Modeling Biochemical Interactions Controlling the Cytoskeleton of Spreading Cells
|
|
|
- Godfrey Maxwell
- 5 years ago
- Views:
Transcription
1 Modeling Biochemical Interactions Controlling the Cytoskeleton of Spreading Cells By Mary Motz, Sophia Rick, and Dr. Magdalena Stolarska Abstract Biological cells respond to their environment. This is done in part by physical attachments through chemical bonds to surfaces with which they interact which then sets of a biochemical signal cascade within the cell. These types of biochemical reactions can be mathematically modeled by systems of differential equations that can then be solved numerically. Such a system of equations often has many parameters that are difficult to determine. However, it is important to correctly set these parameters so that the mathematical model can make accurate predictions about cellular response. In this project, students will work with a mathematical model that describes the chemical reactions that take place when a cell attaches to a surface, and use various pieces of software, including Copasi, MatLab, and ImageJ, to correctly fit parameters to this model. Introduction Cells sense their microenvironments and respond by moving. Generally, cells migrate in four distinct ways: crawling, swimming, extension, and contraction. Cell movement is vital to countless biological processes. Cellular motility is the result of a series of complex biochemical reactions. Recently it has become much clearer that the attachment of a cell to the extracellular surface is imperative to the way the cell moves. Ultimately cell movement can be paired down to the interactions of these critical components: fibronectin, integrin, actin, and myosin. Fibronectin, a binding agent that cells like to stick to, is a protein found on extracellular surfaces. Integrin is a protein found on the cell surface. Each strand of fibronectin on the surface binds with integrin on the cell. When integrin and fibronectin bind, the process of cell movement initializes. Actin, specifically F-actin, is a protein microfilament found within cells that polymerizes and interacts with myosin, the protein that allows cells to contract locally. When actin polymerizes it develops branches, or barbed ends, which push out the edge of the cell, called the leading edge. Barbed ends are typically oriented from the interior toward the cell membrane. The polymerizing, branching, and severing
2 creates more barbed ends which allow the membrane of the cell to be pushed out. Thus, barbed ends are crucial for cell spreading. Figure 1 Figure 2 Figure 1: This is an illustration of the attachments of a cell that is sitting on surface. These attachments are what allow the cell to sense and respond to mechanical properties of the surface and its surrounding microenvironment. Figure 2: Myosin and actin filaments within a cell slide over one another to affect cell movement. While there are many internal structures that affect a cell s movement, a cell s movement is a direct response to the mechanical properties of its microenvironment. Evidence of this can be found in multiple experimental papers. In 2010, Tee and collaborators (2011) did a study regarding the effects of surface stiffness on a cell. They put cells on fibronectin-coated surfaces of varying stiffnesses and found that the stiffer the surface, the more cells spread. Not only that, but they found that the cells sensed the stiffness of the surface, and adjusted their own stiffness to match accordingly. In another study by Engler, researchers found that the fate of a stem cell is dependent on the stiffness of the surface it is on. The stem cells sensed their environment s stiffness via their attachments to the surface and became different types of tissue in response to these connections. Both experiments show the relationship between a cell s attachment to a surface and the cells response. This summer, we took a deeper look at the rate of the cell s biochemical response to its environment. This biochemical response is the series of 14 reactions between 18 intracellular components, including reactants, products, and modifiers. The system initializes when the integrins on the cell bind to the extracellular fibronectin. From there, the cascading reactions activate different proteins within the cell which ultimately sever and nucleate actin. The goal is to have local concentrations of the barbed ends of actin and myosin. Then, these concentrations can be embedded into a model of a moving cell.
3 Model Figure 3: This is the reaction pathway that we studied. It begins with integrin binding to fibronectin yielding activated integrin (A = activated species). The activated integrin then acts as a catalyst in the activation reaction of Src. And the activated Src then acts as a catalyst in the Arp2/3 and Rho activation reactions. Src is also a catalyst in the deactivation of cofilin. Activated Arp2/3 causes the elongation of actin filaments, also called actin nucleation. But actin filaments are also being severed as they grow by activated cofilin. ROCK and activated Rho bind together to get activated ROCK. Finally, activated ROCK acts as a catalyst in the Myosin activation reaction.
4 Reactions The above (Fig 3) reactions can be mathematically described by the system of differential equations below. The individual terms are based on binding, decay, and enzymatic reactions. (Sheridan)
5 Methodology To best examine and analyze the biochemical network, we used a variety of mathematical software. Specifically, we utilized MatLab, Copasi (COmplex PAthway SImulator), and ImageJ. Copasi is advanced software that allowed us to not only create a model of our biological process, but then also run simulations of biochemical pathways. This works by first allowing us to input all reactions we would like to include in our model (Figure 4). Figure 4: Input of all of our reaction in Copasi. Different reactions have different kinetics, so Copasi lets us specify these differences within each reaction (Figure 5). We can choose the rate law for each reaction from a list of pre-programed laws, or, if the rate law we desire does not already exist in Copasi, we are presented with the option of creating our own. This is also where we enter in any known parameter values. Copasi then automatically generates differential equations for each species in the reaction. Figure 5: A closer look at our first reaction in Copasi, integrin and fibronectin combining to activate integrin. Here we have selected the mass action rate law and set our parameters to k_f = and k_r = 0.168
6 Figure 6: A time course graph, measuring the concentration of the species in our reactions, generated by Copasi. Copasi can take the information that we have entered into our reactions, solve the system of equations that it generates, and create a time course graph of the species in the model (Figure 6). We can choose how long we would our time course to run and how often we would like our time steps to be. Copasi then tracks the concentration at each time step and presents us with that information in a savable format after it has completed its analysis. In order for Copasi to create time course graphs that accurately predict what is happening inside the cell, all parameters need to be set to correct values. These values can be determined by matching the model to experimental data. In order to extract experimental data from the literature in a format that can be used by Copasi, we need another piece of software called ImageJ. Once we have pulled the data we need from an image, we can use the parameter fitting tool provided by Copasi. First, we upload the data we have gathered. Then we specify which parameter we would like to fit. Copasi then uses a minimization of weighted sum of squares to find the optimal parameter value (see the formula below). We are also provided with a graph that displays a line made up of the data entered with the text file, the line of best fit generated by Copasi, and the error between the two. Results The data that we obtained was found using specific concentration of each individual species, usually µm, or µmol/l. Then using Copasi s time course tool, we could observe the evolution of each species concentration in the cell (See Figure 6). We were able to obtain data for three species: integrin, Arp2/3, and Rho. To get these results, we needed to pull experimental data from other sources. But this data was not necessarily recorded in concentrations, nor was it in table form necessary for parameter
