Specificity: Induced Fit
|
|
|
- Horatio Barker
- 6 years ago
- Views:
Transcription
1 Specificity: Induced Fit Conformational changes may occur upon ligand binding (Daniel Koshland in 1958) This adaptation is called the induced fit Induced fit allows for tighter binding of the ligand Induced fit allows for high affinity for different ligands Both the ligand and the protein can change their conformations +
2 Recitation Chapter 5
3 Example: Oxygen Binding to Myoglobin When ligand is a gas, binding is expressed in terms of partial pressures. [L] Kd [L] p 50 po2 po 2
4 Globins are oxygen-binding proteins Protein side chains lack affinity for O 2 Some transition metals bind O 2 well but would generate free radicals if free in solution Organometallic compounds such as heme are more suitable, but Fe 2+ in free heme could be oxidized to Fe 3+ Solution Capture the oxygen molecule with heme that is protein bound Myoglobin is the main oxygen storage protein Hemoglobin is a circulating oxygen-binding protein
5 Structures of Porphyrin and Heme Heme. The heme group is present in myoglobin, hemoglobin, and many other proteins, designated heme proteins. Heme consists of a complex organic ring structure, protoporphyrin IX, with a bound iron atom in its ferrous (Fe 2+ ) state. (a) Porphyrins, of which protoporphyrin IX is only one example, consist of four pyrrole rings linked by methene bridges, with substitutions at one or more of the positions denoted X. (b, c) Two representations of heme (derived from PDB ID 1CCR). The iron atom of heme has six coordination bonds: four in the plane of, and bonded to, the flat porphyrin ring system, and (d) two perpendicular to it.
6 Structure of Myoglobin. The eight α-helical segments (shown here as cylinders) are labeled A through H. Nonhelical residues in the bends that connect them are labeled AB, CD, EF, and so forth, indicating the segments they interconnect. A few bends, including BC and DE, are abrupt and do not contain any residues; these are not normally labeled. (The short segment visible between D and E is an artifact of the computer representation.) The heme is bound in a pocket made up largely of the E and F helices, although amino acid residues fromother segments of the protein also participate. The heme group viewed from the side. This view shows the two coordination bonds to Fe 2+ that are perpendicular to the porphyrin ring system. One is occupied by a His residue, sometimes called the proximal His; the other is the binding site for oxygen. The remaining four coordination bonds are in the plane of, and bonded to, the flat porphyrin ring system.
7 Heme binding to protein affects O 2 binding. Steric effects caused by ligand binding to the heme of myoglobin. (c) Another view of the heme of myoglobin, showing the arrangement of key amino acid residues around the heme. The bound O 2 is hydrogen-bonded to the distal His, His E7 (His 64 ), further facilitating the binding of O 2.
8 Could myoglobin transport O 2? po 2 in lungs is about 13 kpa: it sure binds oxygen well po 2 in tissues is about 4 kpa: it will not release it! Would lowering the affinity (P 50 ) of myoglobin to oxygen help? Graphical representations of ligand binding. The fraction of ligandbinding sites occupied, θ, is plotted against the concentration of free ligand. Both curves are rectangular hyperbolas. (b) A curve describing the binding of oxygen to myoglobin. The partial pressure of O 2 in the air above the solution is expressed in kilopascals (kpa). Oxygen binds tightly to myoglobin, with a P 50 of only 0.26 kpa.
9 The Hill Plot of Cooperativity Hill plots for oxygen binding to myoglobin and hemoglobin. When n H 5 1, there is no evident cooperativity. The maximum degree of cooperativity observed for hemoglobin corresponds approximately to n H 5 3. Note that while this indicates a high level of cooperativity, n H is less than n, the number of O 2 -binding sites in hemoglobin. This is normal for a protein that exhibits allosteric binding behavior.
10 Subunit Interactions in Hb Dominant interactions between hemoglobin subunits. In this representation, α subunits are light and β subunits are dark. The strongest subunit interactions (highlighted) occur between unlike subunits. When oxygen binds, the α 1 β 1 contact changes little, but there is a large change at the α 1 β 2 contact, with several ion pairs broken.
11 Subunit Interactions: Details; Some ion pairs that stabilize the T state of deoxyhemoglobin. (b) Interactions between these ion pairs, and between others not shown in (a), are schematized in this representation of the extended polypeptide chains of hemoglobin.
12 R and T States of Hb T = Tense state, More interactions, more stable Lower affinity for O 2 R = Relaxed state, Fewer Interactions, more flexible Higher affinity for O 2 O 2 binding triggers a T R conformational change Conformational change from the T state to the R state involves breaking ion pairs between the α1-2 interface
13 R and T States of Hemoglobin The T R transition. In these depictions of deoxyhemoglobin, as in Figure 5 9, the β subunits are blue and the α subunits are gray. Positively charged side chains and chain termini involved in ion pairs are shown in blue, their negatively charged partners in red. The Lys C5 of each α subunit and Asp FG1 of each β subunit are visible but not labeled. Note that the molecule is oriented slightly differently than in Figure 5 9. The transition from the T state to the R state shifts the subunit pairs substantially, affecting certain ion pairs. Most noticeably, the His HC3 residues at the carboxyl termini of the β subunits, which are involved in ion pairs in the T state, rotate in the R state toward the center of the molecule, where they are no longer in ion pairs. Another dramatic result of the T R transition is a narrowing of the pocket between the β subunits.
14 2,3-BPG binds to the central cavity of Hb Binding of BPG to deoxyhemoglobin. (a) BPG binding stabilizes the T state of deoxyhemoglobin. The negativecharges of BPG interact with several positively charged groups (shown in blue in this surface contour image) that surround the pocket between the β subunits on the surface of deoxyhemoglobin in the T state. (b) The binding pocket for BPG disappears on oxygenation, following transition to the R state. (Compare Fig )
15 Antibodies: Immunoglobulin G Immunoglobulin G. (a) Pairs of heavy and light chains combine to form a Y-shaped molecule. Two antigenbinding sites are formed by the combination of variable domains from one light (V L ) and one heavy (V H ) chain. Cleavage with papain separates the Fab and Fc portions of the protein in the hinge region. The Fc portion of the molecule also contains bound carbohydrate (shown in (b)).
16
17 Antibodies: Immunoglobulin G A ribbon model of the first complete IgG molecule to be crystallized and structurally analyzed. Although the molecule has two identical heavy chains (two shades of blue) and two identical light chains (two shades of red), it crystallized in the asymmetric conformation shown here. Conformational flexibility may be important to the function of immunoglobulins.
