Gene Prediction: Preliminary Results
|
|
|
- Tamsyn Rosemary Daniel
- 6 years ago
- Views:
Transcription
1 Gene Prediction: Preliminary Results
2 Outline Preliminary Pipeline Programs Program Comparison Tests Metrics Gene Prediction Tools: Usage + Results GeneMarkS Glimmer 3.0 Prodigal BLAST ncrna Prediction Tools: Usage + Results trnascan SE RNAmmer RFAM Further Steps
3 Preliminary Pipeline
4 Programs Tested Concerned might be redundant during initial research, but test metrics suggest we should add this to our pipeline Easy Gene was tested at Web Interface stage, but we did not get to test it in depth before this presentation. We plan to test it. During initial research, we wanted to steer clear from programs originally designed for non-bacterial genomes, since training is a major portion of ab initio gene prediction. Since homologous options were not present for our species, we decided it was best to focus on BLAST for homologous Gene Prediction.
5 Log-Odds Log-odds is a common scoring metric used for predicted genes. 1st Matrix shows probability of possible alignment pairs at random 2nd shows probability of possible alignment pairs in your query Final Matrix shows value of 2nd/1st, so likelihood the pair is not observed at random Take log (usually Ln) of likelihoods any values between 0 and 1 become negative any greater than 1 become positive
6 Methods Used RefSeq FTP to obtain annotation for FAM18 A.gff file was obtained with the start, stop, strand and type information for portions of the genome
7 FAM18 summary - Species: Neisseria meningitidis - Serogroup: C Genes CDS Exon trna rrna Total Strand Strand
8 Comparing Tools - Tests Metric True Positive (Rightly Predicted) False Positive (Over Predicted) False Negative (Under Predicted) True Negative Predicted Set Annotated Set
9 Comparing Tools - Metrics Sensitivity: ability to exclude false positives Precision: ability to predict maximum number of genes.
10 Summary PROGRAM Entries Predicted (TP) Unpredicted (FN) Overpredecited (FP) AMIGene GeneMarkS Prodigal trnascan Rfam
11 Summary PROGRAM Sensitivity Precision AMIGene GeneMarkS Prodigal trnascan Rfam
12 Summary
13 GeneMarkS Usage - Sequence Type - Prokaryotic Intronless Eukaryotic Virus Phage EST/cDNA - Output Format (-format) - GFF/LST - Omit Overlaps (-offover) Add to path: /home/yasvanth3/gm/genemark_suite_linux_64/gmsuite/ Example: gmsn.pl -prok <inputfilename.fasta> -format <output format>
14 GeneMarkS Output - GFF File - Gene Name, Start, Stop, Gene ID, Length, Gene Score
15 Glimmer 3.0 Usage 4 steps: Input file : sequence.fa Add to path: /home/vvenkat6/bin/ > long-orfs -n sequence.fa sequence.orf > extract sequence.fa sequence.orf > sequence.train > build-icm -r sequence.icm < sequence.train > glimmer3 sequence.fa sequence.icm out Output file: i) out.predict ii) out.detail
16 out.detail
17 out.predict
18 Prodigal Implements simple log-likelihood scoring functions unlike the previous programs which use complicated HMMs and IMMs Performs well for high GC content Genomes Trade off between # of FPs and TPs
19 Command Used : prodigal.linux -i input_file_name -o output_file_name -f output_format -d nucleotide_sequences_of_all_genes -a protein_sequences_of_all_genes -s potential_genes_with_scores The mode can be specified as well using -p flag. Different output formats can be specified gbk: Genbank-like format (Default) gff: GFF format sqn: Sequin feature table format sco: Simple coordinate output Total No. of Genes Predicted : =1416 in the CISA_all file grep -w "-" out_cisa_all.gff wc -l 645 grep -w "+" out_cisa_all.gff wc -l 771
20 Output File Generated
21 BLAST Step1: Create blast database makeblastdb in FAM18.fasta -dbtype 'nucl' -out FAM18_db RESOURCES: MAKEBLAST: /home/rnagilla3/bin/blast/ncbi-blast /bin/makeblastdb Input file: /home/rnagilla3/assignment_data/fam18.fasta Step2: Run blastn blastn -db FAM18_db query CISA_all.fa outfmt 6 -out BLAST_OUTPUT RESOURCES: BLASTN: /home/rnagilla3/bin/blast/ncbi-blast /bin/blastn QUERY File: /home/yasvanth3/gm/cisa_all.fa
22 BLAST output format
23 Non-coding RNA Prediction
24 trnascan-se command line /home/tmi7/bin/trnascan-se Some Options: -B: search for bacterial trnas (use bacterial trna model) -C: search using Cove analysis only slow, sensitive) -o: save tabular result to... -f: save trna secondary structures to... -m: save statistics summary to...
25 trnascan-se result on CISA_all.fa
26 trnascan-se result on CISA_all.fasta
27 RNAmmer 1.2 /home/akelley35/bin rnammer [-S kingdom] [-m molecules] [-xml xml-file] [-gff gff-file] [-h hmmreport] [-f fasta-file] [sequence] -S Specifies the super kingdom of the input sequence, euk, bac, arc -m Molecule type can be 'tsu' for 5/8s rrna, 'ssu' for 16/18s rrna, 'lsu' for 23/28s rrna or any combination separated by comma -xml,-gff,-h,-f The types of outputs generated.
28 Infernal (Rfam) path to the installed file: /home/tmi7/bin/ Step 1: create an CM database flatfile download from Rfam Step 2: compress and index the flatfile with cmpress cmpress <cmdb> Step 3:search the CM database with cmscan cmscan --noali -E <x> -o <f> --noali <cmdb> <seqfile> : don't output alignments -E <x> : report sequences <= this E-value threshold in output -o <f> : direct output to file <f>
29 Infernal (Rfam) Results
30 Infernal (Rfam) Results
31 ncrna prediction results Prediction Tools FAM18 CISA_all.fasta RNAmmer 12 3 trnascan-se Rfam cmscan (rrna) 12 8 Rfam cmscan (trna) Rfam cmscan (other) 22 25
32 Pipeline Changes AMIgene EasyGene? RSAT - Lower priority Adds non-coding regulatory portions of genome, but want to focus on coding portions first
33 Challenges - Use GenePRIMP to combine GeneMarkS and Prodigal results - Find a way to combine outputs to minimize False Positives and False Negatives - Use a confidence system - Highest confidence is genes confirmed by all relevant outputs that do not contradict - Resolve conflicting results - use database references - compare conflicts and pick higher score or more likely gene prediction - How to determine likely vs unlikely Theoretical Genes (not over predict)
34 Further Steps - Continue testing/comparing programs - Schema for Naming Genes - derive from GeneID, contig # and Sample ID - Finalize Method for Merging Results - i.e. investigate GENE Primp (Gene Prediction Improvement Pipeline) for ideas - Use metrics mentioned previously to filter results of individual programs
35 References Lagesen, Karin, et al. "RNAmmer: consistent and rapid annotation of ribosomal RNA genes." Nucleic acids research 35.9 (2007): Besemer, John, Alexandre Lomsadze, and Mark Borodovsky. "GeneMarkS: a self-training method for prediction of gene starts in microbial genomes. Implications for finding sequence motifs in regulatory regions." Nucleic Acids Research (2001): Burge, Sarah W., et al. "Rfam 11.0: 10 years of RNA families." Nucleic acids research (2012): gks1005. Schattner, Peter, Angela N. Brooks, and Todd M. Lowe. "The trnascan-se, snoscan and snogps web servers for the detection of trnas and snornas." Nucleic acids research 33. suppl 2 (2005): W686-W689. Delcher, Arthur L. et al. Improved microbial gene prediction with GLIMMER. Nucleic Acids Res (23): Delcher, Arthur L. et al. Identifying bacterial genes and endosymbiont DNA with Glimmer. Bioinformatics (2007). 23 (6):
36 LAB ACTIVITY! - In class: - BLAST - GeneMarkS - Rfam - Run programs. Create file on your server folder labeled GenePredHWOut and place output files there - Parts done in groups - one person me before you leave with the files generated together, who worked together, and where I can find these output files - Write answers/observations in a text file marked appropriately. this to me ( below) with subject Gene Prediction HW Answers - Name the file FIRSTNAME_LASTNAME.txt - rachel.kutner06@gmail.com Will provide instructions and grading scheme for those absent/in case you don t finish. Due next Friday at midnight.
