SUPPLEMENTARY DATA. DNAproDB: an interactive tool for structural analysis of DNA-protein complexes
|
|
|
- Rosamond Heath
- 6 years ago
- Views:
Transcription
1 SUPPLEMENTARY DATA DNAproDB: an interactive tool for structural analysis of DNA-protein complexes Jared M. Sagendorf 1, Helen M. Berman 2, *, and Remo Rohs 1, * 1 Molecular and Computational Biology Program, Departments of Biological Sciences, Chemistry, Physics & Astronomy, and Computer Science, University of Southern California, Los Angeles, CA 90089, USA 2 RCSB Protein Data Bank, Center for Integrative Proteomics Research, Rutgers, The State University of New Jersey, Piscataway, NJ 08854, USA *To whom correspondence should be addressed. Tel: ; Fax: ; rohs@usc.edu (R.R.) or Tel: ; berman@rcsb.rutgers.edu (H.M.B.) SUPPLEMENTARY METHODS FEATURE EXTRACTION Protein secondary structure DNAproDB assigns a three-state secondary structure to each residue in the DNA-protein interface using the program DSSP (1). DSSP assigns an eight-state secondary structure; H α helix, G 3 10 helix, I π helix, E extended strand, B isolated β-bridge, S bend, T hydrogen bonded turn, and blank - loop/irregular. To map from eight-state to three-state secondary structure, H, G and I are assigned as helix (H); E and B are assigned as strand (S); and S, T, and blank are assigned as loop (L). Individual strands are treated independently without regard to the formation of β-sheets. Buried solvent-accessible surface area The buried solvent-accessible surface area (BASA) is the difference between the solventaccessible surface area (SASA) (2) of some unit in the structure when it is in the free state and when it is in the bound state. For example, for every residue in the protein, SASA is calculated with all DNA removed from the structure, which is defined as the free state of the protein. All residues with a free SASA (SASA F ) > 0 constitute the protein surface, and may potentially 1
2 interact with the DNA. SASA values are re-calculated with the DNA present to determine the complex SASA (SASA C ). The BASA of each residue is defined as BASA = SASA F SASA C, which will always be greater than or equal to zero. Residues with BASA > 0 are considered to be in contact with the DNA, and the BASA value describes the extent of the contact. The same calculation is performed for each nucleotide, with the free state of the DNA corresponding to the structure with all protein residues removed. Different contributions of the BASA values are determined. For each residue, it is possible to specify how much of the total BASA is due to contact with the DNA major groove, minor groove, or backbone. To calculate these contributions, instead of removing the entire DNA structure for the free-state calculation, only part of the DNA is removed. To calculate the contribution of the DNA major groove to the BASA value of each residue, only the DNA major groove atoms are removed from the structure in the free state. Therefore, BASA wg = SASA F (wg) SASA C, where BASA wg is the surface area in Å 2 of lost SASA for each residue due to contact with the DNA major groove (wg for wide groove ); and SASA F (wg) is the SASA of each residue with only DNA major groove atoms removed. In this way, BASA wg (major groove; wg), BASA sg (minor groove; sg for small groove ) and BASA bb (backbone; bb for backbone ) contributions are determined for each residue. Similarly, for each DNA nucleotide, the BASA contributions due to helices, strands, and loops are determined. This analysis gives a more detailed description of the interface in terms of how much different parts of the structure are contacting each other. However, the sum of each contribution of the BASA may be less than the total BASA (but never greater) because calculating the BASA contributions in this way excludes overlapping surface areas. BASA values are also determined for each residue-nucleotide pair. Here, the entire structure is removed except for the pair, and the free and complex states are calculated as described. Individual contributions of the BASA are not determined for residue-nucleotide pairs. 2
3 SUPPLEMENTARY TABLE PDB ID 5CM DI Modified Nucleotides CHEMICAL NAME PDB ID MSE SEP Modified Residues CHEMICAL NAME SELENOMETHIONINE PHOSPHOSERINE 5-METHYL-2 -DEOXY-CYTIDINE-5-2 -DEOXYINOSINE-5-6OG 6-O-METHYL GUANOSINE-5 - TPO PHOSPHOTHREONINE DOC 2,3 -DIDEOXYCYTIDINE-5 - PTR O-PHOSPHOTYROSINE 8OG 8-OXO-2 -DEOXY-GUANOSINE-5 - S,S-(2- CME HYDROXYETHYL)THIOCYSTEINE DDG 2,3 -DIDEOXY-GUANOSINE-5 - OCS CYSTEINESULFONIC ACID 2DA 2,3 -DIDEOXYADENOSINE-5 - ALY N(6)-ACETYLLYSINE 2DT 3 -DEOXYTHYMIDINE-5-2PR 2-AMINO-9-[2-DEOXYRIBOFURANOSYL]-9H- PURINE-5 - MRG N2-(3-MERCAPTOPROPYL)-2 - DEOXYGUANOSINE-5 - MERCAPTOPROPYL)-2 - DEOXYGUANOSINE-5 - CTG (5R,6S)-5,6-DIHYDRO-5,6- DIHYDROXYTHYMIDINE-5-1CC 5-CARBOXY-2 -DEOXYCYTIDINE Supplementary Table 1. Currently supported chemically modified nucleotides and residues. Any non-standard residues or nucleotide not in this table will be removed from the structure during processing. 3
4 SUPPLEMENTARY FIGURES Supplementary Figure S1. Illustration of the axial coordinate system used to specify the position of each SSE in the DNA-protein interface. Let P be a point in space that indicates the position of an SSE, C be the curve defining the DNA helical axis, and P be a point on C where a perpendicular line can be drawn from P to C. The axial coordinates that define P are (φ, ρ, s). ρ is the distance from P to P ; s is the distance along the curve from a specified point on the curve to P (here, chosen as the end of the helix axis, corresponding to the 5 end of the first DNA strand in the structure); and φ is the angle made in a plane defined by the tangent vector of the curve at P between the line joining P and P which lies in this plane, and the projection of a line joining P and a fixed point in space to the plane. This fixed point is arbitrary; changing the fixed point simply amounts to adding some constant offset to φ relative to the original choice. In practice, the center of mass of the protein is chosen, which allows similar structures to be compared consistently. If this center of mass lies too near the DNA helix axis (which is the case for structures with perfect two-fold symmetry), the point is chosen to lie along a principal axis of the protein. (A) The curve in the figure represents the DNA helix axis (C), which, in this case, is not linear. The red circle is the position of an SSE (P). The length of the line joining P and P is the coordinate ρ. The distance along C from the 5 end to P is the coordinate s. The green plane contains the line joining P and P, and is defined by the tangent vector of C at P. (B) The angle between the projection of the line that joins P and the fixed point to the plane, and the line that joins P and P is the coordinate φ. 4
