J. Fischer, Carolyn
Trasposable elements: Uses of P elements Problem set B at the end
 Trasposable elements: Uses of P elements Problem set B at the end P-elements have revolutionized the way Drosophila geneticists conduct their research. Here, we will discuss just a few of the approaches
Trasposable elements: Uses of P elements Problem set B at the end P-elements have revolutionized the way Drosophila geneticists conduct their research. Here, we will discuss just a few of the approaches
Dual sgrna-directed gene knockout using CRISPR/Cas9 technology in Caenorhabditis. elegans
 Dual sgrna-directed gene knockout using CRISPR/Cas9 technology in Caenorhabditis elegans Xiangyang Chen 1, Fei Xu 1, Chengming Zhu 1, Jiaojiao Ji 1, Xufei Zhou 1, Xuezhu Feng 1*, and Shouhong Guang 1*
Dual sgrna-directed gene knockout using CRISPR/Cas9 technology in Caenorhabditis elegans Xiangyang Chen 1, Fei Xu 1, Chengming Zhu 1, Jiaojiao Ji 1, Xufei Zhou 1, Xuezhu Feng 1*, and Shouhong Guang 1*
Heme utilization in the Caenorhabditis elegans hypodermal cells is facilitated by hemeresponsive
 Supplemental Data Heme utilization in the Caenorhabditis elegans hypodermal cells is facilitated by hemeresponsive gene-2 Caiyong Chen 1, Tamika K. Samuel 1, Michael Krause 2, Harry A. Dailey 3, and Iqbal
Supplemental Data Heme utilization in the Caenorhabditis elegans hypodermal cells is facilitated by hemeresponsive gene-2 Caiyong Chen 1, Tamika K. Samuel 1, Michael Krause 2, Harry A. Dailey 3, and Iqbal
mod-1::mcherry unc-47::gfp RME ser-4::gfp vm2 vm2 vm2 vm2 VNC
 A mod-1::mcherry unc-47::gfp RME B ser-4::gfp vm2 vm2 VNC vm2 vm2 VN Figure S1 Further details of mod- 1 and ser- 4 reporter expression patterns. (A) Adult head region showing mod- 1::mCherry and unc-
A mod-1::mcherry unc-47::gfp RME B ser-4::gfp vm2 vm2 VNC vm2 vm2 VN Figure S1 Further details of mod- 1 and ser- 4 reporter expression patterns. (A) Adult head region showing mod- 1::mCherry and unc-
Transgene-assisted genetic screen identifies rsd-6 and novel genes as key components
 JVI Accepted Manuscript Posted Online 27 June 2018 J. Virol. doi:10.1128/jvi.00416-18 Copyright 2018 Long et al. This is an open-access article distributed under the terms of the Creative Commons Attribution
JVI Accepted Manuscript Posted Online 27 June 2018 J. Virol. doi:10.1128/jvi.00416-18 Copyright 2018 Long et al. This is an open-access article distributed under the terms of the Creative Commons Attribution
Supplementary Figure 1. Homozygous rag2 E450fs mutants are healthy and viable similar to wild-type and heterozygous siblings.
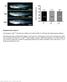 Supplementary Figure 1 Homozygous rag2 E450fs mutants are healthy and viable similar to wild-type and heterozygous siblings. (left) Representative bright-field images of wild type (wt), heterozygous (het)
Supplementary Figure 1 Homozygous rag2 E450fs mutants are healthy and viable similar to wild-type and heterozygous siblings. (left) Representative bright-field images of wild type (wt), heterozygous (het)
Mos1 insertion. MosTIC protocol-11/2006. repair template - Mos1 transposase expression - Mos1 excision - DSB formation. homolog arm.
 MosTIC (Mos1 excision induced Transgene Instructed gene Conversion) Valérie Robert (vrobert@biologie.ens.fr) and Jean-Louis Bessereau (jlbesse@biologie.ens.fr) (November 2006) Introduction: MosTIC (Robert
MosTIC (Mos1 excision induced Transgene Instructed gene Conversion) Valérie Robert (vrobert@biologie.ens.fr) and Jean-Louis Bessereau (jlbesse@biologie.ens.fr) (November 2006) Introduction: MosTIC (Robert
SUPPLEMENTARY INFORMATION
 Figure S1. lev-9 mutants are resistant to levamisole. The levamisole dose-response curves indicate that lev-9 mutants are partially resistant to levamisole similar to lev-10(kr26) mutants. unc-29(x29)
Figure S1. lev-9 mutants are resistant to levamisole. The levamisole dose-response curves indicate that lev-9 mutants are partially resistant to levamisole similar to lev-10(kr26) mutants. unc-29(x29)
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10810 Supplementary Fig. 1: Mutation of the loqs gene leads to shortened lifespan and adult-onset brain degeneration. a. Northern blot of control and loqs f00791 mutant flies. loqs f00791
doi:10.1038/nature10810 Supplementary Fig. 1: Mutation of the loqs gene leads to shortened lifespan and adult-onset brain degeneration. a. Northern blot of control and loqs f00791 mutant flies. loqs f00791
Biol 432L Midterm Oct 6, 2008 Name: 1. Midterm 1, Answer Key Oct. 26, 2009
 Biol 432L Midterm Oct 6, 2008 Name: 1 Midterm 1, Answer Key Oct. 26, 2009 Honor Pledge: I have neither given nor received any unauthorized help on this exam: Name Printed: Signature: 1a. (2 pts) Imagine