37 Grading Attendance in class: 10% Completion of exercise: 30% Proper answers: 40% each question is 5 points Proper output files: 20%
38 BLAST You will have to run Blast using unknown sequence query as query against a known reference database sequence. So, you have to create a blast database with reference.fasta and blast query against this database and submit the results. Both the query files and database reference files are located in /home/rnagilla3/assignment_data/) [ Please write to Roopa, roopareddynagilla@gatech.edu for any permission issues ] MAKEBLAST: /home/rnagilla3/bin/blast/ncbi-blast /bin/makeblastdb BLASTN: /home/rnagilla3/bin/blast/ncbi-blast /bin/blastn Query sequence: query.fasta Reference sequence: reference.fasta (N. meningitidis) 1. How is the output sorted? 2. What is e-value and why is it significant? 3. Pick one of the top homologous sequences for FAM18 and what do you think is the species the sequence is most related to?
39 GeneMarkS You will have to run GeneMarkS with the FAM18 fasta file. The FAM18 file can be found in /home/yasvanth3/fam18.fasta and GeneMarkS (gmsn.pl) can be run from /home/yasvanth3/gm/genemark_suite_linux_64/gmsuite/ Assume the species is unknown and use the appropriate command and parameters to produce a GFF file. Is RBS True or False? (Ribosomal Binding Site) Describe the Format of the Output and List the types of scores that are available.
40 Infernal You will have to run the cmscan program for FAM18 sequence against Rfam CM database. The cmscan program is in /home/tmi7/bin/cmscan The FAM18 can be found in /home/tmi7/fam18.fasta The Rfam CM database is in /home/tmi7/cms/rfam.cm Using parameters of no alignment(--noali) and use a E value of 1E10 (-E), and save a output file(-o). How many cmscan hits did you get? How many ribosomal RNA are there in your output? (There may be some redundant results)
Leonardo Mariño-Ramírez, PhD NCBI / NLM / NIH. BIOL 7210 A Computational Genomics 2/18/2015
 Leonardo Mariño-Ramírez, PhD NCBI / NLM / NIH BIOL 7210 A Computational Genomics 2/18/2015 The $1,000 genome is here! http://www.illumina.com/systems/hiseq-x-sequencing-system.ilmn Bioinformatics bottleneck
Leonardo Mariño-Ramírez, PhD NCBI / NLM / NIH BIOL 7210 A Computational Genomics 2/18/2015 The $1,000 genome is here! http://www.illumina.com/systems/hiseq-x-sequencing-system.ilmn Bioinformatics bottleneck
Genome sequence of Acinetobacter baumannii MDR-TJ
 JB Accepts, published online ahead of print on 11 March 2011 J. Bacteriol. doi:10.1128/jb.00226-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
JB Accepts, published online ahead of print on 11 March 2011 J. Bacteriol. doi:10.1128/jb.00226-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
GeneMarkS-2: Raising Standards of Accuracy in Gene Recognition
 GeneMarkS-2: Raising Standards of Accuracy in Gene Recognition Alexandre Lomsadze 1^, Shiyuyun Tang 2^, Karl Gemayel 3^ and Mark Borodovsky 1,2,3 ^ joint first authors 1 Wallace H. Coulter Department of
GeneMarkS-2: Raising Standards of Accuracy in Gene Recognition Alexandre Lomsadze 1^, Shiyuyun Tang 2^, Karl Gemayel 3^ and Mark Borodovsky 1,2,3 ^ joint first authors 1 Wallace H. Coulter Department of
Why learn sequence database searching? Searching Molecular Databases with BLAST
 Why learn sequence database searching? Searching Molecular Databases with BLAST What have I cloned? Is this really!my gene"? Basic Local Alignment Search Tool How BLAST works Interpreting search results
Why learn sequence database searching? Searching Molecular Databases with BLAST What have I cloned? Is this really!my gene"? Basic Local Alignment Search Tool How BLAST works Interpreting search results
Analysis Report. Institution : Macrogen Japan Name : Macrogen Japan Order Number : 1501APB-0004 Sample Name : 8380 Type of Analysis : De novo assembly
 Analysis Report Institution : Macrogen Japan Name : Macrogen Japan Order Number : 1501APB-0004 Sample Name : 8380 Type of Analysis : De novo assembly 1 Table of Contents 1. Result of Whole Genome Assembly
Analysis Report Institution : Macrogen Japan Name : Macrogen Japan Order Number : 1501APB-0004 Sample Name : 8380 Type of Analysis : De novo assembly 1 Table of Contents 1. Result of Whole Genome Assembly
UCSC Genome Browser. Introduction to ab initio and evidence-based gene finding
 UCSC Genome Browser Introduction to ab initio and evidence-based gene finding Wilson Leung 06/2006 Outline Introduction to annotation ab initio gene finding Basics of the UCSC Browser Evidence-based gene
UCSC Genome Browser Introduction to ab initio and evidence-based gene finding Wilson Leung 06/2006 Outline Introduction to annotation ab initio gene finding Basics of the UCSC Browser Evidence-based gene
COMPUTER RESOURCES II:
 COMPUTER RESOURCES II: Using the computer to analyze data, using the internet, and accessing online databases Bio 210, Fall 2006 Linda S. Huang, Ph.D. University of Massachusetts Boston In the first computer
COMPUTER RESOURCES II: Using the computer to analyze data, using the internet, and accessing online databases Bio 210, Fall 2006 Linda S. Huang, Ph.D. University of Massachusetts Boston In the first computer
Computational analysis of non-coding RNA. Andrew Uzilov BME110 Tue, Nov 16, 2010
 Computational analysis of non-coding RNA Andrew Uzilov auzilov@ucsc.edu BME110 Tue, Nov 16, 2010 1 Corrected/updated talk slides are here: http://tinyurl.com/uzilovrna redirects to: http://users.soe.ucsc.edu/~auzilov/bme110/fall2010/
Computational analysis of non-coding RNA Andrew Uzilov auzilov@ucsc.edu BME110 Tue, Nov 16, 2010 1 Corrected/updated talk slides are here: http://tinyurl.com/uzilovrna redirects to: http://users.soe.ucsc.edu/~auzilov/bme110/fall2010/
Outline. Gene Finding Questions. Recap: Prokaryotic gene finding Eukaryotic gene finding The human gene complement Regulation
 Tues, Nov 29: Gene Finding 1 Online FCE s: Thru Dec 12 Thurs, Dec 1: Gene Finding 2 Tues, Dec 6: PS5 due Project presentations 1 (see course web site for schedule) Thurs, Dec 8 Final papers due Project
Tues, Nov 29: Gene Finding 1 Online FCE s: Thru Dec 12 Thurs, Dec 1: Gene Finding 2 Tues, Dec 6: PS5 due Project presentations 1 (see course web site for schedule) Thurs, Dec 8 Final papers due Project
ELE4120 Bioinformatics. Tutorial 5
 ELE4120 Bioinformatics Tutorial 5 1 1. Database Content GenBank RefSeq TPA UniProt 2. Database Searches 2 Databases A common situation for alignment is to search through a database to retrieve the similar
