Human Gene Regulation
|
|
|
- Belinda Preston
- 6 years ago
- Views:
Transcription
1 Human Gene Regulation 4
2 Transcriptional networks underlying development ~20,000 genes take up 2% of the human genome. 1,000,000 gene regulation regions (promoters, enhancers, silencers, insulators) controlling our genes take up >10%. 99% of gene regulation regions were never seen before the human genome was sequenced. We study these switches. B Gene H H Gene N Gene N B Gene
3 Why study gene regulation? Large, hitherto invisible, disease susceptibility layer Encodes causality in gene regulatory networks Key contributions to genome and phenotype evolution Great promise for human health
4 How to Understand Genomic Datasets? Genomic Regions Enrichment of Annotations Tool job submissions per day, from 7,000 IP addrs >100,000 jobs served, >100 citations
5 Extend From Analysis to Predictions SRF T cell Term ChIP-seq PRISM actin cytoskeleton structural constituent of muscle dilated heart ventricles regulation of insulin secretion Discovers novel functions for hundreds of transcription factors. Known functions supported by hundreds of novel binding sites.
6 interactions appear to be direct and which additional transcriptio work. cesses. Comparing We predict these in data our to preliminary Combine factors may Genetics participate in the with network. Genomics: We predict in our prelim E14.5 network that 4 of Predicted the 5 interactions genomic network. from Black: our genetic netw suggests direct genetic interaction From are direct, Description via novel enhancers. (see to background Understanding We figure), predict red: novel. the 5th genetic inter (Fezf2 repression of Tbr1) is mediated by Satb2. We also predic Tbr1 and Satb2 autoregulation. Lastly, we predict a role for additional interactions, including Tbr1 and Satb2 autoregulation corticothalamic temporally l neocortex and using spatially specific eractions from our genetic network t E14.5 and E16.5 by The genetic network of sets, e predict I will comprehensively the 5th genetic interaction projection neuron fate ill iated then by apply Satb2. the We also specification predict we will extend ites in FACS sorted using functional and d E18.5 by which Sox5 computational genomics. ulations provides a detailed view of all l neocortex Tbr1 measurements Fezf2 Satb2 helps separate e to broader cortical processes. + = t enhancer networks Ctip2 that control Bhlhe22 ic neocortex. genes and enhancers, we predict corticothalamic the enhancers with subcortical callosal d associate is Fig 3. The genetic network of ription projection factors, Classical neuron fate known specification interactions Functional and & T studied by the McConnell lab. Known cers s that Genetics mediate known & genetic + interin projection and preliminary neuron predicted interactions fate specification. direct genetic interactions in black Computational = m get. from our Development data in red. Genomics uron fate specification. nections between transcription factors, on enhancer assays test activity and sgenics test enhancer layer specificity. effect of critical projection neuron Tbr1 Sox5 3 Fezf2 Ctip2 Bhlhe22 subcortical Satb2 callosal Fig 3. The genetic network of projection Rapid neuron Discovery, fate specification studied by the McConnell lab. Known direct Design genetic interactions Principles black and preliminary predicted interactions from our data in red.
7 Gene regulation of tissue development: Measure, Predict, Discover Work with Sue McConnell
8 Analyze Personal Human Genomes Associate to nearest gene(s) Personal cis variant 1 Target gene Self reported medical summary Any relationship? 3 2 Is the group of affected target genes enriched for a particular function or phenotype?
9 Test human disease mutations Section 1 Enhancer / HuC Target gene Work with Philippe Mourrain
10 Identifying regulatory elements uniquely lost in human Work with David Kingsley 17
11 The first human enhancers conserved to protostomes Work with Nadav Ahituv
12 Correlate independent trait loss with gene/enhancer loss trait matching genomic loci [Hiller et al., 2012a,b]
3/15/2016. Genome = Genes + Gene Regulation. Personal Transcription Factor. Binding Site Mutations Point to. Personal Medical Histories
 Personal Transcription Factor GGTGCCAGGGAAAGGGCAGGAGGTGAGTGCTGGGAGGCAGCTGAGGTCAACTTCTTTTGAACTTCCACGTGGTATTTACTCAGAGCAATTGGTGCCAGAG GCTCAGGGCCCTGGAGTATAAAGCAGAATGTCTGCTCTCTGTGCCCAGACGTGAGCAGGTGAGCAGCTGGGGCGAAAGACCTGTTGGAGGCTATGAATGC
Personal Transcription Factor GGTGCCAGGGAAAGGGCAGGAGGTGAGTGCTGGGAGGCAGCTGAGGTCAACTTCTTTTGAACTTCCACGTGGTATTTACTCAGAGCAATTGGTGCCAGAG GCTCAGGGCCCTGGAGTATAAAGCAGAATGTCTGCTCTCTGTGCCCAGACGTGAGCAGGTGAGCAGCTGGGGCGAAAGACCTGTTGGAGGCTATGAATGC
SNPs - GWAS - eqtls. Sebastian Schmeier
 SNPs - GWAS - eqtls s.schmeier@gmail.com http://sschmeier.github.io/bioinf-workshop/ 17.08.2015 Overview Single nucleotide polymorphism (refresh) SNPs effect on genes (refresh) Genome-wide association
SNPs - GWAS - eqtls s.schmeier@gmail.com http://sschmeier.github.io/bioinf-workshop/ 17.08.2015 Overview Single nucleotide polymorphism (refresh) SNPs effect on genes (refresh) Genome-wide association
NGS Approaches to Epigenomics
 I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
What Can the Epigenome Teach Us About Cellular States and Diseases?
 What Can the Epigenome Teach Us About Cellular States and Diseases? (a computer scientist s view) Luca Pinello Outline Epigenetic: the code over the code What can we learn from epigenomic data? Resources
What Can the Epigenome Teach Us About Cellular States and Diseases? (a computer scientist s view) Luca Pinello Outline Epigenetic: the code over the code What can we learn from epigenomic data? Resources
A Genetic Screen to Identify Mammalian Chromatin Modifiers In Vivo.
