Supplementary Data Bioconductor Vignette for DNAshapeR Package. DNAshapeR: an R/Bioconductor package for DNA shape prediction and feature encoding
|
|
|
- Lindsay Flowers
- 6 years ago
- Views:
Transcription
1 Supplementary Data Bioconductor Vignette for DNAshapeR Package DNAshapeR: an R/Bioconductor package for DNA shape prediction and feature encoding Tsu-Pei Chiu 1,#, Federico Comoglio 2,#, Tianyin Zhou 1,&, Lin Yang 1, Renato Paro 2,3 and Remo Rohs 1, * 1 Molecular and Computational Biology Program, Departments of Biological Sciences, Chemistry, Physics, and Computer Science, University of Southern California, Los Angeles, CA 90089, USA, 2 Department of Biosystems Science and Engineering, ETH Zürich, Mattenstrasse 26, 4058 Basel, Switzerland, 3 Faculty of Science, University of Basel, Klingelbergstrasse 50, 4046 Basel, Switzerland & Present address: Google Inc., 1600 Amphitheatre Parkway, Mountain View, CA 94043, USA # These authors contributed equally and are listed in alphabetic order. *To whom correspondence should be addressed: rohs@usc.edu Introduction DNAshapeR predicts DNA shape features in an ultra-fast, high-throughput manner from genomic sequencing data. The package takes either nucleotide sequence or genomic intervals as input, and generates various graphical representations for further analysis. DNAshapeR further encodes DNA sequence and shape features for statistical learning applications by concatenating feature matrices with user-defined combinations of k-mer and DNA shape features that can be readily used as input for machine learning algorithms. In this vignette, you will learn: o o o o how to load/install DNAshapeR how to predict DNA shape features how to visualize DNA shape predictions how to encode sequence and shape features, and apply them Load DNAshapeR library(dnashaper) Predict DNA shape features The core of DNAshapeR, the DNAshape method (Zhou, et al., 2013), uses a sliding pentamer window where structural features unique to each of the 512 distinct pentamers define a vector of minor groove width (MGW), Roll, propeller twist (ProT), and helix twist (HelT) at each nucleotide position. MGW and ProT define base-pair parameters whereas Roll and HelT represent base pairstep parameters. The values for each DNA shape feature as function of its pentamer sequence were derived from all-atom Monte Carlo simulations where DNA structure is sampled in collective and internal degrees of freedom in combination with explicit sodium counter ions (Zhang, et al., 2014).
2 The Monte Carlo simulations were analyzed with a modified Curves approach (Zhou, et al., 2013). Average values of each shape feature for each pentamer were derived from analyzing the ensemble of Monte Carlo predictions for 2,121 DNA fragments of base pairs in length. DNAshapeR predicts the four DNA shape features MGW, HelT, ProT, and Roll, which were observed in various cocrystal structures as playing an important role in specific protein-dna binding. In the latest version, we further added additional 9 DNA shape features beyond our previous set of 4 features, and expanded our available repertoire to a total of 13 features, including 6 inter-base pair or base pair-step parameters (HelT, Rise, Roll, Shift, Slide, and Tilt), 6 intra-base pair or base pairstep parameters (Buckle, Opening, ProT, Shear, Stagger, and Stretch), and MGW. Predict biophysical feature Our previous work explained protein-dna binding specificity based on correlations between MGW and (electrostatic potential) EP observed in experimentally available structures (Joshi, et al., 2007). However, A/T and C/G base pairs carry different partial charge distributions in the minor groove (due primarily to the guanine amino group), which will affect minor-groove EP. We developed a highthroughput method, named DNAphi, to predict minor-groove EP based on data mining of results from solving the nonlinear Poisson-Boltzmann calculations (Honig & Nicholls, 1995) on 2,297 DNA structures derived from Monte Carlo simulations. DNAshapeR includes EP as an additional feature. Predict DNA shape feature due to CpG methylation To achieve a better mechanistic understanding of the effect of CpG methylation on local DNA structure, we developed a high-throughput method, named methyl-dnashape, for predicting the impact of cytosine methylation on DNA shape features. In analogy to the DNAshape method (Zhou, et al., 2013), the method predicts DNA shape features (ProT, HelT, Roll, and MGW) in the context of CpG methylation based on methyl-dnashape Pentamer Query Table (mpqt) derived from the results of all-atom Monte Carlo simulations on a total of 3,518 DNA fragments of lengths varying from 13 to 24 bp. DNAshapeR can predict DNA shape features from custom FASTA files or directly from genomic coordinates in the form of a GRanges object within Bioconductor (see for more information). From FASTA file To predict DNA shape features from a FASTA file library(dnashaper) fn <- system.file("extdata", "CGRsample.fa", package = "DNAshapeR") pred <- getshape(fn)
3 ## Reading the input sequence... ## Reading the input sequence... ## Reading the input sequence... ## Reading the input sequence... ## Parsing files... ## Record length: 2000 ## Record length: 1999 ## Record length: 2000 ## Record length: 1999 ## Done From genomic intervals (e.g. TFs binding sites, CpG islands, replication origins, ) To predict DNA shape from genomic intervals stored as GRanges object, a reference genome is required. Several reference genomes are available within BioConductor as BSgenome objects (see for more information). For example, the saccer3 release of the S.Cerevisiae genome can be retrieved by # Install Bioconductor packages source(" bioclite("bsgenome.scerevisiae.ucsc.saccer3") library(bsgenome.scerevisiae.ucsc.saccer3) Given a reference genome, the getfasta function first extracts the DNA sequences based on the provided genomic coordinates, and then performs shape predictions within a user-defined window (of size equal to width, 100 bp in the example below) computed from the center of each genomic interval: # Create a query GRanges object gr <- GRanges(seqnames = c("chri"), strand = c("+", "-", "+"), ranges = IRanges(start = c(100, 200, 300), width = 100)) getfasta(gr, Scerevisiae, width = 100, filename = "tmp.fa") fn <- "tmp.fa" pred <- getshape(fn)
4 From public domain projects The genomic intervals can also be obtained from public domain projects, including ENCODE, NCBI, Ensembl, etc. The AnnotationHub package (see for more information) provides an interface to retrieve genomic intervals from these multiple online project resources. # Install Bioconductor packages library(bsgenome.hsapiens.ucsc.hg19) library(annotationhub) The genomic intervals of interest can be selected progressively through the functions of sebset and query with keywords, and can be subjected as an input of GRanges object to getfasta function. ah <- AnnotationHub() ah <- subset(ah, species=="homo sapiens") ah <- query(ah, c("h3k4me3", "Gm12878", "Roadmap")) getfasta(ah[[1]], Hsapiens, width = 150, filename = "tmp.fa") fn <- "tmp.fa" pred <- getshape(fn) From FASTA file with methylated DNA sequence To predict DNA shape features in the context of CpG methylation, one can prepare a FASTA file of sequence with symbol Mg : M referring to cytosine of methylated CpG on the leading strand and g referring the cytosine of methylated CpG on lagging strand. For example, > seq1 GTGTCACMgCGTCTATACG notifying the cytosine at position 8 th on the leading strand and the one at position 9 th on the lagging strand are methylated. library(dnashaper) fn <- system.file("extdata", "MethylSample.fa", package = "DNAshapeR") pred <- getshape(fn, methylate = TRUE) From FASTA and methylated position files To predict DNA shape features in the context of CpG methylation, in addition to providing regular FASTA file (without symbolizing Mg ) one can provide an additional input file identifying methylated positions. For example, > seq1 4,16
