Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency.
|
|
|
- Brenda Walker
- 5 years ago
- Views:
Transcription
1 Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency. (a) Transfection of different concentration of GAS5-overexpressing plasmids to hescs. The mrna level of pluripotency gene GAS5, Oct4, Nanog and Sox2 are detected using qpcr. The concentration indicated represents the amount of plasmids transfected into each 6-well. **P<0.01, t-test, n=3. (b) Testing the specificity of GAS5 knockdown using the indicated sirna. KLF4 and GATA3 were predicted to bind none of the sirna. **P<0.01, t-test, n=3. (c) The relative GAS5 RNA level in the cytoplasm and nuclear assessed by qpcr. MTesR1 medium represents undifferentiated state, while bfgf and MEF medium were used to generate differentiation of hescs. **P<0.01, t-test, n=3. (d) FISH analysis of GAS5 in HEK-293T cells and lncrna DANCR (validated nuclear enriched) in hescs (n=2). The analysis of relative transcript in the subcellular level using nuclear-plasma separation were shown in the right panel. The scale bar represents 100nm. **P<0.01,
2 t-test, n=2. (e) The effect of pluripotency genes overexpression on GAS5 RNA level. **P<0.01, t-test, n=3. (f) The inactive histone modification state of the predicted Gas5 promoter region using the ENCODE Ch-IP sequencing data. (g) The gel electrophoresis image of ch-ip assay in Fig. 1i. (h) The gel electrophoresis image of Ch-IP analysis of the binding of OCT4 and SOX2 to Nanog promoter, which served as a positive control for Ch-IP results presented in Fig.1i. (i) EMSA of OCT4 (left panel) and SOX2 (right panel) binding using the probe indicated in Fig. 1h (n=2). The probe designed in the GAS5-3 region was used to detect the binding of SOX2, and probe designed in the GAS5-4 region was used to detect the binding of OCT4 (related to Fig. 1h). Error bars represent standard deviation of the indicated experiment replicates. RNA level of β-actin served as internal reference for qpcr.
3 Supplementary Figure 2, related to Figure 2. GAS5 is Essential for hesc Self-renewal. (a) The expressing efficacy of different stable lentivirus treated cells. **P<0.01, t-test, n=3. (b) The light microscopy showing the growing state of different stably lentivirus treated hescs (upper panel). The lower panel shows the embryo bodies formed by different stably expressed hescs. Lenti-NC represents control lentivirus, Lenti-GAS5 represents GAS5 overexpressing lentivirus, and Lenti-shGAS5 represents short
4 hairpin knockdown lentivirus, n=2. (c) QPCR analysis to measure the effect of GAS5 knockdown in GAS5 stably overexpressed hescs, and GAS5 overexpression in GAS5 stably knockdown hescs on the expression level of pluripotency genes. **P<0.01, t-test, n=3. (d) Quantification of cell cycle analysis to show the effect of GAS5 knockdown in GAS5 stably overexpressed hescs, and GAS5 overexpression in GAS5 stably knockdown hescs on the proliferation of hescs. The percentage of G2 phase cells were statistically analyzed, **P<0.01, t-test, n=3. (e) Cell cycle synchronization assay (n=2) performed in different stably expressed hescs. The cells were treated with NOCODAZOL at 100ng/ml for 12h. 2h before experiment, the cells are washed In PBS and changed for new medium. (f) The analysis of markers of three germ layers in different stably expressed hescs. **P<0.01, t-test, n=3. (g) Alkaline phosphatase staining of different stably expressed hescs in different culture medium (n=2). NC represents normal hescs culture, -bfgf represents using medium that withdraw bfgf, MEF represents using murine embryonic fibroblast (MEF) culture medium, and CM represents using conditioned hescs medium produced by MEF. Error bars represent standard deviation of the indicated experiment replicates. RNA level of β-actin served as internal reference for qpcr.
5 Supplementary Figure 3, related to Figure 3. Global mrna analysis of GAS5-regulated hescs revealed NODAL-SMAD signaling is activated by GAS5 in hescs. (a) Hierarchical clustering showing differentially expressed genes in stably GAS5 overexpressed or knockdown hesc cells. Red color indicates higher expression, whereas green color indicates lower expression. (b) The chart shows the Gene Ontology analysis of down-regulated genes in GAS5 knockdown group indicated in Fig. 3a. (c) The signaling network analysis based on the sequencing data showed that TGFβ and Wnt signaling is up-regulated during GAS5 overexpression, whereas genes
6 involved in basal cell carcinoma is lowered. However, genes involved in Hedgehog signaling showed bilateral change. (d) The graph depicts the number of inversely expressed genes with GAS5 which generated form the sequencing. (e) Gene Ontology analysis of the genes in inverse correlation with GAS5. Each chart represent gene groups indicated in (d). (f) The effect of rhnodal treatment on NODAL signaling related genes. SB431542, an ALK4/7 inhibitor that blocks NODAL and Activin mediated signal transduction. **P<0.01, t-test, n=3. (g) Western blot (n=3) showing the effect of GAS5 on the protein amount of signaling regulators AKT and ERK, and their phosphorylated form pakt and perk. (h) GSEA gene signature analysis of SMAD binding genes in microarray or high through-put sequencing data that involve NODAL or Activin treatment. SB431542, an ALK4/7 inhibitor that blocks NODAL and Activin mediated signal transduction. Related to Fig. 3g. Error bars represent standard deviation of the indicated experiment replicates. RNA level of β-actin served as internal reference for qpcr.
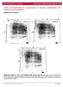
7 Supplementary Figure 4, related to Figure 4. NODAL-Signaling is required for the GAS5-promoted Self-renewal of hescs. (a) Quantification of colony formation assay in Fig. 4b. **P<0.01, t-test, n=3. (b) The effect of different doses of SB and NODAL on pluripotency genes and GAS5 expressions in hescs. (c) The analysis of the mrna level of pluripotency genes in GAS5 overexpressed or control hescs during different doses of fetal bovine serum (FBS) induced differentiation. The bar graph showed that increased doses of FBS (per 12-well) induced down-regulation of the expression of pluriportency genes in either GAS5 overexpressed or control hescs. *P<0.05, **P<0.01, t-test, n=3. (d) qpcr analysis of pluripotency genes in GAS5 or GAS5+rhNODAL treated group compared with negative control (empty vector) group. **P<0.01, t-test, n=3. Error bars represent standard deviation of the indicated experiment replicates. RNA level of β-actin served as internal reference for qpcr.
