Gene Structure & Gene Finding Part II
|
|
|
- Madeleine Marianna Washington
- 6 years ago
- Views:
Transcription
1 Gene Structure & Gene Finding Part II David Wishart 30,000 metabolite
2 Gene Finding in Eukaryotes Eukaryotes Complex gene structure Large genomes (0.1 to 10 billion bp) Exons and Introns (interrupted) Low coding density (<30%) 3% in humans, 25% in Fugu, 60% in yeast Alternate splicing (40-60% of all genes) High abundance of repeat sequence (50% in humans) and pseudo genes Nested genes: overlapping on same or opposite strand or inside an intron
3 Eukaryotic Gene Structure Transcribed Region exon 1 intron 1 exon 2 intron 2 exon3 Upstream Intergenic Region Start codon 5 UTR Stop codon 3 UTR Downstream Intergenic Region Eukaryotic Gene Structure branchpoint site 5 site 3 site exon 1 intron 1 exon 2 intron 2 AG/GT CAG/NT
4 RNA Splicing Exon/Intron Structure (Detail) ATGCTGTTAGGTGG...GCAGATCGATTGAC Exon 1 Intron 1 Exon 2 SPLICE ATGCTGTTAGATCGATTGAC
5 Intron Phase A codon can be interrupted by an intron in one of three places Phase 0: Phase 1: Phase 2: ATGATTGTCAG CAGTAC ATGATGTCAG CAGTTAC ATGAGTCAG CAGTTTAC SPLICE AGTATTTAC Repetitive DNA Moderately Repetitive DNA Tandem gene families (250 copies of rrna, trna gene copies) Pseudogenes (dead genes) Short interspersed elements (SINEs) bp long, 100,000+ copies, scattered Alu repeats are good examples Long interspersed elements (LINEs) bp long 10-10,000 copies per genome
6 Repetitive DNA Highly Repetitive DNA Minisatellite DNA repeats of bp stretching for ~2 kb many different types scattered thru genome Microsatellite DNA repeats of 5-13 bp stretching for 100 s of kb mostly found around centromere Telomeres highly conserved 6 bp repeat (TTAGGG) repeats at end of each chromosome Key Eukaryotic Gene Signals Pol II RNA promoter elements Cap and CCAAT region GC and TATA region Kozak sequence (Ribosome binding site-rbs) Splice donor, acceptor and lariat signals Termination signal Polyadenylation signal
7 Pol II Promoter Elements Exon Intron Exon GC box ~200 bp CCAAT box ~100 bp TATA box ~30 bp Gene Transcription start site (TSS) Pol II Promoter Elements Cap Region/Signal n C A G T n G TATA box (~ 25 bp upstream) T A T A A A n G C C C CCAAT box (~100 bp upstream) T A G C C A A T G GC box (~200 bp upstream) A T A G G C G nga
8 Pol II Promoter Elements TATA box is found in ~70% of promoters WebLogos
9 Kozak (RBS) Sequence A G C C A C C A T G G Splice Signals AG/GT branchpoint site CAG/NT exon 1 intron 1 exon 2
10 Splice Sites Not all splice sites are real ~0.5% of splice sites are non-canonical (i.e. the intron is not GT...AG) It is estimated that 5%of human genes may have non-canonical splice sites ~50% of higher eukaryotes are alternately spliced (different exons are brought together) Miscellaneous Signals Polyadenylation signal A A T A A A or A T T A A A Located 20 bp upstream of poly-a cleavage site Termination Signal A G T G T T C A Located ~30 bp downstream of poly-a cleavage site
11 Polyadenylation CPSF Cleavage & Polyadenylation Specificity Factor PAP Poly-A Polymerase CTsF Cleavage Stimulation Factor Why Polyadenylation is Really Useful Complementary Base Pairing AAAAAAAAAAA TTTTTTTTTTT T TT T T T TTT T TTT A AAA Poly dt Oligo bead T TTT A T A AA T T T T TT T
12 mrna isolation Cell or tissue sample is ground up and lysed with chemicals to release mrna Oligo(dT) beads are added and incubated with mixture to allow A-T annealing Pull down beads with magnet and pull off mrna Making cdna from mrna cdna (i.e. complementary DNA) is a single-stranded DNA segment whose sequence is complementary to that of messenger RNA (mrna) Synthesized by reverse transcriptase
13 Reverse Transcriptase Finding Eukaryotic Genes Experimentally Convert the spliced mrna into cdna cdna mrna A TGCTA TCTGTACTA Reverse transcriptase UACGAUAGACAUGAUAAAAAAAAAA Only expressed genes or expressed sequence tags (EST s) are seen Saves on sequencing effort (97%)
14 Finding Eukaryotic Genes Computationally Content-based Methods GC content, hexamer repeats, composition statistics, codon frequencies Site-based Methods donor sites, acceptor sites, promoter sites, start/stop codons, polya signals, lengths Comparative Methods sequence homology, EST searches Combined Methods Content-Based Methods CpG islands High GC content in 5 ends of genes Codon Bias Some codons are strongly preferred in coding regions, others are not Positional Bias 3rd base tends to be G/C rich in coding regions Ficketts Method looks for unequal base composition in different clusters of i, i+3, i+6 bases - TestCode graph
15 TestCode Plot Comparative Methods Do a BLASTX search of all 6 reading frames against known proteins in GenBank Assumes that the organism under study has genes that are homologous to known genes (used to be a problem, in 2001 analysis of chr. 22 only 50% of genes were similar to known proteins) BLAST against EST database (finds possible or probable 3 end of cdnas)
16 BLASTX Site-Based Methods Based on identifying gene signals (promoter elements, splice sites, start/stop codons, polya sites, etc.) Wide range of methods consensus sequences weight matrices neural networks decision trees hidden markov models (HMMs)
17 Neural Networks Automated method for classification or pattern recognition First described in detail in 1986 Mimic the way the brain works Use Matrix Algebra in calculations Require training on validated data Garbage in = Garbage out Neural Networks nodes Training Layer 1 Hidden Output Set Layer
18 Neural Network Applications Used in Intron/Exon Finding Used in Secondary Structure Prediction Used in Membrane Helix Prediction Used in Phosphorylation Site Prediction Used in Glycosylation Site Prediction Used in Splice Site Prediction Used in Signal Peptide Recognition Neural Network Training Set ACGAAG AGGAAG AGCAAG ACGAAA AGCAAC EEEENN Dersired Output Definitions A = [001] C = [010] G = [100] E = [01] N = [00] Sliding Window ACGAAG [ ] Input Vector [01] Output Vector
19 Neural Network Training [ ] ACGAAG [.6.4.6] e -x [.24.74] [0 1] compare Input Weight Hidden Weight Output Vector Matrix1 Layer Matrix2 Vector Back Propagation [ ] [.6.4.6] e -x [.24.74] [0 1] compare Input Weight Hidden Weight Output Vector Matrix1 Layer Matrix2 Vector