5 Supplementary Figure S2. Polar contact maps of selected structures meeting the search criteria described in Table 1. The complex with PDB ID 1JFI (3) contains a ternary complex of a TATA-binding protein (TBP) with Negative Cofactor 2 (NC2) bound to a TATA-box. The polar contact map shows a characteristic TBP interaction, with a series of strands in the minor groove (mg) in addition to two helices that contact the minor groove belonging to an NC2 domain. The complex with PDB ID 2GKD (4) shows the bacterial resistance protein Calicheamicin gene Cluster (CalC), which binds with a single helix in the minor groove and few other contacts. The complexes with PDB IDs 1J46 (5) and 3U2B (6) contain proteins that predominantly bind with two helices and several loop contacts in the minor groove. 5
6 Supplementary Figure S3. Different visualizations for a TATA-box binding DNA-protein complex (PDB ID 1VTO) (7). Green triangles represent strand SSEs; and blue squares represent loops. (A) Polar contact map for this structure. The DNA helix axis is curved, but the visualization shows the strands as contacting on a single side of the DNA (i.e. confined to one half of the figure). This visualization is intuitive when compared with the 3D structure in (B). (B) 3D representation of the structure, with strand and loop residues that contact the minor groove colored according to SSE type in green and blue, respectively. (C) Linear contact map of this structure, showing minor groove (mg) and major groove (MG) contacts. (D) Nucleotide-residue contact map, showing only DNA backbone (BB) interactions with strand residues. Loop interactions have been turned off. 6
7 SUPPLEMENTARY REFERENCES 1. Kabsch, W. and Sander, C. (1983) Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22, Lee, B. and Richards, F.M. (1971) The interpretation of protein structures: estimation of static accessibility. J. Mol. Biol. 55, Kamada, K., Shu, F., Chen, H., Malik, S., Stelzer, G., Roeder, R.G., Meisterernst, M. and Burley, S.K. (2001) Crystal structure of negative cofactor 2 recognizing the TBP- DNA transcription complex. Cell 106, Singh, S., Hager, M.H., Zhang, C., Griffith, B.R., Lee, M.S., Hallenga, K., Markley, J.L. and Thorson, J.S. (2006) Structural insight into the self-sacrifice mechanism of enediyne resistance. ACS Chem. Biol. 1, Murphy, E.C., Zhurkin, V.B., Louis, J.M., Cornilescu, G. and Clore, G.M. (2001) Structural basis for SRY-dependent 46-X,Y sex reversal: modulation of DNA bending by a naturally occurring point mutation. J. Mol. Biol. 312, Jauch, R., Ng, C.K., Narasimhan, K. and Kolatkar, P.R. (2012) The crystal structure of the Sox4 HMG domain-dna complex suggests a mechanism for positional interdependence in DNA recognition. Biochem. J. 443, Kim, J.L. and Burley, S.K. (1994) 1.9 A resolution refined structure of TBP recognizing the minor groove of TATAAAAG. Nat. Struct. Biol. 1,
Nucleic acids. How DNA works. DNA RNA Protein. DNA (deoxyribonucleic acid) RNA (ribonucleic acid) Central Dogma of Molecular Biology
 Nucleic acid chemistry and basic molecular theory Nucleic acids DNA (deoxyribonucleic acid) RNA (ribonucleic acid) Central Dogma of Molecular Biology Cell cycle DNA RNA Protein Transcription Translation
Nucleic acid chemistry and basic molecular theory Nucleic acids DNA (deoxyribonucleic acid) RNA (ribonucleic acid) Central Dogma of Molecular Biology Cell cycle DNA RNA Protein Transcription Translation
Structural Bioinformatics (C3210) DNA and RNA Structure
 Structural Bioinformatics (C3210) DNA and RNA Structure Importance of DNA/RNA 3D Structure Nucleic acids are essential materials found in all living organisms. Their main function is to maintain and transmit
Structural Bioinformatics (C3210) DNA and RNA Structure Importance of DNA/RNA 3D Structure Nucleic acids are essential materials found in all living organisms. Their main function is to maintain and transmit
Hmwk # 8 : DNA-Binding Proteins : Part II
 The purpose of this exercise is : Hmwk # 8 : DNA-Binding Proteins : Part II 1). to examine the case of a tandem head-to-tail homodimer binding to DNA 2). to view a Zn finger motif 3). to consider the case
The purpose of this exercise is : Hmwk # 8 : DNA-Binding Proteins : Part II 1). to examine the case of a tandem head-to-tail homodimer binding to DNA 2). to view a Zn finger motif 3). to consider the case
Molecular Biology (1)
 Molecular Biology (1) DNA structure and basic applications Mamoun Ahram, PhD Second semester, 2017-2018 Resources This lecture Cooper, pp. 49-52, 118-119, 130 What is molecular biology? Central dogma
Molecular Biology (1) DNA structure and basic applications Mamoun Ahram, PhD Second semester, 2017-2018 Resources This lecture Cooper, pp. 49-52, 118-119, 130 What is molecular biology? Central dogma
DNA & DNA : Protein Interactions BIBC 100
 DNA & DNA : Protein Interactions BIBC 100 Sequence = Information Alphabet = language L,I,F,E LIFE DNA = DNA code A, T, C, G CAC=Histidine CAG=Glutamine GGG=Glycine Protein = Protein code 20 a.a. LIVE EVIL
DNA & DNA : Protein Interactions BIBC 100 Sequence = Information Alphabet = language L,I,F,E LIFE DNA = DNA code A, T, C, G CAC=Histidine CAG=Glutamine GGG=Glycine Protein = Protein code 20 a.a. LIVE EVIL
RNA does not adopt the classic B-DNA helix conformation when it forms a self-complementary double helix
 Reason: RNA has ribose sugar ring, with a hydroxyl group (OH) If RNA in B-from conformation there would be unfavorable steric contact between the hydroxyl group, base, and phosphate backbone. RNA structure
Reason: RNA has ribose sugar ring, with a hydroxyl group (OH) If RNA in B-from conformation there would be unfavorable steric contact between the hydroxyl group, base, and phosphate backbone. RNA structure
BCH222 - Greek Key β Barrels
 BCH222 - Greek Key β Barrels Reading C.I. Branden and J. Tooze (1999) Introduction to Protein Structure, Second Edition, pp. 77-78 & 335-336 (look at the color figures) J.S. Richardson (1981) "The Anatomy
BCH222 - Greek Key β Barrels Reading C.I. Branden and J. Tooze (1999) Introduction to Protein Structure, Second Edition, pp. 77-78 & 335-336 (look at the color figures) J.S. Richardson (1981) "The Anatomy
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs
 Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
CSE : Computational Issues in Molecular Biology. Lecture 19. Spring 2004
 CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
Canonical B-DNA CGCGTTGACAACTGCAGAATC GC AT CG TA AT GC TA TA CG AT 20 Å. Minor Groove 34 Å. Major Groove 3.4 Å. Strands are antiparallel
 DNA Canonical B-DNA 20 Å GC AT CG TA CGCGTTGACAACTGCAGAATC 34 Å AT GC TA Minor Groove 3.4 Å TA CG AT Major Groove Strands are antiparallel CG GC GC Canonical B DNA First determined experimentally by fiber
DNA Canonical B-DNA 20 Å GC AT CG TA CGCGTTGACAACTGCAGAATC 34 Å AT GC TA Minor Groove 3.4 Å TA CG AT Major Groove Strands are antiparallel CG GC GC Canonical B DNA First determined experimentally by fiber
Components of DNA. Components of DNA. Aim: What is the structure of DNA? February 15, DNA_Structure_2011.notebook. Do Now.