8 Supplementary Figure 5, related to Figure 5. NODAL is Post-transcriptionally Regulated by GAS5. (a) Several reports have demonstrated the involvement of GAS5 in mtor pathway. So we first analyzed whether mtor pathway would have effects on GAS5 related pluripotency control using Rapamycin, an mtor inhibitor. The qpcr analysis of the expression of pluripotency genes in Rapamycin treated hescs at different time points. *P<0.05, **P<0.01, t-test, n=3. (b) Other report showed that GAS5 can directly bind Glucocorticoid Receptor (GR) to inhibit its activity. So we use GR sirna to analyze the effect of GR on pluripotency in hescs. The efficacy of GR sirna-mediated knockdown (right panel), and the effect of sigr on the mrna level of pluripotency
9 genes and GAS5 transcript (left panel). **P<0.01, t-test, n=3. (c) The expression change of pluripotency genes and NODAL under different doses of Dexamethasone treatment. DEX represents Dexamethasone. T-test, n=3. (d) Luciferase activities of NODAL promoter in HEK-293T cells with different treatment indicated. *P<0.05, t-test, n=3. (e) Since report showed that enhancer in the first intron of Nodal gene may be more important to the expression of Nodal in mouse EpiSCs (resembles hescs), we analyzed the coordinate ASE in human Nodal gene as indicated in the upper panel. The luciferase reporter assay showed that GAS5 modulation did not significantly influence the reporter activity, which indicates that GAS5 did not modulate NODAL expression through increasing its transcription, t-test, n=4. (f) Sequence alignment of GAS5 and NODAL transcript. Blast results from NCBI showed that only one region is complimentary between two transcripts and the length of which is 12bp long. Indicating there is low possibility for GAS5 and NODAL transcript to form stable RNA-RNA complimentary duplex to increase RNA stability. (g) GO analysis of proteins identified in MS2-GAS5 pull down products using label-free proteomics. Error bars represent standard deviation of the indicated experiment replicates. RNA level of β-actin served as internal reference for qpcr.
10 Supplementary Figure 6, related to Figure 6. Mirnome analysis revealed that GAS5 competes mir-2467, mir-3200, and Let7e with NODAL mrna to sustain NODAL expression in hescs (a) The efficacy of Dicer knockdown by lentiviral shrna. The results showed that Dicer along other mirnas we tested reduced significantly upon Dicer knockdown. **P<0.01, t-test, n=3. (b-c) The effect of different doses of microrna mimics (b) or inhibitors (c) to the mrna level of NODAL and GAS5. The concentration here shows the amount of microrna inhibitors or mimics used for each 12-well transfection. 3200In stands for microrna-3200 inhibitors, and 3200, 2467 and Let7e stands for relative microrna mimics. An Inhibitor that binds to none of these micrornas and a microrna mimic that differs greatly from these microrna were used as respective negative control (NC). *P<0.05, **P<0.01, t-test, n=3.. (d) The
11 effect of overexpressing different doses of precursor micrornas to the mrna level of NODAL and GAS5 in hescs were detected using qpcr. Here, a scramble hairpin precursor RNA serves as negative control named as pre-nc. All other pre-mature microrna were synthesized oligos according to mirbase (v21). **P<0.01, t-test, n=3. (e) Northern Blot analysis (n=2) of mir p, mir p and let7e-5p in hescs using anti-sense probes. The Pre-miRNA lane represents total mirnas that extracted from pre-mirna overexpressed hescs. Each lane is loaded with 1ug of total mirna. (f) The effect of overexpressing GAS5 or GAS5-mut on the expression of mirnas. GAS5-mut represents the mirna binding sites mutated GAS5 transcript (Fig. 6e). (g-h) The degradation assay (n=3) analyzing the turnover of the indicated mirnas upon GAS5 wild type transcript (g) or GAS5-mut (h) transcript transfection. Detection of U6 small RNAs at different time point were served as negative control (NC). (i) The degradation assay analyzing the turnover of the indicated mirnas upon GAS5 transfection (n=3). A microrna mimic that differs greatly from these micrornas was used as negative control (NC). Error bars represent standard deviation of the indicated experiment replicates. RNA level of β-actin served as internal reference for qpcr.
12 Supplementary Figure 7, related to Figure 7. MiR-2467, mir-3200, and Let7e inhibit self-renewal and attenuate GAS5-mediated NODAL signaling in hescs (a) The expression change of pluripotency genes under relative mirna inhibition were measured using qpcr. Suffix -Inh represents relative microrna inhibitors, NC represents scramble inhibitor controls. *P<0.05, **P<0.01, t-test, n=3. (b) Relative NODAL and NANOG levels in different treatment group. **P<0.01, t-test, n=3. (c) The luciferase activities of pluripotency gene 3 UTR reporters under relative mirna transfection (left panle) or inhibition (right panel). Suffix -Inh represents relative microrna inhibitors, groups with no suffix -Inh represent microrna mimic overexpression. NS, non-significant, t-test, n=4. Error bars represent standard deviation of the indicated experiment replicates. RNA level of β-actin served as internal reference for qpcr.
13 Supplementary Figure 8, related to Figure 8. GAS5 inhibited cell growth in other cell types than hescs (a) The effect of GAS5 overpression or knockdown on cell growth assessed by CCK-8 analysis in HEK-293T, Fibroblasts, umsc, DU145 and SH-SY5Y cell lines. s.d. of three independent experiments with three replicate each time are indicated. *P<0.05, **P<0.01, ANOVA was used to measure the significance of each groups.
14 Supplementary Figure 9. The uncropped images of blots shown in the main figures (Red boxes indicated the area shown in the main figures).
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected
 Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
Supplementary Figure 1. (a) The qrt-pcr for lnc-2, lnc-6 and lnc-7 RNA level in DU145, 22Rv1, wild type HCT116 and HCT116 Dicer ex5 cells transfected with the sirna against lnc-2, lnc-6, lnc-7, and the
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 Schematic and results of screening the combinatorial antibody library for Sox2 replacement activity. A single batch of MEFs were plated and transduced with doxycycline inducible
Supplementary Figure 1 Schematic and results of screening the combinatorial antibody library for Sox2 replacement activity. A single batch of MEFs were plated and transduced with doxycycline inducible
Regulation of transcription by the MLL2 complex and MLL complex-associated AKAP95
 Supplementary Information Regulation of transcription by the complex and MLL complex-associated Hao Jiang, Xiangdong Lu, Miho Shimada, Yali Dou, Zhanyun Tang, and Robert G. Roeder Input HeLa NE IP lot:
Supplementary Information Regulation of transcription by the complex and MLL complex-associated Hao Jiang, Xiangdong Lu, Miho Shimada, Yali Dou, Zhanyun Tang, and Robert G. Roeder Input HeLa NE IP lot:
Supplementary Figure 1.
 Supplementary Figure 1. Quantification of western blot analysis of fibroblasts (related to Figure 1) (A-F) Quantification of western blot analysis for control and IR-Mut fibroblasts. Data are expressed
Supplementary Figure 1. Quantification of western blot analysis of fibroblasts (related to Figure 1) (A-F) Quantification of western blot analysis for control and IR-Mut fibroblasts. Data are expressed
Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan).
 1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
Supplementary Figure 1. ips_3y+mir-302b. ips_3y+mir-372. ips_3y+mir-302b + mir-372. ips_4y. ips_4y+mir-302b. ips_4y + mir-372
 Supplementary Figure 1 ips_3y+mir-32b ips_3y+ ips_3y+mir-32b + ips_4y ips_4y+mir-32b ips_4y + Nature Biotechnology: doi:1.138/nb.t.1862 TRA-1-6 DAPI Oct3/4 Supplementary Figure 1: ips cells derived from
Supplementary Figure 1 ips_3y+mir-32b ips_3y+ ips_3y+mir-32b + ips_4y ips_4y+mir-32b ips_4y + Nature Biotechnology: doi:1.138/nb.t.1862 TRA-1-6 DAPI Oct3/4 Supplementary Figure 1: ips cells derived from
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector.
 Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector. (a) Western blotting analysis and (b) qpcr analysis of eif6 expression in HEK293 T cells transfected with either
Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector. (a) Western blotting analysis and (b) qpcr analysis of eif6 expression in HEK293 T cells transfected with either
Li et al., Supplemental Figures
 Li et al., Supplemental Figures Fig. S1. Suppressing TGM2 expression with TGM2 sirnas inhibits migration and invasion in A549-TR cells. A, A549-TR cells transfected with negative control sirna (NC sirna)
Li et al., Supplemental Figures Fig. S1. Suppressing TGM2 expression with TGM2 sirnas inhibits migration and invasion in A549-TR cells. A, A549-TR cells transfected with negative control sirna (NC sirna)
Supplementary Figure 1 Validate the expression of mir-302b after bacterial infection by northern
 Supplementary Figure 1 Validate the expression of mir-302b after bacterial infection by northern blot. Northern blot analysis of mir-302b expression following infection with PAO1, PAK and Kp in (A) lung
Supplementary Figure 1 Validate the expression of mir-302b after bacterial infection by northern blot. Northern blot analysis of mir-302b expression following infection with PAO1, PAK and Kp in (A) lung
1. Primers for PCR to amplify hairpin stem-loop precursor mir-145 plus different flanking sequence from human genomic DNA.
 Supplemental data: 1. Primers for PCR to amplify hairpin stem-loop precursor mir-145 plus different flanking sequence from human genomic DNA. Strategy#1: 20nt at both sides: #1_BglII-Fd primer : 5 -gga
Supplemental data: 1. Primers for PCR to amplify hairpin stem-loop precursor mir-145 plus different flanking sequence from human genomic DNA. Strategy#1: 20nt at both sides: #1_BglII-Fd primer : 5 -gga
Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate
 Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
OmicsLink shrna Clones guaranteed knockdown even in difficult-to-transfect cells
 OmicsLink shrna Clones guaranteed knockdown even in difficult-to-transfect cells OmicsLink shrna clone collections consist of lentiviral, and other mammalian expression vector based small hairpin RNA (shrna)
OmicsLink shrna Clones guaranteed knockdown even in difficult-to-transfect cells OmicsLink shrna clone collections consist of lentiviral, and other mammalian expression vector based small hairpin RNA (shrna)
Transcriptional regulation of BRCA1 expression by a metabolic switch: Di, Fernandez, De Siervi, Longo, and Gardner. H3K4Me3
 ChIP H3K4Me3 enrichment.25.2.15.1.5 H3K4Me3 H3K4Me3 ctrl H3K4Me3 + E2 NS + E2 1. kb kb +82 kb Figure S1. Estrogen promotes entry of MCF-7 into the cell cycle but does not significantly change activation-associated
ChIP H3K4Me3 enrichment.25.2.15.1.5 H3K4Me3 H3K4Me3 ctrl H3K4Me3 + E2 NS + E2 1. kb kb +82 kb Figure S1. Estrogen promotes entry of MCF-7 into the cell cycle but does not significantly change activation-associated
TE5 KYSE510 TE7 KYSE70 KYSE140
 TE5 KYSE5 TT KYSE7 KYSE4 Supplementary Figure. Hockey stick plots showing input normalized, rank ordered H3K7ac signals for the candidate SE-associated lncrnas in this study. Rpm Rpm Rpm Chip-seq H3K7ac
TE5 KYSE5 TT KYSE7 KYSE4 Supplementary Figure. Hockey stick plots showing input normalized, rank ordered H3K7ac signals for the candidate SE-associated lncrnas in this study. Rpm Rpm Rpm Chip-seq H3K7ac
supplementary information
 DOI: 10.1038/ncb2116 Figure S1 CDK phosphorylation of EZH2 in cells. (a) Comparison of candidate CDK phosphorylation sites on EZH2 with known CDK substrates by multiple sequence alignments. (b) CDK1 and
DOI: 10.1038/ncb2116 Figure S1 CDK phosphorylation of EZH2 in cells. (a) Comparison of candidate CDK phosphorylation sites on EZH2 with known CDK substrates by multiple sequence alignments. (b) CDK1 and
14_integrins_EGFR
 α1 Integrin α1 -/- fibroblasts from integrin α1 knockout animals Figure S1. Serum-starved fibroblasts from α -/- 1 and +/+ mice were stimulated with 10% FBS for 30 min. (FBS) or plated on collagen I (CI)
α1 Integrin α1 -/- fibroblasts from integrin α1 knockout animals Figure S1. Serum-starved fibroblasts from α -/- 1 and +/+ mice were stimulated with 10% FBS for 30 min. (FBS) or plated on collagen I (CI)
Thermo Scientific Dharmacon SMARTvector 2.0 Lentiviral shrna Particles
 Thermo Scientific Dharmacon SMARTvector 2.0 Lentiviral shrna Particles Long-term gene silencing shrna-specific design algorithm High titer, purified particles Thermo Scientific Dharmacon SMARTvector shrna
Thermo Scientific Dharmacon SMARTvector 2.0 Lentiviral shrna Particles Long-term gene silencing shrna-specific design algorithm High titer, purified particles Thermo Scientific Dharmacon SMARTvector shrna
Revision Checklist for Science Signaling Research Manuscripts: Data Requirements and Style Guidelines
 Revision Checklist for Science Signaling Research Manuscripts: Data Requirements and Style Guidelines Further information can be found at: http://stke.sciencemag.org/sites/default/files/researcharticlerevmsinstructions_0.pdf.
Revision Checklist for Science Signaling Research Manuscripts: Data Requirements and Style Guidelines Further information can be found at: http://stke.sciencemag.org/sites/default/files/researcharticlerevmsinstructions_0.pdf.
Investigating the regulation of mirna biogenesis and Argonaute2 by RNA binding proteins
 Investigating the regulation of mirna biogenesis and Argonaute2 by RNA binding proteins Patrick Peter Connerty Supervisor: Gyorgy Hutvagner Thesis Submitted for the Degree of Doctor of Philosophy (Science)
Investigating the regulation of mirna biogenesis and Argonaute2 by RNA binding proteins Patrick Peter Connerty Supervisor: Gyorgy Hutvagner Thesis Submitted for the Degree of Doctor of Philosophy (Science)
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna
 Supplemental Materials and Methods Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna (Mission shrna, Sigma) against mouse MMP14 were transfected into HEK293 cells using FuGene6
Supplemental Materials and Methods Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna (Mission shrna, Sigma) against mouse MMP14 were transfected into HEK293 cells using FuGene6
Supplemental Fig. 1: PEA-15 knockdown efficiency assessed by immunohistochemistry and qpcr
 Supplemental figure legends Supplemental Fig. 1: PEA-15 knockdown efficiency assessed by immunohistochemistry and qpcr A, LβT2 cells were transfected with either scrambled or PEA-15 sirna. Cells were then
Supplemental figure legends Supplemental Fig. 1: PEA-15 knockdown efficiency assessed by immunohistochemistry and qpcr A, LβT2 cells were transfected with either scrambled or PEA-15 sirna. Cells were then
Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells.