18 Antigens bind via induced fit; Antigen binding causes significant structural changes to the antibody Induced fit in the binding of an antigen to IgG. The molecule here, shown in surface contour, is the Fab fragment of an IgG. The antigen is a small peptide derived from HIV. Two residues in the heavy chain (blue) and one in the light chain (red) are colored to provide visual points of reference. (a) View of the Fab fragment in the absence of antigen, looking down on the antigen-binding site. (b) The same view, but with the Fab fragment in the bound conformation; the antigen is omitted to provide an unobstructed view of the altered binding site. Note how the binding cavity has enlarged and several groups have shifted position. (c) The same view as (b), but with the antigen in the binding site, pictured as a red stick structure.
19 Myosin thick filaments slide along actin thin filaments
20 Actomyosin Cycle Molecular mechanism of muscle contraction. Conformational changes in the myosin head that are coupled to stages in the ATP hydrolytic cycle cause myosin to successively dissociate from one actin subunit, then associate with another farther along the actin filament. In this way the myosin heads slide along the thin filaments, drawing the thick filament array into the thin filament array (see Figure 5-30).
21 Regulation of muscle contraction Availability of myosin-binding sites on actin is regulated by troponin and tropomyosin avoids continuous muscle contraction Nerve impulse triggers release of Ca 2+ Causes conformational changes to tropomyosin-troponin complex exposing myosin-binding sites
2013 W. H. Freeman and Company. 5 Function of Globular Proteins
 2013 W. H. Freeman and Company 5 Function of Globular Proteins CHAPTER 5: Function of Globular Proteins Key topics in protein function: Reversible binding of ligands is essential Specificity of ligands
2013 W. H. Freeman and Company 5 Function of Globular Proteins CHAPTER 5: Function of Globular Proteins Key topics in protein function: Reversible binding of ligands is essential Specificity of ligands
Proteins the primary biological macromolecules of living organisms
 Proteins the primary biological macromolecules of living organisms Protein structure and folding Primary Secondary Tertiary Quaternary structure of proteins Structure of Proteins Protein molecules adopt
Proteins the primary biological macromolecules of living organisms Protein structure and folding Primary Secondary Tertiary Quaternary structure of proteins Structure of Proteins Protein molecules adopt
Hmwk # 8 : DNA-Binding Proteins : Part II
 The purpose of this exercise is : Hmwk # 8 : DNA-Binding Proteins : Part II 1). to examine the case of a tandem head-to-tail homodimer binding to DNA 2). to view a Zn finger motif 3). to consider the case
The purpose of this exercise is : Hmwk # 8 : DNA-Binding Proteins : Part II 1). to examine the case of a tandem head-to-tail homodimer binding to DNA 2). to view a Zn finger motif 3). to consider the case
Chapter 4. Antigen Recognition by B-cell and T-cell Receptors
 Chapter 4 Antigen Recognition by B-cell and T-cell Receptors Antigen recognition by BCR and TCR B cells 2 separate functions of immunoglobulin (Ig) bind pathogen & induce immune responses recruit cells
Chapter 4 Antigen Recognition by B-cell and T-cell Receptors Antigen recognition by BCR and TCR B cells 2 separate functions of immunoglobulin (Ig) bind pathogen & induce immune responses recruit cells
Proteins Higher Order Structures
 Proteins Higher Order Structures Dr. Mohammad Alsenaidy Department of Pharmaceutics College of Pharmacy King Saud University Office: AA 101 msenaidy@ksu.edu.sa Previously on PHT 426!! Protein Structures
Proteins Higher Order Structures Dr. Mohammad Alsenaidy Department of Pharmaceutics College of Pharmacy King Saud University Office: AA 101 msenaidy@ksu.edu.sa Previously on PHT 426!! Protein Structures
Ensemble refinement shows conformational flexibility in crystal structures of human complement factor D
 Supplementary Information for Ensemble refinement shows conformational flexibility in crystal structures of human complement factor D Federico Forneris a,b, B. Tom Burnley a,b,c and Piet Gros a * a Crystal
Supplementary Information for Ensemble refinement shows conformational flexibility in crystal structures of human complement factor D Federico Forneris a,b, B. Tom Burnley a,b,c and Piet Gros a * a Crystal
Diversity of proteins
 BCMB 3100: Partial notes Chapter 4 (Part 1) Diversity of proteins 3D structure of proteins Fibrous vs globular proteins Conformation vs configuration 1, 2, 3 and 4 structure Peptide groups in polypeptide
BCMB 3100: Partial notes Chapter 4 (Part 1) Diversity of proteins 3D structure of proteins Fibrous vs globular proteins Conformation vs configuration 1, 2, 3 and 4 structure Peptide groups in polypeptide
Immunoglobulins. Light chain ~22-23 KDa whereas the heavy chain ~55-60 KDa
 Immunoglobulins Immunoglobulin (Ig) has a common name which is "Antibody (Ab)", but actually we should say Ig, why? Because the proteins, which are involved, are actually globular proteins "known as globulins"
Immunoglobulins Immunoglobulin (Ig) has a common name which is "Antibody (Ab)", but actually we should say Ig, why? Because the proteins, which are involved, are actually globular proteins "known as globulins"
Enzymes Part III: regulation I. Dr. Mamoun Ahram Summer, 2017
 Enzymes Part III: regulation I Dr. Mamoun Ahram Summer, 2017 Mechanisms of regulation Expression of isoenzymes Regulation of enzymatic activity Inhibitors Conformational changes Allostery Modulators Reversible
Enzymes Part III: regulation I Dr. Mamoun Ahram Summer, 2017 Mechanisms of regulation Expression of isoenzymes Regulation of enzymatic activity Inhibitors Conformational changes Allostery Modulators Reversible
SDS-PAGE and Western Blot. Molecular Basis of Evolution
 1 SDS-PAGE and Western Blot Molecular Basis of Evolution Homology high level of DNA and protein sequence similarity due to common ancestry. Evidence Genomes of related organisms are very similar. Even
1 SDS-PAGE and Western Blot Molecular Basis of Evolution Homology high level of DNA and protein sequence similarity due to common ancestry. Evidence Genomes of related organisms are very similar. Even
CSE : Computational Issues in Molecular Biology. Lecture 19. Spring 2004
 CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
Protein homology. Antigens & Antibodies I. Administrative issues:
 Administrative issues: Recommended text: Goldsby/Kuby Immunology, 6th edition (Note that Innate Immunity is not adequately covered in the 5th edition.) Text book reading assignments are to supplement the
Administrative issues: Recommended text: Goldsby/Kuby Immunology, 6th edition (Note that Innate Immunity is not adequately covered in the 5th edition.) Text book reading assignments are to supplement the
H zone narrows; light band narrows; outer darker regions of A / dark band widen; 2 max
 M. (a) (i) A / dark band is mainly due to myosin filaments; H zone only myosin filaments; darker band has both types of filament; light band has only actin filaments; max H zone narrows; light band narrows;
M. (a) (i) A / dark band is mainly due to myosin filaments; H zone only myosin filaments; darker band has both types of filament; light band has only actin filaments; max H zone narrows; light band narrows;
Bi 8 Lecture 7. Ellen Rothenberg 26 January Reading: Ch. 3, pp ; panel 3-1
 Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Introduction to Proteins
 Introduction to Proteins Lecture 4 Module I: Molecular Structure & Metabolism Molecular Cell Biology Core Course (GSND5200) Matthew Neiditch - Room E450U ICPH matthew.neiditch@umdnj.edu What is a protein?
