The modified RNAfold program was used with molecule specific and molecule independent
|
|
|
- Bartholomew Johnathan Wilson
- 6 years ago
- Views:
Transcription
1 Supplemental Data The modified RNAfold program was used with molecule specific and molecule independent hairpin loop statistical potentials to predict the secondary structure for sixteen RNA molecular classes. The experimentally derived energetic parameters for base pair stacks and multi stem loops were used with both hairpin and internal loop statistical potentials. Molecule specific Using the molecule specific hairpin loop statistical potentials increased the prediction accuracy relative to the unmodified RNAfold program (TURNER99) for fourteen of the sixteen RNA molecular classes. The increase in accuracy for the fourteen RNA classes ranged from 2% (bacterial SRP) to 25% (HCV IRES); the TPP riboswitch showed no improvement and the SAM riboswitch showed a decrease of 1% (Fig. S1). Molecule specific internal loop statistical potentials increased prediction accuracy for all sixteen RNA molecular classes. The extent of the increase varied from 8% (TPP riboswitch) to 34% (bacterial 5S and 16S rrna) (Fig S1). With the exception of trna and U1 spliceosomal RNA, all of the RNA molecular classes had larger increases in their prediction accuracy with moleculespecific internal loop statistical potentials compared to molecule specific hairpin loop statistical potentials (Supplemental Data, see Excel file Accuracies.xlsx). 1
2 Figure S1: RNA secondary structure prediction accuracies for sixteen RNA molecular classes for RNAfold using 1) unmodified RNAfold, 2) molecule specific hairpin loop statistical potentials; 3) molecule specific internal loop statistical potentials. The experimentally derived energetic parameters for base pair stacks and multi stem loops were used with both hairpin and internal loop statistical potentials. Results for sixteen RNA molecular classes are divided into a) bacterial 5S, 16S, and 23S rrna, eukaryotic 5S and 16S rrna, trna, RNase P A and bacterial SRP; b) U1 spliceosomal RNA, hepatitis C virus internal ribosome entry site (HCV IRES), ykok leader, TPP and SAM riboswitches, iron response element (IRE), HIV type 1 dimerisation initiation site (HIV DIS) and UnaL2 Line 3 element. 2
3 Molecule independent Molecule independent hairpin loop statistical potentials increased overall prediction accuracy for twelve of the sixteen molecular classes from 1% (eukaryotic nuclear 16S rrna) to 19% (trna) (Fig. S2). Of the four molecular classes with a prediction accuracy for the statistical potential lower than RNAFold (TURNER99): bacterial SRP was 2% lower and ykok leader, TPP and SAM riboswitches were all 3% lower. The prediction accuracies with the statistical potentials for the molecule independent values were only 0 5% lower than the moleculespecific for twelve of the sixteen molecular classes. Ykok leader and bacterial 23S rrna were only 6% lower, bacterial 16S rrna 9% lower and HCV IRES was 13% less. Fifteen of the sixteen RNA molecular classes had improvements in their prediction accuracy when using the molecule independent statistical potentials. The increases ranged from 1% (eukaryotic 16S rrna and RNase P A) to 25% (bacterial 5S rrna) (Fig. S2). The ykok leader showed no change in accuracy. But the decrease in the prediction accuracy between the molecule specific vs. independent specific was much greater than for hairpin loops with decreases from 2% to 20%. Similar to the molecule specific results, all of the RNA molecular classes showed larger increases with molecule independent internal loop statistical potentials versus the hairpin loop statistical potentials except for eukaryotic 16S rrna, trna and U1 spliceosomal RNA (Supplemental Data, see Excel file Accuracies.xlsx). 3
4 Figure S2: RNA secondary structure prediction accuracies for sixteen RNA molecular classes for RNAfold using 1) unmodified RNAfold, 2) molecule independent hairpin loop statistical potentials; 3) molecule independent internal loop statistical potentials. The experimentally derived energetic parameters for base pair stacks and multi stem loops were used with both the hairpin and internal loop statistical potentials. Results for sixteen RNA molecular classes are divided into a) bacterial 5S rrna, eukaryotic 5S rrna, bacterial 16S rrna, bacterial 23S rrna, trna, eukaryotic 16S rrna, RNase P A and bacterial SRP; b) U1 spliceosomal RNA, hepatitis C virus internal ribosome entry site (HCV IRES), ykok leader, TPP and SAM riboswitches, iron response element (IRE), HIV type 1 dimerisation initiation site (HIV DIS) and UnaL2 Line 3 element. 4
5 The average standard deviations for the histogram in Figure 2 of the paper for each program for each of the sixteen training RNA molecular classes are shown as error bars in Figures S3. On average, our new statistical potentials had smaller standard deviations than all five of the competing programs (Supplemental Data, see Excel file Accuracies.xlsx). 5
6 Figure S3: RNA secondary structure prediction accuracies plus standard deviations for five RNA folding programs: RNAfold, RNAstructure (TURNER04 and (TURNER04+), ContraFold, MultiFold and RNAfold using statistical potentials. Results for sixteen RNA molecular classes are divided into a) bacterial 5S rrna, eukaryotic 5S rrna, bacterial 16S rrna, bacterial 23S rrna, trna, eukaryotic 16S rrna, RNase P A and bacterial SRP; b) U1 spliceosomal RNA, hepatitis C virus internal ribosome entry site (HCV IRES), ykok leader, TPP and SAM riboswitches, iron response element (IRE), HIV type 1 dimerisation initiation site (HIV DIS) and UnaL2 Line 3 element. 6
7 Andronescu et al. 1 followed a standard approach for training and testing; 80% of the sequences were used for training and the remaining 20% for testing. When the statistical potentials are generated and tested following this approach, the results are very similar to those seen when using the full set of sequences for generation and testing. Overall, the combined molecule independent statistical potentials still outperformed the other four programs (Figs. S4a and S4b). On average, over the sixteen RNA molecular classes, our statistical potentials scored 15% higher than RNAfold (TURNER99), 14% for RNAstructure (TURNER04), 14% higher for RNAstructure (TURNER04 Plus), 12% for CONTRAfold and 13% for MultiFold. Our statistical potentials outperformed all four programs for all sixteen RNA molecular classes with the exception of the ykok leader RNA where RNAfold (TURNER99) matched our score and RNase P A where CONTRAfold scored 2% higher. The difference in accuracy between our statistical potentials and the program with the best results for a given molecule ranged from 2% (RNase P A) to 16% (IRE) (Figs. S4a and S4b). On average, our statistical potentials outperformed the program with the best results for a given RNA molecule by 6% (Supplemental Data, see Excel file Accuracies.xlsx). 7
