Time Course Global Gene Expression Analysis of an In Vivo Candida Biofilm
|
|
|
- Beverly Owen
- 6 years ago
- Views:
Transcription
1 MAJOR ARTICLE Time Course Global Gene Expression Analysis of an In Vivo Candida Biofilm Jeniel E. Nett, 1,3 Alexander J. Lepak, 1 Karen Marchillo, 1 and David R. Andes 1,2 Departments of 1 Medicine, 2 Medical Microbiology and Immunology, and 3 Cellular and Molecular Biology, University of Wisconsin, Madison, Wisconsin Candida infection of devices is common and invariably associated with biofilm growth. Exploratory microarray studies were undertaken to identify target genes associated with biofilm formation from an in vivo catheter model over time. We compared messenger RNA levels from Candida albicans grown in an in vivo central venous catheter biofilm model at 12 h (intermediate growth) and 24 h (mature) to in vitro planktonic cells without a biofilm substrate, using C. albicans oligo arrays. A total of 124 transcripts were similarly up-regulated at the 12- and 24-h time points. Ontology categories most highly represented included energy/metabolism (12%), carbohydrate (10%), and protein (13%) synthesis and modification, and transport (6%). Numerous genes were previously identified from in vitro biofilm studies. These genes included those associated with hyphal growth, amino acid metabolism, adherence, drug resistance, ergosterol biosynthesis, and b-glucan synthesis. In the current data set, adherence genes were unique to those from the earlier time point. Differences between the current in vivo biofilm expression data and that previously reported from in vitro models, including alterations in metabolism and carbohydrate processing, may be due to the continuous availability of nutrients from host serum and the incorporation of the host-pathogen interaction. Candida albicans is an important human pathogen causing a wide spectrum of diseases and is a leading cause of nosocomial bloodstream infection. Candidemia is frequently associated with implantation of a medical device. When Candida organisms grow on surfaces, such as a venous catheter or urinary catheter, they adapt to a biofilm lifestyle [1, 2]. Biofilm growth involves phenotypic changes distinct from planktonic growth [3 5]. Two such characteristics, drug resistance and reduced susceptibility to the host immune system, contribute to the difficulty in treating Candida biofilm infections. Greater understanding of the gene expression patterns present in Candida biofilms is one method Received 8 September 2008; accepted 21 October 2008; electronically published 15 June Potential conflicts of interest: none reported. Financial support: National Institutes of Health (grant RO1 AI ). The content of this article is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. Reprints or correspondence: Dr. David R. Andes, 600 Highland Ave., Rm. H4.572, Madison, WI (dra@medicine.wisc.edu). The Journal of Infectious Diseases 2009; 200: by the Infectious Diseases Society of America. All rights reserved /2009/ $15.00 DOI: / to begin to discern the pathways important for the biofilm phenotype and identify potential antifungal targets. Many in vitro model systems have been developed to mimic the biofilm growth found on infected medical devices. These models have provided the foundation for investigation of biofilm composition, architecture, and mechanisms of drug resistance [3, 5, 6]. More recently, transcription profiling experiments have identified potential biochemical pathways required for biofilm formation [7 9]. Although the substratum common to these clinical infection sites (i.e., catheter and denture material) can be used in vitro, the models lack other environmental conditions potentially important in human infections [10]. In device-associated infections, the substratum has been preconditioned with host proteins soon after implantation [1, 10]. This preconditioning film impacts biofilm formation and extracellular matrix production. In vitro models are not necessarily exposed to all the host proteins needed for the appropriate preconditioning film. Also, in many in vitro models, nutrients are depleted and waste products accumulate. However, in vivo biofilm cells are exposed Gene Expression Analysis of In Vivo Candida Biofilm JID 2009:200 (15 July) 307
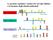
2 Table 1. Primers used for RT-PCR experiments. to a continuous nutrient supply and waste products are eliminated while in the vasculature or other host tissue site [11]. Imaging of in vivo biofilms has also demonstrated incorporation of host cells, which may potentially impact biofilm formation. This host-pathogen interaction is not represented in these in vitro models. Finally, flow dynamics of host vasculature are difficult to reproduce in vitro. In the current experiments, we use an in vivo rat catheter biofilm infection model [11]. This central venous catheter rat model mimics conditions encountered in human infections, including exposure to host proteins and immune components, host vasculature, and biofilm substratum. Inclusion of these variables in the model system allows us to account for the hostpathogen interaction and examine its impact on the transcriptional profile of Candida during biofilm development. The importance of the host-pathogen interaction has been previously demonstrated by use of in vivo model system in both the host and pathogen transcriptomes [12]. The current study demonstrates the relevance and importance of incorporating the host infection site and immune system for the biofilm process. METHODS In vivo venous catheter biofilm formation. A C. albicans central venous catheter biofilm model was used for in vivo experiments as previously described [11, 13, 14]. After catheter placement for 24 h, a C. albicans SC5314 inoculum of 10 6 cells/ ml in a sodium chloride concentration of 0.15 mol/l was instilled in the catheter. Catheters were removed for RNA isolation or imaging after a 12- or 24-h incubation period. To yield a sufficient amount of RNA for the microarray analysis, Candida cells from 2 5 catheters were pooled. Collected cells were flash frozen with liquid nitrogen in AE buffer (sodium acetate concentration, 50 mmol/l; ph 5.2; EDTA concentration, 10 mmol/l) before RNA processing as previously described [11]. RNA was collected using the hot method as previously described [11]. The RNA integrity was assessed using Agilent Bioanalyzer 2100 with an RNA nanochip before use in microarrays. Planktonic culture conditions. C. albicans SC5314 was propagated overnight in YPD medium at 37 C on an orbital shaker at 250 rpm and used for inoculation of planktonic cultures. Planktonic cultures were obtained in parallel with in vivo cultures by incubated the cells in 20 ml of RPMI in a glass flask at 37 C on an orbital shaker at 250 rpm. Log-phase planktonic cells were harvested by centrifugation, and the cells were flash frozen with liquid nitrogen in AE buffer until RNA processing. RNA was collected and its integrity investigated as described above. Biofilm scanning electron microscopy (SEM). Catheters were harvested at each time point for imaging as previously described [11]. Samples were imaged in a JEOL JSM-6100 in the high-vacuum mode at 10 kv. The images were assembled using Adobe Photoshop 7.0. Microarray and reverse transcription polymerase chain reaction (RT-PCR). C. albicans oligo microarray slides were purchased from the Biotechnology Research Institute, National Research Council, Montreal (Version Oligo Trial 2). Three biologic replicates and dye swaps were performed for each condition and time point as previously described [12, 15, 16]. The microarrays were scanned with an Agilent DNA microarray scanner (G2565BA). Lowess normalization and statistical analysis were performed using Genespring v.7 (Agilent Technologies). Genes whose expression was either increased 1.5-fold (log 2 ) or decreased 0.67-fold for C. albicans biofilm samples relative to planktonic samples were considered for further analysis. A 2-sided Student t test was used determine the statistical significance of the log ratios. Quantitative real-time RT-PCR was used to confirm messenger RNA abundances for a subset of up-regulated transcripts from the microarray studies. The TaqMan probe and primer sets were designed using Primer Express (Applied Biosystems) (table 1). The QuantiTect probe RT-PCR kit (Qiagen) was used in an ABI Prism 7700 v1.7 sequence detection system (Applied Biosystems) as previously described [11]. Reactions were performed in triplicate. The quantitative data analysis was completed using the C t (2 C t) method [17]. The comparative expression method generated data as transcript fold-change normalized to a constitutive reference gene transcript (ACT1) and relative to a control or baseline condition. The statistical significance of differences in expression between biofilm and planktonic cells was analyzed by analysis of variance. RESULTS In vivo Candida biofilm model. Images of the intermediate biofilm (12 h) demonstrated yeast and hyphal cells attached to the substratum. At 24 h, the mature biofilm consisted of an extensive network of yeast and hyphae embedded in strands of heterogeneous matrix material (figure 1). Host cells, including red and white blood cells, were also embedded in the Candida biofilm. Transcript profiles of intermediate and mature in vivo Candida biofilms. The genome-wide changes in gene expression 308 JID 2009:200 (15 July) Nett et al.