Introduction to Proteins Lecture 4 Module I: Molecular Structure & Metabolism Molecular Cell Biology Core Course (GSND5200) Matthew Neiditch - Room E450U ICPH matthew.neiditch@umdnj.edu What is a protein?
 The replication of DNA Kornberg 1957 Meselson and Stahl 1958 Cairns 1963 Okazaki 1968 DNA Replication The driving force for DNA synthesis. The addition of a nucleotide to a growing polynucleotide
The replication of DNA Kornberg 1957 Meselson and Stahl 1958 Cairns 1963 Okazaki 1968 DNA Replication The driving force for DNA synthesis. The addition of a nucleotide to a growing polynucleotide
Nucleic Acids, Proteins, and Enzymes
 3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
Structural bioinformatics
 Structural bioinformatics Why structures? The representation of the molecules in 3D is more informative New properties of the molecules are revealed, which can not be detected by sequences Eran Eyal Plant
Structural bioinformatics Why structures? The representation of the molecules in 3D is more informative New properties of the molecules are revealed, which can not be detected by sequences Eran Eyal Plant
BCH222 - Greek Key β Barrels
 BCH222 - Greek Key β Barrels Reading C.I. Branden and J. Tooze (1999) Introduction to Protein Structure, Second Edition, pp. 77-78 & 335-336 (look at the color figures) J.S. Richardson (1981) "The Anatomy
BCH222 - Greek Key β Barrels Reading C.I. Branden and J. Tooze (1999) Introduction to Protein Structure, Second Edition, pp. 77-78 & 335-336 (look at the color figures) J.S. Richardson (1981) "The Anatomy
Nature Structural & Molecular Biology: doi: /nsmb.2548
 Supplementary Figure 1. Structure of GltPhout. (a) Stereo view of a slice through a single GltPhout protomer shown in stick representation along with 2Fo-Fc and anomalous difference electron maps. The
Supplementary Figure 1. Structure of GltPhout. (a) Stereo view of a slice through a single GltPhout protomer shown in stick representation along with 2Fo-Fc and anomalous difference electron maps. The
has only one nucleotide, U20, between G19 and A21, while trna Glu CUC has two
 SPPLEMENTRY INFORMTION doi:1.138/nature9411 Supplementary Discussion The structural characteristics of trn Gln G in comparison to trn Glu 16 in trn Gln G is directed towards the G19 56 pair, or the outer
SPPLEMENTRY INFORMTION doi:1.138/nature9411 Supplementary Discussion The structural characteristics of trn Gln G in comparison to trn Glu 16 in trn Gln G is directed towards the G19 56 pair, or the outer
Chromatographic Separation of the three forms of RNA Polymerase II.
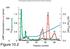 Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Structural Bioinformatics (C3210) DNA and RNA Structure
 Structural Bioinformatics (C3210) DNA and RNA Structure Importance of DNA/RNA 3D Structure Nucleic acids are essential materials found in all living organisms. Their main function is to maintain and transmit
Structural Bioinformatics (C3210) DNA and RNA Structure Importance of DNA/RNA 3D Structure Nucleic acids are essential materials found in all living organisms. Their main function is to maintain and transmit
Τάσος Οικονόµου ιαλεξη 8. Kινηση, λειτουργια, ελεγχος.
 Τάσος Οικονόµου ιαλεξη 8 Kινηση, λειτουργια, ελεγχος http://ecoserver.imbb.forth.gr/bio321.htm εν ξεχνω. Cell The peptide bond Polypeptides are stabilized by: 1. Covalent bonds= amide bond 2. Noncovalent,
Τάσος Οικονόµου ιαλεξη 8 Kινηση, λειτουργια, ελεγχος http://ecoserver.imbb.forth.gr/bio321.htm εν ξεχνω. Cell The peptide bond Polypeptides are stabilized by: 1. Covalent bonds= amide bond 2. Noncovalent,
Immunoglobulins: Structure and Function
 Immunoglobulins: Structure and Function Immunoglobulins:Structure and Function Definition: Glycoprotein molecules that are produced by plasma cells in response to an immunogen and which function as antibodies
Immunoglobulins: Structure and Function Immunoglobulins:Structure and Function Definition: Glycoprotein molecules that are produced by plasma cells in response to an immunogen and which function as antibodies
X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction
 X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction Tomohisa Shimasaki 1, Hiromi Yoshida 2, Shigehiro Kamitori 2 & Koji
X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction Tomohisa Shimasaki 1, Hiromi Yoshida 2, Shigehiro Kamitori 2 & Koji
Protein and RNA Review
 Protein and RNA Review Protein and RNA Review 1 Proteins: Introduction Proteins are the versatile building blocks and active molecules that form the basis of living systems. Function follows structure
Protein and RNA Review Protein and RNA Review 1 Proteins: Introduction Proteins are the versatile building blocks and active molecules that form the basis of living systems. Function follows structure
Protein design. CS/CME/BioE/Biophys/BMI 279 Oct. 24, 2017 Ron Dror
 Protein design CS/CME/BioE/Biophys/BMI 279 Oct. 24, 2017 Ron Dror 1 Outline Why design proteins? Overall approach: Simplifying the protein design problem Protein design methodology Designing the backbone
Protein design CS/CME/BioE/Biophys/BMI 279 Oct. 24, 2017 Ron Dror 1 Outline Why design proteins? Overall approach: Simplifying the protein design problem Protein design methodology Designing the backbone
Cambridge International Examinations Cambridge International Advanced Subsidiary and Advanced Level
 Cambridge International Examinations Cambridge International Advanced Subsidiary and Advanced Level *2249654089* BIOLOGY 9700/21 Paper 2 AS Level Structured Questions October/November 2016 1 hour 15 minutes
Cambridge International Examinations Cambridge International Advanced Subsidiary and Advanced Level *2249654089* BIOLOGY 9700/21 Paper 2 AS Level Structured Questions October/November 2016 1 hour 15 minutes
green B 1 ) into a single unit to model the substrate in this reaction. enzyme
 Teacher Key Objectives You will use the model pieces in the kit to: Simulate enzymatic actions. Explain enzymatic specificity. Investigate two types of enzyme inhibitors used in regulating enzymatic activity.