20 Calculate New Output [ ] [.7.4.7] e -x [.16.91] [0 1] Converged! Input Weight Hidden Weight Output Vector Matrix1 Layer Matrix2 Vector Train on Second Input Vector [ ] ACGAAG [.8.6.5] e -x [.12.95] [0 1] Compare Input Weight Hidden Weight Output Vector Matrix1 Layer Matrix2 Vector
21 Back Propagation e -x [ ] [.8.6.5] [.12.95] [0 1] compare Input Weight Hidden Weight Output Vector Matrix1 Layer Matrix2 Vector After Many Iterations Two Generalized Weight Matrices
22 Neural Networks Matrix1 Matrix2 ACGAGG New pattern EEEENN Prediction Input Layer 1 Hidden Output Layer HMM for Gene Finding
23 Combined Methods Bring 2 or more methods together (usually site detection + composition) GRAIL ( FGENEH ( HMMgene ( GENSCAN( Gene Parser ( GRPL (GeneTool/BioTools) Genscan
24 GENSCAN How Do They Work? 5th order Hidden Markov Model Hexamer composition statistics of exons vs. introns Exon/intron length distributions Scan of promoter and polya signals Weight matrices of 5 splice signals and start codon region (12 bp) Uses dynamic programming to optimize gene model using above data How Well Do They Do? Correlation (x100) Method Xpound GeneID GRAIL 2 Genie GeneP3 RPL RPL+ Burset & Guigio test set (1996)
25 How Well Do They Do? "Evaluation of gene finding programs" S. Rogic, A. K. Mackworth and B. F. F. Ouellette. Genome Research, 11: (2001). Easy vs. Hard Predictions 3 equally abundant states (easy) BUT random prediction = 33% correct Rare events, unequal distribution (hard) BUT biased random prediction = 90% correct
26 Gene Prediction (Evaluation) TP FP TN FN TP FN TN Actual Predicted Sensitivity Measure of the % of false negative results (sn = means 0.4% false negatives) Specificity Precision Correlation Measure of the % of false positive results Measure of the % positive results Combined measure of sensitivity and specificity Gene Prediction (Evaluation) TP FP TN FN TP FN TN Actual Predicted Sensitivity or Recall Specificity Precision Sn=TP/(TP + FN) Sp=TN/(TN + FP) Pr=TP/(TP + FP) Correlation CC=(TP*TN-FP*FN)/[(TP+FP)(TN+FN)(TP+FN)(TN+FP)] 0.5 This is a better way of evaluating
27 Different Strokes for Different Folks Precision and specificity statistics favor conservative predictors that make no prediction when there is doubt about the correctness of a prediction, while the sensitivity (recall) statistic favors liberal predictors that make a prediction if there is a chance of success. Information retrieval papers report precision and recall,while bioinformaticspapers tend to report specificity and sensitivity. Gene Prediction Accuracy at the Exon Level WRONG EXON CORRECT EXON MISSING EXON Actual Predicted Sensitivity S n = number of correct exons number of actual exons Specificity S p = number of correct exons number of predicted exons
28 Better Approaches Are Emerging... Programs that combine site, comparative and composition (3 in 1) GenomeScan, FGENESH++, Twinscan Programs that use synteny between organisms ROSETTA, SLAM, SGP Programs that combine predictions from multiple predictors GeneComber, DIGIT GenomeScan -
29 TwinScan - SLAM -
30 GeneComber - submit.php Outstanding Issues Most Gene finders don t handle UTRs (untranslated regions) ~40% of human genes have non-coding 1st exons (UTRs) Most gene finders don t handle alternative splicing Most gene finders don t handle overlapping or nested genes Most can t find non-protein genes (trnas)
31 Bottom Line... Gene finding in eukaryotes is not yet a solved problem Accuracy of the best methods approaches 80% at the exon level (90% at the nucleotide level) in coding-rich regions (much lower for whole genomes) Gene predictions should always be verified by other means (cdna sequencing, BLAST search, Mass spec.)
GenBank Growth. In 2003 ~ 31 million sequences ~ 37 billion base pairs
 Gene Finding GenBank Growth GenBank Growth In 2003 ~ 31 million sequences ~ 37 billion base pairs GenBank: Exponential Growth Growth of GenBank in billions of base pairs from release 3 in April of 1994
Gene Finding GenBank Growth GenBank Growth In 2003 ~ 31 million sequences ~ 37 billion base pairs GenBank: Exponential Growth Growth of GenBank in billions of base pairs from release 3 in April of 1994
Outline. Gene Finding Questions. Recap: Prokaryotic gene finding Eukaryotic gene finding The human gene complement Regulation
 Tues, Nov 29: Gene Finding 1 Online FCE s: Thru Dec 12 Thurs, Dec 1: Gene Finding 2 Tues, Dec 6: PS5 due Project presentations 1 (see course web site for schedule) Thurs, Dec 8 Final papers due Project
Tues, Nov 29: Gene Finding 1 Online FCE s: Thru Dec 12 Thurs, Dec 1: Gene Finding 2 Tues, Dec 6: PS5 due Project presentations 1 (see course web site for schedule) Thurs, Dec 8 Final papers due Project
Genscan. The Genscan HMM model Training Genscan Validating Genscan. (c) Devika Subramanian,
 Genscan The Genscan HMM model Training Genscan Validating Genscan (c) Devika Subramanian, 2009 96 Gene structure assumed by Genscan donor site acceptor site (c) Devika Subramanian, 2009 97 A simple model
Genscan The Genscan HMM model Training Genscan Validating Genscan (c) Devika Subramanian, 2009 96 Gene structure assumed by Genscan donor site acceptor site (c) Devika Subramanian, 2009 97 A simple model
How to design an HMM for a new problem. HMM model structure. Inherent limitation of HMMs. Duration modeling. Duration modeling
 How to design an HMM for a new problem Architecture/topology design: What are the states, observation symbols, and the topology of the state transition graph? Learning/Training: Fully annotated or partially
How to design an HMM for a new problem Architecture/topology design: What are the states, observation symbols, and the topology of the state transition graph? Learning/Training: Fully annotated or partially
Gene Identification in silico
 Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Genome annotation & EST
 Genome annotation & EST What is genome annotation? The process of taking the raw DNA sequence produced by the genome sequence projects and adding the layers of analysis and interpretation necessary
Genome annotation & EST What is genome annotation? The process of taking the raw DNA sequence produced by the genome sequence projects and adding the layers of analysis and interpretation necessary