 Aim: What is the structure of DNA? Do Now: Explain the Hershey Chase experiment and what was its conclusion? Homework Read pp. 298 299 P.299 3,4,6.7 Do Now Paperclip Combos Material: 8 paperclips, 2 each
Aim: What is the structure of DNA? Do Now: Explain the Hershey Chase experiment and what was its conclusion? Homework Read pp. 298 299 P.299 3,4,6.7 Do Now Paperclip Combos Material: 8 paperclips, 2 each
Diversity in DNA recognition by p53 revealed by crystal structures with Hoogsteen base pairs
 SUPPLEMENTARY INFORMATION Diversity in DNA recognition by p53 revealed by crystal structures with Hoogsteen base pairs Malka Kitayner 1, 3, Haim Rozenberg 1, 3, Remo Rohs 2, 3, Oded Suad 1, Dov Rabinovich
SUPPLEMENTARY INFORMATION Diversity in DNA recognition by p53 revealed by crystal structures with Hoogsteen base pairs Malka Kitayner 1, 3, Haim Rozenberg 1, 3, Remo Rohs 2, 3, Oded Suad 1, Dov Rabinovich
Structure/function relationship in DNA-binding proteins
 PHRM 836 September 22, 2015 Structure/function relationship in DNA-binding proteins Devlin Chapter 8.8-9 u General description of transcription factors (TFs) u Sequence-specific interactions between DNA
PHRM 836 September 22, 2015 Structure/function relationship in DNA-binding proteins Devlin Chapter 8.8-9 u General description of transcription factors (TFs) u Sequence-specific interactions between DNA
MCB 110:Biochemistry of the Central Dogma of MB. MCB 110:Biochemistry of the Central Dogma of MB
 MCB 110:Biochemistry of the Central Dogma of MB Part 1. DNA replication, repair and genomics (Prof. Alber) Part 2. RNA & protein synthesis. Prof. Zhou Part 3. Membranes, protein secretion, trafficking
MCB 110:Biochemistry of the Central Dogma of MB Part 1. DNA replication, repair and genomics (Prof. Alber) Part 2. RNA & protein synthesis. Prof. Zhou Part 3. Membranes, protein secretion, trafficking
CMPS 6630: Introduction to Computational Biology and Bioinformatics. Secondary Structure Prediction
 CMPS 6630: Introduction to Computational Biology and Bioinformatics Secondary Structure Prediction Secondary Structure Annotation Given a macromolecular structure Identify the regions of secondary structure
CMPS 6630: Introduction to Computational Biology and Bioinformatics Secondary Structure Prediction Secondary Structure Annotation Given a macromolecular structure Identify the regions of secondary structure
Nucleic Acid Structure. Nucleic Acid Sequence Abbreviations. Sequence Abbreviations, con t.
 BC 4054 Spring 2001 Chapter 11 & 12 Review Lecture otes Slide 1 ucleic Acid Structure Linear polymer of nucleotides Phosphodiester linkage between 3 and 5 positions See Figure 11.17 Slide 2 ucleic Acid
BC 4054 Spring 2001 Chapter 11 & 12 Review Lecture otes Slide 1 ucleic Acid Structure Linear polymer of nucleotides Phosphodiester linkage between 3 and 5 positions See Figure 11.17 Slide 2 ucleic Acid
All Rights Reserved. U.S. Patents 6,471,520B1; 5,498,190; 5,916, North Market Street, Suite CC130A, Milwaukee, WI 53202
 Secondary Structure In the previous protein folding activity, you created a hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously folds
Secondary Structure In the previous protein folding activity, you created a hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously folds
BETA STRAND Prof. Alejandro Hochkoeppler Department of Pharmaceutical Sciences and Biotechnology University of Bologna
 Prof. Alejandro Hochkoeppler Department of Pharmaceutical Sciences and Biotechnology University of Bologna E-mail: a.hochkoeppler@unibo.it C-ter NH and CO groups: right, left, right (plane of the slide)
Prof. Alejandro Hochkoeppler Department of Pharmaceutical Sciences and Biotechnology University of Bologna E-mail: a.hochkoeppler@unibo.it C-ter NH and CO groups: right, left, right (plane of the slide)
Chapter 5: Nucleic Acids, etc.
 Chapter 5: Nucleic Acids, etc. Voet & Voet: Sections 1 & 3 Pages 82-84 & 88-93 Any introductory Biochemistry textbook will have an introductory chapter on nucleic acids Slide 1 Nucleotides and Derivatives
Chapter 5: Nucleic Acids, etc. Voet & Voet: Sections 1 & 3 Pages 82-84 & 88-93 Any introductory Biochemistry textbook will have an introductory chapter on nucleic acids Slide 1 Nucleotides and Derivatives
Sequence Analysis '17 -- lecture Secondary structure 3. Sequence similarity and homology 2. Secondary structure prediction
 Sequence Analysis '17 -- lecture 16 1. Secondary structure 3. Sequence similarity and homology 2. Secondary structure prediction Alpha helix Right-handed helix. H-bond is from the oxygen at i to the nitrogen
Sequence Analysis '17 -- lecture 16 1. Secondary structure 3. Sequence similarity and homology 2. Secondary structure prediction Alpha helix Right-handed helix. H-bond is from the oxygen at i to the nitrogen
Protein Folding Problem I400: Introduction to Bioinformatics
 Protein Folding Problem I400: Introduction to Bioinformatics November 29, 2004 Protein biomolecule, macromolecule more than 50% of the dry weight of cells is proteins polymer of amino acids connected into
Protein Folding Problem I400: Introduction to Bioinformatics November 29, 2004 Protein biomolecule, macromolecule more than 50% of the dry weight of cells is proteins polymer of amino acids connected into
STRUCTURAL BIOLOGY. α/β structures Closed barrels Open twisted sheets Horseshoe folds
 STRUCTURAL BIOLOGY α/β structures Closed barrels Open twisted sheets Horseshoe folds The α/β domains Most frequent domain structures are α/β domains: A central parallel or mixed β sheet Surrounded by α
STRUCTURAL BIOLOGY α/β structures Closed barrels Open twisted sheets Horseshoe folds The α/β domains Most frequent domain structures are α/β domains: A central parallel or mixed β sheet Surrounded by α
Name 10 Molecular Biology of the Gene Test Date Study Guide You must know: The structure of DNA. The major steps to replication.