Teacher Key Objectives You will use the model pieces in the kit to: Simulate enzymatic actions. Explain enzymatic specificity. Investigate two types of enzyme inhibitors used in regulating enzymatic activity.
MOLECULAR RECOGNITION
 MOLECULAR RECOGNITION Bioanalytical Methods Classification 1. Biassay: molecular recognition, signal generation and detection in solution or on inert solid phase 2. Biosensor: molecular recognition system
MOLECULAR RECOGNITION Bioanalytical Methods Classification 1. Biassay: molecular recognition, signal generation and detection in solution or on inert solid phase 2. Biosensor: molecular recognition system
MOLEBIO LAB #3: Electrophoretic Separation of Proteins
 MOLEBIO LAB #3: Electrophoretic Separation of Proteins Introduction: Proteins occupy a central position in the structure and function of all living organisms. Some proteins serve as structural components
MOLEBIO LAB #3: Electrophoretic Separation of Proteins Introduction: Proteins occupy a central position in the structure and function of all living organisms. Some proteins serve as structural components
BETA STRAND Prof. Alejandro Hochkoeppler Department of Pharmaceutical Sciences and Biotechnology University of Bologna
 Prof. Alejandro Hochkoeppler Department of Pharmaceutical Sciences and Biotechnology University of Bologna E-mail: a.hochkoeppler@unibo.it C-ter NH and CO groups: right, left, right (plane of the slide)
Prof. Alejandro Hochkoeppler Department of Pharmaceutical Sciences and Biotechnology University of Bologna E-mail: a.hochkoeppler@unibo.it C-ter NH and CO groups: right, left, right (plane of the slide)
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs
 Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
6- Important Molecules of Living Systems. Proteins Nucleic Acids Taft College Human Physiology
 6- Important Molecules of Living Systems Proteins Nucleic Acids Taft College Human Physiology Proteins Proteins- made from: C, H, O, N, and S. Proteins are very large molecules composed of long chains
6- Important Molecules of Living Systems Proteins Nucleic Acids Taft College Human Physiology Proteins Proteins- made from: C, H, O, N, and S. Proteins are very large molecules composed of long chains
BIRKBECK COLLEGE (University of London)
 BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
Chapter 3 Nucleic Acids, Proteins, and Enzymes
 3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
Supplementary Figure 1.
 Supplementary Figure 1. Assessment of quaternary structure of soluble RSV F proteins. Soluble variants of F proteins from A2 and B1 RSV strains were expressed in HEK293 cells. The cell culture supernatants
Supplementary Figure 1. Assessment of quaternary structure of soluble RSV F proteins. Soluble variants of F proteins from A2 and B1 RSV strains were expressed in HEK293 cells. The cell culture supernatants
PROTEINS & NUCLEIC ACIDS
 Chapter 3 Part 2 The Molecules of Cells PROTEINS & NUCLEIC ACIDS Lecture by Dr. Fernando Prince 3.11 Nucleic Acids are the blueprints of life Proteins are the machines of life We have already learned that
Chapter 3 Part 2 The Molecules of Cells PROTEINS & NUCLEIC ACIDS Lecture by Dr. Fernando Prince 3.11 Nucleic Acids are the blueprints of life Proteins are the machines of life We have already learned that
SUPPLEMENTAL FIGURE LEGENDS. Figure S1: Homology alignment of DDR2 amino acid sequence. Shown are
 SUPPLEMENTAL FIGURE LEGENDS Figure S1: Homology alignment of DDR2 amino acid sequence. Shown are the amino acid sequences of human DDR2, mouse DDR2 and the closest homologs in zebrafish and C. Elegans.
SUPPLEMENTAL FIGURE LEGENDS Figure S1: Homology alignment of DDR2 amino acid sequence. Shown are the amino acid sequences of human DDR2, mouse DDR2 and the closest homologs in zebrafish and C. Elegans.
Genes and Proteins in Health. and Disease
 Genes and Health and I can describe the structure of proteins All proteins contain the chemical elements Carbon, Hydrogen, Oxygen and Nitrogen. Some also contain sulphur. Proteins are built from subunits
Genes and Health and I can describe the structure of proteins All proteins contain the chemical elements Carbon, Hydrogen, Oxygen and Nitrogen. Some also contain sulphur. Proteins are built from subunits
Immunoglobulins. Harper s biochemistry Chapter 49
 Immunoglobulins Harper s biochemistry Chapter 49 Immune system Detects and inactivates foreign molecules, viruses, bacteria and microorganisms Two components with 2 strategies B Lymphocytes (humoral immune
Immunoglobulins Harper s biochemistry Chapter 49 Immune system Detects and inactivates foreign molecules, viruses, bacteria and microorganisms Two components with 2 strategies B Lymphocytes (humoral immune
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than. SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with
 Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Basic Antibody Structure. Multiple myeloma = cancerous plasma cells Monomer = 150,000. Chapter 4. Immunoglobulin Structure and Function
 Chapter 4. Immunoglobulin Structure and Function. Functional Regions. Types of chains. Constant & Variable regions 4. Glycoprotein * * * Heavy chain= 446 aa Light chain= 4aa Each heavy and light chain