Computational gene finding
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Genomics and Gene Recognition Genes and Blue Genes
 Genomics and Gene Recognition Genes and Blue Genes November 3, 2004 Eukaryotic Gene Structure eukaryotic genomes are considerably more complex than those of prokaryotes eukaryotic cells have organelles
Genomics and Gene Recognition Genes and Blue Genes November 3, 2004 Eukaryotic Gene Structure eukaryotic genomes are considerably more complex than those of prokaryotes eukaryotic cells have organelles
Computational gene finding
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding. Devika Subramanian Comp 470
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) The biological context Lec 1 Lec 2 Lec 3 Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) The biological context Lec 1 Lec 2 Lec 3 Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
CSE 527 Computational Biology Autumn Lectures ~14-15 Gene Prediction
 CSE 527 Computational Biology Autumn 2004 Lectures ~14-15 Gene Prediction Some References A great online bib http://www.nslij-genetics.org/gene/ A good intro survey JM Claverie (1997) "Computational methods
CSE 527 Computational Biology Autumn 2004 Lectures ~14-15 Gene Prediction Some References A great online bib http://www.nslij-genetics.org/gene/ A good intro survey JM Claverie (1997) "Computational methods
Transcription in Eukaryotes
 Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Annotating the Genome (H)
 Annotating the Genome (H) Annotation principles (H1) What is annotation? In general: annotation = explanatory note* What could be useful as an annotation of a DNA sequence? an amino acid sequence? What
Annotating the Genome (H) Annotation principles (H1) What is annotation? In general: annotation = explanatory note* What could be useful as an annotation of a DNA sequence? an amino acid sequence? What
I. Gene Expression Figure 1: Central Dogma of Molecular Biology
 I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
Homework 4. Due in class, Wednesday, November 10, 2004
 1 GCB 535 / CIS 535 Fall 2004 Homework 4 Due in class, Wednesday, November 10, 2004 Comparative genomics 1. (6 pts) In Loots s paper (http://www.seas.upenn.edu/~cis535/lab/sciences-loots.pdf), the authors
1 GCB 535 / CIS 535 Fall 2004 Homework 4 Due in class, Wednesday, November 10, 2004 Comparative genomics 1. (6 pts) In Loots s paper (http://www.seas.upenn.edu/~cis535/lab/sciences-loots.pdf), the authors
How Computing Science Saved The Human Genome Project Athabasca Hall Sept. 9, 2013
 How Computing Science Saved The Human Genome Project david.wishart@ualberta.ca 3-41 Athabasca Hall Sept. 9, 2013 The Human Genome Project HGP Announcement June 26, 2000*** J. Craig Venter Francis Collins
How Computing Science Saved The Human Genome Project david.wishart@ualberta.ca 3-41 Athabasca Hall Sept. 9, 2013 The Human Genome Project HGP Announcement June 26, 2000*** J. Craig Venter Francis Collins
Make the protein through the genetic dogma process.
 Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Reading Lecture 8: Lecture 9: Lecture 8. DNA Libraries. Definition Types Construction
 Lecture 8 Reading Lecture 8: 96-110 Lecture 9: 111-120 DNA Libraries Definition Types Construction 142 DNA Libraries A DNA library is a collection of clones of genomic fragments or cdnas from a certain
Lecture 8 Reading Lecture 8: 96-110 Lecture 9: 111-120 DNA Libraries Definition Types Construction 142 DNA Libraries A DNA library is a collection of clones of genomic fragments or cdnas from a certain
EGPred: Prediction of Eukaryotic Genes Using Ab Initio Methods After Combining With Sequence Similarity Approaches
 Methods EGPred: Prediction of Eukaryotic Genes Using Ab Initio Methods After Combining With Sequence Similarity Approaches Biju Issac and Gajendra Pal Singh Raghava 1 Institute of Microbial Technology,
Methods EGPred: Prediction of Eukaryotic Genes Using Ab Initio Methods After Combining With Sequence Similarity Approaches Biju Issac and Gajendra Pal Singh Raghava 1 Institute of Microbial Technology,
Genome annotation. Erwin Datema (2011) Sandra Smit (2012, 2013)
 Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
Improved Splice Site Detection in Genie
 Improved Splice Site Detection in Genie Martin Reese Informatics Group Human Genome Center Lawrence Berkeley National Laboratory MGReese@lbl.gov http://www-hgc.lbl.gov/inf Santa Fe, 1/23/97 Database Homologies
Improved Splice Site Detection in Genie Martin Reese Informatics Group Human Genome Center Lawrence Berkeley National Laboratory MGReese@lbl.gov http://www-hgc.lbl.gov/inf Santa Fe, 1/23/97 Database Homologies
Gene Prediction in Eukaryotes
 Gene Prediction in Eukaryotes Jan-Jaap Wesselink Biomol Informatics, S.L. jjw@biomol-informatics.com June 2010/Madrid jjw@biomol-informatics.com (BI) Gene Prediction June 2010/Madrid 1 / 34 Outline 1 Gene
Gene Prediction in Eukaryotes Jan-Jaap Wesselink Biomol Informatics, S.L. jjw@biomol-informatics.com June 2010/Madrid jjw@biomol-informatics.com (BI) Gene Prediction June 2010/Madrid 1 / 34 Outline 1 Gene
Lecture 7 Motif Databases and Gene Finding
 Introduction to Bioinformatics for Medical Research Gideon Greenspan gdg@cs.technion.ac.il Lecture 7 Motif Databases and Gene Finding Motif Databases & Gene Finding Motifs Recap Motif Databases TRANSFAC
Introduction to Bioinformatics for Medical Research Gideon Greenspan gdg@cs.technion.ac.il Lecture 7 Motif Databases and Gene Finding Motif Databases & Gene Finding Motifs Recap Motif Databases TRANSFAC
MODULE 5: TRANSLATION
 MODULE 5: TRANSLATION Lesson Plan: CARINA ENDRES HOWELL, LEOCADIA PALIULIS Title Translation Objectives Determine the codons for specific amino acids and identify reading frames by looking at the Base
MODULE 5: TRANSLATION Lesson Plan: CARINA ENDRES HOWELL, LEOCADIA PALIULIS Title Translation Objectives Determine the codons for specific amino acids and identify reading frames by looking at the Base
SSA Signal Search Analysis II
 SSA Signal Search Analysis II SSA other applications - translation In contrast to translation initiation in bacteria, translation initiation in eukaryotes is not guided by a Shine-Dalgarno like motif.
