Expanded View Figures
|
|
|
- Harvey Ray
- 5 years ago
- Views:
Transcription
1 yako Yachie-Kinoshita et al Predictive control of pluripotency Molecular Systems iology Expanded View Figures Figure EV1. General strategy of the simulation. The properties of synchronous (i) and asynchronous (ii) oolean simulation. Importantly in S, a single starting profile can give rise to multiple different resultant profiles and therefore all transitional profiles are represented with probabilities. Note that we used random S where the genes updated in each transition are set randomly to reduce the calculation cost. The background of our simulation framework: heterogeneity derived from single-cell fluctuations and stabilization of PSC populations showing robust gene expression pattern under a certain signaling input condition (depicted with black geometric shapes). ª 218 The uthors Molecular Systems iology 14: e EV1
2 Molecular Systems iology Predictive control of pluripotency yako Yachie-Kinoshita et al Lrh1 Pitx2 Mycn SignalCT Dnmt3b ctivin/nodal Cdx2 Klf2 Lefty1 Nanog Jarid2 Pecam1 Oct4 Tcf3 Rex1 EpiTF Gcnf SignalWNT igsk3b Sox2 Smad7 Tbx3 EpiTFs Smad6 Gata6 Esrrb Myc Klf4 SignalPI3K SignalMP Dusp6 Fgf4 Gbx2 SignalLIF mp4 SignalERK Fgfr2 SignalFGF LIF bfgf Figure EV2. Modeling of mesc-grn; related to Fig 2. network view of the mesc-grn model where rectangles indicate model components including genes (white and green white and green for genes with and without outgoing regulatory interactions), signaling activities (gray), and cytokines or small molecules as inputs to control signaling activities (dark gray). Edges between rectangles represent the regulatory relationships between genes (solid lines) and within signaling pathways (dotted lines). The edge color indicates either literature curation-based (black) or inferred/predicted (red) regulations. EV2 Molecular Systems iology 14: e ª 218 The uthors
3 yako Yachie-Kinoshita et al Predictive control of pluripotency Molecular Systems iology Figure EV3. Comparison of predicted and experimentally observed data on gene expression patterns in distinct PSCs; related to Fig 3. Predicted population-averaged expression level (mean of five independent simulations) for each pluripotency-associated gene in the control LS condition is comparable to the frequency of gene-expressing cells from the reported single-cell measurements using RN-seq (triangle; Kolodziejczyk et al, 215) and fluidigmqpcr (circle; Macrthur et al, 212). The gene expression levels were binarized into ON or OFF for each single cell to calculate frequencies. The error bars for the simulation resutls represent s.d. of five independent simulations, whereas those for the single-cell RN-seq data represent s.d. of two replicates. Comparison of predicted co-expressions of epiblast-specific genes (EpiTFs Eomes, Otx2, and Fgf5) with O/S/N and those measured by qpcr-based single-cell mrn data. Oct4 and Nanog are likely to be co-expressed with EpiTFs, whereas Sox2 is not, as it shows a negative correlation with EpiTFs (i.e., EpiTF expression in Sox2 cells is higher) in both our simulation and in published, single-cell expression data (Hayashi et al, 28; Macrthur et al, 212). P-value was calculated by Fisher s exact test. The error bars represent s.d. of five independent simulations. C Simulation of distinct PSCs recapitulates population-level gene expression measurements. ll expression data are scaled relative to respective gene expression in control LS conditions. The hierarchical clustering with U (approximately unbiased) P-value and P (bootstrap probability) value based on multiscale bootstrap resampling was carried out with the pvclust package in R. The results are displayed in the pinwheel view in Fig 3C, where upper and lower limits are set to +5 and 1, respectively, in the simulation to avoid singularities caused by null expression. The experimental data were taken from indicated GEO entries, and RN-seq data for EpiSCs were kindly provided by Dr. Ronald Mckay. Cell lines used are 129 (GSE1563 and GSE62155), J1 (GSE58735), ESF175/1, ESF58/2, ESF122 (GSE792) cell lines, or mescs were derived from C57L/6J strain blastocysts (GSE53275). D In silico GOF/LOF study in EpiSC (bf+) or mesc (LS) conditions. ll SCCs above ten profiles and sustainability >.7 are shown, and the gene expression levels of each component in each SCC are color-coded between blue (.) to yellow (1.). The population-averaged expression levels based on the GOF/LOF results were shown in the PC metrics in Fig 3D. E Predictions (left) and measurements (right) of population-averaged expression levels of OSN in EpiSC conditions (bf+) increased in response to extrinsic manipulation of MP4. MP4 was set as continuous-on (EpiSC + MP4) and random (EpiSC). The frequencies reported for Oct4, Sox2, and Nanog-positive cells, assessed by Cellomics high content screening, represent the mean and s.d. of four replicates. sterisk indicates the significant difference (P <.5 unpaired, two-sided t-test for total OSN). The error bars for the simulation results represent s.d. of five independent simulations. Source data are available online for this figure. ª 218 The uthors Molecular Systems iology 14: e EV3
4 Molecular Systems iology Predictive control of pluripotency yako Yachie-Kinoshita et al C E D Figure EV3. EV4 Molecular Systems iology 14: e ª 218 The uthors
5 yako Yachie-Kinoshita et al Predictive control of pluripotency Molecular Systems iology Figure EV4. LIF stabilizes the pluripotent population while 2i up-regulates OSN; related to Fig 4. Summation of frequencies ( 1) of O/S/N-positive cells assessed by high content screening at 2 days after medium replacement (n = 6 for Oct4 and Sox2, n = 4 for Nanog; the error bars represent s.d.). sterisk indicates the significant difference for total OSN assessed by unpaired, two-sided student t-test (**P <.5, *P <.1). Upper panel: measure of Pearson s correlation (PCC) for OSN in 2i-supplemented conditions, quantified using high content screening (n = 6 for Oct4 and Sox2, n = 4 for Nanog; the error bars represent s.d.). Lower panel: measure of PCC for OSN in the results of a minimal perturbation-sensitivity analysis described in (C). The significance was assessed by Wilcoxon exact rank test (*P <.5, **P <.1). C The results of minimal perturbation-sensitivity analysis of the model where the model network was perturbed by removing single regulatory edges. Purple indicates down-regulation of the genes by removing the regulatory edge, which means the regulation had a positive role on the expression level of O/S/N, while green indicates the reverse. Note that the results shown are the effects of regulatory edge removal: the removal of inhibitory regulation from Cdx2 to Oct4 increases Oct4, which means that the regulation edge acts as a negative effector for Oct4 level. D The difference in PSC population stability between and 2i L conditions was assessed via published single gene LOF and double genes LOF in vitro (Dunn et al, 214; EXP) and in silico (SIM). The upper panel depicts the experimental results for the relative count of alkaline phosphatase (P) positive cells to untreated colonies upon gene manipulations. The simulation data (lower panel) shows population-averaged expression level of Oct4 relative to controls ( and 2i L) where the manipulated gene was set as continuously OFF. lue nodes indicate the response in 2i L, while red nodes indicate those in (n = 5 for simulation and n = 3 5 for experimental). E The experimental inputs (additives of the conditions) corresponding to the simulation inputs shown in Fig 4E. bbreviations used are as follows: Jaki for Janus kinase (JK) inhibitor, Dkk1 for Dickkopf1, and lki for chemical inhibitor selective for ctivin receptor-like kinase (LK) 4/5/7. F Predicted levels (left) and measured gene-expressing cell frequencies (right) of O/S/N for each condition group (L, LIF; W, WNT). The average value of four distinct signal conditions (MPctivin/Nodal) in each group for each gene was calculated and summed up into an OSN score (n = 6 for Oct4 and Sox2, n = 4 for Nanog; the error bars represent s.d.). sterisk indicates the significant difference for total OSN assessed by Wilcoxon exact rank test (**P <.5, *P <.1). G Susceptibility of O/S/N expression levels of PSCs to ctivin and MP signal perturbations was predicted (upper panel) and measured (lower panel). Standard deviation per mean of predicted expression levels or gene-expressing frequencies from immunostaining was calculated as a coefficient of variation for each group (LIFWNT) including four distinct signal conditions (MPctivin/Nodal). H Frequencies of O/S/N-expressing cells in the 19 combinatorial signal conditions with and without serum assessed by immunostaining (n = 2). Most conditions were equivalent except for L W + and +L W. The notable down-regulation of OSN in L+W+ (=+ ) was seen in both with- and without-serum conditions. The error bars represent s.d. of five independent simulations. Source data are available online for this figure. ª 218 The uthors Molecular Systems iology 14: e EV5