8 Figure S4: RNA secondary structure prediction accuracies for five RNA folding programs: RNAfold, RNAstructure (TURNER04 and (TURNER04+), ContraFold, MultiFold and RNAfold using statistical potentials. 80% of the sequences of each RNA molecular class were used for the training set of the statistical potentials. Testing results shown here are for the remaining 20%. Results for sixteen RNA molecular classes are divided into a) bacterial 5S rrna, eukaryotic 5S rrna, bacterial 16S rrna, bacterial 23S rrna, trna, eukaryotic 16S rrna, RNase P A and bacterial SRP; b) U1 spliceosomal RNA, hepatitis C virus internal ribosome entry site (HCV IRES), ykok leader, TPP and SAM riboswitches, iron response element (IRE), HIV type 1 dimerisation initiation site (HIV DIS) and UnaL2 Line 3 element. 8
9 The second method for testing our statistical potentials involved using nine control RNA molecular classes (see methods) that were not used in the generation of the statistical potentials and comparing our results against the results of the four other RNA folding programs. The hairpin statistical potentials had only small changes on most of molecular classes (Fig. S5). Seven of the nine showed changes from 3% lower (GEMM) to no change relative to the original RNAfold. RNase P B (5% lower) and R2 (11% higher) experienced larger changes. The results for internal loop statistical potentials showed slightly larger changes (Fig. S5). Four of the nine showed no changes or slight decreases up to %3. HDV, RNase P B and archaeal 16S rrna showed increases from 5% to 6%, but GEMM cis regulatory element and R2 RNA element showed large decreases of 12% and 14%, respectively (Supplemental Data, see Excel file Accuracies.xlsx). 9
10 Figure S5: RNA secondary structure prediction accuracies for nine RNA molecular classes for RNAfold using 1) unmodified RNAfold, 2) molecule independent hairpin loop statistical potentials; 3) molecule independent internal loop statistical potentials. The experimentally derived energetic parameters for base pair stacks and multi stem loops were used with both the hairpin internal loop statistical potentials. The results are for RNase P B, Hammerhead III ribozyme, purine riboswitch, hepatitis delta virus ribozyme (HDV), HIV ribosomal frameshift signal, GEMM cis regulatory element, R2 RNA element and mitochondrial and archaeal 16S rrna. 10
11 The next step for testing involved combining the hairpin and internal loop statistical potentials and testing them on the control sequences. On average, over these nine RNA molecular classes, our combined statistical potentials slightly outperformed RNAfold (TURNER99) by 2%, RNAstructure (TURNER04) by 3%, RNAstructure (TURNER04+) by 3% and CONTRAfold (.1%) (Fig. S6). MultiFold scored 1% better than our statistical potentials. Our statistical potentials predicted the RNA secondary structure better in 26 out of 45 direct comparisons and equaled in two more comparisons (Supplemental Data, see Excel file Accuracies.xlsx). 11
12 Figure S6: RNA secondary structure prediction accuracies for four RNA folding programs: RNAfold, RNAstructure (TURNER04 & TURNER04 plus newer tri and tetraloop thermodynamic parameters), CONTRAfold, MultiFold and RNAfold using statistical potentials. Results for nine RNA molecular classes: RNase P B, Hammerhead III ribozyme, purine riboswitch, hepatitis delta virus ribozyme (HDV), HIV ribosomal frameshift signal, GEMM cis regulatory element, R2 RNA element and mitochondrial and archaeal 16S rrna. 12
13 The average standard deviations for the histogram in Figure S6 of the paper for each program for each of the nine control RNA molecular classes are shown as error bars in Figure S7. On average, our new statistical potentials had a similar standard deviation as RNAstructure and slightly higher for the other three programs (Supplemental Data, see Excel file Accuracies.xlsx). 13
14 Figure S7: RNA secondary structure prediction accuracies plus standard deviations for five RNA folding programs: RNAfold, RNAstructure (TURNER04 and (TURNER04+), ContraFold, MultiFold and RNAfold using statistical potentials. The results are for RNase P B, Hammerhead III ribozyme, purine riboswitch, hepatitis delta virus ribozyme (HDV), HIV ribosomal frameshift signal, GEMM cis regulatory element, R2 RNA element and mitochondrial and archaeal 16S rrna. 14
15 Table S1: Number of sequences and their average length for eight RNA molecular classes used in determining prediction accuracy of RNA secondary structure folding programs. # of Sequences Average Length Bac 5S rrna Euk 5S rrna Bac 16S rrna Bac 23S rrna trna Euk 16S rrna RNase P A Bac SRP Table S2: Number of sequences and their average length for eight RNA molecular classes used in determining prediction accuracy of RNA secondary structure folding programs. # of Sequences Average Length U1 HCV IRES ykok TPP SAM IRE HIV DIS UnaL Table S3: Number of sequences and their average length for four RNA molecular classes used in determining prediction accuracy of RNA secondary structure folding programs. # of Sequences Average Length RNase P B Hammerhead 3 Purine HDV Table S4: Number of sequences and their average length for five RNA molecular classes used in determining prediction accuracy of RNA secondary structure folding programs. # of Sequences Average Length HIVFE GEMM R2 Mito 16S Arc 16S
16 1. Andronescu, M., Condon, A., Hoos, H. H., Mathews, D. H. & Murphy, K. P. (2010). Computational approaches for RNA energy parameter estimation. RNA 16,
RNA is a single strand molecule composed of subunits called nucleotides joined by phosphodiester bonds.
 The Versatility of RNA Primary structure of RNA RNA is a single strand molecule composed of subunits called nucleotides joined by phosphodiester bonds. Each nucleotide subunit is composed of a ribose sugar,
The Versatility of RNA Primary structure of RNA RNA is a single strand molecule composed of subunits called nucleotides joined by phosphodiester bonds. Each nucleotide subunit is composed of a ribose sugar,
90 Algorithms in Bioinformatics I, WS 06, ZBIT, D. Huson, December 4, 2006
 90 Algorithms in Bioinformatics I, WS 06, ZBIT, D. Huson, December 4, 2006 8 RNA Secondary Structure Sources for this lecture: R. Durbin, S. Eddy, A. Krogh und G. Mitchison. Biological sequence analysis,
90 Algorithms in Bioinformatics I, WS 06, ZBIT, D. Huson, December 4, 2006 8 RNA Secondary Structure Sources for this lecture: R. Durbin, S. Eddy, A. Krogh und G. Mitchison. Biological sequence analysis,
RNA secondary structure prediction and analysis
 RNA secondary structure prediction and analysis 1 Resources Lecture Notes from previous years: Takis Benos Covariance algorithm: Eddy and Durbin, Nucleic Acids Research, v22: 11, 2079 Useful lecture slides
RNA secondary structure prediction and analysis 1 Resources Lecture Notes from previous years: Takis Benos Covariance algorithm: Eddy and Durbin, Nucleic Acids Research, v22: 11, 2079 Useful lecture slides
Chapter 31. Transcription and RNA processing
 Chapter 31 Transcription and RNA processing RNA polymerase (RNAP) E. coli promoters Components of E. coli RNA Polymerase Holoenzyme (α 2 ββ'ωσ) Structure of prokaryotic RNAP The closed and open state of
Chapter 31 Transcription and RNA processing RNA polymerase (RNAP) E. coli promoters Components of E. coli RNA Polymerase Holoenzyme (α 2 ββ'ωσ) Structure of prokaryotic RNAP The closed and open state of
8/21/2014. From Gene to Protein
 From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
Review of Protein (one or more polypeptide) A polypeptide is a long chain of..
 Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Protein Synthesis. DNA to RNA to Protein
 Protein Synthesis DNA to RNA to Protein From Genes to Proteins Processing the information contained in DNA into proteins involves a sequence of events known as gene expression and results in protein synthesis.
Protein Synthesis DNA to RNA to Protein From Genes to Proteins Processing the information contained in DNA into proteins involves a sequence of events known as gene expression and results in protein synthesis.
DNA makes RNA makes Proteins. The Central Dogma
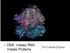 DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
RNA Genomics II. BME 110: CompBio Tools Todd Lowe & Andrew Uzilov May 17, 2011
 RNA Genomics II BME 110: CompBio Tools Todd Lowe & Andrew Uzilov May 17, 2011 1 TIME Why RNA? An evolutionary perspective The RNA World hypotheses: life arose as self-replicating non-coding RNA (ncrna)
RNA Genomics II BME 110: CompBio Tools Todd Lowe & Andrew Uzilov May 17, 2011 1 TIME Why RNA? An evolutionary perspective The RNA World hypotheses: life arose as self-replicating non-coding RNA (ncrna)
RNA Structure and the Versatility of RNA. Mitesh Shrestha
 RNA Structure and the Versatility of RNA Mitesh Shrestha Ribonucleic Acid (RNA) Nitrogenous Bases (Adenine, Uracil, Guanine, Cytosine) Ribose Sugar Ribonucleic Acid (RNA) Phosphate Group RNA world Hypothesis
RNA Structure and the Versatility of RNA Mitesh Shrestha Ribonucleic Acid (RNA) Nitrogenous Bases (Adenine, Uracil, Guanine, Cytosine) Ribose Sugar Ribonucleic Acid (RNA) Phosphate Group RNA world Hypothesis
Chapter 8 Lecture Outline. Transcription, Translation, and Bioinformatics
 Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
MOLECULAR GENETICS PROTEIN SYNTHESIS. Molecular Genetics Activity #2 page 1
 AP BIOLOGY MOLECULAR GENETICS ACTIVITY #2 NAME DATE HOUR PROTEIN SYNTHESIS Molecular Genetics Activity #2 page 1 GENETIC CODE PROTEIN SYNTHESIS OVERVIEW Molecular Genetics Activity #2 page 2 PROTEIN SYNTHESIS
AP BIOLOGY MOLECULAR GENETICS ACTIVITY #2 NAME DATE HOUR PROTEIN SYNTHESIS Molecular Genetics Activity #2 page 1 GENETIC CODE PROTEIN SYNTHESIS OVERVIEW Molecular Genetics Activity #2 page 2 PROTEIN SYNTHESIS
RNA Secondary Structure Prediction Computational Genomics Seyoung Kim
 RNA Secondary Structure Prediction 02-710 Computational Genomics Seyoung Kim Outline RNA folding Dynamic programming for RNA secondary structure prediction Covariance model for RNA structure prediction
RNA Secondary Structure Prediction 02-710 Computational Genomics Seyoung Kim Outline RNA folding Dynamic programming for RNA secondary structure prediction Covariance model for RNA structure prediction
Chapter 14 Active Reading Guide From Gene to Protein
 Name: AP Biology Mr. Croft Chapter 14 Active Reading Guide From Gene to Protein This is going to be a very long journey, but it is crucial to your understanding of biology. Work on this chapter a single
Name: AP Biology Mr. Croft Chapter 14 Active Reading Guide From Gene to Protein This is going to be a very long journey, but it is crucial to your understanding of biology. Work on this chapter a single
RNA Structure Prediction and Comparison. RNA Biology Background
 RN Structure Prediction and omparison Session 1 RN Biology Background Robert iegerich Faculty of Technology robert@techfak.ni-bielefeld.de October 13, 2013 Robert iegerich Overview of lecture topics The
RN Structure Prediction and omparison Session 1 RN Biology Background Robert iegerich Faculty of Technology robert@techfak.ni-bielefeld.de October 13, 2013 Robert iegerich Overview of lecture topics The
Graphical methods in RNA structure matching
 Purdue University Purdue e-pubs Open Access Dissertations Theses and Dissertations 8-2016 Graphical methods in RNA structure matching Jiajie Huang Purdue University Follow this and additional works at:
Purdue University Purdue e-pubs Open Access Dissertations Theses and Dissertations 8-2016 Graphical methods in RNA structure matching Jiajie Huang Purdue University Follow this and additional works at:
Types of nucleic acid
 RNA STRUCTURE 1 Types of nucleic acid DNA Deoxyribonucleic acid RNA ribonucleic acid HOCH 2 O OH HOCH 2 O OH OH OH OH (no O) ribose deoxyribose 2 Nucleic acids consist of repeating nucleotide that have
RNA STRUCTURE 1 Types of nucleic acid DNA Deoxyribonucleic acid RNA ribonucleic acid HOCH 2 O OH HOCH 2 O OH OH OH OH (no O) ribose deoxyribose 2 Nucleic acids consist of repeating nucleotide that have
Unit 6: Molecular Genetics & DNA Technology Guided Reading Questions (100 pts total)
 Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 16 The Molecular Basis of Inheritance Unit 6: Molecular Genetics
Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 16 The Molecular Basis of Inheritance Unit 6: Molecular Genetics
RNA metabolism. DNA dependent synthesis of RNA RNA processing RNA dependent synthesis of RNA and DNA.
