Supplemental Information. Macrophages Mediate the Repair of Brain. Vascular Rupture through Direct Physical Adhesion. and Mechanical Traction
|
|
|
- Mervyn Golden
- 6 years ago
- Views:
Transcription
1 Immunity, Volume 44 Supplemental Information Macrophages Mediate the Repair of Brain Vascular Rupture through Direct Physical Adhesion and Mechanical Traction Chi Liu, Chuan Wu, Qifen Yang, Jing Gao, Li Li, Deqin Yang, and Lingfei Luo
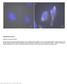
2 Figure S1
3 Figure S1, related to Figure 2. Repair of cerebrovascular ruptures can be mediated by peripheral macrophages. (A and B) Combination of fluorescent in situ hybridizations (FISH) and immunohistochemistry indicates that the majority of cerebrovascular ruptures are repaired by Lcp1 and p2y12 double positive macrophages (A, n=56/58). However, in rare cases, ruptures can be repaired by p2y12-negative macrophages (B, n=2/58). (C and D) The majority of cerebrovascular ruptures are repaired by Lcp1 and slc7a7 double positive macrophages (C, n=44/45). However, in rare cases, ruptures can be repaired by slc7a7-negative macrophages (D, n=1/45). (E H) In contrast to the un-injected control (E and G), brain-resident microglia can be efficiently depleted by the injection of slc7a7mo as indicated by the mpeg1:gfp transgene (F, n=33/40) and FISH-p2y12 (H, n=28/36). (I and J) In slc7a7 morphants, a small portion of cerebrovascular ruptures can be repaired by macrophages (I, n=17/81). All these repairing macrophages are negative for p2y12 (J, n=17/17). (I) and (J) exhibit the same macrophage. The time elapsed since the laser ablation is indicated in hours:minutes. Scale bar, 10 μm.
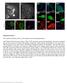
4 Figure S2 Figure S2, related to Figure 3. Macrophages in the brain express Cdh5 and other adhesion molecules. (A and B) Under the Tg(cdh5:gal4; UAS:GFP) transgenic background, strong and weak GFP epifluorescences were detected in brain vascular ECs and macrophages, respectively (A, n=25/25). Expression of GFP in the macrophages was confirmed by antibodies against Lcp1 and GFP (B, n=20/20). Arrowheads indicate macrophages. Scale bar, 10 μm. (C) RT-PCRs indicated that the sorted brain macrophages expressed cdh5, pecam1, icam1, and vcam1, but not kdrl or zo1.
5 Figure S3 Figure S3, related to Figure 5. Sorting of the working macrophages and endothelial end cells. (A and B) The Kaede-labeled working macrophage was photoconverted from green to red epifluorescence (A). About eighty macrophages were photoconverted and proceeded to sorting, and about twenty of them were finally confirmed and collected (B, R6) for further transcriptional profile analysis. (C and D) The Dendra2-labeled endothelial end cells were photoconverted from green to
6 red epifluorescence (C). About eighty cells were photoconverted and proceeded to sorting, and about twenty of them were finally confirmed and collected (D, R6) for further transcriptional profile analysis. The purple circles indicate irradiation points by the 405-nm laser for photoconversion.
7 Figure S4
8 Figure S4, related to Figure 5. Statistics of pathway enrichment in the working macrophages and endothelial end cells. Transcriptional profile analyses of the working macrophages (A) and of the endothelial end cells (B) at three different lesion stages indicate statistics of pathway enrichment. Rich-ratio represents the ratio of significantly different expression genes in a pathway / the ratio of significantly different expression genes in all pathways. The size of the color dot indicates the number of significantly different expression genes in a pathway. The darkness of the color dot indicates the p-val.
9 Figure S5 Figure S5, related to Figure 5. KEGG pathway map of the cell adhesion and mechanical force regulatory genes. Schematic based on KEGG pathway map displays comparison of cell adhesion and
10 mechanical force regulatory genes between three different lesion stages, macrophage arrival vs uninjured controls (A), macrophage traction vs macrophage arrival (B), and macrophage traction vs uninjured controls (C), in both macrophages and endothelial end cells. Red and green boxes represent up-regulated and down-regulated genes, respectively.
11 Figure S6 Figure S6, related to Figure 1. Macrophages mediate the repair of fin vascular ruptures. Dynamic cellular events that occur in the repair of fin vascular rupture. Note that the macrophage also extends filopodia or lamellipodia, physically adheres to both endothelial ends and mediates their ligation (n=10/20). Note that the neutrophils that are fluorescently labeled in yellow arrive at and leave the lesion earlier than macrophages, but never physically adhere to the endothelial ends (n=20/20). Scale bar, 10 μm.
12 Supplemental Experimental Procedures Zebrafish Strains and Generation of Transgenic Lines Zebrafish of the Tg(kdrl:Dendra2-NTR) cq10 abbreviated as Tg(kdrl:DenNTR) cq10, Tg(kdrl:Lifeact-DsRed) cq11, Tg(coronin1a:Lifeact-DsRed) cq12 abbreviated as Tg(coro1a:Lifeact-DsRed) cq12, Tg(kdrl:eb3-GFP) cq13, Tg(kdrl:Rac1-FRET) cq17 (Chen et al., 2012), Tg(coronin1a:Rac1-FRET) cq18 abbreviated as Tg(coro1a:Rac1-FRET) cq18 (Li et al., 2012b), Tg(coro1a:GFP; lysc:dsred) (Li et al., 2012a), Tg(kdrl:GFP; gata1:dsred), Tg(kdrl:DsRed), Tg(kdrl:GcaMP5) cq19, Tg(coronin1a:hCRIB-RasCT) cq20 abbreviated as Tg(coro1a:hCRIB-RasCT) cq20, Tg(kdrl:hCRIB-RasCT) cq21, Tg(coronin1a:Kaede) cq22 abbreviated as Tg(coro1a:Kaede) cq22, Tg(kdrl:pecam1-GFP) cq23, Tg(cdh5 BAC :gal4ff) mu101 abbreviated as Tg(cdh5:gal4) (Bussmann et al., 2011), Tg(UAS:GFP) nkuasgfp1a (Bussmann et al., 2011), and Tg(mpeg1:GFP) (Ellett et al., 2011) lines were raised and maintained under standard laboratory conditions according to Institutional Animal Care and Use Committee protocols. kdrl-denntr construct was generated by replacing the lfabp promoter with the kdrl promoter in the pbluescript vector. Lifeact-DsRed was generated by the direct addition of the Lifeact DNA sequence before DsRed with the following primers: 5'-TCCCCCGGGATGGGTGTCGCAGATTTGATCAAGAAATTCGAAAGCATCTCAAAG GAAGAAATGGCCTCCTCCGAGAACGT-3' and 5'-CACCTGTTCCTGTAATCGATA-3'.