5 notifying the cytosine at position 4 th and 16 th on leading strand, and 5 th and 17 th on lagging strand are methylated. library(dnashaper) fn <- system.file("extdata", "SingleSeqsample.fa", package = "DNAshapeR") fn_pos <- system.file("extdata", "MethylSamplePos.fa", package = "DNAshapeR") pred <- getshape(fn, methylate = TRUE, methylatedposfile = fn_pos) Visualize DNA shape prediction DNAshapeR can be used to generate various graphical representations for further analyses. The prediction result can be visualized in the form of scatter plots (as introduced in Comoglio, et al., 2015), heat maps (as introduced in Yang, et al., 2014), or genome browser tracks (as introduced in Chiu, et al., 2014). Ensemble representation: metashape plot The prediction result can be visualized in the metaprofiles of DNA shape features. plotshape(pred$mgw)
6 # MGW can be replaced with HelT, Rise, Roll, Shift, Slide, Tilt, Buckle, Opening, ProT, Shear, Stagger, Stretch or EP Ensemble representation: heat map The prediction result can be visualized in the heat map of DNA shape features. library(fields) heatshape(pred$prot, 20)
7 # ProT can be replaced with MGW can be replaced with MGW, Buckle, Opening, ProT, Shear, Stagger, Stretch or EP #heatshape(pred$roll[1:500, 1:1980], 20) # Roll can be replaced with HelT, Rise, Roll, Shift, Slide or Tilt Individual representation: genome browser-like tracks The prediction result can be visualized in the form of genome browser tracks. *Note that the input data should only contain one sequence. fn2 <- system.file("extdata", "SingleSeqsample.fa", package = "DNAshapeR") pred2 <- getshape(fn2) trackshape(fn2, pred2) # Only for single sequence files
8 Encode sequence and shape features DNAshapeR can be used to generate feature vectors for a user-defined model. These models can consist of either sequence features (1-mer, 2-mer, 3-mer), shape features (MGW, HelT, Rise, Roll, Shift, Slide, Tilt, Buckle, Opening, ProT, Shear, Stagger, Stretch and EP), or any combination of those two. For 1-mer features, sequence is encoded in form of four binary numbers (i.e., in terms of 1-mers, 1000 for adenine, 0100 for cytosine, 0010 for guanine, and 0001 for thymine) at each nucleotide position (Zhou, et al., 2015). The feature encoding function of the DNAshapeR package
9 enables the determination of higher order sequence features, for example, 2-mers and 3-mers (16 and 64 binary features at each position, respectively). User can also choose to include second order shape features in the generated feature vector. The second order shape features are product terms of values for the same category of shape features (MGW, HelT, Rise, Roll, Shift, Slide, Tilt, Buckle, Opening, ProT, Shear, Stagger, Stretch or EP) at adjacent positions. They were introduced to encode the tendency of, for instance, a narrow minor groove region exhibiting an enhanced narrowing if adjacent positions are also characterized by a narrow groove (Zhou, et al., 2015). The feature encoding function of DNAshapeR enables the generation of any subset of these features, either only a selected shape category or first order shape features, and any combination with shape or sequence features. The result of feature encoding for each sequence is a chimera feature vector. Encoding process A feature type vector should be defined before encoding. The vector can be any combination of characters of k-mer, n-shape, n-mgw, n-prot, n-roll, n-helt, n-rise, n-shift, n-slide, n-tilt, n-buckle, n-opening, n-shear, n-stagger, n-stretch, n-ep (k, n are integers) where 1-shape refers to first order and 2-shape to second order shape features. Notice that n-shape represents the default 4 shape features (MGW, ProT, Roll and HelT). library(biostrings) featuretype <- c("1-mer", "1-shape") featurevector <- encodeseqshape(fn, pred, featuretype) featurevector Showcase of statistical machine learning application Feature encoding of multiple sequences thus results in a feature matrix, which can be used as input for variety of statistical machine learning methods. For example, an application is the quantitative modeling of PBM derived protein-dna binding by linear regression as demonstrated below. First, the experimental binding affinity values are combined with the feature matrix in a data frame structure. filename3 <- system.file("extdata", "PBMsample.s", package = "DNAshapeR") experimentaldata <- read.table(filename3) df <- data.frame(affinity=experimentaldata$v1, featurevector) Then, a machine learning package (which can be any learning tools) is used to train a multiple linear regression (MLR) model based on 10-fold cross-validation. In this example, we used the caret package (see for more information).
10 library(caret) traincontrol <- traincontrol(method = "cv", number = 10, savepredictions = TRUE) model <- train (affinity~., data = df, trcontrol=traincontrol, method="lm", preprocess=null) summary(model) Author contributions T.P.C., F.C., and R.R. designed and executed this project with the help of T.Z., L.Y., and R.P. based on methods developed by T.P.C., T.Z., and L.Y. J.L. introduced additional DNA shape features. The project was conceived by F.C. and directed by R.R. Acknowledgements The authors acknowledge comments and suggestions by members of the Rohs lab. This work was supported by the NIH (R01GM106056, U01GM103804, and R01HG to R.R.). Release of open-source software and open-access publication were supported by the NSF (MCB to R.R.). References Chiu, T.-P., et al. GBshape: a genome browser database for DNA shape annotations. Nucleic Acids Res. 2015;43:D Comoglio, F., et al. High-resolution profiling of Drosophila replication start sites reveals a DNA shape and chromatin signature of metazoan origins. Cell Rep. 2015;11(5): Yang, L., et al. TFBSshape: a motif database for DNA shape features of transcription factor binding sites. Nucleic Acids Res. 2014;42:D Zhou, T., et al. Quantitative modeling of transcription factor binding specificities using DNA shape. Proc. Natl. Acad. Sci. U S A 2015;112(15): Zhou, T., et al. DNAshape: a method for the high-throughput prediction of DNA structural features on a genomic scale. Nucleic Acids Res. 2013;41:W Zhang, X., et al. Conformations of p53 response elements in solution deduced using site-directed spin labeling and Monte Carlo sampling. Nucleic Acids Res. 2014;42(4):
11 Honig, B. and Nicholls, A. Classical electrostatics in biology and chemistry. Science, 1995;268: Joshi, R., et al. Functional specificity of a Hox protein mediated by the recognition of minor groove structure. Cell, 2007;131:
Supplementary Data for DNA sequence+shape kernel enables alignment-free modeling of transcription factor binding.