 Gus Frangou, Stephanie Palmer & Mark Groudine Fred Hutchinson Cancer Research Center, Seattle- USA A Genetic Screen to Identify Mammalian Chromatin Modifiers In Vivo. During mammalian development and differentiation
Gus Frangou, Stephanie Palmer & Mark Groudine Fred Hutchinson Cancer Research Center, Seattle- USA A Genetic Screen to Identify Mammalian Chromatin Modifiers In Vivo. During mammalian development and differentiation
Title: Genome-Wide Predictions of Transcription Factor Binding Events using Multi- Dimensional Genomic and Epigenomic Features Background
 Title: Genome-Wide Predictions of Transcription Factor Binding Events using Multi- Dimensional Genomic and Epigenomic Features Team members: David Moskowitz and Emily Tsang Background Transcription factors
Title: Genome-Wide Predictions of Transcription Factor Binding Events using Multi- Dimensional Genomic and Epigenomic Features Team members: David Moskowitz and Emily Tsang Background Transcription factors
Whole Transcriptome Analysis of Illumina RNA- Seq Data. Ryan Peters Field Application Specialist
 Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Practice Exam A. Briefly describe how IL-25 treatment might be able to help this responder subgroup of liver cancer patients.
 Practice Exam 2007 1. A special JAK-STAT signaling system (JAK5-STAT5) was recently identified in which a gene called TS5 becomes selectively transcribed and expressed in the liver upon induction by a
Practice Exam 2007 1. A special JAK-STAT signaling system (JAK5-STAT5) was recently identified in which a gene called TS5 becomes selectively transcribed and expressed in the liver upon induction by a
From Machine Learning to Personalized Medicine
 1101100000 0010101101 Karsten Borgwardt KRUPP SYMPOSIUM From Machine Learning to Personalized Medicine Max Planck Institute of Psychiatry Munich, Germany October 21, 2016 0010111011 The non-profit Alfried
1101100000 0010101101 Karsten Borgwardt KRUPP SYMPOSIUM From Machine Learning to Personalized Medicine Max Planck Institute of Psychiatry Munich, Germany October 21, 2016 0010111011 The non-profit Alfried
Measuring Protein-DNA interactions
 Measuring Protein-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Transcription Factors are genetic switches 3 Regulation of Gene Expression by Transcription
Measuring Protein-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Transcription Factors are genetic switches 3 Regulation of Gene Expression by Transcription
Using GREAT.stanford.edu to interpret cis-regulatory rich datasets including ChIP-Seq, Epi markers, GWAS etc.
 Using GREAT.stanford.edu to interpret cis-regulatory rich datasets including ChIP-Seq, Epi markers, GWAS etc. Gill Bejerano Dept. of Developmental Biology & Dept. of Computer Science Stanford University
Using GREAT.stanford.edu to interpret cis-regulatory rich datasets including ChIP-Seq, Epi markers, GWAS etc. Gill Bejerano Dept. of Developmental Biology & Dept. of Computer Science Stanford University
Genetics Journal Club YoSon Park Feb 11, Announcement: Marco Trizzino s seminar March 25 th, 2016!
 Genetics Journal Club YoSon Park Feb 11, 2016 Announcement: Marco Trizzino s seminar March 25 th, 2016! Craniofacial development Implications in brain development, bipedal postures, etc Most craniofacial
Genetics Journal Club YoSon Park Feb 11, 2016 Announcement: Marco Trizzino s seminar March 25 th, 2016! Craniofacial development Implications in brain development, bipedal postures, etc Most craniofacial
Trilateral Project WM4 Comparative studies in new technologies (biotechnology, business methods, etc.)
 Trilateral Project WM4 Comparative studies in new technologies (biotechnology, business methods, etc.) Report on comparative study on Examination Practice Relating to Single Nucleotide Polymorphisms (SNPs)
Trilateral Project WM4 Comparative studies in new technologies (biotechnology, business methods, etc.) Report on comparative study on Examination Practice Relating to Single Nucleotide Polymorphisms (SNPs)
: Genomic Regions Enrichment of Annotations Tool
 http://great.stanford.edu/ : Genomic Regions Enrichment of Annotations Tool Gill Bejerano Dept. of Developmental Biology & Dept. of Computer Science Stanford University 1 Human Gene Regulation 10 13 different
http://great.stanford.edu/ : Genomic Regions Enrichment of Annotations Tool Gill Bejerano Dept. of Developmental Biology & Dept. of Computer Science Stanford University 1 Human Gene Regulation 10 13 different
Personal and population genomics of human regulatory variation
 Personal and population genomics of human regulatory variation Benjamin Vernot,, and Joshua M. Akey Department of Genome Sciences, University of Washington, Seattle, Washington 98195, USA Ting WANG 1 Brief
Personal and population genomics of human regulatory variation Benjamin Vernot,, and Joshua M. Akey Department of Genome Sciences, University of Washington, Seattle, Washington 98195, USA Ting WANG 1 Brief
ChIP-seq analysis 2/28/2018
 ChIP-seq analysis 2/28/2018 Acknowledgements Much of the content of this lecture is from: Furey (2012) ChIP-seq and beyond Park (2009) ChIP-seq advantages + challenges Landt et al. (2012) ChIP-seq guidelines
ChIP-seq analysis 2/28/2018 Acknowledgements Much of the content of this lecture is from: Furey (2012) ChIP-seq and beyond Park (2009) ChIP-seq advantages + challenges Landt et al. (2012) ChIP-seq guidelines
BTRY 7210: Topics in Quantitative Genomics and Genetics
 BTRY 7210: Topics in Quantitative Genomics and Genetics Jason Mezey Biological Statistics and Computational Biology (BSCB) Department of Genetic Medicine jgm45@cornell.edu Spring 2015, Thurs.,12:20-1:10
BTRY 7210: Topics in Quantitative Genomics and Genetics Jason Mezey Biological Statistics and Computational Biology (BSCB) Department of Genetic Medicine jgm45@cornell.edu Spring 2015, Thurs.,12:20-1:10
Lecture 14. ENCODE. Michael Schatz. March 15, 2017 JHU : Applied Comparative Genomics
 Lecture 14. ENCODE Michael Schatz March 15, 2017 JHU 601.749: Applied Comparative Genomics Project Proposal! Due March 15 HW6: Due March 29 *-seq in 4 short vignettes RNA-seq Methyl-seq ChIP-seq Hi-C Chromatin
Lecture 14. ENCODE Michael Schatz March 15, 2017 JHU 601.749: Applied Comparative Genomics Project Proposal! Due March 15 HW6: Due March 29 *-seq in 4 short vignettes RNA-seq Methyl-seq ChIP-seq Hi-C Chromatin
DNA. bioinformatics. epigenetics methylation structural variation. custom. assembly. gene. tumor-normal. mendelian. BS-seq. prediction.