ELE4120 Bioinformatics Tutorial 5 1 1. Database Content GenBank RefSeq TPA UniProt 2. Database Searches 2 Databases A common situation for alignment is to search through a database to retrieve the similar
RNA Genomics II. BME 110: CompBio Tools Todd Lowe & Andrew Uzilov May 17, 2011
 RNA Genomics II BME 110: CompBio Tools Todd Lowe & Andrew Uzilov May 17, 2011 1 TIME Why RNA? An evolutionary perspective The RNA World hypotheses: life arose as self-replicating non-coding RNA (ncrna)
RNA Genomics II BME 110: CompBio Tools Todd Lowe & Andrew Uzilov May 17, 2011 1 TIME Why RNA? An evolutionary perspective The RNA World hypotheses: life arose as self-replicating non-coding RNA (ncrna)
Genome annotation. Erwin Datema (2011) Sandra Smit (2012, 2013)
 Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
Small Genome Annotation and Data Management at TIGR
 Small Genome Annotation and Data Management at TIGR Michelle Gwinn, William Nelson, Robert Dodson, Steven Salzberg, Owen White Abstract TIGR has developed, and continues to refine, a comprehensive, efficient
Small Genome Annotation and Data Management at TIGR Michelle Gwinn, William Nelson, Robert Dodson, Steven Salzberg, Owen White Abstract TIGR has developed, and continues to refine, a comprehensive, efficient
Gene-centered resources at NCBI
 COURSE OF BIOINFORMATICS a.a. 2014-2015 Gene-centered resources at NCBI We searched Accession Number: M60495 AT NCBI Nucleotide Gene has been implemented at NCBI to organize information about genes, serving
COURSE OF BIOINFORMATICS a.a. 2014-2015 Gene-centered resources at NCBI We searched Accession Number: M60495 AT NCBI Nucleotide Gene has been implemented at NCBI to organize information about genes, serving
Gene Identification in silico
 Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Glossary of Commonly used Annotation Terms
 Glossary of Commonly used Annotation Terms Akela a general use server for the annotation group as well as other groups throughout TIGR. Annotation Notebook a link from the gene list page that is associated
Glossary of Commonly used Annotation Terms Akela a general use server for the annotation group as well as other groups throughout TIGR. Annotation Notebook a link from the gene list page that is associated
RNA-seq Data Analysis
 Lecture 3. Clustering; Function/Pathway Enrichment analysis RNA-seq Data Analysis Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Lecture 1. Map RNA-seq read to genome Lecture
Lecture 3. Clustering; Function/Pathway Enrichment analysis RNA-seq Data Analysis Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Lecture 1. Map RNA-seq read to genome Lecture
 Genome Annotation Genome annotation What is the function of each part of the genome? Where are the genes? What is the mrna sequence (transcription, splicing) What is the protein sequence? What does
Genome Annotation Genome annotation What is the function of each part of the genome? Where are the genes? What is the mrna sequence (transcription, splicing) What is the protein sequence? What does
Sequence Based Function Annotation. Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University
 Sequence Based Function Annotation Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Usage scenarios for sequence based function annotation Function prediction of newly cloned
Sequence Based Function Annotation Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Usage scenarios for sequence based function annotation Function prediction of newly cloned
RNA folding & ncrna discovery
 I519 Introduction to Bioinformatics RNA folding & ncrna discovery Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Non-coding RNAs and their functions RNA structures RNA folding
I519 Introduction to Bioinformatics RNA folding & ncrna discovery Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Non-coding RNAs and their functions RNA structures RNA folding
Genes and gene finding
 Genes and gene finding Ben Langmead Department of Computer Science You are free to use these slides. If you do, please sign the guestbook (www.langmead-lab.org/teaching-materials), or email me (ben.langmead@gmail.com)
Genes and gene finding Ben Langmead Department of Computer Science You are free to use these slides. If you do, please sign the guestbook (www.langmead-lab.org/teaching-materials), or email me (ben.langmead@gmail.com)
Applied Bioinformatics
 Applied Bioinformatics Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Course overview What is bioinformatics Data driven science: the creation and advancement
Applied Bioinformatics Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Course overview What is bioinformatics Data driven science: the creation and advancement
Product Applications for the Sequence Analysis Collection
 Product Applications for the Sequence Analysis Collection Pipeline Pilot Contents Introduction... 1 Pipeline Pilot and Bioinformatics... 2 Sequence Searching with Profile HMM...2 Integrating Data in a
Product Applications for the Sequence Analysis Collection Pipeline Pilot Contents Introduction... 1 Pipeline Pilot and Bioinformatics... 2 Sequence Searching with Profile HMM...2 Integrating Data in a
BIOINFORMATICS Introduction
 BIOINFORMATICS Introduction Mark Gerstein, Yale University bioinfo.mbb.yale.edu/mbb452a 1 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu What is Bioinformatics? (Molecular) Bio -informatics One idea
BIOINFORMATICS Introduction Mark Gerstein, Yale University bioinfo.mbb.yale.edu/mbb452a 1 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu What is Bioinformatics? (Molecular) Bio -informatics One idea
Protein Sequence Analysis. BME 110: CompBio Tools Todd Lowe April 19, 2007 (Slide Presentation: Carol Rohl)
 Protein Sequence Analysis BME 110: CompBio Tools Todd Lowe April 19, 2007 (Slide Presentation: Carol Rohl) Linear Sequence Analysis What can you learn from a (single) protein sequence? Calculate it s physical
Protein Sequence Analysis BME 110: CompBio Tools Todd Lowe April 19, 2007 (Slide Presentation: Carol Rohl) Linear Sequence Analysis What can you learn from a (single) protein sequence? Calculate it s physical
Post-assembly Data Analysis
 Assembled transcriptome Post-assembly Data Analysis Quantification: the expression level of each gene in each sample DE genes: genes differentially expressed between samples Clustering/network analysis
Assembled transcriptome Post-assembly Data Analysis Quantification: the expression level of each gene in each sample DE genes: genes differentially expressed between samples Clustering/network analysis