Biol 432L Midterm Oct 6, 2008 Name: 1 Midterm 1, Answer Key Oct. 26, 2009 Honor Pledge: I have neither given nor received any unauthorized help on this exam: Name Printed: Signature: 1a. (2 pts) Imagine
Sol narae (Sona) is a Drosophila ADAMTS involved in Wg signaling
 Sol narae (Sona) is a Drosophila ADAMTS involved in Wg signaling Go-Woon Kim, Jong-Hoon Won, Ok-Kyung Lee, Sang-Soo Lee, Jeong-Hoon Han, Orkhon Tsogtbaatar, Sujin Nam, Yeon Kim, and Kyung-Ok Cho* Supplementary
Sol narae (Sona) is a Drosophila ADAMTS involved in Wg signaling Go-Woon Kim, Jong-Hoon Won, Ok-Kyung Lee, Sang-Soo Lee, Jeong-Hoon Han, Orkhon Tsogtbaatar, Sujin Nam, Yeon Kim, and Kyung-Ok Cho* Supplementary
An antibiotic selection marker for nematode transgenesis
 nature methods An antibiotic selection marker for nematode transgenesis Rosina Giordano-Santini, Stuart Milstein, Nenad Svrzikapa, Domena Tu, Robert Johnsen, David Baillie, Marc Vidal & Denis Dupuy Supplementary
nature methods An antibiotic selection marker for nematode transgenesis Rosina Giordano-Santini, Stuart Milstein, Nenad Svrzikapa, Domena Tu, Robert Johnsen, David Baillie, Marc Vidal & Denis Dupuy Supplementary
Supplemental Data. Borg et al. Plant Cell (2014) /tpc
 Supplementary Figure 1 - Alignment of selected angiosperm DAZ1 and DAZ2 homologs Multiple sequence alignment of selected DAZ1 and DAZ2 homologs. A consensus sequence built using default parameters is shown
Supplementary Figure 1 - Alignment of selected angiosperm DAZ1 and DAZ2 homologs Multiple sequence alignment of selected DAZ1 and DAZ2 homologs. A consensus sequence built using default parameters is shown
Repression of Germline RNAi Pathways in Somatic Cells by Retinoblastoma Pathway Chromatin Complexes
 Repression of Germline RNAi Pathways in Somatic Cells by Retinoblastoma Pathway Chromatin Complexes The Harvard community has made this article openly available. Please share how this access benefits you.
Repression of Germline RNAi Pathways in Somatic Cells by Retinoblastoma Pathway Chromatin Complexes The Harvard community has made this article openly available. Please share how this access benefits you.
Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product.
 Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Xu et al., Supplementary Figures 1-7
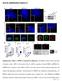 Xu et al., Supplementary Figures 1-7 Supplementary Figure 1. PIPKI is required for ciliogenesis. (a) PIPKI localizes at the basal body of primary cilium. RPE-1 cells treated with two sirnas targeting to
Xu et al., Supplementary Figures 1-7 Supplementary Figure 1. PIPKI is required for ciliogenesis. (a) PIPKI localizes at the basal body of primary cilium. RPE-1 cells treated with two sirnas targeting to
PIWI Associated sirnas and pirnas Specifically Require the Caenorhabditis elegans HEN1 Ortholog henn-1
 PIWI Associated sirnas and pirnas Specifically Require the Caenorhabditis elegans HEN1 Ortholog henn-1 Taiowa A. Montgomery, Young-Soo Rim, Chi Zhang, Robert H. Dowen, Carolyn M. Phillips, Sylvia E. J.
PIWI Associated sirnas and pirnas Specifically Require the Caenorhabditis elegans HEN1 Ortholog henn-1 Taiowa A. Montgomery, Young-Soo Rim, Chi Zhang, Robert H. Dowen, Carolyn M. Phillips, Sylvia E. J.
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature09937 a Name Position Primersets 1a 1b 2 3 4 b2 Phenotype Genotype b Primerset 1a D T C R I E 10000 8000 6000 5000 4000 3000 2500 2000 1500 1000 800 Donor (D)
SUPPLEMENTARY INFORMATION doi:10.1038/nature09937 a Name Position Primersets 1a 1b 2 3 4 b2 Phenotype Genotype b Primerset 1a D T C R I E 10000 8000 6000 5000 4000 3000 2500 2000 1500 1000 800 Donor (D)
The Silence of the Genes
 The Silence of the Genes Initial Observation: Plant geneticists aim to turn pink petunia to purple by over-expression of chalcone synthetase wt wt + Chalcone synthetase cdna driven by cauliflower mosaic
The Silence of the Genes Initial Observation: Plant geneticists aim to turn pink petunia to purple by over-expression of chalcone synthetase wt wt + Chalcone synthetase cdna driven by cauliflower mosaic
Nature Biotechnology: doi: /nbt.4166
 Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
RNA Helicase 1 interacts with an ABC RNAi Transporter: Genetic Interactions with haf-6
 RNA Helicase 1 interacts with an ABC RNAi Transporter: Genetic Interactions with haf-6 By Laticia Rivera Bachelor of Science in Biochemistry and Molecular Biology Oklahoma State University Stillwater,
RNA Helicase 1 interacts with an ABC RNAi Transporter: Genetic Interactions with haf-6 By Laticia Rivera Bachelor of Science in Biochemistry and Molecular Biology Oklahoma State University Stillwater,
No, because expression of the P elements and hence transposase in suppressed in the F1.