SSA Signal Search Analysis II SSA other applications - translation In contrast to translation initiation in bacteria, translation initiation in eukaryotes is not guided by a Shine-Dalgarno like motif.
Transcription and Post Transcript Modification
 Transcription and Post Transcript Modification You Should Be Able To 1. Describe transcription. 2. Compare and contrast eukaryotic + prokaryotic transcription. 3. Explain mrna processing in eukaryotes.
Transcription and Post Transcript Modification You Should Be Able To 1. Describe transcription. 2. Compare and contrast eukaryotic + prokaryotic transcription. 3. Explain mrna processing in eukaryotes.
Eukaryotic Gene Prediction. Wei Zhu May 2007
 Eukaryotic Gene Prediction Wei Zhu May 2007 In nature, nothing is perfect... - Alice Walker Gene Structure What is Gene Prediction? Gene prediction is the problem of parsing a sequence into nonoverlapping
Eukaryotic Gene Prediction Wei Zhu May 2007 In nature, nothing is perfect... - Alice Walker Gene Structure What is Gene Prediction? Gene prediction is the problem of parsing a sequence into nonoverlapping
Lecture Summary: Regulation of transcription. General mechanisms-what are the major regulatory points?
 BCH 401G Lecture 37 Andres Lecture Summary: Regulation of transcription. General mechanisms-what are the major regulatory points? RNA processing: Capping, polyadenylation, splicing. Why process mammalian
BCH 401G Lecture 37 Andres Lecture Summary: Regulation of transcription. General mechanisms-what are the major regulatory points? RNA processing: Capping, polyadenylation, splicing. Why process mammalian
UCSC Genome Browser. Introduction to ab initio and evidence-based gene finding
 UCSC Genome Browser Introduction to ab initio and evidence-based gene finding Wilson Leung 06/2006 Outline Introduction to annotation ab initio gene finding Basics of the UCSC Browser Evidence-based gene
UCSC Genome Browser Introduction to ab initio and evidence-based gene finding Wilson Leung 06/2006 Outline Introduction to annotation ab initio gene finding Basics of the UCSC Browser Evidence-based gene
Eukaryotic Gene Structure
 Eukaryotic Gene Structure Terminology Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Gene Basic physical and
Eukaryotic Gene Structure Terminology Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Gene Basic physical and
Chapter 2. An Introduction to Genes and Genomes
 PowerPoint Lectures for Introduction to Biotechnology, Second Edition William J.Thieman and Michael A.Palladino Chapter 2 An Introduction to Genes and Genomes Lectures by Lara Dowland Chapter Contents
PowerPoint Lectures for Introduction to Biotechnology, Second Edition William J.Thieman and Michael A.Palladino Chapter 2 An Introduction to Genes and Genomes Lectures by Lara Dowland Chapter Contents
Outline. Introduction to ab initio and evidence-based gene finding. Prokaryotic gene predictions
 Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
30 Gene expression: Transcription
 30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
Lecture for Wednesday. Dr. Prince BIOL 1408
 Lecture for Wednesday Dr. Prince BIOL 1408 THE FLOW OF GENETIC INFORMATION FROM DNA TO RNA TO PROTEIN Copyright 2009 Pearson Education, Inc. Genes are expressed as proteins A gene is a segment of DNA that
Lecture for Wednesday Dr. Prince BIOL 1408 THE FLOW OF GENETIC INFORMATION FROM DNA TO RNA TO PROTEIN Copyright 2009 Pearson Education, Inc. Genes are expressed as proteins A gene is a segment of DNA that
Transcription is the first stage of gene expression
 Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
ab initio and Evidence-Based Gene Finding
 ab initio and Evidence-Based Gene Finding A basic introduction to annotation Outline What is annotation? ab initio gene finding Genome databases on the web Basics of the UCSC browser Evidence-based gene
ab initio and Evidence-Based Gene Finding A basic introduction to annotation Outline What is annotation? ab initio gene finding Genome databases on the web Basics of the UCSC browser Evidence-based gene
CSE 527 Computational Biology" Gene Prediction"
 CSE 527 Computational Biology" Gene Prediction" Gene Finding: Motivation" Sequence data flooding in" What does it mean?" "protein genes, RNA genes, mitochondria, chloroplast, regulation, replication, structure,
CSE 527 Computational Biology" Gene Prediction" Gene Finding: Motivation" Sequence data flooding in" What does it mean?" "protein genes, RNA genes, mitochondria, chloroplast, regulation, replication, structure,
Gene & genome organisation. Computational gene identification
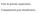 Gene & genome organisation Computational gene identification Eubacterial gene Eukaryotic gene Regulatory elements Promoter Translation start Introns polya signal Transcription stop DNA Transcription
Gene & genome organisation Computational gene identification Eubacterial gene Eukaryotic gene Regulatory elements Promoter Translation start Introns polya signal Transcription stop DNA Transcription
The Nature of Genes. The Nature of Genes. Genes and How They Work. Chapter 15/16
 Genes and How They Work Chapter 15/16 The Nature of Genes Beadle and Tatum proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes
Genes and How They Work Chapter 15/16 The Nature of Genes Beadle and Tatum proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes
Collect, analyze and synthesize. Annotation. Annotation for D. virilis. Evidence Based Annotation. GEP goals: Evidence for Gene Models 08/22/2017
 Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
PLNT2530 (2018) Unit 6b Sequence Libraries
 PLNT2530 (2018) Unit 6b Sequence Libraries Molecular Biotechnology (Ch 4) Analysis of Genes and Genomes (Ch 5) Unless otherwise cited or referenced, all content of this presenataion is licensed under the
PLNT2530 (2018) Unit 6b Sequence Libraries Molecular Biotechnology (Ch 4) Analysis of Genes and Genomes (Ch 5) Unless otherwise cited or referenced, all content of this presenataion is licensed under the
Collect, analyze and synthesize. Annotation. Annotation for D. virilis. GEP goals: Evidence Based Annotation. Evidence for Gene Models 12/26/2018
 Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
HUMAN GENOME BIOINFORMATICS. Tore Samuelsson, Dec 2009
 HUMAN GENOME BIOINFORMATICS Tore Samuelsson, Dec 2009 The sequenced (gray filled) and unsequenced (white) portions of the human genome. Peter F.R. Little Genome Res. 2005; 15: 1759-1766 Human genome organisation
HUMAN GENOME BIOINFORMATICS Tore Samuelsson, Dec 2009 The sequenced (gray filled) and unsequenced (white) portions of the human genome. Peter F.R. Little Genome Res. 2005; 15: 1759-1766 Human genome organisation
GENETICS - CLUTCH CH.10 TRANSCRIPTION.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF TRANSCRIPTION Transcription is the process of using DNA as a template to RNA RNA polymerase is the enzyme that transcribes DNA - There are many different types
!! www.clutchprep.com CONCEPT: OVERVIEW OF TRANSCRIPTION Transcription is the process of using DNA as a template to RNA RNA polymerase is the enzyme that transcribes DNA - There are many different types
Review of Protein (one or more polypeptide) A polypeptide is a long chain of..
 Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
BIOLOGY - CLUTCH CH.17 - GENE EXPRESSION.
 !! www.clutchprep.com CONCEPT: GENES Beadle and Tatum develop the one gene one enzyme hypothesis through their work with Neurospora (bread mold). This idea was later revised as the one gene one polypeptide
!! www.clutchprep.com CONCEPT: GENES Beadle and Tatum develop the one gene one enzyme hypothesis through their work with Neurospora (bread mold). This idea was later revised as the one gene one polypeptide
Chapter 3. DNA, RNA, and Protein Synthesis
 Chapter 3. DNA, RNA, and Protein Synthesis 4. Transcription Gene Expression Regulatory region (promoter) 5 flanking region Upstream region Coding region 3 flanking region Downstream region Transcription
Chapter 3. DNA, RNA, and Protein Synthesis 4. Transcription Gene Expression Regulatory region (promoter) 5 flanking region Upstream region Coding region 3 flanking region Downstream region Transcription
BS 50 Genetics and Genomics Week of Oct 24
 BS 50 Genetics and Genomics Week of Oct 24 Additional Practice Problems for Section Question 1: The following table contains a list of statements that apply to replication, transcription, both, or neither.
BS 50 Genetics and Genomics Week of Oct 24 Additional Practice Problems for Section Question 1: The following table contains a list of statements that apply to replication, transcription, both, or neither.
Genome Annotation. What Does Annotation Describe??? Genome duplications Genes Mobile genetic elements Small repeats Genetic diversity
 Genome Annotation Genome Sequencing Costliest aspect of sequencing the genome o But Devoid of content Genome must be annotated o Annotation definition Analyzing the raw sequence of a genome and describing
Genome Annotation Genome Sequencing Costliest aspect of sequencing the genome o But Devoid of content Genome must be annotated o Annotation definition Analyzing the raw sequence of a genome and describing
Chapter 6: Transcription and RNA Processing in Eukaryotes
 3. Basic Genetics Plant Molecular Biology Chapter 6: Transcription and RNA Processing in Eukaryotes - Genetic organization in eukaryote - Transcription in eukaryote - - RNA processing in eukaryote - Translation
3. Basic Genetics Plant Molecular Biology Chapter 6: Transcription and RNA Processing in Eukaryotes - Genetic organization in eukaryote - Transcription in eukaryote - - RNA processing in eukaryote - Translation
CH 17 :From Gene to Protein
 CH 17 :From Gene to Protein Defining a gene gene gene Defining a gene is problematic because one gene can code for several protein products, some genes code only for RNA, two genes can overlap, and there
CH 17 :From Gene to Protein Defining a gene gene gene Defining a gene is problematic because one gene can code for several protein products, some genes code only for RNA, two genes can overlap, and there
Sequence Analysis. II: Sequence Patterns and Matrices. George Bell, Ph.D. WIBR Bioinformatics and Research Computing
 Sequence Analysis II: Sequence Patterns and Matrices George Bell, Ph.D. WIBR Bioinformatics and Research Computing Sequence Patterns and Matrices Multiple sequence alignments Sequence patterns Sequence
Sequence Analysis II: Sequence Patterns and Matrices George Bell, Ph.D. WIBR Bioinformatics and Research Computing Sequence Patterns and Matrices Multiple sequence alignments Sequence patterns Sequence
Fig Ch 17: From Gene to Protein
 Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Genome 373: Gene Predic/on I. Doug Fowler
 Genome 373: Gene Predic/on I Doug Fowler Outline Review of gene structure Scale of the problem Solu;ons Empirical methods Ab ini&o predic;on What is a gene? A locatable region of genomic sequence, corresponding
Genome 373: Gene Predic/on I Doug Fowler Outline Review of gene structure Scale of the problem Solu;ons Empirical methods Ab ini&o predic;on What is a gene? A locatable region of genomic sequence, corresponding
Gene Expression: Transcription
 Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
Profile HMMs. 2/10/05 CAP5510/CGS5166 (Lec 10) 1 START STATE 1 STATE 2 STATE 3 STATE 4 STATE 5 STATE 6 END
 Profile HMMs START STATE 1 STATE 2 STATE 3 STATE 4 STATE 5 STATE 6 END 2/10/05 CAP5510/CGS5166 (Lec 10) 1 Profile HMMs with InDels Insertions Deletions Insertions & Deletions DELETE 1 DELETE 2 DELETE 3
Profile HMMs START STATE 1 STATE 2 STATE 3 STATE 4 STATE 5 STATE 6 END 2/10/05 CAP5510/CGS5166 (Lec 10) 1 Profile HMMs with InDels Insertions Deletions Insertions & Deletions DELETE 1 DELETE 2 DELETE 3
Chapter 4 METHOD FOR PREDICTING GENES IN EUKARYOTIC GENOMES
 qene Prediction Chapter 4 METHOD FOR PREDICTING GENES IN EUKARYOTIC GENOMES 4.1 INTRODUCTION Evaluation of seven ab initio methods that were evaluated on a non-homologous mammalian data set (Rogic et al.
qene Prediction Chapter 4 METHOD FOR PREDICTING GENES IN EUKARYOTIC GENOMES 4.1 INTRODUCTION Evaluation of seven ab initio methods that were evaluated on a non-homologous mammalian data set (Rogic et al.