Introduction to Bioinformatics CPSC 265. What is bioinformatics? Textbooks
 Introduction to Bioinformatics CPSC 265 Thanks to Jonathan Pevsner, Ph.D. Textbooks Johnathan Pevsner, who I stole most of these slides from (thanks!) has written a textbook, Bioinformatics and Functional
Introduction to Bioinformatics CPSC 265 Thanks to Jonathan Pevsner, Ph.D. Textbooks Johnathan Pevsner, who I stole most of these slides from (thanks!) has written a textbook, Bioinformatics and Functional
Gene Signal Estimates from Exon Arrays
 Gene Signal Estimates from Exon Arrays I. Introduction: With exon arrays like the GeneChip Human Exon 1.0 ST Array, researchers can examine the transcriptional profile of an entire gene (Figure 1). Being
Gene Signal Estimates from Exon Arrays I. Introduction: With exon arrays like the GeneChip Human Exon 1.0 ST Array, researchers can examine the transcriptional profile of an entire gene (Figure 1). Being
Files for this Tutorial: All files needed for this tutorial are compressed into a single archive: [BLAST_Intro.tar.gz]
![Files for this Tutorial: All files needed for this tutorial are compressed into a single archive: [BLAST_Intro.tar.gz] Files for this Tutorial: All files needed for this tutorial are compressed into a single archive: [BLAST_Intro.tar.gz]](/thumbs/75/71877651.jpg) BLAST Exercise: Detecting and Interpreting Genetic Homology Adapted by W. Leung and SCR Elgin from Detecting and Interpreting Genetic Homology by Dr. J. Buhler Prequisites: None Resources: The BLAST web
BLAST Exercise: Detecting and Interpreting Genetic Homology Adapted by W. Leung and SCR Elgin from Detecting and Interpreting Genetic Homology by Dr. J. Buhler Prequisites: None Resources: The BLAST web
Computational gene finding. Devika Subramanian Comp 470
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) The biological context Lec 1 Lec 2 Lec 3 Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) The biological context Lec 1 Lec 2 Lec 3 Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Outline. Evolution. Adaptive convergence. Common similarity problems. Chapter 7: Similarity searches on sequence databases
 Chapter 7: Similarity searches on sequence databases All science is either physics or stamp collection. Ernest Rutherford Outline Why is similarity important BLAST Protein and DNA Interpreting BLAST Individualizing
Chapter 7: Similarity searches on sequence databases All science is either physics or stamp collection. Ernest Rutherford Outline Why is similarity important BLAST Protein and DNA Interpreting BLAST Individualizing
Efficient and Accurate Analysis of non coding RNAs with InSyBio ncrnaseq
 Efficient and Accurate Analysis of non coding RNAs with InSyBio ncrnaseq WHITE PAPER By InSyBio Ltd Aigli Korfiati Computer Engineer, MSc, PhD candidate InSyBio Product Development Manager October 2015
Efficient and Accurate Analysis of non coding RNAs with InSyBio ncrnaseq WHITE PAPER By InSyBio Ltd Aigli Korfiati Computer Engineer, MSc, PhD candidate InSyBio Product Development Manager October 2015
Comparative Bioinformatics. BSCI348S Fall 2003 Midterm 1
 BSCI348S Fall 2003 Midterm 1 Multiple Choice: select the single best answer to the question or completion of the phrase. (5 points each) 1. The field of bioinformatics a. uses biomimetic algorithms to
BSCI348S Fall 2003 Midterm 1 Multiple Choice: select the single best answer to the question or completion of the phrase. (5 points each) 1. The field of bioinformatics a. uses biomimetic algorithms to
Last Update: 12/31/2017. Recommended Background Tutorial: An Introduction to NCBI BLAST
 BLAST Exercise: Detecting and Interpreting Genetic Homology Adapted by T. Cordonnier, C. Shaffer, W. Leung and SCR Elgin from Detecting and Interpreting Genetic Homology by Dr. J. Buhler Recommended Background
BLAST Exercise: Detecting and Interpreting Genetic Homology Adapted by T. Cordonnier, C. Shaffer, W. Leung and SCR Elgin from Detecting and Interpreting Genetic Homology by Dr. J. Buhler Recommended Background
1 Abstract. 2 Introduction. 3 Requirements. 4 Procedure
 1 Abstract 2 Introduction This SOP describes the procedure for creating the reference genomes database used for HMP WGS read mapping. The database is comprised of all archaeal, bacterial, lower eukaryote
1 Abstract 2 Introduction This SOP describes the procedure for creating the reference genomes database used for HMP WGS read mapping. The database is comprised of all archaeal, bacterial, lower eukaryote
Computational aspects of ncrna research. Mihaela Zavolan Biozentrum, Basel Swiss Institute of Bioinformatics
 Computational aspects of ncrna research Mihaela Zavolan Biozentrum, Basel Swiss Institute of Bioinformatics Computational aspects on ncrna Bacterial ncrnas research Gene discovery Target discovery Discovery
Computational aspects of ncrna research Mihaela Zavolan Biozentrum, Basel Swiss Institute of Bioinformatics Computational aspects on ncrna Bacterial ncrnas research Gene discovery Target discovery Discovery
Outline. Introduction to ab initio and evidence-based gene finding. Prokaryotic gene predictions
 Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
Chimp Sequence Annotation: Region 2_3
 Chimp Sequence Annotation: Region 2_3 Jeff Howenstein March 30, 2007 BIO434W Genomics 1 Introduction We received region 2_3 of the ChimpChunk sequence, and the first step we performed was to run RepeatMasker
Chimp Sequence Annotation: Region 2_3 Jeff Howenstein March 30, 2007 BIO434W Genomics 1 Introduction We received region 2_3 of the ChimpChunk sequence, and the first step we performed was to run RepeatMasker
Bioinformatic analysis of phage AB3, a phikmv-like virus infecting Acinetobacter baumannii
 Bioinformatic analysis of phage AB3, a phikmv-like virus infecting Acinetobacter baumannii J. Zhang 1 *, X. Liu 1 * and X.-J. Li 2 1 Department of Geriatrics Medicine, The Third People s Hospital of Chongqing,
Bioinformatic analysis of phage AB3, a phikmv-like virus infecting Acinetobacter baumannii J. Zhang 1 *, X. Liu 1 * and X.-J. Li 2 1 Department of Geriatrics Medicine, The Third People s Hospital of Chongqing,
Typically, to be biologically related means to share a common ancestor. In biology, we call this homologous
 Typically, to be biologically related means to share a common ancestor. In biology, we call this homologous. Two proteins sharing a common ancestor are said to be homologs. Homologyoften implies structural