 Problem set B 1. A wild-type ry+ (rosy) gene was introduced into a ry mutant using P element-mediated gene transformation, and a strain containing a stable ry+ gene was established. If a transformed male
Problem set B 1. A wild-type ry+ (rosy) gene was introduced into a ry mutant using P element-mediated gene transformation, and a strain containing a stable ry+ gene was established. If a transformed male
Nature Structural & Molecular Biology: doi: /nsmb.3308
 Supplementary Figure 1 Analysis of CED-3 autoactivation and CED-4-induced CED-3 activation. (a) Repeat experiments of Fig. 1a. (b) Repeat experiments of Fig. 1b. (c) Quantitative analysis of three independent
Supplementary Figure 1 Analysis of CED-3 autoactivation and CED-4-induced CED-3 activation. (a) Repeat experiments of Fig. 1a. (b) Repeat experiments of Fig. 1b. (c) Quantitative analysis of three independent
Supplemental Figure legends Figure S1. (A) (B) (C) (D) Figure S2. Figure S3. (A-E) Figure S4. Figure S5. (A, C, E, G, I) (B, D, F, H, Figure S6.
 Supplemental Figure legends Figure S1. Map-based cloning and complementation testing for ZOP1. (A) ZOP1 was mapped to a ~273-kb interval on Chromosome 1. In the interval, a single-nucleotide G to A substitution
Supplemental Figure legends Figure S1. Map-based cloning and complementation testing for ZOP1. (A) ZOP1 was mapped to a ~273-kb interval on Chromosome 1. In the interval, a single-nucleotide G to A substitution
University of Groningen
 University of Groningen Mapping determinants of gene expression plasticity by genetical genomics in C-elegans Li, Yang; Gutteling, E.W.; Tijsterman, M.; Fu, Jingyuan; Riksen, J.A.G.; Prins, P.; Plasterk,
University of Groningen Mapping determinants of gene expression plasticity by genetical genomics in C-elegans Li, Yang; Gutteling, E.W.; Tijsterman, M.; Fu, Jingyuan; Riksen, J.A.G.; Prins, P.; Plasterk,
Supplementary Methods
 Supplementary Methods MARCM-based forward genetic screen We used ethylmethane sulfonate (EMS) to mutagenize flies carrying FRT 2A and FRT 82B transgenes, which are sites for the FLP-mediated recombination
Supplementary Methods MARCM-based forward genetic screen We used ethylmethane sulfonate (EMS) to mutagenize flies carrying FRT 2A and FRT 82B transgenes, which are sites for the FLP-mediated recombination
7.22 ANSWERS to Problem set #1
 7.22 ANSWERS to Problem set #1 1. List whether the pairs marked by the arrows are orthologous or paralogous. α tubulin (mouse) α tubulin (fly) α tubulin (worm) α tubulin (yeast) paralogous Tubulin β tubulin
7.22 ANSWERS to Problem set #1 1. List whether the pairs marked by the arrows are orthologous or paralogous. α tubulin (mouse) α tubulin (fly) α tubulin (worm) α tubulin (yeast) paralogous Tubulin β tubulin
To investigate the heredity of the WFP gene, we selected plants that were homozygous
 Supplementary information Supplementary Note ST-12 WFP allele is semi-dominant To investigate the heredity of the WFP gene, we selected plants that were homozygous for chromosome 1 of Nipponbare and heterozygous
Supplementary information Supplementary Note ST-12 WFP allele is semi-dominant To investigate the heredity of the WFP gene, we selected plants that were homozygous for chromosome 1 of Nipponbare and heterozygous
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1119481/dc1 Supporting Online Material for LIN-12/Notch Activation Leads to MicroRNA-Mediated Down-Regulation of Vav in C. elegans Andrew S. Yoo and Iva Greenwald* *To
www.sciencemag.org/cgi/content/full/1119481/dc1 Supporting Online Material for LIN-12/Notch Activation Leads to MicroRNA-Mediated Down-Regulation of Vav in C. elegans Andrew S. Yoo and Iva Greenwald* *To
Growth factor, augmenter of liver regeneration
 Supplemental Table 1: Human and mouse PC1 sequence equivalencies Human Mouse Domain Clinical significance; Score* PolyPhen prediction; PSIC score difference C210G C210G WSC Highly likely pathogenic; 15
Supplemental Table 1: Human and mouse PC1 sequence equivalencies Human Mouse Domain Clinical significance; Score* PolyPhen prediction; PSIC score difference C210G C210G WSC Highly likely pathogenic; 15
Supplementary Figure 1. Phenotype and genotype of cultured and transplanted S1 KCST (A) Brightfield and mcherry fluorescence images of the spheres
 Supplementary Figure 1. Phenotype and genotype of cultured and transplanted S1 KCST (A) Brightfield and mcherry fluorescence images of the spheres generated from the CD133-positive cells infected with
Supplementary Figure 1. Phenotype and genotype of cultured and transplanted S1 KCST (A) Brightfield and mcherry fluorescence images of the spheres generated from the CD133-positive cells infected with
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/404/ra120/dc1 Supplementary Materials for The subcellular localization and activity of cortactin is regulated by acetylation and interaction with Keap1 Akihiro
www.sciencesignaling.org/cgi/content/full/8/404/ra120/dc1 Supplementary Materials for The subcellular localization and activity of cortactin is regulated by acetylation and interaction with Keap1 Akihiro
Biology 163 Laboratory in Genetics, Final Exam, Dec. 10, 2005
 1 Biology 163 Laboratory in Genetics, Final Exam, Dec. 10, 2005 Honor Pledge: I have neither given nor received any unauthorized help on this exam: Name Printed: Signature: 1. (2 pts) If you see the following
1 Biology 163 Laboratory in Genetics, Final Exam, Dec. 10, 2005 Honor Pledge: I have neither given nor received any unauthorized help on this exam: Name Printed: Signature: 1. (2 pts) If you see the following
Cambridge University Press
 Figure 1.1. Model of RNAi pathway in C. elegans. Transmembrane protein SID-1 allows dsrna to enter the cell. In the cytoplasm,dsrna gets processed by DCR-1,existing in a complex with RDE-4,RDE-1 and DRH-1.