 Name 10 Molecular Biology of the Gene Test Date Study Guide You must know: The structure of DNA. The major steps to replication. The difference between replication, transcription, and translation. How
Name 10 Molecular Biology of the Gene Test Date Study Guide You must know: The structure of DNA. The major steps to replication. The difference between replication, transcription, and translation. How
Globins. The Backbone structure of Myoglobin 2. The Heme complex in myoglobin. Lecture 10/01/2009. Role of the Globin.
 Globins Lecture 10/01/009 The Backbone structure of Myoglobin Myoglobin: 44 x 44 x 5 Å single subunit 153 amino acid residues 11 residues are in an a helix. Helices are named A, B, C, F. The heme pocket
Globins Lecture 10/01/009 The Backbone structure of Myoglobin Myoglobin: 44 x 44 x 5 Å single subunit 153 amino acid residues 11 residues are in an a helix. Helices are named A, B, C, F. The heme pocket
Chapter Fundamental Molecular Genetic Mechanisms
 Chapter 5-1 - Fundamental Molecular Genetic Mechanisms 5.1 Structure of Nucleic Acids 5.2 Transcription of Protein-Coding Genes and Formation of Functional mrna 5.3 The Decoding of mrna by trnas 5.4 Stepwise
Chapter 5-1 - Fundamental Molecular Genetic Mechanisms 5.1 Structure of Nucleic Acids 5.2 Transcription of Protein-Coding Genes and Formation of Functional mrna 5.3 The Decoding of mrna by trnas 5.4 Stepwise
Packing of Secondary Structures
 7.88 Lecture Notes - 5 7.24/7.88J/5.48J The Protein Folding and Human Disease Packing of Secondary Structures Packing of Helices against sheets Packing of sheets against sheets Parallel Orthogonal Table:
7.88 Lecture Notes - 5 7.24/7.88J/5.48J The Protein Folding and Human Disease Packing of Secondary Structures Packing of Helices against sheets Packing of sheets against sheets Parallel Orthogonal Table:
Understanding DNA Structure
 Understanding DNA Structure I619 Structural Bioinformatics Molecular Biology Basics + Scale total length of DNA in a human cell is about 2m DNA is compacted in length by a factor of 10000 the compaction
Understanding DNA Structure I619 Structural Bioinformatics Molecular Biology Basics + Scale total length of DNA in a human cell is about 2m DNA is compacted in length by a factor of 10000 the compaction
Specificity: Induced Fit
 Specificity: Induced Fit Conformational changes may occur upon ligand binding (Daniel Koshland in 1958) This adaptation is called the induced fit Induced fit allows for tighter binding of the ligand Induced
Specificity: Induced Fit Conformational changes may occur upon ligand binding (Daniel Koshland in 1958) This adaptation is called the induced fit Induced fit allows for tighter binding of the ligand Induced
Bi 8 Lecture 7. Ellen Rothenberg 26 January Reading: Ch. 3, pp ; panel 3-1
 Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
BIOLOGICAL SCIENCE. Lecture Presentation by Cindy S. Malone, PhD, California State University Northridge. FIFTH EDITION Freeman Quillin Allison
 BIOLOGICAL SCIENCE FIFTH EDITION Freeman Quillin Allison 4 Lecture Presentation by Cindy S. Malone, PhD, California State University Northridge In this chapter you will learn that Nucleic acids store the
BIOLOGICAL SCIENCE FIFTH EDITION Freeman Quillin Allison 4 Lecture Presentation by Cindy S. Malone, PhD, California State University Northridge In this chapter you will learn that Nucleic acids store the
RNA Part I: Chemical Structure of RNA
 RA Part I: Chemical Structure of RA Structural differences between RA and DA Resistance of phosphate esters to basic hydrolysis The 2 - group of RA facilitates chemical cleavage in aqueous a by forming
RA Part I: Chemical Structure of RA Structural differences between RA and DA Resistance of phosphate esters to basic hydrolysis The 2 - group of RA facilitates chemical cleavage in aqueous a by forming
Paper : 03 Structure and Function of Biomolecules II Module: 02 Nucleosides, Nucleotides and type of Nucleic Acids
 Paper : 03 Structure and Function of Biomolecules II Module: 02 Nucleosides, Nucleotides and type of Nucleic Acids Principal Investigator Paper Coordinators Prof. Sunil Kumar Khare, Professor, Department
Paper : 03 Structure and Function of Biomolecules II Module: 02 Nucleosides, Nucleotides and type of Nucleic Acids Principal Investigator Paper Coordinators Prof. Sunil Kumar Khare, Professor, Department
Molecular Genetics Quiz #1 SBI4U K T/I A C TOTAL
 Name: Molecular Genetics Quiz #1 SBI4U K T/I A C TOTAL Part A: Multiple Choice (15 marks) Circle the letter of choice that best completes the statement or answers the question. One mark for each correct
Name: Molecular Genetics Quiz #1 SBI4U K T/I A C TOTAL Part A: Multiple Choice (15 marks) Circle the letter of choice that best completes the statement or answers the question. One mark for each correct
Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a
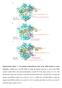 Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a HNH-state 1 in PDB 4OO8, in which the distance from the C atom of the HNH catalytic residue 840 to the
Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a HNH-state 1 in PDB 4OO8, in which the distance from the C atom of the HNH catalytic residue 840 to the
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
PrimePCR Assay Validation Report
 Gene Information Gene Name SRY (sex determining region Y)-box 6 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID SOX6 Human This gene encodes a member of the
Gene Information Gene Name SRY (sex determining region Y)-box 6 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID SOX6 Human This gene encodes a member of the
BIRKBECK COLLEGE (University of London)
 BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
Part IV => DNA and RNA. 4.1 Nucleotide Properties 4.1a Nucleotide Nomenclature 4.1b DNA Sequencing
 Part IV => DNA and RNA 4.1 Nucleotide Properties 4.1a Nucleotide Nomenclature 4.1b DNA Sequencing The Molecular Dogma of Life DNA carries genetic information in the form of its sequence of nucleotides
Part IV => DNA and RNA 4.1 Nucleotide Properties 4.1a Nucleotide Nomenclature 4.1b DNA Sequencing The Molecular Dogma of Life DNA carries genetic information in the form of its sequence of nucleotides
What Are the Chemical Structures and Functions of Nucleic Acids?