Typically, to be biologically related means to share a common ancestor. In biology, we call this homologous. Two proteins sharing a common ancestor are said to be homologs. Homologyoften implies structural
Basic Bioinformatics: Homology, Sequence Alignment,
 Basic Bioinformatics: Homology, Sequence Alignment, and BLAST William S. Sanders Institute for Genomics, Biocomputing, and Biotechnology (IGBB) High Performance Computing Collaboratory (HPC 2 ) Mississippi
Basic Bioinformatics: Homology, Sequence Alignment, and BLAST William S. Sanders Institute for Genomics, Biocomputing, and Biotechnology (IGBB) High Performance Computing Collaboratory (HPC 2 ) Mississippi
Bioinformatics for Proteomics. Ann Loraine
 Bioinformatics for Proteomics Ann Loraine aloraine@uab.edu What is bioinformatics? The science of collecting, processing, organizing, storing, analyzing, and mining biological information, especially data
Bioinformatics for Proteomics Ann Loraine aloraine@uab.edu What is bioinformatics? The science of collecting, processing, organizing, storing, analyzing, and mining biological information, especially data
Mapping strategies for sequence reads
 Mapping strategies for sequence reads Ernest Turro University of Cambridge 21 Oct 2013 Quantification A basic aim in genomics is working out the contents of a biological sample. 1. What distinct elements
Mapping strategies for sequence reads Ernest Turro University of Cambridge 21 Oct 2013 Quantification A basic aim in genomics is working out the contents of a biological sample. 1. What distinct elements
Why Use BLAST? David Form - August 15,
 Wolbachia Workshop 2017 Bioinformatics BLAST Basic Local Alignment Search Tool Finding Model Organisms for Study of Disease Can yeast be used as a model organism to study cystic fibrosis? BLAST Why Use
Wolbachia Workshop 2017 Bioinformatics BLAST Basic Local Alignment Search Tool Finding Model Organisms for Study of Disease Can yeast be used as a model organism to study cystic fibrosis? BLAST Why Use
Agenda. Web Databases for Drosophila. Gene annotation workflow. GEP Drosophila annotation projects 01/01/2018. Annotation adding labels to a sequence
 Agenda GEP annotation project overview Web Databases for Drosophila An introduction to web tools, databases and NCBI BLAST Web databases for Drosophila annotation UCSC Genome Browser NCBI / BLAST FlyBase
Agenda GEP annotation project overview Web Databases for Drosophila An introduction to web tools, databases and NCBI BLAST Web databases for Drosophila annotation UCSC Genome Browser NCBI / BLAST FlyBase
MicroSEQ Rapid Microbial Identification System
 MicroSEQ Rapid Microbial Identification System Giving you complete control over microbial identification using the gold-standard genotypic method The MicroSEQ ID microbial identification system, based
MicroSEQ Rapid Microbial Identification System Giving you complete control over microbial identification using the gold-standard genotypic method The MicroSEQ ID microbial identification system, based
BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers
 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html UCSC
BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html UCSC
Genomics and Transcriptomics of Spirodela polyrhiza
 Genomics and Transcriptomics of Spirodela polyrhiza Doug Bryant Bioinformatics Core Facility & Todd Mockler Group, Donald Danforth Plant Science Center Desired Outcomes High-quality genomic reference sequence
Genomics and Transcriptomics of Spirodela polyrhiza Doug Bryant Bioinformatics Core Facility & Todd Mockler Group, Donald Danforth Plant Science Center Desired Outcomes High-quality genomic reference sequence
Bioinformatic tools for metagenomic data analysis
 Bioinformatic tools for metagenomic data analysis MEGAN - blast-based tool for exploring taxonomic content MG-RAST (SEED, FIG) - rapid annotation of metagenomic data, phylogenetic classification and metabolic
Bioinformatic tools for metagenomic data analysis MEGAN - blast-based tool for exploring taxonomic content MG-RAST (SEED, FIG) - rapid annotation of metagenomic data, phylogenetic classification and metabolic
The String Alignment Problem. Comparative Sequence Sizes. The String Alignment Problem. The String Alignment Problem.
 Dec-82 Oct-84 Aug-86 Jun-88 Apr-90 Feb-92 Nov-93 Sep-95 Jul-97 May-99 Mar-01 Jan-03 Nov-04 Sep-06 Jul-08 May-10 Mar-12 Growth of GenBank 160,000,000,000 180,000,000 Introduction to Bioinformatics Iosif
Dec-82 Oct-84 Aug-86 Jun-88 Apr-90 Feb-92 Nov-93 Sep-95 Jul-97 May-99 Mar-01 Jan-03 Nov-04 Sep-06 Jul-08 May-10 Mar-12 Growth of GenBank 160,000,000,000 180,000,000 Introduction to Bioinformatics Iosif
a-dB. Code assigned:
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
FACULTY OF BIOCHEMISTRY AND MOLECULAR MEDICINE
 FACULTY OF BIOCHEMISTRY AND MOLECULAR MEDICINE BIOMOLECULES COURSE: COMPUTER PRACTICAL 1 Author of the exercise: Prof. Lloyd Ruddock Edited by Dr. Leila Tajedin 2017-2018 Assistant: Leila Tajedin (leila.tajedin@oulu.fi)
FACULTY OF BIOCHEMISTRY AND MOLECULAR MEDICINE BIOMOLECULES COURSE: COMPUTER PRACTICAL 1 Author of the exercise: Prof. Lloyd Ruddock Edited by Dr. Leila Tajedin 2017-2018 Assistant: Leila Tajedin (leila.tajedin@oulu.fi)
Host : Dr. Nobuyuki Nukina Tutor : Dr. Fumitaka Oyama
 Method to assign the coding regions of ESTs Céline Becquet Summer Program 2002 Structural Neuropathology Lab Molecular Neuropathology Group RIKEN Brain Science Institute Host : Dr. Nobuyuki Nukina Tutor
Method to assign the coding regions of ESTs Céline Becquet Summer Program 2002 Structural Neuropathology Lab Molecular Neuropathology Group RIKEN Brain Science Institute Host : Dr. Nobuyuki Nukina Tutor
ABSTRACT METHODS FOR MICROBIAL GENOMICS. Professor Steven L. Salzberg Department of Computer Science
 ABSTRACT Title of Dissertation: COMPARATIVE AND COMPUTATIONAL METHODS FOR MICROBIAL GENOMICS Derrick Edward Wood, Doctor of Philosophy, 2014 Directed by: Professor Steven L. Salzberg Department of Computer
ABSTRACT Title of Dissertation: COMPARATIVE AND COMPUTATIONAL METHODS FOR MICROBIAL GENOMICS Derrick Edward Wood, Doctor of Philosophy, 2014 Directed by: Professor Steven L. Salzberg Department of Computer
Reference genomes and common file formats
 Reference genomes and common file formats Overview Reference genomes and GRC Fasta and FastQ (unaligned sequences) SAM/BAM (aligned sequences) Summarized genomic features BED (genomic intervals) GFF/GTF