Figure 1.1. Model of RNAi pathway in C. elegans. Transmembrane protein SID-1 allows dsrna to enter the cell. In the cytoplasm,dsrna gets processed by DCR-1,existing in a complex with RDE-4,RDE-1 and DRH-1.
DEPS-1 promotes P-granule assembly and RNA interference in C. elegans germ cells
 Development epress online publication date 30 January 2008 983 Development 135, 983-993 (2008) doi:10.1242/dev.015552 DEPS-1 promotes P-granule assembly and RNA interference in C. elegans germ cells Caroline
Development epress online publication date 30 January 2008 983 Development 135, 983-993 (2008) doi:10.1242/dev.015552 DEPS-1 promotes P-granule assembly and RNA interference in C. elegans germ cells Caroline
Mannen et al., http :// /cgi /content /full /jcb /DC1
 Supplemental material JCB Mannen et al., http ://www.jcb.org /cgi /content /full /jcb.201601024 /DC1 THE JOURNAL OF CELL BIOLOGY Figure S1. Characterization of SNB components. (A) SNB localization of Venus-tagged
Supplemental material JCB Mannen et al., http ://www.jcb.org /cgi /content /full /jcb.201601024 /DC1 THE JOURNAL OF CELL BIOLOGY Figure S1. Characterization of SNB components. (A) SNB localization of Venus-tagged
Heterochromatin Silencing
 Heterochromatin Silencing Heterochromatin silencing Most DNA in eukaryotes consists of repetitive DNA, including retrotransposons, transposable elements. Packaged into a condensed form: Heterochromatin:
Heterochromatin Silencing Heterochromatin silencing Most DNA in eukaryotes consists of repetitive DNA, including retrotransposons, transposable elements. Packaged into a condensed form: Heterochromatin:
University of Groningen
 University of Groningen Differential impact of the HEN1 homolog HENN-1 on 21U and 26G RNAs in the germline of Caenorhabditis elegans Kamminga, Leonie M; van Wolfswinkel, Josien C; Luteijn, Maartje J; Kaaij,
University of Groningen Differential impact of the HEN1 homolog HENN-1 on 21U and 26G RNAs in the germline of Caenorhabditis elegans Kamminga, Leonie M; van Wolfswinkel, Josien C; Luteijn, Maartje J; Kaaij,
(a) Scheme depicting the strategy used to generate the ko and conditional alleles. (b) RT-PCR for
 Supplementary Figure 1 Generation of Diaph3 ko mice. (a) Scheme depicting the strategy used to generate the ko and conditional alleles. (b) RT-PCR for different regions of Diaph3 mrna from WT, heterozygote
Supplementary Figure 1 Generation of Diaph3 ko mice. (a) Scheme depicting the strategy used to generate the ko and conditional alleles. (b) RT-PCR for different regions of Diaph3 mrna from WT, heterozygote
SAS6-like protein in Plasmodium indicates that conoid-associated apical complex proteins persist in invasive stages within the mosquito vector
 SAS6-like protein in Plasmodium indicates that conoid-associated apical complex proteins persist in invasive stages within the mosquito vector Richard J. Wall 1,#, Magali Roques 1,+, Nicholas J. Katris
SAS6-like protein in Plasmodium indicates that conoid-associated apical complex proteins persist in invasive stages within the mosquito vector Richard J. Wall 1,#, Magali Roques 1,+, Nicholas J. Katris
Supplemental Data. Cui et al. (2012). Plant Cell /tpc a b c d. Stem UBC32 ACTIN
 A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
a) Stock 4: 252 flies total: 120 are P[w+]: 64 are CyO; 56 are TM3; all are females. 132 are w-: 68 are CyO; 64 are TM3; all are males.
![a) Stock 4: 252 flies total: 120 are P[w+]: 64 are CyO; 56 are TM3; all are females. 132 are w-: 68 are CyO; 64 are TM3; all are males. a) Stock 4: 252 flies total: 120 are P[w+]: 64 are CyO; 56 are TM3; all are females. 132 are w-: 68 are CyO; 64 are TM3; all are males.](/thumbs/91/106088469.jpg) Drosophila Problem Set 2018 1) You have created P element transformants of a construct that contains the mini-white gene, which confers an orange eye color in a homozygous white mutant background. For
Drosophila Problem Set 2018 1) You have created P element transformants of a construct that contains the mini-white gene, which confers an orange eye color in a homozygous white mutant background. For
RNAi mechanisms in Caenorhabditis elegans
 FEBS Letters 579 (2005) 5932 5939 FEBS 29870 Minireview RNAi mechanisms in Caenorhabditis elegans Alla Grishok * Center for Cancer Research, MIT, E17-526, 40 Ames Street, Cambridge, MA 02139, USA Received
FEBS Letters 579 (2005) 5932 5939 FEBS 29870 Minireview RNAi mechanisms in Caenorhabditis elegans Alla Grishok * Center for Cancer Research, MIT, E17-526, 40 Ames Street, Cambridge, MA 02139, USA Received
Watson BM Gene Capitolo 11
 Watson BM Gene Capitolo 11 Le proteine della trasposizione ? target-site primed reverse transcription Watson et al., BIOLOGIA MOLECOLARE DEL GENE, Zanichelli editore S.p.A. Trimeric structure for an
Watson BM Gene Capitolo 11 Le proteine della trasposizione ? target-site primed reverse transcription Watson et al., BIOLOGIA MOLECOLARE DEL GENE, Zanichelli editore S.p.A. Trimeric structure for an
At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in
 Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
Supplementary Data 1.