6 Molecular Systems iology Predictive control of pluripotency yako Yachie-Kinoshita et al C D E H G F Figure EV4. EV6 Molecular Systems iology 14: e ª 218 The uthors
7 yako Yachie-Kinoshita et al Predictive control of pluripotency Molecular Systems iology Proportion D Lineage contribution (% Chimera) 5 (%) Epiblast ME Coexist PSC PE TE H2-Tomato Oct4 EpiTFs Gata6 Cdx2 2i L+- 2i L LIF EpiSC (bf+) TE 8 7 EPI Total Chimera efficiency (%) # Chimeras C Cdx2(+) Density #Cells (LS) LS +- d5 Cdx2 (a.u.) Cdx2(+) Frequency (%) d * LS mesc CD Oct4(+) ** Oct4(-) 2i-L 2i-L TSC mesc TSC CDCP EPI-position TE-position Figure EV5. In silico subpopulation analysis reveals mescs exit pluripotency toward TE-like lineage in + condition; related to Fig 5. In silico subpopulation analysis of possible signaling input combinations with and 2i L. The threshold values for predicted expression levels of lineage specifiers in each SCC are set as follows: Oct4 =.3, EpiTFs =.2, and Gata6 =.5 and Cdx2 =.7. Frequency of Cdx2 + population including TE-like subpopulation (Cdx2 + /Oct4 ) after 5 days in culture measured by immunostaining. Data represent the means and the error bars represent s.d. of six replicates consisting of two biological replicates with three technical replicates in each. The representative density plot of immunostaining of Cdx2 on day 5 is shown in the left panel. The significance was assessed by Wilcoxon exact rank test (*P <.5, **P <.5) for the frequencies of total Cdx2 + and Cdx2 + Oct4 comparing + and +. C Histograms for expression of TE-enriched cell surface markers, CD4 and CDCP1. Shown are mescs treated in (1 st column), 2i L (2 nd column), or (3 rd column), in unsupplemented medium (1 st row) or MP4 and LKi supplemented medium (+ ; 2 nd row). Control mescs (kept in control LS) and TSCs are shown at the bottom. D In vivo lineage contribution frequency and chimera efficiency of H2-Tomato ESCs (Morgani et al, 213) treated with either or in the presence of serum. Source data are available online for this figure. ª 218 The uthors Molecular Systems iology 14: e EV7
Modeling signaling-dependent pluripotency with Boolean logic to predict cell fate transitions
 Published online: January 29, 218 rticle Modeling signaling-dependent pluripotency with oolean logic to predict cell fate transitions yako Yachie-Kinoshita 1,2,3, Kento Onishi 1,2, Joel Ostblom 1,2, Matthew
Published online: January 29, 218 rticle Modeling signaling-dependent pluripotency with oolean logic to predict cell fate transitions yako Yachie-Kinoshita 1,2,3, Kento Onishi 1,2, Joel Ostblom 1,2, Matthew
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 Schematic and results of screening the combinatorial antibody library for Sox2 replacement activity. A single batch of MEFs were plated and transduced with doxycycline inducible
Supplementary Figure 1 Schematic and results of screening the combinatorial antibody library for Sox2 replacement activity. A single batch of MEFs were plated and transduced with doxycycline inducible
Nature Biotechnology: doi: /nbt.4166
 Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency.
 Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency. (a) Transfection of different concentration of GAS5-overexpressing
Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency. (a) Transfection of different concentration of GAS5-overexpressing
Development 142: doi: /dev : Supplementary Material
 Development 142: doi:1.42/dev.11687: Supplementary Material Figure S1 - Relative expression of lineage markers in single ES cell and cardiomyocyte by real-time quantitative PCR analysis. Single ES cell
Development 142: doi:1.42/dev.11687: Supplementary Material Figure S1 - Relative expression of lineage markers in single ES cell and cardiomyocyte by real-time quantitative PCR analysis. Single ES cell
Construction and Validation of a Regulatory Network for Pluripotency and Self-Renewal of Mouse Embryonic Stem Cells
 Construction and Validation of a Regulatory Network for Pluripotency and Self-Renewal of Mouse Embryonic Stem Cells Huilei Xu 1,2,3, Yen-Sin Ang 2,3, Ana Sevilla 2,3, Ihor R. Lemischka 1,2,3 *, Avi Ma
Construction and Validation of a Regulatory Network for Pluripotency and Self-Renewal of Mouse Embryonic Stem Cells Huilei Xu 1,2,3, Yen-Sin Ang 2,3, Ana Sevilla 2,3, Ihor R. Lemischka 1,2,3 *, Avi Ma
The many faces of Pluripotency: in vitro adaptations of a continuum of in vivo states
 Morgani et al. BMC Developmental Biology (2017) 17:7 DOI 10.1186/s12861-017-0150-4 REVIEW The many faces of Pluripotency: in vitro adaptations of a continuum of in vivo states Sophie Morgani 1,2, Jennifer
Morgani et al. BMC Developmental Biology (2017) 17:7 DOI 10.1186/s12861-017-0150-4 REVIEW The many faces of Pluripotency: in vitro adaptations of a continuum of in vivo states Sophie Morgani 1,2, Jennifer
Supplementary Figure 1. Soft fibrin gels promote growth and organized mesodermal differentiation. Representative images of single OGTR1 ESCs cultured
 Supplementary Figure 1. Soft fibrin gels promote growth and organized mesodermal differentiation. Representative images of single OGTR1 ESCs cultured in 90-Pa 3D fibrin gels for 5 days in the presence
Supplementary Figure 1. Soft fibrin gels promote growth and organized mesodermal differentiation. Representative images of single OGTR1 ESCs cultured in 90-Pa 3D fibrin gels for 5 days in the presence
V10: Core Pluripotency Network
 V10: Core Pluripotency Network Mouse ES cells were isolated for the first time in 1981 from mouse blastocysts. Maintenance of the self-renewing state of mouse ES cells requires the cytokine leukemia inhibitory
V10: Core Pluripotency Network Mouse ES cells were isolated for the first time in 1981 from mouse blastocysts. Maintenance of the self-renewing state of mouse ES cells requires the cytokine leukemia inhibitory
T-iPSC. Gra-iPSC. B-iPSC. TTF-iPSC. Supplementary Figure 1. Nature Biotechnology: doi: /nbt.1667
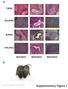 a T-iPSC Gra-iPSC B-iPSC TTF-iPSC Ectoderm Endoderm Mesoderm b Nature Biotechnology: doi:.38/nbt.667 Supplementary Figure Klf4 transgene expression Oct4 transgene expression.7 Fold GAPDH.6.5.4.3 Fold GAPDH
a T-iPSC Gra-iPSC B-iPSC TTF-iPSC Ectoderm Endoderm Mesoderm b Nature Biotechnology: doi:.38/nbt.667 Supplementary Figure Klf4 transgene expression Oct4 transgene expression.7 Fold GAPDH.6.5.4.3 Fold GAPDH
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Multiplet rate and sensitivity of the GemCode single cell platform from scrna-seq of 50:50 mixing of 293Ts and 3T3s. (a) Inferred multiplet rate as a function
Supplementary Figures Supplementary Figure 1. Multiplet rate and sensitivity of the GemCode single cell platform from scrna-seq of 50:50 mixing of 293Ts and 3T3s. (a) Inferred multiplet rate as a function
Nature Methods: doi: /nmeth.4396
 Supplementary Figure 1 Comparison of technical replicate consistency between and across the standard ATAC-seq method, DNase-seq, and Omni-ATAC. (a) Heatmap-based representation of ATAC-seq quality control
Supplementary Figure 1 Comparison of technical replicate consistency between and across the standard ATAC-seq method, DNase-seq, and Omni-ATAC. (a) Heatmap-based representation of ATAC-seq quality control
Single-cell analysis reveals the continuum of human lympho-myeloid progenitor cells
 SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41590-017-0001-2 In the format provided by the authors and unedited. Single-cell analysis reveals the continuum of human lympho-myeloid progenitor
SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41590-017-0001-2 In the format provided by the authors and unedited. Single-cell analysis reveals the continuum of human lympho-myeloid progenitor
SUPPLEMENTARY INFORMATION
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION ARTICLE NUMBER: 16206 DOI: 10.1038/NMICROBIOL.2016.206 Single cell RNA seq ties macrophage polarization to growth rate of intracellular
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION ARTICLE NUMBER: 16206 DOI: 10.1038/NMICROBIOL.2016.206 Single cell RNA seq ties macrophage polarization to growth rate of intracellular
SUPPLEMENTARY FIG. S9. Morphology and DNA staining of cipsc-differentiated trophoblast cell-like cell. Scale bar: 100 mm. SUPPLEMENTARY FIG. S10.