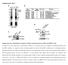 Supplementary Fig. 1 Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells. (a) FRTL-5 cells were treated with 1 mm dibutyryl camp for 24 h, and the lysates
Supplementary Fig. 1 Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells. (a) FRTL-5 cells were treated with 1 mm dibutyryl camp for 24 h, and the lysates
Supplementary Fig. 1. (A) Working model. The pluripotency transcription factor OCT4
 SUPPLEMENTARY FIGURE LEGENDS Supplementary Fig. 1. (A) Working model. The pluripotency transcription factor OCT4 directly up-regulates the expression of NIPP1 and CCNF that together inhibit protein phosphatase
SUPPLEMENTARY FIGURE LEGENDS Supplementary Fig. 1. (A) Working model. The pluripotency transcription factor OCT4 directly up-regulates the expression of NIPP1 and CCNF that together inhibit protein phosphatase
PI3K/mTORC2 regulates TGFβ/Activin signalling by modulating Smad2/3
 Supplementary Information PI3K/mTORC2 regulates TGFβ/Activin signalling by modulating Smad2/3 activity via linker phosphorylation Jason S.L. Yu, 1 Thamil Selvee Ramasamy, 1,2 Nick Murphy, 1 Marie Holt,
Supplementary Information PI3K/mTORC2 regulates TGFβ/Activin signalling by modulating Smad2/3 activity via linker phosphorylation Jason S.L. Yu, 1 Thamil Selvee Ramasamy, 1,2 Nick Murphy, 1 Marie Holt,
Thyroid peroxidase gene expression is induced by lipopolysaccharide involving Nuclear Factor (NF)-κB p65 subunit phosphorylation
 1 2 3 4 5 SUPPLEMENTAL DATA Thyroid peroxidase gene expression is induced by lipopolysaccharide involving Nuclear Factor (NF)-κB p65 subunit phosphorylation Magalí Nazar, Juan Pablo Nicola, María Laura
1 2 3 4 5 SUPPLEMENTAL DATA Thyroid peroxidase gene expression is induced by lipopolysaccharide involving Nuclear Factor (NF)-κB p65 subunit phosphorylation Magalí Nazar, Juan Pablo Nicola, María Laura
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Transcriptional Regulation (Gene Regulation)
 Experimental Techniques in Biomedical Sciences 의생명과학실험기법 Transcriptional Regulation (Gene Regulation) 4/17/13 Jeong Hoon Kim (jeongkim@skku.edu) Department of Health Sciences and Technology, SKKU Graduate
Experimental Techniques in Biomedical Sciences 의생명과학실험기법 Transcriptional Regulation (Gene Regulation) 4/17/13 Jeong Hoon Kim (jeongkim@skku.edu) Department of Health Sciences and Technology, SKKU Graduate
Therapeutic & Prevention Application of Nucleic Acids
 Therapeutic & Prevention Application of Nucleic Acids Seyed Amir Hossein Jalali Institute of Biotechnology and Bioengineering, Isfahan University Of Technology (IUT). 30.7.2015 * Plasmids * DNA Aptamers
Therapeutic & Prevention Application of Nucleic Acids Seyed Amir Hossein Jalali Institute of Biotechnology and Bioengineering, Isfahan University Of Technology (IUT). 30.7.2015 * Plasmids * DNA Aptamers
Site-Directed Mutagenesis. Mutations in four Smad4 sites of mouse Gat1 promoter
 Supplement Supporting Materials and Methods Site-Directed Mutagenesis. Mutations in four Smad4 sites of mouse Gat1 promoter were independently generated using a two-step PCR method. The Smad4 binding site
Supplement Supporting Materials and Methods Site-Directed Mutagenesis. Mutations in four Smad4 sites of mouse Gat1 promoter were independently generated using a two-step PCR method. The Smad4 binding site
Dual PI3K/ERK inhibition induces necroptotic cell death of Hodgkin Lymphoma cells through IER3 downregulation
 Dual PI3K/ERK inhibition induces necroptotic cell death of Hodgkin Lymphoma cells through IER3 downregulation *Silvia Laura Locatelli, 1 Giuseppa Careddu, 1 Giuliano Giuseppe Stirparo, 1 Luca Castagna,
Dual PI3K/ERK inhibition induces necroptotic cell death of Hodgkin Lymphoma cells through IER3 downregulation *Silvia Laura Locatelli, 1 Giuseppa Careddu, 1 Giuliano Giuseppe Stirparo, 1 Luca Castagna,
Table 1. Primers, annealing temperatures, and product sizes for PCR amplification.
 Table 1. Primers, annealing temperatures, and product sizes for PCR amplification. Gene Direction Primer sequence (5 3 ) Annealing Temperature Size (bp) BRCA1 Forward TTGCGGGAGGAAAATGGGTAGTTA 50 o C 292
Table 1. Primers, annealing temperatures, and product sizes for PCR amplification. Gene Direction Primer sequence (5 3 ) Annealing Temperature Size (bp) BRCA1 Forward TTGCGGGAGGAAAATGGGTAGTTA 50 o C 292
Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat
 Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat units in the human genome. Annotated transposable elements
Supplementary Figure 1. NORAD expression in mouse (A) and dog (B). The black boxes indicate the position of the regions alignable to the 12 repeat units in the human genome. Annotated transposable elements
Nanog-Luc. R-Luc. 40 HA-Klf Relative protein level of Klf USP21. Vector USP2 USP21 *** *** *** ***
 :Flag Relative protein level of Relative protein level of : Flag Vec 21LV 21SV 2 : HA HA- HA- HA- HA-Klf4 Relative mrna expression Relative protein level of Relative protein level of Relative protein level
:Flag Relative protein level of Relative protein level of : Flag Vec 21LV 21SV 2 : HA HA- HA- HA- HA-Klf4 Relative mrna expression Relative protein level of Relative protein level of Relative protein level
Description of Supplementary Files. File name: Supplementary Information Description: Supplementary figures and supplementary tables.