 Supplementary Data 1. Evaluation of the effects of number of F2 progeny to be bulked (n) and average sequencing coverage (depth) of the genome (G) on the levels of false positive SNPs (SNP index = 1).
Supplementary Data 1. Evaluation of the effects of number of F2 progeny to be bulked (n) and average sequencing coverage (depth) of the genome (G) on the levels of false positive SNPs (SNP index = 1).
1a. What is the ratio of feathered to unfeathered shanks in the offspring of the above cross?
 1. Whether or not the shanks of chickens contains feathers is due to two independently assorting genes. Individuals have unfeathered shanks when they are homozygous for recessive genes at two loci; the
1. Whether or not the shanks of chickens contains feathers is due to two independently assorting genes. Individuals have unfeathered shanks when they are homozygous for recessive genes at two loci; the
Learning Objectives. Define RNA interference. Define basic terminology. Describe molecular mechanism. Define VSP and relevance
 Learning Objectives Define RNA interference Define basic terminology Describe molecular mechanism Define VSP and relevance Describe role of RNAi in antigenic variation A Nobel Way to Regulate Gene Expression
Learning Objectives Define RNA interference Define basic terminology Describe molecular mechanism Define VSP and relevance Describe role of RNAi in antigenic variation A Nobel Way to Regulate Gene Expression
SUPPLEMENTARY INFORMATION
 Materials and Methods Transgenic Plant Materials and DNA Constructs. VEX1::H2B-GFP, ACA3::H2B-GFP, KRP6::H2B-GFP, KRP6::mock21ts-GFP, KRP6::TE21ts- GFP and KRP6::miR161ts-GFP constructs were generated
Materials and Methods Transgenic Plant Materials and DNA Constructs. VEX1::H2B-GFP, ACA3::H2B-GFP, KRP6::H2B-GFP, KRP6::mock21ts-GFP, KRP6::TE21ts- GFP and KRP6::miR161ts-GFP constructs were generated
Department of Physiology, Tokyo Women's Medical University School of Medicine, Tokyo, Japan 2
 Supplementary Information Endomembrane-associated RSD-3 is important for RNAi induced by extracellular silencing RNA in both somatic and germ cells of Caenorhabditis elegans! Rieko Imae 1,4, Katsufumi
Supplementary Information Endomembrane-associated RSD-3 is important for RNAi induced by extracellular silencing RNA in both somatic and germ cells of Caenorhabditis elegans! Rieko Imae 1,4, Katsufumi
Bi Lecture 3 Loss-of-function (Ch. 4A) Monday, April 8, 13
 Bi190-2013 Lecture 3 Loss-of-function (Ch. 4A) Infer Gene activity from type of allele Loss-of-Function alleles are Gold Standard If organism deficient in gene A fails to accomplish process B, then gene
Bi190-2013 Lecture 3 Loss-of-function (Ch. 4A) Infer Gene activity from type of allele Loss-of-Function alleles are Gold Standard If organism deficient in gene A fails to accomplish process B, then gene
Two classes of silencing RNAs move between Caenorhabditis elegans tissues
 A r t i c l e s Two classes of silencing RNAs move between Caenorhabditis elegans tissues Antony M Jose 1,2, Giancarlo A Garcia 1 & Craig P Hunter 1 2011 Nature America, Inc. All rights reserved. Organism-wide
A r t i c l e s Two classes of silencing RNAs move between Caenorhabditis elegans tissues Antony M Jose 1,2, Giancarlo A Garcia 1 & Craig P Hunter 1 2011 Nature America, Inc. All rights reserved. Organism-wide
The RRPA knock-in allele was generated by homologous recombination in TC1 ES cells.
 Supplemental Materials Materials & Methods Generation of RRPA and RAPA Knock-in Mice The RRPA knock-in allele was generated by homologous recombination in TC1 ES cells. Targeted ES clones in which the
Supplemental Materials Materials & Methods Generation of RRPA and RAPA Knock-in Mice The RRPA knock-in allele was generated by homologous recombination in TC1 ES cells. Targeted ES clones in which the
Supplementary Data Supplementary Figures
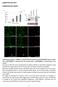 Supplementary Data Supplementary Figures Supplementary Figure 1. Pi04314 is expressed during infection, each GFP-Pi04314 fusion is stable and myr GFP-Pi04314 is removed from the nucleus while NLS GFP-Pi04314
Supplementary Data Supplementary Figures Supplementary Figure 1. Pi04314 is expressed during infection, each GFP-Pi04314 fusion is stable and myr GFP-Pi04314 is removed from the nucleus while NLS GFP-Pi04314
Characterization of a novel gene involved in border cell migration
 Characterization of a novel gene involved in border cell migration Wei-Hao Li, He-Yen Chou and Li-Mei Pai Graduate Institute of Basic Medical Science Chang-Gung University, Tao-Yuan, Taiwan, R.O.C. Abstract
Characterization of a novel gene involved in border cell migration Wei-Hao Li, He-Yen Chou and Li-Mei Pai Graduate Institute of Basic Medical Science Chang-Gung University, Tao-Yuan, Taiwan, R.O.C. Abstract
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Figure S2. Verification of ihog and boi
 Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
Caenorhabditis elegans: The Heavyweight Champ of Gene Knockout Technology