13 kdrl-lifeact-dsred and coro1a-lifeact-dsred were generate by inserting Lifeact-DsRed downstream of the kdrl promoter or the coronin1a promoter. eb3 was amplified from human cdna using the primers 5'-GTCGACATGGCCGTCAATGTGTACTC-3' and 5'-GGATCCCGTACTCGTCCTGGTCTTCTTG-3' and then cloned into the pegfp-n2 plasmid. The eb3-gfp sequence was then excised and subcloned downstream of the kdrl promoter to generate the kdrl-eb3-gfp construct. pecam1 was amplified from zebrafish cdna, and the same subcloning procedure that was used for eb3 was applied to generate the kdrl-pecam1-gfp construct. kdrl-gcamp5 and kdrl-hcrib-rasct were constructed by subcloning GcaMP5 and hcrib-rasct, respectively, downstream of the kdrl promoter. The constructs were injected into zebrafish embryos of the AB genetic background at the one-cell stage for transgenesis. Laser Microsurgical Cell Ablation, Photoconversion, In Vivo Time-lapse and FRET Imaging For laser microsurgical cell ablation, the larvae were mounted in 1.2% low melting point agarose and viewed using a LSM780NLO confocal microscope (Carl Zeiss). The target cerebrovascular endothelial cell or macrophage was focused in a single confocal plane and irradiated for three seconds by a multi-photon laser at 800 nm, which was generated using a Chameleon Vision2 Laser System (Coherent). Effective ablations were identified by cellular debris. Whole larvae photoconversion was performed by exposing the larvae to UV light for
14 4 minutes using a M165FC Stereo Microscope (Leica). Most of the Dendra2 epifluorescence was converted from green to red. Focused photoconversion of the endothelial ends was achieved by irradiation with a 405-nm laser equipped on an LSM780NLO confocal microscope for 30 seconds. For time-lapse live imaging, zebrafish embryos were grown in the presence of 0.003% 1-phenyl-2-thiourea (PTU) to avoid pigmentation. Larvae were mounted in 1.2% low melting point agarose in 35-mm glass bottom dishes. Time-lapse images were captured using a 20 water immersion objective mounted on an LSM780NLO confocal microscope that was equipped with a heating stage to maintain 28.5 C. Z-stack images were collected every 3 10 minutes, and three-dimensional data sets were compiled using ZEN2010 software (Carl Zeiss). To measure Rac1 activity, FRET ratio images were acquired using a Zeiss LSM780 confocal microscope, as previously described (Kardash et al., 2010; Kardash et al., 2011; Li et al., 2012b). RNA Sequencing Focused photoconversions of endothelial end cells or macrophages at three lesion stages (uninjured control, macrophage arrival, and macrophage traction) were achieved by irradiating Tg(kdrl:DenNTR; coro1a:egfp) or Tg(coro1a:Kaeda; kdrl:egfp) transgenic larvae with a 405-nm laser equipped on an LSM780NLO confocal microscope for 30 seconds. About eighty cells (repairing macrophages or endothelial end cells) in
15 twenty animals were photoconverted. After photoconversion, zebrafish heads were dissected, collected, and homogenized in 1 ml 0.5% trypsin at 4 C. The homogenized cell suspension was centrifuged at 1000x g at 4 C for 5 minutes. Then, the supernatant was removed and the cell pellet was resuspended in PBS, followed by cell sorting by flow cytometry (Moflo XDP, Beckman). cdna libraries were generated from these sorted cells using the Smart-seq2 protocol (Picelli et al., 2013). Then, RNA sequencing was performed using the PE100 strategy (HiSeq 2500, Illumina). Raw data were filtered to obtain clean data using Perl scripts. Reference genome and annotations were downloaded from the ENSEMBL website ( Reference genome library was built using the Bowtie2 v2.2.3 software, and clean data was aligned to the library using the TopHat v software. Gene expressions were calculated by the RPKM method using the HTSeq v0.6.0 software. Genes with P value<0.05 and log2 ratio 1 were identified as differentially expressed genes using the DESeq (v1.16.0) software. These data were then subject to GO (gene ontology, analyses and aligned to KEGG (Kyoto Encyclopedia of Genes and Genomes, database to build pathway maps. Immunohistochemistry and Fluorescent In Situ Hybridizations Whole-mount antibody staining was performed as previously described (Liu et al., 2009). To detect the expression of Cdh5, the larvae were fixed 2.5 hours and 4 hours after laser
16 ablation. Anti-zfCdh5 antibodies (Blum et al.,2008) (1:400, a gift from Markus Affolter) and anti-lcp1 antibodies(li et al., 2011) (1:400) were visualized using donkey-anti-rabbit AlexaFluor 647 (Invitrogen). Anti-GFP antibodies (1:400, Santa Cruz) were visualized using goat-anti-mouse AlexaFluor 488 (Invitrogen). Combination of fluorescent in situ hybridizations and immunohistochemistry was carried out as previously described (He et al., 2014) using anti-lcp1 antibodies (1:400) and p2y12 and slc7a7 antisense RNA probes. Cell Sorting, RNA Isolation, and RT-PCR The heads of five hundred Tg(mpeg1:GFP) transgenic larvae were dissected at 84 hpf, and their cells were dissociated and sorted as previously described (Lu et al., 2013). Total RNA was isolated from sorted macrophages using TRIzol reagent (Invitrogen), and cdna was obtained via reverse transcription using OmniScript RT Kits (Qiagen). PCR was then performed to detect the transcriptions of cdh5, pecam1, icam1, vcam1, kdrl, and zo1. Primer sequences are available on request.
17 Supplemental References Blum, Y., Belting, H.G., Ellertsdottir, E., Herwig, L., Lüders, F., and Affolter, M. (2008). Complex cell rearrangements during intersegmental vessel sprouting and vessel fusion in the zebrafish embryo. Dev. Biol. 316, Picelli, S., Bjorklund, A.K., Faridani, O.R., Sagasser, S., Winberg, G., and Sandberg, R. (2013). Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10,
Supplementary Information
 Supplementary Information Live imaging reveals the dynamics and regulation of mitochondrial nucleoids during the cell cycle in Fucci2-HeLa cells Taeko Sasaki 1, Yoshikatsu Sato 2, Tetsuya Higashiyama 1,2,
Supplementary Information Live imaging reveals the dynamics and regulation of mitochondrial nucleoids during the cell cycle in Fucci2-HeLa cells Taeko Sasaki 1, Yoshikatsu Sato 2, Tetsuya Higashiyama 1,2,
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product.
 Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish
 Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
Developmental Cell Supplemental Information A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish Julien Ablain, Ellen M. Durand, Song Yang, Yi Zhou, and Leonard I. Zon % larvae
Confocal immunofluorescence microscopy
 Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
Supplemental Materials and Methods
 Supplemental Materials and Methods 125 I-CXCL12 binding assay KG1 cells (2 10 6 ) were preincubated on ice with cold CXCL12 (1.6µg/mL corresponding to 200nM), CXCL11 (1.66µg/mL corresponding to 200nM),
Supplemental Materials and Methods 125 I-CXCL12 binding assay KG1 cells (2 10 6 ) were preincubated on ice with cold CXCL12 (1.6µg/mL corresponding to 200nM), CXCL11 (1.66µg/mL corresponding to 200nM),
Myers Lab ChIP-seq Protocol v Modified January 10, 2014
 Myers Lab ChIP-seq Protocol V011014 1 Contact information: Dr. Florencia Pauli Behn HudsonAlpha Institute for Biotechnology 601 Genome Way Huntsville, AL 35806 Telephone: 256-327-5229 Email: fpauli@hudsonalpha.org
Myers Lab ChIP-seq Protocol V011014 1 Contact information: Dr. Florencia Pauli Behn HudsonAlpha Institute for Biotechnology 601 Genome Way Huntsville, AL 35806 Telephone: 256-327-5229 Email: fpauli@hudsonalpha.org
Fatchiyah
 Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
Molecular Cell Biology - Problem Drill 11: Recombinant DNA
 Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
SMARTer Ultra Low RNA Kit for Illumina Sequencing Two powerful technologies combine to enable sequencing with ultra-low levels of RNA
 SMARTer Ultra Low RNA Kit for Illumina Sequencing Two powerful technologies combine to enable sequencing with ultra-low levels of RNA The most sensitive cdna synthesis technology, combined with next-generation
SMARTer Ultra Low RNA Kit for Illumina Sequencing Two powerful technologies combine to enable sequencing with ultra-low levels of RNA The most sensitive cdna synthesis technology, combined with next-generation
Phosphate buffered saline (PBS) for washing the cells TE buffer (nuclease-free) ph 7.5 for resuspending the SingleShot RNA control template
 Catalog # Description 172-5085 SingleShot SYBR Green Kit, 100 x 50 µl reactions For research purposes only. Introduction The SingleShot SYBR Green Kit prepares genomic DNA (gdna) free RNA directly from
Catalog # Description 172-5085 SingleShot SYBR Green Kit, 100 x 50 µl reactions For research purposes only. Introduction The SingleShot SYBR Green Kit prepares genomic DNA (gdna) free RNA directly from
Protocol for tissue-specific gene disruption in zebrafish
 Protocol for tissue-specific gene disruption in zebrafish Overview This protocol describes a method to inactivate genes in zebrafish in a tissue-specific manner. It can be used to analyze mosaic loss-of-function
Protocol for tissue-specific gene disruption in zebrafish Overview This protocol describes a method to inactivate genes in zebrafish in a tissue-specific manner. It can be used to analyze mosaic loss-of-function
SANTA CRUZ BIOTECHNOLOGY, INC.
 TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
To isolate single GNS 144 cell clones, cells were plated at a density of 1cell/well
 Supplemental Information: Supplemental Methods: Cell culture To isolate single GNS 144 cell clones, cells were plated at a density of 1cell/well in 96 well Primaria plates in GNS media and incubated at
Supplemental Information: Supplemental Methods: Cell culture To isolate single GNS 144 cell clones, cells were plated at a density of 1cell/well in 96 well Primaria plates in GNS media and incubated at
In vitro Human Umbilical Vein Endothelial Cells (HUVEC) Tube-formation Assay. Josephine MY Ko and Maria Li Lung *
 In vitro Human Umbilical Vein Endothelial Cells (HUVEC) Tube-formation Assay Josephine MY Ko and Maria Li Lung * Clinical Oncology Department, The Univerisity of Hong Kong, Hong Kong, Hong Kong SAR *For
In vitro Human Umbilical Vein Endothelial Cells (HUVEC) Tube-formation Assay Josephine MY Ko and Maria Li Lung * Clinical Oncology Department, The Univerisity of Hong Kong, Hong Kong, Hong Kong SAR *For
TruSeq ChIP Sample Preparation
 FOR RESEARCH USE ONLY Date: Illumina Kit Description: NOTE Unless familiar with the protocol in the latest version of the TruSeq ChIP Sample Preparation Guide (part # 15023092), new or less experienced
FOR RESEARCH USE ONLY Date: Illumina Kit Description: NOTE Unless familiar with the protocol in the latest version of the TruSeq ChIP Sample Preparation Guide (part # 15023092), new or less experienced
Full-length single-cell RNA-seq applied to a viral human. cancer: Applications to HPV expression and splicing analysis. Supplementary Information
 Full-length single-cell RNA-seq applied to a viral human cancer: Applications to HPV expression and splicing analysis in HeLa S3 cells Liang Wu 1, Xiaolong Zhang 1,2, Zhikun Zhao 1,3,4, Ling Wang 5, Bo
Full-length single-cell RNA-seq applied to a viral human cancer: Applications to HPV expression and splicing analysis in HeLa S3 cells Liang Wu 1, Xiaolong Zhang 1,2, Zhikun Zhao 1,3,4, Ling Wang 5, Bo
Nature Immunology: doi: /ni Supplementary Figure 1
 Supplementary Figure 1 BALB/c LYVE1-deficient mice exhibited reduced lymphatic trafficking of all DC subsets after oxazolone-induced sensitization. (a) Schematic overview of the mouse skin oxazolone contact
Supplementary Figure 1 BALB/c LYVE1-deficient mice exhibited reduced lymphatic trafficking of all DC subsets after oxazolone-induced sensitization. (a) Schematic overview of the mouse skin oxazolone contact
Confocal Microscopy Analyzes Cells
 Choosing Filters for Fluorescence A Laurin Publication Photonic Solutions for Biotechnology and Medicine November 2002 Confocal Microscopy Analyzes Cells Reprinted from the November 2002 issue of Biophotonics
Choosing Filters for Fluorescence A Laurin Publication Photonic Solutions for Biotechnology and Medicine November 2002 Confocal Microscopy Analyzes Cells Reprinted from the November 2002 issue of Biophotonics
jetcrispr RNP transfection reagent PROTOCOL
 jetcrispr RNP transfection reagent PROTOCOL DESCRIPTION jetcrispr is a RiboNucleoProtein (RNP) transfection reagent designed to perform CRISPR-Cas9 genome editing in mammalian cells. This reagent has been
jetcrispr RNP transfection reagent PROTOCOL DESCRIPTION jetcrispr is a RiboNucleoProtein (RNP) transfection reagent designed to perform CRISPR-Cas9 genome editing in mammalian cells. This reagent has been
Technical Review. Real time PCR
 Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Methods of Biomaterials Testing Lesson 3-5. Biochemical Methods - Molecular Biology -
 Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Cycles of vascular plexus formation within the nephrogenic zone of the developing mouse kidney
 1 Supplementary text and data for: 2 3 4 5 Cycles of vascular plexus formation within the nephrogenic zone of the developing mouse kidney Authors: David A. D. Munro 1*, Peter Hohenstein 2, and Jamie A.