 THE NUCLEIC ACIDS What Are the Chemical Structures and Functions of Nucleic Acids? Nucleic acids are polymers specialized for the storage, transmission, and use of genetic information. DNA = deoxyribonucleic
THE NUCLEIC ACIDS What Are the Chemical Structures and Functions of Nucleic Acids? Nucleic acids are polymers specialized for the storage, transmission, and use of genetic information. DNA = deoxyribonucleic
Proteins Higher Order Structures
 Proteins Higher Order Structures Dr. Mohammad Alsenaidy Department of Pharmaceutics College of Pharmacy King Saud University Office: AA 101 msenaidy@ksu.edu.sa Previously on PHT 426!! Protein Structures
Proteins Higher Order Structures Dr. Mohammad Alsenaidy Department of Pharmaceutics College of Pharmacy King Saud University Office: AA 101 msenaidy@ksu.edu.sa Previously on PHT 426!! Protein Structures
The Double Helix. DNA and RNA, part 2. Part A. Hint 1. The difference between purines and pyrimidines. Hint 2. Distinguish purines from pyrimidines
 DNA and RNA, part 2 Due: 3:00pm on Wednesday, September 24, 2014 You will receive no credit for items you complete after the assignment is due. Grading Policy The Double Helix DNA, or deoxyribonucleic
DNA and RNA, part 2 Due: 3:00pm on Wednesday, September 24, 2014 You will receive no credit for items you complete after the assignment is due. Grading Policy The Double Helix DNA, or deoxyribonucleic
Nucleotides: structure and functions. Prof. Dalė Vieželienė Biochemistry department Room No
 Nucleotides: structure and functions Prof. Dalė Vieželienė Biochemistry department Room No. 229 Email: daleveze@med.kmu.lt Composition of Nucleic Acids Nucleotide structure Two types of nucleic acids:
Nucleotides: structure and functions Prof. Dalė Vieželienė Biochemistry department Room No. 229 Email: daleveze@med.kmu.lt Composition of Nucleic Acids Nucleotide structure Two types of nucleic acids:
Nucleic Acids, Proteins, and Enzymes
 3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
Protein Synthesis Notes
 Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
Answers to Module 1. An obligate aerobe is an organism that has an absolute requirement of oxygen for growth.
 Answers to Module 1 Short Answers 1) What is an obligate aerobe? An obligate aerobe is an organism that has an absolute requirement of oxygen for growth. What about facultative anaerobe? 2) Distinguish
Answers to Module 1 Short Answers 1) What is an obligate aerobe? An obligate aerobe is an organism that has an absolute requirement of oxygen for growth. What about facultative anaerobe? 2) Distinguish
Super Models. Nucleotides Molecular Model Kit Copyright 2015 Ryler Enterprises, Inc. Recommended for ages 10 - adult
 Super Models! ucleotides Molecular Model Kit Copyright 015 Ryler Enterprises, Inc. Recommended for ages 10 - adult Caution: Atom centers and vinyl tubing are a choking hazard. Do not eat or chew model
Super Models! ucleotides Molecular Model Kit Copyright 015 Ryler Enterprises, Inc. Recommended for ages 10 - adult Caution: Atom centers and vinyl tubing are a choking hazard. Do not eat or chew model
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
38. Inter-basepair Hydrogen Bonds in DNA
 190 Proc. Japan Acad., 70, Ser. B (1994) [Vol. 70(B), 38. Inter-basepair Hydrogen Bonds in DNA By Masashi SUZUKI*),t) and Naoto YAGI**) (Communicated by Setsuro EBASHI, M. J. A., Dec. 12, 1994) Abstract:
190 Proc. Japan Acad., 70, Ser. B (1994) [Vol. 70(B), 38. Inter-basepair Hydrogen Bonds in DNA By Masashi SUZUKI*),t) and Naoto YAGI**) (Communicated by Setsuro EBASHI, M. J. A., Dec. 12, 1994) Abstract:
Nature Structural & Molecular Biology: doi: /nsmb.3175
 Supplementary Figure 1 Medium- and low-fret groups probably representing varying positions on the DNA show similar changes between H3 and CENP-A nucleosomes with and without CENP-C CD, as does the high-fret
Supplementary Figure 1 Medium- and low-fret groups probably representing varying positions on the DNA show similar changes between H3 and CENP-A nucleosomes with and without CENP-C CD, as does the high-fret
Protocol S1: Supporting Information
 Protocol S1: Supporting Information Basis for the specificity of the kinase domain of Abl for peptide substrates The crystal structures reported in this work were obtained using two different ATP analog-peptide
Protocol S1: Supporting Information Basis for the specificity of the kinase domain of Abl for peptide substrates The crystal structures reported in this work were obtained using two different ATP analog-peptide
MOLECULAR STRUCTURE OF DNA
 MOLECULAR STRUCTURE OF DNA Characteristics of the Genetic Material 1. Replication Reproduced and transmitted faithfully from cell to cell (generation to generation) 2. Information Storage Biologically
MOLECULAR STRUCTURE OF DNA Characteristics of the Genetic Material 1. Replication Reproduced and transmitted faithfully from cell to cell (generation to generation) 2. Information Storage Biologically
NUCLEIC ACIDS Genetic material of all known organisms DNA: deoxyribonucleic acid RNA: ribonucleic acid (e.g., some viruses)
 NUCLEIC ACIDS Genetic material of all known organisms DNA: deoxyribonucleic acid RNA: ribonucleic acid (e.g., some viruses) Consist of chemically linked sequences of nucleotides Nitrogenous base Pentose-
NUCLEIC ACIDS Genetic material of all known organisms DNA: deoxyribonucleic acid RNA: ribonucleic acid (e.g., some viruses) Consist of chemically linked sequences of nucleotides Nitrogenous base Pentose-
Chapter 12 Packet DNA 1. What did Griffith conclude from his experiment? 2. Describe the process of transformation.