 Supplementary Data for DNA sequence+shape kernel enables alignment-free modeling of transcription factor binding. Wenxiu Ma 1, Lin Yang 2, Remo Rohs 2, and William Stafford Noble 3 1 Department of Statistics,
Supplementary Data for DNA sequence+shape kernel enables alignment-free modeling of transcription factor binding. Wenxiu Ma 1, Lin Yang 2, Remo Rohs 2, and William Stafford Noble 3 1 Department of Statistics,
Transcription factor binding site prediction in vivo using DNA sequence and shape features
 Transcription factor binding site prediction in vivo using DNA sequence and shape features Anthony Mathelier, Lin Yang, Tsu-Pei Chiu, Remo Rohs, and Wyeth Wasserman anthony.mathelier@gmail.com @AMathelier
Transcription factor binding site prediction in vivo using DNA sequence and shape features Anthony Mathelier, Lin Yang, Tsu-Pei Chiu, Remo Rohs, and Wyeth Wasserman anthony.mathelier@gmail.com @AMathelier
Representation in Supervised Machine Learning Application to Biological Problems
 Representation in Supervised Machine Learning Application to Biological Problems Frank Lab Howard Hughes Medical Institute & Columbia University 2010 Robert Howard Langlois Hughes Medical Institute What
Representation in Supervised Machine Learning Application to Biological Problems Frank Lab Howard Hughes Medical Institute & Columbia University 2010 Robert Howard Langlois Hughes Medical Institute What
Nucleic acids. How DNA works. DNA RNA Protein. DNA (deoxyribonucleic acid) RNA (ribonucleic acid) Central Dogma of Molecular Biology
 Nucleic acid chemistry and basic molecular theory Nucleic acids DNA (deoxyribonucleic acid) RNA (ribonucleic acid) Central Dogma of Molecular Biology Cell cycle DNA RNA Protein Transcription Translation
Nucleic acid chemistry and basic molecular theory Nucleic acids DNA (deoxyribonucleic acid) RNA (ribonucleic acid) Central Dogma of Molecular Biology Cell cycle DNA RNA Protein Transcription Translation
MCB 110:Biochemistry of the Central Dogma of MB. MCB 110:Biochemistry of the Central Dogma of MB
 MCB 110:Biochemistry of the Central Dogma of MB Part 1. DNA replication, repair and genomics (Prof. Alber) Part 2. RNA & protein synthesis. Prof. Zhou Part 3. Membranes, protein secretion, trafficking
MCB 110:Biochemistry of the Central Dogma of MB Part 1. DNA replication, repair and genomics (Prof. Alber) Part 2. RNA & protein synthesis. Prof. Zhou Part 3. Membranes, protein secretion, trafficking
Basic concepts of molecular biology
 Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it Life The main actors in the chemistry of life are molecules called proteins nucleic acids Proteins: many different
Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it Life The main actors in the chemistry of life are molecules called proteins nucleic acids Proteins: many different
Gene and DNA structure. Dr Saeb Aliwaini
 Gene and DNA structure Dr Saeb Aliwaini 2016 DNA during cell cycle Cell cycle for different cell types Molecular Biology - "Study of the synthesis, structure, and function of macromolecules (DNA, RNA,
Gene and DNA structure Dr Saeb Aliwaini 2016 DNA during cell cycle Cell cycle for different cell types Molecular Biology - "Study of the synthesis, structure, and function of macromolecules (DNA, RNA,
Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid
 Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Lecture 2: Central Dogma of Molecular Biology & Intro to Programming
 Lecture 2: Central Dogma of Molecular Biology & Intro to Programming Central Dogma of Molecular Biology Proteins: workhorse molecules of biological systems Proteins are synthesized from the genetic blueprints
Lecture 2: Central Dogma of Molecular Biology & Intro to Programming Central Dogma of Molecular Biology Proteins: workhorse molecules of biological systems Proteins are synthesized from the genetic blueprints
Biochemistry 302, February 11, 2004 Exam 1 (100 points) 1. What form of DNA is shown on this Nature Genetics cover? Z-DNA or left-handed DNA
 1 Biochemistry 302, February 11, 2004 Exam 1 (100 points) Name I. Structural recognition (very short answer, 2 points each) 1. What form of DNA is shown on this Nature Genetics cover? Z-DNA or left-handed
1 Biochemistry 302, February 11, 2004 Exam 1 (100 points) Name I. Structural recognition (very short answer, 2 points each) 1. What form of DNA is shown on this Nature Genetics cover? Z-DNA or left-handed
Characterizing DNA binding sites high throughput approaches Biol4230 Tues, April 24, 2018 Bill Pearson Pinn 6-057
 Characterizing DNA binding sites high throughput approaches Biol4230 Tues, April 24, 2018 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Reviewing sites: affinity and specificity representation binding
Characterizing DNA binding sites high throughput approaches Biol4230 Tues, April 24, 2018 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Reviewing sites: affinity and specificity representation binding
Biology 644: Bioinformatics
 Processes Activation Repression Initiation Elongation.... Processes Splicing Editing Degradation Translation.... Transcription Translation DNA Regulators DNA-Binding Transcription Factors Chromatin Remodelers....
Processes Activation Repression Initiation Elongation.... Processes Splicing Editing Degradation Translation.... Transcription Translation DNA Regulators DNA-Binding Transcription Factors Chromatin Remodelers....
1.1 Chemical structure and conformational flexibility of single-stranded DNA
 1 DNA structures 1.1 Chemical structure and conformational flexibility of single-stranded DNA Single-stranded DNA (ssdna) is the building base for the double helix and other DNA structures. All these structures
1 DNA structures 1.1 Chemical structure and conformational flexibility of single-stranded DNA Single-stranded DNA (ssdna) is the building base for the double helix and other DNA structures. All these structures
Suppl. Table S1. Characteristics of DHS regions analyzed by bisulfite sequencing. No. CpGs analyzed in the amplicon. Genomic location specificity
 Suppl. Table S1. Characteristics of DHS regions analyzed by bisulfite sequencing. DHS/GRE Genomic location Tissue specificity DHS type CpG density (per 100 bp) No. CpGs analyzed in the amplicon CpG within
Suppl. Table S1. Characteristics of DHS regions analyzed by bisulfite sequencing. DHS/GRE Genomic location Tissue specificity DHS type CpG density (per 100 bp) No. CpGs analyzed in the amplicon CpG within
This place covers: Methods or systems for genetic or protein-related data processing in computational molecular biology.
 G16B BIOINFORMATICS, i.e. INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR GENETIC OR PROTEIN-RELATED DATA PROCESSING IN COMPUTATIONAL MOLECULAR BIOLOGY Methods or systems for genetic
G16B BIOINFORMATICS, i.e. INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR GENETIC OR PROTEIN-RELATED DATA PROCESSING IN COMPUTATIONAL MOLECULAR BIOLOGY Methods or systems for genetic
(Very) Basic Molecular Biology
 (Very) Basic Molecular Biology (Very) Basic Molecular Biology Each human cell has 46 chromosomes --double-helix DNA molecule (Very) Basic Molecular Biology Each human cell has 46 chromosomes --double-helix
(Very) Basic Molecular Biology (Very) Basic Molecular Biology Each human cell has 46 chromosomes --double-helix DNA molecule (Very) Basic Molecular Biology Each human cell has 46 chromosomes --double-helix
The ChIP-Seq project. Giovanna Ambrosini, Philipp Bucher. April 19, 2010 Lausanne. EPFL-SV Bucher Group
 The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
2/10/17. Contents. Applications of HMMs in Epigenomics
 2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
Epigenetics and DNase-Seq
 Epigenetics and DNase-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by Anthony
Epigenetics and DNase-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by Anthony
What Are the Chemical Structures and Functions of Nucleic Acids?
 THE NUCLEIC ACIDS What Are the Chemical Structures and Functions of Nucleic Acids? Nucleic acids are polymers specialized for the storage, transmission, and use of genetic information. DNA = deoxyribonucleic
THE NUCLEIC ACIDS What Are the Chemical Structures and Functions of Nucleic Acids? Nucleic acids are polymers specialized for the storage, transmission, and use of genetic information. DNA = deoxyribonucleic
Chapter 8. Microbial Genetics. Lectures prepared by Christine L. Case. Copyright 2010 Pearson Education, Inc.