7 estimation in Copasi. So we needed to use ImageJ to not only quantify the data, but also convert the data into the kind of data we can use (concentrations). Using ImageJ, we scaled and integrated graphs to obtain three data sets that can then be individually imported into Copasi s parameter estimation tool. a. b. c. Figure 7a-c: (a) The data for integrin evolution is given in fluorescence intensities. In extracting this data from a publication by Mould and collaborators in 2014, we scaled the intensities so that their maximum was 15. We also only used the first 100 sec in our parameter fitting. (b) The Arp2/3 experimental data is taken from a study by Goode, Drubin, and Rodal in 2001, we needed to analyze a shorter time range, so we scaled it to go from 0 to 1 s. (c) Most interestingly, the data we analyzed for the activation of Rho is taken from an article by Dubash and collaborators in 2007.This came in the form of a Western Blot. ImageJ is able to integrate the intensities of darkness to obtain a fluorescence intensity such as the one shown in (a). These integrated intensities were then scaled to a maximum value of 1. Using the data-sets we obtained from ImageJ, we then used the parameter estimation tool in Copasi. On each of the following graphs the dotted line is the experimental data that we entered into Copasi, the solid line is the line of best fit generated by Copasi, and the small circles line is the error between the two. Figure 8: Integrin Results - kcat = Figure 9: Arp Results - kcat =
8 All other parameters in the model are approximated based off of earlier work. Discussion and Conclusions This summer, we had a significant number of accomplishments. We learned how to use unknown software, Copasi and ImageJ, in order to achieve our goals. And in doing so, we found three parameter values for our reaction Figure 10: Rho Results - kcat = pathway. Unfortunately, there were many times that reaching our goal was very difficult due to the lack of existing experimental data and the nonlinearity of the reaction equations. This nonlinearity of the equations had adverse effects on the analytical tools used by Copasi so that it yielded varying results at times. Moving forward, we hope to get more viable experimental data on more of the intercellular species. This would allow us to find parameter values for the remaining approximations. Ultimately, we hope that this model will be integrated into a model of a cell that moves. References Dubash, Adi D., et al. "A novel role for Lsc/p115 RhoGEF and LARG in regulating RhoA activity downstream of adhesion to fibronectin." Journal of cell science (2007): Engler, Adam J., et al. "Matrix elasticity directs stem cell lineage specification." Cell (2006): Goode, Bruce L., et al. "Activation of the Arp2/3 complex by the actin filament binding protein Abp1p." The Journal of cell biology (2001): Hoops, Stefan, et al. "COPASI a complex pathway simulator." Bioinformatics (2006): Mould, A. Paul, et al. "Disruption of integrin fibronectin complexes by allosteric but not ligand-mimetic inhibitors." Biochemical Journal (2014):
9 Sheridan, Nicholas. Modeling biochemical interactions controlling the cytoskeleton of spreading using the Finite Element Method, CAM Summer Research Report, Tania, Nessy et al. Modeling the Synergy of Cofilin and Arp2/3 in Lamellipodial Protrusive Activity Biophysical Journal, Volume Tee, Shang-You, et al. "Cell shape and substrate rigidity both regulate cell stiffness." Biophysical journal (2011): L25-L27. Appendix Given the parameter values above, here we show time course plots of all critical species of the model. Ultimately, the goal is that the attachment of the cell to the surface leads to an increase in barbed ends, which are the primary pushing machinery in the cell. Integrin and Activated Integrin Src and Activated Src Arp2/3 and Activated Arp2/3 Rho and Activated Rho
10 ROCK and Activated ROCK Myosin and Activated Myosin Cofilin and Activated Cofilin F-Actin New and F-Actin Old Barbed Ends
11 These are tables of the parameter values and intial concentrations that we used in our Copasi model. The highlighted parameters are the values obtained using Copasi s parameter estimating tool. Parameter Values used in Copasi Integrin + Fib = Integrin Active Cofilin -> Cofilin Active k_f l/(µmol*s) k_cat /s k_r /s K_M 4 µmol/l Src = Src Active Cofilin Active -> Cofilin k_cat /s k_cat /s k_cat /s K_M 4 µmol/l K_M 6.7 µmol/l Cofilin Active -> K_M2 4 µmol/l k_sev 0.01 Arp2/3 -> Arp2/3 Active Cof_0 0.1 k_cat /s n 4 K_M 2.2 µmol/l l k_f /s FActinNew -> FActinOld Arp2/3 -> k_age 0.1 1/s k_nuc 60 -> FActinNew l V_ /s K_MA 2 µmol/l J_f 0.01 µmol/(l*s) Rho -> RhoA FActinOld -> k_cat /s k_deg /s K_M 6.7 µmol/l -> BarbedEnds RhoA + ROCK = ROCKA k_cap 0 k_f 10 l/(µmol*s) kappa 106 k_r /s k_sev 0.01 Myosin -> Myosin Active n 4 k_cat 1.8 1/s l K_M 2.47 µmol/l k_nuc 60 K_MA 2 µmol/l Cof_0 0.1 Initial Concentrations Species Concentration (µmol/l) Integrin 15 IntegrinA 0 Fibronectin 0.15 Src 1 SrcA 0 Csk 0.05 Arp2/3 1 Arp2/3A 0 Rho 1 RhoA 0 ROCK 1 ROCKA 0 Myosin 1 MyosinA 0 Cofilin 1 CofilinA 0 FActinNew 1 FActinOld 1 BarbedEnds 1
me239 mechanics of the cell me239 mechanics of the cell 3.1 biopolymers - motivation 3.2 biopolymers - polymerization
 4. the cytoskeleton - fiber bundle models bathe, heussinger, claessens, bausch, frey [2008] me239 mechanics of the cell 1 me239 mechanics of the cell 2 biopolymers polymerization of actin and tubulin Figure
4. the cytoskeleton - fiber bundle models bathe, heussinger, claessens, bausch, frey [2008] me239 mechanics of the cell 1 me239 mechanics of the cell 2 biopolymers polymerization of actin and tubulin Figure
ANAT Cell Biology Lecture 11 School of Medical Sciences The University of New South Wales. UNSW Copyright Notice
 ANAT3231 - Cell Biology Lecture 11 School of Medical Sciences The University of New South Wales The actin cytoskeleton Prof Peter Gunning Oncology Research Unit Room 502A Wallace Wurth Building Email:
ANAT3231 - Cell Biology Lecture 11 School of Medical Sciences The University of New South Wales The actin cytoskeleton Prof Peter Gunning Oncology Research Unit Room 502A Wallace Wurth Building Email:
Sélection Internationale Ens Ulm 2012, Cell Biology
 Sélection Internationale Ens Ulm 2012, Cell Biology Please read the whole subject before starting. Questions 1-8 and 12-14 are general biology questions. In questions 9, 10, 11 and15, we ask you to interpret
Sélection Internationale Ens Ulm 2012, Cell Biology Please read the whole subject before starting. Questions 1-8 and 12-14 are general biology questions. In questions 9, 10, 11 and15, we ask you to interpret
Lab Module 7: Cell Adhesion
 Lab Module 7: Cell Adhesion Tissues are made of cells and materials secreted by cells that occupy the spaces between the individual cells. This material outside of cells is called the Extracellular Matrix
Lab Module 7: Cell Adhesion Tissues are made of cells and materials secreted by cells that occupy the spaces between the individual cells. This material outside of cells is called the Extracellular Matrix
Lecture 13. Motor Proteins I
 Lecture 13 Motor Proteins I Introduction: The study of motor proteins has become a major focus in cell and molecular biology. Motor proteins are very interesting because they do what no man-made engines
Lecture 13 Motor Proteins I Introduction: The study of motor proteins has become a major focus in cell and molecular biology. Motor proteins are very interesting because they do what no man-made engines
Sabrina Jedlicka. Interactions of Neurons and Materials
 Sabrina Jedlicka Interactions of Neurons and Materials Developmental Cell Biology Some Perspective Stem cells: Undifferentiated cells that are capable of proliferation, self-maintenance and differentiation
Sabrina Jedlicka Interactions of Neurons and Materials Developmental Cell Biology Some Perspective Stem cells: Undifferentiated cells that are capable of proliferation, self-maintenance and differentiation
Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question in Section B and ONE question from Section C.
 UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 2017-18 CELL BIOLOGY BIO-5005B Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question in Section
UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 2017-18 CELL BIOLOGY BIO-5005B Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question in Section
me239 mechanics of the cell homework biopolymers - motivation 3.1 biopolymers - motivation 3. biopolymers biopolymers biopolymers
 3. biopolymers the inner life of a cell, viel & lue, harvard [2006] me239 mechanics of the cell 1 2 biopolymers Figure 3.1. Biopolymers. Characteristic length scales on the cellular and sucellular level..