Chapter 4. Immunoglobulin Structure and Function. Functional Regions. Types of chains. Constant & Variable regions 4. Glycoprotein * * * Heavy chain= 446 aa Light chain= 4aa Each heavy and light chain
Fundamentals of Protein Structure
 Outline Fundamentals of Protein Structure Yu (Julie) Chen and Thomas Funkhouser Princeton University CS597A, Fall 2005 Protein structure Primary Secondary Tertiary Quaternary Forces and factors Levels
Outline Fundamentals of Protein Structure Yu (Julie) Chen and Thomas Funkhouser Princeton University CS597A, Fall 2005 Protein structure Primary Secondary Tertiary Quaternary Forces and factors Levels
Chapter 2. Antibodies
 Chapter 2. Antibodies An iddy-biddy antibody Just nanometers long Saved the butt of a sumo man Hundreds of kilos strong Anonymous The main elements of the immune system are firstly antibodies, secondly
Chapter 2. Antibodies An iddy-biddy antibody Just nanometers long Saved the butt of a sumo man Hundreds of kilos strong Anonymous The main elements of the immune system are firstly antibodies, secondly
Molecular Dynamics of Proteins
 Molecular Dynamics of Proteins Ubiquitin Myoglobin Equilibrium Properties of Proteins Ubiquitin Root Mean Squared Deviation: measure for equilibration and protein flexibility RMSD constant protein equilibrated
Molecular Dynamics of Proteins Ubiquitin Myoglobin Equilibrium Properties of Proteins Ubiquitin Root Mean Squared Deviation: measure for equilibration and protein flexibility RMSD constant protein equilibrated
Molecular Forces in Antibody Maturation*
 Molecular Forces in Antibody Maturation* Melik Demirel 1,2 1 Allen Pearce Assistant Professor, College of Engineering, The Pennsylvania State University, University Park, PA, USA, E-mail: mcd18@psu.edu
Molecular Forces in Antibody Maturation* Melik Demirel 1,2 1 Allen Pearce Assistant Professor, College of Engineering, The Pennsylvania State University, University Park, PA, USA, E-mail: mcd18@psu.edu
BCH 462. Single Radial Immunodiffusion and Immuno-electrophoresis
 BCH 462 Single Radial Immunodiffusion and Immuno-electrophoresis Immunoassays tests include: 1. Precipitation. 2. Agglutination. 3. Immunofluorescence. 4. Radioimmunoassay (RIA). 5. Enzyme-Linked Immuno
BCH 462 Single Radial Immunodiffusion and Immuno-electrophoresis Immunoassays tests include: 1. Precipitation. 2. Agglutination. 3. Immunofluorescence. 4. Radioimmunoassay (RIA). 5. Enzyme-Linked Immuno
STRUCTURAL BIOLOGY. α/β structures Closed barrels Open twisted sheets Horseshoe folds
 STRUCTURAL BIOLOGY α/β structures Closed barrels Open twisted sheets Horseshoe folds The α/β domains Most frequent domain structures are α/β domains: A central parallel or mixed β sheet Surrounded by α
STRUCTURAL BIOLOGY α/β structures Closed barrels Open twisted sheets Horseshoe folds The α/β domains Most frequent domain structures are α/β domains: A central parallel or mixed β sheet Surrounded by α
Chapter 5: Nucleic Acids, etc.
 Chapter 5: Nucleic Acids, etc. Voet & Voet: Sections 1 & 3 Pages 82-84 & 88-93 Any introductory Biochemistry textbook will have an introductory chapter on nucleic acids Slide 1 Nucleotides and Derivatives
Chapter 5: Nucleic Acids, etc. Voet & Voet: Sections 1 & 3 Pages 82-84 & 88-93 Any introductory Biochemistry textbook will have an introductory chapter on nucleic acids Slide 1 Nucleotides and Derivatives
Protein Folding Problem I400: Introduction to Bioinformatics
 Protein Folding Problem I400: Introduction to Bioinformatics November 29, 2004 Protein biomolecule, macromolecule more than 50% of the dry weight of cells is proteins polymer of amino acids connected into
Protein Folding Problem I400: Introduction to Bioinformatics November 29, 2004 Protein biomolecule, macromolecule more than 50% of the dry weight of cells is proteins polymer of amino acids connected into
Module I: Introduction
 Module I: Introduction 20.109 Lecture 1 3 February, 2011 Introduction to: Module Overview Fundamental concepts and techniques in molecular biology Appreciating nucleic acids (RNA in particular) as more
Module I: Introduction 20.109 Lecture 1 3 February, 2011 Introduction to: Module Overview Fundamental concepts and techniques in molecular biology Appreciating nucleic acids (RNA in particular) as more
Protein Structure and Function: From Sequence to Consequence 1. From Sequence to Structure
 Protein Structure and Function: From Sequence to Consequence Gregory A. Petsko and Dagmar Ringe, Brandeis University Published by New Science Press Ltd and distributed in the United States and Canada by
Protein Structure and Function: From Sequence to Consequence Gregory A. Petsko and Dagmar Ringe, Brandeis University Published by New Science Press Ltd and distributed in the United States and Canada by
Structure of Hemoglobin M Boston, a Variant with a Five-Coordinated
 Proc. Nat. Acad. Sci. USA Vol. 70, No. 12, Part II, pp. 3870-3874, December 1973 Structure of Hemoglobin M Boston, a Variant with a Five-Coordinated Ferric Heme (x-ray analysis/molecular pathology/inherited
Proc. Nat. Acad. Sci. USA Vol. 70, No. 12, Part II, pp. 3870-3874, December 1973 Structure of Hemoglobin M Boston, a Variant with a Five-Coordinated Ferric Heme (x-ray analysis/molecular pathology/inherited
Final exam. Please write your name on the exam and keep an ID card ready.