 Chapter 8 Microbial Genetics Lectures prepared by Christine L. Case Structure and Function of Genetic Material Learning Objectives 8-1 Define genetics, genome, chromosome, gene, genetic code, genotype,
Chapter 8 Microbial Genetics Lectures prepared by Christine L. Case Structure and Function of Genetic Material Learning Objectives 8-1 Define genetics, genome, chromosome, gene, genetic code, genotype,
Basic concepts of molecular biology
 Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it What is life made of? 1665: Robert Hooke discovered that organisms are composed of individual compartments called cells
Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it What is life made of? 1665: Robert Hooke discovered that organisms are composed of individual compartments called cells
Outline. Structure of DNA DNA Functions Transcription Translation Mutation Cytogenetics Mendelian Genetics Quantitative Traits Linkage
 Genetics Outline Structure of DNA DNA Functions Transcription Translation Mutation Cytogenetics Mendelian Genetics Quantitative Traits Linkage Chromosomes are composed of chromatin, which is DNA and associated
Genetics Outline Structure of DNA DNA Functions Transcription Translation Mutation Cytogenetics Mendelian Genetics Quantitative Traits Linkage Chromosomes are composed of chromatin, which is DNA and associated
Computational Analysis of Ultra-high-throughput sequencing data: ChIP-Seq
 Computational Analysis of Ultra-high-throughput sequencing data: ChIP-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne Data flow in ChIP-Seq data analysis Level 1:
Computational Analysis of Ultra-high-throughput sequencing data: ChIP-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne Data flow in ChIP-Seq data analysis Level 1:
Gene Identification in silico
 Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
DNA Structures. Biochemistry 201 Molecular Biology January 5, 2000 Doug Brutlag. The Structural Conformations of DNA
 DNA Structures Biochemistry 201 Molecular Biology January 5, 2000 Doug Brutlag The Structural Conformations of DNA 1. The principle message of this lecture is that the structure of DNA is much more flexible
DNA Structures Biochemistry 201 Molecular Biology January 5, 2000 Doug Brutlag The Structural Conformations of DNA 1. The principle message of this lecture is that the structure of DNA is much more flexible
Molecular Structures
 Molecular Structures 1 Molecular structures 2 Why is it important? Answers to scientific questions such as: What does the structure of protein X look like? Can we predict the binding of molecule X to Y?
Molecular Structures 1 Molecular structures 2 Why is it important? Answers to scientific questions such as: What does the structure of protein X look like? Can we predict the binding of molecule X to Y?
DNA RNA PROTEIN. Professor Andrea Garrison Biology 11 Illustrations 2010 Pearson Education, Inc. unless otherwise noted
 DNA RNA PROTEIN Professor Andrea Garrison Biology 11 Illustrations 2010 Pearson Education, Inc. unless otherwise noted DNA Molecule of heredity Contains all the genetic info our cells inherit Determines
DNA RNA PROTEIN Professor Andrea Garrison Biology 11 Illustrations 2010 Pearson Education, Inc. unless otherwise noted DNA Molecule of heredity Contains all the genetic info our cells inherit Determines
The RNA tools registry
 university of copenhagen f a c u lt y o f h e a lt h a n d m e d i c a l s c i e n c e s The RNA tools registry A community effort to catalog RNA bioinformatics resources and their relationships Anne Wenzel
university of copenhagen f a c u lt y o f h e a lt h a n d m e d i c a l s c i e n c e s The RNA tools registry A community effort to catalog RNA bioinformatics resources and their relationships Anne Wenzel
Bio-inspired Computing for Network Modelling
 ISSN (online): 2289-7984 Vol. 6, No.. Pages -, 25 Bio-inspired Computing for Network Modelling N. Rajaee *,a, A. A. S. A Hussaini b, A. Zulkharnain c and S. M. W Masra d Faculty of Engineering, Universiti
ISSN (online): 2289-7984 Vol. 6, No.. Pages -, 25 Bio-inspired Computing for Network Modelling N. Rajaee *,a, A. A. S. A Hussaini b, A. Zulkharnain c and S. M. W Masra d Faculty of Engineering, Universiti
Drug DNA interaction. Modeling DNA ligand interaction of intercalating ligands
 Drug DNA interaction DNA as carrier of genetic information is a major target for drug interaction because of the ability to interfere with transcription (gene expression and protein synthesis) and DNA
Drug DNA interaction DNA as carrier of genetic information is a major target for drug interaction because of the ability to interfere with transcription (gene expression and protein synthesis) and DNA
NUCLEIC ACIDS. DNA (Deoxyribonucleic Acid) and RNA (Ribonucleic Acid): information storage molecules made up of nucleotides.
 NUCLEIC ACIDS DNA (Deoxyribonucleic Acid) and RNA (Ribonucleic Acid): information storage molecules made up of nucleotides. Base Adenine Guanine Cytosine Uracil Thymine Abbreviation A G C U T DNA RNA 2
NUCLEIC ACIDS DNA (Deoxyribonucleic Acid) and RNA (Ribonucleic Acid): information storage molecules made up of nucleotides. Base Adenine Guanine Cytosine Uracil Thymine Abbreviation A G C U T DNA RNA 2
Algorithms in Bioinformatics
 Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline Central Dogma of Molecular
Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline Central Dogma of Molecular
Molecular Structures
 Molecular Structures 1 Molecular structures 2 Why is it important? Answers to scientific questions such as: What does the structure of protein X look like? Can we predict the binding of molecule X to Y?
Molecular Structures 1 Molecular structures 2 Why is it important? Answers to scientific questions such as: What does the structure of protein X look like? Can we predict the binding of molecule X to Y?
DNA & DNA : Protein Interactions BIBC 100
 DNA & DNA : Protein Interactions BIBC 100 Sequence = Information Alphabet = language L,I,F,E LIFE DNA = DNA code A, T, C, G CAC=Histidine CAG=Glutamine GGG=Glycine Protein = Protein code 20 a.a. LIVE EVIL
DNA & DNA : Protein Interactions BIBC 100 Sequence = Information Alphabet = language L,I,F,E LIFE DNA = DNA code A, T, C, G CAC=Histidine CAG=Glutamine GGG=Glycine Protein = Protein code 20 a.a. LIVE EVIL
DOUBLE-STRAND DNA BREAK PREDICTION USING EPIGENOME MARKS AT KILOBASE RESOLUTION
 DOUBLE-STRAND DNA BREAK PREDICTION USING EPIGENOME MARKS AT KILOBASE RESOLUTION Raphaël MOURAD, Assist. Prof. Centre de Biologie Intégrative Université Paul Sabatier, Toulouse III INTRODUCTION Double-strand
DOUBLE-STRAND DNA BREAK PREDICTION USING EPIGENOME MARKS AT KILOBASE RESOLUTION Raphaël MOURAD, Assist. Prof. Centre de Biologie Intégrative Université Paul Sabatier, Toulouse III INTRODUCTION Double-strand
Computational Genomics. Irit Gat-Viks & Ron Shamir & Haim Wolfson Fall
 Computational Genomics Irit Gat-Viks & Ron Shamir & Haim Wolfson Fall 2015-16 1 What s in class this week Motivation Administrata Some very basic biology Some very basic biotechnology Examples of our type
Computational Genomics Irit Gat-Viks & Ron Shamir & Haim Wolfson Fall 2015-16 1 What s in class this week Motivation Administrata Some very basic biology Some very basic biotechnology Examples of our type
NGS Approaches to Epigenomics
 I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
TERTIARY MOTIF INTERACTIONS ON RNA STRUCTURE
 1 TERTIARY MOTIF INTERACTIONS ON RNA STRUCTURE Bioinformatics Senior Project Wasay Hussain Spring 2009 Overview of RNA 2 The central Dogma of Molecular biology is DNA RNA Proteins The RNA (Ribonucleic
1 TERTIARY MOTIF INTERACTIONS ON RNA STRUCTURE Bioinformatics Senior Project Wasay Hussain Spring 2009 Overview of RNA 2 The central Dogma of Molecular biology is DNA RNA Proteins The RNA (Ribonucleic
DNA Structure & the Genome. Bio160 General Biology
 DNA Structure & the Genome Bio160 General Biology Lecture Outline I. DNA A nucleic acid II. Chromosome Structure III. Chromosomes and Genes IV. DNA vs. RNA I. DNA A Nucleic Acid Structure of DNA: Remember:
DNA Structure & the Genome Bio160 General Biology Lecture Outline I. DNA A nucleic acid II. Chromosome Structure III. Chromosomes and Genes IV. DNA vs. RNA I. DNA A Nucleic Acid Structure of DNA: Remember:
DNA and RNA are both made of nucleotides. Proteins are made of amino acids. Transcription can be reversed but translation cannot.