3. biopolymers the inner life of a cell, viel & lue, harvard [2006] me239 mechanics of the cell 1 2 biopolymers Figure 3.1. Biopolymers. Characteristic length scales on the cellular and sucellular level..
Cell Migration, the Cytoskeleton, Chemotaxis, and Haptotaxis. 3/9/17 ChE 575
 Cell Migration, the Cytoskeleton, Chemotaxis, and Haptotaxis 3/9/17 ChE 575 When, Where, Why do cells migrate? 1. Neutrophil Migration to Battle Infection 2. Development 3. Wound Healing 4. Disease 2 Wound
Cell Migration, the Cytoskeleton, Chemotaxis, and Haptotaxis 3/9/17 ChE 575 When, Where, Why do cells migrate? 1. Neutrophil Migration to Battle Infection 2. Development 3. Wound Healing 4. Disease 2 Wound
Single Cell Motility in Artificial Tissue
 Single Cell Motility in Artificial Tissue Rhoda J. Hawkins Lecturer in Physics, University of Sheffield Raphaël Voituriez, Jean-François Joanny, Jacques Prost Matthieu Piel, Ana-Maria Lennon-Dumenil, Philippe
Single Cell Motility in Artificial Tissue Rhoda J. Hawkins Lecturer in Physics, University of Sheffield Raphaël Voituriez, Jean-François Joanny, Jacques Prost Matthieu Piel, Ana-Maria Lennon-Dumenil, Philippe
cell fusion live and fixed imaging genetics biochemistry in vitro systems inhibitors of cellular processes (transcription, replication, microtubules)
 DISCUSSION SECTIONS BY STUDENT NUMBER ENDING IN ODD NUMBERS 2-3, EVEN 3-4 Methods for Studying the Cell Cycle cell fusion live and fixed imaging genetics biochemistry in vitro systems inhibitors of cellular
DISCUSSION SECTIONS BY STUDENT NUMBER ENDING IN ODD NUMBERS 2-3, EVEN 3-4 Methods for Studying the Cell Cycle cell fusion live and fixed imaging genetics biochemistry in vitro systems inhibitors of cellular
Evidence that Pseudopodia Formation Decreases with Cytochalasin B
 Evidence that Pseudopodia Formation Decreases with Cytochalasin B Jessica Emory Independent Research Project Report Bio 219- Cell Biology December 3, 2008 Introduction Amoeboid movement is the result of
Evidence that Pseudopodia Formation Decreases with Cytochalasin B Jessica Emory Independent Research Project Report Bio 219- Cell Biology December 3, 2008 Introduction Amoeboid movement is the result of
6 pts. B) State two mechanisms to explain the effect shown in the figure.
 Question 1. (10 points) The rate of uptake of glucose into rat adipocytes is increased by addition of insulin to the medium. The increase reaches a maximum within 30 seconds at 37 C and is energy dependent.
Question 1. (10 points) The rate of uptake of glucose into rat adipocytes is increased by addition of insulin to the medium. The increase reaches a maximum within 30 seconds at 37 C and is energy dependent.
Changes in E-cadherin rigidity sensing regulate cell adhesion
 Changes in E-cadherin rigidity sensing regulate cell adhesion Caitlin Collins a, Aleksandra K. Denisin b,c, Beth L. Pruitt b,c, and W. James Nelson a,1 a Department of Biology, Stanford University, Stanford,
Changes in E-cadherin rigidity sensing regulate cell adhesion Caitlin Collins a, Aleksandra K. Denisin b,c, Beth L. Pruitt b,c, and W. James Nelson a,1 a Department of Biology, Stanford University, Stanford,
Supplemental Information. A Computational Model of YAP/TAZ Mechanosensing. Meng Sun, Fabian Spill, and Muhammad H. Zaman
 Biophysical Journal, Volume 110 Supplemental Information A Computational Model of YAP/TAZ Mechanosensing Meng Sun, Fabian Spill, and Muhammad H. Zaman Supporting Methods Molecular intervention When latrunculin
Biophysical Journal, Volume 110 Supplemental Information A Computational Model of YAP/TAZ Mechanosensing Meng Sun, Fabian Spill, and Muhammad H. Zaman Supporting Methods Molecular intervention When latrunculin
MCBII. Points this page
 1. What makes intermediate filaments (IFs) an inefficient track for motor proteins (4 pts)? A. The outer surface of IFs is hydrophobic. B. IFs are nonpolar structures. C. IFs contain coiled coil domains.
1. What makes intermediate filaments (IFs) an inefficient track for motor proteins (4 pts)? A. The outer surface of IFs is hydrophobic. B. IFs are nonpolar structures. C. IFs contain coiled coil domains.
Cell-Extracellular Matrix Interactions
 Cell-Extracellular Matrix Interactions Extracellular Matrix (ECM) Organised network outside of the cell s plasma membrane Between cells Composition varies throughout the ECM Continuous sheet of ECM = Basement
Cell-Extracellular Matrix Interactions Extracellular Matrix (ECM) Organised network outside of the cell s plasma membrane Between cells Composition varies throughout the ECM Continuous sheet of ECM = Basement
Dimeric WH2 domains in Vibrio VopF promote actin filament barbed end uncapping and assisted elongation
 Dimeric WH2 domains in Vibrio VopF promote actin filament barbed end uncapping and assisted elongation Julien Pernier, Jozsef Orban 1, Balendu Sankara Avvaru, Antoine Jégou, Guillaume Romet- Lemonne, Bérengère
Dimeric WH2 domains in Vibrio VopF promote actin filament barbed end uncapping and assisted elongation Julien Pernier, Jozsef Orban 1, Balendu Sankara Avvaru, Antoine Jégou, Guillaume Romet- Lemonne, Bérengère
Ac(n cytoskeleton 12/7/11. The Cytoskeleton. Func0ons of the Cytoskeleton. B Visegrady. Elements of the Cytoskeleton
 The Cytoskeleton Ac(n cytoskeleton B Visegrady The cytoskeleton is an intricate network of protein filaments that extends throughout the cytoplasm. A skin cell (fibroblast) stained with Coomassie blue.
The Cytoskeleton Ac(n cytoskeleton B Visegrady The cytoskeleton is an intricate network of protein filaments that extends throughout the cytoplasm. A skin cell (fibroblast) stained with Coomassie blue.
Supplemental figures (1-9) Gu et al. ADF/Cofilin-Mediated Actin Dynamics Regulate AMPA Receptor Trafficking during Synaptic Plasticity
 Supplemental figures (1-9) Gu et al. ADF/Cofilin-Mediated Actin Dynamics Regulate AMPA Receptor Trafficking during Synaptic Plasticity Supplemental figure 1. The quantification method to determine the
Supplemental figures (1-9) Gu et al. ADF/Cofilin-Mediated Actin Dynamics Regulate AMPA Receptor Trafficking during Synaptic Plasticity Supplemental figure 1. The quantification method to determine the
Rapid Chromatin Condensation Increases Stem Cell Nuclear Mechanics and Mechanosensitivity
 Rapid Chromatin Condensation Increases Stem Cell Nuclear Mechanics and Mechanosensitivity Su-Jin Heo, MS 1, Tristan P. Driscoll, BS 1, Stephen Thorpe, PhD 2, Woojin M. Han, MS 1, Dawn M. Elliott, PhD 3,
Rapid Chromatin Condensation Increases Stem Cell Nuclear Mechanics and Mechanosensitivity Su-Jin Heo, MS 1, Tristan P. Driscoll, BS 1, Stephen Thorpe, PhD 2, Woojin M. Han, MS 1, Dawn M. Elliott, PhD 3,
Cell-Environment Interactions. Chieh-Chun Chen
 Cell-Environment Interactions Chieh-Chun Chen Part 1: Soft Lithography in Biology and Biochemistry Chieh-Chun Chen Outlines Introduction Key features of soft lithography Applications In microscopic biochemical
Cell-Environment Interactions Chieh-Chun Chen Part 1: Soft Lithography in Biology and Biochemistry Chieh-Chun Chen Outlines Introduction Key features of soft lithography Applications In microscopic biochemical
Sabrina Jedlicka. Interactions of Neurons and Materials
 Sabrina Jedlicka Interactions of Neurons and Materials Neurological Disorders Over 600 known neurological disorders ú Diseases of the CNS and PNS Epilepsy Alzheimer s Parkinson s Etc. ú Diseases that attack
Sabrina Jedlicka Interactions of Neurons and Materials Neurological Disorders Over 600 known neurological disorders ú Diseases of the CNS and PNS Epilepsy Alzheimer s Parkinson s Etc. ú Diseases that attack
System of protein filaments in the cytoplasm of. eukaryotic cell that gives the cell shape and capacity for
 Cytoskeleton System of protein filaments in the cytoplasm of eukaryotic cell that gives the cell shape and capacity for directed movement. The protein filaments are responsible for the shaping, moving,
Cytoskeleton System of protein filaments in the cytoplasm of eukaryotic cell that gives the cell shape and capacity for directed movement. The protein filaments are responsible for the shaping, moving,
Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question in Section B and ONE question from Section C.
 UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 2013-2014 CELL BIOLOGY BIO-2B06 Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question in
UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 2013-2014 CELL BIOLOGY BIO-2B06 Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question in
BME Engineering Molecular Cell Biology. The Cytoskeleton (I): Actin The Cytoskeleton (II): Microtubule & Intermediate Filament
 BME 42-620 Engineering Molecular Cell Biology Lecture 09: The Cytoskeleton (I): Actin The Cytoskeleton (II): Microtubule & Intermediate Filament BME42-620 Lecture 09, September 27, 2011 1 Outline Overviewofcytoskeletal
BME 42-620 Engineering Molecular Cell Biology Lecture 09: The Cytoskeleton (I): Actin The Cytoskeleton (II): Microtubule & Intermediate Filament BME42-620 Lecture 09, September 27, 2011 1 Outline Overviewofcytoskeletal
Arterial Tissue Mimics for Studying Cerebral Aneurysm Formation
 Arterial Tissue Mimics for Studying Cerebral Aneurysm Formation Emily N. Sevcik Faculty Mentor: Patrick W. Alford Undergraduate Research Opportunities Program Project Final Report Spring 2014 Project and
Arterial Tissue Mimics for Studying Cerebral Aneurysm Formation Emily N. Sevcik Faculty Mentor: Patrick W. Alford Undergraduate Research Opportunities Program Project Final Report Spring 2014 Project and
General Education Learning Outcomes
 BOROUGH OF MANHATTAN COMMUNITY COLLEGE City University of New York Department of Science Title of Course: Cell Biology Class hours 3 BIO Section: 260 Lab hours 3 Semester Spring 2018 Credits 4 Schedule:
BOROUGH OF MANHATTAN COMMUNITY COLLEGE City University of New York Department of Science Title of Course: Cell Biology Class hours 3 BIO Section: 260 Lab hours 3 Semester Spring 2018 Credits 4 Schedule:
Michaelis Menten Kinetics -Enzyme Kinetics, Binding and Cooperativity
 Michaelis Menten Kinetics -Enzyme Kinetics, Binding and Cooperativity Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by MHRD Page 1
Michaelis Menten Kinetics -Enzyme Kinetics, Binding and Cooperativity Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by MHRD Page 1
A Computational Model of Cell Movement Linked to Substrate Rigidity
 Lehigh University Lehigh Preserve Theses and Dissertations 2015 A Computational Model of Cell Movement Linked to Substrate Rigidity Jie Sheng Lehigh University Follow this and additional works at: http://preserve.lehigh.edu/etd
Lehigh University Lehigh Preserve Theses and Dissertations 2015 A Computational Model of Cell Movement Linked to Substrate Rigidity Jie Sheng Lehigh University Follow this and additional works at: http://preserve.lehigh.edu/etd
LIFE Formation III. Morphogenesis: building 3D structures START FUTURE. How-to 2 PROBLEMS FORMATION SYSTEMS SYSTEMS.
 7.013 4.6.07 Formation III Morphogenesis: building 3D structures VIRUSES CANCER HUMAN DISEASE START SYSTEMS BIOLOGY PROBLEMS FUTURE LIFE IMMUNE NERVOUS SYSTEMS How-to 1 FOUNDATIONS How-to 2 REC. DNA BIOCHEM
7.013 4.6.07 Formation III Morphogenesis: building 3D structures VIRUSES CANCER HUMAN DISEASE START SYSTEMS BIOLOGY PROBLEMS FUTURE LIFE IMMUNE NERVOUS SYSTEMS How-to 1 FOUNDATIONS How-to 2 REC. DNA BIOCHEM
Three major types of cytoskeleton
 The Cytoskeleton Organizes and stabilizes cells Pulls chromosomes apart Drives intracellular traffic Supports plasma membrane and nuclear envelope Enables cellular movement Guides growth of the plant cell
The Cytoskeleton Organizes and stabilizes cells Pulls chromosomes apart Drives intracellular traffic Supports plasma membrane and nuclear envelope Enables cellular movement Guides growth of the plant cell
Molecular Cell Biology - Problem Drill 01: Introduction to Molecular Cell Biology
 Molecular Cell Biology - Problem Drill 01: Introduction to Molecular Cell Biology Question No. 1 of 10 1. Which statement describes how an organism is organized from most simple to most complex? Question
Molecular Cell Biology - Problem Drill 01: Introduction to Molecular Cell Biology Question No. 1 of 10 1. Which statement describes how an organism is organized from most simple to most complex? Question
G = H (T S) I. Cellular Metabolism & Reaction Coupling Figure 1: Metabolism
 I. Cellular Metabolism & Reaction Coupling Figure 1: Metabolism Metabolism represents the sum total of ALL chemical reactions within the cell. These reactions can be regarded as either catabolic or anabolic
I. Cellular Metabolism & Reaction Coupling Figure 1: Metabolism Metabolism represents the sum total of ALL chemical reactions within the cell. These reactions can be regarded as either catabolic or anabolic
RESEARCH OPPORTUNITY PROGRAM 299Y/399Y PROJECT DESCRIPTIONS SUMMER
 Project Code: CSB 1S Joshua Currie, Assistant Professor TITLE OF RESEARCH PROJECT: Self Organization of the Regenerative Blastema Number of 299Y Spots: 1 Number of 399Y Spots: 1 We are interested in the
Project Code: CSB 1S Joshua Currie, Assistant Professor TITLE OF RESEARCH PROJECT: Self Organization of the Regenerative Blastema Number of 299Y Spots: 1 Number of 399Y Spots: 1 We are interested in the
Final exam. Please write your name on the exam and keep an ID card ready.
 Biophysics of Macromolecules Prof. R. Jungmann and Prof. J. Lipfert SS 2017 Final exam Final exam First name: Last name: Student number ( Matrikelnummer ): Please write your name on the exam and keep an
Biophysics of Macromolecules Prof. R. Jungmann and Prof. J. Lipfert SS 2017 Final exam Final exam First name: Last name: Student number ( Matrikelnummer ): Please write your name on the exam and keep an
6 Enzymes II W. H. Freeman and Company
 6 Enzymes II 2017 W. H. Freeman and Company The role of an enzyme in an enzyme-catalyzed reaction is to: A. bind a transition state intermediate, such that it cannot be converted back to substrate. B.
6 Enzymes II 2017 W. H. Freeman and Company The role of an enzyme in an enzyme-catalyzed reaction is to: A. bind a transition state intermediate, such that it cannot be converted back to substrate. B.