 Biophysics of Macromolecules Prof. R. Jungmann and Prof. J. Lipfert SS 2017 Final exam Final exam First name: Last name: Student number ( Matrikelnummer ): Please write your name on the exam and keep an
Biophysics of Macromolecules Prof. R. Jungmann and Prof. J. Lipfert SS 2017 Final exam Final exam First name: Last name: Student number ( Matrikelnummer ): Please write your name on the exam and keep an
Learning to Use PyMOL (includes instructions for PS #2)
 Learning to Use PyMOL (includes instructions for PS #2) To begin, download the saved PyMOL session file, 4kyz.pse from the Chem 391 Assignments web page: http://people.reed.edu/~glasfeld/chem391/assign.html
Learning to Use PyMOL (includes instructions for PS #2) To begin, download the saved PyMOL session file, 4kyz.pse from the Chem 391 Assignments web page: http://people.reed.edu/~glasfeld/chem391/assign.html
Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a
 Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a HNH-state 1 in PDB 4OO8, in which the distance from the C atom of the HNH catalytic residue 840 to the
Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a HNH-state 1 in PDB 4OO8, in which the distance from the C atom of the HNH catalytic residue 840 to the
Nature Structural & Molecular Biology: doi: /nsmb.3175
 Supplementary Figure 1 Medium- and low-fret groups probably representing varying positions on the DNA show similar changes between H3 and CENP-A nucleosomes with and without CENP-C CD, as does the high-fret
Supplementary Figure 1 Medium- and low-fret groups probably representing varying positions on the DNA show similar changes between H3 and CENP-A nucleosomes with and without CENP-C CD, as does the high-fret
Amino Acids. Amino Acid Structure
 Amino Acids Pratt & Cornely Chapter 4 Alpha carbon Sidechain Proteins peptides Amino Acid Structure 1 L amino acids Glycine R/S vs D/L L isoleucine racemization Stereochemisty Common Amino Acids 2 Which
Amino Acids Pratt & Cornely Chapter 4 Alpha carbon Sidechain Proteins peptides Amino Acid Structure 1 L amino acids Glycine R/S vs D/L L isoleucine racemization Stereochemisty Common Amino Acids 2 Which
Proteins. Amino Acids (APK) Peptides (APK) 5/23/2012. Peptide bond. Acid. Amino
 Proteins Amino Acids (APK) Acid Amino Image courtesy of Biotech (biotech.chem.indiana.edu/pages/ protein_intro.html) Peptides (APK) Peptide bond 1 Proteins (polypeptides) Segment of a protein Peptide bonds
Proteins Amino Acids (APK) Acid Amino Image courtesy of Biotech (biotech.chem.indiana.edu/pages/ protein_intro.html) Peptides (APK) Peptide bond 1 Proteins (polypeptides) Segment of a protein Peptide bonds
CHAPTER 3 ANTIBODY STRUCTURE I
 CHAPTER 3 ANTIBODY STRUCTURE I See APPENDIX: (3) OUCHTERLONY ANALYSIS; (6), EQUILIBRIUM DIALYSIS; (7) CROSS-REACTIVITY Electrophoretic separation of serum proteins identifies the GAMMA-GLOBULIN fraction
CHAPTER 3 ANTIBODY STRUCTURE I See APPENDIX: (3) OUCHTERLONY ANALYSIS; (6), EQUILIBRIUM DIALYSIS; (7) CROSS-REACTIVITY Electrophoretic separation of serum proteins identifies the GAMMA-GLOBULIN fraction
Gene Expression - Transcription
 DNA Gene Expression - Transcription Genes are expressed as encoded proteins in a 2 step process: transcription + translation Central dogma of biology: DNA RNA protein Transcription: copy DNA strand making
DNA Gene Expression - Transcription Genes are expressed as encoded proteins in a 2 step process: transcription + translation Central dogma of biology: DNA RNA protein Transcription: copy DNA strand making
CHAPTER 21 LECTURE SLIDES
 CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
CHAPTER 21 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
IRON METABOLISM. Harper s Illustrated Biochemistry chapter 50
 IRON METABOLISM Harper s Illustrated Biochemistry chapter 50 IRON 26th element in the periodic table Chemical Symbol: Fe MW = 55.85 Electron Configuration: 1s 2 2s 2 2p 6 3s 2 3p 6 4s 2 3d 6 Fourth most
IRON METABOLISM Harper s Illustrated Biochemistry chapter 50 IRON 26th element in the periodic table Chemical Symbol: Fe MW = 55.85 Electron Configuration: 1s 2 2s 2 2p 6 3s 2 3p 6 4s 2 3d 6 Fourth most
ANTIBODIES. Agents of Immunity
 ANTIBODIES Agents of Immunity - Antibodies are: The Organization What are they? Protective agents of the immune system Neutralize foreign agents called antigens Essential part of the Adaptive Immune System
ANTIBODIES Agents of Immunity - Antibodies are: The Organization What are they? Protective agents of the immune system Neutralize foreign agents called antigens Essential part of the Adaptive Immune System
Lecture for Wednesday. Dr. Prince BIOL 1408
 Lecture for Wednesday Dr. Prince BIOL 1408 THE FLOW OF GENETIC INFORMATION FROM DNA TO RNA TO PROTEIN Copyright 2009 Pearson Education, Inc. Genes are expressed as proteins A gene is a segment of DNA that
Lecture for Wednesday Dr. Prince BIOL 1408 THE FLOW OF GENETIC INFORMATION FROM DNA TO RNA TO PROTEIN Copyright 2009 Pearson Education, Inc. Genes are expressed as proteins A gene is a segment of DNA that
Label-free interaction analysis in realtime using surface plasmon resonance
 GE Healthcare Technology Note 23 Biacore systems Label-free interaction analysis in realtime using surface plasmon resonance Providing quantitative data on: report point Specificity sensorgram To what
GE Healthcare Technology Note 23 Biacore systems Label-free interaction analysis in realtime using surface plasmon resonance Providing quantitative data on: report point Specificity sensorgram To what
MCB 110:Biochemistry of the Central Dogma of MB. MCB 110:Biochemistry of the Central Dogma of MB
 MCB 110:Biochemistry of the Central Dogma of MB Part 1. DNA replication, repair and genomics (Prof. Alber) Part 2. RNA & protein synthesis. Prof. Zhou Part 3. Membranes, protein secretion, trafficking
MCB 110:Biochemistry of the Central Dogma of MB Part 1. DNA replication, repair and genomics (Prof. Alber) Part 2. RNA & protein synthesis. Prof. Zhou Part 3. Membranes, protein secretion, trafficking
Chapter 8: DNA and RNA
 Chapter 8: DNA and RNA Lecture Outline Enger, E. D., Ross, F. C., & Bailey, D. B. (2012). Concepts in biology (14th ed.). New York: McGraw- Hill. 1 8-1 DNA and the Importance of Proteins Proteins play
Chapter 8: DNA and RNA Lecture Outline Enger, E. D., Ross, F. C., & Bailey, D. B. (2012). Concepts in biology (14th ed.). New York: McGraw- Hill. 1 8-1 DNA and the Importance of Proteins Proteins play
UV Fluorescence Polarization as a Means to Investigate Protein Conformational and Mass Change