 Supplementary Data SUPPLEMENTARY FIG. S1. Lentivirus-infected canine fibroblasts show YFP expression. (A, B) Uninfected canine skin fibroblasts (CSFs); (C, D) infected CSFs; (E, F) uninfected CTFs; (G,
Supplementary Data SUPPLEMENTARY FIG. S1. Lentivirus-infected canine fibroblasts show YFP expression. (A, B) Uninfected canine skin fibroblasts (CSFs); (C, D) infected CSFs; (E, F) uninfected CTFs; (G,
Supplementary Figure 1 Characterization of OKSM-iNSCs
 Supplementary Figure 1 Characterization of OKSM-iNSCs (A) Gene expression levels of Sox1 and Sox2 in the indicated cell types by microarray gene expression analysis. (B) Dendrogram cluster analysis of
Supplementary Figure 1 Characterization of OKSM-iNSCs (A) Gene expression levels of Sox1 and Sox2 in the indicated cell types by microarray gene expression analysis. (B) Dendrogram cluster analysis of
A common molecular logic determines embryonic stem cell self-renewal and reprogramming
 Resource A common molecular logic determines embryonic stem cell self-renewal and reprogramming Sara-Jane Dunn 1,2,, Meng Amy Li 2,, Elena Carbognin 3, Austin Smith 2,4,* & Graziano Martello 3,** Abstract
Resource A common molecular logic determines embryonic stem cell self-renewal and reprogramming Sara-Jane Dunn 1,2,, Meng Amy Li 2,, Elena Carbognin 3, Austin Smith 2,4,* & Graziano Martello 3,** Abstract
Supplementary Figures Supplementary Figure 1
 Supplementary Figures Supplementary Figure 1 Supplementary Figure 1 COMSOL simulation demonstrating flow characteristics in hydrodynamic trap structures. (a) Flow field within the hydrodynamic traps before
Supplementary Figures Supplementary Figure 1 Supplementary Figure 1 COMSOL simulation demonstrating flow characteristics in hydrodynamic trap structures. (a) Flow field within the hydrodynamic traps before
Understanding Transcripts in UCSC Genome Browser
 Understanding Transcripts in UCSC Genome Browser Cline and Kent, Nature Biotechnology 27, 153-155 (2009) 1 Gene Annotations 2 Gene Annotations 2 3 Galaxy Databases are not analyses tools. Databases are
Understanding Transcripts in UCSC Genome Browser Cline and Kent, Nature Biotechnology 27, 153-155 (2009) 1 Gene Annotations 2 Gene Annotations 2 3 Galaxy Databases are not analyses tools. Databases are
Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Supplementary Material
 Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions Joshua N. Burton 1, Andrew Adey 1, Rupali P. Patwardhan 1, Ruolan Qiu 1, Jacob O. Kitzman 1, Jay Shendure 1 1 Department
Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions Joshua N. Burton 1, Andrew Adey 1, Rupali P. Patwardhan 1, Ruolan Qiu 1, Jacob O. Kitzman 1, Jay Shendure 1 1 Department
gacgacgaggagaccaccgctttg aggcacattgaaggtctcaaacatg
 Supplementary information Supplementary table 1: primers for cloning and sequencing cloning for E- Ras ggg aat tcc ctt gag ctg ctg ggg aat ggc ttt gcc ggt cta gag tat aaa gga agc ttt gaa tcc Tpbp Oct3/4
Supplementary information Supplementary table 1: primers for cloning and sequencing cloning for E- Ras ggg aat tcc ctt gag ctg ctg ggg aat ggc ttt gcc ggt cta gag tat aaa gga agc ttt gaa tcc Tpbp Oct3/4
Extrinsic regulation of pluripotent stem cells Martin F. Pera 1 & Patrick P. L. Tam 2,3
 NATURE Vol 465 10 June 2010 doi:10.1038/nature09228 REVIEW INSIGHT Extrinsic regulation of pluripotent stem s Martin F. Pera 1 & Patrick P. L. Tam 2,3 During early mammalian development, as the pluripotent
NATURE Vol 465 10 June 2010 doi:10.1038/nature09228 REVIEW INSIGHT Extrinsic regulation of pluripotent stem s Martin F. Pera 1 & Patrick P. L. Tam 2,3 During early mammalian development, as the pluripotent
Cell Stem Cell, volume 9 Supplemental Information
 Cell Stem Cell, volume 9 Supplemental Information Large-Scale Analysis Reveals Acquisition of Lineage-Specific Chromosomal Aberrations in Human Adult Stem Cells Uri Ben-David, Yoav Mayshar, and Nissim
Cell Stem Cell, volume 9 Supplemental Information Large-Scale Analysis Reveals Acquisition of Lineage-Specific Chromosomal Aberrations in Human Adult Stem Cells Uri Ben-David, Yoav Mayshar, and Nissim
Cell-to-Cell Variability and Culture Conditions during Self-Renewal Reversibly Affect Subsequent Differentiation of Mouse Embryonic Stem Cells
 Cell-to-Cell Variability and Culture Conditions during Self-Renewal Reversibly Affect Subsequent Differentiation of Mouse Embryonic Stem Cells by Jit Hin Tan Bachelor of Science in Chemical Engineering
Cell-to-Cell Variability and Culture Conditions during Self-Renewal Reversibly Affect Subsequent Differentiation of Mouse Embryonic Stem Cells by Jit Hin Tan Bachelor of Science in Chemical Engineering
Supplementary Figure 1.