 INFORMATION TRANSFER Information in cells Properties of information Information must be able to be stored, accessed, retrieved, transferred, read and used. Information is about order, it is basically the
INFORMATION TRANSFER Information in cells Properties of information Information must be able to be stored, accessed, retrieved, transferred, read and used. Information is about order, it is basically the
DNA & Genetics. Chapter Introduction DNA 6/12/2012. How are traits passed from parents to offspring?
 Section 5.3 DNA & Genetics Chapter Introduction How are traits passed from parents to offspring? Chromatin- DNA in the nucleus loose strands Chromosome- When DNA gets organized before cell division Gene-
Section 5.3 DNA & Genetics Chapter Introduction How are traits passed from parents to offspring? Chromatin- DNA in the nucleus loose strands Chromosome- When DNA gets organized before cell division Gene-
2/19/13. Contents. Applications of HMMs in Epigenomics
 2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
AGENDA for 10/11/13 AGENDA: HOMEWORK: Due end of the period OBJECTIVES:
 AGENDA for 10/11/13 AGENDA: 1. Finish 1.2.3 DNA Analysis Analyzing DNA Samples Using Current Forensic Methods OBJECTIVES: 1. Demonstrate the steps of gel electrophoresis 2. Analyze restriction fragment
AGENDA for 10/11/13 AGENDA: 1. Finish 1.2.3 DNA Analysis Analyzing DNA Samples Using Current Forensic Methods OBJECTIVES: 1. Demonstrate the steps of gel electrophoresis 2. Analyze restriction fragment
Discovery of Transcription Factor Binding Sites with Deep Convolutional Neural Networks
 Discovery of Transcription Factor Binding Sites with Deep Convolutional Neural Networks Reesab Pathak Dept. of Computer Science Stanford University rpathak@stanford.edu Abstract Transcription factors are
Discovery of Transcription Factor Binding Sites with Deep Convolutional Neural Networks Reesab Pathak Dept. of Computer Science Stanford University rpathak@stanford.edu Abstract Transcription factors are
Applications of HMMs in Epigenomics
 I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
SUPPLEMENTARY METHODS. Retrieval of experimentally determined structures from Protein Data Bank (PDB)
 SUPPLEMENTARY METHODS Retrieval of experimentally determined structures from Protein Data Bank (PDB) To generate count statistics, we used an advanced search interface to query the PDB [1] for occurrences
SUPPLEMENTARY METHODS Retrieval of experimentally determined structures from Protein Data Bank (PDB) To generate count statistics, we used an advanced search interface to query the PDB [1] for occurrences
DNA. Branden & Tooze, Ch. 7 Deoxyribose nucleic acids are made of three parts
 DNA Branden & Tooze, Ch. 7 Deoxyribose nucleic acids are made of three parts base: adenine, cytosine, guanine, thymine sugar: deoxyribose phosphate: will form the phosphate backbone wide narrow DNA binding
DNA Branden & Tooze, Ch. 7 Deoxyribose nucleic acids are made of three parts base: adenine, cytosine, guanine, thymine sugar: deoxyribose phosphate: will form the phosphate backbone wide narrow DNA binding
Introduction to Bioinformatics
 Introduction to Bioinformatics If the 19 th century was the century of chemistry and 20 th century was the century of physic, the 21 st century promises to be the century of biology...professor Dr. Satoru
Introduction to Bioinformatics If the 19 th century was the century of chemistry and 20 th century was the century of physic, the 21 st century promises to be the century of biology...professor Dr. Satoru
A. Incorrect! A sugar residue is only part of a nucleotide. Go back and review the structure of nucleotides.
 Organic Chemistry - Problem Drill 24: ucleic Acids o. 1 of 10 1. What are the components of a nucleotide? (A) A sugar residue (B) A sugar residue + a nitrogenous base (C) A sugar residue + a nitrogenous
Organic Chemistry - Problem Drill 24: ucleic Acids o. 1 of 10 1. What are the components of a nucleotide? (A) A sugar residue (B) A sugar residue + a nitrogenous base (C) A sugar residue + a nitrogenous
Introduction to Microarray Data Analysis and Gene Networks. Alvis Brazma European Bioinformatics Institute
 Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
Mass Spectrometry. Kyle Chau and Andrew Gioe
 Mass Spectrometry Kyle Chau and Andrew Gioe Computation of Molecular Mass - Mass Spectrum is a plot of intensity as a function of masscharge ratio, m/z. - Mass-charge ratio can be determined through accelerating
Mass Spectrometry Kyle Chau and Andrew Gioe Computation of Molecular Mass - Mass Spectrum is a plot of intensity as a function of masscharge ratio, m/z. - Mass-charge ratio can be determined through accelerating
Non-coding Function & Variation, MPRAs. Mike White Bio5488 3/5/18
 Non-coding Function & Variation, MPRAs Mike White Bio5488 3/5/18 Outline MONDAY Non-coding function and variation The barcode Basic versions of MRPA technology WEDNESDAY More varieties of MRPAs Some key
Non-coding Function & Variation, MPRAs Mike White Bio5488 3/5/18 Outline MONDAY Non-coding function and variation The barcode Basic versions of MRPA technology WEDNESDAY More varieties of MRPAs Some key
Name 10 Molecular Biology of the Gene Test Date Study Guide You must know: The structure of DNA. The major steps to replication.