 Description of Supplementary Files File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Supplementary Data 1 Description: Differential expression
Description of Supplementary Files File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Supplementary Data 1 Description: Differential expression
TOOLS sirna and mirna. User guide
 TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
Figure S2. Response of mouse ES cells to GSK3 inhibition. Mentioned in discussion
 Stem Cell Reports, Volume 1 Supplemental Information Robust Self-Renewal of Rat Embryonic Stem Cells Requires Fine-Tuning of Glycogen Synthase Kinase-3 Inhibition Yaoyao Chen, Kathryn Blair, and Austin
Stem Cell Reports, Volume 1 Supplemental Information Robust Self-Renewal of Rat Embryonic Stem Cells Requires Fine-Tuning of Glycogen Synthase Kinase-3 Inhibition Yaoyao Chen, Kathryn Blair, and Austin
a KYSE270-CON KYSE270-Id1
 a KYSE27-CON KYSE27- shcon shcon sh b Human Mouse CD31 Relative MVD 3.5 3 2.5 2 1.5 1.5 *** *** c KYSE15 KYSE27 sirna (nm) 5 1 Id2 Id2 sirna 5 1 sirna (nm) 5 1 Id2 sirna 5 1 Id2 [h] (pg per ml) d 3 2 1
a KYSE27-CON KYSE27- shcon shcon sh b Human Mouse CD31 Relative MVD 3.5 3 2.5 2 1.5 1.5 *** *** c KYSE15 KYSE27 sirna (nm) 5 1 Id2 Id2 sirna 5 1 sirna (nm) 5 1 Id2 sirna 5 1 Id2 [h] (pg per ml) d 3 2 1
Premade Lentiviral Particles for ips Stem Factors: Single vector system. For generation of induced pluripotent stem (ips) cells and other applications
 Premade Lentiviral Particles for ips Stem Factors: Single vector system For generation of induced pluripotent stem (ips) cells and other applications RESEARCH USE ONLY, not for diagnostics or therapeutics
Premade Lentiviral Particles for ips Stem Factors: Single vector system For generation of induced pluripotent stem (ips) cells and other applications RESEARCH USE ONLY, not for diagnostics or therapeutics
SUPPLEMENTARY INFORMATION
 SUPPLEMENTRY INFORMTION DOI:.38/ncb Kdmb locus kb Long isoform Short isoform Long isoform Jmj XX PHD F-box LRR Short isoform XX PHD F-box LRR Target Vector 3xFlag Left H Neo Right H TK LoxP LoxP Kdmb Locus
SUPPLEMENTRY INFORMTION DOI:.38/ncb Kdmb locus kb Long isoform Short isoform Long isoform Jmj XX PHD F-box LRR Short isoform XX PHD F-box LRR Target Vector 3xFlag Left H Neo Right H TK LoxP LoxP Kdmb Locus
Supplementary Figure 1. Soft fibrin gels promote growth and organized mesodermal differentiation. Representative images of single OGTR1 ESCs cultured
 Supplementary Figure 1. Soft fibrin gels promote growth and organized mesodermal differentiation. Representative images of single OGTR1 ESCs cultured in 90-Pa 3D fibrin gels for 5 days in the presence
Supplementary Figure 1. Soft fibrin gels promote growth and organized mesodermal differentiation. Representative images of single OGTR1 ESCs cultured in 90-Pa 3D fibrin gels for 5 days in the presence
Supplementary
 Supplementary information Supplementary Material and Methods Plasmid construction The transposable element vectors for inducible expression of RFP-FUS wt and EGFP-FUS R521C and EGFP-FUS P525L were derived
Supplementary information Supplementary Material and Methods Plasmid construction The transposable element vectors for inducible expression of RFP-FUS wt and EGFP-FUS R521C and EGFP-FUS P525L were derived
SUPPLEMENTARY INFORMATION FILE
 SUPPLEMENTARY INFORMATION FILE Existence of a microrna pathway in anucleate platelets Patricia Landry, Isabelle Plante, Dominique L. Ouellet, Marjorie P. Perron, Guy Rousseau & Patrick Provost 1. SUPPLEMENTARY
SUPPLEMENTARY INFORMATION FILE Existence of a microrna pathway in anucleate platelets Patricia Landry, Isabelle Plante, Dominique L. Ouellet, Marjorie P. Perron, Guy Rousseau & Patrick Provost 1. SUPPLEMENTARY
SUPPLEMENTARY INFORMATION
 doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
SUPPLEMENTARY INFORMATION Dynamic Phosphorylation of HP1 Regulates Mitotic Progression in Human Cells Supplementary Figures Supplementary Figure 1. NDR1 interacts with HP1. (a) Immunoprecipitation using
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1154040/dc1 Supporting Online Material for Selective Blockade of MicroRNA Processing by Lin-28 Srinivas R. Viswanathan, George Q. Daley,* Richard I. Gregory* *To whom
www.sciencemag.org/cgi/content/full/1154040/dc1 Supporting Online Material for Selective Blockade of MicroRNA Processing by Lin-28 Srinivas R. Viswanathan, George Q. Daley,* Richard I. Gregory* *To whom
SUPPLEMENTARY INFORMATION
 (Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
(Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
Supplementary Table, Figures and Videos
 Supplementary Table, Figures and Videos Table S1. Oligonucleotides used for different approaches. (A) RT-qPCR study. (B) qpcr study after ChIP assay. (C) Probes used for EMSA. Figure S1. Notch activation
Supplementary Table, Figures and Videos Table S1. Oligonucleotides used for different approaches. (A) RT-qPCR study. (B) qpcr study after ChIP assay. (C) Probes used for EMSA. Figure S1. Notch activation
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11070 Supplementary Figure 1 Purification of FLAG-tagged proteins. a, Purification of FLAG-RNF12 by FLAG-affinity from nuclear extracts of wild-type (WT) and two FLAG- RNF12 transgenic
doi:10.1038/nature11070 Supplementary Figure 1 Purification of FLAG-tagged proteins. a, Purification of FLAG-RNF12 by FLAG-affinity from nuclear extracts of wild-type (WT) and two FLAG- RNF12 transgenic
Premade Lentiviral Particles for human ips Stem Factors
 Premade Lentiviral Particles for human ips Stem Factors For generating induced pluripotent stem (ips) cells or other applications RESEARCH USE ONLY, not for diagnostics or therapeutics Cat# Product Name
Premade Lentiviral Particles for human ips Stem Factors For generating induced pluripotent stem (ips) cells or other applications RESEARCH USE ONLY, not for diagnostics or therapeutics Cat# Product Name
microrna Shifra Ben-Dor March 2010
 microrna Shifra Ben-Dor March 2010 Outline Biology of mirna Prediction of mirna mirna Databases Prediction of mirna Targets micrornas (mirna) Naturally expressed small RNAs Involved in regulation of target
microrna Shifra Ben-Dor March 2010 Outline Biology of mirna Prediction of mirna mirna Databases Prediction of mirna Targets micrornas (mirna) Naturally expressed small RNAs Involved in regulation of target
HPV E6 oncoprotein targets histone methyltransferases for modulating specific. Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu,
 1 HPV E oncoprotein targets histone methyltransferases for modulating specific gene transcription 3 5 Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu, Cheng-Ming Chiang, Sheng-Chung
1 HPV E oncoprotein targets histone methyltransferases for modulating specific gene transcription 3 5 Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu, Cheng-Ming Chiang, Sheng-Chung
Fig. S1 TGF RI inhibitor SB effectively blocks phosphorylation of Smad2 induced by TGF. FET cells were treated with TGF in the presence of
 Fig. S1 TGF RI inhibitor SB525334 effectively blocks phosphorylation of Smad2 induced by TGF. FET cells were treated with TGF in the presence of different concentrations of SB525334. Cells were lysed and
Fig. S1 TGF RI inhibitor SB525334 effectively blocks phosphorylation of Smad2 induced by TGF. FET cells were treated with TGF in the presence of different concentrations of SB525334. Cells were lysed and
GE Healthcare. DharmaconTM. Gene Editing, RNAi & Gene Expression. RNAi Application Guide 2016/17
 GE Healthcare DharmaconTM Gene Editing, RNAi & Gene Expression RNAi Application Guide 2016/17 Every discovery is a stepping stone toward clarity, firm conclusions, and ultimately an amazing breakthrough.
GE Healthcare DharmaconTM Gene Editing, RNAi & Gene Expression RNAi Application Guide 2016/17 Every discovery is a stepping stone toward clarity, firm conclusions, and ultimately an amazing breakthrough.