Reference genomes and common file formats Overview Reference genomes and GRC Fasta and FastQ (unaligned sequences) SAM/BAM (aligned sequences) Summarized genomic features BED (genomic intervals) GFF/GTF
RNA-Seq Software, Tools, and Workflows
 RNA-Seq Software, Tools, and Workflows Monica Britton, Ph.D. Sr. Bioinformatics Analyst September 1, 2016 Some mrna-seq Applications Differential gene expression analysis Transcriptional profiling Assumption:
RNA-Seq Software, Tools, and Workflows Monica Britton, Ph.D. Sr. Bioinformatics Analyst September 1, 2016 Some mrna-seq Applications Differential gene expression analysis Transcriptional profiling Assumption:
RNA Secondary Structure Prediction Computational Genomics Seyoung Kim
 RNA Secondary Structure Prediction 02-710 Computational Genomics Seyoung Kim Outline RNA folding Dynamic programming for RNA secondary structure prediction Covariance model for RNA structure prediction
RNA Secondary Structure Prediction 02-710 Computational Genomics Seyoung Kim Outline RNA folding Dynamic programming for RNA secondary structure prediction Covariance model for RNA structure prediction
Regulation of eukaryotic transcription:
 Promoter definition by mass genome annotation data: in silico primer extension EMBNET course Bioinformatics of transcriptional regulation Jan 28 2008 Christoph Schmid Regulation of eukaryotic transcription:
Promoter definition by mass genome annotation data: in silico primer extension EMBNET course Bioinformatics of transcriptional regulation Jan 28 2008 Christoph Schmid Regulation of eukaryotic transcription:
NCBI web resources I: databases and Entrez
 NCBI web resources I: databases and Entrez Yanbin Yin Most materials are downloaded from ftp://ftp.ncbi.nih.gov/pub/education/ 1 Homework assignment 1 Two parts: Extract the gene IDs reported in table
NCBI web resources I: databases and Entrez Yanbin Yin Most materials are downloaded from ftp://ftp.ncbi.nih.gov/pub/education/ 1 Homework assignment 1 Two parts: Extract the gene IDs reported in table
BLAST. compared with database sequences Sequences with many matches to high- scoring words are used for final alignments
 BLAST 100 times faster than dynamic programming. Good for database searches. Derive a list of words of length w from query (e.g., 3 for protein, 11 for DNA) High-scoring words are compared with database
BLAST 100 times faster than dynamic programming. Good for database searches. Derive a list of words of length w from query (e.g., 3 for protein, 11 for DNA) High-scoring words are compared with database
Analysis of Microarray Data
 Analysis of Microarray Data Lecture 3: Visualization and Functional Analysis George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Review
Analysis of Microarray Data Lecture 3: Visualization and Functional Analysis George Bell, Ph.D. Senior Bioinformatics Scientist Bioinformatics and Research Computing Whitehead Institute Outline Review
Genome Annotation. What Does Annotation Describe??? Genome duplications Genes Mobile genetic elements Small repeats Genetic diversity
 Genome Annotation Genome Sequencing Costliest aspect of sequencing the genome o But Devoid of content Genome must be annotated o Annotation definition Analyzing the raw sequence of a genome and describing
Genome Annotation Genome Sequencing Costliest aspect of sequencing the genome o But Devoid of content Genome must be annotated o Annotation definition Analyzing the raw sequence of a genome and describing
Bionano Access 1.1 Software User Guide
 Bionano Access 1.1 Software User Guide Document Number: 30142 Document Revision: B For Research Use Only. Not for use in diagnostic procedures. Copyright 2017 Bionano Genomics, Inc. All Rights Reserved.
Bionano Access 1.1 Software User Guide Document Number: 30142 Document Revision: B For Research Use Only. Not for use in diagnostic procedures. Copyright 2017 Bionano Genomics, Inc. All Rights Reserved.
Optimization of RNAi Targets on the Human Transcriptome Ahmet Arslan Kurdoglu Computational Biosciences Program Arizona State University
 Optimization of RNAi Targets on the Human Transcriptome Ahmet Arslan Kurdoglu Computational Biosciences Program Arizona State University my background Undergraduate Degree computer systems engineer (ASU
Optimization of RNAi Targets on the Human Transcriptome Ahmet Arslan Kurdoglu Computational Biosciences Program Arizona State University my background Undergraduate Degree computer systems engineer (ASU
Genome and DNA Sequence Databases. BME 110: CompBio Tools Todd Lowe April 5, 2007
 Genome and DNA Sequence Databases BME 110: CompBio Tools Todd Lowe April 5, 2007 Admin Reading: Chapters 2 & 3 Notes available in PDF format on-line (see class calendar page): http://www.soe.ucsc.edu/classes/bme110/spring07/bme110-calendar.html
Genome and DNA Sequence Databases BME 110: CompBio Tools Todd Lowe April 5, 2007 Admin Reading: Chapters 2 & 3 Notes available in PDF format on-line (see class calendar page): http://www.soe.ucsc.edu/classes/bme110/spring07/bme110-calendar.html
Ensembl Tools. EBI is an Outstation of the European Molecular Biology Laboratory.
 Ensembl Tools EBI is an Outstation of the European Molecular Biology Laboratory. Questions? We ve muted all the mics Ask questions in the Chat box in the webinar interface I will check the Chat box periodically
Ensembl Tools EBI is an Outstation of the European Molecular Biology Laboratory. Questions? We ve muted all the mics Ask questions in the Chat box in the webinar interface I will check the Chat box periodically
Ab initio gene identification in metagenomic sequences
 Nucleic Acids Research Advance Access published April 19, 2010 Nucleic Acids Research, 2010, 1 15 doi:10.1093/nar/gkq275 Ab initio gene identification in metagenomic sequences Wenhan Zhu 1, Alexandre Lomsadze
Nucleic Acids Research Advance Access published April 19, 2010 Nucleic Acids Research, 2010, 1 15 doi:10.1093/nar/gkq275 Ab initio gene identification in metagenomic sequences Wenhan Zhu 1, Alexandre Lomsadze
axe Documentation Release g6d4d1b6-dirty Kevin Murray
 axe Documentation Release 0.3.2-5-g6d4d1b6-dirty Kevin Murray Jul 17, 2017 Contents 1 Axe Usage 3 1.1 Inputs and Outputs..................................... 4 1.2 The barcode file......................................
axe Documentation Release 0.3.2-5-g6d4d1b6-dirty Kevin Murray Jul 17, 2017 Contents 1 Axe Usage 3 1.1 Inputs and Outputs..................................... 4 1.2 The barcode file......................................
MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE?
 MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE? Lesson Plan: Title Introduction to the Genome Browser: what is a gene? JOYCE STAMM Objectives Demonstrate basic skills in using the UCSC Genome
MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE? Lesson Plan: Title Introduction to the Genome Browser: what is a gene? JOYCE STAMM Objectives Demonstrate basic skills in using the UCSC Genome
Variant calling in NGS experiments
 Variant calling in NGS experiments Jorge Jiménez jjimeneza@cipf.es BIER CIBERER Genomics Department Centro de Investigacion Principe Felipe (CIPF) (Valencia, Spain) 1 Index 1. NGS workflow 2. Variant calling
Variant calling in NGS experiments Jorge Jiménez jjimeneza@cipf.es BIER CIBERER Genomics Department Centro de Investigacion Principe Felipe (CIPF) (Valencia, Spain) 1 Index 1. NGS workflow 2. Variant calling
RNA-Seq with the Tuxedo Suite
 RNA-Seq with the Tuxedo Suite Monica Britton, Ph.D. Sr. Bioinformatics Analyst September 2015 Workshop The Basic Tuxedo Suite References Trapnell C, et al. 2009 TopHat: discovering splice junctions with
RNA-Seq with the Tuxedo Suite Monica Britton, Ph.D. Sr. Bioinformatics Analyst September 2015 Workshop The Basic Tuxedo Suite References Trapnell C, et al. 2009 TopHat: discovering splice junctions with
SVMerge Output File Format Specification Sheet
 SVMerge Output File Format Specification Sheet Document Number: 30165 Document Revision: C For Research Use Only. Not for use in diagnostic procedures. Copyright 2017 Bionano Genomics, Inc. All Rights
SVMerge Output File Format Specification Sheet Document Number: 30165 Document Revision: C For Research Use Only. Not for use in diagnostic procedures. Copyright 2017 Bionano Genomics, Inc. All Rights
SAMPLE LITERATURE Please refer to included weblink for correct version.
 Edvo-Kit #340 DNA Informatics Experiment Objective: In this experiment, students will explore the popular bioninformatics tool BLAST. First they will read sequences from autoradiographs of automated gel
Edvo-Kit #340 DNA Informatics Experiment Objective: In this experiment, students will explore the popular bioninformatics tool BLAST. First they will read sequences from autoradiographs of automated gel
Genomics AGRY Michael Gribskov Hock 331
 Genomics AGRY 60000 Michael Gribskov gribskov@purdue.edu Hock 331 Computing Essentials Resources In this course we will assemble and annotate both genomic and transcriptomic sequence assemblies We will
Genomics AGRY 60000 Michael Gribskov gribskov@purdue.edu Hock 331 Computing Essentials Resources In this course we will assemble and annotate both genomic and transcriptomic sequence assemblies We will
The use of bioinformatic analysis in support of HGT from plants to microorganisms. Meeting with applicants Parma, 26 November 2015
 The use of bioinformatic analysis in support of HGT from plants to microorganisms Meeting with applicants Parma, 26 November 2015 WHY WE NEED TO CONSIDER HGT IN GM PLANT RA Directive 2001/18/EC As general
The use of bioinformatic analysis in support of HGT from plants to microorganisms Meeting with applicants Parma, 26 November 2015 WHY WE NEED TO CONSIDER HGT IN GM PLANT RA Directive 2001/18/EC As general
Types of Databases - By Scope
 Biological Databases Bioinformatics Workshop 2009 Chi-Cheng Lin, Ph.D. Department of Computer Science Winona State University clin@winona.edu Biological Databases Data Domains - By Scope - By Level of
Biological Databases Bioinformatics Workshop 2009 Chi-Cheng Lin, Ph.D. Department of Computer Science Winona State University clin@winona.edu Biological Databases Data Domains - By Scope - By Level of
RNA secondary structure prediction and analysis
 RNA secondary structure prediction and analysis 1 Resources Lecture Notes from previous years: Takis Benos Covariance algorithm: Eddy and Durbin, Nucleic Acids Research, v22: 11, 2079 Useful lecture slides
RNA secondary structure prediction and analysis 1 Resources Lecture Notes from previous years: Takis Benos Covariance algorithm: Eddy and Durbin, Nucleic Acids Research, v22: 11, 2079 Useful lecture slides
Ab Initio SERVER PROTOTYPE FOR PREDICTION OF PHOSPHORYLATION SITES IN PROTEINS*
 COMPUTATIONAL METHODS IN SCIENCE AND TECHNOLOGY 9(1-2) 93-100 (2003/2004) Ab Initio SERVER PROTOTYPE FOR PREDICTION OF PHOSPHORYLATION SITES IN PROTEINS* DARIUSZ PLEWCZYNSKI AND LESZEK RYCHLEWSKI BiolnfoBank
COMPUTATIONAL METHODS IN SCIENCE AND TECHNOLOGY 9(1-2) 93-100 (2003/2004) Ab Initio SERVER PROTOTYPE FOR PREDICTION OF PHOSPHORYLATION SITES IN PROTEINS* DARIUSZ PLEWCZYNSKI AND LESZEK RYCHLEWSKI BiolnfoBank
Gene Structure & Gene Finding Part II
 Gene Structure & Gene Finding Part II David Wishart david.wishart@ualberta.ca 30,000 metabolite Gene Finding in Eukaryotes Eukaryotes Complex gene structure Large genomes (0.1 to 10 billion bp) Exons and
Gene Structure & Gene Finding Part II David Wishart david.wishart@ualberta.ca 30,000 metabolite Gene Finding in Eukaryotes Eukaryotes Complex gene structure Large genomes (0.1 to 10 billion bp) Exons and
Complete Genome Sequence of Pathogenic Bacterium
 JB Accepts, published online ahead of print on 25 March 2011 J. Bacteriol. doi:10.1128/jb.00301-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
JB Accepts, published online ahead of print on 25 March 2011 J. Bacteriol. doi:10.1128/jb.00301-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
ORTHOMINE - A dataset of Drosophila core promoters and its analysis. Sumit Middha Advisor: Dr. Peter Cherbas
 ORTHOMINE - A dataset of Drosophila core promoters and its analysis Sumit Middha Advisor: Dr. Peter Cherbas Introduction Challenges and Motivation D melanogaster Promoter Dataset Expanding promoter sequences
ORTHOMINE - A dataset of Drosophila core promoters and its analysis Sumit Middha Advisor: Dr. Peter Cherbas Introduction Challenges and Motivation D melanogaster Promoter Dataset Expanding promoter sequences
Sequence Annotation & Designing Gene-specific qpcr Primers (computational)
 James Madison University From the SelectedWorks of Ray Enke Ph.D. Fall October 31, 2016 Sequence Annotation & Designing Gene-specific qpcr Primers (computational) Raymond A Enke This work is licensed under
James Madison University From the SelectedWorks of Ray Enke Ph.D. Fall October 31, 2016 Sequence Annotation & Designing Gene-specific qpcr Primers (computational) Raymond A Enke This work is licensed under
PRESENTING SEQUENCES 5 GAATGCGGCTTAGACTGGTACGATGGAAC 3 3 CTTACGCCGAATCTGACCATGCTACCTTG 5
 Molecular Biology-2017 1 PRESENTING SEQUENCES As you know, sequences may either be double stranded or single stranded and have a polarity described as 5 and 3. The 5 end always contains a free phosphate