 Name 10 Molecular Biology of the Gene Test Date Study Guide You must know: The structure of DNA. The major steps to replication. The difference between replication, transcription, and translation. How
Name 10 Molecular Biology of the Gene Test Date Study Guide You must know: The structure of DNA. The major steps to replication. The difference between replication, transcription, and translation. How
Learning Methods for DNA Binding in Computational Biology
 Learning Methods for DNA Binding in Computational Biology Mark Kon Dustin Holloway Yue Fan Chaitanya Sai Charles DeLisi Boston University IJCNN Orlando August 16, 2007 Outline Background on Transcription
Learning Methods for DNA Binding in Computational Biology Mark Kon Dustin Holloway Yue Fan Chaitanya Sai Charles DeLisi Boston University IJCNN Orlando August 16, 2007 Outline Background on Transcription
Homework 4. Due in class, Wednesday, November 10, 2004
 1 GCB 535 / CIS 535 Fall 2004 Homework 4 Due in class, Wednesday, November 10, 2004 Comparative genomics 1. (6 pts) In Loots s paper (http://www.seas.upenn.edu/~cis535/lab/sciences-loots.pdf), the authors
1 GCB 535 / CIS 535 Fall 2004 Homework 4 Due in class, Wednesday, November 10, 2004 Comparative genomics 1. (6 pts) In Loots s paper (http://www.seas.upenn.edu/~cis535/lab/sciences-loots.pdf), the authors
AGENDA for 10/10/13 AGENDA: HOMEWORK: Due end of the period OBJECTIVES: Due Fri, 10-11
 AGENDA for 10/10/13 AGENDA: 1. 1.2.3 DNA Analysis Analyzing DNA Samples Using Current Forensic Methods OBJECTIVES: 1. Demonstrate the steps of gel electrophoresis 2. Analyze restriction fragment length
AGENDA for 10/10/13 AGENDA: 1. 1.2.3 DNA Analysis Analyzing DNA Samples Using Current Forensic Methods OBJECTIVES: 1. Demonstrate the steps of gel electrophoresis 2. Analyze restriction fragment length
Sections 12.3, 13.1, 13.2
 Sections 12.3, 13.1, 13.2 Background: Watson & Crick recognized that base pairing in the double helix allows DNA to be copied, or replicated Each strand in the double helix has all the information to remake
Sections 12.3, 13.1, 13.2 Background: Watson & Crick recognized that base pairing in the double helix allows DNA to be copied, or replicated Each strand in the double helix has all the information to remake
BIOLOGICAL SCIENCE. Lecture Presentation by Cindy S. Malone, PhD, California State University Northridge. FIFTH EDITION Freeman Quillin Allison
 BIOLOGICAL SCIENCE FIFTH EDITION Freeman Quillin Allison 4 Lecture Presentation by Cindy S. Malone, PhD, California State University Northridge In this chapter you will learn that Nucleic acids store the
BIOLOGICAL SCIENCE FIFTH EDITION Freeman Quillin Allison 4 Lecture Presentation by Cindy S. Malone, PhD, California State University Northridge In this chapter you will learn that Nucleic acids store the
ON USING DNA DISTANCES AND CONSENSUS IN REPEATS DETECTION
 ON USING DNA DISTANCES AND CONSENSUS IN REPEATS DETECTION Petre G. POP Technical University of Cluj-Napoca, Romania petre.pop@com.utcluj.ro Abstract: Sequence repeats are the simplest form of regularity
ON USING DNA DISTANCES AND CONSENSUS IN REPEATS DETECTION Petre G. POP Technical University of Cluj-Napoca, Romania petre.pop@com.utcluj.ro Abstract: Sequence repeats are the simplest form of regularity
Roadmap. The Cell. Introduction to Molecular Biology. DNA RNA Protein Central dogma Genetic code Gene structure Human Genome
 Introduction to Molecular Biology Lodish et al Ch1-4 http://www.ncbi.nlm.nih.gov/books EECS 458 CWRU Fall 2004 DNA RNA Protein Central dogma Genetic code Gene structure Human Genome Roadmap The Cell Lodish
Introduction to Molecular Biology Lodish et al Ch1-4 http://www.ncbi.nlm.nih.gov/books EECS 458 CWRU Fall 2004 DNA RNA Protein Central dogma Genetic code Gene structure Human Genome Roadmap The Cell Lodish
Bioinformatics Course AA 2017/2018 Tutorial 2
 UNIVERSITÀ DEGLI STUDI DI PAVIA - FACOLTÀ DI SCIENZE MM.FF.NN. - LM MOLECULAR BIOLOGY AND GENETICS Bioinformatics Course AA 2017/2018 Tutorial 2 Anna Maria Floriano annamaria.floriano01@universitadipavia.it
UNIVERSITÀ DEGLI STUDI DI PAVIA - FACOLTÀ DI SCIENZE MM.FF.NN. - LM MOLECULAR BIOLOGY AND GENETICS Bioinformatics Course AA 2017/2018 Tutorial 2 Anna Maria Floriano annamaria.floriano01@universitadipavia.it
Molecular Biology. IMBB 2017 RAB, Kigali - Rwanda May 02 13, Francesca Stomeo
 Molecular Biology IMBB 2017 RAB, Kigali - Rwanda May 02 13, 2017 Francesca Stomeo Molecular biology is the study of biology at a molecular level, especially DNA and RNA - replication, transcription, translation,
Molecular Biology IMBB 2017 RAB, Kigali - Rwanda May 02 13, 2017 Francesca Stomeo Molecular biology is the study of biology at a molecular level, especially DNA and RNA - replication, transcription, translation,
Genome 541 Gene regulation and epigenomics Lecture 3 Integrative analysis of genomics assays
 Genome 541 Gene regulation and epigenomics Lecture 3 Integrative analysis of genomics assays Please consider both the forward and reverse strands (i.e. reverse compliment sequence). You do not need to
Genome 541 Gene regulation and epigenomics Lecture 3 Integrative analysis of genomics assays Please consider both the forward and reverse strands (i.e. reverse compliment sequence). You do not need to
Genomic Regions Flanking E-Box Binding Sites Influence DNA Binding Specificity of bhlh Transcription Factors through DNA Shape
 Cell Reports Article Genomic Regions Flanking E-Box Binding Sites Influence DNA Binding Specificity of bhlh Transcription Factors through DNA Shape Raluca Gordân, 1,7 Ning Shen, 3,6 Iris Dror, 5,6 Tianyin
Cell Reports Article Genomic Regions Flanking E-Box Binding Sites Influence DNA Binding Specificity of bhlh Transcription Factors through DNA Shape Raluca Gordân, 1,7 Ning Shen, 3,6 Iris Dror, 5,6 Tianyin
Basic Bioinformatics: Homology, Sequence Alignment,
 Basic Bioinformatics: Homology, Sequence Alignment, and BLAST William S. Sanders Institute for Genomics, Biocomputing, and Biotechnology (IGBB) High Performance Computing Collaboratory (HPC 2 ) Mississippi
Basic Bioinformatics: Homology, Sequence Alignment, and BLAST William S. Sanders Institute for Genomics, Biocomputing, and Biotechnology (IGBB) High Performance Computing Collaboratory (HPC 2 ) Mississippi
PRESENTING SEQUENCES 5 GAATGCGGCTTAGACTGGTACGATGGAAC 3 3 CTTACGCCGAATCTGACCATGCTACCTTG 5
 Molecular Biology-2017 1 PRESENTING SEQUENCES As you know, sequences may either be double stranded or single stranded and have a polarity described as 5 and 3. The 5 end always contains a free phosphate
Molecular Biology-2017 1 PRESENTING SEQUENCES As you know, sequences may either be double stranded or single stranded and have a polarity described as 5 and 3. The 5 end always contains a free phosphate
RNA is a single strand molecule composed of subunits called nucleotides joined by phosphodiester bonds.