3 Figure 1. Scanning electron micrograph of Candida albicans in vivo biofilm (original magnification 1000). C. albicans SC5314 was inoculated into a rat venous catheter and allowed to dwell for 24 h. Catheters were placed in a fixative, treated with osmium tetroxide, and dehydrated through a series of ethanol washes and critical point drying. in intermediate and mature Candida biofilms are shown in tables 1 and 2. From the intermediate biofilm and planktonic comparison, we identified 545 transcripts (8.6%) of the 6354 ORFs as differentially regulated at least 1.5-fold (table 2). Within this group, 457 transcripts were up-regulated during biofilm formation (biofilm:planktonic ratio, 11.5) and 88 transcripts were down-regulated (biofilm:planktonic ratio,!0.67). In the comparison of mature (24 h) biofilm cells and planktonic cells, 1034 transcripts (16.3%) were differentially regulated (table 2). A total of 523 transcripts were up-regulated during mature biofilm formation, and 511 transcripts were downregulated. Of the genes differentially expressed in intermediate biofilm formation, multiple functional categories were represented (table 2). During intermediate biofilm development, transcripts involved in protein synthesis (17%), transport (5%), stress (4%), amino acid metabolism (4%), cell wall metabolism (3%), energy and general metabolism (9%), and carbohydrate processing (7%) were abundant. The functions of 44% of the genes with transcriptional up-regulation in intermediate biofilm formation have not yet been identified. Transcripts reduced during intermediate biofilm formation were primarily involved with DNA processing and cell cycle (19%), although changes in transcription and protein synthesis (10%), transport (5%), cytoskeleton (5%), and carbohydrate processing (6%) were also evident. In the mature biofilm, we noted similar changes in gene regulation (table 2). Again, multiple functional categories were represented. Transcription of genes associated with protein processing (17%), energy and metabolism (8%), transport (5%), DNA/cell cycle (5%), and carbohydrate processing (5%) was up-regulated in the mature biofilm cells. Similar to the intermediate biofilm, mature biofilm formation was associated with reduction of transcripts involved in DNA processing and cell cycle (7%), transcription and protein synthesis (13%), and transport (7%). The functions of nearly 40% of the differentially regulated genes are not yet known. Next, we identified transcript abundance in both the intermediate biofilm and mature biofilm cells as a marker for relevance throughout the majority of biofilm formation. Transcript abundance of 124 genes was found at both the 12- and 24-h time points, compared with cells growing in planktonic conditions (table 3). Genes involved in carbohydrate synthesis and processing (10%), transcription and protein synthesis (13%), and energy and metabolism (12%) were among those highly expressed at both time points. Transcripts involved in cell wall metabolism (4%), iron metabolism (3%) and amino acid metabolism (5%) were also abundant. Transcription of Gene Expression Analysis of In Vivo Candida Biofilm JID 2009:200 (15 July) 309
4 Table 2. Frequency distribution of differentially expressed genes among functional categories. only 27 genes was down-regulated in both the intermediate and mature biofilms (table 3). Genes involved in DNA processing (30%) were most represented among the reduced transcripts. Validation of microarray data by real-time RT-PCR. Thirteen genes of interest were chosen to include several functional categories, including those involved in adherence, drug resistance, ergosterol synthesis, transport, amino acid metabolism, cell wall synthesis, and iron metabolism. We found that the expression direction for each gene chosen was consistent with the array data (table 4). DISCUSSION Nearly all device-associated infections are associated with organisms growing as biofilms [1, 2, 18]. Most of our knowledge about Candida biofilms is based on in vitro studies, but such systems cannot completely simulate the environment of the infection site and host immune system. The in vivo central venous catheter model was developed to incorporate these experimental variables and has been shown to be useful for the investigation of Candida biofilm development, diagnosis, drug treatment and resistance, and study of the role of specific individual gene products [11, 13, 19, 20]. The current investigation expands the examination of the molecular basis for biofilm formation by assessing the temporal global transcript profile during C. albicans biofilm development. These experiments identified differential regulation of many genes previously shown important in biofilm formation in vitro [4, 21 23]. Also, a subset of differentially expressed transcripts in the current in vivo study has been previously identified in genomic investigation of the planktonic Candida response to the mammalian host and specific host cells [12, 24]. Among the similarities to prior biofilm study was the up-regulation of genes involved in adherence (ALS1 andals4), cell wall metabolism (ECE1 and SAP5), carbohydrate and general metabolism (CGT, ICL1, MLS1, PCK1, and PDK1), and hyphal formation (HWP2). Noteworthy, in vivo biofilm formation involved differential regulation of the glyoxylate cycle, which allows organisms to use 2 carbon molecules as an energy source. Transcripts for 2 enzymes of the glyoxylate cycle, MSL1 and ICL1, encoding malate synthase and isocitrate lyase, were 2.4- and 2.3-fold more abundant in the mature biofilm. Increased expression of these genes has been described upon inoculation and infection of animal models in vivo and by phagocytosis [24, 25]. Mature biofilms are heterogeneous, with cells presumably exposed to gradients of nutrients. Biofilm cells closest to the catheter surface may experience a decreased supply of glucose. Increased expression of the glyoxylate cycle potentially allows the cells to use additional carbon sources. The current analysis identified several expression differences between the 12- and 24-h biofilm time points. These differences may be accounted for by the stepwise morphologic and architectural changes during biofilm development or perhaps by a change in metabolic state or a quorum sensing process. Initial biofilm formation requires the attachment of yeast cells to a substratum [26, 27]. As expected, expression of a greater number and intensity of adherence genes was observed at the earlier 12-h time point. ALS (agglutinin-like sequence) genes are a family of adhesins recognized to play a role in adherence and early biofilm formation [21, 