 Chapter 12 Packet DNA and RNA Name Period California State Standards covered by this chapter: Cell Biology 1. The fundamental life processes of plants and animals depend on a variety of chemical reactions
Chapter 12 Packet DNA and RNA Name Period California State Standards covered by this chapter: Cell Biology 1. The fundamental life processes of plants and animals depend on a variety of chemical reactions
Retrieving and Viewing Protein Structures from the Protein Data Base
 Retrieving and Viewing Protein Structures from the Protein Data Base 7.88J Protein Folding Prof. David Gossard September 15, 2003 1 PDB Acknowledgements The Protein Data Bank (PDB - http://www.pdb.org/)
Retrieving and Viewing Protein Structures from the Protein Data Base 7.88J Protein Folding Prof. David Gossard September 15, 2003 1 PDB Acknowledgements The Protein Data Bank (PDB - http://www.pdb.org/)
Final exam. Please write your name on the exam and keep an ID card ready.
 Biophysics of Macromolecules Prof. R. Jungmann and Prof. J. Lipfert SS 2017 Final exam Final exam First name: Last name: Student number ( Matrikelnummer ): Please write your name on the exam and keep an
Biophysics of Macromolecules Prof. R. Jungmann and Prof. J. Lipfert SS 2017 Final exam Final exam First name: Last name: Student number ( Matrikelnummer ): Please write your name on the exam and keep an
Structure formation and association of biomolecules. Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München
 Structure formation and association of biomolecules Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München Motivation Many biomolecules are chemically synthesized
Structure formation and association of biomolecules Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München Motivation Many biomolecules are chemically synthesized
Gene Expression - Transcription
 DNA Gene Expression - Transcription Genes are expressed as encoded proteins in a 2 step process: transcription + translation Central dogma of biology: DNA RNA protein Transcription: copy DNA strand making
DNA Gene Expression - Transcription Genes are expressed as encoded proteins in a 2 step process: transcription + translation Central dogma of biology: DNA RNA protein Transcription: copy DNA strand making
Syllabus for GUTS Lecture on DNA and Nucleotides
 Syllabus for GUTS Lecture on DNA and Nucleotides I. Introduction. DNA is the instruction manual for how to build a living organism here on earth. The instructions in DNA are propagated to future generations
Syllabus for GUTS Lecture on DNA and Nucleotides I. Introduction. DNA is the instruction manual for how to build a living organism here on earth. The instructions in DNA are propagated to future generations
Lesson Overview DNA Replication
 12.3 THINK ABOUT IT Before a cell divides, its DNA must first be copied. How might the double-helix structure of DNA make that possible? Review Question! At what stage of the cell cycle do cells duplicate
12.3 THINK ABOUT IT Before a cell divides, its DNA must first be copied. How might the double-helix structure of DNA make that possible? Review Question! At what stage of the cell cycle do cells duplicate
The Genetic Code and Transcription. Chapter 12 Honors Genetics Ms. Susan Chabot
 The Genetic Code and Transcription Chapter 12 Honors Genetics Ms. Susan Chabot TRANSCRIPTION Copy SAME language DNA to RNA Nucleic Acid to Nucleic Acid TRANSLATION Copy DIFFERENT language RNA to Amino
The Genetic Code and Transcription Chapter 12 Honors Genetics Ms. Susan Chabot TRANSCRIPTION Copy SAME language DNA to RNA Nucleic Acid to Nucleic Acid TRANSLATION Copy DIFFERENT language RNA to Amino
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than. SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with
 Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Ch Molecular Biology of the Gene
 Ch. 12 - Molecular Biology of the Gene AP BIOLOGY CHAPTER GUIDE 1. In the middle of the unraveling the mysteries of DNA, researchers knew that genetic material must be able to. It must be stable so it
Ch. 12 - Molecular Biology of the Gene AP BIOLOGY CHAPTER GUIDE 1. In the middle of the unraveling the mysteries of DNA, researchers knew that genetic material must be able to. It must be stable so it
DNA AND CHROMOSOMES. Genetica per Scienze Naturali a.a prof S. Presciuttini
 DNA AND CHROMOSOMES This document is licensed under the Attribution-NonCommercial-ShareAlike 2.5 Italy license, available at http://creativecommons.org/licenses/by-nc-sa/2.5/it/ 1. The Building Blocks
DNA AND CHROMOSOMES This document is licensed under the Attribution-NonCommercial-ShareAlike 2.5 Italy license, available at http://creativecommons.org/licenses/by-nc-sa/2.5/it/ 1. The Building Blocks
Table S1. Crystallographic Data and Refinement Statistics, Related to Experimental Procedures hsddb1-drddb2 CPD
 Data collection Table S1. Crystallographic Data and Refinement Statistics, Related to Experimental Procedures hsddb1-drddb2 CPD #1 hsddb1-drddb2 CPD #2 Values in parentheses are for the highest resolution
Data collection Table S1. Crystallographic Data and Refinement Statistics, Related to Experimental Procedures hsddb1-drddb2 CPD #1 hsddb1-drddb2 CPD #2 Values in parentheses are for the highest resolution
Chapter 1 Structure of Nucleic Acids DNA The structure of part of a DNA double helix
 Chapter 1 Structure of Nucleic Acids DNA The structure of part of a DNA double helix Deoxyribonucleic acid ) (DNA) is a nucleic acid that contains the genetic instructions used in the development and functioning
Chapter 1 Structure of Nucleic Acids DNA The structure of part of a DNA double helix Deoxyribonucleic acid ) (DNA) is a nucleic acid that contains the genetic instructions used in the development and functioning
Your Name: MID TERM ANSWER SHEET SIN: ( )
 MIDTERM EXAMINATION (October 23, 2008) BIOE150. Introduction to Bio-Nanoscience & Bio-Nanotechnology Professor Seung-Wuk Lee Fall Semester, 2008 0. Write down your name and the last digit of your SIN in
MIDTERM EXAMINATION (October 23, 2008) BIOE150. Introduction to Bio-Nanoscience & Bio-Nanotechnology Professor Seung-Wuk Lee Fall Semester, 2008 0. Write down your name and the last digit of your SIN in
Algorithms in Bioinformatics
 Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline Central Dogma of Molecular
Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline Central Dogma of Molecular
Fig Ch 17: From Gene to Protein
 Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Structural bioinformatics
 Structural bioinformatics Why structures? The representation of the molecules in 3D is more informative New properties of the molecules are revealed, which can not be detected by sequences Eran Eyal Plant
Structural bioinformatics Why structures? The representation of the molecules in 3D is more informative New properties of the molecules are revealed, which can not be detected by sequences Eran Eyal Plant
RNA synthesis/transcription I Biochemistry 302. February 6, 2004 Bob Kelm
 RNA synthesis/transcription I Biochemistry 302 February 6, 2004 Bob Kelm Overview of RNA classes Messenger RNA (mrna) Encodes protein Relatively short half-life ( 3 min in E. coli, 30 min in eukaryotic
RNA synthesis/transcription I Biochemistry 302 February 6, 2004 Bob Kelm Overview of RNA classes Messenger RNA (mrna) Encodes protein Relatively short half-life ( 3 min in E. coli, 30 min in eukaryotic
Nucleic Acids and the RNA World. Pages Chapter 4
 Nucleic Acids and the RNA World Pages 74-89 Chapter 4 RNA vs. Protein Chemical Evolution stated that life evolved from a polymer called a protein. HOWEVER, now many scientists question this. There is currently
Nucleic Acids and the RNA World Pages 74-89 Chapter 4 RNA vs. Protein Chemical Evolution stated that life evolved from a polymer called a protein. HOWEVER, now many scientists question this. There is currently
has only one nucleotide, U20, between G19 and A21, while trna Glu CUC has two
 SPPLEMENTRY INFORMTION doi:1.138/nature9411 Supplementary Discussion The structural characteristics of trn Gln G in comparison to trn Glu 16 in trn Gln G is directed towards the G19 56 pair, or the outer
SPPLEMENTRY INFORMTION doi:1.138/nature9411 Supplementary Discussion The structural characteristics of trn Gln G in comparison to trn Glu 16 in trn Gln G is directed towards the G19 56 pair, or the outer