1 Supplementary text and data for: 2 3 4 5 Cycles of vascular plexus formation within the nephrogenic zone of the developing mouse kidney Authors: David A. D. Munro 1*, Peter Hohenstein 2, and Jamie A.
Construction of plant complementation vector and generation of transgenic plants
 MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
Supplementary File 3: DNA and RNA isolation
 Supplementary File 3: DNA and RNA isolation Q-CROC-02 Biopsy protocol For the purposes of this protocol, four needle core biopsies (NCBs) of lymph node tissue are isolated from each patient using a 16G
Supplementary File 3: DNA and RNA isolation Q-CROC-02 Biopsy protocol For the purposes of this protocol, four needle core biopsies (NCBs) of lymph node tissue are isolated from each patient using a 16G
Recent technology allow production of microarrays composed of 70-mers (essentially a hybrid of the two techniques)
 Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
CHAPTER 9 DNA Technologies
 CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
Real-time PCR. Total RNA was isolated from purified splenic or LP macrophages using
 Supplementary Methods Real-time PCR. Total RNA was isolated from purified splenic or LP macrophages using the Qiagen RNeasy Mini Kit, according to the manufacturer s protocol with on-column DNase digestion
Supplementary Methods Real-time PCR. Total RNA was isolated from purified splenic or LP macrophages using the Qiagen RNeasy Mini Kit, according to the manufacturer s protocol with on-column DNase digestion
ab Hypoxic Response Human Flow Cytometry Kit
 ab126585 Hypoxic Response Human Flow Cytometry Kit Instructions for Use For measuring protein levels by flow cytometry: hypoxia-inducible factor 1-alpha (HIF1A) and BCL2/adenovirus E1B 19 kda proteininteracting
ab126585 Hypoxic Response Human Flow Cytometry Kit Instructions for Use For measuring protein levels by flow cytometry: hypoxia-inducible factor 1-alpha (HIF1A) and BCL2/adenovirus E1B 19 kda proteininteracting
scgem Workflow Experimental Design Single cell DNA methylation primer design
 scgem Workflow Experimental Design Single cell DNA methylation primer design The scgem DNA methylation assay uses qpcr to measure digestion of target loci by the methylation sensitive restriction endonuclease
scgem Workflow Experimental Design Single cell DNA methylation primer design The scgem DNA methylation assay uses qpcr to measure digestion of target loci by the methylation sensitive restriction endonuclease
Figure S2. Response of mouse ES cells to GSK3 inhibition. Mentioned in discussion
 Stem Cell Reports, Volume 1 Supplemental Information Robust Self-Renewal of Rat Embryonic Stem Cells Requires Fine-Tuning of Glycogen Synthase Kinase-3 Inhibition Yaoyao Chen, Kathryn Blair, and Austin
Stem Cell Reports, Volume 1 Supplemental Information Robust Self-Renewal of Rat Embryonic Stem Cells Requires Fine-Tuning of Glycogen Synthase Kinase-3 Inhibition Yaoyao Chen, Kathryn Blair, and Austin
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
Contents. 1 Basic Molecular Microbiology of Bacteria... 1 Exp. 1.1 Isolation of Genomic DNA Introduction Principle...
 Contents 1 Basic Molecular Microbiology of Bacteria... 1 Exp. 1.1 Isolation of Genomic DNA... 1 Introduction... 1 Principle... 1 Reagents Required and Their Role... 2 Procedure... 3 Observation... 4 Result
Contents 1 Basic Molecular Microbiology of Bacteria... 1 Exp. 1.1 Isolation of Genomic DNA... 1 Introduction... 1 Principle... 1 Reagents Required and Their Role... 2 Procedure... 3 Observation... 4 Result
QuickExtract RNA Extraction Kit
 Cat. Nos. QER09015 and QER090150 Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook (facebook.com/epicentrebio), and Twitter (@EpicentreBio). www.epicentre.com Lit. # 299 12/2012 1
Cat. Nos. QER09015 and QER090150 Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook (facebook.com/epicentrebio), and Twitter (@EpicentreBio). www.epicentre.com Lit. # 299 12/2012 1
Recombinant DNA Technology. The Role of Recombinant DNA Technology in Biotechnology. yeast. Biotechnology. Recombinant DNA technology.
 PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
Use of Drosophila Melanogaster as a Model System in the Study of Human Sodium- Dependent Multivitamin Transporter. Michael Brinton BIOL 230W.