 Supplementary Figure 1. Quantification of western blot analysis of fibroblasts (related to Figure 1) (A-F) Quantification of western blot analysis for control and IR-Mut fibroblasts. Data are expressed
Supplementary Figure 1. Quantification of western blot analysis of fibroblasts (related to Figure 1) (A-F) Quantification of western blot analysis for control and IR-Mut fibroblasts. Data are expressed
The bimodally expressed microrna mir-142 gates exit from pluripotency
 Article The bimodally expressed microrna mir-42 gates exit from pluripotency Hanna L Sladitschek & Pierre A Neveu * Abstract A stem cell s decision to self-renew or differentiate is thought to critically
Article The bimodally expressed microrna mir-42 gates exit from pluripotency Hanna L Sladitschek & Pierre A Neveu * Abstract A stem cell s decision to self-renew or differentiate is thought to critically
Quantitative analysis of Bidirectional Signaling (qbids)
 Quantitative analysis of Bidirectional Signaling (qbids) EphB2 + cells were labeled independently with light (C 12 N 14 ) arginine and lysine or heavy (C 13 N 15 ) arginine and lysine ephrin-b1 + cells
Quantitative analysis of Bidirectional Signaling (qbids) EphB2 + cells were labeled independently with light (C 12 N 14 ) arginine and lysine or heavy (C 13 N 15 ) arginine and lysine ephrin-b1 + cells
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Workflow for multimodal analysis using sc-gem on a programmable microfluidic device (Fluidigm). 1) Cells are captured and lysed, 2) RNA from lysed single cell is reverse-transcribed
Supplementary Figure 1 Workflow for multimodal analysis using sc-gem on a programmable microfluidic device (Fluidigm). 1) Cells are captured and lysed, 2) RNA from lysed single cell is reverse-transcribed
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Integrative Biology. This journal is The Royal Society of Chemistry 2015 Electronic Supplementary Information Table S1. Definition of quantitative cellular features
Electronic Supplementary Material (ESI) for Integrative Biology. This journal is The Royal Society of Chemistry 2015 Electronic Supplementary Information Table S1. Definition of quantitative cellular features
initial single-cell analysis, with a pragmatic focus on surface markers with the highest potential for
 Supplementary Figure 1: Summary of the exclusionary approach to surface marker selection for initial single-cell analysis, with a pragmatic focus on surface markers with the highest potential for protein
Supplementary Figure 1: Summary of the exclusionary approach to surface marker selection for initial single-cell analysis, with a pragmatic focus on surface markers with the highest potential for protein
Biology Open (2015): doi: /bio : Supplementary information
 Fig. S1. Verification of optimal conditions for the generation of LIF-dependent inscs Representative images (from five repeat experiments) of human primary fibroblasts undergoing reprogramming for 21 days
Fig. S1. Verification of optimal conditions for the generation of LIF-dependent inscs Representative images (from five repeat experiments) of human primary fibroblasts undergoing reprogramming for 21 days
mrna Sequencing Quality Control (V6)
 mrna Sequencing Quality Control (V6) Notes: the following analyses are based on 8 adult brains sequenced in USC and Yale 1. Error Rates The error rates of each sequencing cycle are reported for 120 tiles
mrna Sequencing Quality Control (V6) Notes: the following analyses are based on 8 adult brains sequenced in USC and Yale 1. Error Rates The error rates of each sequencing cycle are reported for 120 tiles
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature09944 Supplementary Figure 1. Establishing DNA sequence similarity thresholds for phylum and genus levels Sequence similarity distributions of pairwise alignments of 40 universal single
doi:10.1038/nature09944 Supplementary Figure 1. Establishing DNA sequence similarity thresholds for phylum and genus levels Sequence similarity distributions of pairwise alignments of 40 universal single
Directed Differentiation of Human Induced Pluripotent Stem Cells. into Fallopian Tube Epithelium
 Directed Differentiation of Human Induced Pluripotent Stem Cells into Fallopian Tube Epithelium Nur Yucer 1,2, Marie Holzapfel 2, Tilley Jenkins Vogel 2, Lindsay Lenaeus 1, Loren Ornelas 1, Anna Laury
Directed Differentiation of Human Induced Pluripotent Stem Cells into Fallopian Tube Epithelium Nur Yucer 1,2, Marie Holzapfel 2, Tilley Jenkins Vogel 2, Lindsay Lenaeus 1, Loren Ornelas 1, Anna Laury
SUPPLEMENTARY INFORMATION
 SUPPLEMENTRY INFORMTION DOI:.38/ncb Kdmb locus kb Long isoform Short isoform Long isoform Jmj XX PHD F-box LRR Short isoform XX PHD F-box LRR Target Vector 3xFlag Left H Neo Right H TK LoxP LoxP Kdmb Locus
SUPPLEMENTRY INFORMTION DOI:.38/ncb Kdmb locus kb Long isoform Short isoform Long isoform Jmj XX PHD F-box LRR Short isoform XX PHD F-box LRR Target Vector 3xFlag Left H Neo Right H TK LoxP LoxP Kdmb Locus
Nature Biotechnology: doi: /nbt.4103
 Supplementary Figure 1 Cell types, spatial context and gene expression patterns within the juvenile zebrafish brain. a. Number of cells per cluster from each of the six biological replicates of whole brains
Supplementary Figure 1 Cell types, spatial context and gene expression patterns within the juvenile zebrafish brain. a. Number of cells per cluster from each of the six biological replicates of whole brains
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11272 Supplementary Figure 1. Experimental platform for epigenetic reversion of murine EpiSCs back to naïve ground state pluripotency. a, NGFP1 naïve-ipscs (40XY) expanded in 2i/LIF defined
doi:10.1038/nature11272 Supplementary Figure 1. Experimental platform for epigenetic reversion of murine EpiSCs back to naïve ground state pluripotency. a, NGFP1 naïve-ipscs (40XY) expanded in 2i/LIF defined
The bimodally expressed microrna mir-142 gates exit from pluripotency
 The bimodally expressed microrna mir-142 gates exit from pluripotency Hanna L. Sladitschek and Pierre A. Neveu Corresponding author: Pierre A. Neveu, European Molecular Biology Laboratory Review timeline:
The bimodally expressed microrna mir-142 gates exit from pluripotency Hanna L. Sladitschek and Pierre A. Neveu Corresponding author: Pierre A. Neveu, European Molecular Biology Laboratory Review timeline:
Decoding the Pluripotency Network: The Emergence of New Transcription Factors
 Biomedicines 2013, 1, 49-78; doi:10.3390/biomedicines1010049 OPEN ACCESS biomedicines ISSN 2075-4426 www.mdpi.com/journal/biomedicines/ Review Decoding the Pluripotency Network: The Emergence of New Transcription
Biomedicines 2013, 1, 49-78; doi:10.3390/biomedicines1010049 OPEN ACCESS biomedicines ISSN 2075-4426 www.mdpi.com/journal/biomedicines/ Review Decoding the Pluripotency Network: The Emergence of New Transcription
Nature Methods: doi: /nmeth Supplementary Figure 1. Pilot CrY2H-seq experiments to confirm strain and plasmid functionality.