 RNA metabolism DNA dependent synthesis of RNA RNA processing RNA dependent synthesis of RNA and DNA http://www.youtube.com/watch?v=ovc8nxobxmq DNA dependent synthesis of RNA : production of an RNA molecule
RNA metabolism DNA dependent synthesis of RNA RNA processing RNA dependent synthesis of RNA and DNA http://www.youtube.com/watch?v=ovc8nxobxmq DNA dependent synthesis of RNA : production of an RNA molecule
Computational analysis of non-coding RNA. Andrew Uzilov BME110 Tue, Nov 16, 2010
 Computational analysis of non-coding RNA Andrew Uzilov auzilov@ucsc.edu BME110 Tue, Nov 16, 2010 1 Corrected/updated talk slides are here: http://tinyurl.com/uzilovrna redirects to: http://users.soe.ucsc.edu/~auzilov/bme110/fall2010/
Computational analysis of non-coding RNA Andrew Uzilov auzilov@ucsc.edu BME110 Tue, Nov 16, 2010 1 Corrected/updated talk slides are here: http://tinyurl.com/uzilovrna redirects to: http://users.soe.ucsc.edu/~auzilov/bme110/fall2010/
Computational Biology I LSM5191 (2003/4)
 Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
Name 10 Molecular Biology of the Gene Test Date Study Guide You must know: The structure of DNA. The major steps to replication.
 Name 10 Molecular Biology of the Gene Test Date Study Guide You must know: The structure of DNA. The major steps to replication. The difference between replication, transcription, and translation. How
Name 10 Molecular Biology of the Gene Test Date Study Guide You must know: The structure of DNA. The major steps to replication. The difference between replication, transcription, and translation. How
DNA Transcription. Dr Aliwaini
 DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
2. From the first paragraph in this section, find three ways in which RNA differs from DNA.
 Name Chapter 17: From Gene to Protein Begin reading at page 328 Basic Principles of Transcription and Translation. Work on this chapter a single concept at a time, and expect to spend at least 6 hours
Name Chapter 17: From Gene to Protein Begin reading at page 328 Basic Principles of Transcription and Translation. Work on this chapter a single concept at a time, and expect to spend at least 6 hours
Chapter 17: From Gene to Protein
 Name Period This is going to be a very long journey, but it is crucial to your understanding of biology. Work on this chapter a single concept at a time, and expect to spend at least 6 hours to truly master
Name Period This is going to be a very long journey, but it is crucial to your understanding of biology. Work on this chapter a single concept at a time, and expect to spend at least 6 hours to truly master
Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype)
 Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype) Question#1: One-Gene, One-Polypeptide The figure below shows the results of feeding trials with one auxotroph strain of Neurospora
Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype) Question#1: One-Gene, One-Polypeptide The figure below shows the results of feeding trials with one auxotroph strain of Neurospora
DNA Structure & the Genome. Bio160 General Biology
 DNA Structure & the Genome Bio160 General Biology Lecture Outline I. DNA A nucleic acid II. Chromosome Structure III. Chromosomes and Genes IV. DNA vs. RNA I. DNA A Nucleic Acid Structure of DNA: Remember:
DNA Structure & the Genome Bio160 General Biology Lecture Outline I. DNA A nucleic acid II. Chromosome Structure III. Chromosomes and Genes IV. DNA vs. RNA I. DNA A Nucleic Acid Structure of DNA: Remember:
DNA and RNA. Chapter 12
 DNA and RNA Chapter 12 History of DNA Late 1800 s scientists discovered that DNA is in the nucleus of the cell 1902 Walter Sutton proposed that hereditary material resided in the chromosomes in the nucleus
DNA and RNA Chapter 12 History of DNA Late 1800 s scientists discovered that DNA is in the nucleus of the cell 1902 Walter Sutton proposed that hereditary material resided in the chromosomes in the nucleus
DNA RNA PROTEIN SYNTHESIS -NOTES-
 DNA RNA PROTEIN SYNTHESIS -NOTES- THE COMPONENTS AND STRUCTURE OF DNA DNA is made up of units called nucleotides. Nucleotides are made up of three basic components:, called deoxyribose in DNA In DNA, there
DNA RNA PROTEIN SYNTHESIS -NOTES- THE COMPONENTS AND STRUCTURE OF DNA DNA is made up of units called nucleotides. Nucleotides are made up of three basic components:, called deoxyribose in DNA In DNA, there
Themes: RNA and RNA Processing. Messenger RNA (mrna) What is a gene? RNA is very versatile! RNA-RNA interactions are very important!
 Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Transcription Eukaryotic Cells
 Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
DNA Transcription. Visualizing Transcription. The Transcription Process
 DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
Nucleic Acids: DNA and RNA
 Nucleic Acids: DNA and RNA Living organisms are complex systems. Hundreds of thousands of proteins exist inside each one of us to help carry out our daily functions. These proteins are produced locally,
Nucleic Acids: DNA and RNA Living organisms are complex systems. Hundreds of thousands of proteins exist inside each one of us to help carry out our daily functions. These proteins are produced locally,
5 -GAC-3 5 -GTC-3 5 -CAG Which of these are NOT important for RNA Polymerase interacting with DNA?