Supplementary information
 Supplementary information Supplementary Figure 1 (a) EM image of the pyramidal layer of wt mice. CA3 pyramidal neurons were selected according to their typical alignment, size and shape for subsequent
Supplementary information Supplementary Figure 1 (a) EM image of the pyramidal layer of wt mice. CA3 pyramidal neurons were selected according to their typical alignment, size and shape for subsequent
Marilyn G. Rimando, Hao-Hsiang Wu, Yu-An Liu, Chien-Wei Lee, Shu-Wen Kuo, Yin-
 Supplementary Information Glucocorticoid receptor and Histone Deacetylase 6 mediate the differential effect of dexamethasone during osteogenesis of Mesenchymal stromal cells (MSCs) Marilyn G. Rimando,
Supplementary Information Glucocorticoid receptor and Histone Deacetylase 6 mediate the differential effect of dexamethasone during osteogenesis of Mesenchymal stromal cells (MSCs) Marilyn G. Rimando,
SUPPLEMENTARY INFORMATION
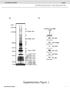 doi: 10.1038/nature05841 SUPPLEMENTARY INFORMATION www.nature.com/nature 1 www.nature.com/nature 2 www.nature.com/nature 3 www.nature.com/nature 4 a Hours of Development: eif-6(rnai): 4 wildtype 30 4 lin-4
doi: 10.1038/nature05841 SUPPLEMENTARY INFORMATION www.nature.com/nature 1 www.nature.com/nature 2 www.nature.com/nature 3 www.nature.com/nature 4 a Hours of Development: eif-6(rnai): 4 wildtype 30 4 lin-4
b alternative classical none
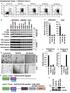 Supplementary Figure. 1: Related to Figure.1 a d e b alternative classical none NIK P-IkBa Total IkBa Tubulin P52 (Lighter) P52 (Darker) RelB (Lighter) RelB (Darker) HDAC1 Control-Sh RelB-Sh NF-kB2-Sh
Supplementary Figure. 1: Related to Figure.1 a d e b alternative classical none NIK P-IkBa Total IkBa Tubulin P52 (Lighter) P52 (Darker) RelB (Lighter) RelB (Darker) HDAC1 Control-Sh RelB-Sh NF-kB2-Sh
Embryonic germ cell formation. Maintainance of pluripotency
 hvasa 2.5 kb promoter egfp 3 UTR(1kb) 1. Stable integration of germ cell reporter into hescs 2. Adherent differentiation and FACS isolation for VASA:GFP+ cells 4. Silencing or overexpression of the human
hvasa 2.5 kb promoter egfp 3 UTR(1kb) 1. Stable integration of germ cell reporter into hescs 2. Adherent differentiation and FACS isolation for VASA:GFP+ cells 4. Silencing or overexpression of the human
This is the author's accepted version of the manuscript.
 This is the author's accepted version of the manuscript. The definitive version is published in Nature Communications Online Edition: 2015/4/16 (Japan time), doi:10.1038/ncomms7780. The final version published
This is the author's accepted version of the manuscript. The definitive version is published in Nature Communications Online Edition: 2015/4/16 (Japan time), doi:10.1038/ncomms7780. The final version published
Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and
 Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
Supplementary Figure 1 A
 Supplementary Figure A B M. mullata p53, 3 UTR Luciferase activity (%) mir-5b 8 Le et al. Supplementary Information NC-DP - + - - - - - NC-DP - - + - - - - NC-DP3 - - - + - - - 5b-DP - - - - + + + NC-AS
Supplementary Figure A B M. mullata p53, 3 UTR Luciferase activity (%) mir-5b 8 Le et al. Supplementary Information NC-DP - + - - - - - NC-DP - - + - - - - NC-DP3 - - - + - - - 5b-DP - - - - + + + NC-AS
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Kawai and Amano, http://www.jcb.org/cgi/content/full/jcb.201110008/dc1 Figure S1. Regulation of mirna expression by BRCA1. (A) Confirmation
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Kawai and Amano, http://www.jcb.org/cgi/content/full/jcb.201110008/dc1 Figure S1. Regulation of mirna expression by BRCA1. (A) Confirmation
Supplementary Figure 1 PARP1 is involved in regulating the stability of mrnas from pro-inflammatory cytokine/chemokine mediators.
 Supplementary Figure 1 PARP1 is involved in regulating the stability of mrnas from pro-inflammatory cytokine/chemokine mediators. (a) A graphic depiction of the approach to determining the stability of
Supplementary Figure 1 PARP1 is involved in regulating the stability of mrnas from pro-inflammatory cytokine/chemokine mediators. (a) A graphic depiction of the approach to determining the stability of
Supplementary Fig. 1
 a FL (1-2266) NL (1-1190) CL (1191-2266) HA-ICE1: - HA-ICE1: - - - FLAG-ICE2: + + + + FLAG-ELL: + + + + + + IP: anti-ha FLAG-ICE2 HA-ICE1-FL HA-ICE1-NL HA-ICE1-CL FLAG-ICE2 b IP: anti-ha FL (1-2266) NL
a FL (1-2266) NL (1-1190) CL (1191-2266) HA-ICE1: - HA-ICE1: - - - FLAG-ICE2: + + + + FLAG-ELL: + + + + + + IP: anti-ha FLAG-ICE2 HA-ICE1-FL HA-ICE1-NL HA-ICE1-CL FLAG-ICE2 b IP: anti-ha FL (1-2266) NL
Supplementary Figure 1. IFN-γ induces TRC dormancy. a, IFN-γ induced dormancy
 Supplementary Figure 1. IFN-γ induces TRC dormancy. a, IFN-γ induced dormancy of various tumor type TRCs, including H22 (murine hepatocarcinoma) and CT26 (murine colon cancer). Bar, 50 µm. b, B16 cells
Supplementary Figure 1. IFN-γ induces TRC dormancy. a, IFN-γ induced dormancy of various tumor type TRCs, including H22 (murine hepatocarcinoma) and CT26 (murine colon cancer). Bar, 50 µm. b, B16 cells
Effects of metformin on Sirt1, Nrf2 and AhR expression in cancer cells
 A Dissertation for the Degree of Doctor of Philosophy Effects of metformin on Sirt1, Nrf2 and AhR expression in cancer cells Department of Pharmacy Graduate School Chungnam National University By Minh
A Dissertation for the Degree of Doctor of Philosophy Effects of metformin on Sirt1, Nrf2 and AhR expression in cancer cells Department of Pharmacy Graduate School Chungnam National University By Minh
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
supplementary information
 Figure S1 ZEB1 full length mrna. (a) Analysis of the ZEB1 mrna using the UCSC genome browser (http://genome.ucsc.edu) revealed truncation of the annotated Refseq sequence (NM_030751). The probable terminus
Figure S1 ZEB1 full length mrna. (a) Analysis of the ZEB1 mrna using the UCSC genome browser (http://genome.ucsc.edu) revealed truncation of the annotated Refseq sequence (NM_030751). The probable terminus
SUPPLEMENTARY INFORMATION
 ARTICLE NUMBER: 0 DOI: 0.0/NMICROBIOL.0. The host protein CLUH participates in the subnuclear transport of influenza virus ribonucleoprotein complexes Tomomi Ando,, Seiya Yamayoshi, Yuriko Tomita,, Shinji
ARTICLE NUMBER: 0 DOI: 0.0/NMICROBIOL.0. The host protein CLUH participates in the subnuclear transport of influenza virus ribonucleoprotein complexes Tomomi Ando,, Seiya Yamayoshi, Yuriko Tomita,, Shinji
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/259/ra5/dc1 Supplementary Materials for Systems Biology Approach Identifies the Kinase Csnk1a1 as a Regulator of the DNA Damage Response in Embryonic Stem Cells
www.sciencesignaling.org/cgi/content/full/6/259/ra5/dc1 Supplementary Materials for Systems Biology Approach Identifies the Kinase Csnk1a1 as a Regulator of the DNA Damage Response in Embryonic Stem Cells