 Supplementary Figure 1 Pilot CrY2H-seq experiments to confirm strain and plasmid functionality. (a) RT-PCR on HIS3 positive diploid cell lysate containing known interaction partners AT3G62420 (bzip53)
Supplementary Figure 1 Pilot CrY2H-seq experiments to confirm strain and plasmid functionality. (a) RT-PCR on HIS3 positive diploid cell lysate containing known interaction partners AT3G62420 (bzip53)
Human induced Pluripotent Stem Cell (ipsc) Line: GM23225*B
 Certificate of Analysis NIGMS Human tic Cell Repository Human induced Pluripotent Stem Cell (ipsc) Line: GM23225*B Diagnosis Mutation Reprogramming method Publication(s) describing ipsc line establishment
Certificate of Analysis NIGMS Human tic Cell Repository Human induced Pluripotent Stem Cell (ipsc) Line: GM23225*B Diagnosis Mutation Reprogramming method Publication(s) describing ipsc line establishment
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/259/ra5/dc1 Supplementary Materials for Systems Biology Approach Identifies the Kinase Csnk1a1 as a Regulator of the DNA Damage Response in Embryonic Stem Cells
www.sciencesignaling.org/cgi/content/full/6/259/ra5/dc1 Supplementary Materials for Systems Biology Approach Identifies the Kinase Csnk1a1 as a Regulator of the DNA Damage Response in Embryonic Stem Cells
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11435 Supplementary Figures Figure S1. The scheme of androgenetic haploid embryonic stem (ahes) cells derivation. WWW.NATURE.COM/NATURE 1 RESEARCH SUPPLEMENTARY INFORMATION Figure S2.
doi:10.1038/nature11435 Supplementary Figures Figure S1. The scheme of androgenetic haploid embryonic stem (ahes) cells derivation. WWW.NATURE.COM/NATURE 1 RESEARCH SUPPLEMENTARY INFORMATION Figure S2.
Supplementary Data. Supplementary Methods Three-step protocol for spontaneous differentiation of mouse induced pluripotent stem (embryonic stem) cells
 Supplementary Data Supplementary Methods Three-step protocol for spontaneous differentiation of mouse induced pluripotent stem (embryonic stem) cells Mouse induced pluripotent stem cells (ipscs) were cultured
Supplementary Data Supplementary Methods Three-step protocol for spontaneous differentiation of mouse induced pluripotent stem (embryonic stem) cells Mouse induced pluripotent stem cells (ipscs) were cultured
INTESTINAL ORGANOIDS. IntestiCult and STEMdiff Media for Intestinal Organoid Culture. Scientists Helping Scientists
 INTESTINAL ORGANOIDS IntestiCult and STEMdiff Media for Intestinal Organoid Culture Scientists Helping Scientists WWW.STEMCELL.COM CULTURING INTESTINAL ORGANOIDS TABLE OF CONTENTS 3 Introduction 4 IntestiCult
INTESTINAL ORGANOIDS IntestiCult and STEMdiff Media for Intestinal Organoid Culture Scientists Helping Scientists WWW.STEMCELL.COM CULTURING INTESTINAL ORGANOIDS TABLE OF CONTENTS 3 Introduction 4 IntestiCult
Lecture 8: Predicting and analyzing metagenomic composition from 16S survey data
 Lecture 8: Predicting and analyzing metagenomic composition from 16S survey data What can we tell about the taxonomic and functional stability of microbiota? Why? Nature. 2012; 486(7402): 207 214. doi:10.1038/nature11234
Lecture 8: Predicting and analyzing metagenomic composition from 16S survey data What can we tell about the taxonomic and functional stability of microbiota? Why? Nature. 2012; 486(7402): 207 214. doi:10.1038/nature11234
Nature Biotechnology: doi: /nbt.4086
 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3
Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3 p815 p815-hcd2 Ag (-) anti-cd3
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1. Functional assays of hepatocyte-like cells induced from H1 hescs (a) P450 and AAT staining, PAS staining, LDL uptake assay, and ICG uptake and release assay
Supplementary Figure 1 Supplementary Figure 1. Functional assays of hepatocyte-like cells induced from H1 hescs (a) P450 and AAT staining, PAS staining, LDL uptake assay, and ICG uptake and release assay
SUPPLEMENTARY INFORMATION
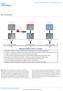 DOI: 10.1038/ncb2239 Hepatocytes (Endoderm) Foreskin fibroblasts (Mesoderm) Melanocytes (Ectoderm) +Dox rtta TetO CMV O,S,M,K & N H1 hes H7 hes H9 hes H1 hes-derived EBs H7 hes-derived EBs H9 hes-derived
DOI: 10.1038/ncb2239 Hepatocytes (Endoderm) Foreskin fibroblasts (Mesoderm) Melanocytes (Ectoderm) +Dox rtta TetO CMV O,S,M,K & N H1 hes H7 hes H9 hes H1 hes-derived EBs H7 hes-derived EBs H9 hes-derived
Electromagnetic Fields Mediate Efficient Cell Reprogramming Into a Pluripotent State
 Electromagnetic Fields Mediate Efficient Cell Reprogramming Into a Pluripotent State Soonbong Baek, 1 Xiaoyuan Quan, 1 Soochan Kim, 2 Christopher Lengner, 3 Jung- Keug Park, 4 and Jongpil Kim 1* 1 Lab
Electromagnetic Fields Mediate Efficient Cell Reprogramming Into a Pluripotent State Soonbong Baek, 1 Xiaoyuan Quan, 1 Soochan Kim, 2 Christopher Lengner, 3 Jung- Keug Park, 4 and Jongpil Kim 1* 1 Lab
Supplementary Data. Generation and Characterization of Mixl1- Inducible Embryonic Stem Cells Under the Control of Oct3/4 Promoter or CAG Promoter
 Supplementary Data Generation and Characterization of Mixl1- Inducible Embryonic Stem Cells Under the Control of Oct3/4 Promoter or CAG Promoter We first introduced an expression unit composed of a tetresponse
Supplementary Data Generation and Characterization of Mixl1- Inducible Embryonic Stem Cells Under the Control of Oct3/4 Promoter or CAG Promoter We first introduced an expression unit composed of a tetresponse
A molecular roadmap for the emergence of early-embryonic-like cells in culture
 SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41588-017-0016-5 In the format provided by the authors and unedited. A molecular roadmap for the emergence of early-embryonic-like cells in culture
SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41588-017-0016-5 In the format provided by the authors and unedited. A molecular roadmap for the emergence of early-embryonic-like cells in culture
HUMAN EMBRYONIC STEM CELL (hesc) ASSESSMENT BY REAL-TIME POLYMERASE CHAIN REACTION (PCR)
 HUMAN EMBRYONIC STEM CELL (hesc) ASSESSMENT BY REAL-TIME POLYMERASE CHAIN REACTION (PCR) OBJECTIVE: is performed to analyze gene expression of human Embryonic Stem Cells (hescs) in culture. Real-Time PCR
HUMAN EMBRYONIC STEM CELL (hesc) ASSESSMENT BY REAL-TIME POLYMERASE CHAIN REACTION (PCR) OBJECTIVE: is performed to analyze gene expression of human Embryonic Stem Cells (hescs) in culture. Real-Time PCR
Supplementary Figure 1. Details of the component modelling and steps in generating and validating a PTGS. The steps (boxes) and the flow of
 Supplementary Figure 1. Details of the component modelling and steps in generating and validating a PTGS. The steps (boxes) and the flow of information (arrows) through the modelling process, including
Supplementary Figure 1. Details of the component modelling and steps in generating and validating a PTGS. The steps (boxes) and the flow of information (arrows) through the modelling process, including
Supplementary Data. Supplementary Materials and Methods Twenty-cell gene expression profiling. Stem cell transcriptome analysis
 Supplementary Data The Supplementary Data includes Supplementary Materials and Methods, and Figures (Supplementary Figs. S1 S10). Supplementary Materials and Methods Twenty-cell gene expression profiling
Supplementary Data The Supplementary Data includes Supplementary Materials and Methods, and Figures (Supplementary Figs. S1 S10). Supplementary Materials and Methods Twenty-cell gene expression profiling
Expanded View Figures
 Vasco Meneghini et al Effective lentiviral GT in NHP brains EMO Molecular Medicine Expanded View Figures Figure EV1. VN and GFP distribution in LV.GFPinjected NHPs. Vector copy number (VN) cartography
Vasco Meneghini et al Effective lentiviral GT in NHP brains EMO Molecular Medicine Expanded View Figures Figure EV1. VN and GFP distribution in LV.GFPinjected NHPs. Vector copy number (VN) cartography
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Generation of NSCs from hpscs in SDC medium.