 A p p l i c a t i o n N o t e UV Fluorescence Polarization as a Means to Investigate Protein Conformational and Mass Change Using Intrinsic Tryptophan Fluorescence in Conjunction with UV-capable Polarizers
A p p l i c a t i o n N o t e UV Fluorescence Polarization as a Means to Investigate Protein Conformational and Mass Change Using Intrinsic Tryptophan Fluorescence in Conjunction with UV-capable Polarizers
466 Asn (N) to Ala (A) Generate beta dimer Interface
 Table S1: Amino acid changes to the HexA α-subunit to convert the dimer interface from α to β and to introduce the putative GM2A binding surface from β- onto the α- subunit Residue position (α-numbering)
Table S1: Amino acid changes to the HexA α-subunit to convert the dimer interface from α to β and to introduce the putative GM2A binding surface from β- onto the α- subunit Residue position (α-numbering)
Application of Biacore Technology
 Principles and typical results Application of Biacore Technology Common types of Biacore analyses Specificity analysis Is my molecule of interest specific for its target? Multiple binding analysis In which
Principles and typical results Application of Biacore Technology Common types of Biacore analyses Specificity analysis Is my molecule of interest specific for its target? Multiple binding analysis In which
Fig Ch 17: From Gene to Protein
 Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
BIOLOGY. Chapter 15 Genes & Proteins
 BIOLOGY Chapter 15 Genes & Proteins CMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 17 Protein Synthesis 2014 Pearson Education, Inc. Fig. 17-1 Figure 17.1a n albino racoon Condition
BIOLOGY Chapter 15 Genes & Proteins CMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 17 Protein Synthesis 2014 Pearson Education, Inc. Fig. 17-1 Figure 17.1a n albino racoon Condition
IDTutorial: DNA Replication
 IDTutorial: DNA Replication Introduction In their report announcing the structure of the DNA molecule, Watson and Crick (Nature, 171: 737-738, 1953) observe, It has not escaped our notice that the specific
IDTutorial: DNA Replication Introduction In their report announcing the structure of the DNA molecule, Watson and Crick (Nature, 171: 737-738, 1953) observe, It has not escaped our notice that the specific
Proteins Amides from Amino Acids
 Chapter 26 and Chapter 28 Proteins Amides from Amino Acids Amino acids contain a basic amino group and an acidic carboxyl group Joined as amides between the ¾NH 2 of one amino acid and the ¾CO 2 H to the
Chapter 26 and Chapter 28 Proteins Amides from Amino Acids Amino acids contain a basic amino group and an acidic carboxyl group Joined as amides between the ¾NH 2 of one amino acid and the ¾CO 2 H to the
Blood is 55% Plasma (Liquid)
 Blood is 55% Plasma (Liquid) The plasma portion of blood is: 91% Water Maintains blood volume Transports molecules 7% Proteins (ie: clotting proteins, albumin, immunoglobulins ) 2 % Salts, gases (O 2,
Blood is 55% Plasma (Liquid) The plasma portion of blood is: 91% Water Maintains blood volume Transports molecules 7% Proteins (ie: clotting proteins, albumin, immunoglobulins ) 2 % Salts, gases (O 2,
COMPUTATIONAL CHEMISTRY OF THE PORPHYRIN RING SYSTEM
 Proceedings of the South Dakota Academy of Science, Vol. 80 (2001) 129 COMPUTATIONAL CHEMISTRY OF THE PORPHYRIN RING SYSTEM Kari Lunder and Arlen Viste Department of Chemistry Augustana College Sioux Falls,
Proceedings of the South Dakota Academy of Science, Vol. 80 (2001) 129 COMPUTATIONAL CHEMISTRY OF THE PORPHYRIN RING SYSTEM Kari Lunder and Arlen Viste Department of Chemistry Augustana College Sioux Falls,
Table S1. Crystallographic Data and Refinement Statistics, Related to Experimental Procedures hsddb1-drddb2 CPD
 Data collection Table S1. Crystallographic Data and Refinement Statistics, Related to Experimental Procedures hsddb1-drddb2 CPD #1 hsddb1-drddb2 CPD #2 Values in parentheses are for the highest resolution
Data collection Table S1. Crystallographic Data and Refinement Statistics, Related to Experimental Procedures hsddb1-drddb2 CPD #1 hsddb1-drddb2 CPD #2 Values in parentheses are for the highest resolution
C. Tight Turns. = -30, φ 3. = 0, and type II approximately = 120, φ 3. = -60, ψ 2. = -90, ψ 3. = +90, ψ 3
 Tight turns (also known as reverse turns, β turns, β bends, hairpin bends, 310 bends, kinks, widgets, etc.) are the first and most prevalent type of nonrepetitive structure that has been recognized. While
Tight turns (also known as reverse turns, β turns, β bends, hairpin bends, 310 bends, kinks, widgets, etc.) are the first and most prevalent type of nonrepetitive structure that has been recognized. While
Protein Synthesis Notes
 Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
Super Models. Deoxyribonucleic Acid (DNA) Molecular Model Kit. Copyright 2015 Ryler Enterprises, Inc. Recommended for ages 10-adult
 Super Models Deoxyribonucleic Acid (DNA) Molecular Model Kit Copyright 2015 Ryler Enterprises, Inc. Recommended for ages 10-adult! Caution: Atom centers and vinyl tubing are a choking hazard. Do not eat
Super Models Deoxyribonucleic Acid (DNA) Molecular Model Kit Copyright 2015 Ryler Enterprises, Inc. Recommended for ages 10-adult! Caution: Atom centers and vinyl tubing are a choking hazard. Do not eat
Chapter I.1: Hemocyanin (Hc)
 Chapter I.1: Hemocyanin (Hc) 2 is essential for animals Low 2 solubility in aqueous solution (0.2 ml in 100 ml plasma) need some ways to carry 2 Exception otothenoids/family Channichthyidae Antarctic Icefish
Chapter I.1: Hemocyanin (Hc) 2 is essential for animals Low 2 solubility in aqueous solution (0.2 ml in 100 ml plasma) need some ways to carry 2 Exception otothenoids/family Channichthyidae Antarctic Icefish
Addison Ault, Department of Chemistry, Cornell College, Mount Vernon, IA. The isoelectric point is a point on the ph scale. It is the ph at which the
 1 Isoelectric Points On the Back of the Envelope Addison Ault, Department of Chemistry, Cornell College, Mount Vernon, IA The isoelectric point is a point on the ph scale. It is the ph at which the average
1 Isoelectric Points On the Back of the Envelope Addison Ault, Department of Chemistry, Cornell College, Mount Vernon, IA The isoelectric point is a point on the ph scale. It is the ph at which the average
Antibodies and Antigens in the Blood Bank 9/7/2015 NAHLA BAKHAMIS 1
 Antibodies and Antigens in the Blood Bank NAHLA BAKHAMIS 9/7/2015 NAHLA BAKHAMIS 1 Outline Antibodies structure, classes and functions Most important Abs in the blood bank effective roles of Abs Zeta potential