 Epigenomics T TM activation SNP target ncrna validation metagenomics genetics private RRBS-seq de novo trio RIP-seq exome mendelian comparative genomics DNA NGS ChIP-seq bioinformatics assembly tumor-normal
Epigenomics T TM activation SNP target ncrna validation metagenomics genetics private RRBS-seq de novo trio RIP-seq exome mendelian comparative genomics DNA NGS ChIP-seq bioinformatics assembly tumor-normal
Reviewers' Comments: Reviewer #1 (Remarks to the Author)
 Reviewers' Comments: Reviewer #1 (Remarks to the Author) In this study, Rosenbluh et al reported direct comparison of two screening approaches: one is genome editing-based method using CRISPR-Cas9 (cutting,
Reviewers' Comments: Reviewer #1 (Remarks to the Author) In this study, Rosenbluh et al reported direct comparison of two screening approaches: one is genome editing-based method using CRISPR-Cas9 (cutting,
latestdevelopments relevant for the Ag sector André Eggen Agriculture Segment Manager, Europe
 Overviewof Illumina s latestdevelopments relevant for the Ag sector André Eggen Agriculture Segment Manager, Europe Seminar der Studienrichtung Tierwissenschaften, TÜM, July 1, 2009 Overviewof Illumina
Overviewof Illumina s latestdevelopments relevant for the Ag sector André Eggen Agriculture Segment Manager, Europe Seminar der Studienrichtung Tierwissenschaften, TÜM, July 1, 2009 Overviewof Illumina
Computational Workflows for Genome-Wide Association Study: I
 Computational Workflows for Genome-Wide Association Study: I Department of Computer Science Brown University, Providence sorin@cs.brown.edu October 16, 2014 Outline 1 Outline 2 3 Monogenic Mendelian Diseases
Computational Workflows for Genome-Wide Association Study: I Department of Computer Science Brown University, Providence sorin@cs.brown.edu October 16, 2014 Outline 1 Outline 2 3 Monogenic Mendelian Diseases
RP1: AN EXAMPLE OF REVERSE GENETICS APPROACH TO DESCRIBE COMMON RECESSIVE DEFECTS
 RP1: AN EXAMPLE OF REVERSE GENETICS APPROACH TO DESCRIBE COMMON RECESSIVE DEFECTS C. Grohs GABI, INRA, AgroParisTech, Université Paris Saclay 28 / 08 / 2017 Outbreaks of recessive defects as a consequence
RP1: AN EXAMPLE OF REVERSE GENETICS APPROACH TO DESCRIBE COMMON RECESSIVE DEFECTS C. Grohs GABI, INRA, AgroParisTech, Université Paris Saclay 28 / 08 / 2017 Outbreaks of recessive defects as a consequence
Non-coding Function & Variation, MPRAs. Mike White Bio5488 3/5/18
 Non-coding Function & Variation, MPRAs Mike White Bio5488 3/5/18 Outline MONDAY Non-coding function and variation The barcode Basic versions of MRPA technology WEDNESDAY More varieties of MRPAs Some key
Non-coding Function & Variation, MPRAs Mike White Bio5488 3/5/18 Outline MONDAY Non-coding function and variation The barcode Basic versions of MRPA technology WEDNESDAY More varieties of MRPAs Some key
SUPPLEMENTARY INFORMATION
 18Pura Brain Liver 2 kb 9Mtmr Heart Brain 1.4 kb 15Ppara 1.5 kb IPTpcc 1.3 kb GAPDH 1.5 kb IPGpr1 2 kb GAPDH 1.5 kb 17Tcte 13Elov12 Lung Thymus 6 kb 5 kb 17Tbcc Kidney Embryo 4.5 kb 3.7 kb GAPDH 1.5 kb
18Pura Brain Liver 2 kb 9Mtmr Heart Brain 1.4 kb 15Ppara 1.5 kb IPTpcc 1.3 kb GAPDH 1.5 kb IPGpr1 2 kb GAPDH 1.5 kb 17Tcte 13Elov12 Lung Thymus 6 kb 5 kb 17Tbcc Kidney Embryo 4.5 kb 3.7 kb GAPDH 1.5 kb
Pharmacogenetics: A SNPshot of the Future. Ani Khondkaryan Genomics, Bioinformatics, and Medicine Spring 2001
 Pharmacogenetics: A SNPshot of the Future Ani Khondkaryan Genomics, Bioinformatics, and Medicine Spring 2001 1 I. What is pharmacogenetics? It is the study of how genetic variation affects drug response
Pharmacogenetics: A SNPshot of the Future Ani Khondkaryan Genomics, Bioinformatics, and Medicine Spring 2001 1 I. What is pharmacogenetics? It is the study of how genetic variation affects drug response
FY10 INVITATION PROPOSAL SUMMARY PAGE
 FY10 INVITATION PROPOSAL SUMMARY PAGE Project Title: Pilot Sequence: High Density DNA Sequencing to Detect Population Structure of Pacific Herring (Project Number: 10100165) Project Period: October 1,
FY10 INVITATION PROPOSAL SUMMARY PAGE Project Title: Pilot Sequence: High Density DNA Sequencing to Detect Population Structure of Pacific Herring (Project Number: 10100165) Project Period: October 1,
Ecological genomics and molecular adaptation: state of the Union and some research goals for the near future.
 Ecological genomics and molecular adaptation: state of the Union and some research goals for the near future. Louis Bernatchez Genomics and Conservation of Aquatic Resources Université LAVAL! Molecular
Ecological genomics and molecular adaptation: state of the Union and some research goals for the near future. Louis Bernatchez Genomics and Conservation of Aquatic Resources Université LAVAL! Molecular
9. What proteins will be affected by mutations in the trans-acting elements? Cis-acting elements?
 6. What regulates the expression of a gene? 7. What are the cis- and trans-acting elements? 8. Can a deficiency in a trans-acting element be overcome by the addition of another copy of the gene to a cell?
6. What regulates the expression of a gene? 7. What are the cis- and trans-acting elements? 8. Can a deficiency in a trans-acting element be overcome by the addition of another copy of the gene to a cell?