28, 29]. In the current studies, transcript abundance of ALS1 (12-fold) and ALS2 (4-fold) was present only at the earlier time point. The current study did not observe a transcript abundance of ALS3. This result supports the hypothesis that adhesins may have overlapping functions in biofilm formation in vivo. Another possibility is that ALS3 may have been up-regulated at a time point other than those examined in the current microarray analysis. Upon initial cell adherence to a solid surface and the instigation of biofilm formation, genes involved in amino acid biosynthesis have been shown to be differentially regulated in vitro [7, 30]. Pathways involved in sulfur metabolism and regulation of methionine and cysteine biosynthesis are among those upregulated as early as 30 min after adhesion. In the current study, increased expression of MET3 (1.6-fold), MET10 (1.7-fold), CYS3 (3-fold), andcys4 (1.7-fold) was detected in the intermediate biofilm. The specific role of these pathways in early biofilm formation has not yet been elucidated. MET3 is a primary activator of sulfur assimilation in planktonic cells and is generally repressed in the presence of extracellular cysteine and methionine. However, transcript abundance of MET3 has been identified in biofilm systems containing methionine and cysteine concentrations greater than that normally needed for repression [30]. Although it is possible that amino acids are limited within microenvironments in a heterogeneous biofilm, the swift up-regulation of these pathways upon adherence to a solid surface suggests a role in sensing and responding to the surface [30]. In contrast to the intermediate biofilm, changes in expression of genes involved in methionine and cysteine biosynthesis were not predominant in the mature phase of biofilm growth. Instead, abundance of transcripts encoding amino acid 310 JID 2009:200 (15 July) Nett et al.
5 Table 3. biofilms. Transcripts differentially regulated in both intermediate and mature permeases (DAO2, DIP5, GAP6, and GNP1) was noted [31, 32]. Mature Candida biofilms are marked by a basal layer of yeast cells with subsequent layering of filamentous morphotypes and extensive matrix production [33]. Transition to the hyphal morphology intuitively requires differential regulation of hyphalassociated genes [20, 34]. The current studies identified expression of a number of hyphal-associated genes in both the intermediate biofilm (15 genes) and the mature biofilm (40 genes). Furthermore, transcription of TEC1, a transcription factor involved in hyphal morphogenesis, was up-regulated (2.4-fold) in the mature biofilm [35]. Of note, we also identified down-regulation (2.5-fold) of TUP1, a negative regulator of hyphal formation [9, 36]. Recent studies have described quorum sensing molecules in Candida organisms similar to those described in bacterial biofilm systems [37, 38]. Farnesol acts to inhibit formation of biofilm, whereas tyrosol induces replication and hyphal morphogenesis at low cell densities [39, 40]. CHK1, a histidine kinase gene, appears to be a component of the farnesol quorum sensing pathway because the chk1/chk1 mutant is unresponsive to the action of farnesol [41]. In the current study, the CHK1 transcript was abundant (1.8-fold) in the mature biofilm but not the intermediate biofilm. This timing is consistent with CHK1 involvement in the regulation of mature biofilms and, perhaps, not surprising, given the much greater burden of organisms in the milieu at this later phase of development. A recent microarray analysis compared the transcriptome of a 24- h mature biofilm exposed to farnesol with an unexposed biofilm and identified 274 differentially regulated transcripts [9]. Genes involved in hyphal morphology, drug resistance, and cell wall maintenance were highly represented. Sixty-nine of these farnesol-responsive genes were differentially regulated in the mature biofilm of the current study (table 3) [9]. We also identified altered regulation of ergosterol and b- glucan pathways associated with in vivo biofilm growth. Similar changes in gene regulation have been recently described in the most basal layer of in vitro biofilm cells, blastospores [23]. When compared with the current studies, both identified increased transcripts of ERG25 and b-1,6 glucan synthesis genes KRE1 and SKN1 [25]. ERG25 is putative C-4 methyl sterol oxidase with a role in C4-demethylation of ergosterol biosynthesis intermediates. It has been proposed that this up-regulation may allow for increased conversion of lanosterol to nonergosterol intermediates, including eburicol and 14-methyl fecosterol, at the expense of conversion to ergosterol [23, 42]. Additionally, altered expression of the b-1,3 glucan synthesis and modification pathways (FKS1, BGL2, XOG1, PHR2, FEN1, GDB1, andsga1) was prominent in the current study [43, 44]. It has been hypothesized that the glucan pathway may restructure the biofilm cell wall and contribute to the drug-resistant phenotype [23]. The b-glucan pathway may also play a role in biofilm matrix production. Formation of a mature biofilm requires production of an extracellular polymeric matrix, which is composed primarily of carbohydrate, glucose, and protein with smaller amounts of hexosamine and phosphorus [3, 18]. The presence of glucan molecules in secreted Candida biofilm material and their possible role in biofilm resistance has recently been reported [45]. A number of transcripts involved in carbohydrate processing and synthesis [44] and cell wall metabolism [40] were differentially expressed in the mature biofilm. These gene products may participate in production of matrix material. Interestingly, transcripts involved in the b-1,3 glucan degradation pathway (ENG1 and SWC1) were reduced at the 24-h time point. Both gene products, ENG1 and SWC1, have b-1,3 glucosidase activity. Disruption of CaENG1 results in decreased extracellular b-1,3 glucanase activity [8]. It is possible that down-regulation of these b-1,3 glucosidases and altered regulation of the b-1,3 glucan pathway may serve to conserve glucans for construction of a mature biofilm matrix. Many factors have been proposed to contribute to Candida biofilm drug resistance, including up-regulation of efflux pumps, decreased perfusion of antimicrobials through the matrix, slow growth, and alterations in plasma membrane ergosterol content [46, 47]. Early biofilm resistance coincides with increased transcript levels of the efflux pump genes MDR1 and CDR1 [48]. The current studies identified transcript up-regulation of CDR2 at 12 h (1.5-fold) and MDR1 at both 12 h (2.1- fold) and 24 h (1.9-fold). PDR16 transcript abundance was also noted in both the intermediate biofilm (2.2-fold) and mature biofilm (3.7-fold). PDR16 encodes a phosphatidylinositol trans- Table 4. Comparison of RT-PCR and microarray results for genes of interest. Gene Expression Analysis of In Vivo Candida Biofilm JID 2009:200 (15 July) 311