 Name This exam is schedule for 75 minutes and I anticipate it to take the full time allotted. You are free to leave if you finish. The exam is split into two sections. Part 1 is multiple choice select
Name This exam is schedule for 75 minutes and I anticipate it to take the full time allotted. You are free to leave if you finish. The exam is split into two sections. Part 1 is multiple choice select
Chapter 14: Gene Expression: From Gene to Protein
 Chapter 14: Gene Expression: From Gene to Protein This is going to be a very long journey, but it is crucial to your understanding of biology. Work on this chapter a single concept at a time, and expect
Chapter 14: Gene Expression: From Gene to Protein This is going to be a very long journey, but it is crucial to your understanding of biology. Work on this chapter a single concept at a time, and expect
Protein Synthesis Notes
 Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
DNA Structure and Replication, and Virus Structure and Replication Test Review
 DNA Structure and Replication, and Virus Structure and Replication Test Review What does DNA stand for? Deoxyribonucleic Acid DNA is what type of macromolecule? DNA is a nucleic acid The building blocks
DNA Structure and Replication, and Virus Structure and Replication Test Review What does DNA stand for? Deoxyribonucleic Acid DNA is what type of macromolecule? DNA is a nucleic acid The building blocks
CHAPTER 17 FROM GENE TO PROTEIN. Section C: The Synthesis of Protein
 CHAPTER 17 FROM GENE TO PROTEIN Section C: The Synthesis of Protein 1. Translation is the RNA-directed synthesis of a polypeptide: a closer look 2. Signal peptides target some eukaryotic polypeptides to
CHAPTER 17 FROM GENE TO PROTEIN Section C: The Synthesis of Protein 1. Translation is the RNA-directed synthesis of a polypeptide: a closer look 2. Signal peptides target some eukaryotic polypeptides to
AP BIOLOGY RNA, DNA, & Proteins Chapters 16 & 17 Review
 AP BIOLOGY RNA, DNA, & Proteins Chapters 16 & 17 Review Enzyme that adds nucleotide subunits to an RNA primer during replication DNA polymerase III Another name for protein synthesis translation Sugar
AP BIOLOGY RNA, DNA, & Proteins Chapters 16 & 17 Review Enzyme that adds nucleotide subunits to an RNA primer during replication DNA polymerase III Another name for protein synthesis translation Sugar
RNA does not adopt the classic B-DNA helix conformation when it forms a self-complementary double helix
 Reason: RNA has ribose sugar ring, with a hydroxyl group (OH) If RNA in B-from conformation there would be unfavorable steric contact between the hydroxyl group, base, and phosphate backbone. RNA structure
Reason: RNA has ribose sugar ring, with a hydroxyl group (OH) If RNA in B-from conformation there would be unfavorable steric contact between the hydroxyl group, base, and phosphate backbone. RNA structure
From Gene to Protein. Chapter 17. Biology Eighth Edition Neil Campbell and Jane Reece. PowerPoint Lecture Presentations for
 Chapter 17 From Gene to Protein PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp
Chapter 17 From Gene to Protein PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp
microrna-122 stimulates translation of Hepatitis C Virus RNA
 Supplementary information for: microrna-122 stimulates translation of Hepatitis C Virus RNA Jura Inga Henke 1,5, Dagmar Goergen 1,5, Junfeng Zheng 3, Yutong Song 4, Christian G. Schüttler 2, Carmen Fehr
Supplementary information for: microrna-122 stimulates translation of Hepatitis C Virus RNA Jura Inga Henke 1,5, Dagmar Goergen 1,5, Junfeng Zheng 3, Yutong Song 4, Christian G. Schüttler 2, Carmen Fehr
Structural Bioinformatics (C3210) DNA and RNA Structure
 Structural Bioinformatics (C3210) DNA and RNA Structure Importance of DNA/RNA 3D Structure Nucleic acids are essential materials found in all living organisms. Their main function is to maintain and transmit
Structural Bioinformatics (C3210) DNA and RNA Structure Importance of DNA/RNA 3D Structure Nucleic acids are essential materials found in all living organisms. Their main function is to maintain and transmit
From Gene to Protein transcription, messenger RNA (mrna) translation, RNA processing triplet code, template strand, codons,
 From Gene to Protein I. Transcription and translation are the two main processes linking gene to protein. A. RNA is chemically similar to DNA, except that it contains ribose as its sugar and substitutes
From Gene to Protein I. Transcription and translation are the two main processes linking gene to protein. A. RNA is chemically similar to DNA, except that it contains ribose as its sugar and substitutes
Chem 465 Biochemistry II
 Chem 465 Biochemistry II Name: 2 points Multiple choice (4 points apiece): 1. Which of the following is not true of trna molecules? A) The 3'-terminal sequence is -CCA. B) Their anticodons are complementary
Chem 465 Biochemistry II Name: 2 points Multiple choice (4 points apiece): 1. Which of the following is not true of trna molecules? A) The 3'-terminal sequence is -CCA. B) Their anticodons are complementary
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION
 M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
The Genetic Code and Transcription. Chapter 12 Honors Genetics Ms. Susan Chabot
 The Genetic Code and Transcription Chapter 12 Honors Genetics Ms. Susan Chabot TRANSCRIPTION Copy SAME language DNA to RNA Nucleic Acid to Nucleic Acid TRANSLATION Copy DIFFERENT language RNA to Amino
The Genetic Code and Transcription Chapter 12 Honors Genetics Ms. Susan Chabot TRANSCRIPTION Copy SAME language DNA to RNA Nucleic Acid to Nucleic Acid TRANSLATION Copy DIFFERENT language RNA to Amino
Name: Class: Date: ID: A
 Class: _ Date: _ CH 12 Review Multiple Choice Identify the choice that best completes the statement or answers the question. 1. How many codons are needed to specify three amino acids? a. 6 c. 3 b. 12
Class: _ Date: _ CH 12 Review Multiple Choice Identify the choice that best completes the statement or answers the question. 1. How many codons are needed to specify three amino acids? a. 6 c. 3 b. 12
Prokaryotic Transcription
 Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
Protein Synthesis: Transcription and Translation
 Protein Synthesis: Transcription and Translation Proteins In living things, proteins are in charge of the expression of our traits (hair/eye color, ability to make insulin, predisposition for cancer, etc.)
Protein Synthesis: Transcription and Translation Proteins In living things, proteins are in charge of the expression of our traits (hair/eye color, ability to make insulin, predisposition for cancer, etc.)
Independent Study Guide The Blueprint of Life, from DNA to Protein (Chapter 7)
 Independent Study Guide The Blueprint of Life, from DNA to Protein (Chapter 7) I. General Principles (Chapter 7 introduction) a. Morse code distinct series of dots and dashes encode the 26 letters of the
Independent Study Guide The Blueprint of Life, from DNA to Protein (Chapter 7) I. General Principles (Chapter 7 introduction) a. Morse code distinct series of dots and dashes encode the 26 letters of the
NUCLEIC ACIDS: DNA AND RNA. HLeeYu Jsuico Junsay Department of Chemistry School of Science and Engineering Ateneo de Manila University
 NUCLEIC ACIDS: DNA AND RNA HLeeYu Jsuico Junsay Department of Chemistry School of Science and Engineering Ateneo de Manila University 1 BUILDING BLOCKS OF NUCLEIC ACIDS 2 Nucleic Acids are important for
NUCLEIC ACIDS: DNA AND RNA HLeeYu Jsuico Junsay Department of Chemistry School of Science and Engineering Ateneo de Manila University 1 BUILDING BLOCKS OF NUCLEIC ACIDS 2 Nucleic Acids are important for
STUDY GUIDE SECTION 10-1 Discovery of DNA
 STUDY GUIDE SECTION 10-1 Discovery of DNA Name Period Date Multiple Choice-Write the correct letter in the blank. 1. The virulent strain of the bacterium S. pneumoniae causes disease because it a. has
STUDY GUIDE SECTION 10-1 Discovery of DNA Name Period Date Multiple Choice-Write the correct letter in the blank. 1. The virulent strain of the bacterium S. pneumoniae causes disease because it a. has
Bundle 5 Test Review
 Bundle 5 Test Review DNA vs. RNA DNA Replication Gene Mutations- Protein Synthesis 1. Label the different components and complete the complimentary base pairing. What is this molecule called? _Nucleic
Bundle 5 Test Review DNA vs. RNA DNA Replication Gene Mutations- Protein Synthesis 1. Label the different components and complete the complimentary base pairing. What is this molecule called? _Nucleic
Ch. 10 Notes DNA: Transcription and Translation
 Ch. 10 Notes DNA: Transcription and Translation GOALS Compare the structure of RNA with that of DNA Summarize the process of transcription Relate the role of codons to the sequence of amino acids that
Ch. 10 Notes DNA: Transcription and Translation GOALS Compare the structure of RNA with that of DNA Summarize the process of transcription Relate the role of codons to the sequence of amino acids that
Module 7: The Central Dogma. CSE590: Molecular programming and neural computation. All slides by Eric Klavins.