Tracking Cellular Protein Localization and Movement in Cells with a Flexible Fluorescent Labeling Technology. Chad Zimprich January 2015
 Tracking Cellular Protein Localization and Movement in Cells with a Flexible Fluorescent Labeling Technology Chad Zimprich January 2015 Presentation verview HaloTag Fusion Technology Design Functionality
Tracking Cellular Protein Localization and Movement in Cells with a Flexible Fluorescent Labeling Technology Chad Zimprich January 2015 Presentation verview HaloTag Fusion Technology Design Functionality
Practice Exam 3 MCBII
 1. Which is not a feature of lamin intermediate filaments (IFs)? (4 pts) A. They are protein polymers. B. They are used by motor proteins to traffic cargo. C. They contain coiled coil domains. D. They
1. Which is not a feature of lamin intermediate filaments (IFs)? (4 pts) A. They are protein polymers. B. They are used by motor proteins to traffic cargo. C. They contain coiled coil domains. D. They
High-Throughput Method for Microfluidic Placement of Cells in Micropatterned Tissues
 High-Throughput Method for Microfluidic Placement of Cells in Micropatterned Tissues Emily N. Sevcik Faculty Mentor: Patrick W. Alford Undergraduate Research Opportunities Program Project Final Report
High-Throughput Method for Microfluidic Placement of Cells in Micropatterned Tissues Emily N. Sevcik Faculty Mentor: Patrick W. Alford Undergraduate Research Opportunities Program Project Final Report
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Integrative Biology. This journal is The Royal Society of Chemistry 2015 Electronic Supplementary Information Table S1. Definition of quantitative cellular features
Electronic Supplementary Material (ESI) for Integrative Biology. This journal is The Royal Society of Chemistry 2015 Electronic Supplementary Information Table S1. Definition of quantitative cellular features
Supplementary Figure 1. Soft fibrin gels promote growth and organized mesodermal differentiation. Representative images of single OGTR1 ESCs cultured
 Supplementary Figure 1. Soft fibrin gels promote growth and organized mesodermal differentiation. Representative images of single OGTR1 ESCs cultured in 90-Pa 3D fibrin gels for 5 days in the presence
Supplementary Figure 1. Soft fibrin gels promote growth and organized mesodermal differentiation. Representative images of single OGTR1 ESCs cultured in 90-Pa 3D fibrin gels for 5 days in the presence
Actin cap associated focal adhesions and their distinct role in cellular mechanosensing
 Actin cap associated focal adhesions and their distinct role in cellular mechanosensing Dong-Hwee Kim 1,2, Shyam B. Khatau 1,2, Yunfeng Feng 1,3, Sam Walcott 1,4, Sean X. Sun 1,2,4, Gregory D. Longmore
Actin cap associated focal adhesions and their distinct role in cellular mechanosensing Dong-Hwee Kim 1,2, Shyam B. Khatau 1,2, Yunfeng Feng 1,3, Sam Walcott 1,4, Sean X. Sun 1,2,4, Gregory D. Longmore
Analysis of cytoskeletal features in dried and living cells and slow dynamic experiments using Atomic Force Microscopy
 Analysis of cytoskeletal features in dried and living cells and slow dynamic experiments using Atomic Force Microscopy Author Watson, Gregory, Watson, Jolanta Published 008 Journal Title Journal of Physics:
Analysis of cytoskeletal features in dried and living cells and slow dynamic experiments using Atomic Force Microscopy Author Watson, Gregory, Watson, Jolanta Published 008 Journal Title Journal of Physics:
CHAPTER 16 THE CYTOSKELETON
 CHAPTER 16 THE CYTOSKELETON THE SELF-ASSEMBLY AND DYNAMIC STRUCTURE OF CYTOSKELETAL FILAMENTS HOW CELLS REGULATE THEIR CYTOSKELETAL FILAMENTS MOLECULAR MOTORS THE CYTOSKELETON AND CELL BEHAVIOR THE SELF-ASSEMBLY
CHAPTER 16 THE CYTOSKELETON THE SELF-ASSEMBLY AND DYNAMIC STRUCTURE OF CYTOSKELETAL FILAMENTS HOW CELLS REGULATE THEIR CYTOSKELETAL FILAMENTS MOLECULAR MOTORS THE CYTOSKELETON AND CELL BEHAVIOR THE SELF-ASSEMBLY
An Up-Close Model Characterizing a Highly Nonlinear, Fluffy-Cored, Skinned, Composite Material Used in a Local Buckling Analysis
 An Up-Close Model Characterizing a Highly Nonlinear, Fluffy-Cored, Skinned, Composite Material Used in a Local Buckling Analysis Ted Diehl 1, Jay G. Sloan 1, Delisia L. Dickerson 1, Mark A. Lamontia 1,
An Up-Close Model Characterizing a Highly Nonlinear, Fluffy-Cored, Skinned, Composite Material Used in a Local Buckling Analysis Ted Diehl 1, Jay G. Sloan 1, Delisia L. Dickerson 1, Mark A. Lamontia 1,
Printing of biochemically-patterned slides with the InnoStamp40 for deterministic cell immobilization.
 Printing of biochemically-patterned slides with the InnoStamp40 for deterministic cell immobilization. Jean christophe CAU 1 1 Innopsys, Parc activestre, Carbonne France contact@innopsys.fr www.innopsys.com
Printing of biochemically-patterned slides with the InnoStamp40 for deterministic cell immobilization. Jean christophe CAU 1 1 Innopsys, Parc activestre, Carbonne France contact@innopsys.fr www.innopsys.com
JCB. Supplemental material THE JOURNAL OF CELL BIOLOGY. Paul et al.,
 Supplemental material JCB Paul et al., http://www.jcb.org/cgi/content/full/jcb.201502040/dc1 THE JOURNAL OF CELL BIOLOGY Figure S1. Mutant p53-expressing cells display limited retrograde actin flow at
Supplemental material JCB Paul et al., http://www.jcb.org/cgi/content/full/jcb.201502040/dc1 THE JOURNAL OF CELL BIOLOGY Figure S1. Mutant p53-expressing cells display limited retrograde actin flow at
Michal Reichman-Fried
 Zebrafish Development and Genetics course 2013 Cell labeling and migration Imaging primordial germ cell migration Instructors: Erez Raz (erez.raz@uni-muenster.de) Michal Reichman-Fried (mreichm@uni-muenster.de)
Zebrafish Development and Genetics course 2013 Cell labeling and migration Imaging primordial germ cell migration Instructors: Erez Raz (erez.raz@uni-muenster.de) Michal Reichman-Fried (mreichm@uni-muenster.de)
Unit title: Biochemistry: Theory and Laboratory Skills (SCQF level 7)
 Higher National Unit specification General information Unit code: H922 34 Superclass: RH Publication date: May 2015 Source: Scottish Qualifications Authority Version: 01 Unit purpose This Unit is designed
Higher National Unit specification General information Unit code: H922 34 Superclass: RH Publication date: May 2015 Source: Scottish Qualifications Authority Version: 01 Unit purpose This Unit is designed
What can you remember about enzymes? Mr W
 What can you remember about enzymes? Mr W Human Cells (f) Enzymes and Metabolism Learning Intentions Describe metabolism, synthetic (energy requiring) and breakdown (energy releasing) pathways Cell Metabolism
What can you remember about enzymes? Mr W Human Cells (f) Enzymes and Metabolism Learning Intentions Describe metabolism, synthetic (energy requiring) and breakdown (energy releasing) pathways Cell Metabolism
Golgi are located near the MTOC while the ER is spread throughout the cytoplasm, so which of the following is probably true?