 Use of Drosophila Melanogaster as a Model System in the Study of Human Sodium- Dependent Multivitamin Transporter Michael Brinton BIOL 230W.001 28 October 2013 TA: Sashi Gollapudi Introduction Many human
Use of Drosophila Melanogaster as a Model System in the Study of Human Sodium- Dependent Multivitamin Transporter Michael Brinton BIOL 230W.001 28 October 2013 TA: Sashi Gollapudi Introduction Many human
PreAnalytiX Supplementary Protocol
 PreAnalytiX Supplementary Protocol Purification of total RNA from microdissected PAXgene Tissue fixed, paraffin-embedded (PFPE) and PAXgene Tissue fixed, cryo-embedded (PFCE) tissues This protocol is for
PreAnalytiX Supplementary Protocol Purification of total RNA from microdissected PAXgene Tissue fixed, paraffin-embedded (PFPE) and PAXgene Tissue fixed, cryo-embedded (PFCE) tissues This protocol is for
Incorporating Molecular ID Technology. Accel-NGS 2S MID Indexing Kits
 Incorporating Molecular ID Technology Accel-NGS 2S MID Indexing Kits Molecular Identifiers (MIDs) MIDs are indices used to label unique library molecules MIDs can assess duplicate molecules in sequencing
Incorporating Molecular ID Technology Accel-NGS 2S MID Indexing Kits Molecular Identifiers (MIDs) MIDs are indices used to label unique library molecules MIDs can assess duplicate molecules in sequencing
Chapter 10: Classification of Microorganisms
 Chapter 10: Classification of Microorganisms 1. The Taxonomic Hierarchy 2. Methods of Identification 1. The Taxonomic Hierarchy Phylogenetic Tree of the 3 Domains Taxonomic Hierarchy 8 successive taxa
Chapter 10: Classification of Microorganisms 1. The Taxonomic Hierarchy 2. Methods of Identification 1. The Taxonomic Hierarchy Phylogenetic Tree of the 3 Domains Taxonomic Hierarchy 8 successive taxa
Mayumi Egawa, Kaori Mukai, Soichiro Yoshikawa, Misako Iki, Naofumi Mukaida, Yohei Kawano, Yoshiyuki Minegishi, and Hajime Karasuyama
 Immunity, Volume 38 Supplemental Information Inflammatory Monocytes Recruited to Allergic Skin Acquire an Anti-inflammatory M2 Phenotype via Basophil-Derived Interleukin-4 Mayumi Egawa, Kaori Mukai, Soichiro
Immunity, Volume 38 Supplemental Information Inflammatory Monocytes Recruited to Allergic Skin Acquire an Anti-inflammatory M2 Phenotype via Basophil-Derived Interleukin-4 Mayumi Egawa, Kaori Mukai, Soichiro
Nature Methods: doi: /nmeth Supplementary Figure 1. Retention of RNA with LabelX.
 Supplementary Figure 1 Retention of RNA with LabelX. (a) Epi-fluorescence image of single molecule FISH (smfish) against GAPDH on HeLa cells expanded without LabelX treatment. (b) Epi-fluorescence image
Supplementary Figure 1 Retention of RNA with LabelX. (a) Epi-fluorescence image of single molecule FISH (smfish) against GAPDH on HeLa cells expanded without LabelX treatment. (b) Epi-fluorescence image
Supplemental Data. LMO4 Controls the Balance between Excitatory. and Inhibitory Spinal V2 Interneurons
 Neuron, Volume 61 Supplemental Data LMO4 Controls the Balance between Excitatory and Inhibitory Spinal V2 Interneurons Kaumudi Joshi, Seunghee Lee, Bora Lee, Jae W. Lee, and Soo-Kyung Lee Supplemental
Neuron, Volume 61 Supplemental Data LMO4 Controls the Balance between Excitatory and Inhibitory Spinal V2 Interneurons Kaumudi Joshi, Seunghee Lee, Bora Lee, Jae W. Lee, and Soo-Kyung Lee Supplemental
SUPPLEMENTARY MATERIAL AND METHODS
 SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
DNA Arrays Affymetrix GeneChip System
 DNA Arrays Affymetrix GeneChip System chip scanner Affymetrix Inc. hybridization Affymetrix Inc. data analysis Affymetrix Inc. mrna 5' 3' TGTGATGGTGGGAATTGGGTCAGAAGGACTGTGGGCGCTGCC... GGAATTGGGTCAGAAGGACTGTGGC
DNA Arrays Affymetrix GeneChip System chip scanner Affymetrix Inc. hybridization Affymetrix Inc. data analysis Affymetrix Inc. mrna 5' 3' TGTGATGGTGGGAATTGGGTCAGAAGGACTGTGGGCGCTGCC... GGAATTGGGTCAGAAGGACTGTGGC
Genetic Engineering & Recombinant DNA
 Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr.
 MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
Gene Expression Profiling and Validation Using Agilent SurePrint G3 Gene Expression Arrays
 Gene Expression Profiling and Validation Using Agilent SurePrint G3 Gene Expression Arrays Application Note Authors Bahram Arezi, Nilanjan Guha and Anne Bergstrom Lucas Agilent Technologies Inc. Santa
Gene Expression Profiling and Validation Using Agilent SurePrint G3 Gene Expression Arrays Application Note Authors Bahram Arezi, Nilanjan Guha and Anne Bergstrom Lucas Agilent Technologies Inc. Santa
RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by. 5 AGACACAAACACCAUUGUCACACUCCACAGC; Rand-2 OMe,
 Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Materials and methods. by University of Washington Yeast Resource Center) from several promoters, including
 Supporting online material for Elowitz et al. report Materials and methods Strains and plasmids. Plasmids expressing CFP or YFP (wild-type codons, developed by University of Washington Yeast Resource Center)
Supporting online material for Elowitz et al. report Materials and methods Strains and plasmids. Plasmids expressing CFP or YFP (wild-type codons, developed by University of Washington Yeast Resource Center)
Long and short/small RNA-seq data analysis
 Long and short/small RNA-seq data analysis GEF5, 4.9.2015 Sami Heikkinen, PhD, Dos. Topics 1. RNA-seq in a nutshell 2. Long vs short/small RNA-seq 3. Bioinformatic analysis work flows GEF5 / Heikkinen
Long and short/small RNA-seq data analysis GEF5, 4.9.2015 Sami Heikkinen, PhD, Dos. Topics 1. RNA-seq in a nutshell 2. Long vs short/small RNA-seq 3. Bioinformatic analysis work flows GEF5 / Heikkinen