 The Versatility of RNA Primary structure of RNA RNA is a single strand molecule composed of subunits called nucleotides joined by phosphodiester bonds. Each nucleotide subunit is composed of a ribose sugar,
The Versatility of RNA Primary structure of RNA RNA is a single strand molecule composed of subunits called nucleotides joined by phosphodiester bonds. Each nucleotide subunit is composed of a ribose sugar,
DNAFSMiner: A Web-Based Software Toolbox to Recognize Two Types of Functional Sites in DNA Sequences
 DNAFSMiner: A Web-Based Software Toolbox to Recognize Two Types of Functional Sites in DNA Sequences Huiqing Liu Hao Han Jinyan Li Limsoon Wong Institute for Infocomm Research, 21 Heng Mui Keng Terrace,
DNAFSMiner: A Web-Based Software Toolbox to Recognize Two Types of Functional Sites in DNA Sequences Huiqing Liu Hao Han Jinyan Li Limsoon Wong Institute for Infocomm Research, 21 Heng Mui Keng Terrace,
Introduction to Bioinformatics
 Introduction to Bioinformatics Dan Lopresti Computer Science and Engineering Office PL 350 dal9@lehigh.edu BioS 10 November 2018 Slide 1 Last year when I gave this talk BioS 10 November 2018 Slide 2 Motivation
Introduction to Bioinformatics Dan Lopresti Computer Science and Engineering Office PL 350 dal9@lehigh.edu BioS 10 November 2018 Slide 1 Last year when I gave this talk BioS 10 November 2018 Slide 2 Motivation
Many transcription factors! recognize DNA shape
 Many transcription factors! recognize DN shape Katie Pollard! Gladstone Institutes USF Division of Biostatistics, Institute for Human Genetics, and Institute for omputational Health Sciences ENODE Users
Many transcription factors! recognize DN shape Katie Pollard! Gladstone Institutes USF Division of Biostatistics, Institute for Human Genetics, and Institute for omputational Health Sciences ENODE Users
btd_getpdb Retrieve protein structure data from Protein Data Bank (PDB) database btd_getuniprot Retrieve sequence information from UniProt database
 BTD btd.nwalign Needleman-Wunsch Global alignment of two nucleotide sequences btd.swalign Smith Waterman Local alignment of two nucleotide sequences btd_cpg CpG Islands (counts 'GC' dinucleotides in a
BTD btd.nwalign Needleman-Wunsch Global alignment of two nucleotide sequences btd.swalign Smith Waterman Local alignment of two nucleotide sequences btd_cpg CpG Islands (counts 'GC' dinucleotides in a
The Structure and Func.on of Macromolecules Nucleic Acids
 The Structure and Func.on of Macromolecules Nucleic Acids The FOUR Classes of Large Biomolecules All living things are made up of four classes of large biological molecules: Carbohydrates Lipids Protein
The Structure and Func.on of Macromolecules Nucleic Acids The FOUR Classes of Large Biomolecules All living things are made up of four classes of large biological molecules: Carbohydrates Lipids Protein
CMSE 520 BIOMOLECULAR STRUCTURE, FUNCTION AND DYNAMICS
 CMSE 520 BIOMOLECULAR STRUCTURE, FUNCTION AND DYNAMICS (Computational Structural Biology) OUTLINE Review: Molecular biology Proteins: structure, conformation and function(5 lectures) Generalized coordinates,
CMSE 520 BIOMOLECULAR STRUCTURE, FUNCTION AND DYNAMICS (Computational Structural Biology) OUTLINE Review: Molecular biology Proteins: structure, conformation and function(5 lectures) Generalized coordinates,
Nucleic acids and protein synthesis
 THE FUNCTIONS OF DNA Nucleic acids and protein synthesis The full name of DNA is deoxyribonucleic acid. Every nucleotide has the same sugar molecule and phosphate group, but each nucleotide contains one
THE FUNCTIONS OF DNA Nucleic acids and protein synthesis The full name of DNA is deoxyribonucleic acid. Every nucleotide has the same sugar molecule and phosphate group, but each nucleotide contains one
Nucleic Acids. Information specifying protein structure
 Nucleic Acids Nucleic acids represent the fourth major class of biomolecules (other major classes of biomolecules are proteins, carbohydrates, fats) Genome - the genetic information of an organism Information
Nucleic Acids Nucleic acids represent the fourth major class of biomolecules (other major classes of biomolecules are proteins, carbohydrates, fats) Genome - the genetic information of an organism Information
Information specifying protein structure. Chapter 19 Nucleic Acids Nucleotides Are the Building Blocks of Nucleic Acids
 Chapter 19 Nucleic Acids Information specifying protein structure Nucleic acids represent the fourth major class of biomolecules (other major classes of biomolecules are proteins, carbohydrates, fats)
Chapter 19 Nucleic Acids Information specifying protein structure Nucleic acids represent the fourth major class of biomolecules (other major classes of biomolecules are proteins, carbohydrates, fats)
Independent Study Guide The Blueprint of Life, from DNA to Protein (Chapter 7)
 Independent Study Guide The Blueprint of Life, from DNA to Protein (Chapter 7) I. General Principles (Chapter 7 introduction) a. Morse code distinct series of dots and dashes encode the 26 letters of the
Independent Study Guide The Blueprint of Life, from DNA to Protein (Chapter 7) I. General Principles (Chapter 7 introduction) a. Morse code distinct series of dots and dashes encode the 26 letters of the
Discovering gene regulatory control using ChIP-chip and ChIP-seq. An introduction to gene regulatory control, concepts and methodologies
 Discovering gene regulatory control using ChIP-chip and ChIP-seq An introduction to gene regulatory control, concepts and methodologies Ian Simpson ian.simpson@.ed.ac.uk bit.ly/bio2_2012 The Central Dogma
Discovering gene regulatory control using ChIP-chip and ChIP-seq An introduction to gene regulatory control, concepts and methodologies Ian Simpson ian.simpson@.ed.ac.uk bit.ly/bio2_2012 The Central Dogma
Exploring Similarities of Conserved Domains/Motifs
 Exploring Similarities of Conserved Domains/Motifs Sotiria Palioura Abstract Traditionally, proteins are represented as amino acid sequences. There are, though, other (potentially more exciting) representations;
Exploring Similarities of Conserved Domains/Motifs Sotiria Palioura Abstract Traditionally, proteins are represented as amino acid sequences. There are, though, other (potentially more exciting) representations;
Student Manual. Pre-Lab Introduction to DNA Fingerprinting STUDENT MANUAL BACKGROUND
 BACKGROUND Pre-Lab Introduction to DNA Fingerprinting You are about to perform a procedure known as DNA fingerprinting. The data obtained may allow you to determine if the samples of DNA that you will
BACKGROUND Pre-Lab Introduction to DNA Fingerprinting You are about to perform a procedure known as DNA fingerprinting. The data obtained may allow you to determine if the samples of DNA that you will
A Brief Introduction to Bioinformatics
 A Brief Introduction to Bioinformatics Dan Lopresti Associate Professor Office PL 404B dal9@lehigh.edu February 2007 Slide 1 Motivation Biology easily has 500 years of exciting problems to work on. Donald
A Brief Introduction to Bioinformatics Dan Lopresti Associate Professor Office PL 404B dal9@lehigh.edu February 2007 Slide 1 Motivation Biology easily has 500 years of exciting problems to work on. Donald
Laboratory 3. Molecular Genetics
 Laboratory 3. Molecular Genetics I. The Basics of DNA Structure DNA molecules are composed of small building blocks called nucleotides: A. Each DNA nucleotide is composed of three smaller molecules bonded
Laboratory 3. Molecular Genetics I. The Basics of DNA Structure DNA molecules are composed of small building blocks called nucleotides: A. Each DNA nucleotide is composed of three smaller molecules bonded
Sequence logos for DNA sequence alignments
 Sequence logos for DNA sequence alignments Oliver Bembom Division of Biostatistics, University of California, Berkeley October, 202 Introduction An alignment of DNA or amino acid sequences is commonly
Sequence logos for DNA sequence alignments Oliver Bembom Division of Biostatistics, University of California, Berkeley October, 202 Introduction An alignment of DNA or amino acid sequences is commonly
What happens after DNA Replication??? Transcription, translation, gene expression/protein synthesis!!!!