6 fer protein of the Sec14p family and is up-regulated in fluconazole-resistant cells overexpressing CDR1 and CDR2 [49]. Three prior studies have examined the global transcriptional response during Candida biofilm development [7, 9, 30]. Similar to the in vitro models, the current study found differential regulation of numerous amino acid synthesis genes in both the intermediate biofilms (18 genes) and mature biofilms (21 genes), as well as transcripts involved in sulfur metabolism (Met3 and Met10) (table 3). Analysis of the in vivo data set also identified differences from the in vitro transcriptional studies. For example, transcripts of genes encoding glucose transporters were among those most abundant at both time points. Increased expression of both HGT1 (5.4-fold) and HGT2 (52- fold) was observed in intermediate biofilms. Mature biofilm formation was associated with transcriptional abundance of HGT1 (4.7-fold), HGT2 (7.6-fold), HGT14 (2.2-fold), HGT15 (5-fold), and HGT19 (4.5-fold). One possible explanation for this expression is the need for scavenging glucose for nutrientstarved biofilm cells. Another possibility is a glucose requirement for production of the carbohydrate matrix. A sizable subset of genes was differentially regulated in both the intermediate (8.5%) and mature (16%) biofilms. Genes involved in amino acid metabolism (AGP2, ARG1, CAN1, CDG1, CPA1, CPA2, andsmm1), cell wall metabolism (BGL2 and ECE1), iron metabolism (CFL2, FRE10, ISU1, andsit1), and drug resistance (ERG25, PDR16, andmdr1) were among those abundant during the time periods examined (table 3). We suspect these gene products may play a role in the biofilm phenotypes exhibited throughout the biofilm lifecycle, including drug resistance, cell wall changes, altered metabolism, and maintaining cell attachment. Of note, the study did not identify differential expression of several genes of demonstrated importance in biofilm formation (ADH1, BCR1, CPH1, EGF1, MKC1, and YWP1) [19, 21, 34, 50]. The timing of expression of these genes may not correlate with the time points examined in our study. For example, BCR1 and ALS3 may play a role in earlier biofilm formation. Also, the magnitude of the differences in transcript level may be below that detectable by our microarray analysis or regulation may occur at the posttranscriptional level. The absence of these genes in the current data set points to the importance of time course experimentation. The identification of transcripts unique to the current study demonstrates the value of in vivo models that more closely mimic the disease state. The data set provides a resource for laboratories to guide directed biofilm investigations and target specific gene products. References 1. Kojic EM, Darouiche RO. Candida infections of medical devices. Clin Microbiol Rev 2004; 17: Douglas LJ. Candida biofilms and their role in infection. Trends Microbiol 2003; 11: Baillie GS, Douglas LJ. Matrix polymers of Candida biofilms and their possible role in biofilm resistance to antifungal agents. J Antimicrob Chemother 2000; 46: Mukherjee PK, Chandra J, Kuhn DM, Ghannoum MA. Mechanism of fluconazole resistance in Candida albicans biofilms: phase-specific role of efflux pumps and membrane sterols. Infect Immun 2003;71: Ramage G, Bachmann S, Patterson TF, Wickes BL, Lopez-Ribot JL. Investigation of multidrug efflux pumps in relation to fluconazole resistance in Candida albicans biofilms. J Antimicrob Chemother 2002; 49: Chandra J, Kuhn DM, Mukherjee PK, Hoyer LL, McCormick T, Ghannoum MA. Biofilm formation by the fungal pathogen Candida albicans: development, architecture, and drug resistance. J Bacteriol 2001; 183: Garcia-Sanchez S, Aubert S, Iraqui I, Janbon G, Ghigo JM, d Enfert C. Candida albicans biofilms: a developmental state associated with specific and stable gene expression patterns. Eukaryot Cell 2004;3: Esteban PF, Rios I, Garcia R, et al. Characterization of the CaENG1 gene encoding an endo-1,3-beta-glucanase involved in cell separation in Candida albicans. Curr Microbiol 2005; 51: Cao YY, Cao YB, Xu Z, et al. cdna microarray analysis of differential gene expression in Candida albicans biofilm exposed to farnesol. Antimicrob Agents Chemother 2005; 49: Nett J, Andes D. Candida albicans biofilm development, modeling a host-pathogen interaction. Curr Opin Microbiol 2006; 9: Andes D, Nett J, Oschel P, Albrecht R, Marchillo K, Pitula A. Development and characterization of an in vivo central venous catheter Candida albicans biofilm model. Infect Immun 2004; 72: Andes D, Lepak A, Pitula A, Marchillo K, Clark J. A simple approach for estimating gene expression in Candida albicans directly from a systemic infection site. J Infect Dis 2005; 192: Nett J, Lincoln L, Marchillo K, Andes D. b -1,3 glucan as a test for central venous catheter biofilm infection. J Infect Dis 2007;195: Nett J, Lincoln L, Marchillo K, et al. Putative role of beta-1,3 glucans in Candida albicans biofilm resistance. Antimicrob Agents Chemother 2007; 51: Lepak A, Nett J, Lincoln L, Marchillo K, Andes D. Time course of microbiologic outcome and gene expression in Candida albicans during and following in vitro and in vivo exposure to fluconazole. Antimicrob Agents Chemother 2006; 50: Nobile CJ, Solis N, Myers CL, et al. Candida albicans transcription factor Rim101 mediates pathogenic interactions through cell wall functions. Cell Microbiol 2008; 10: Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2 DDCT method. Methods 2001; 25: Donlan RM. Biofilm formation: a clinically relevant microbiological process. Clin Infect Dis 2001; 33: Mukherjee PK, Mohamed S, Chandra J, et al. Alcohol dehydrogenase restricts the ability of the pathogen Candida albicans to form a biofilm on catheter surfaces through an ethanol-based mechanism. Infect Immun 2006; 74: Nobile CJ, Nett JE, Andes DR, Mitchell AP. Function of Candida albicans adhesin Hwp1 in biofilm formation. Eukaryot Cell 2006;5: Nobile CJ, Andes DR, Nett JE, et al. Critical role of Bcr1-dependent adhesins in C. albicans biofilm formation in vitro and in vivo. PLoS Pathog 2006; 2:e Zhao X, Oh SH, Yeater KM, Hoyer LL. Analysis of the Candida albicans Als2p and Als4p adhesins suggests the potential for compensatory function within the Als family. Microbiology 2005; 151: Khot PD, Suci PA, Miller RL, Nelson RD, Tyler BJ. A small subpop- 312 JID 2009:200 (15 July) Nett et al.