Linking Genetic Variation to Important Phenotypes: SNPs, CNVs, GWAS, and eqtls
 Linking Genetic Variation to Important Phenotypes: SNPs, CNVs, GWAS, and eqtls BMI/CS 776 www.biostat.wisc.edu/bmi776/ Colin Dewey cdewey@biostat.wisc.edu Spring 2012 1. Understanding Human Genetic Variation
Linking Genetic Variation to Important Phenotypes: SNPs, CNVs, GWAS, and eqtls BMI/CS 776 www.biostat.wisc.edu/bmi776/ Colin Dewey cdewey@biostat.wisc.edu Spring 2012 1. Understanding Human Genetic Variation
The ENCODE Encyclopedia. & Variant Annotation Using RegulomeDB and HaploReg
 The ENCODE Encyclopedia & Variant Annotation Using RegulomeDB and HaploReg Jill E. Moore Weng Lab University of Massachusetts Medical School October 10, 2015 Where s the Encyclopedia? ENCODE: Encyclopedia
The ENCODE Encyclopedia & Variant Annotation Using RegulomeDB and HaploReg Jill E. Moore Weng Lab University of Massachusetts Medical School October 10, 2015 Where s the Encyclopedia? ENCODE: Encyclopedia
Crash-course in genomics
 Crash-course in genomics Molecular biology : How does the genome code for function? Genetics: How is the genome passed on from parent to child? Genetic variation: How does the genome change when it is
Crash-course in genomics Molecular biology : How does the genome code for function? Genetics: How is the genome passed on from parent to child? Genetic variation: How does the genome change when it is
Nature Genetics: doi: /ng.3556 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE. Supplementary Figure 1
 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
resequencing storage SNP ncrna metagenomics private trio de novo exome ncrna RNA DNA bioinformatics RNA-seq comparative genomics
 RNA Sequencing T TM variation genetics validation SNP ncrna metagenomics private trio de novo exome mendelian ChIP-seq RNA DNA bioinformatics custom target high-throughput resequencing storage ncrna comparative
RNA Sequencing T TM variation genetics validation SNP ncrna metagenomics private trio de novo exome mendelian ChIP-seq RNA DNA bioinformatics custom target high-throughput resequencing storage ncrna comparative
BTRY 7210: Topics in Quantitative Genomics and Genetics
 BTRY 7210: Topics in Quantitative Genomics and Genetics Jason Mezey Biological Statistics and Computational Biology (BSCB) Department of Genetic Medicine jgm45@cornell.edu January 29, 2015 Why you re here
BTRY 7210: Topics in Quantitative Genomics and Genetics Jason Mezey Biological Statistics and Computational Biology (BSCB) Department of Genetic Medicine jgm45@cornell.edu January 29, 2015 Why you re here
elin grundberg JULY 2018 JK - PRESENTATION- 11 MINS [MIXED RESPONDENTS] [Other comments:]
![elin grundberg JULY 2018 JK - PRESENTATION- 11 MINS [MIXED RESPONDENTS] [Other comments:] elin grundberg JULY 2018 JK - PRESENTATION- 11 MINS [MIXED RESPONDENTS] [Other comments:]](/thumbs/82/86384595.jpg) elin grundberg JULY 2018 JK - PRESENTATION- 11 MINS [MIXED RESPONDENTS] [Other comments:] So, we're going to be talking about epigenomic research, so, Elin, are you around? Come over. You can introduce
elin grundberg JULY 2018 JK - PRESENTATION- 11 MINS [MIXED RESPONDENTS] [Other comments:] So, we're going to be talking about epigenomic research, so, Elin, are you around? Come over. You can introduce
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Flowchart illustrating the three complementary strategies for gene prioritization used in this study.. Supplementary Figure 2 Flow diagram illustrating calcaneal quantitative ultrasound
Supplementary Figure 1 Flowchart illustrating the three complementary strategies for gene prioritization used in this study.. Supplementary Figure 2 Flow diagram illustrating calcaneal quantitative ultrasound
Introduction to Bioinformatics and Gene Expression Technologies
 Introduction to Bioinformatics and Gene Expression Technologies Utah State University Fall 2017 Statistical Bioinformatics (Biomedical Big Data) Notes 1 1 Vocabulary Gene: hereditary DNA sequence at a
Introduction to Bioinformatics and Gene Expression Technologies Utah State University Fall 2017 Statistical Bioinformatics (Biomedical Big Data) Notes 1 1 Vocabulary Gene: hereditary DNA sequence at a
Introduction to Bioinformatics and Gene Expression Technologies
 Vocabulary Introduction to Bioinformatics and Gene Expression Technologies Utah State University Fall 2017 Statistical Bioinformatics (Biomedical Big Data) Notes 1 Gene: Genetics: Genome: Genomics: hereditary
Vocabulary Introduction to Bioinformatics and Gene Expression Technologies Utah State University Fall 2017 Statistical Bioinformatics (Biomedical Big Data) Notes 1 Gene: Genetics: Genome: Genomics: hereditary
Genetics Lecture Notes Lectures 17 19
 Genetics Lecture Notes 7.03 2005 Lectures 17 19 Lecture 17 Gene Regulation We are now going to look at ways that genetics can be used to study gene regulation. The issue is how cells adjust the expression
Genetics Lecture Notes 7.03 2005 Lectures 17 19 Lecture 17 Gene Regulation We are now going to look at ways that genetics can be used to study gene regulation. The issue is how cells adjust the expression
RNA-Seq Now What? BIS180L Professor Maloof May 24, 2018
 RNA-Seq Now What? BIS180L Professor Maloof May 24, 2018 We have differentially expressed genes, what do we want to know about them? We have differentially expressed genes, what do we want to know about
RNA-Seq Now What? BIS180L Professor Maloof May 24, 2018 We have differentially expressed genes, what do we want to know about them? We have differentially expressed genes, what do we want to know about
File S1. Program overview and features
 File S1 Program overview and features Query list filtering. Further filtering may be applied through user selected query lists (Figure. 2B, Table S3) that restrict the results and/or report specifically
File S1 Program overview and features Query list filtering. Further filtering may be applied through user selected query lists (Figure. 2B, Table S3) that restrict the results and/or report specifically
Computational Systems Biology Deep Learning in the Life Sciences
 Computational Systems Biology Deep Learning in the Life Sciences 6.802 6.874 20.390 20.490 HST.506 Christina Ji April 6, 2017 DanQ: a hybrid convolutional and recurrent deep neural network for quantifying
Computational Systems Biology Deep Learning in the Life Sciences 6.802 6.874 20.390 20.490 HST.506 Christina Ji April 6, 2017 DanQ: a hybrid convolutional and recurrent deep neural network for quantifying
Huijuan Feng, Shining Ma,Chao Ye & Zhixing Feng
 Huijuan Feng, Shining Ma,Chao Ye & Zhixing Feng Background-Author introduction Research interest: Methods for gene mapping of complex traits Inference of population structure from genetic data Genome variation
Huijuan Feng, Shining Ma,Chao Ye & Zhixing Feng Background-Author introduction Research interest: Methods for gene mapping of complex traits Inference of population structure from genetic data Genome variation
Deep learning sequence-based ab initio prediction of variant effects on expression and disease risk
 Summer Review 7 Deep learning sequence-based ab initio prediction of variant effects on expression and disease risk Jian Zhou 1,2,3, Chandra L. Theesfeld 1, Kevin Yao 3, Kathleen M. Chen 3, Aaron K. Wong