Antibodies and Antigens in the Blood Bank NAHLA BAKHAMIS 9/7/2015 NAHLA BAKHAMIS 1 Outline Antibodies structure, classes and functions Most important Abs in the blood bank effective roles of Abs Zeta potential
2012 GENERAL [5 points]
![2012 GENERAL [5 points] 2012 GENERAL [5 points]](/thumbs/75/72452386.jpg) GENERAL [5 points] 2012 Mark all processes that are part of the 'standard dogma of molecular' [ ] DNA replication [ ] transcription [ ] translation [ ] reverse transposition [ ] DNA restriction [ ] DNA
GENERAL [5 points] 2012 Mark all processes that are part of the 'standard dogma of molecular' [ ] DNA replication [ ] transcription [ ] translation [ ] reverse transposition [ ] DNA restriction [ ] DNA
Supplementary material for Ab initio structure modeling of complex thin-film oxides: thermodynamical stability of TiC/thin-film
 Supplementary material for Ab initio structure modeling of complex thin-film oxides: thermodynamical stability of TiC/thin-film alumina (Rohrer et al 2009 J. Phys.: Condens. Matter 22 015004) J Rohrer
Supplementary material for Ab initio structure modeling of complex thin-film oxides: thermodynamical stability of TiC/thin-film alumina (Rohrer et al 2009 J. Phys.: Condens. Matter 22 015004) J Rohrer
Biomolecular chemistry. 7. Antibodies: structure and function
 154 Biomolecular chemistry 7. Antibodies: structure and function Suggested reading: Sections 5.1 to 5.3 of Mikkelsen and Cortón, Bioanalytical Chemistry Primary Source Material Biochemistry Chapter 33:
154 Biomolecular chemistry 7. Antibodies: structure and function Suggested reading: Sections 5.1 to 5.3 of Mikkelsen and Cortón, Bioanalytical Chemistry Primary Source Material Biochemistry Chapter 33:
Chapter 2 Molecules to enzymes - Short answer [72 marks]
![Chapter 2 Molecules to enzymes - Short answer [72 marks] Chapter 2 Molecules to enzymes - Short answer [72 marks]](/thumbs/72/67260206.jpg) Chapter 2 Molecules to enzymes - Short answer [72 marks] 1a. Outline primary and quaternary protein structures. Primary protein structure: Quaternary protein structure: a. (primary structure) is sequence
Chapter 2 Molecules to enzymes - Short answer [72 marks] 1a. Outline primary and quaternary protein structures. Primary protein structure: Quaternary protein structure: a. (primary structure) is sequence
Characterization of Aptamer Binding using SensíQ SPR Platforms
 Characterization of Aptamer Binding using SensíQ SPR Platforms APPLICATION NOTE INTRODUCTION Aptamers have the potential to provide a better solution in diagnostics and other research areas than traditional
Characterization of Aptamer Binding using SensíQ SPR Platforms APPLICATION NOTE INTRODUCTION Aptamers have the potential to provide a better solution in diagnostics and other research areas than traditional
DENATURATION OF HEMOGLOBIN IN THE PRESENCE OF TANNIC ACID
 PROCEEDINGS OF THE YEREVAN STATE UNIVERSITY C h e m i s t r y a n d B i o l o g y 2014, 1, p. 23 27 C h emistr y DENATURATION OF HEMOGLOBIN IN THE PRESENCE OF TANNIC ACID K. R. GRIGORYAN, L. S. SARGSYAN
PROCEEDINGS OF THE YEREVAN STATE UNIVERSITY C h e m i s t r y a n d B i o l o g y 2014, 1, p. 23 27 C h emistr y DENATURATION OF HEMOGLOBIN IN THE PRESENCE OF TANNIC ACID K. R. GRIGORYAN, L. S. SARGSYAN
UNIT V -CRYSTAL STRUCTURE
 UNIT V -CRYSTAL STRUCTURE Solids are of two types: Amorphous and crystalline. In amorphous solids, there is no order in the arrangement of their constituent atoms (molecules). Hence no definite structure
UNIT V -CRYSTAL STRUCTURE Solids are of two types: Amorphous and crystalline. In amorphous solids, there is no order in the arrangement of their constituent atoms (molecules). Hence no definite structure
Chapter 11. Gene Expression and Regulation. Lectures by Gregory Ahearn. University of North Florida. Copyright 2009 Pearson Education, Inc..
 Chapter 11 Gene Expression and Regulation Lectures by Gregory Ahearn University of North Florida Copyright 2009 Pearson Education, Inc.. 11.1 How Is The Information In DNA Used In A Cell? Most genes contain
Chapter 11 Gene Expression and Regulation Lectures by Gregory Ahearn University of North Florida Copyright 2009 Pearson Education, Inc.. 11.1 How Is The Information In DNA Used In A Cell? Most genes contain
Introduction to molecular docking
 Introduction to molecular docking Alvaro Cortes Cabrera, Javier Klett, Antonio Morreale Centro de Biología Molecular Severo Ochoa. UAM CSIC C/Nicolas Cabrera, 1. Campos Cantoblanco UAM. 28049, Madrid.
Introduction to molecular docking Alvaro Cortes Cabrera, Javier Klett, Antonio Morreale Centro de Biología Molecular Severo Ochoa. UAM CSIC C/Nicolas Cabrera, 1. Campos Cantoblanco UAM. 28049, Madrid.
Masayoshi Honda, Jeehae Park, Robert A. Pugh, Taekjip Ha, and Maria Spies
 Molecular Cell, Volume 35 Supplemental Data Single-Molecule Analysis Reveals Differential Effect of ssdna-binding Proteins on DNA Translocation by XPD Helicase Masayoshi Honda, Jeehae Park, Robert A. Pugh,
Molecular Cell, Volume 35 Supplemental Data Single-Molecule Analysis Reveals Differential Effect of ssdna-binding Proteins on DNA Translocation by XPD Helicase Masayoshi Honda, Jeehae Park, Robert A. Pugh,
Electrophoresis and transfer
 Electrophoresis and transfer Electrophoresis Cation = positively charged ion, it moves toward the cathode (-) Anion = negatively charged ion, it moves toward the anode (+) Amphoteric substance = can have
Electrophoresis and transfer Electrophoresis Cation = positively charged ion, it moves toward the cathode (-) Anion = negatively charged ion, it moves toward the anode (+) Amphoteric substance = can have
Movement at the Molecular Level
 Movement at the Molecular Level Diffusion: = 6 D t (D 6 π µ a) Typical numbers: 10 nm protein in water D= 10-10 m 2 /s.in cells D= 10-12 m 2 /s (D= 10-14 m 2 /s lipids) [] 1/2 =1 µm, t ~0.2
Movement at the Molecular Level Diffusion: = 6 D t (D 6 π µ a) Typical numbers: 10 nm protein in water D= 10-10 m 2 /s.in cells D= 10-12 m 2 /s (D= 10-14 m 2 /s lipids) [] 1/2 =1 µm, t ~0.2
DNA AND CHROMOSOMES. Genetica per Scienze Naturali a.a prof S. Presciuttini
 DNA AND CHROMOSOMES This document is licensed under the Attribution-NonCommercial-ShareAlike 2.5 Italy license, available at http://creativecommons.org/licenses/by-nc-sa/2.5/it/ 1. The Building Blocks
DNA AND CHROMOSOMES This document is licensed under the Attribution-NonCommercial-ShareAlike 2.5 Italy license, available at http://creativecommons.org/licenses/by-nc-sa/2.5/it/ 1. The Building Blocks