 Caenorhabditis elegans: The Heavyweight Champ of Gene Knockout Technology Rachel Pan The nematode (worm) Caenorhabditis elegans was the first multicellular organism to have its genome, or complete DNA
Caenorhabditis elegans: The Heavyweight Champ of Gene Knockout Technology Rachel Pan The nematode (worm) Caenorhabditis elegans was the first multicellular organism to have its genome, or complete DNA
gypsy fold change in steady state RNA levels RnpS1 Acn mago tsu GFP armi sirna mediated depletion against indicated genes in OSCs
 Hayashi_FigS1 fold change in steady state RNA levels 1000 100 10 1 gypsy GFP armi mago tsu Acn RnpS1 btz sirna mediated depletion against indicated genes in OSCs Supplementary Figure S1. The EJC is required
Hayashi_FigS1 fold change in steady state RNA levels 1000 100 10 1 gypsy GFP armi mago tsu Acn RnpS1 btz sirna mediated depletion against indicated genes in OSCs Supplementary Figure S1. The EJC is required
Supplementary Information
 Supplementary Information MED18 interaction with distinct transcription factors regulates plant immunity, flowering time and responses to hormones Supplementary Figure 1. Diagram showing T-DNA insertion
Supplementary Information MED18 interaction with distinct transcription factors regulates plant immunity, flowering time and responses to hormones Supplementary Figure 1. Diagram showing T-DNA insertion
SUPPORTING ONLINE MATERIAL
 SUPPORTING ONLINE MATERIAL MATERIAL AND METHODS Fly stocks All Drosophila stocks were grown at 25 C on yeast-containing corn meal/molasses medium. The nanos (nos) alleles nos 18 and nos 53 have been previously
SUPPORTING ONLINE MATERIAL MATERIAL AND METHODS Fly stocks All Drosophila stocks were grown at 25 C on yeast-containing corn meal/molasses medium. The nanos (nos) alleles nos 18 and nos 53 have been previously
Two Classes of Silencing RNAs Move between Caenorhabditis elegans Tissues
 Two Classes of Silencing RNAs Move between Caenorhabditis elegans Tissues The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation
Two Classes of Silencing RNAs Move between Caenorhabditis elegans Tissues The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation
Experimental Tools and Resources Available in Arabidopsis. Manish Raizada, University of Guelph, Canada
 Experimental Tools and Resources Available in Arabidopsis Manish Raizada, University of Guelph, Canada Community website: The Arabidopsis Information Resource (TAIR) at http://www.arabidopsis.org Can order
Experimental Tools and Resources Available in Arabidopsis Manish Raizada, University of Guelph, Canada Community website: The Arabidopsis Information Resource (TAIR) at http://www.arabidopsis.org Can order
Supplemental Information
 Supplemental Information Itemized List Materials and Methods, Related to Supplemental Figures S5A-C and S6. Supplemental Figure S1, Related to Figures 1 and 2. Supplemental Figure S2, Related to Figure
Supplemental Information Itemized List Materials and Methods, Related to Supplemental Figures S5A-C and S6. Supplemental Figure S1, Related to Figures 1 and 2. Supplemental Figure S2, Related to Figure
Genome research in eukaryotes
 Functional Genomics Genome and EST sequencing can tell us how many POTENTIAL genes are present in the genome Proteomics can tell us about proteins and their interactions The goal of functional genomics
Functional Genomics Genome and EST sequencing can tell us how many POTENTIAL genes are present in the genome Proteomics can tell us about proteins and their interactions The goal of functional genomics
Biology 4361 Developmental Biology Lecture 4. The Genetic Core of Development
 Biology 4361 Developmental Biology Lecture 4. The Genetic Core of Development The only way to get from genotype to phenotype is through developmental processes. - Remember the analogy that the zygote contains
Biology 4361 Developmental Biology Lecture 4. The Genetic Core of Development The only way to get from genotype to phenotype is through developmental processes. - Remember the analogy that the zygote contains
Supplemental Figure Legends:
 Supplemental Figure Legends: Fig S1. GFP-ABRO1 localization. U2OS cells were infected with retrovirus expressing GFP- ABRO1. The cells were fixed with 3.6% formaldehyde and stained with antibodies against
Supplemental Figure Legends: Fig S1. GFP-ABRO1 localization. U2OS cells were infected with retrovirus expressing GFP- ABRO1. The cells were fixed with 3.6% formaldehyde and stained with antibodies against
Automated analysis of embryonic gene expression with cellular resolution in C. elegans
 Automated analysis of embryonic gene expression with cellular resolution in C. elegans John Isaac Murray, Zhirong Bao, Thomas J Boyle, Max E Boeck, Barbara L Mericle, Thomas J Nicholas, Zhongying Zhao,
Automated analysis of embryonic gene expression with cellular resolution in C. elegans John Isaac Murray, Zhirong Bao, Thomas J Boyle, Max E Boeck, Barbara L Mericle, Thomas J Nicholas, Zhongying Zhao,
a) What is heteochromatin? highly condensed, inactive chromatin c) What is RNAi? Briefly explain how it works.