 Golgi are located near the MTOC while the ER is spread throughout the cytoplasm, so which of the following is probably true? A. COPII and COPI vesicles are transported with dynein. B. COPII and COPI vesicles
Golgi are located near the MTOC while the ER is spread throughout the cytoplasm, so which of the following is probably true? A. COPII and COPI vesicles are transported with dynein. B. COPII and COPI vesicles
The Thermo Scientific Nunclon Sphera surface supports formation of embryoid bodies from pluripotent stem cells
 1 The Thermo Scientific surface supports formation of embryoid bodies from pluripotent stem cells Louise Gaarn, Amy Sinor-Anderson, Robert Scott, Cindy Neeley, Joseph Granchelli, and Tina Marwood Application
1 The Thermo Scientific surface supports formation of embryoid bodies from pluripotent stem cells Louise Gaarn, Amy Sinor-Anderson, Robert Scott, Cindy Neeley, Joseph Granchelli, and Tina Marwood Application
Pathway Tools Schema and Semantic Inference Layer: Pathways and the Overview. SRI International Bioinformatics
 Pathway Tools Schema and Semantic Inference Layer: Pathways and the Overview 1 Outline Pathways Representation of Pathways Querying Pathways Programmatically How Pathway Diagrams are Generated Future Work:
Pathway Tools Schema and Semantic Inference Layer: Pathways and the Overview 1 Outline Pathways Representation of Pathways Querying Pathways Programmatically How Pathway Diagrams are Generated Future Work:
Biophysics of contractile ring assembly
 Biophysics of contractile ring assembly Dimitrios Vavylonis Department of Physics, Lehigh University October 1, 2007 Physical biology of the cell Physical processes in cell organization and function: Transport
Biophysics of contractile ring assembly Dimitrios Vavylonis Department of Physics, Lehigh University October 1, 2007 Physical biology of the cell Physical processes in cell organization and function: Transport
ANAT Cell Biology Lecture 12 School of Medical Sciences The University of New South Wales
 ANAT3231 - Cell Biology Lecture 12 School of Medical Sciences The University of New South Wales Prof Peter Gunning Microfilaments, 2010 Sllide 1 UNSW Copyright Notice COMMONWEALTH OF AUSTRALIA Copyright
ANAT3231 - Cell Biology Lecture 12 School of Medical Sciences The University of New South Wales Prof Peter Gunning Microfilaments, 2010 Sllide 1 UNSW Copyright Notice COMMONWEALTH OF AUSTRALIA Copyright
Resolving Macromolecular Organization in Cells Using Super Resolution Microscopy Karen Porter-Davis Chamblee Charter High School
 Resolving Macromolecular Organization in Cells Using Super Resolution Microscopy Karen Porter-Davis Chamblee Charter High School STEP-UP Program 2015 Super Resolution Imaging } Two Projects to Image }
Resolving Macromolecular Organization in Cells Using Super Resolution Microscopy Karen Porter-Davis Chamblee Charter High School STEP-UP Program 2015 Super Resolution Imaging } Two Projects to Image }
Journal of Biological Engineering is an open access, peer-reviewed online journal that encompasses all aspects biological engineering.
 Journal of Biological Engineering is an open access, peer-reviewed online journal that encompasses all aspects biological engineering. The journal welcomes manuscripts on the following topics: synthetic
Journal of Biological Engineering is an open access, peer-reviewed online journal that encompasses all aspects biological engineering. The journal welcomes manuscripts on the following topics: synthetic
Introduction. Figure 1. Oris Cell Migration Assay Principle
 Optimizing Performance of the Membrane-free, Oris Cell Migration Assay for High Throughput Screening using the BioTek Synergy HT Multi-Mode Microplate Reader Keren I. Hulkower, Renee L. Herber, and Scott
Optimizing Performance of the Membrane-free, Oris Cell Migration Assay for High Throughput Screening using the BioTek Synergy HT Multi-Mode Microplate Reader Keren I. Hulkower, Renee L. Herber, and Scott
System. Dynamic Monitoring of Receptor Tyrosine Kinase Activation in Living Cells. Application Note No. 4 / March
 System Application Note No. 4 / March 2008 Dynamic Monitoring of Receptor Tyrosine Kinase Activation in Living Cells www.roche-applied-science.com Dynamic Monitoring of Receptor Tyrosine Kinase Activation
System Application Note No. 4 / March 2008 Dynamic Monitoring of Receptor Tyrosine Kinase Activation in Living Cells www.roche-applied-science.com Dynamic Monitoring of Receptor Tyrosine Kinase Activation
The cytoskeleton. The cytoskeleton, the motor proteins, the muscle and its regulation. The cytoskeleton. The cytoskeleton.
 , the motor proteins, the muscle and its regulation Dept. of Biophysics, University of Pécs Zoltán Ujfalusi January-February 2012 Dynamic framework of the Eukaryotes Three main filament-class: 1. Intermedier
, the motor proteins, the muscle and its regulation Dept. of Biophysics, University of Pécs Zoltán Ujfalusi January-February 2012 Dynamic framework of the Eukaryotes Three main filament-class: 1. Intermedier
ENZYMES. Unit 3 - Energy
 ENZYMES Unit 3 - Energy What is an enzyme? What do they do? What is an enzyme? What do they do? Key Things to remember: They are proteins They are catalysts They are reusable - not consumed in reaction
ENZYMES Unit 3 - Energy What is an enzyme? What do they do? What is an enzyme? What do they do? Key Things to remember: They are proteins They are catalysts They are reusable - not consumed in reaction
SUPPLEMENTARY INFORMATION
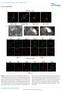 DOI: 10.1038/ncb2386 Figure 1 Src-containing puncta are not focal adhesions, podosomes or endosomes. (a) FAK-/- were stained with anti-py416 Src (green) and either (in red) the focal adhesion protein paxillin,
DOI: 10.1038/ncb2386 Figure 1 Src-containing puncta are not focal adhesions, podosomes or endosomes. (a) FAK-/- were stained with anti-py416 Src (green) and either (in red) the focal adhesion protein paxillin,
Cellular Biomechanics. Linda Lowe-Krentz Bioscience in the 21 st Century October 17, 2011
 Cellular Biomechanics Linda Lowe-Krentz Bioscience in the 21 st Century October 17, 2011 Outline for today Tubes Vessel mechanics and disease Models Lung mechanics Technology integration Atherosclerosis
Cellular Biomechanics Linda Lowe-Krentz Bioscience in the 21 st Century October 17, 2011 Outline for today Tubes Vessel mechanics and disease Models Lung mechanics Technology integration Atherosclerosis
Cells Culture Techniques Marta Czernik
 Cells Culture Techniques 13.03.2018 Marta Czernik Why we need the cell/tissue culture Research To overcome problems in studying cellular behaviour such as: - confounding effects of the surrounding tissues
Cells Culture Techniques 13.03.2018 Marta Czernik Why we need the cell/tissue culture Research To overcome problems in studying cellular behaviour such as: - confounding effects of the surrounding tissues
Title: The cleaved FAS ligand activates the Na + /H + exchanger NHE1 through. Akt/ROCK1 to stimulate cell motility.