 Supplementary Figure 1 Generation of NSCs from hpscs in SDC medium. (a) Q-PCR of pluripotent markers (OCT4, NANOG), neural markers (SOX1, PAX6, N-Cadherin), markers for the other germ layers (T, EOMES,
Supplementary Figure 1 Generation of NSCs from hpscs in SDC medium. (a) Q-PCR of pluripotent markers (OCT4, NANOG), neural markers (SOX1, PAX6, N-Cadherin), markers for the other germ layers (T, EOMES,
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full//7/e6/dc Supplementary Materials for Heterogeneity of cellular circadian clocks in intact plants and its correction under light-dark cycles Tomoaki Muranaka and
advances.sciencemag.org/cgi/content/full//7/e6/dc Supplementary Materials for Heterogeneity of cellular circadian clocks in intact plants and its correction under light-dark cycles Tomoaki Muranaka and
TITLE: Identification of different mechanisms leading to PAX6 down-regulation, as potential
 TITLE: Identification of different mechanisms leading to PAX6 down-regulation, as potential events contributing to the onset of Hirschsprung disease AUTHOR: María Valle Enguix-Riego 1,2,#, Ana Torroglosa
TITLE: Identification of different mechanisms leading to PAX6 down-regulation, as potential events contributing to the onset of Hirschsprung disease AUTHOR: María Valle Enguix-Riego 1,2,#, Ana Torroglosa
Disease Clinical features Genotype
 Disease Clinical features Genotype Alpha 1 Antitrypsin deficiency (A1ATD) Patient 1 65 year old caucasian male, liver transplant recipient, explant biopsy confirmed A1ATD induced liver cirrhosis Patient
Disease Clinical features Genotype Alpha 1 Antitrypsin deficiency (A1ATD) Patient 1 65 year old caucasian male, liver transplant recipient, explant biopsy confirmed A1ATD induced liver cirrhosis Patient
Knowledge-Guided Analysis with KnowEnG Lab
 Han Sinha Song Weinshilboum Knowledge-Guided Analysis with KnowEnG Lab KnowEnG Center Powerpoint by Charles Blatti Knowledge-Guided Analysis KnowEnG Center 2017 1 Exercise In this exercise we will be doing
Han Sinha Song Weinshilboum Knowledge-Guided Analysis with KnowEnG Lab KnowEnG Center Powerpoint by Charles Blatti Knowledge-Guided Analysis KnowEnG Center 2017 1 Exercise In this exercise we will be doing
Long-term dynamics of CA1 hippocampal place codes
 Long-term dynamics of CA1 hippocampal place codes Yaniv Ziv, Laurie D. Burns, Eric D. Cocker, Elizabeth O. Hamel, Kunal K. Ghosh, Lacey J. Kitch, Abbas El Gamal, and Mark J. Schnitzer Supplementary Fig.
Long-term dynamics of CA1 hippocampal place codes Yaniv Ziv, Laurie D. Burns, Eric D. Cocker, Elizabeth O. Hamel, Kunal K. Ghosh, Lacey J. Kitch, Abbas El Gamal, and Mark J. Schnitzer Supplementary Fig.
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 An extended version of Figure 2a, depicting multi-model training and reverse-complement mode To use the GPU s full computational power, we train several independent models in parallel
Supplementary Figure 1 An extended version of Figure 2a, depicting multi-model training and reverse-complement mode To use the GPU s full computational power, we train several independent models in parallel
Nature Biotechnology: doi: /nbt Supplementary Figure 1. sndrop-seq overview.
 Supplementary Figure 1 sndrop-seq overview. A. sndrop-seq method showing modifications needed to process nuclei, including bovine serum albumin (BSA) coating and droplet heating to ensure complete nuclear
Supplementary Figure 1 sndrop-seq overview. A. sndrop-seq method showing modifications needed to process nuclei, including bovine serum albumin (BSA) coating and droplet heating to ensure complete nuclear
SUPPLEMENTARY INFORMATION
 Supplementary Discussion Interestingly, a recent study demonstrated that knockdown of Tet1 alone would lead to dysregulation of differentiation genes, but only minor defects in ES maintenance and no change
Supplementary Discussion Interestingly, a recent study demonstrated that knockdown of Tet1 alone would lead to dysregulation of differentiation genes, but only minor defects in ES maintenance and no change
DEPArray Technology. Sorting and Recovery of Rare Cells
 DEPArray Technology Sorting and Recovery of Rare Cells Delivering pure, single, viable cells The DEPArray system from Silicon Biosystems is the only automated instrument that can identify, quantify, and
DEPArray Technology Sorting and Recovery of Rare Cells Delivering pure, single, viable cells The DEPArray system from Silicon Biosystems is the only automated instrument that can identify, quantify, and
Supplementary Fig. 5
 Supplementary Fig. 5 Supplemental Figures legends Supplementary Figure 1 (A) Additional dot plots from CyTOF analysis from untreated group. (B) Gating strategy for assessment of CD11c + NK cells frequency
Supplementary Fig. 5 Supplemental Figures legends Supplementary Figure 1 (A) Additional dot plots from CyTOF analysis from untreated group. (B) Gating strategy for assessment of CD11c + NK cells frequency
Supplementary Table, Figures and Videos
 Supplementary Table, Figures and Videos Table S1. Oligonucleotides used for different approaches. (A) RT-qPCR study. (B) qpcr study after ChIP assay. (C) Probes used for EMSA. Figure S1. Notch activation
Supplementary Table, Figures and Videos Table S1. Oligonucleotides used for different approaches. (A) RT-qPCR study. (B) qpcr study after ChIP assay. (C) Probes used for EMSA. Figure S1. Notch activation
SUPPLEMENTARY INFORMATION
 doi:.38/nature08267 Figure S: Karyotype analysis of ips cell line One hundred individual chromosomal spreads were counted for each of the ips samples. Shown here is an spread at passage 5, predominantly
doi:.38/nature08267 Figure S: Karyotype analysis of ips cell line One hundred individual chromosomal spreads were counted for each of the ips samples. Shown here is an spread at passage 5, predominantly
Result Tables The Result Table, which indicates chromosomal positions and annotated gene names, promoter regions and CpG islands, is the best way for
 Result Tables The Result Table, which indicates chromosomal positions and annotated gene names, promoter regions and CpG islands, is the best way for you to discover methylation changes at specific genomic
Result Tables The Result Table, which indicates chromosomal positions and annotated gene names, promoter regions and CpG islands, is the best way for you to discover methylation changes at specific genomic
ChIPXpress: using publicly available gene expression data to improve ChIP-seq and ChIPchip target gene ranking
 Wu and Ji BMC Bioinformatics 2013, 14:188 METHODOLOGY ARTICLE Open Access ChIPXpress: using publicly available gene expression data to improve ChIP-seq and ChIPchip target gene ranking George Wu and Hongkai
Wu and Ji BMC Bioinformatics 2013, 14:188 METHODOLOGY ARTICLE Open Access ChIPXpress: using publicly available gene expression data to improve ChIP-seq and ChIPchip target gene ranking George Wu and Hongkai
David M. Rocke Division of Biostatistics and Department of Biomedical Engineering University of California, Davis
 David M. Rocke Division of Biostatistics and Department of Biomedical Engineering University of California, Davis Outline RNA-Seq for differential expression analysis Statistical methods for RNA-Seq: Structure