 GENE 603 Exam II, Fall 2016 1A Research has shown that histone acetylation is triggered by neuronal activity in rat brains during fear conditioning and memory formation. a) What enzymes are involved in
GENE 603 Exam II, Fall 2016 1A Research has shown that histone acetylation is triggered by neuronal activity in rat brains during fear conditioning and memory formation. a) What enzymes are involved in
Regulation of transcription by the MLL2 complex and MLL complex-associated AKAP95
 Supplementary Information Regulation of transcription by the complex and MLL complex-associated Hao Jiang, Xiangdong Lu, Miho Shimada, Yali Dou, Zhanyun Tang, and Robert G. Roeder Input HeLa NE IP lot:
Supplementary Information Regulation of transcription by the complex and MLL complex-associated Hao Jiang, Xiangdong Lu, Miho Shimada, Yali Dou, Zhanyun Tang, and Robert G. Roeder Input HeLa NE IP lot:
The Human Protein PRR14 Tethers Heterochromatin to the Nuclear Lamina During Interphase and Mitotic Exit
 Cell Reports, Volume 5 Supplemental Information The Human Protein PRR14 Tethers Heterochromatin to the Nuclear Lamina During Interphase and Mitotic Exit Andrey Poleshko, Katelyn M. Mansfield, Caroline
Cell Reports, Volume 5 Supplemental Information The Human Protein PRR14 Tethers Heterochromatin to the Nuclear Lamina During Interphase and Mitotic Exit Andrey Poleshko, Katelyn M. Mansfield, Caroline
Figure S1a. The modular dcas9 fusion system works efficiently to suppress and activate endogenous gene expression in C. elegans
 Figure S1a. The modular dcas9 fusion system works efficiently to suppress and activate endogenous gene expression in C. elegans A, a normal wild-type hermaphrodite N2 is shown. B, a dpy-5-supressed F1
Figure S1a. The modular dcas9 fusion system works efficiently to suppress and activate endogenous gene expression in C. elegans A, a normal wild-type hermaphrodite N2 is shown. B, a dpy-5-supressed F1
1a. What is the ratio of feathered to unfeathered shanks in the offspring of the above cross?
 Problem Set 5 answers 1. Whether or not the shanks of chickens contains feathers is due to two independently assorting genes. Individuals have unfeathered shanks when they are homozygous for recessive
Problem Set 5 answers 1. Whether or not the shanks of chickens contains feathers is due to two independently assorting genes. Individuals have unfeathered shanks when they are homozygous for recessive
Lecture 8: Transgenic Model Systems and RNAi
 Lecture 8: Transgenic Model Systems and RNAi I. Model systems 1. Caenorhabditis elegans Caenorhabditis elegans is a microscopic (~1 mm) nematode (roundworm) that normally lives in soil. It has become one
Lecture 8: Transgenic Model Systems and RNAi I. Model systems 1. Caenorhabditis elegans Caenorhabditis elegans is a microscopic (~1 mm) nematode (roundworm) that normally lives in soil. It has become one
To assess the localization of Citrine fusion proteins, we performed antibody staining to
 Trinh et al 1 SUPPLEMENTAL MATERIAL FlipTraps recapitulate endogenous protein localization To assess the localization of Citrine fusion proteins, we performed antibody staining to compare the expression
Trinh et al 1 SUPPLEMENTAL MATERIAL FlipTraps recapitulate endogenous protein localization To assess the localization of Citrine fusion proteins, we performed antibody staining to compare the expression
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity.
 Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
University of Dundee. Published in: Scientific Reports. DOI: /srep Publication date: 2016
 University of Dundee SAS6-like protein in Plasmodium indicates that conoid-associated apical complex proteins persist in invasive stages within the mosquito vector Wall, Richard J.; Roques, Magali; Katris,
University of Dundee SAS6-like protein in Plasmodium indicates that conoid-associated apical complex proteins persist in invasive stages within the mosquito vector Wall, Richard J.; Roques, Magali; Katris,
Prof. Fahd M. Nasr. Faculty of Sciences Lebanese University Beirut, Lebanon.
 Prof. Fahd M. Nasr Faculty of Sciences Lebanese University Beirut, Lebanon https://yeastwonderfulworld.wordpress.com/ Biol328 - B3212 Molecular Biotechnology Partial Exam Question I Explain the procedure
Prof. Fahd M. Nasr Faculty of Sciences Lebanese University Beirut, Lebanon https://yeastwonderfulworld.wordpress.com/ Biol328 - B3212 Molecular Biotechnology Partial Exam Question I Explain the procedure
ASPP1 Fw GGTTGGGAATCCACGTGTTG ASPP1 Rv GCCATATCTTGGAGCTCTGAGAG
 Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or
 Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Sperm cells are passive cargo of the pollen tube in plant fertilization
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 3 ARTICLE NUMBER: 17079 Sperm cells are passive cargo of the pollen tube in plant fertilization Jun Zhang 1, Qingpei
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 3 ARTICLE NUMBER: 17079 Sperm cells are passive cargo of the pollen tube in plant fertilization Jun Zhang 1, Qingpei
Lihua Julie Zhu. Integrated Analysis Of ChIP-seq/chip using ChIPpeakAnno and GeneNetworkBuilder
 April 2008 Integrated Analysis Of ChIP-seq/chip using ChIPpeakAnno and GeneNetworkBuilder Lihua Julie Zhu Bioconductor Developer Meeting Zurich, Switzerland December 13th -14th 2012 Outline Introduction
April 2008 Integrated Analysis Of ChIP-seq/chip using ChIPpeakAnno and GeneNetworkBuilder Lihua Julie Zhu Bioconductor Developer Meeting Zurich, Switzerland December 13th -14th 2012 Outline Introduction
A RRM1 H2AX DAPI. RRM1 H2AX DAPI Merge. Cont. sirna RRM1
 A H2AX DAPI H2AX DAPI Merge Cont sirna Figure S1: Accumulation of RRM1 at DNA damage sites (A) HeLa cells were subjected to in situ detergent extraction without IR irradiation, and immunostained with the
A H2AX DAPI H2AX DAPI Merge Cont sirna Figure S1: Accumulation of RRM1 at DNA damage sites (A) HeLa cells were subjected to in situ detergent extraction without IR irradiation, and immunostained with the