Themes: RNA and RNA Processing. Messenger RNA (mrna) What is a gene? RNA is very versatile! RNA-RNA interactions are very important!
 Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
May 16. Gene Finding
 Gene Finding j T[j,k] k i Q is a set of states T is a matrix of transition probabilities T[j,k]: probability of moving from state j to state k Σ is a set of symbols e j (S) is the probability of emitting
Gene Finding j T[j,k] k i Q is a set of states T is a matrix of transition probabilities T[j,k]: probability of moving from state j to state k Σ is a set of symbols e j (S) is the probability of emitting
Applications of HMMs in Computational Biology. BMI/CS Colin Dewey
 Applications of HMMs in Computational Biology BMI/CS 576 www.biostat.wisc.edu/bmi576.html Colin Dewey cdewey@biostat.wisc.edu Fall 2008 The Gene Finding Task Given: an uncharacterized DNA sequence Do:
Applications of HMMs in Computational Biology BMI/CS 576 www.biostat.wisc.edu/bmi576.html Colin Dewey cdewey@biostat.wisc.edu Fall 2008 The Gene Finding Task Given: an uncharacterized DNA sequence Do:
Annotation of contig27 in the Muller F Element of D. elegans. Contig27 is a 60,000 bp region located in the Muller F element of the D. elegans.
 David Wang Bio 434W 4/27/15 Annotation of contig27 in the Muller F Element of D. elegans Abstract Contig27 is a 60,000 bp region located in the Muller F element of the D. elegans. Genscan predicted six
David Wang Bio 434W 4/27/15 Annotation of contig27 in the Muller F Element of D. elegans Abstract Contig27 is a 60,000 bp region located in the Muller F element of the D. elegans. Genscan predicted six
Transcription. The sugar molecule found in RNA is ribose, rather than the deoxyribose found in DNA.
 Transcription RNA (ribonucleic acid) is a key intermediary between a DNA sequence and a polypeptide. RNA is an informational polynucleotide similar to DNA, but it differs from DNA in three ways: RNA generally
Transcription RNA (ribonucleic acid) is a key intermediary between a DNA sequence and a polypeptide. RNA is an informational polynucleotide similar to DNA, but it differs from DNA in three ways: RNA generally
Transcription steps. Transcription steps. Eukaryote RNA processing
 Transcription steps Initiation at 5 end of gene binding of RNA polymerase to promoter unwinding of DNA Elongation addition of nucleotides to 3 end rules of base pairing requires Mg 2+ energy from NTP substrates
Transcription steps Initiation at 5 end of gene binding of RNA polymerase to promoter unwinding of DNA Elongation addition of nucleotides to 3 end rules of base pairing requires Mg 2+ energy from NTP substrates
Year III Pharm.D Dr. V. Chitra
 Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
The Genetic Code and Transcription. Chapter 12 Honors Genetics Ms. Susan Chabot
 The Genetic Code and Transcription Chapter 12 Honors Genetics Ms. Susan Chabot TRANSCRIPTION Copy SAME language DNA to RNA Nucleic Acid to Nucleic Acid TRANSLATION Copy DIFFERENT language RNA to Amino
The Genetic Code and Transcription Chapter 12 Honors Genetics Ms. Susan Chabot TRANSCRIPTION Copy SAME language DNA to RNA Nucleic Acid to Nucleic Acid TRANSLATION Copy DIFFERENT language RNA to Amino
Gene Prediction. Mario Stanke. Institut für Mikrobiologie und Genetik Abteilung Bioinformatik. Gene Prediction p.
 Gene Prediction Mario Stanke mstanke@gwdg.de Institut für Mikrobiologie und Genetik Abteilung Bioinformatik Gene Prediction p.1/23 Why Predict Genes with a Computer? tons of data 39/250 eukaryotic/prokaryotic
Gene Prediction Mario Stanke mstanke@gwdg.de Institut für Mikrobiologie und Genetik Abteilung Bioinformatik Gene Prediction p.1/23 Why Predict Genes with a Computer? tons of data 39/250 eukaryotic/prokaryotic
Annotating Fosmid 14p24 of D. Virilis chromosome 4
 Lo 1 Annotating Fosmid 14p24 of D. Virilis chromosome 4 Lo, Louis April 20, 2006 Annotation Report Introduction In the first half of Research Explorations in Genomics I finished a 38kb fragment of chromosome
Lo 1 Annotating Fosmid 14p24 of D. Virilis chromosome 4 Lo, Louis April 20, 2006 Annotation Report Introduction In the first half of Research Explorations in Genomics I finished a 38kb fragment of chromosome
B. Incorrect! Centromeric DNA is largely heterochromatin, which is inactive DNA.
 MCAT Biology - Problem Drill 06: Molecular Biology of Eukaryotes Question No. 1 of 10 1. Which type of DNA would have the highest level of expression? Question #01 (A) Heterochromatin. (B) Centromeric
MCAT Biology - Problem Drill 06: Molecular Biology of Eukaryotes Question No. 1 of 10 1. Which type of DNA would have the highest level of expression? Question #01 (A) Heterochromatin. (B) Centromeric
Fermentation. Lesson Overview. Lesson Overview 13.1 RNA
 13.1 RNA THINK ABOUT IT DNA is the genetic material of cells. The sequence of nucleotide bases in the strands of DNA carries some sort of code. In order for that code to work, the cell must be able to
13.1 RNA THINK ABOUT IT DNA is the genetic material of cells. The sequence of nucleotide bases in the strands of DNA carries some sort of code. In order for that code to work, the cell must be able to
Computational Biology I LSM5191 (2003/4)
 Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
Genes and How They Work. Chapter 15
 Genes and How They Work Chapter 15 The Nature of Genes They proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes The central
Genes and How They Work Chapter 15 The Nature of Genes They proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes The central
DNA Replication and Repair
 DNA Replication and Repair http://hyperphysics.phy-astr.gsu.edu/hbase/organic/imgorg/cendog.gif Overview of DNA Replication SWYK CNs 1, 2, 30 Explain how specific base pairing enables existing DNA strands
DNA Replication and Repair http://hyperphysics.phy-astr.gsu.edu/hbase/organic/imgorg/cendog.gif Overview of DNA Replication SWYK CNs 1, 2, 30 Explain how specific base pairing enables existing DNA strands
BIO 311C Spring Lecture 36 Wednesday 28 Apr.