David M. Rocke Division of Biostatistics and Department of Biomedical Engineering University of California, Davis Outline RNA-Seq for differential expression analysis Statistical methods for RNA-Seq: Structure
Heterogeneities in Nanog Expression Drive Stable Commitment to Pluripotency in the Mouse Blastocyst
 Article Heterogeneities in Nanog Expression Drive Stable Commitment to Pluripotency in the Mouse Blastocyst Graphical Abstract Authors Panagiotis Xenopoulos, Minjung Kang,..., Stefano Di Talia, Anna-Katerina
Article Heterogeneities in Nanog Expression Drive Stable Commitment to Pluripotency in the Mouse Blastocyst Graphical Abstract Authors Panagiotis Xenopoulos, Minjung Kang,..., Stefano Di Talia, Anna-Katerina
Figure S2. Response of mouse ES cells to GSK3 inhibition. Mentioned in discussion
 Stem Cell Reports, Volume 1 Supplemental Information Robust Self-Renewal of Rat Embryonic Stem Cells Requires Fine-Tuning of Glycogen Synthase Kinase-3 Inhibition Yaoyao Chen, Kathryn Blair, and Austin
Stem Cell Reports, Volume 1 Supplemental Information Robust Self-Renewal of Rat Embryonic Stem Cells Requires Fine-Tuning of Glycogen Synthase Kinase-3 Inhibition Yaoyao Chen, Kathryn Blair, and Austin
Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat
 Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat units in the human genome. Annotated transposable elements
Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat units in the human genome. Annotated transposable elements
Edinburgh Research Explorer
 Edinburgh Research Explorer Esrrb extinction triggers dismantling of naïve pluripotency and marks commitment to differentiation Citation for published version: Festuccia, N, Halbritter, F, Corsinotti,
Edinburgh Research Explorer Esrrb extinction triggers dismantling of naïve pluripotency and marks commitment to differentiation Citation for published version: Festuccia, N, Halbritter, F, Corsinotti,
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
SUPPLEMENTARY INFORMATION
 18Pura Brain Liver 2 kb 9Mtmr Heart Brain 1.4 kb 15Ppara 1.5 kb IPTpcc 1.3 kb GAPDH 1.5 kb IPGpr1 2 kb GAPDH 1.5 kb 17Tcte 13Elov12 Lung Thymus 6 kb 5 kb 17Tbcc Kidney Embryo 4.5 kb 3.7 kb GAPDH 1.5 kb
18Pura Brain Liver 2 kb 9Mtmr Heart Brain 1.4 kb 15Ppara 1.5 kb IPTpcc 1.3 kb GAPDH 1.5 kb IPGpr1 2 kb GAPDH 1.5 kb 17Tcte 13Elov12 Lung Thymus 6 kb 5 kb 17Tbcc Kidney Embryo 4.5 kb 3.7 kb GAPDH 1.5 kb
NIA Aging Cell Repository
 Certificate of Analysis NIA Aging Cell Repository Human induced Pluripotent Stem Cell (ipsc) Line: AG25370*B Diagnosis Alzheimer Disease, Familial, Type 4 Mutation Reprogramming method Publication(s) describing
Certificate of Analysis NIA Aging Cell Repository Human induced Pluripotent Stem Cell (ipsc) Line: AG25370*B Diagnosis Alzheimer Disease, Familial, Type 4 Mutation Reprogramming method Publication(s) describing
Comprehensive Cell Surface Protein Profiling Identifies Specific Markers of Human Naive and Primed Pluripotent States
 Comprehensive Cell Surface Protein Profiling Identifies Specific Markers of Human Naive and Primed Pluripotent States Resource Graphical Abstract Cell surface screen in hpscs 486 antibodies Primed cell
Comprehensive Cell Surface Protein Profiling Identifies Specific Markers of Human Naive and Primed Pluripotent States Resource Graphical Abstract Cell surface screen in hpscs 486 antibodies Primed cell
Systematic comparison of CRISPR/Cas9 and RNAi screens for essential genes
 CORRECTION NOTICE Nat. Biotechnol. doi:10.1038/nbt. 3567 Systematic comparison of CRISPR/Cas9 and RNAi screens for essential genes David W Morgens, Richard M Deans, Amy Li & Michael C Bassik In the version
CORRECTION NOTICE Nat. Biotechnol. doi:10.1038/nbt. 3567 Systematic comparison of CRISPR/Cas9 and RNAi screens for essential genes David W Morgens, Richard M Deans, Amy Li & Michael C Bassik In the version
a previous report, Lee and colleagues showed that Oct4 and Sox2 bind to DXPas34 and Xite
 SUPPLEMENTARY INFORMATION doi:1.138/nature9496 a 1 Relative RNA levels D12 female ES cells 24h Control sirna 24h sirna Oct4 5 b,4 Oct4 Xist (wt) Xist (mut) Tsix 3' (wt) Tsix 5' (wt) Beads Oct4,2 c,2 Xist
SUPPLEMENTARY INFORMATION doi:1.138/nature9496 a 1 Relative RNA levels D12 female ES cells 24h Control sirna 24h sirna Oct4 5 b,4 Oct4 Xist (wt) Xist (mut) Tsix 3' (wt) Tsix 5' (wt) Beads Oct4,2 c,2 Xist
Single-Cell Profiling of Epigenetic Modifiers Identifies PRDM14 as an Inducer of Cell Fate in the Mammalian Embryo
 Cell Reports Article Single-Cell Profiling of Epigenetic Modifiers Identifies PRDM14 as an Inducer of Cell Fate in the Mammalian Embryo Adam Burton, 1,6 Julius Muller, 2,6 Shengjiang Tu, 4 Pablo Padilla-Longoria,
Cell Reports Article Single-Cell Profiling of Epigenetic Modifiers Identifies PRDM14 as an Inducer of Cell Fate in the Mammalian Embryo Adam Burton, 1,6 Julius Muller, 2,6 Shengjiang Tu, 4 Pablo Padilla-Longoria,
Supplementary Figure 1. Characterization of hipscs derived from primary human fibroblasts. a,b. Morphology of hipscs. hipscs exhibit hesc-like
 Supplementary Figure 1. Characterization of hipscs derived from primary human fibroblasts. a,b. Morphology of hipscs. hipscs exhibit hesc-like morphology in co-culture with mouse embryonic feeder fibroblasts
Supplementary Figure 1. Characterization of hipscs derived from primary human fibroblasts. a,b. Morphology of hipscs. hipscs exhibit hesc-like morphology in co-culture with mouse embryonic feeder fibroblasts
Nanog-Luc. R-Luc. 40 HA-Klf Relative protein level of Klf USP21. Vector USP2 USP21 *** *** *** ***
 :Flag Relative protein level of Relative protein level of : Flag Vec 21LV 21SV 2 : HA HA- HA- HA- HA-Klf4 Relative mrna expression Relative protein level of Relative protein level of Relative protein level
:Flag Relative protein level of Relative protein level of : Flag Vec 21LV 21SV 2 : HA HA- HA- HA- HA-Klf4 Relative mrna expression Relative protein level of Relative protein level of Relative protein level
A Late Transition in Somatic Cell Reprogramming Requires Regulators Distinct from the Pluripotency Network
 Article A Late Transition in Somatic Cell Reprogramming Requires Regulators Distinct from the Pluripotency Network Azadeh Golipour, 1,, Laurent David,,, Yu Liu, 1, Gowtham Jayakumaran, 1, Calley L. Hirsch,
Article A Late Transition in Somatic Cell Reprogramming Requires Regulators Distinct from the Pluripotency Network Azadeh Golipour, 1,, Laurent David,,, Yu Liu, 1, Gowtham Jayakumaran, 1, Calley L. Hirsch,
PI3K/mTORC2 regulates TGFβ/Activin signalling by modulating Smad2/3
 Supplementary Information PI3K/mTORC2 regulates TGFβ/Activin signalling by modulating Smad2/3 activity via linker phosphorylation Jason S.L. Yu, 1 Thamil Selvee Ramasamy, 1,2 Nick Murphy, 1 Marie Holt,
Supplementary Information PI3K/mTORC2 regulates TGFβ/Activin signalling by modulating Smad2/3 activity via linker phosphorylation Jason S.L. Yu, 1 Thamil Selvee Ramasamy, 1,2 Nick Murphy, 1 Marie Holt,
Figure S1, related to Figure 1. Characterization of biosensor behavior in vivo.