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3363 Supplementary Figure 1 Several WNTs bind to the extracellular domains of PKD1. (a) HEK293T cells were co-transfected with indicated plasmids. Flag-tagged proteins were immunoprecipiated
DOI: 10.1038/ncb3363 Supplementary Figure 1 Several WNTs bind to the extracellular domains of PKD1. (a) HEK293T cells were co-transfected with indicated plasmids. Flag-tagged proteins were immunoprecipiated
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/496/eaam6291/dc1 Supplementary Materials for Regulation of autophagy, NF-κB signaling, and cell viability by mir-124 in KRAS mutant mesenchymal-like NSCLC cells
www.sciencesignaling.org/cgi/content/full/10/496/eaam6291/dc1 Supplementary Materials for Regulation of autophagy, NF-κB signaling, and cell viability by mir-124 in KRAS mutant mesenchymal-like NSCLC cells
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature09732 Supplementary Figure 1: Depletion of Fbw7 results in elevated Mcl-1 abundance. a, Total thymocytes from 8-wk-old Lck-Cre/Fbw7 +/fl (Control) or Lck-Cre/Fbw7 fl/fl (Fbw7 KO) mice
doi:10.1038/nature09732 Supplementary Figure 1: Depletion of Fbw7 results in elevated Mcl-1 abundance. a, Total thymocytes from 8-wk-old Lck-Cre/Fbw7 +/fl (Control) or Lck-Cre/Fbw7 fl/fl (Fbw7 KO) mice
Design. Construction. Characterization
 Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
A Survey of Genetic Methods
 IBS 8102 Cell, Molecular, and Developmental Biology A Survey of Genetic Methods January 24, 2008 DNA RNA Hybridization ** * radioactive probe reverse transcriptase polymerase chain reaction RT PCR DNA
IBS 8102 Cell, Molecular, and Developmental Biology A Survey of Genetic Methods January 24, 2008 DNA RNA Hybridization ** * radioactive probe reverse transcriptase polymerase chain reaction RT PCR DNA
Data Sheet. SBE Reporter Kit (TGFβ/SMAD signaling pathway) Catalog #: 60654
 Data Sheet SBE Reporter Kit (TGFβ/SMAD signaling pathway) Catalog #: 60654 Background The transforming growth factor beta (TGFβ) signaling pathway is involved in a diverse range of cell processes such
Data Sheet SBE Reporter Kit (TGFβ/SMAD signaling pathway) Catalog #: 60654 Background The transforming growth factor beta (TGFβ) signaling pathway is involved in a diverse range of cell processes such
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
Sarker et al. Supplementary Material. Subcellular Fractionation
 Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Estradiol-Estrogen Receptor α Mediates the Expression of the CXXC5 Gene through the Estrogen Response Element-Dependent Signaling Pathway
 Estradiol-Estrogen Receptor α Mediates the Expression of the CXXC5 Gene through the Estrogen Response Element-Dependent Signaling Pathway Pelin Yaşar, Gamze Ayaz and Mesut Muyan SUPPLEMENTARY INFORMATION
Estradiol-Estrogen Receptor α Mediates the Expression of the CXXC5 Gene through the Estrogen Response Element-Dependent Signaling Pathway Pelin Yaşar, Gamze Ayaz and Mesut Muyan SUPPLEMENTARY INFORMATION
Xu et al., Supplementary Figures 1-7
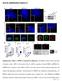 Xu et al., Supplementary Figures 1-7 Supplementary Figure 1. PIPKI is required for ciliogenesis. (a) PIPKI localizes at the basal body of primary cilium. RPE-1 cells treated with two sirnas targeting to
Xu et al., Supplementary Figures 1-7 Supplementary Figure 1. PIPKI is required for ciliogenesis. (a) PIPKI localizes at the basal body of primary cilium. RPE-1 cells treated with two sirnas targeting to
Supplementary Figure 1. TSA (10 nmol/l), non-class-selective HDAC inhibitor, potentiates
 Supplementary Figure 1. TSA (10 nmol/l), non-class-selective HDAC inhibitor, potentiates vascular calcification (VC). (a) Von Kossa staining shows that TSA potentiated the Pi-induced VC. Scale bar, 100
Supplementary Figure 1. TSA (10 nmol/l), non-class-selective HDAC inhibitor, potentiates vascular calcification (VC). (a) Von Kossa staining shows that TSA potentiated the Pi-induced VC. Scale bar, 100
Suppl. Table S1. Characteristics of DHS regions analyzed by bisulfite sequencing. No. CpGs analyzed in the amplicon. Genomic location specificity
 Suppl. Table S1. Characteristics of DHS regions analyzed by bisulfite sequencing. DHS/GRE Genomic location Tissue specificity DHS type CpG density (per 100 bp) No. CpGs analyzed in the amplicon CpG within
Suppl. Table S1. Characteristics of DHS regions analyzed by bisulfite sequencing. DHS/GRE Genomic location Tissue specificity DHS type CpG density (per 100 bp) No. CpGs analyzed in the amplicon CpG within
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 Generation of the AARE-Gene system construct. (a) Position and sequence alignment of AAREs extracted from human Trb3, Chop or Atf3 promoters. AARE core sequences are boxed in grey.
Supplementary Figure 1 Generation of the AARE-Gene system construct. (a) Position and sequence alignment of AAREs extracted from human Trb3, Chop or Atf3 promoters. AARE core sequences are boxed in grey.
SUPPLEMENTARY INFORMATION
 DOI: 1.138/ncb37 a mrna relative expression 1.3 1..8.5.3. CTR NGPS OCT4 SOX2 KLF4 c-myc b BANF1 mrna levels 1.2 1..8.6.4.2. plko.1 shbanf1 c Tra-1-6+ colonies 3 1 plko.1 shbanf1 d BANF1 locus Plasmid donor
DOI: 1.138/ncb37 a mrna relative expression 1.3 1..8.5.3. CTR NGPS OCT4 SOX2 KLF4 c-myc b BANF1 mrna levels 1.2 1..8.6.4.2. plko.1 shbanf1 c Tra-1-6+ colonies 3 1 plko.1 shbanf1 d BANF1 locus Plasmid donor
Chemically defined conditions for human ipsc derivation and culture
 Nature Methods Chemically defined conditions for human ipsc derivation and culture Guokai Chen, Daniel R Gulbranson, Zhonggang Hou, Jennifer M Bolin, Victor Ruotti, Mitchell D Probasco, Kimberly Smuga-Otto,
Nature Methods Chemically defined conditions for human ipsc derivation and culture Guokai Chen, Daniel R Gulbranson, Zhonggang Hou, Jennifer M Bolin, Victor Ruotti, Mitchell D Probasco, Kimberly Smuga-Otto,
(a) Immunoblotting to show the migration position of Flag-tagged MAVS
 Supplementary Figure 1 Characterization of six MAVS isoforms. (a) Immunoblotting to show the migration position of Flag-tagged MAVS isoforms. HEK293T Mavs -/- cells were transfected with constructs expressing
Supplementary Figure 1 Characterization of six MAVS isoforms. (a) Immunoblotting to show the migration position of Flag-tagged MAVS isoforms. HEK293T Mavs -/- cells were transfected with constructs expressing
Khaled_Fig. S MITF TYROSINASE R²= MITF PDE4D R²= Variance from mean mrna expression
 Khaled_Fig. S Variance from mean mrna expression.8 2 MITF.6 TYROSINASE.4 R²=.73.2.8.6.4.2.8.6.4.2.8.6.4.2 MALME3M SKMEL28 UACC257 MITF PDE4D R²=.63 4.5 5 MITF 3.5 4 PDE4B 2.5 3 R²=.2.5 2.5.8.6.4.2.8.6.4.2
Khaled_Fig. S Variance from mean mrna expression.8 2 MITF.6 TYROSINASE.4 R²=.73.2.8.6.4.2.8.6.4.2.8.6.4.2 MALME3M SKMEL28 UACC257 MITF PDE4D R²=.63 4.5 5 MITF 3.5 4 PDE4B 2.5 3 R²=.2.5 2.5.8.6.4.2.8.6.4.2
NTM486-04, NTM174-04,
 Transfection of transformed human trabecular meshwork TM5, and primary human NTM210-05, NTM486-04, NTM174-04, and NTM153-00 cells with Metafectene Easy Adnan Dibas1A,C, Ming Jiang1A,C, Thomas Yorio1A,C.