Summer Review 7 Deep learning sequence-based ab initio prediction of variant effects on expression and disease risk Jian Zhou 1,2,3, Chandra L. Theesfeld 1, Kevin Yao 3, Kathleen M. Chen 3, Aaron K. Wong
Association Mapping in Plants PLSC 731 Plant Molecular Genetics Phil McClean April, 2010
 Association Mapping in Plants PLSC 731 Plant Molecular Genetics Phil McClean April, 2010 Traditional QTL approach Uses standard bi-parental mapping populations o F2 or RI These have a limited number of
Association Mapping in Plants PLSC 731 Plant Molecular Genetics Phil McClean April, 2010 Traditional QTL approach Uses standard bi-parental mapping populations o F2 or RI These have a limited number of
Non-coding Function & Variation, MPRAs II. Mike White Bio /5/18
 Non-coding Function & Variation, MPRAs II Mike White Bio 5488 3/5/18 MPRA Review Problem 1: Where does your CRE DNA come from? DNA synthesis Genomic fragments Targeted regulome capture Problem 2: How do
Non-coding Function & Variation, MPRAs II Mike White Bio 5488 3/5/18 MPRA Review Problem 1: Where does your CRE DNA come from? DNA synthesis Genomic fragments Targeted regulome capture Problem 2: How do
2/19/13. Contents. Applications of HMMs in Epigenomics
 2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
Microarray Informatics
 Microarray Informatics Donald Dunbar MSc Seminar 31 st January 2007 Aims To give a biologist s view of microarray experiments To explain the technologies involved To describe typical microarray experiments
Microarray Informatics Donald Dunbar MSc Seminar 31 st January 2007 Aims To give a biologist s view of microarray experiments To explain the technologies involved To describe typical microarray experiments
Applications of HMMs in Epigenomics
 I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
Traditional Genetic Improvement. Genetic variation is due to differences in DNA sequence. Adding DNA sequence data to traditional breeding.
 1 Introduction What is Genomic selection and how does it work? How can we best use DNA data in the selection of cattle? Mike Goddard 5/1/9 University of Melbourne and Victorian DPI of genomic selection
1 Introduction What is Genomic selection and how does it work? How can we best use DNA data in the selection of cattle? Mike Goddard 5/1/9 University of Melbourne and Victorian DPI of genomic selection
11/22/13. Proteomics, functional genomics, and systems biology. Biosciences 741: Genomics Fall, 2013 Week 11
 Proteomics, functional genomics, and systems biology Biosciences 741: Genomics Fall, 2013 Week 11 1 Figure 6.1 The future of genomics Functional Genomics The field of functional genomics represents the
Proteomics, functional genomics, and systems biology Biosciences 741: Genomics Fall, 2013 Week 11 1 Figure 6.1 The future of genomics Functional Genomics The field of functional genomics represents the
MicroRNAs and Genome Editing Two Evolving Protagonists in Medicine
 MicroRNAs and Genome Editing Two Evolving Protagonists in Medicine Gerald W Dorn II, MD, FACC Needleman Professor and Associate Chair Department of Internal Medicine Washington University in St Louis The
MicroRNAs and Genome Editing Two Evolving Protagonists in Medicine Gerald W Dorn II, MD, FACC Needleman Professor and Associate Chair Department of Internal Medicine Washington University in St Louis The
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Characterization of Hi-C/CHi-C dynamics and enhancer identification. (a) Scatterplot of Hi-C read counts supporting contacts between domain boundaries. Contacts enclosing domains
Supplementary Figure 1 Characterization of Hi-C/CHi-C dynamics and enhancer identification. (a) Scatterplot of Hi-C read counts supporting contacts between domain boundaries. Contacts enclosing domains
Whole genome sequencing in the UK Biobank
 Whole genome sequencing in the UK Biobank Part of the UK Government s Industrial Strategy Challenge Fund (ISCF) for the Data to Early Diagnosis and Precision Medicine initiative Aim to produce deep characterisation
Whole genome sequencing in the UK Biobank Part of the UK Government s Industrial Strategy Challenge Fund (ISCF) for the Data to Early Diagnosis and Precision Medicine initiative Aim to produce deep characterisation
Functional Genomics and Single-Cell Genomics. Christine Queitsch Department of Genome Sciences
 Functional Genomics and Single-Cell Genomics Christine Queitsch Department of Genome Sciences queitsch@uw.edu 1 Outline Challenges Combining methods for better genome assembly Functional genomics let s
Functional Genomics and Single-Cell Genomics Christine Queitsch Department of Genome Sciences queitsch@uw.edu 1 Outline Challenges Combining methods for better genome assembly Functional genomics let s
Comparative eqtl analyses within and between seven tissue types suggest mechanisms underlying cell type specificity of eqtls
 Comparative eqtl analyses within and between seven tissue types suggest mechanisms underlying cell type specificity of eqtls, Duke University Christopher D Brown, University of Pennsylvania November 9th,
Comparative eqtl analyses within and between seven tissue types suggest mechanisms underlying cell type specificity of eqtls, Duke University Christopher D Brown, University of Pennsylvania November 9th,
2/10/17. Contents. Applications of HMMs in Epigenomics
 2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
Problem Set 8. Answer Key
 MCB 102 University of California, Berkeley August 11, 2009 Isabelle Philipp Online Document Problem Set 8 Answer Key 1. The Genetic Code (a) Are all amino acids encoded by the same number of codons? no
MCB 102 University of California, Berkeley August 11, 2009 Isabelle Philipp Online Document Problem Set 8 Answer Key 1. The Genetic Code (a) Are all amino acids encoded by the same number of codons? no
Year III Pharm.D Dr. V. Chitra
 Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
MCB 102 University of California, Berkeley August 11 13, Problem Set 8
 MCB 102 University of California, Berkeley August 11 13, 2009 Isabelle Philipp Handout Problem Set 8 The answer key will be posted by Tuesday August 11. Try to solve the problem sets always first without
MCB 102 University of California, Berkeley August 11 13, 2009 Isabelle Philipp Handout Problem Set 8 The answer key will be posted by Tuesday August 11. Try to solve the problem sets always first without
Using human genetics to make new medicines
 Using human genetics to make new medicines Dr. Maya Ghoussaini Open targets, Wellcome Trust Sanger Institute, UK ELRIG, Drug Discovery, Liverpool, 3 rd -4 th October 2017 Drug development rarely succeeds
Using human genetics to make new medicines Dr. Maya Ghoussaini Open targets, Wellcome Trust Sanger Institute, UK ELRIG, Drug Discovery, Liverpool, 3 rd -4 th October 2017 Drug development rarely succeeds
Bi8 Lecture 19. Review and Practice Questions March 8th 2016
 Bi8 Lecture 19 Review and Practice Questions March 8th 2016 Common Misconceptions Where Words Matter Common Misconceptions DNA vs RNA vs Protein Su(H) vs Su(H) Replication Transcription Transcribed vs
Bi8 Lecture 19 Review and Practice Questions March 8th 2016 Common Misconceptions Where Words Matter Common Misconceptions DNA vs RNA vs Protein Su(H) vs Su(H) Replication Transcription Transcribed vs
What is Bioinformatics?