Supplementary Figure 1: Derivation and characterization of RN ips cell lines. (a) RN ips cells maintain expression of pluripotency markers OCT4 and
 Supplementary Figure 1: Derivation and characterization of RN ips cell lines. (a) RN ips cells maintain expression of pluripotency markers OCT4 and SSEA4 after 10 passages in mtesr 1 medium. (b) Schematic
Supplementary Figure 1: Derivation and characterization of RN ips cell lines. (a) RN ips cells maintain expression of pluripotency markers OCT4 and SSEA4 after 10 passages in mtesr 1 medium. (b) Schematic
Transfection of Mouse ES Cells and Mouse ips cells using the Stemfect 2.0 -mesc Transfection Reagent
 APPLICATION NOTE Page 1 Transfection of Mouse ES Cells and Mouse ips cells using the Stemfect 2.0 -mesc Transfection Reagent Authors: Amelia L. Cianci 1, Xun Cheng 1 and Kerry P. Mahon 1,2 1 Stemgent Inc.,
APPLICATION NOTE Page 1 Transfection of Mouse ES Cells and Mouse ips cells using the Stemfect 2.0 -mesc Transfection Reagent Authors: Amelia L. Cianci 1, Xun Cheng 1 and Kerry P. Mahon 1,2 1 Stemgent Inc.,
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Peter Dy, Weihuan Wang, Pallavi Bhattaram, Qiuqing Wang, Lai Wang, R. Tracy Ballock, and Véronique Lefebvre
 Developmental Cell, Volume 22 Supplemental Information Sox9 Directs Hypertrophic Maturation and Blocks Osteoblast Differentiation of Growth Plate Chondrocytes Peter Dy, Weihuan Wang, Pallavi Bhattaram,
Developmental Cell, Volume 22 Supplemental Information Sox9 Directs Hypertrophic Maturation and Blocks Osteoblast Differentiation of Growth Plate Chondrocytes Peter Dy, Weihuan Wang, Pallavi Bhattaram,
Computational Biology I LSM5191
 Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
To assess the localization of Citrine fusion proteins, we performed antibody staining to
 Trinh et al 1 SUPPLEMENTAL MATERIAL FlipTraps recapitulate endogenous protein localization To assess the localization of Citrine fusion proteins, we performed antibody staining to compare the expression
Trinh et al 1 SUPPLEMENTAL MATERIAL FlipTraps recapitulate endogenous protein localization To assess the localization of Citrine fusion proteins, we performed antibody staining to compare the expression
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX
 Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup
 Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup 1. Introduction The data produced by IHEC is illustrated in Figure 1. Figure 1. The space of epigenomic
Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup 1. Introduction The data produced by IHEC is illustrated in Figure 1. Figure 1. The space of epigenomic
Using Low Input of Poly (A) + RNA and Total RNA for Oligonucleotide Microarrays Application
 Using Low Input of Poly (A) + RNA and Total RNA for Oligonucleotide Microarrays Application Gene Expression Author Michelle M. Chen Agilent Technologies, Inc. 3500 Deer Creek Road, MS 25U-7 Palo Alto,
Using Low Input of Poly (A) + RNA and Total RNA for Oligonucleotide Microarrays Application Gene Expression Author Michelle M. Chen Agilent Technologies, Inc. 3500 Deer Creek Road, MS 25U-7 Palo Alto,
Product Name : Simple mirna Detection Kit
 Product Name : Simple mirna Detection Kit Code No. : DS700 This product is for research use only Kit Contents This kit provides sufficient reagents to perform 20 reactions for detecting microrna. Components
Product Name : Simple mirna Detection Kit Code No. : DS700 This product is for research use only Kit Contents This kit provides sufficient reagents to perform 20 reactions for detecting microrna. Components
GM130 Is Required for Compartmental Organization of Dendritic Golgi Outposts
 Current Biology, Volume 24 Supplemental Information GM130 Is Required for Compartmental Organization of Dendritic Golgi Outposts Wei Zhou, Jin Chang, Xin Wang, Masha G. Savelieff, Yinyin Zhao, Shanshan
Current Biology, Volume 24 Supplemental Information GM130 Is Required for Compartmental Organization of Dendritic Golgi Outposts Wei Zhou, Jin Chang, Xin Wang, Masha G. Savelieff, Yinyin Zhao, Shanshan
A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
Roche Molecular Biochemicals Technical Note No. LC 10/2000
 Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Expression Array System
 Integrated Science for Gene Expression Applied Biosystems Expression Array System Expression Array System SEE MORE GENES The most complete, most sensitive system for whole genome expression analysis. The
Integrated Science for Gene Expression Applied Biosystems Expression Array System Expression Array System SEE MORE GENES The most complete, most sensitive system for whole genome expression analysis. The
BENG 183 Trey Ideker. Genome Assembly and Physical Mapping
 BENG 183 Trey Ideker Genome Assembly and Physical Mapping Reasons for sequencing Complete genome sequencing!!! Resequencing (Confirmatory) E.g., short regions containing single nucleotide polymorphisms
BENG 183 Trey Ideker Genome Assembly and Physical Mapping Reasons for sequencing Complete genome sequencing!!! Resequencing (Confirmatory) E.g., short regions containing single nucleotide polymorphisms
Archer ALK, RET, ROS1 Fusion Detection v1 Illumina Platform
 Archer ALK, RET, ROS1 Fusion Detection v1 Illumina Platform P/N AK0001-8 Archer ALK, RET, ROS1 Fusion Detection v1 for Illumina Platform IFU-AK0001-8 Rev. A Table of Contents Product Description..3 Workflow
Archer ALK, RET, ROS1 Fusion Detection v1 Illumina Platform P/N AK0001-8 Archer ALK, RET, ROS1 Fusion Detection v1 for Illumina Platform IFU-AK0001-8 Rev. A Table of Contents Product Description..3 Workflow
SIMS2003. Instructors:Rus Yukhananov, Alex Loguinov BWH, Harvard Medical School. Introduction to Microarray Technology.