Supplementary Figure 1. Chromosome 3 is devoid of the telomere-proximal subtelomeric
 Supplementary Figure 1. Chromosome 3 is devoid of the telomere-proximal subtelomeric common sequences. (a) Schematic illustration of the telomere-proximal site of subtelomeres. The restriction map is based
Supplementary Figure 1. Chromosome 3 is devoid of the telomere-proximal subtelomeric common sequences. (a) Schematic illustration of the telomere-proximal site of subtelomeres. The restriction map is based
Fig. S1. Endocytosis of extracellular cargo does not depend on dia. (A) To visualize endocytosis, fluorescently labelled wheat germ agglutinin
 Fig. S1. Endocytosis of extracellular cargo does not depend on dia. (A) To visualize endocytosis, fluorescently labelled wheat germ agglutinin (WGA-Alexa555) was injected into the extracellular perivitteline
Fig. S1. Endocytosis of extracellular cargo does not depend on dia. (A) To visualize endocytosis, fluorescently labelled wheat germ agglutinin (WGA-Alexa555) was injected into the extracellular perivitteline
Two classes of genetic pathways I) Developmental/synthetic pathways II) Regulatory/binary switch pathways
 Two classes of genetic pathways I) Developmental/synthetic pathways II) Regulatory/binary switch pathways I) Developmental pathways Synthesis and/or assembly of molecule(s), subcellular, cellular or multicellular
Two classes of genetic pathways I) Developmental/synthetic pathways II) Regulatory/binary switch pathways I) Developmental pathways Synthesis and/or assembly of molecule(s), subcellular, cellular or multicellular
SUPPLEMENTARY INFORMATION
 SULEMENTARY INFORMATION doi:10.1038/nature12117 Supplementary Discussion Although loss of leads to initiation of postembryonic development even under starvation conditions, ectopic overexpression of under
SULEMENTARY INFORMATION doi:10.1038/nature12117 Supplementary Discussion Although loss of leads to initiation of postembryonic development even under starvation conditions, ectopic overexpression of under
Enzyme that uses RNA as a template to synthesize a complementary DNA
 Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Biology 163 Laboratory in Genetics Midterm 2, Nov. 14, Honor Pledge: I have neither given nor received any unauthorized help on this exam:
 1 Biology 163 Laboratory in Genetics Midterm 2, Nov. 14, 2005 Honor Pledge: I have neither given nor received any unauthorized help on this exam: Name Printed: ignature: 1. Normally you need to cross two
1 Biology 163 Laboratory in Genetics Midterm 2, Nov. 14, 2005 Honor Pledge: I have neither given nor received any unauthorized help on this exam: Name Printed: ignature: 1. Normally you need to cross two
Lecture 2-3: Using Mutants to study Biological processes
 Lecture 2-3: Using Mutants to study Biological processes Objectives: 1. Why use mutants? 2. How are mutants isolated? 3. What important genetic analyses must be done immediately after a genetic screen
Lecture 2-3: Using Mutants to study Biological processes Objectives: 1. Why use mutants? 2. How are mutants isolated? 3. What important genetic analyses must be done immediately after a genetic screen
 DOI: 10.1038/ncb3259 A Ismail et al. Supplementary Figure 1 B 60000 45000 SSC 30000 15000 Live cells 0 0 15000 30000 45000 60000 FSC- PARR 60000 45000 PARR Width 30000 FSC- 15000 Single cells 0 0 15000
DOI: 10.1038/ncb3259 A Ismail et al. Supplementary Figure 1 B 60000 45000 SSC 30000 15000 Live cells 0 0 15000 30000 45000 60000 FSC- PARR 60000 45000 PARR Width 30000 FSC- 15000 Single cells 0 0 15000
An H4K16 histone acetyltransferase mediates decondensation of the X chromosome in C. elegans males
 DOI 10.1186/s13072-016-0097-x Epigenetics & Chromatin RESEARCH Open Access An H4K16 histone acetyltransferase mediates decondensation of the X chromosome in C. elegans males Alyssa C. Lau 1,2, Kevin P.
DOI 10.1186/s13072-016-0097-x Epigenetics & Chromatin RESEARCH Open Access An H4K16 histone acetyltransferase mediates decondensation of the X chromosome in C. elegans males Alyssa C. Lau 1,2, Kevin P.
Biotechnology Explorer
 Biotechnology Explorer C. elegans Behavior Kit Bioinformatics Supplement explorer.bio-rad.com Catalog #166-5120EDU This kit contains temperature-sensitive reagents. Open immediately and see individual
Biotechnology Explorer C. elegans Behavior Kit Bioinformatics Supplement explorer.bio-rad.com Catalog #166-5120EDU This kit contains temperature-sensitive reagents. Open immediately and see individual
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2880 Supplementary Figure 1 Sequence alignment of Deup1 and Cep63. The protein sequence alignment was generated by the Clustal X 2.0 multiple sequence alignment program using default parameters.
DOI: 10.1038/ncb2880 Supplementary Figure 1 Sequence alignment of Deup1 and Cep63. The protein sequence alignment was generated by the Clustal X 2.0 multiple sequence alignment program using default parameters.
Fig. S1. Negative controls for in situ hybridization results using bam antisense probe. Fig. S2. Overexpression of either mir-275 or mir306
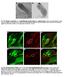 Fig. S1. Negative controls for in situ hybridization results using bam antisense probe. (A) In situ hybridization using bam sense probe in wild-type testis. (B) In situ hybridization using bam antisense
Fig. S1. Negative controls for in situ hybridization results using bam antisense probe. (A) In situ hybridization using bam sense probe in wild-type testis. (B) In situ hybridization using bam antisense
Biological Sciences 50 Practice Final Exam. Allocate your time wisely.
 NAME: Fall 2005 TF: Biological Sciences 50 Practice Final Exam A. Be sure to write your name on the top of each of page of the examination. B. Write each answer only on the same page as the pertinent question.
NAME: Fall 2005 TF: Biological Sciences 50 Practice Final Exam A. Be sure to write your name on the top of each of page of the examination. B. Write each answer only on the same page as the pertinent question.