 What is Bioinformatics? Bioinformatics is the field of science in which biology, computer science, and information technology merge to form a single discipline. - NCBI The ultimate goal of the field is
What is Bioinformatics? Bioinformatics is the field of science in which biology, computer science, and information technology merge to form a single discipline. - NCBI The ultimate goal of the field is
Finding Enrichments of Functional Annotations for Disease- Associated Single-Nucleotide Polymorphisms
 Finding Enrichments of Functional Annotations for Disease- Associated Single-Nucleotide Polymorphisms Steven Homberg 11/10/13 1 Abstract Computational analysis of SNP-disease associations from GWAS as
Finding Enrichments of Functional Annotations for Disease- Associated Single-Nucleotide Polymorphisms Steven Homberg 11/10/13 1 Abstract Computational analysis of SNP-disease associations from GWAS as
res2)=)colocalisation(eqtl_file="icoslg_eqtl.txt",) gwas_file="uc_gwas_icoslg_locus.txt",)p1)=)1e+4,)p2)=)1e+4,)p12)=)1e+5,) region)=)200000)$
 Homework 7 res1)=)colocalisation(eqtl_file="ptk2b_eqtl.txt",) gwas_file="ad_gwas_ptk2b_locus.txt",)p1)=)1e+4,)p2)=)1e+4,)p12)=)1e+5,) region)=)200000)$ ##))1.02e+06))2.43e+05))6.80e+04))1.51e+02))9.84e+01)$
Homework 7 res1)=)colocalisation(eqtl_file="ptk2b_eqtl.txt",) gwas_file="ad_gwas_ptk2b_locus.txt",)p1)=)1e+4,)p2)=)1e+4,)p12)=)1e+5,) region)=)200000)$ ##))1.02e+06))2.43e+05))6.80e+04))1.51e+02))9.84e+01)$
Stefano Monti. Workshop Format
 Gad Getz Stefano Monti Michael Reich {gadgetz,smonti,mreich}@broad.mit.edu http://www.broad.mit.edu/~smonti/aws Broad Institute of MIT & Harvard October 18-20, 2006 Cambridge, MA Workshop Format Morning
Gad Getz Stefano Monti Michael Reich {gadgetz,smonti,mreich}@broad.mit.edu http://www.broad.mit.edu/~smonti/aws Broad Institute of MIT & Harvard October 18-20, 2006 Cambridge, MA Workshop Format Morning
Toward a cohesive understanding of glyphosate resistance evolution in the common morning glory
 Toward a cohesive understanding of glyphosate resistance evolution in the common morning glory Regina Baucom Dept of Ecology and Evolutionary Biology University of Michigan Ann Arbor, MI, USA Treated
Toward a cohesive understanding of glyphosate resistance evolution in the common morning glory Regina Baucom Dept of Ecology and Evolutionary Biology University of Michigan Ann Arbor, MI, USA Treated
Decoding Chromatin States with Epigenome Data Advanced Topics in Computa8onal Genomics
 Decoding Chromatin States with Epigenome Data 02-715 Advanced Topics in Computa8onal Genomics HMMs for Decoding Chromatin States Epigene8c modifica8ons of the genome have been associated with Establishing
Decoding Chromatin States with Epigenome Data 02-715 Advanced Topics in Computa8onal Genomics HMMs for Decoding Chromatin States Epigene8c modifica8ons of the genome have been associated with Establishing
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs Deep transcritpomes for traditional model species from ENCODE (and modencode) Deep RNA-seq and chromatin analysis on 147 human cell types, as well as tissues,
Introduction to genome biology Lisa Stubbs Deep transcritpomes for traditional model species from ENCODE (and modencode) Deep RNA-seq and chromatin analysis on 147 human cell types, as well as tissues,
Supplementary Table 1: Oligo designs. A list of ATAC-seq oligos used for PCR.
 Ad1_noMX: Ad2.1_TAAGGCGA Ad2.2_CGTACTAG Ad2.3_AGGCAGAA Ad2.4_TCCTGAGC Ad2.5_GGACTCCT Ad2.6_TAGGCATG Ad2.7_CTCTCTAC Ad2.8_CAGAGAGG Ad2.9_GCTACGCT Ad2.10_CGAGGCTG Ad2.11_AAGAGGCA Ad2.12_GTAGAGGA Ad2.13_GTCGTGAT
Ad1_noMX: Ad2.1_TAAGGCGA Ad2.2_CGTACTAG Ad2.3_AGGCAGAA Ad2.4_TCCTGAGC Ad2.5_GGACTCCT Ad2.6_TAGGCATG Ad2.7_CTCTCTAC Ad2.8_CAGAGAGG Ad2.9_GCTACGCT Ad2.10_CGAGGCTG Ad2.11_AAGAGGCA Ad2.12_GTAGAGGA Ad2.13_GTCGTGAT
Lecture 18. 2) Check to see whether the mutation is recessive and trans-acting (most will be).