7 ulation of blastospores in Candida albicans biofilms exhibit resistance to amphotericin B associated with differential regulation of ergosterol and beta-1,6-glucan pathway genes. Antimicrob Agents Chemother 2006; 50: Fradin C, Kretschmar M, Nichterlein T, Gaillardin C, d Enfert C, Hube B. Stage-specific gene expression of Candida albicans in human blood. Mol Microbiol 2003; 47: Lorenz MC, Fink GR. The glyoxylate cycle is required for fungal virulence. Nature 2001; 412: Kumamoto CA, Vinces MD. Alternative Candida albicans lifestyles: growth on surfaces. Annu Rev Microbiol 2005; 59: Hawser SP, Douglas LJ. Biofilm formation by Candida species on the surface of catheter materials in vitro. Infect Immun 1994; 62: Richard ML, Nobile CJ, Bruno VM, Mitchell AP. Candida albicans biofilm-defective mutants. Eukaryot Cell 2005; 4: Nobile CJ, Schneider HA, Nett JE, et al. Complementary adhesin function in C. albicans biofilm formation. Curr Biol 2008; 18: Murillo LA, Newport G, Lan CY, Habelitz S, Dungan J, Agabian NM. Genome-wide transcription profiling of the early phase of biofilm formation by Candida albicans. Eukaryot Cell 2005; 4: Regenberg B, Holmberg S, Olsen LD, Kielland-Brandt MC. Dip5p mediates high-affinity and high-capacity transport of L-glutamate and L-aspartate in Saccharomyces cerevisiae. Curr Genet 1998; 33: Regenberg B, During-Olsen L, Kielland-Brandt MC, Holmberg S. Substrate specificity and gene expression of the amino-acid permeases in Saccharomyces cerevisiae. CurrGenet1999; 36: Hawser SP, Baillie GS, Douglas LJ. Production of extracellular matrix by Candida albicans biofilms. J Med Microbiol 1998; 47: Ramage G, VandeWalle K, Lopez-Ribot JL, Wickes BL. The filamentation pathway controlled by the Efg1 regulator protein is required for normal biofilm formation and development in Candida albicans.fems Microbiol Lett 2002; 214: Schweizer A, Rupp S, Taylor BN, Rollinghoff M, Schroppel K. The TEA/ATTS transcription factor CaTec1p regulates hyphal development and virulence in Candida albicans. Mol Microbiol 2000; 38: Braun BR, Johnson AD. Control of filament formation in Candida albicans by the transcriptional repressor TUP1. Science 1997; 277: Hornby JM, Jensen EC, Lisec AD, et al. Quorum sensing in the dimorphic fungus Candida albicans is mediated by farnesol. Appl Environ Microbiol 2001; 67: Oh KB, Miyazawa H, Naito T, Matsuoka H. Purification and characterization of an autoregulatory substance capable of regulating the morphological transition incandida albicans. Proc Natl Acad Sci U S A 2001; 98: Ramage G, Saville SP, Wickes BL, Lopez-Ribot JL. Inhibition of Candida albicans biofilm formation by farnesol, a quorum-sensing molecule. Appl Environ Microbiol 2002; 68: Chen H, Fujita M, Feng Q, Clardy J, Fink GR. Tyrosol is a quorumsensing molecule in Candida albicans. Proc Natl Acad Sci U S A 2004; 101: Kruppa M, Krom BP, Chauhan N, Bambach AV, Cihlar RL, Calderone RA. The two-component signal transduction protein Chk1p regulates quorum sensing in Candida albicans. Eukaryot Cell 2004; 3: Barker KS, Crisp S, Wiederhold N, et al. Genome-wide expression profiling reveals genes associated with amphotericin B and fluconazole resistance in experimentally induced antifungal resistant isolates of Candida albicans. J Antimicrob Chemother 2004; 54: Sarthy AV, McGonigal T, Coen M, Frost DJ, Meulbroek JA, Goldman RC. Phenotype in Candida albicans of a disruption of the BGL2 gene encoding a 1,3-beta-glucosyltransferase. Microbiology 1997; 143: Mio T, Adachi-Shimizu M, Tachibana Y, et al. Cloning of the Candida albicans homolog of Saccharomyces cerevisiae GSC1/FKS1 and its involvement in beta-1,3-glucan synthesis. J Bacteriol 1997; 179: Nett J, Lincoln L, Marchillo K, et al. Putative role of beta-1,3 glucans in Candida albicans biofilm resistance. Antimicrob Agents Chemother 2007; 51: Ramage G, Saville SP, Thomas DP, Lopez-Ribot JL. Candida biofilms: an update. Eukaryot Cell 2005; 4: Kuhn DM, Ghannoum MA. Candida biofilms: antifungal resistance and emerging therapeutic options. Curr Opin Investig Drugs 2004;5: Mateus C, Crow SA Jr, Ahearn DG. Adherence of Candida albicans to silicone induces immediate enhanced tolerance to fluconazole. Antimicrob Agents Chemother 2004; 48: Saidane S, Weber S, De Deken X, St-Germain G, Raymond M. PDR16- mediated azole resistance in Candida albicans. Mol Microbiol 2006; 60: Kumamoto CA. A contact-activated kinase signals Candida albicans invasive growth and biofilm development. Proc Natl Acad Sci U S A 2005; 102: Gene Expression Analysis of In Vivo Candida Biofilm JID 2009:200 (15 July) 313
NIH Public Access Author Manuscript Med Mycol. Author manuscript; available in PMC 2012 February 1.