 SIMS2003 Instructors:Rus Yukhananov, Alex Loguinov BWH, Harvard Medical School Introduction to Microarray Technology. Lecture 1 I. EXPERIMENTAL DETAILS II. ARRAY CONSTRUCTION III. IMAGE ANALYSIS Lecture
SIMS2003 Instructors:Rus Yukhananov, Alex Loguinov BWH, Harvard Medical School Introduction to Microarray Technology. Lecture 1 I. EXPERIMENTAL DETAILS II. ARRAY CONSTRUCTION III. IMAGE ANALYSIS Lecture
3.1.4 DNA Microarray Technology
 3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
De novo metatranscriptome assembly and coral gene expression profile of Montipora capitata with growth anomaly
 Additional File 1 De novo metatranscriptome assembly and coral gene expression profile of Montipora capitata with growth anomaly Monika Frazier, Martin Helmkampf, M. Renee Bellinger, Scott Geib, Misaki
Additional File 1 De novo metatranscriptome assembly and coral gene expression profile of Montipora capitata with growth anomaly Monika Frazier, Martin Helmkampf, M. Renee Bellinger, Scott Geib, Misaki
Stellaris RNA FISH Protocol for Simultaneous IF + FISH in Adherent Cells
 Stellaris RNA FISH Protocol for Simultaneous IF + FISH in Adherent Cells General Protocol & Storage Product Description A set of Stellaris RNA FISH Probes is comprised of up to 48 singly labeled oligonucleotides
Stellaris RNA FISH Protocol for Simultaneous IF + FISH in Adherent Cells General Protocol & Storage Product Description A set of Stellaris RNA FISH Probes is comprised of up to 48 singly labeled oligonucleotides
Welcome to the NGS webinar series
 Welcome to the NGS webinar series Webinar 1 NGS: Introduction to technology, and applications NGS Technology Webinar 2 Targeted NGS for Cancer Research NGS in cancer Webinar 3 NGS: Data analysis for genetic
Welcome to the NGS webinar series Webinar 1 NGS: Introduction to technology, and applications NGS Technology Webinar 2 Targeted NGS for Cancer Research NGS in cancer Webinar 3 NGS: Data analysis for genetic
Guide-it Indel Identification Kit User Manual
 Clontech Laboratories, Inc. Guide-it Indel Identification Kit User Manual Cat. No. 631444 (120114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra Bella Avenue, Mountain View, CA 94043, USA
Clontech Laboratories, Inc. Guide-it Indel Identification Kit User Manual Cat. No. 631444 (120114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra Bella Avenue, Mountain View, CA 94043, USA
Supplementary information, Figure S1
 Supplementary information, Figure S1 (A) Schematic diagram of the sgrna and hspcas9 expression cassettes in a single binary vector designed for Agrobacterium-mediated stable transformation of Arabidopsis
Supplementary information, Figure S1 (A) Schematic diagram of the sgrna and hspcas9 expression cassettes in a single binary vector designed for Agrobacterium-mediated stable transformation of Arabidopsis
Cloning small RNAs for Solexa Sequencing version 2.0 by Nelson Lau Page 1 of 5 (Modified from Solexa sequencing protocol from Bartel lab)
 Cloning small RNAs for Solexa Sequencing version 2.0 by Nelson Lau 09162008 Page 1 of 5 General Cloning Protocol: Gel-purification 1. Pour 1mm thick, urea denaturing 10% or 15% polyacrylamide gels, with
Cloning small RNAs for Solexa Sequencing version 2.0 by Nelson Lau 09162008 Page 1 of 5 General Cloning Protocol: Gel-purification 1. Pour 1mm thick, urea denaturing 10% or 15% polyacrylamide gels, with
PrimeScript RT reagent Kit (Perfect Real Time)
 Cat. # RR037A For Research Use PrimeScript RT reagent Kit (Perfect Real Time) Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Storage... 3 IV. Features... 4 V. Precautions...
Cat. # RR037A For Research Use PrimeScript RT reagent Kit (Perfect Real Time) Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Storage... 3 IV. Features... 4 V. Precautions...
Dengue Virus subtypes 1,2 3 and 4
 TM Primerdesign Ltd Dengue Virus subtypes 1,2 3 and 4 3 Untranslated Region (3 UTR) genesig Standard Kit 150 tests For general laboratory and research use only 1 Introduction to Dengue Virus subtypes 1,2
TM Primerdesign Ltd Dengue Virus subtypes 1,2 3 and 4 3 Untranslated Region (3 UTR) genesig Standard Kit 150 tests For general laboratory and research use only 1 Introduction to Dengue Virus subtypes 1,2
Supplementary Information
 Supplementary Information Supplementary Figure 1. ZBTB20 expression in the developing DRG. ZBTB20 expression in the developing DRG was detected by immunohistochemistry using anti-zbtb20 antibody 9A10 on
Supplementary Information Supplementary Figure 1. ZBTB20 expression in the developing DRG. ZBTB20 expression in the developing DRG was detected by immunohistochemistry using anti-zbtb20 antibody 9A10 on
ab CytoPainter ER Staining Kit Red Fluorescence
 ab139482 CytoPainter ER Staining Kit Red Fluorescence Instructions for Use Designed to detect Human endoplasmic reticulum by microscopy. This product is for research use only and is not intended for diagnostic
ab139482 CytoPainter ER Staining Kit Red Fluorescence Instructions for Use Designed to detect Human endoplasmic reticulum by microscopy. This product is for research use only and is not intended for diagnostic
Targeted modification of gene function exploiting homology directed repair of TALENmediated double strand breaks in barley
 Targeted modification of gene function exploiting homology directed repair of TALENmediated double strand breaks in barley Nagaveni Budhagatapalli a, Twan Rutten b, Maia Gurushidze a, Jochen Kumlehn a,
Targeted modification of gene function exploiting homology directed repair of TALENmediated double strand breaks in barley Nagaveni Budhagatapalli a, Twan Rutten b, Maia Gurushidze a, Jochen Kumlehn a,
CytoPainter Golgi Staining Kit Green Fluorescence
 ab139483 CytoPainter Golgi Staining Kit Green Fluorescence Instructions for Use Designed for the detection of Golgi bodies by microscopy This product is for research use only and is not intended for diagnostic
ab139483 CytoPainter Golgi Staining Kit Green Fluorescence Instructions for Use Designed for the detection of Golgi bodies by microscopy This product is for research use only and is not intended for diagnostic
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling
 Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
EdU Flow Cytometry Kit. User Manual
 User Manual Ordering information: (for detailed kit content see Table 2) EdU Flow Cytometry Kits for 50 assays: Product number EdU Used fluorescent dye BCK-FC488-50 10 mg 6-FAM Azide BCK-FC555-50 10 mg