 Lecture 18 In the preceding examples of bacterial gene regulation, we have used known regulatory mechanisms to see how mutations in different elements of the system would behave in dominance tests and
Lecture 18 In the preceding examples of bacterial gene regulation, we have used known regulatory mechanisms to see how mutations in different elements of the system would behave in dominance tests and
ZBED6 the birth of a transcription factor in placental mammals. Wild Boar X domestic pig intercross initiated 1989
 the birth of a transcription factor in placental mammals Wild Boar X domestic pig intercross initiated 1989 X Göran Andersson F1 X F1 Swedish University of Agricultural Sciences ALLBIO- Scilife-UPPNEX-BILS
the birth of a transcription factor in placental mammals Wild Boar X domestic pig intercross initiated 1989 X Göran Andersson F1 X F1 Swedish University of Agricultural Sciences ALLBIO- Scilife-UPPNEX-BILS
Linking Genetic Variation to Important Phenotypes: SNPs, CNVs, GWAS, and eqtls
 Linking Genetic Variation to Important Phenotypes: SNPs, CNVs, GWAS, and eqtls BMI/CS 776 www.biostat.wisc.edu/bmi776/ Mark Craven craven@biostat.wisc.edu Spring 2011 1. Understanding Human Genetic Variation!
Linking Genetic Variation to Important Phenotypes: SNPs, CNVs, GWAS, and eqtls BMI/CS 776 www.biostat.wisc.edu/bmi776/ Mark Craven craven@biostat.wisc.edu Spring 2011 1. Understanding Human Genetic Variation!
a rj13 ORNLf CP CONF-~~/!CI~~--The Functional Genomics Initiative at Oak Ridge National Laboratory*
 I I d ORNLf CP-94806 CONF-~~/!CI~~--The Functional Genomics Initiative at Oak Ridge National Laboratory* Dabney Johnson t, Monica Justice t; Ken Beattie t, Michelle Buchanan $, Michael Ramsey 0,Rose Ramsey
I I d ORNLf CP-94806 CONF-~~/!CI~~--The Functional Genomics Initiative at Oak Ridge National Laboratory* Dabney Johnson t, Monica Justice t; Ken Beattie t, Michelle Buchanan $, Michael Ramsey 0,Rose Ramsey
EPIB 668 Genetic association studies. Aurélie LABBE - Winter 2011
 EPIB 668 Genetic association studies Aurélie LABBE - Winter 2011 1 / 71 OUTLINE Linkage vs association Linkage disequilibrium Case control studies Family-based association 2 / 71 RECAP ON GENETIC VARIANTS
EPIB 668 Genetic association studies Aurélie LABBE - Winter 2011 1 / 71 OUTLINE Linkage vs association Linkage disequilibrium Case control studies Family-based association 2 / 71 RECAP ON GENETIC VARIANTS
The genetic code of gene regulatory elements
 The genetic code of gene regulatory elements Ivan Ovcharenko Computational Biology Branch National Center for Biotechnology Information National Institutes of Health October 23, 2008 Outline Gene deserts
The genetic code of gene regulatory elements Ivan Ovcharenko Computational Biology Branch National Center for Biotechnology Information National Institutes of Health October 23, 2008 Outline Gene deserts
From Genotype to Phenotype
 From Genotype to Phenotype Johanna Vilkki Green technology, Natural Resources Institute Finland Systems biology Genome Transcriptome genes mrna Genotyping methodology SNP TOOLS, WG SEQUENCING Functional
From Genotype to Phenotype Johanna Vilkki Green technology, Natural Resources Institute Finland Systems biology Genome Transcriptome genes mrna Genotyping methodology SNP TOOLS, WG SEQUENCING Functional
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description:
 File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. dcas9-mq1 fusion protein induces de novo
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. dcas9-mq1 fusion protein induces de novo
Can Semantic Web Technologies enable Translational Medicine? (Or Can Translational Medicine help enrich the Semantic Web?)
 Can Semantic Web Technologies enable Translational Medicine? (Or Can Translational Medicine help enrich the Semantic Web?) Vipul Kashyap 1, Tonya Hongsermeier 1, Samuel Aronson 2 1 Clinical Informatics
Can Semantic Web Technologies enable Translational Medicine? (Or Can Translational Medicine help enrich the Semantic Web?) Vipul Kashyap 1, Tonya Hongsermeier 1, Samuel Aronson 2 1 Clinical Informatics
Experimental validation of candidates of tissuespecific and CpG-island-mediated alternative polyadenylation in mouse
 Karin Fleischhanderl; Martina Fondi Experimental validation of candidates of tissuespecific and CpG-island-mediated alternative polyadenylation in mouse 108 - Biotechnologie Abstract --- Keywords: Alternative
Karin Fleischhanderl; Martina Fondi Experimental validation of candidates of tissuespecific and CpG-island-mediated alternative polyadenylation in mouse 108 - Biotechnologie Abstract --- Keywords: Alternative
Nature Genetics: doi: /ng Supplementary Figure 1. H3K27ac HiChIP enriches enhancer promoter-associated chromatin contacts.
 Supplementary Figure 1 H3K27ac HiChIP enriches enhancer promoter-associated chromatin contacts. (a) Schematic of chromatin contacts captured in H3K27ac HiChIP. (b) Loop call overlap for cohesin HiChIP
Supplementary Figure 1 H3K27ac HiChIP enriches enhancer promoter-associated chromatin contacts. (a) Schematic of chromatin contacts captured in H3K27ac HiChIP. (b) Loop call overlap for cohesin HiChIP
This summary statement was issued in 2004 for a PHS 398 application. We will post a more recent sample summary statement soon.