 NIH Public Access Author Manuscript Published in final edited form as: Med Mycol. 2012 February ; 50(2): 214 218. doi:10.3109/13693786.2011.580016. Comparative analysis of Candida biofilm quantitation
NIH Public Access Author Manuscript Published in final edited form as: Med Mycol. 2012 February ; 50(2): 214 218. doi:10.3109/13693786.2011.580016. Comparative analysis of Candida biofilm quantitation
Genetic Basis of Candida Biofilm Resistance Due to Drug-Sequestering Matrix Glucan
 BRIEF REPORT Genetic Basis of Candida Biofilm Resistance Due to Drug-Sequestering Matrix Glucan Jeniel E. Nett, 1,3 Hiram Sanchez, 1 Michael T Cain, 1 and David R. Andes 1,2 Departments of 1 Medicine,
BRIEF REPORT Genetic Basis of Candida Biofilm Resistance Due to Drug-Sequestering Matrix Glucan Jeniel E. Nett, 1,3 Hiram Sanchez, 1 Michael T Cain, 1 and David R. Andes 1,2 Departments of 1 Medicine,
Candida albicans biofilm development, modeling a host pathogen interaction Jeniel Nett and David Andes
 Candida albicans biofilm development, modeling a host pathogen interaction Jeniel Nett and David Andes Medical device-associated infections involve the attachment of cells to a surface, production of an
Candida albicans biofilm development, modeling a host pathogen interaction Jeniel Nett and David Andes Medical device-associated infections involve the attachment of cells to a surface, production of an
Optimizing a Candida Biofilm Microtiter Plate Model for Measurement of Antifungal Susceptibility by Tetrazolium Salt Assay
 JOURNAL OF CLINICAL MICROBIOLOGY, Apr. 2011, p. 1426 1433 Vol. 49, No. 4 0095-1137/11/$12.00 doi:10.1128/jcm.02273-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Optimizing
JOURNAL OF CLINICAL MICROBIOLOGY, Apr. 2011, p. 1426 1433 Vol. 49, No. 4 0095-1137/11/$12.00 doi:10.1128/jcm.02273-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Optimizing
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
A Candida Biofilm-Induced Pathway for Matrix Glucan Delivery: Implications for Drug Resistance
 A Candida Biofilm-Induced Pathway for Matrix Glucan Delivery: Implications for Drug Resistance Heather T. Taff 1, Jeniel E. Nett 1, Robert Zarnowski 1, Kelly M. Ross 1, Hiram Sanchez 1, Mike T. Cain 1,
A Candida Biofilm-Induced Pathway for Matrix Glucan Delivery: Implications for Drug Resistance Heather T. Taff 1, Jeniel E. Nett 1, Robert Zarnowski 1, Kelly M. Ross 1, Hiram Sanchez 1, Mike T. Cain 1,
An expanded regulatory network temporally controls Candida albicans biofilm formation
 Molecular Microbiology (2015) 96(6), 1226 1239 doi:10.1111/mmi.13002 First published online 23 April 2015 An expanded regulatory network temporally controls Candida albicans biofilm formation Emily P.
Molecular Microbiology (2015) 96(6), 1226 1239 doi:10.1111/mmi.13002 First published online 23 April 2015 An expanded regulatory network temporally controls Candida albicans biofilm formation Emily P.
Fungal biofilms. Research CMU. Carnegie Mellon University. Saranna Fanning Carnegie Mellon University
 Carnegie Mellon University Research Showcase @ CMU Department of Biological Sciences Mellon College of Science 1-1-2012 Fungal biofilms. Saranna Fanning Carnegie Mellon University Aaron P. Mitchell Carnegie
Carnegie Mellon University Research Showcase @ CMU Department of Biological Sciences Mellon College of Science 1-1-2012 Fungal biofilms. Saranna Fanning Carnegie Mellon University Aaron P. Mitchell Carnegie
AP Biology Gene Expression/Biotechnology REVIEW
 AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Recombinant DNA Technology. The Role of Recombinant DNA Technology in Biotechnology. yeast. Biotechnology. Recombinant DNA technology.
 PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
cdna Microarray Analysis of Differential Gene Expression in Candida albicans Biofilm Exposed to Farnesol
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Feb. 2005, p. 584 589 Vol. 49, No. 2 0066-4804/05/$08.00 0 doi:10.1128/aac.49.2.584 589.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved.
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Feb. 2005, p. 584 589 Vol. 49, No. 2 0066-4804/05/$08.00 0 doi:10.1128/aac.49.2.584 589.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved.
Antifungal PK/PD Made Simple. David Andes, MD University of Wisconsin
 Antifungal PK/PD Made Simple David Andes, MD University of Wisconsin PK/PD Concept Serum Drug Concentration Peak:MIC AUC:MIC Ratio Time Above MIC MIC Time Hypothesis and Concept There is an optimal drug
Antifungal PK/PD Made Simple David Andes, MD University of Wisconsin PK/PD Concept Serum Drug Concentration Peak:MIC AUC:MIC Ratio Time Above MIC MIC Time Hypothesis and Concept There is an optimal drug
Molecular Cell Biology - Problem Drill 11: Recombinant DNA
 Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Molecular Cell Biology - Problem Drill 11: Recombinant DNA Question No. 1 of 10 1. Which of the following statements about the sources of DNA used for molecular cloning is correct? Question #1 (A) cdna
Microarray Technique. Some background. M. Nath
 Microarray Technique Some background M. Nath Outline Introduction Spotting Array Technique GeneChip Technique Data analysis Applications Conclusion Now Blind Guess? Functional Pathway Microarray Technique
Microarray Technique Some background M. Nath Outline Introduction Spotting Array Technique GeneChip Technique Data analysis Applications Conclusion Now Blind Guess? Functional Pathway Microarray Technique
CHAPTER 20 DNA TECHNOLOGY AND GENOMICS. Section A: DNA Cloning
 Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
HiPer RT-PCR Teaching Kit
 HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
Case. Case. Case. Case. Reference lab AST. Nelesh Govender, NICD 2013/03/08. Candida species: Antifungal susceptibility testing in 2013
 Nelesh Govender, NICD 13/3/8 se ndida species: Antifungal susceptibility testing in 13 Nelesh Govender National Institute for Communicable Diseases and University of the Witwatersrand, Johannesburg Elderly
Nelesh Govender, NICD 13/3/8 se ndida species: Antifungal susceptibility testing in 13 Nelesh Govender National Institute for Communicable Diseases and University of the Witwatersrand, Johannesburg Elderly
Lecture #1. Introduction to microarray technology
 Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
Phenotype switching affects biofilm formation by Candida parapsilosis
 Microbiology (2005), 151, 1073 1081 DOI 10.1099/mic.0.27739-0 Phenotype switching affects biofilm formation by Candida parapsilosis Sean F. Laffey and Geraldine Butler Correspondence Geraldine Butler Geraldine.Butler@ucd.ie
Microbiology (2005), 151, 1073 1081 DOI 10.1099/mic.0.27739-0 Phenotype switching affects biofilm formation by Candida parapsilosis Sean F. Laffey and Geraldine Butler Correspondence Geraldine Butler Geraldine.Butler@ucd.ie
SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes
 WHITE PAPER SuperScript IV Reverse Transcriptase SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes Abstract Reverse transcriptases (RTs) from avian myeloblastosis virus
WHITE PAPER SuperScript IV Reverse Transcriptase SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes Abstract Reverse transcriptases (RTs) from avian myeloblastosis virus
Use of Drosophila Melanogaster as a Model System in the Study of Human Sodium- Dependent Multivitamin Transporter. Michael Brinton BIOL 230W.