 Module 7: The Central Dogma CSE590: Molecular programming and neural computation. All slides by Eric Klavins. 1 The Central Dogma RNA Polymerase The Ribosome m Note: We will look mainly at prokaryo=c (e.g.
Module 7: The Central Dogma CSE590: Molecular programming and neural computation. All slides by Eric Klavins. 1 The Central Dogma RNA Polymerase The Ribosome m Note: We will look mainly at prokaryo=c (e.g.
Replication Review. 1. What is DNA Replication? 2. Where does DNA Replication take place in eukaryotic cells?
 Replication Review 1. What is DNA Replication? 2. Where does DNA Replication take place in eukaryotic cells? 3. Where does DNA Replication take place in the cell cycle? 4. 4. What guides DNA Replication?
Replication Review 1. What is DNA Replication? 2. Where does DNA Replication take place in eukaryotic cells? 3. Where does DNA Replication take place in the cell cycle? 4. 4. What guides DNA Replication?
Nucleic Acids and the RNA World. Pages Chapter 4
 Nucleic Acids and the RNA World Pages 74-89 Chapter 4 RNA vs. Protein Chemical Evolution stated that life evolved from a polymer called a protein. HOWEVER, now many scientists question this. There is currently
Nucleic Acids and the RNA World Pages 74-89 Chapter 4 RNA vs. Protein Chemical Evolution stated that life evolved from a polymer called a protein. HOWEVER, now many scientists question this. There is currently
DNA Structure and Analysis. Chapter 4: Background
 DNA Structure and Analysis Chapter 4: Background Molecular Biology Three main disciplines of biotechnology Biochemistry Genetics Molecular Biology # Biotechnology: A Laboratory Skills Course explorer.bio-rad.com
DNA Structure and Analysis Chapter 4: Background Molecular Biology Three main disciplines of biotechnology Biochemistry Genetics Molecular Biology # Biotechnology: A Laboratory Skills Course explorer.bio-rad.com
RNA synthesis/transcription I Biochemistry 302. February 6, 2004 Bob Kelm
 RNA synthesis/transcription I Biochemistry 302 February 6, 2004 Bob Kelm Overview of RNA classes Messenger RNA (mrna) Encodes protein Relatively short half-life ( 3 min in E. coli, 30 min in eukaryotic
RNA synthesis/transcription I Biochemistry 302 February 6, 2004 Bob Kelm Overview of RNA classes Messenger RNA (mrna) Encodes protein Relatively short half-life ( 3 min in E. coli, 30 min in eukaryotic
BEADLE & TATUM EXPERIMENT
 FROM DNA TO PROTEINS: gene expression Chapter 14 LECTURE OBJECTIVES What Is the Evidence that Genes Code for Proteins? How Does Information Flow from Genes to Proteins? How Is the Information Content in
FROM DNA TO PROTEINS: gene expression Chapter 14 LECTURE OBJECTIVES What Is the Evidence that Genes Code for Proteins? How Does Information Flow from Genes to Proteins? How Is the Information Content in
Chapter 1 Structure of Nucleic Acids DNA The structure of part of a DNA double helix
 Chapter 1 Structure of Nucleic Acids DNA The structure of part of a DNA double helix Deoxyribonucleic acid ) (DNA) is a nucleic acid that contains the genetic instructions used in the development and functioning
Chapter 1 Structure of Nucleic Acids DNA The structure of part of a DNA double helix Deoxyribonucleic acid ) (DNA) is a nucleic acid that contains the genetic instructions used in the development and functioning
DNA vs. RNA B-4.1. Compare DNA and RNA in terms of structure, nucleotides and base pairs.
 DNA vs. RNA B-4.1 Compare DNA and RNA in terms of structure, nucleotides and base pairs. Key Concepts l Nucleic Acids: l deoxyribonucleic acid (DNA) l ribonucleic acid (RNA) l Nucleotides: l nitrogen base,
DNA vs. RNA B-4.1 Compare DNA and RNA in terms of structure, nucleotides and base pairs. Key Concepts l Nucleic Acids: l deoxyribonucleic acid (DNA) l ribonucleic acid (RNA) l Nucleotides: l nitrogen base,
Transcription in Eukaryotes
 Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Maximizing Expected Base Pair Accuracy in RNA Secondary Structure Prediction by Joining Stochastic Context-Free Grammars Method
 www.ijcsi.org 533 Maximizing Expected Base Pair Accuracy in RNA Secondary Structure Prediction by Joining Stochastic Context-Free Grammars Method Shahira M. Habashy Faculty of Engineering, Helwan University,
www.ijcsi.org 533 Maximizing Expected Base Pair Accuracy in RNA Secondary Structure Prediction by Joining Stochastic Context-Free Grammars Method Shahira M. Habashy Faculty of Engineering, Helwan University,
PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein
 PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein This is also known as: The central dogma of molecular biology Protein Proteins are made
PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein This is also known as: The central dogma of molecular biology Protein Proteins are made
Fig Ch 17: From Gene to Protein
 Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Protein Synthesis & Gene Expression
 DNA provides the instructions for how to build proteins Each gene dictates how to build a single protein in prokaryotes The sequence of nucleotides (AGCT) in DNA dictates the order of amino acids that
DNA provides the instructions for how to build proteins Each gene dictates how to build a single protein in prokaryotes The sequence of nucleotides (AGCT) in DNA dictates the order of amino acids that
C. Incorrect! Threonine is an amino acid, not a nucleotide base.
 MCAT Biology - Problem Drill 05: RNA and Protein Biosynthesis Question No. 1 of 10 1. Which of the following bases are only found in RNA? Question #01 (A) Ribose. (B) Uracil. (C) Threonine. (D) Adenine.
MCAT Biology - Problem Drill 05: RNA and Protein Biosynthesis Question No. 1 of 10 1. Which of the following bases are only found in RNA? Question #01 (A) Ribose. (B) Uracil. (C) Threonine. (D) Adenine.
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
Assembling Protein Molecules
 How Does Dna Provide Instructions For Assembling Protein Molecules What does the information in DNA molecules provide instructions for? A. Assembling B. Assembling protein molecules into amino acids. C.