Transfection of transformed human trabecular meshwork TM5, and primary human NTM210-05, NTM486-04, NTM174-04, and NTM153-00 cells with Metafectene Easy Adnan Dibas1A,C, Ming Jiang1A,C, Thomas Yorio1A,C.
MISSION shrna Library: Next Generation RNA Interference
 Page 1 of 6 Page 1 of 6 Return to Web Version MISSION shrna Library: Next Generation RNA Interference By: Stephanie Uder, Henry George, Betsy Boedeker, LSI Volume 6 Article 2 Introduction The technology
Page 1 of 6 Page 1 of 6 Return to Web Version MISSION shrna Library: Next Generation RNA Interference By: Stephanie Uder, Henry George, Betsy Boedeker, LSI Volume 6 Article 2 Introduction The technology
Lullaby sirna Transfection Reagent - Results
 sirna Transfection Reagent - Results OZ Biosciences is delighted to announce the launching of a new sirna transfection reagent: -sirna. This lipid based transfection reagent is specifically designed for
sirna Transfection Reagent - Results OZ Biosciences is delighted to announce the launching of a new sirna transfection reagent: -sirna. This lipid based transfection reagent is specifically designed for
CD93 and dystroglycan cooperation in human endothelial cell adhesion and migration
 /, Supplementary Advance Publications Materials 2016 CD93 and dystroglycan cooperation in human endothelial cell adhesion and migration Supplementary Materials Supplementary Figure S1: In ECs CD93 silencing
/, Supplementary Advance Publications Materials 2016 CD93 and dystroglycan cooperation in human endothelial cell adhesion and migration Supplementary Materials Supplementary Figure S1: In ECs CD93 silencing
LINGO-1, A TRANSMEMBRANE SIGNALING PROTEIN, INHIBITS OLIGODENDROCYTE DIFFERENTIATION AND MYELINATION THROUGH INTERCELLULAR SELF- INTERACTIONS.
 Supplemental Data: LINGO-1, A TRANSMEMBRANE SIGNALING PROTEIN, INHIBITS OLIGODENDROCYTE DIFFERENTIATION AND MYELINATION THROUGH INTERCELLULAR SELF- INTERACTIONS. Scott Jepson, Bryan Vought, Christian H.
Supplemental Data: LINGO-1, A TRANSMEMBRANE SIGNALING PROTEIN, INHIBITS OLIGODENDROCYTE DIFFERENTIATION AND MYELINATION THROUGH INTERCELLULAR SELF- INTERACTIONS. Scott Jepson, Bryan Vought, Christian H.
Supplementary Information
 Supplementary Information Sam68 modulates the promoter specificity of NF-κB and mediates expression of CD25 in activated T cells Kai Fu 1, 6, Xin Sun 1, 6, Wenxin Zheng 1, 6, Eric M. Wier 1, Andrea Hodgson
Supplementary Information Sam68 modulates the promoter specificity of NF-κB and mediates expression of CD25 in activated T cells Kai Fu 1, 6, Xin Sun 1, 6, Wenxin Zheng 1, 6, Eric M. Wier 1, Andrea Hodgson
Supplementary Figure 1: Overexpression of EBV-encoded proteins Western blot analysis of the expression levels of EBV-encoded latency III proteins in
 Supplementary Figure 1: Overexpression of EBV-encoded proteins Western blot analysis of the expression levels of EBV-encoded latency III proteins in BL2 cells. The Ponceau S staining of the membranes or
Supplementary Figure 1: Overexpression of EBV-encoded proteins Western blot analysis of the expression levels of EBV-encoded latency III proteins in BL2 cells. The Ponceau S staining of the membranes or
Supplementary Information
 Supplementary Information Peroxiredoxin-2 and STAT3 form a redox relay for H 2 O 2 signaling Mirko C. Sobotta 1, Willy Liou 1, Sarah Stöcker 1, Deepti Talwar 1, Michael Oehler 1, Thomas Ruppert 2, Annette
Supplementary Information Peroxiredoxin-2 and STAT3 form a redox relay for H 2 O 2 signaling Mirko C. Sobotta 1, Willy Liou 1, Sarah Stöcker 1, Deepti Talwar 1, Michael Oehler 1, Thomas Ruppert 2, Annette
B. Transgenic plants with strong phenotype (%)
 A. TCTAGTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT AAATATGGTCTAAAGAAGAAGAATATGGTCTAAAGAAGAAGAATATGG 2XP35S STTM165 5 GGGGGATGAAGctaCCTGGTCCGA3 3 CCCCCUACUUC---GGACCAGGCU5 mir165 HindIII mir165 96 nt GTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT
A. TCTAGTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT AAATATGGTCTAAAGAAGAAGAATATGGTCTAAAGAAGAAGAATATGG 2XP35S STTM165 5 GGGGGATGAAGctaCCTGGTCCGA3 3 CCCCCUACUUC---GGACCAGGCU5 mir165 HindIII mir165 96 nt GTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT
MicroRNAs Sequencing, analysis and then what? Click to edit Master subtitle. Pamela Mukhopadhyay Winter School 5 th July 2016
 MicroRNAs Sequencing, analysis and then what? Click to edit Master subtitle Pamela Mukhopadhyay Winter School 5 th July 2016 Presentation overview Introduction sequencing analysis Identifying targets biogenesis
MicroRNAs Sequencing, analysis and then what? Click to edit Master subtitle Pamela Mukhopadhyay Winter School 5 th July 2016 Presentation overview Introduction sequencing analysis Identifying targets biogenesis
CIGNAL REPORTER ASSAYS & ARRAYS
 CIGNAL REPORTER ASSAYS & ARRAYS TM Cell-Based for Measuring Activity Cancer Cell Cycle Cytokines / Inflammation Neuroscience Stem Cell / Development Toxicology Focus on your CIGNAL TM REPORTER ASSAY SYSTEM
CIGNAL REPORTER ASSAYS & ARRAYS TM Cell-Based for Measuring Activity Cancer Cell Cycle Cytokines / Inflammation Neuroscience Stem Cell / Development Toxicology Focus on your CIGNAL TM REPORTER ASSAY SYSTEM