 Use of Drosophila Melanogaster as a Model System in the Study of Human Sodium- Dependent Multivitamin Transporter Michael Brinton BIOL 230W.001 28 October 2013 TA: Sashi Gollapudi Introduction Many human
Use of Drosophila Melanogaster as a Model System in the Study of Human Sodium- Dependent Multivitamin Transporter Michael Brinton BIOL 230W.001 28 October 2013 TA: Sashi Gollapudi Introduction Many human
Penetration of Candida Biofilms by Antifungal Agents
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Sept. 2004, p. 3291 3297 Vol. 48, No. 9 0066-4804/04/$08.00 0 DOI: 10.1128/AAC.48.9.3291 3297.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved.
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Sept. 2004, p. 3291 3297 Vol. 48, No. 9 0066-4804/04/$08.00 0 DOI: 10.1128/AAC.48.9.3291 3297.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved.
Goals of pharmacogenomics
 Goals of pharmacogenomics Use drugs better and use better drugs! People inherit/exhibit differences in drug: Absorption Metabolism and degradation of the drug Transport of drug to the target molecule Excretion
Goals of pharmacogenomics Use drugs better and use better drugs! People inherit/exhibit differences in drug: Absorption Metabolism and degradation of the drug Transport of drug to the target molecule Excretion
2054, Chap. 14, page 1
 2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
Transmission Electron Microscopic Study of Antibiotic Action on Klebsiella pneumoniae Biofilm
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Aug. 2002, p. 2679 2683 Vol. 46, No. 8 0066-4804/02/$04.00 0 DOI: 10.1128/AAC.46.8.2679 2683.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved.
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Aug. 2002, p. 2679 2683 Vol. 46, No. 8 0066-4804/02/$04.00 0 DOI: 10.1128/AAC.46.8.2679 2683.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved.
ESCMID Online Lecture Library. by author
 Eric DANNAOUI ESCMID Postgraduate Education Course 20-22 June 2013, Sibiu Antifungal susceptibility testing and detection of resistance: principles and practices Unité de Parasitologie-Mycologie, Laboratoire
Eric DANNAOUI ESCMID Postgraduate Education Course 20-22 June 2013, Sibiu Antifungal susceptibility testing and detection of resistance: principles and practices Unité de Parasitologie-Mycologie, Laboratoire
The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity
 Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
REGULATION OF GENE EXPRESSION
 REGULATION OF GENE EXPRESSION Each cell of a living organism contains thousands of genes. But all genes do not function at a time. Genes function according to requirements of the cell. Genes control the
REGULATION OF GENE EXPRESSION Each cell of a living organism contains thousands of genes. But all genes do not function at a time. Genes function according to requirements of the cell. Genes control the
Rat Indwelling Urinary Catheter Model of Candida albicans
 IAI Accepts, published online ahead of print on 2 September 2014 Infect. Immun. doi:10.1128/iai.02284-14 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 2 Rat Indwelling Urinary
IAI Accepts, published online ahead of print on 2 September 2014 Infect. Immun. doi:10.1128/iai.02284-14 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 2 Rat Indwelling Urinary
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 14: Microarray Some slides were adapted from Dr. Luke Huan (University of Kansas), Dr. Shaojie Zhang (University of Central Florida), and Dr. Dong Xu and
EECS730: Introduction to Bioinformatics Lecture 14: Microarray Some slides were adapted from Dr. Luke Huan (University of Kansas), Dr. Shaojie Zhang (University of Central Florida), and Dr. Dong Xu and
BIOTECHNOLOGY. Course Syllabus. Section A: Engineering Mathematics. Subject Code: BT. Course Structure. Engineering Mathematics. General Biotechnology
 BIOTECHNOLOGY Subject Code: BT Course Structure Sections/Units Section A Section B Unit 1 Unit 2 Unit 3 Unit 4 Unit 5 Unit 6 Unit 7 Section C Section D Section E Topics Engineering Mathematics General
BIOTECHNOLOGY Subject Code: BT Course Structure Sections/Units Section A Section B Unit 1 Unit 2 Unit 3 Unit 4 Unit 5 Unit 6 Unit 7 Section C Section D Section E Topics Engineering Mathematics General
Using Low Input of Poly (A) + RNA and Total RNA for Oligonucleotide Microarrays Application
 Using Low Input of Poly (A) + RNA and Total RNA for Oligonucleotide Microarrays Application Gene Expression Author Michelle M. Chen Agilent Technologies, Inc. 3500 Deer Creek Road, MS 25U-7 Palo Alto,
Using Low Input of Poly (A) + RNA and Total RNA for Oligonucleotide Microarrays Application Gene Expression Author Michelle M. Chen Agilent Technologies, Inc. 3500 Deer Creek Road, MS 25U-7 Palo Alto,
Biofilm Matrix Regulation by Candida albicans Zap1
 Biofilm Matrix Regulation by Candida albicans Zap1 Clarissa J. Nobile 1,2, Jeniel E. Nett 3, Aaron D. Hernday 2, Oliver R. Homann 2, Jean-Sebastien Deneault 4, Andre Nantel 4, David R. Andes 3, Alexander
Biofilm Matrix Regulation by Candida albicans Zap1 Clarissa J. Nobile 1,2, Jeniel E. Nett 3, Aaron D. Hernday 2, Oliver R. Homann 2, Jean-Sebastien Deneault 4, Andre Nantel 4, David R. Andes 3, Alexander
Learning Objectives :
 Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
TransIT -mrna Transfection Kit
 INTRODUCTION TransIT -mrna Transfection Kit is designed to transfect RNA into a broad range of cell types with minimal cellular toxicity. RNA delivery avoids transcriptional regulation effects by directly
INTRODUCTION TransIT -mrna Transfection Kit is designed to transfect RNA into a broad range of cell types with minimal cellular toxicity. RNA delivery avoids transcriptional regulation effects by directly
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION
 M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
Genetics Lecture 21 Recombinant DNA
 Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
disaccharides = two mono-s linked together e.g. lactose = glucose + galactose sucrose = glucose + fructose