C. Incorrect! Threonine is an amino acid, not a nucleotide base.
 MCAT Biology - Problem Drill 05: RNA and Protein Biosynthesis Question No. 1 of 10 1. Which of the following bases are only found in RNA? Question #01 (A) Ribose. (B) Uracil. (C) Threonine. (D) Adenine.
MCAT Biology - Problem Drill 05: RNA and Protein Biosynthesis Question No. 1 of 10 1. Which of the following bases are only found in RNA? Question #01 (A) Ribose. (B) Uracil. (C) Threonine. (D) Adenine.
5. Structure and Replication of DNA
 BIO2310 General and Molecular Genetics 5. Structure and Replication of DNA Key questions: How was DNA shown to be the genetic material? How about RNA? How was the structure of DNA determined to be the
BIO2310 General and Molecular Genetics 5. Structure and Replication of DNA Key questions: How was DNA shown to be the genetic material? How about RNA? How was the structure of DNA determined to be the
Study Guide for Chapter 12 Exam DNA, RNA, & Protein Synthesis
 Name: Date: Period: Study Guide for Chapter 12 Exam DNA, RNA, & Protein Synthesis ***Completing this study guide in its entirety will result in extra credit on the exam. You must show me the DAY OF the
Name: Date: Period: Study Guide for Chapter 12 Exam DNA, RNA, & Protein Synthesis ***Completing this study guide in its entirety will result in extra credit on the exam. You must show me the DAY OF the
TERTIARY MOTIF INTERACTIONS ON RNA STRUCTURE
 1 TERTIARY MOTIF INTERACTIONS ON RNA STRUCTURE Bioinformatics Senior Project Wasay Hussain Spring 2009 Overview of RNA 2 The central Dogma of Molecular biology is DNA RNA Proteins The RNA (Ribonucleic
1 TERTIARY MOTIF INTERACTIONS ON RNA STRUCTURE Bioinformatics Senior Project Wasay Hussain Spring 2009 Overview of RNA 2 The central Dogma of Molecular biology is DNA RNA Proteins The RNA (Ribonucleic
MBMB451A Section1 Fall 2008 KEY These questions may have more than one correct answer
 MBMB451A Section1 Fall 2008 KEY These questions may have more than one correct answer 1. In a double stranded molecule of DNA, the ratio of purines : pyrimidines is (a) variable (b) determined by the base
MBMB451A Section1 Fall 2008 KEY These questions may have more than one correct answer 1. In a double stranded molecule of DNA, the ratio of purines : pyrimidines is (a) variable (b) determined by the base
Prot-SSP: A Tool for Amino Acid Pairing Pattern Analysis in Secondary Structures
 Mol2Net, 2015, 1(Section F), pages 1-6, Proceedings 1 SciForum Mol2Net Prot-SSP: A Tool for Amino Acid Pairing Pattern Analysis in Secondary Structures Miguel de Sousa 1, *, Cristian R. Munteanu 2 and
Mol2Net, 2015, 1(Section F), pages 1-6, Proceedings 1 SciForum Mol2Net Prot-SSP: A Tool for Amino Acid Pairing Pattern Analysis in Secondary Structures Miguel de Sousa 1, *, Cristian R. Munteanu 2 and
Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators
 Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators At least 5 potential gene expression control points Superfamily of Gene Regulators Activation of gene structure Initiation
Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators At least 5 potential gene expression control points Superfamily of Gene Regulators Activation of gene structure Initiation
Lecture 2: Central Dogma of Molecular Biology & Intro to Programming
 Lecture 2: Central Dogma of Molecular Biology & Intro to Programming Central Dogma of Molecular Biology Proteins: workhorse molecules of biological systems Proteins are synthesized from the genetic blueprints
Lecture 2: Central Dogma of Molecular Biology & Intro to Programming Central Dogma of Molecular Biology Proteins: workhorse molecules of biological systems Proteins are synthesized from the genetic blueprints
Chapter 13 Section 2: DNA Replication
 Chapter 13 Section 2: DNA Replication Opening Activity DNA is considered to be a relatively stable molecule. What gives it this stability, even though the hydrogen bonds between the nitrogen bases are
Chapter 13 Section 2: DNA Replication Opening Activity DNA is considered to be a relatively stable molecule. What gives it this stability, even though the hydrogen bonds between the nitrogen bases are
Problem Set 8. Answer Key
 MCB 102 University of California, Berkeley August 11, 2009 Isabelle Philipp Online Document Problem Set 8 Answer Key 1. The Genetic Code (a) Are all amino acids encoded by the same number of codons? no
MCB 102 University of California, Berkeley August 11, 2009 Isabelle Philipp Online Document Problem Set 8 Answer Key 1. The Genetic Code (a) Are all amino acids encoded by the same number of codons? no
Genes - DNA - Chromosome. Chutima Talabnin Ph.D. School of Biochemistry,Institute of Science, Suranaree University of Technology
 Genes - DNA - Chromosome Chutima Talabnin Ph.D. School of Biochemistry,Institute of Science, Suranaree University of Technology DNA Cellular DNA contains genes and intragenic regions both of which may
Genes - DNA - Chromosome Chutima Talabnin Ph.D. School of Biochemistry,Institute of Science, Suranaree University of Technology DNA Cellular DNA contains genes and intragenic regions both of which may
DNA Transcription. Visualizing Transcription. The Transcription Process
 DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
Introduction to Proteins
 Introduction to Proteins Lecture 4 Module I: Molecular Structure & Metabolism Molecular Cell Biology Core Course (GSND5200) Matthew Neiditch - Room E450U ICPH matthew.neiditch@umdnj.edu What is a protein?