 Posted May 22, 2007. See more samples here: http://www.niaid.nih.gov/ncn/grant/app/default.htm This summary statement was issued in 2004 for a PHS 398 application. We will post a more recent sample summary
Posted May 22, 2007. See more samples here: http://www.niaid.nih.gov/ncn/grant/app/default.htm This summary statement was issued in 2004 for a PHS 398 application. We will post a more recent sample summary
ChIP-Seq Tools. J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015
 ChIP-Seq Tools J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA or
ChIP-Seq Tools J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA or
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
with different microfluidic device parameters. Distribution of number of transcripts (y-axis) detected across nuclei, ranked in decreasing
 Supplementary Figure 1 DroNc-seq benchmarking experiments. (a) Overview of DroNc-seq. (b) CAD schema of DroNc-seq microfluidic device. (c) Comparison of library complexity from experiments with different
Supplementary Figure 1 DroNc-seq benchmarking experiments. (a) Overview of DroNc-seq. (b) CAD schema of DroNc-seq microfluidic device. (c) Comparison of library complexity from experiments with different
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
Review of Transposable elements have rewired the core regulatory network of human embryonic stem cells
 Review of Transposable elements have rewired the core regulatory network of human embryonic stem cells RNA structure Mutagenic screens using transposons Nature Genetics, 42(7), 631-635 (2010). Bradly Alicea
Review of Transposable elements have rewired the core regulatory network of human embryonic stem cells RNA structure Mutagenic screens using transposons Nature Genetics, 42(7), 631-635 (2010). Bradly Alicea
Many transcription factors! recognize DNA shape
 Many transcription factors! recognize DN shape Katie Pollard! Gladstone Institutes USF Division of Biostatistics, Institute for Human Genetics, and Institute for omputational Health Sciences ENODE Users
Many transcription factors! recognize DN shape Katie Pollard! Gladstone Institutes USF Division of Biostatistics, Institute for Human Genetics, and Institute for omputational Health Sciences ENODE Users
Experimental genetics - 2 Partha Roy
 Partha Roy Experimental genetics - 2 Making genetically altered animal 1) Gene knock-out k from: a) the entire animal b) selected cell-type/ tissue c) selected cell-type/tissue at certain time 2) Transgenic
Partha Roy Experimental genetics - 2 Making genetically altered animal 1) Gene knock-out k from: a) the entire animal b) selected cell-type/ tissue c) selected cell-type/tissue at certain time 2) Transgenic
CyVerse Overview. National Academies Special Topics Summer Institute on Quantitative Biology
 Transforming Science Through Data-driven Discovery CyVerse Overview National Academies Special Topics Summer Institute on Quantitative Biology Jason Williams Lead, CyVerse Education, Outreach, Training
Transforming Science Through Data-driven Discovery CyVerse Overview National Academies Special Topics Summer Institute on Quantitative Biology Jason Williams Lead, CyVerse Education, Outreach, Training
 www.illumina.com/hiseq www.illumina.com FOR RESEARCH USE ONLY 2012 2014 Illumina, Inc. All rights reserved. Illumina, BaseSpace, cbot, CSPro, Genetic Energy, HiSeq, Nextera, TruSeq, the pumpkin orange
www.illumina.com/hiseq www.illumina.com FOR RESEARCH USE ONLY 2012 2014 Illumina, Inc. All rights reserved. Illumina, BaseSpace, cbot, CSPro, Genetic Energy, HiSeq, Nextera, TruSeq, the pumpkin orange
Applications of ChIP. November 05, David Grotsky, PhD Scientific Support Specialist - Epigenetics
 Applications of ChIP November 05, 2014 David Grotsky, PhD Scientific Support Specialist - Epigenetics Overview Introduction to chromatin and ChIP The histone code hypothesis What we learn from ChIP-on-chip
Applications of ChIP November 05, 2014 David Grotsky, PhD Scientific Support Specialist - Epigenetics Overview Introduction to chromatin and ChIP The histone code hypothesis What we learn from ChIP-on-chip
Our website:
 Biomedical Informatics Summer Internship Program (BMI SIP) The Department of Biomedical Informatics hosts an annual internship program each summer which provides high school, undergraduate, and graduate
Biomedical Informatics Summer Internship Program (BMI SIP) The Department of Biomedical Informatics hosts an annual internship program each summer which provides high school, undergraduate, and graduate
Stalking the Genetic Basis of a Trait
 OVERVIEW This activity supplements the short film Popped Secret: The Mysterious Origin of Corn. It focuses on the tb1 gene, whose expression is related to phenotypic changes associated with the evolution
OVERVIEW This activity supplements the short film Popped Secret: The Mysterious Origin of Corn. It focuses on the tb1 gene, whose expression is related to phenotypic changes associated with the evolution
GENETICS - CLUTCH CH.15 GENOMES AND GENOMICS.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
!! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
HLA and other tales: The different perspectives of Celiac Disease Gutierrez Achury, Henry Javier
 University of Groningen HLA and other tales: The different perspectives of Celiac Disease Gutierrez Achury, Henry Javier IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's
University of Groningen HLA and other tales: The different perspectives of Celiac Disease Gutierrez Achury, Henry Javier IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's
SUPPLEMENTARY INFORMATION
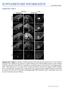 doi:10.1038/nature11094 Supplementary Figures Supplementary Figure 1: Analysis of GFP expression in P0 wild type and mutant ( E1-4) Fezf2-Gfp BAC transgenic mice. Epifluorescent images of sagittal forebrain
doi:10.1038/nature11094 Supplementary Figures Supplementary Figure 1: Analysis of GFP expression in P0 wild type and mutant ( E1-4) Fezf2-Gfp BAC transgenic mice. Epifluorescent images of sagittal forebrain
Transcriptomics. Marta Puig Institut de Biotecnologia i Biomedicina Universitat Autònoma de Barcelona
 Transcriptomics Marta Puig Institut de Biotecnologia i Biomedicina Universitat Autònoma de Barcelona Central dogma of molecular biology Central dogma of molecular biology Genome Complete DNA content of
Transcriptomics Marta Puig Institut de Biotecnologia i Biomedicina Universitat Autònoma de Barcelona Central dogma of molecular biology Central dogma of molecular biology Genome Complete DNA content of
Lecture 2: Height in Plants, Animals, and Humans. Michael Gore lecture notes Tucson Winter Institute version 18 Jan 2013
 Lecture 2: Height in Plants, Animals, and Humans Michael Gore lecture notes Tucson Winter Institute version 18 Jan 2013 Is height a polygenic trait? http://en.wikipedia.org/wiki/gregor_mendel Case Study
Lecture 2: Height in Plants, Animals, and Humans Michael Gore lecture notes Tucson Winter Institute version 18 Jan 2013 Is height a polygenic trait? http://en.wikipedia.org/wiki/gregor_mendel Case Study