 Developmental Cell, Volume 23 Supplemental Information Separase Biosensor Reveals that Cohesin Cleavage Timing Depends on Phosphatase PP2A Cdc55 Regulation Gilad Yaakov, Kurt Thorn, and David O. Morgan
Developmental Cell, Volume 23 Supplemental Information Separase Biosensor Reveals that Cohesin Cleavage Timing Depends on Phosphatase PP2A Cdc55 Regulation Gilad Yaakov, Kurt Thorn, and David O. Morgan
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Characterization of Hi-C/CHi-C dynamics and enhancer identification. (a) Scatterplot of Hi-C read counts supporting contacts between domain boundaries. Contacts enclosing domains
Supplementary Figure 1 Characterization of Hi-C/CHi-C dynamics and enhancer identification. (a) Scatterplot of Hi-C read counts supporting contacts between domain boundaries. Contacts enclosing domains
Supplemental Information. T Follicular Helper Cell-Germinal Center B Cell. Interaction Strength Regulates Entry into Plasma
 Immunity, Volume 48 Supplemental Information T Follicular Helper ell-germinal enter B ell Interaction Strength Regulates Entry into Plasma ell or Recycling Germinal enter ell Fate Wataru Ise, Kentaro Fujii,
Immunity, Volume 48 Supplemental Information T Follicular Helper ell-germinal enter B ell Interaction Strength Regulates Entry into Plasma ell or Recycling Germinal enter ell Fate Wataru Ise, Kentaro Fujii,
The role of FGF/Erk signaling in pluripotent cells
 REVIEW 3351 Development 137, 3351-3360 (2010) doi:10.1242/dev.050146 2010. Published by The Company of Biologists Ltd The role of FGF/Erk signaling in pluripotent cells Fredrik Lanner 1 and Janet Rossant
REVIEW 3351 Development 137, 3351-3360 (2010) doi:10.1242/dev.050146 2010. Published by The Company of Biologists Ltd The role of FGF/Erk signaling in pluripotent cells Fredrik Lanner 1 and Janet Rossant
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Optimization of Patch-seq protocol.
 Supplementary Figure 1 Optimization of Patch-seq protocol. Pilot experiments were carried out to test the effect of various protocol modifications on cdna yield, including addition of RNase inhibitor to
Supplementary Figure 1 Optimization of Patch-seq protocol. Pilot experiments were carried out to test the effect of various protocol modifications on cdna yield, including addition of RNase inhibitor to
Supplemental material JCB. Pitaval et al., Cell shape and ciliogenesis Pitaval et al.
 Supplemental material THE JOURNAL OF CELL BIOLOGY Pitaval et al., http://www.jcb.org/cgi/content/full/jcb.201004003/dc1 S1 Figure S1. Cell cycle exit analysis. (A) Experimental procedure. RPE1 cells were
Supplemental material THE JOURNAL OF CELL BIOLOGY Pitaval et al., http://www.jcb.org/cgi/content/full/jcb.201004003/dc1 S1 Figure S1. Cell cycle exit analysis. (A) Experimental procedure. RPE1 cells were
Fundamental properties of Stem Cells
 Stem cells Learning Goals: Define what a stem cell is and describe its general properties, using hematopoietic stem cells as an example. Describe to a non-scientist the current progress of human stem cell
Stem cells Learning Goals: Define what a stem cell is and describe its general properties, using hematopoietic stem cells as an example. Describe to a non-scientist the current progress of human stem cell
Upstream/Downstream Relation Detection of Signaling Molecules using Microarray Data
 Vol 1 no 1 2005 Pages 1 5 Upstream/Downstream Relation Detection of Signaling Molecules using Microarray Data Ozgun Babur 1 1 Center for Bioinformatics, Computer Engineering Department, Bilkent University,
Vol 1 no 1 2005 Pages 1 5 Upstream/Downstream Relation Detection of Signaling Molecules using Microarray Data Ozgun Babur 1 1 Center for Bioinformatics, Computer Engineering Department, Bilkent University,
Esrrb extinction triggers dismantling of naïve pluripotency and marks commitment to differentiation
 Esrrb extinction triggers dismantling of naïve pluripotency and marks commitment to differentiation Nicola Festuccia, Florian Halbritter, Andrea Corsinotti, Alessia Gagliardi, Douglas Colby, Simon R. Tomlinson
Esrrb extinction triggers dismantling of naïve pluripotency and marks commitment to differentiation Nicola Festuccia, Florian Halbritter, Andrea Corsinotti, Alessia Gagliardi, Douglas Colby, Simon R. Tomlinson
Nature Genetics: doi: /ng Supplementary Figure 1. Neighbor-joining tree of the 183 wild, cultivated, and weedy rice accessions.
 Supplementary Figure 1 Neighbor-joining tree of the 183 wild, cultivated, and weedy rice accessions. Relationships of cultivated and wild rice correspond to previously observed relationships 40. Wild rice
Supplementary Figure 1 Neighbor-joining tree of the 183 wild, cultivated, and weedy rice accessions. Relationships of cultivated and wild rice correspond to previously observed relationships 40. Wild rice
Modeling cell fate specification during early embryonic development in mouse
 Université Libre de Bruxelles Modeling cell fate specification during early embryonic development in mouse Didier Gonze Unité de Chronobiologie Théorique Faculté des Sciences Université Libre de Bruxelles
Université Libre de Bruxelles Modeling cell fate specification during early embryonic development in mouse Didier Gonze Unité de Chronobiologie Théorique Faculté des Sciences Université Libre de Bruxelles
Figure S1. nuclear extracts. HeLa cell nuclear extract. Input IgG IP:ORC2 ORC2 ORC2. MCM4 origin. ORC2 occupancy
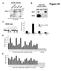 A nuclear extracts B HeLa cell nuclear extract Figure S1 ORC2 (in kda) 21 132 7 ORC2 Input IgG IP:ORC2 32 ORC C D PRKDC ORC2 occupancy Directed against ORC2 C-terminus (sc-272) MCM origin 2 2 1-1 -1kb
A nuclear extracts B HeLa cell nuclear extract Figure S1 ORC2 (in kda) 21 132 7 ORC2 Input IgG IP:ORC2 32 ORC C D PRKDC ORC2 occupancy Directed against ORC2 C-terminus (sc-272) MCM origin 2 2 1-1 -1kb
Gene Expression Data Analysis (I)
 Gene Expression Data Analysis (I) Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Bioinformatics tasks Biological question Experiment design Microarray experiment
Gene Expression Data Analysis (I) Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Bioinformatics tasks Biological question Experiment design Microarray experiment