Molecular Biology-2017 1 PRESENTING SEQUENCES As you know, sequences may either be double stranded or single stranded and have a polarity described as 5 and 3. The 5 end always contains a free phosphate
Introduction to Bioinformatics
 Introduction to Bioinformatics Dortmund, 16.-20.07.2007 Lectures: Sven Rahmann Exercises: Udo Feldkamp, Michael Wurst 1 Goals of this course Learn about Software tools Databases Methods (Algorithms) in
Introduction to Bioinformatics Dortmund, 16.-20.07.2007 Lectures: Sven Rahmann Exercises: Udo Feldkamp, Michael Wurst 1 Goals of this course Learn about Software tools Databases Methods (Algorithms) in
Bundle 5 Test Review
 Bundle 5 Test Review DNA vs. RNA DNA Replication Gene Mutations- Protein Synthesis 1. Label the different components and complete the complimentary base pairing. What is this molecule called? _Nucleic
Bundle 5 Test Review DNA vs. RNA DNA Replication Gene Mutations- Protein Synthesis 1. Label the different components and complete the complimentary base pairing. What is this molecule called? _Nucleic
The modified RNAfold program was used with molecule specific and molecule independent
 Supplemental Data The modified RNAfold program was used with molecule specific and molecule independent hairpin loop statistical potentials to predict the secondary structure for sixteen RNA molecular
Supplemental Data The modified RNAfold program was used with molecule specific and molecule independent hairpin loop statistical potentials to predict the secondary structure for sixteen RNA molecular
European Union Reference Laboratory for Genetically Modified Food and Feed (EURL GMFF)
 Guideline for the submission of DNA sequences derived from genetically modified organisms and associated annotations within the framework of Directive 2001/18/EC and Regulation (EC) No 1829/2003 European
Guideline for the submission of DNA sequences derived from genetically modified organisms and associated annotations within the framework of Directive 2001/18/EC and Regulation (EC) No 1829/2003 European
DNA makes RNA makes Proteins. The Central Dogma
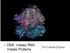 DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
Sequence Databases and database scanning
 Sequence Databases and database scanning Marjolein Thunnissen Lund, 2012 Types of databases: Primary sequence databases (proteins and nucleic acids). Composite protein sequence databases. Secondary databases.
Sequence Databases and database scanning Marjolein Thunnissen Lund, 2012 Types of databases: Primary sequence databases (proteins and nucleic acids). Composite protein sequence databases. Secondary databases.
Functional Genomics Research Stream. Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update
 Functional Genomics Research Stream Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update Updates Alternate Lab Meeting Fridays 11:30-1:00 WEL 4.224 Welcome to attend either one Lab Log thanks
Functional Genomics Research Stream Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update Updates Alternate Lab Meeting Fridays 11:30-1:00 WEL 4.224 Welcome to attend either one Lab Log thanks
Whole Transcriptome Analysis of Illumina RNA- Seq Data. Ryan Peters Field Application Specialist
 Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Complete Genome Sequence of the Polycyclic Aromatic Hydrocarbon-Degrading. Bacterium Alteromonas sp. Strain SN2
 JB Accepts, published online ahead of print on 24 June 2011 J. Bacteriol. doi:10.1128/jb.05252-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
JB Accepts, published online ahead of print on 24 June 2011 J. Bacteriol. doi:10.1128/jb.05252-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
Post-assembly Data Analysis
 Assembled transcriptome Post-assembly Data Analysis Quantification: get expression for each gene in each sample Genes differentially expressed between samples Clustering/network analysis Identifying over-represented
Assembled transcriptome Post-assembly Data Analysis Quantification: get expression for each gene in each sample Genes differentially expressed between samples Clustering/network analysis Identifying over-represented
Genscan. The Genscan HMM model Training Genscan Validating Genscan. (c) Devika Subramanian,
 Genscan The Genscan HMM model Training Genscan Validating Genscan (c) Devika Subramanian, 2009 96 Gene structure assumed by Genscan donor site acceptor site (c) Devika Subramanian, 2009 97 A simple model
Genscan The Genscan HMM model Training Genscan Validating Genscan (c) Devika Subramanian, 2009 96 Gene structure assumed by Genscan donor site acceptor site (c) Devika Subramanian, 2009 97 A simple model
Tutorial. Whole Metagenome Functional Analysis (beta) Sample to Insight. November 21, 2017
 Whole Metagenome Functional Analysis (beta) November 21, 2017 Sample to Insight QIAGEN Aarhus Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.qiagenbioinformatics.com AdvancedGenomicsSupport@qiagen.com
Whole Metagenome Functional Analysis (beta) November 21, 2017 Sample to Insight QIAGEN Aarhus Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.qiagenbioinformatics.com AdvancedGenomicsSupport@qiagen.com
CHAPTER 21 LECTURE SLIDES
 CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
Protein Bioinformatics Part I: Access to information
 Protein Bioinformatics Part I: Access to information 260.655 April 6, 2006 Jonathan Pevsner, Ph.D. pevsner@kennedykrieger.org Outline [1] Proteins at NCBI RefSeq accession numbers Cn3D to visualize structures
Protein Bioinformatics Part I: Access to information 260.655 April 6, 2006 Jonathan Pevsner, Ph.D. pevsner@kennedykrieger.org Outline [1] Proteins at NCBI RefSeq accession numbers Cn3D to visualize structures
Assignment 9: Genetic Variation
 Assignment 9: Genetic Variation Due Date: Friday, March 30 th, 2018, 10 am In this assignment, you will profile genome variation information and attempt to answer biologically relevant questions. The variant
Assignment 9: Genetic Variation Due Date: Friday, March 30 th, 2018, 10 am In this assignment, you will profile genome variation information and attempt to answer biologically relevant questions. The variant
Three-Way Comparison and Investigation of Annotated Halorhabdus utahensis Genome
 Three-Way Comparison and Investigation of Annotated Halorhabdus utahensis Genome Peter Bakke, Nick Carney, Will DeLoache, Mary Gearing, Matt Lotz, Jay McNair, Pallavi Penumetcha, Samantha Simpson, Laura
Three-Way Comparison and Investigation of Annotated Halorhabdus utahensis Genome Peter Bakke, Nick Carney, Will DeLoache, Mary Gearing, Matt Lotz, Jay McNair, Pallavi Penumetcha, Samantha Simpson, Laura
MATH 5610, Computational Biology
 MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