How Does Dna Provide Instructions For Assembling Protein Molecules What does the information in DNA molecules provide instructions for? A. Assembling B. Assembling protein molecules into amino acids. C.
O C. 5 th C. 3 rd C. the national health museum
 Elements of Molecular Biology Cells Cells is a basic unit of all living organisms. It stores all information to replicate itself Nucleus, chromosomes, genes, All living things are made of cells Prokaryote,
Elements of Molecular Biology Cells Cells is a basic unit of all living organisms. It stores all information to replicate itself Nucleus, chromosomes, genes, All living things are made of cells Prokaryote,
Chapter 10 - Molecular Biology of the Gene
 Bio 100 - Molecular Genetics 1 A. Bacterial Transformation Chapter 10 - Molecular Biology of the Gene Researchers found that they could transfer an inherited characteristic (e.g. the ability to cause pneumonia),
Bio 100 - Molecular Genetics 1 A. Bacterial Transformation Chapter 10 - Molecular Biology of the Gene Researchers found that they could transfer an inherited characteristic (e.g. the ability to cause pneumonia),
Pauling/Itano Experiment
 Chapter 12 Pauling/Itano Experiment Linus Pauling and Harvey Itano knew that hemoglobin, a molecule in red blood cells, contained an electrical charge. They wanted to see if the hemoglobin in normal RBC
Chapter 12 Pauling/Itano Experiment Linus Pauling and Harvey Itano knew that hemoglobin, a molecule in red blood cells, contained an electrical charge. They wanted to see if the hemoglobin in normal RBC
RNAiFold: Complete Inverse Folding for Synthetic Design
 RNAiFold: Complete Inverse Folding for Synthetic Design Juan Antonio García Martín 23/07/2015 Benasque 2015: Computational Analysis of RNA Structure and Function RNA folding Sequence + Structure Folding
RNAiFold: Complete Inverse Folding for Synthetic Design Juan Antonio García Martín 23/07/2015 Benasque 2015: Computational Analysis of RNA Structure and Function RNA folding Sequence + Structure Folding
DNA is the genetic material. DNA structure. Chapter 7: DNA Replication, Transcription & Translation; Mutations & Ames test
 DNA is the genetic material Chapter 7: DNA Replication, Transcription & Translation; Mutations & Ames test Dr. Amy Rogers Bio 139 General Microbiology Hereditary information is carried by DNA Griffith/Avery
DNA is the genetic material Chapter 7: DNA Replication, Transcription & Translation; Mutations & Ames test Dr. Amy Rogers Bio 139 General Microbiology Hereditary information is carried by DNA Griffith/Avery
RNA: Transcription and Triplet Code
 RNA: Transcription and Triplet Code There are Five Kinds of RNA, All of Which are Templated from DNA. The first type of RNA is trna. The "t" stands for "transfer". This RNA is the RNA that transfers amino
RNA: Transcription and Triplet Code There are Five Kinds of RNA, All of Which are Templated from DNA. The first type of RNA is trna. The "t" stands for "transfer". This RNA is the RNA that transfers amino
Structural motif detection in high-throughput structural probing of RNA molecules
 Structural motif detection in high-throughput structural probing of RNA molecules Diego Muñoz Medina, Sergio Pablo Sánchez Cordero December 11, 009 Abstract In recent years, the advances in structural
Structural motif detection in high-throughput structural probing of RNA molecules Diego Muñoz Medina, Sergio Pablo Sánchez Cordero December 11, 009 Abstract In recent years, the advances in structural
What is RNA? Another type of nucleic acid A working copy of DNA Does not matter if it is damaged or destroyed
 RNA Section 3.1 What is RNA? Another type of nucleic acid A working copy of DNA Does not matter if it is damaged or destroyed Used to direct the production of proteins that determines an organisms characteristics
RNA Section 3.1 What is RNA? Another type of nucleic acid A working copy of DNA Does not matter if it is damaged or destroyed Used to direct the production of proteins that determines an organisms characteristics
Bundle 6 Test Review
 Bundle 6 Test Review DNA vs. RNA DNA Replication Gene Mutations- Protein Synthesis 1. Label the different components and complete the complimentary base pairing. What is this molecule called? Deoxyribonucleic
Bundle 6 Test Review DNA vs. RNA DNA Replication Gene Mutations- Protein Synthesis 1. Label the different components and complete the complimentary base pairing. What is this molecule called? Deoxyribonucleic
Optimizing Synthetic DNA for Metabolic Engineering Applications. Howard Salis Penn State University
 Optimizing Synthetic DNA for Metabolic Engineering Applications Howard Salis Penn State University Synthetic Biology Specify a function Build a genetic system (a DNA molecule) Genetic Pseudocode call producequorumsignal(luxi
Optimizing Synthetic DNA for Metabolic Engineering Applications Howard Salis Penn State University Synthetic Biology Specify a function Build a genetic system (a DNA molecule) Genetic Pseudocode call producequorumsignal(luxi
DNA and RNA. Chapter 12
 DNA and RNA Chapter 12 Warm Up Exercise Test Corrections Make sure to indicate your new answer and provide an explanation for why this is the correct answer. Do this with a red pen in the margins of your
DNA and RNA Chapter 12 Warm Up Exercise Test Corrections Make sure to indicate your new answer and provide an explanation for why this is the correct answer. Do this with a red pen in the margins of your
Chapter 12. DNA TRANSCRIPTION and TRANSLATION
 Chapter 12 DNA TRANSCRIPTION and TRANSLATION 12-3 RNA and Protein Synthesis WARM UP What are proteins? Where do they come from? From DNA to RNA to Protein DNA in our cells carry the instructions for making
Chapter 12 DNA TRANSCRIPTION and TRANSLATION 12-3 RNA and Protein Synthesis WARM UP What are proteins? Where do they come from? From DNA to RNA to Protein DNA in our cells carry the instructions for making
TERTIARY MOTIF INTERACTIONS ON RNA STRUCTURE
 1 TERTIARY MOTIF INTERACTIONS ON RNA STRUCTURE Bioinformatics Senior Project Wasay Hussain Spring 2009 Overview of RNA 2 The central Dogma of Molecular biology is DNA RNA Proteins The RNA (Ribonucleic
1 TERTIARY MOTIF INTERACTIONS ON RNA STRUCTURE Bioinformatics Senior Project Wasay Hussain Spring 2009 Overview of RNA 2 The central Dogma of Molecular biology is DNA RNA Proteins The RNA (Ribonucleic
RNA Secondary Structure Prediction
 RNA Secondary Structure Prediction Outline 1) Introduction: RNA structure basics 2) Dynamic programming for RNA secondary structure prediction The Central Dogma of Molecular Biology DNA CCTGAGCCAACTATTGATGAA
RNA Secondary Structure Prediction Outline 1) Introduction: RNA structure basics 2) Dynamic programming for RNA secondary structure prediction The Central Dogma of Molecular Biology DNA CCTGAGCCAACTATTGATGAA
DNA. Is a molecule that encodes the genetic instructions used in the development and functioning of all known living organisms and many viruses.