 BIO 311C Spring 2010 1 Lecture 36 Wednesday 28 Apr. Synthesis of a Polypeptide Chain 5 direction of ribosome movement along the mrna 3 ribosome mrna NH 2 polypeptide chain direction of mrna movement through
BIO 311C Spring 2010 1 Lecture 36 Wednesday 28 Apr. Synthesis of a Polypeptide Chain 5 direction of ribosome movement along the mrna 3 ribosome mrna NH 2 polypeptide chain direction of mrna movement through
Transcription and Translation
 Transcription and Translation Central Dogma of Molecular The flow of information in the cell starts at DNA, which replicates to form more DNA. Information is then transcribed into RNA, and then it is translated
Transcription and Translation Central Dogma of Molecular The flow of information in the cell starts at DNA, which replicates to form more DNA. Information is then transcribed into RNA, and then it is translated
Regulation of bacterial gene expression
 Regulation of bacterial gene expression Gene Expression Gene Expression: RNA and protein synthesis DNA ----------> RNA ----------> Protein transcription translation! DNA replication only occurs in cells
Regulation of bacterial gene expression Gene Expression Gene Expression: RNA and protein synthesis DNA ----------> RNA ----------> Protein transcription translation! DNA replication only occurs in cells
Molecular Cell Biology - Problem Drill 08: Transcription, Translation and the Genetic Code
 Molecular Cell Biology - Problem Drill 08: Transcription, Translation and the Genetic Code Question No. 1 of 10 1. Which of the following statements about how genes function is correct? Question #1 (A)
Molecular Cell Biology - Problem Drill 08: Transcription, Translation and the Genetic Code Question No. 1 of 10 1. Which of the following statements about how genes function is correct? Question #1 (A)
Finding Eukaryotic Genes Computationally
 Gene Identification Finding Eukaryotic Genes Computationally ü Content-based Methods GC content, hexamer repeats, composition statistics, codon frequencies ü Site-based Methods donor sites, acceptor sites,
Gene Identification Finding Eukaryotic Genes Computationally ü Content-based Methods GC content, hexamer repeats, composition statistics, codon frequencies ü Site-based Methods donor sites, acceptor sites,
DNAFSMiner: A Web-Based Software Toolbox to Recognize Two Types of Functional Sites in DNA Sequences
 DNAFSMiner: A Web-Based Software Toolbox to Recognize Two Types of Functional Sites in DNA Sequences Huiqing Liu Hao Han Jinyan Li Limsoon Wong Institute for Infocomm Research, 21 Heng Mui Keng Terrace,
DNAFSMiner: A Web-Based Software Toolbox to Recognize Two Types of Functional Sites in DNA Sequences Huiqing Liu Hao Han Jinyan Li Limsoon Wong Institute for Infocomm Research, 21 Heng Mui Keng Terrace,
132 Grundlagen der Bioinformatik, SoSe 14, D. Huson, June 22, This exposition is based on the following source, which is recommended reading:
 132 Grundlagen der Bioinformatik, SoSe 14, D. Huson, June 22, 214 1 Gene Prediction Using HMMs This exposition is based on the following source, which is recommended reading: 1. Chris Burge and Samuel
132 Grundlagen der Bioinformatik, SoSe 14, D. Huson, June 22, 214 1 Gene Prediction Using HMMs This exposition is based on the following source, which is recommended reading: 1. Chris Burge and Samuel
SCBC203 Gene Expression. Assoc. Prof. Rutaiwan Tohtong Department of Biochemistry Faculty of Science PR318
 SCBC203 Gene Expression Assoc. Prof. Rutaiwan Tohtong Department of Biochemistry Faculty of Science PR318 Rutaiwan.toh@mahidol.ac.th 1 Gene Expression Gene expression is a process where by the genetic
SCBC203 Gene Expression Assoc. Prof. Rutaiwan Tohtong Department of Biochemistry Faculty of Science PR318 Rutaiwan.toh@mahidol.ac.th 1 Gene Expression Gene expression is a process where by the genetic
BIOCHEMISTRY REVIEW. Overview of Biomolecules. Chapter 12 Transcription
 BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 12 Transcription 2 3 4 5 Are You Getting It?? Which are general characteristics of transcription? (multiple answers) a) An entire DNA molecule is transcribed
BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 12 Transcription 2 3 4 5 Are You Getting It?? Which are general characteristics of transcription? (multiple answers) a) An entire DNA molecule is transcribed
Grundlagen der Bioinformatik, SoSe 11, D. Huson, July 4, This exposition is based on the following source, which is recommended reading:
 Grundlagen der Bioinformatik, SoSe 11, D. Huson, July 4, 211 155 12 Gene Prediction Using HMMs This exposition is based on the following source, which is recommended reading: 1. Chris Burge and Samuel
Grundlagen der Bioinformatik, SoSe 11, D. Huson, July 4, 211 155 12 Gene Prediction Using HMMs This exposition is based on the following source, which is recommended reading: 1. Chris Burge and Samuel
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Prediction 10/21/05
 Gene Prediction 1/21/5 1/21/5 Gene Prediction Announcements Eam 2 - net Friday Posted online: Eam 2 Study Guide 544 Reading Assignment (2 papers) (formerly Gene Prediction - ) 1/21/5 D Dobbs ISU - BCB
Gene Prediction 1/21/5 1/21/5 Gene Prediction Announcements Eam 2 - net Friday Posted online: Eam 2 Study Guide 544 Reading Assignment (2 papers) (formerly Gene Prediction - ) 1/21/5 D Dobbs ISU - BCB
Lecture 11. Initiation of RNA Pol II transcription. Transcription Initiation Complex
 Lecture 11 *Eukaryotic Transcription Gene Organization RNA Processing 5 cap 3 polyadenylation splicing Translation Initiation of RNA Pol II transcription Consensus sequence of promoter TATA Transcription
Lecture 11 *Eukaryotic Transcription Gene Organization RNA Processing 5 cap 3 polyadenylation splicing Translation Initiation of RNA Pol II transcription Consensus sequence of promoter TATA Transcription
Transcription. By : Lucia Dhiantika Witasari M.Biotech., Apt
 Transcription By : Lucia Dhiantika Witasari M.Biotech., Apt REGULATION OF GENE EXPRESSION 11/26/2010 2 RNA Messenger RNAs (mrnas) encode the amino acid sequence of one or more polypeptides specified by
Transcription By : Lucia Dhiantika Witasari M.Biotech., Apt REGULATION OF GENE EXPRESSION 11/26/2010 2 RNA Messenger RNAs (mrnas) encode the amino acid sequence of one or more polypeptides specified by
There are four major types of introns. Group I introns, found in some rrna genes, are self-splicing: they can catalyze their own removal.