User Manual Ordering information: (for detailed kit content see Table 2) EdU Flow Cytometry Kits for 50 assays: Product number EdU Used fluorescent dye BCK-FC488-50 10 mg 6-FAM Azide BCK-FC555-50 10 mg
Supporting Information
 Supporting Information Liu et al. 10.1073/pnas.0901216106 SI Materials and Methods Reagents. Thioglycolate and LPS from Escherichia coli 0111:B4 were from Sigma-Aldrich. PAM3CSK4 and poly(i:c) were from
Supporting Information Liu et al. 10.1073/pnas.0901216106 SI Materials and Methods Reagents. Thioglycolate and LPS from Escherichia coli 0111:B4 were from Sigma-Aldrich. PAM3CSK4 and poly(i:c) were from
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Convoy TM Transfection Reagent
 Convoy TM Transfection Reagent Catalog No.11103 0.25ml (40-80 transfections in 35mm dishes) Catalog No.11105 0.5 ml (80-165 transfections in 35mm dishes) Catalog No.11110 1.0 ml (165-330 transfections
Convoy TM Transfection Reagent Catalog No.11103 0.25ml (40-80 transfections in 35mm dishes) Catalog No.11105 0.5 ml (80-165 transfections in 35mm dishes) Catalog No.11110 1.0 ml (165-330 transfections
Globin Block Modules for QuantSeq Instruction Manual
 Globin Block Modules for QuantSeq Instruction Manual Catalog Numbers: 070 (RS-Globin Block, Homo sapiens, 96 rxn) 071 (RS-Globin Block, Sus scrofa, 96 rxn) 015 (QuantSeq 3 mrna-seq Library Prep Kit for
Globin Block Modules for QuantSeq Instruction Manual Catalog Numbers: 070 (RS-Globin Block, Homo sapiens, 96 rxn) 071 (RS-Globin Block, Sus scrofa, 96 rxn) 015 (QuantSeq 3 mrna-seq Library Prep Kit for
MMLV Reverse Transcriptase 1st-Strand cdna Synthesis Kit
 MMLV Reverse Transcriptase 1st-Strand cdna Synthesis Kit Cat. No. MM070150 Available exclusively thru Lucigen. lucigen.com/epibio www.lucigen.com MA265E MMLV Reverse Transcriptase 1st-Strand cdna Synthesis
MMLV Reverse Transcriptase 1st-Strand cdna Synthesis Kit Cat. No. MM070150 Available exclusively thru Lucigen. lucigen.com/epibio www.lucigen.com MA265E MMLV Reverse Transcriptase 1st-Strand cdna Synthesis
SUPPLEMENTARY INFORMATION. Transcriptional output transiently spikes upon mitotic exit
 SUPPLEMENTARY INFORMATION Transcriptional output transiently spikes upon mitotic exit Viola Vaňková Hausnerová 1, 2, Christian Lanctôt 1* 1 BIOCEV and Department of Cell Biology, Faculty of Science, Charles
SUPPLEMENTARY INFORMATION Transcriptional output transiently spikes upon mitotic exit Viola Vaňková Hausnerová 1, 2, Christian Lanctôt 1* 1 BIOCEV and Department of Cell Biology, Faculty of Science, Charles
HiPer RT-PCR Teaching Kit
 HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing
 Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
Supplementary Information Design of small molecule-responsive micrornas based on structural requirements for Drosha processing Chase L. Beisel, Yvonne Y. Chen, Stephanie J. Culler, Kevin G. Hoff, & Christina
Return to Web Version
 Return to Web Version PKH Linker Kits BioFiles 2007, 2.5, 22. PKH Linker Kits for Fluorescent Cell Labeling Features Achieve stable, uniform, intense, and reproducible fluorescent labeling of live cells
Return to Web Version PKH Linker Kits BioFiles 2007, 2.5, 22. PKH Linker Kits for Fluorescent Cell Labeling Features Achieve stable, uniform, intense, and reproducible fluorescent labeling of live cells
SOLiD Total RNA-Seq Kit SOLiD RNA Barcoding Kit
 SOLiD Total RNA-Seq Kit SOLiD RNA Barcoding Kit Agenda SOLiD Total RNAseq Kit Overview Kit Configurations Barcoding Kit Introduction New Small RNA and WT Workflow Small RNA Workflow Step-by-step Workflow
SOLiD Total RNA-Seq Kit SOLiD RNA Barcoding Kit Agenda SOLiD Total RNAseq Kit Overview Kit Configurations Barcoding Kit Introduction New Small RNA and WT Workflow Small RNA Workflow Step-by-step Workflow
BD Single-Cell Multiplexing Kit Human Protocol
 BD Single-Cell Multiplexing Kit Protocol For use with the BD Rhapsody system 11/2017 Doc ID: 54478 Rev. 1.0 Contents Overview on page 3 Sample Tag library preparation workflow on page 4 Reference for BD
BD Single-Cell Multiplexing Kit Protocol For use with the BD Rhapsody system 11/2017 Doc ID: 54478 Rev. 1.0 Contents Overview on page 3 Sample Tag library preparation workflow on page 4 Reference for BD
Post-expansion antibody delivery, after epitope-preserving homogenization.
 Supplementary Figure 1 Post-expansion antibody delivery, after epitope-preserving homogenization. (a, b) Wide-field fluorescence images of Thy1-YFP-expressing mouse brain hemisphere slice before expansion
Supplementary Figure 1 Post-expansion antibody delivery, after epitope-preserving homogenization. (a, b) Wide-field fluorescence images of Thy1-YFP-expressing mouse brain hemisphere slice before expansion
Figure S1. Purity of primary cultures of renal proximal tubular epithelial culture ascertained by cytokeratin staining.
 Supplementary information Supplementary figures Figure S1. Purity of primary cultures of renal proximal tubular epithelial culture ascertained by cytokeratin staining. Figure S2. Induction of Nur77 in
Supplementary information Supplementary figures Figure S1. Purity of primary cultures of renal proximal tubular epithelial culture ascertained by cytokeratin staining. Figure S2. Induction of Nur77 in
Supplementary Figures Montero et al._supplementary Figure 1
 Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
SUPPORTING INFORMATION. Optical Control of CRISPR/Cas9 Gene Editing
 SUPPORTING INFORMATION Optical Control of CRISPR/ Gene Editing James Hemphill 1, Erin K. Borchardt 2, Kalyn Brown 1, Aravind Asokan 2, and Alexander Deiters 1 * 1 University of Pittsburgh, Department of
SUPPORTING INFORMATION Optical Control of CRISPR/ Gene Editing James Hemphill 1, Erin K. Borchardt 2, Kalyn Brown 1, Aravind Asokan 2, and Alexander Deiters 1 * 1 University of Pittsburgh, Department of
Plasmid DNA transfection of SW480 human colorectal cancer cells with the Biontex K2 Transfection System
 Plasmid DNA transfection of human colorectal cancer cells with the Biontex K2 Transfection System Stephanie Hehlgans and Franz Rödel, Department of Radiotherapy and Oncology, Goethe- University Frankfurt,
Plasmid DNA transfection of human colorectal cancer cells with the Biontex K2 Transfection System Stephanie Hehlgans and Franz Rödel, Department of Radiotherapy and Oncology, Goethe- University Frankfurt,