Introduction to Proteins Lecture 4 Module I: Molecular Structure & Metabolism Molecular Cell Biology Core Course (GSND5200) Matthew Neiditch - Room E450U ICPH matthew.neiditch@umdnj.edu What is a protein?
Regulation of gene expression. (Lehninger pg )
 Regulation of gene expression (Lehninger pg. 1072-1085) Today s lecture Gene expression Constitutive, inducible, repressible genes Specificity factors, activators, repressors Negative and positive gene
Regulation of gene expression (Lehninger pg. 1072-1085) Today s lecture Gene expression Constitutive, inducible, repressible genes Specificity factors, activators, repressors Negative and positive gene
Protein Structure Prediction. christian studer , EPFL
 Protein Structure Prediction christian studer 17.11.2004, EPFL Content Definition of the problem Possible approaches DSSP / PSI-BLAST Generalization Results Definition of the problem Massive amounts of
Protein Structure Prediction christian studer 17.11.2004, EPFL Content Definition of the problem Possible approaches DSSP / PSI-BLAST Generalization Results Definition of the problem Massive amounts of
RNA is a single strand molecule composed of subunits called nucleotides joined by phosphodiester bonds.
 The Versatility of RNA Primary structure of RNA RNA is a single strand molecule composed of subunits called nucleotides joined by phosphodiester bonds. Each nucleotide subunit is composed of a ribose sugar,
The Versatility of RNA Primary structure of RNA RNA is a single strand molecule composed of subunits called nucleotides joined by phosphodiester bonds. Each nucleotide subunit is composed of a ribose sugar,
Protein-Protein Interactions I
 Biochemistry 412 Protein-Protein Interactions I March 11, 2008 Macromolecular Recognition by Proteins Protein folding is a process governed by intramolecular recognition. Protein-protein association is
Biochemistry 412 Protein-Protein Interactions I March 11, 2008 Macromolecular Recognition by Proteins Protein folding is a process governed by intramolecular recognition. Protein-protein association is
KEY CONCEPT DNA was identified as the genetic material through a series of experiments. Found live S with R bacteria and injected
 Section 1: Identifying DNA as the Genetic Material KEY CONCEPT DNA was identified as the genetic material through a series of experiments. VOCABULARY bacteriophage MAIN IDEA: Griffith finds a transforming
Section 1: Identifying DNA as the Genetic Material KEY CONCEPT DNA was identified as the genetic material through a series of experiments. VOCABULARY bacteriophage MAIN IDEA: Griffith finds a transforming
Unit Description: The unit on DNA replication will include the following activities:
 Contact Information Retha Prescod Title: Viruses Not Welcome Abstract: The author proposes that an initial virtual exploration on DNA replication will serve as an introduction to a difficult yet integral
Contact Information Retha Prescod Title: Viruses Not Welcome Abstract: The author proposes that an initial virtual exploration on DNA replication will serve as an introduction to a difficult yet integral
Molecular Biology I. The Chemical Nature of DNA. Dr. Obaidur rahman
 Molecular Biology I The Chemical Nature of DNA Dr. Obaidur rahman Characteristics of Genetic Material The coding instructions of all living organisms are written in the same genetic language that of nucleic
Molecular Biology I The Chemical Nature of DNA Dr. Obaidur rahman Characteristics of Genetic Material The coding instructions of all living organisms are written in the same genetic language that of nucleic
6-Foot Mini Toober Activity
 Big Idea The interaction between the substrate and enzyme is highly specific. Even a slight change in shape of either the substrate or the enzyme may alter the efficient and selective ability of the enzyme
Big Idea The interaction between the substrate and enzyme is highly specific. Even a slight change in shape of either the substrate or the enzyme may alter the efficient and selective ability of the enzyme
RNA Structure Prediction and Comparison. RNA Biology Background
 RN Structure Prediction and omparison Session 1 RN Biology Background Robert iegerich Faculty of Technology robert@techfak.ni-bielefeld.de October 13, 2013 Robert iegerich Overview of lecture topics The
RN Structure Prediction and omparison Session 1 RN Biology Background Robert iegerich Faculty of Technology robert@techfak.ni-bielefeld.de October 13, 2013 Robert iegerich Overview of lecture topics The
Solution Structure of the DNA-binding Domain of GAL4 from Saccharomyces cerevisiae
 Vol. 14, No. 1-4 175 Solution Structure of the DNA-binding Domain of GAL4 from Saccharomyces cerevisiae James D. Baleja, V. Thanabal, Ted Mau, and Gerhard Wagner Department of Biological Chemistry and
Vol. 14, No. 1-4 175 Solution Structure of the DNA-binding Domain of GAL4 from Saccharomyces cerevisiae James D. Baleja, V. Thanabal, Ted Mau, and Gerhard Wagner Department of Biological Chemistry and
Spring 2006 Biochemistry 302 Exam 1
 1 Name Spring 2006 Biochemistry 302 Exam 1 Directions: This exam has 36 questions/problems totaling 90 points. Check to make sure you have all six pages. Some questions have multiple parts so read each
1 Name Spring 2006 Biochemistry 302 Exam 1 Directions: This exam has 36 questions/problems totaling 90 points. Check to make sure you have all six pages. Some questions have multiple parts so read each
Lectures 9 and 10: Random Walks and the Structure of Macromolecules (contd.)
 Lectures 9 and 10: Random Walks and the Structure of Macromolecules (contd.) Lecturer: Brigita Urbanc Office: 12 909 (E mail: brigita@drexel.edu) Course website: www.physics.drexel.edu/~brigita/courses/biophys_2011
Lectures 9 and 10: Random Walks and the Structure of Macromolecules (contd.) Lecturer: Brigita Urbanc Office: 12 909 (E mail: brigita@drexel.edu) Course website: www.physics.drexel.edu/~brigita/courses/biophys_2011
Transcription in Eukaryotes
 Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
CHEM 4420 Exam I Spring 2013 Page 1 of 6
 CHEM 4420 Exam I Spring 2013 Page 1 of 6 Name Use complete sentences when requested. There are 100 possible points on this exam. The multiple choice questions are worth 2 points each. All other questions
CHEM 4420 Exam I Spring 2013 Page 1 of 6 Name Use complete sentences when requested. There are 100 possible points on this exam. The multiple choice questions are worth 2 points each. All other questions
Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and
 Supplementary Tables Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and their side chain contacting residues in the second chain (B) a Interface Res. in Contacting
Supplementary Tables Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and their side chain contacting residues in the second chain (B) a Interface Res. in Contacting