 Title: The cleaved FAS ligand activates the Na + /H + exchanger NHE through Akt/ROCK to stimulate cell motility. Authors : Monet Michael, Poët Mallorie, Tauzin Sébastien 2,#, Fouqué Amélie 3, Cophignon
Title: The cleaved FAS ligand activates the Na + /H + exchanger NHE through Akt/ROCK to stimulate cell motility. Authors : Monet Michael, Poët Mallorie, Tauzin Sébastien 2,#, Fouqué Amélie 3, Cophignon
The cytoskeletal system
 The cytoskeletal system Types, polymerization, characteristics and mechanical properties of cytoskeletal filaments. Associated protein. 24.09.2013. Cytoskeleton The cellular "scaffold" or "skeleton" within
The cytoskeletal system Types, polymerization, characteristics and mechanical properties of cytoskeletal filaments. Associated protein. 24.09.2013. Cytoskeleton The cellular "scaffold" or "skeleton" within
TEKS and S.E.s. B.9C identify and investigate the role of enzymes
 Enzymes TEKS and S.E.s B.9C identify and investigate the role of enzymes Vocabulary Enzyme Catalyst Substrate Active site Substrate-enzyme complex Activation energy Inhibitor Catabolic Anabolic Reactant
Enzymes TEKS and S.E.s B.9C identify and investigate the role of enzymes Vocabulary Enzyme Catalyst Substrate Active site Substrate-enzyme complex Activation energy Inhibitor Catabolic Anabolic Reactant
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
DNA Circuits for Analog Computing
 DNA Circuits for Analog Computing Tianqi Song Department of Computer Science Duke University 1 Outline Motivation What is DNA computing? Why are we interested in DNA computing? What has been done? Why
DNA Circuits for Analog Computing Tianqi Song Department of Computer Science Duke University 1 Outline Motivation What is DNA computing? Why are we interested in DNA computing? What has been done? Why
BIOLOGY 1 SCIENCE AND ENGINEERING PRACTICES
 OVERVIEW The academic standards and performance indicators establish the practices and core content for all Biology courses in South Carolina high schools. The core ideas within the standards are not meant
OVERVIEW The academic standards and performance indicators establish the practices and core content for all Biology courses in South Carolina high schools. The core ideas within the standards are not meant
MICB688L/MOCB639 Advanced Cell Biology Exam II
 MICB688L/MOCB639 Advanced Cell Biology Exam II May 10, 2001 Name: 1. Briefly describe the four major classes of cell surface receptors and their modes of action (immediate downstream only) (8) 2. Please
MICB688L/MOCB639 Advanced Cell Biology Exam II May 10, 2001 Name: 1. Briefly describe the four major classes of cell surface receptors and their modes of action (immediate downstream only) (8) 2. Please
Structural Study of Actin Bundles
 Structural Study of Actin Bundles Danielle Aranda Mentor: Dr. Linda Hirst Advisor: Professor Cyrus Safinya Major Funding: National Science Foundation UCSB Materials Research Lab Vocabulary α-actinin G-actin
Structural Study of Actin Bundles Danielle Aranda Mentor: Dr. Linda Hirst Advisor: Professor Cyrus Safinya Major Funding: National Science Foundation UCSB Materials Research Lab Vocabulary α-actinin G-actin
I. Enzyme Action Figure 1: Activation Energy (Ea) Activation Energy (Ea):
 I. Enzyme Action Figure 1: Activation Energy (Ea) *Activation energy represents an energy barrier that reactant molecules must overcome in order to react & form products. Think of an Olympic track star
I. Enzyme Action Figure 1: Activation Energy (Ea) *Activation energy represents an energy barrier that reactant molecules must overcome in order to react & form products. Think of an Olympic track star
Substrate rigidity and force define form through tyrosine phosphatase and kinase pathways
 Review TRENDS in Cell Biology Vol.16 No.4 April 2006 Substrate rigidity and force define form through tyrosine phosphatase and kinase pathways Grégory Giannone 1,2 and Michael. Sheetz 1 1 Department of
Review TRENDS in Cell Biology Vol.16 No.4 April 2006 Substrate rigidity and force define form through tyrosine phosphatase and kinase pathways Grégory Giannone 1,2 and Michael. Sheetz 1 1 Department of
The Integrated Biomedical Sciences Graduate Program
 The Integrated Biomedical Sciences Graduate Program at the university of notre dame Cutting-edge biomedical research and training that transcends traditional departmental and disciplinary boundaries to
The Integrated Biomedical Sciences Graduate Program at the university of notre dame Cutting-edge biomedical research and training that transcends traditional departmental and disciplinary boundaries to
1 Scaffolds. 2 Extracellular matrix
 This lecture describes the components of the extracellular matrix and their influence on the cell properties and functions. 1 Scaffolds One of the key components of the tissue engineering triad is the
This lecture describes the components of the extracellular matrix and their influence on the cell properties and functions. 1 Scaffolds One of the key components of the tissue engineering triad is the
Promotion of HDF Cell Attachment and Proliferation
 Promotion of HDF Cell Attachment and Proliferation Objectives To qualitatively assess the effect of fibronectin (Fn) on HDF cell attachment Fn Attachment Assay To observe HDF cell proliferation and position
Promotion of HDF Cell Attachment and Proliferation Objectives To qualitatively assess the effect of fibronectin (Fn) on HDF cell attachment Fn Attachment Assay To observe HDF cell proliferation and position
Biology Eighth Edition Neil Campbell and Jane Reece
 BIG IDEA IV Biological systems interact, and these systems and their interactions possess complex properties. Enduring Understanding 4.B Competition and cooperation are important aspects of biological
BIG IDEA IV Biological systems interact, and these systems and their interactions possess complex properties. Enduring Understanding 4.B Competition and cooperation are important aspects of biological
Cell-Biomaterial Mechanical Interaction in the Framework of Tissue Engineering: Insights, Computational Modeling and Perspectives
 Int. J. Mol. Sci. 2011, 12, 8217-8244; doi:10.3390/ijms12118217 Review OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Cell-Biomaterial Mechanical Interaction
Int. J. Mol. Sci. 2011, 12, 8217-8244; doi:10.3390/ijms12118217 Review OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Cell-Biomaterial Mechanical Interaction
1. The microtubule wall is composed of globular proteins arranged in longitudinal rows called.
 Name: Quiz name: Quiz 7 ate: 1. The microtubule wall is composed of globular proteins arranged in longitudinal rows called. microfilaments protofilaments prototubules microtubular subunits 2. Which of
Name: Quiz name: Quiz 7 ate: 1. The microtubule wall is composed of globular proteins arranged in longitudinal rows called. microfilaments protofilaments prototubules microtubular subunits 2. Which of
Force Measurement Software Module
 Microscopy from Carl Zeiss Force Measurement Software Module Optical tweezers are used to manipulate microscopic particles such as cells or sub-cellular structures. The Force Measurement (FM) module expands
Microscopy from Carl Zeiss Force Measurement Software Module Optical tweezers are used to manipulate microscopic particles such as cells or sub-cellular structures. The Force Measurement (FM) module expands
Effect of Cytochalasin on Average Pseudopodia Length in Amoeba Proteus
 Effect of Cytochalasin on Average Pseudopodia Length in Amoeba Proteus Alex Waciega Independent Research Project Report Cell Biology 219 December 3, 2008 Introduction In our experiment we studied the effects
Effect of Cytochalasin on Average Pseudopodia Length in Amoeba Proteus Alex Waciega Independent Research Project Report Cell Biology 219 December 3, 2008 Introduction In our experiment we studied the effects
SDS-PAGE and Western Blot. Molecular Basis of Evolution
 1 SDS-PAGE and Western Blot Molecular Basis of Evolution Homology high level of DNA and protein sequence similarity due to common ancestry. Evidence Genomes of related organisms are very similar. Even
1 SDS-PAGE and Western Blot Molecular Basis of Evolution Homology high level of DNA and protein sequence similarity due to common ancestry. Evidence Genomes of related organisms are very similar. Even
20.320, notes for 9/18
 20.320 Notes Page 1 20.320, notes for 9/18 Tuesday, September 18, 2012 9:37 AM Last time We covered the graph of RU vs. time during an SPR experiment. We see a different graph from the association and
20.320 Notes Page 1 20.320, notes for 9/18 Tuesday, September 18, 2012 9:37 AM Last time We covered the graph of RU vs. time during an SPR experiment. We see a different graph from the association and
Performance of a Label-Free Image-Based 2D Scratch Wound Healing Assay to monitor Cell Migration and its Inhibition
 A p p l i c a t i o n N o t e Performance of a Label-Free Image-Based 2D Scratch Wound Healing Assay to monitor Cell Migration and its Inhibition Leonie Rieger and Brad Larson, Applications Department,
A p p l i c a t i o n N o t e Performance of a Label-Free Image-Based 2D Scratch Wound Healing Assay to monitor Cell Migration and its Inhibition Leonie Rieger and Brad Larson, Applications Department,
MEMS based sensors for cellular studies
 MEMS based sensors for cellular studies Taher Saif Mechanical Science and Engineering University of Illinois at Urbana-Champaign Part of GEM4 Summer School lectures on instruments for cell mechanics studies
MEMS based sensors for cellular studies Taher Saif Mechanical Science and Engineering University of Illinois at Urbana-Champaign Part of GEM4 Summer School lectures on instruments for cell mechanics studies
Title. Adhesive lamination: new technology in Solvent Less lamination.