 Is a molecule that encodes the genetic instructions used in the development and functioning of all known living organisms and many viruses. Genetic information is encoded as a sequence of nucleotides (guanine,
Is a molecule that encodes the genetic instructions used in the development and functioning of all known living organisms and many viruses. Genetic information is encoded as a sequence of nucleotides (guanine,
Year III Pharm.D Dr. V. Chitra
 Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Chapter 13 - Concept Mapping
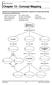 Chapter 13 - Concept Mapping Using the terms and phrases provided below, complete the concept map showing the discovery of DNA structure. amount of base pairs five-carbon sugar purine DNA polymerases Franklin
Chapter 13 - Concept Mapping Using the terms and phrases provided below, complete the concept map showing the discovery of DNA structure. amount of base pairs five-carbon sugar purine DNA polymerases Franklin
RNA Processing. Voet-Voet 1254 RNA molecules. Post-transcriptional processing. Enzymes. Most bacterial RNA molecules All eukaryotic RNA molecules
 RNA Processing Voet-Voet 1254 RNA molecules Post-transcriptional processing Most bacterial RNA molecules All eukaryotic RNA molecules Enzymes Catalytic RNA Ribozymes Catalytic protein Summary RNA processing
RNA Processing Voet-Voet 1254 RNA molecules Post-transcriptional processing Most bacterial RNA molecules All eukaryotic RNA molecules Enzymes Catalytic RNA Ribozymes Catalytic protein Summary RNA processing
Protein Synthesis. Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy
 Protein Synthesis Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy STRUCTURE OF RNA RNA, adenine forms a base pair with
Protein Synthesis Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy STRUCTURE OF RNA RNA, adenine forms a base pair with
Chapter Fundamental Molecular Genetic Mechanisms
 Chapter 5-1 - Fundamental Molecular Genetic Mechanisms 5.1 Structure of Nucleic Acids 5.2 Transcription of Protein-Coding Genes and Formation of Functional mrna 5.3 The Decoding of mrna by trnas 5.4 Stepwise
Chapter 5-1 - Fundamental Molecular Genetic Mechanisms 5.1 Structure of Nucleic Acids 5.2 Transcription of Protein-Coding Genes and Formation of Functional mrna 5.3 The Decoding of mrna by trnas 5.4 Stepwise
DNA Structure and Properties Basic Properties Predicting Melting Temperature. Dinesh Yadav
 DNA Structure and Properties Basic Properties Predicting Melting Temperature Dinesh Yadav Nucleic Acid Structure Question: Is this RNA or DNA? Molecules of Life, pp. 15 2 Nucleic Acid Bases Molecules of
DNA Structure and Properties Basic Properties Predicting Melting Temperature Dinesh Yadav Nucleic Acid Structure Question: Is this RNA or DNA? Molecules of Life, pp. 15 2 Nucleic Acid Bases Molecules of
RNA Part I: Chemical Structure of RNA
 RA Part I: Chemical Structure of RA Structural differences between RA and DA Resistance of phosphate esters to basic hydrolysis The 2 - group of RA facilitates chemical cleavage in aqueous a by forming
RA Part I: Chemical Structure of RA Structural differences between RA and DA Resistance of phosphate esters to basic hydrolysis The 2 - group of RA facilitates chemical cleavage in aqueous a by forming
American Society of Cytopathology Core Curriculum in Molecular Biology
 Brought American Society of Cytopathology Core Curriculum in Molecular Biology Brought American Society of Cytopathology Core Curriculum in Molecular Biology Chapter 3 Molecular Techniques Alternatives
Brought American Society of Cytopathology Core Curriculum in Molecular Biology Brought American Society of Cytopathology Core Curriculum in Molecular Biology Chapter 3 Molecular Techniques Alternatives
TRANSCRIPTION AND PROCESSING OF RNA
 TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
Frederick Griffith. Dead Smooth Bacteria. Live Smooth Bacteria. Live Rough Bacteria. Live R+ dead S Bacteria
 Frederick Griffith Live Smooth Bacteria Live Rough Bacteria Dead Smooth Bacteria Live R+ dead S Bacteria Live Smooth Bacteria Frederick Griffith Live Rough Bacteria Dead Smooth Bacteria Live R+ dead S
Frederick Griffith Live Smooth Bacteria Live Rough Bacteria Dead Smooth Bacteria Live R+ dead S Bacteria Live Smooth Bacteria Frederick Griffith Live Rough Bacteria Dead Smooth Bacteria Live R+ dead S
Click here to read the case study about protein synthesis.
 Click here to read the case study about protein synthesis. Big Question: How do cells use the genetic information stored in DNA to make millions of different proteins the body needs? Key Concept: Genetics
Click here to read the case study about protein synthesis. Big Question: How do cells use the genetic information stored in DNA to make millions of different proteins the body needs? Key Concept: Genetics
DNA, RNA, PROTEIN SYNTHESIS, AND MUTATIONS UNIT GUIDE Due December 9 th. Monday Tuesday Wednesday Thursday Friday 16 CBA History of DNA video
 DNA, RNA, PROTEIN SYNTHESIS, AND MUTATIONS UNIT GUIDE Due December 9 th Monday Tuesday Wednesday Thursday Friday 16 CBA History of DNA video 17 History of DNA 18 Lecture: DNA Structure Worksheet 19 Lecture:
DNA, RNA, PROTEIN SYNTHESIS, AND MUTATIONS UNIT GUIDE Due December 9 th Monday Tuesday Wednesday Thursday Friday 16 CBA History of DNA video 17 History of DNA 18 Lecture: DNA Structure Worksheet 19 Lecture:
Efficient and Accurate Analysis of non coding RNAs with InSyBio ncrnaseq
 Efficient and Accurate Analysis of non coding RNAs with InSyBio ncrnaseq WHITE PAPER By InSyBio Ltd Aigli Korfiati Computer Engineer, MSc, PhD candidate InSyBio Product Development Manager October 2015
Efficient and Accurate Analysis of non coding RNAs with InSyBio ncrnaseq WHITE PAPER By InSyBio Ltd Aigli Korfiati Computer Engineer, MSc, PhD candidate InSyBio Product Development Manager October 2015