 What happens after DNA Replication??? Transcription, translation, gene expression/protein synthesis!!!! Protein Synthesis/Gene Expression Why do we need to make proteins? To build parts for our body as
What happens after DNA Replication??? Transcription, translation, gene expression/protein synthesis!!!! Protein Synthesis/Gene Expression Why do we need to make proteins? To build parts for our body as
Molecular Biology (1)
 Molecular Biology (1) DNA structure and basic applications Mamoun Ahram, PhD Second semester, 2017-2018 Resources This lecture Cooper, pp. 49-52, 118-119, 130 What is molecular biology? Central dogma
Molecular Biology (1) DNA structure and basic applications Mamoun Ahram, PhD Second semester, 2017-2018 Resources This lecture Cooper, pp. 49-52, 118-119, 130 What is molecular biology? Central dogma
A Protein Secondary Structure Prediction Method Based on BP Neural Network Ru-xi YIN, Li-zhen LIU*, Wei SONG, Xin-lei ZHAO and Chao DU
 2017 2nd International Conference on Artificial Intelligence: Techniques and Applications (AITA 2017 ISBN: 978-1-60595-491-2 A Protein Secondary Structure Prediction Method Based on BP Neural Network Ru-xi
2017 2nd International Conference on Artificial Intelligence: Techniques and Applications (AITA 2017 ISBN: 978-1-60595-491-2 A Protein Secondary Structure Prediction Method Based on BP Neural Network Ru-xi
Name Date Class. The Central Dogma of Biology
 Concept Mapping The Central Dogma of Biology Complete the events chain showing the events that occur as DNA codes for RNA, which guides the synthesis of proteins, the central dogma of biology. These terms
Concept Mapping The Central Dogma of Biology Complete the events chain showing the events that occur as DNA codes for RNA, which guides the synthesis of proteins, the central dogma of biology. These terms
Structural Bioinformatics (C3210) DNA and RNA Structure
 Structural Bioinformatics (C3210) DNA and RNA Structure Importance of DNA/RNA 3D Structure Nucleic acids are essential materials found in all living organisms. Their main function is to maintain and transmit
Structural Bioinformatics (C3210) DNA and RNA Structure Importance of DNA/RNA 3D Structure Nucleic acids are essential materials found in all living organisms. Their main function is to maintain and transmit
ChIP-seq analysis 2/28/2018
 ChIP-seq analysis 2/28/2018 Acknowledgements Much of the content of this lecture is from: Furey (2012) ChIP-seq and beyond Park (2009) ChIP-seq advantages + challenges Landt et al. (2012) ChIP-seq guidelines
ChIP-seq analysis 2/28/2018 Acknowledgements Much of the content of this lecture is from: Furey (2012) ChIP-seq and beyond Park (2009) ChIP-seq advantages + challenges Landt et al. (2012) ChIP-seq guidelines
I. Gene Expression Figure 1: Central Dogma of Molecular Biology
 I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
Chapter 13 - Concept Mapping
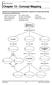 Chapter 13 - Concept Mapping Using the terms and phrases provided below, complete the concept map showing the discovery of DNA structure. amount of base pairs five-carbon sugar purine DNA polymerases Franklin
Chapter 13 - Concept Mapping Using the terms and phrases provided below, complete the concept map showing the discovery of DNA structure. amount of base pairs five-carbon sugar purine DNA polymerases Franklin
CRISPRseek Workshop Design of target-specific guide RNAs in CRISPR-Cas9 genome-editing systems
 April 2008 CRISPRseek Workshop Design of target-specific guide RNAs in CRISPR-Cas9 genome-editing systems Lihua Julie Zhu August 1st 2014 Outline Background and Motives CRISPRseek Functionality Dependency
April 2008 CRISPRseek Workshop Design of target-specific guide RNAs in CRISPR-Cas9 genome-editing systems Lihua Julie Zhu August 1st 2014 Outline Background and Motives CRISPRseek Functionality Dependency
Reading for lecture 2
 Reading for lecture 2 1. Structure of DNA and RNA 2. Information storage by DNA 3. The Central Dogma Voet and Voet, Chapters 28 (29,30) Alberts et al, Chapters 5 (3) 1 5 4 1 3 2 3 3 Structure of DNA and
Reading for lecture 2 1. Structure of DNA and RNA 2. Information storage by DNA 3. The Central Dogma Voet and Voet, Chapters 28 (29,30) Alberts et al, Chapters 5 (3) 1 5 4 1 3 2 3 3 Structure of DNA and
What is Life? David Martin Degner Degner Scientific and Engineering Anchorage, Alaska
 What is Life? David Martin Degner Degner Scientific and Engineering Anchorage, Alaska davidmartindegner@gmail.com A static and dynamic physical model is presented for the Gram (+) prokaryotic cell, the
What is Life? David Martin Degner Degner Scientific and Engineering Anchorage, Alaska davidmartindegner@gmail.com A static and dynamic physical model is presented for the Gram (+) prokaryotic cell, the
March 9, Hidden Markov Models and. BioInformatics, Part I. Steven R. Dunbar. Intro. BioInformatics Problem. Hidden Markov.
 and, and, March 9, 2017 1 / 30 Outline and, 1 2 3 4 2 / 30 Background and, Prof E. Moriyama (SBS) has a Seminar SBS, Math, Computer Science, Statistics Extensive use of program "HMMer" Britney (Hinds)
and, and, March 9, 2017 1 / 30 Outline and, 1 2 3 4 2 / 30 Background and, Prof E. Moriyama (SBS) has a Seminar SBS, Math, Computer Science, Statistics Extensive use of program "HMMer" Britney (Hinds)
DNA. Using DNA to solve crimes
 DNA Using DNA to solve crimes Physical characteristics are inherited from both parents DNA contains all the inherited information for each person DNA is contained in the nucleus of every cell in your body
DNA Using DNA to solve crimes Physical characteristics are inherited from both parents DNA contains all the inherited information for each person DNA is contained in the nucleus of every cell in your body
Resources. How to Use This Presentation. Chapter 10. Objectives. Table of Contents. Griffith s Discovery of Transformation. Griffith s Experiments
 How to Use This Presentation To View the presentation as a slideshow with effects select View on the menu bar and click on Slide Show. To advance through the presentation, click the right-arrow key or
How to Use This Presentation To View the presentation as a slideshow with effects select View on the menu bar and click on Slide Show. To advance through the presentation, click the right-arrow key or
Introduction to Bioinformatics
 Introduction to Bioinformatics Dr. Taysir Hassan Abdel Hamid Lecturer, Information Systems Department Faculty of Computer and Information Assiut University taysirhs@aun.edu.eg taysir_soliman@hotmail.com
Introduction to Bioinformatics Dr. Taysir Hassan Abdel Hamid Lecturer, Information Systems Department Faculty of Computer and Information Assiut University taysirhs@aun.edu.eg taysir_soliman@hotmail.com
BIMM 143: Introduction to Bioinformatics (Winter 2018)
 BIMM 143: Introduction to Bioinformatics (Winter 2018) Course Instructor: Dr. Barry J. Grant ( bjgrant@ucsd.edu ) Course Website: https://bioboot.github.io/bimm143_w18/ DRAFT: 2017-12-02 (20:48:10 PST
BIMM 143: Introduction to Bioinformatics (Winter 2018) Course Instructor: Dr. Barry J. Grant ( bjgrant@ucsd.edu ) Course Website: https://bioboot.github.io/bimm143_w18/ DRAFT: 2017-12-02 (20:48:10 PST