 AIMCAL CONFERENCE 2016, Memphis, TN. Title. Adhesive lamination: new technology in Solvent Less lamination. Giancarlo Caimmi, Nordmeccanica NA Ltd. Short abstracts: A new development in solvent less lamination
AIMCAL CONFERENCE 2016, Memphis, TN. Title. Adhesive lamination: new technology in Solvent Less lamination. Giancarlo Caimmi, Nordmeccanica NA Ltd. Short abstracts: A new development in solvent less lamination
INVESTIGATION OF MSC DIFFERENTIATION ON ELECTROSPUN NANOFIBROUS SCAFFOLDS
 With support of NSF Award no. EEC-0754741 INVESTIGATION OF MSC DIFFERENTIATION ON ELECTROSPUN NANOFIBROUS SCAFFOLDS NSF Summer Undergraduate Fellowship in Sensor Technologies Emily Wible (Bioengineering)
With support of NSF Award no. EEC-0754741 INVESTIGATION OF MSC DIFFERENTIATION ON ELECTROSPUN NANOFIBROUS SCAFFOLDS NSF Summer Undergraduate Fellowship in Sensor Technologies Emily Wible (Bioengineering)
Supplemental material JCB. Pitaval et al., Cell shape and ciliogenesis Pitaval et al.
 Supplemental material THE JOURNAL OF CELL BIOLOGY Pitaval et al., http://www.jcb.org/cgi/content/full/jcb.201004003/dc1 S1 Figure S1. Cell cycle exit analysis. (A) Experimental procedure. RPE1 cells were
Supplemental material THE JOURNAL OF CELL BIOLOGY Pitaval et al., http://www.jcb.org/cgi/content/full/jcb.201004003/dc1 S1 Figure S1. Cell cycle exit analysis. (A) Experimental procedure. RPE1 cells were
Engage the student to formulate a Researchable Question (RQ) by making observations. principle or concept. Write it down.
 Science Lead Teacher Institute Summer 2010 Connecting Biology to Chemistry through Inquiry-based Learning General Approach Using the Scientific Method to develop better conceptual knowledge and reasoning
Science Lead Teacher Institute Summer 2010 Connecting Biology to Chemistry through Inquiry-based Learning General Approach Using the Scientific Method to develop better conceptual knowledge and reasoning
Supporting Information
 Supporting Information Signal mingle: Micropatterns of BMP-2 and fibronectin on soft biopolymeric films regulate myoblast shape and SMAD signaling. Vincent Fitzpatrick, Laure Fourel, Olivier Destaing,
Supporting Information Signal mingle: Micropatterns of BMP-2 and fibronectin on soft biopolymeric films regulate myoblast shape and SMAD signaling. Vincent Fitzpatrick, Laure Fourel, Olivier Destaing,
A New Cellular and Molecular Engineering Curriculum at Rice University
 Session A New Cellular and Molecular Engineering Curriculum at Rice University Ka-Yiu San, Larry V. McIntire, Ann Saterbak Department of Bioengineering, Rice University Houston, Texas 77005 Abstract The
Session A New Cellular and Molecular Engineering Curriculum at Rice University Ka-Yiu San, Larry V. McIntire, Ann Saterbak Department of Bioengineering, Rice University Houston, Texas 77005 Abstract The
Chapter 3. DNA Replication & The Cell Cycle
 Chapter 3 DNA Replication & The Cell Cycle DNA Replication and the Cell Cycle Before cells divide, they must duplicate their DNA // the genetic material DNA is organized into strands called chromosomes
Chapter 3 DNA Replication & The Cell Cycle DNA Replication and the Cell Cycle Before cells divide, they must duplicate their DNA // the genetic material DNA is organized into strands called chromosomes
Fluorescence Imaging with One Nanometer Accuracy Lab
 I. Introduction. Fluorescence Imaging with One Nanometer Accuracy Lab Traditional light microscope is limited by the diffraction limit of light, typically around 250 nm. However, many biological processes
I. Introduction. Fluorescence Imaging with One Nanometer Accuracy Lab Traditional light microscope is limited by the diffraction limit of light, typically around 250 nm. However, many biological processes
green B 1 ) into a single unit to model the substrate in this reaction. enzyme
 Teacher Key Objectives You will use the model pieces in the kit to: Simulate enzymatic actions. Explain enzymatic specificity. Investigate two types of enzyme inhibitors used in regulating enzymatic activity.
Teacher Key Objectives You will use the model pieces in the kit to: Simulate enzymatic actions. Explain enzymatic specificity. Investigate two types of enzyme inhibitors used in regulating enzymatic activity.
Thermal Management of LEDs: Looking Beyond Thermal Conductivity Values
 Thermal Management of LEDs: Looking Beyond Thermal Conductivity Values Specifically designed and formulated chemical products are widely used in the electronics industry for a vast array of applications.
Thermal Management of LEDs: Looking Beyond Thermal Conductivity Values Specifically designed and formulated chemical products are widely used in the electronics industry for a vast array of applications.
Active biomaterials/scaffolds for stem cell-based soft tissue engineering (in a nutshell )
 Active biomaterials/scaffolds for stem cell-based soft tissue engineering (in a nutshell ) Emanuele Giordano Responsabile Lab ICM BioEngLab DEI CIRI SdV-TS Università di Bologna emanuele.giordano@unibo.it
Active biomaterials/scaffolds for stem cell-based soft tissue engineering (in a nutshell ) Emanuele Giordano Responsabile Lab ICM BioEngLab DEI CIRI SdV-TS Università di Bologna emanuele.giordano@unibo.it
CELL BIOLOGY - CLUTCH CH CYTOSKELETON AND CELL MOVEMENT.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF THE CYTOSKELETON The cytoskeleton is an intricate of protein filaments that extend throughout the cytoplasm It is a highly organized and dynamic structure (constantly
!! www.clutchprep.com CONCEPT: OVERVIEW OF THE CYTOSKELETON The cytoskeleton is an intricate of protein filaments that extend throughout the cytoplasm It is a highly organized and dynamic structure (constantly
Design Principles of Length Control of Cytoskeletal Structures
 Design Principles of Length Control of Cytoskeletal Structures Lishibanya Mohapatra 1, Bruce L. Goode 2, Predrag Jelenkovic 3, Rob Phillips 4, Jane Kondev 5,* 1 Department of Physics, Brandeis University,
Design Principles of Length Control of Cytoskeletal Structures Lishibanya Mohapatra 1, Bruce L. Goode 2, Predrag Jelenkovic 3, Rob Phillips 4, Jane Kondev 5,* 1 Department of Physics, Brandeis University,