 1 2 Continuous genes - Intron: Many eukaryotic genes contain coding regions called exons and noncoding regions called intervening sequences or introns. The average human gene contains from eight to nine
1 2 Continuous genes - Intron: Many eukaryotic genes contain coding regions called exons and noncoding regions called intervening sequences or introns. The average human gene contains from eight to nine
What is a Gene? Snyder and Gerstein, Science Gene Finding Questions
 ues, Nov 28: ene Finding 1 Online FE s: hru Dec 11 hurs, Nov 30: ene Finding 2 ues, Dec 5: PS5 due in my office at 5pm. Project presentation preparation no class hurs, Dec 7 Final papers due Project presentations
ues, Nov 28: ene Finding 1 Online FE s: hru Dec 11 hurs, Nov 30: ene Finding 2 ues, Dec 5: PS5 due in my office at 5pm. Project presentation preparation no class hurs, Dec 7 Final papers due Project presentations
Chapters 31-32: Ribonucleic Acid (RNA)
 Chapters 31-32: Ribonucleic Acid (RNA) Short segments from the transcription, processing and translation sections of each chapter Slide 1 RNA In comparison with DNA RNA utilizes uracil in place of thymine
Chapters 31-32: Ribonucleic Acid (RNA) Short segments from the transcription, processing and translation sections of each chapter Slide 1 RNA In comparison with DNA RNA utilizes uracil in place of thymine
Gene Expression: Transcription, Translation, RNAs and the Genetic Code
 Lecture 28-29 Gene Expression: Transcription, Translation, RNAs and the Genetic Code Central dogma of molecular biology During transcription, the information in a DNA sequence (a gene) is copied into a
Lecture 28-29 Gene Expression: Transcription, Translation, RNAs and the Genetic Code Central dogma of molecular biology During transcription, the information in a DNA sequence (a gene) is copied into a
The Flow of Genetic Information
 Chapter 17 The Flow of Genetic Information The DNA inherited by an organism leads to specific traits by dictating the synthesis of proteins and of RNA molecules involved in protein synthesis. Proteins
Chapter 17 The Flow of Genetic Information The DNA inherited by an organism leads to specific traits by dictating the synthesis of proteins and of RNA molecules involved in protein synthesis. Proteins
Methods and Algorithms for Gene Prediction
 Methods and Algorithms for Gene Prediction Chaochun Wei 韦朝春 Sc.D. ccwei@sjtu.edu.cn http://cbb.sjtu.edu.cn/~ccwei Shanghai Jiao Tong University Shanghai Center for Bioinformation Technology 5/12/2011 K-J-C
Methods and Algorithms for Gene Prediction Chaochun Wei 韦朝春 Sc.D. ccwei@sjtu.edu.cn http://cbb.sjtu.edu.cn/~ccwei Shanghai Jiao Tong University Shanghai Center for Bioinformation Technology 5/12/2011 K-J-C
#28 - Promoter Prediction 10/29/07
 BCB 444/544 Required Reading (before lecture) Lecture 28 Mon Oct 29 - Lecture 28 Promoter & Regulatory Element Prediction Chp 9 - pp 113-126 Gene Prediction - finish it Wed Oct 30 - Lecture 29 Phylogenetics
BCB 444/544 Required Reading (before lecture) Lecture 28 Mon Oct 29 - Lecture 28 Promoter & Regulatory Element Prediction Chp 9 - pp 113-126 Gene Prediction - finish it Wed Oct 30 - Lecture 29 Phylogenetics
RNA : functional role
 RNA : functional role Hamad Yaseen, PhD MLS Department, FAHS Hamad.ali@hsc.edu.kw RNA mrna rrna trna 1 From DNA to Protein -Outline- From DNA to RNA From RNA to Protein From DNA to RNA Transcription: Copying
RNA : functional role Hamad Yaseen, PhD MLS Department, FAHS Hamad.ali@hsc.edu.kw RNA mrna rrna trna 1 From DNA to Protein -Outline- From DNA to RNA From RNA to Protein From DNA to RNA Transcription: Copying
We have. Learned the structure of DNA. Talked about DNA replication and all of the complicated vocabulary that goes with it
 Do Now 1. What enzyme inserts new DNA nucleotides during replication? 2. Name 2 other important players in DNA replication and their function. 3. In what direction can DNA polymerase add nucleotides? 4.
Do Now 1. What enzyme inserts new DNA nucleotides during replication? 2. Name 2 other important players in DNA replication and their function. 3. In what direction can DNA polymerase add nucleotides? 4.
Biology. Biology. Slide 1 of 39. End Show. Copyright Pearson Prentice Hall
 Biology Biology 1 of 39 12-3 RNA and Protein Synthesis 2 of 39 Essential Question What is transcription and translation and how do they take place? 3 of 39 12 3 RNA and Protein Synthesis Genes are coded
Biology Biology 1 of 39 12-3 RNA and Protein Synthesis 2 of 39 Essential Question What is transcription and translation and how do they take place? 3 of 39 12 3 RNA and Protein Synthesis Genes are coded
Biology. Biology. Slide 1 of 39. End Show. Copyright Pearson Prentice Hall
 Biology Biology 1 of 39 12-3 RNA and Protein Synthesis 2 of 39 12 3 RNA and Protein Synthesis Genes are coded DNA instructions that control the production of proteins. Genetic messages can be decoded by
Biology Biology 1 of 39 12-3 RNA and Protein Synthesis 2 of 39 12 3 RNA and Protein Synthesis Genes are coded DNA instructions that control the production of proteins. Genetic messages can be decoded by
Branches of Genetics
 Branches of Genetics 1. Transmission genetics Classical genetics or Mendelian genetics 2. Molecular genetics chromosomes, DNA, regulation of gene expression recombinant DNA, biotechnology, bioinformatics,
Branches of Genetics 1. Transmission genetics Classical genetics or Mendelian genetics 2. Molecular genetics chromosomes, DNA, regulation of gene expression recombinant DNA, biotechnology, bioinformatics,
Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site.
 Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site. Still having trouble understanding the material? Check
Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site. Still having trouble understanding the material? Check