 involved in the degradation of molecules found in animal cells membrane limited varies in shape and size contains acid hydrolases (phosphatase, nucleases, proteases, etc.), enzymes that work only at acid
involved in the degradation of molecules found in animal cells membrane limited varies in shape and size contains acid hydrolases (phosphatase, nucleases, proteases, etc.), enzymes that work only at acid
M Keramatipour 2. M Keramatipour 1. M Keramatipour 4. M Keramatipour 3. M Keramatipour 5. M Keramatipour
 Molecular Cloning Methods Mohammad Keramatipour MD, PhD keramatipour@tums.ac.ir Outline DNA recombinant technology DNA cloning co Cell based PCR PCR-based Some application of DNA cloning Genomic libraries
Molecular Cloning Methods Mohammad Keramatipour MD, PhD keramatipour@tums.ac.ir Outline DNA recombinant technology DNA cloning co Cell based PCR PCR-based Some application of DNA cloning Genomic libraries
Biofilm Protocol Optimization For Pseudomonas aeruginosa. Introduction. Materials and Methods. Culture Media, Incubation Time, and Biofilm Measurement
 Biofilm Protocol Optimization For Pseudomonas aeruginosa Culture Media, Incubation Time, and Biofilm Measurement Introduction In addition to the conventional arsenal of antibiotic resistance mechanisms
Biofilm Protocol Optimization For Pseudomonas aeruginosa Culture Media, Incubation Time, and Biofilm Measurement Introduction In addition to the conventional arsenal of antibiotic resistance mechanisms
Real-Time PCR Workshop Gene Expression. Applications Absolute and Relative Quantitation
 Real-Time PCR Workshop Gene Expression Applications Absolute and Relative Quantitation Absolute Quantitation Easy to understand the data, difficult to develop/qualify the standards Relative Quantitation
Real-Time PCR Workshop Gene Expression Applications Absolute and Relative Quantitation Absolute Quantitation Easy to understand the data, difficult to develop/qualify the standards Relative Quantitation
SIMS2003. Instructors:Rus Yukhananov, Alex Loguinov BWH, Harvard Medical School. Introduction to Microarray Technology.
 SIMS2003 Instructors:Rus Yukhananov, Alex Loguinov BWH, Harvard Medical School Introduction to Microarray Technology. Lecture 1 I. EXPERIMENTAL DETAILS II. ARRAY CONSTRUCTION III. IMAGE ANALYSIS Lecture
SIMS2003 Instructors:Rus Yukhananov, Alex Loguinov BWH, Harvard Medical School Introduction to Microarray Technology. Lecture 1 I. EXPERIMENTAL DETAILS II. ARRAY CONSTRUCTION III. IMAGE ANALYSIS Lecture
SYBR Green Realtime PCR Master Mix
 Instruction manual SYBR Green Realtime PCR Master Mix 0810 F0924K SYBR Green Realtime PCR Master Mix QPK-201T 1 ml x 1 QPK-201 1 ml x 5 Contents [1] Introduction [2] Components [3] Primer design [4] Detection
Instruction manual SYBR Green Realtime PCR Master Mix 0810 F0924K SYBR Green Realtime PCR Master Mix QPK-201T 1 ml x 1 QPK-201 1 ml x 5 Contents [1] Introduction [2] Components [3] Primer design [4] Detection
The Role of Candida albicans Biofilms in Human Disease
 In: Candida albicans: Symptoms, Causes and Treatment Options ISBN: 978-1-62808-882-3 Editors: Leon A. Dietrich and Tim S. Friedmann 2013 Nova Science Publishers, Inc. The license for this PDF is unlimited
In: Candida albicans: Symptoms, Causes and Treatment Options ISBN: 978-1-62808-882-3 Editors: Leon A. Dietrich and Tim S. Friedmann 2013 Nova Science Publishers, Inc. The license for this PDF is unlimited
Introduction to BioMEMS & Medical Microdevices DNA Microarrays and Lab-on-a-Chip Methods
 Introduction to BioMEMS & Medical Microdevices DNA Microarrays and Lab-on-a-Chip Methods Companion lecture to the textbook: Fundamentals of BioMEMS and Medical Microdevices, by Prof., http://saliterman.umn.edu/
Introduction to BioMEMS & Medical Microdevices DNA Microarrays and Lab-on-a-Chip Methods Companion lecture to the textbook: Fundamentals of BioMEMS and Medical Microdevices, by Prof., http://saliterman.umn.edu/
ZYMOLYASE PROTOCOLS. 7. Spin 2 minutes in microfuge, pour super into a fresh tube and repeat spin. Remove 500 ul to a fresh tube.
 1 ZYMOLYASE PROTOCOLS Smash and Grab Zymolyase PROVIDED BY: DAVID AMBERG 1. Grow cells in 3mls selective media o/n 2. Pellet cells by 2 quick spins in a microfuge 3. Re-suspend cells in 200 u1 of the following
1 ZYMOLYASE PROTOCOLS Smash and Grab Zymolyase PROVIDED BY: DAVID AMBERG 1. Grow cells in 3mls selective media o/n 2. Pellet cells by 2 quick spins in a microfuge 3. Re-suspend cells in 200 u1 of the following
The Integrated Biomedical Sciences Graduate Program
 The Integrated Biomedical Sciences Graduate Program at the university of notre dame Cutting-edge biomedical research and training that transcends traditional departmental and disciplinary boundaries to
The Integrated Biomedical Sciences Graduate Program at the university of notre dame Cutting-edge biomedical research and training that transcends traditional departmental and disciplinary boundaries to
SI Results Peculiarities in the consumption of ammonium and glucose by AmtB- strains SI Materials and Methods Strain Construction
 SI Results Peculiarities in the consumption of ammonium and glucose by AmtB - strains. The AmtB - strain failed to consume all the ammonium in the medium under NH 3 -limiting conditions (.5 mm total ammonium
SI Results Peculiarities in the consumption of ammonium and glucose by AmtB - strains. The AmtB - strain failed to consume all the ammonium in the medium under NH 3 -limiting conditions (.5 mm total ammonium
How antimicrobial agents work
 Physical and Chemical Control of Microbes Physical Agents heat or radiation Chemical Agents disinfectants or antiseptics Important Terms 1. Sterilization process of killing all viable microbes 2. Bactericide
Physical and Chemical Control of Microbes Physical Agents heat or radiation Chemical Agents disinfectants or antiseptics Important Terms 1. Sterilization process of killing all viable microbes 2. Bactericide
PureSpin DNA Clean Up Kit
 PureSpin DNA Clean Up Kit Micro Columns INSTRUCTION MANUAL KIT COMPONENTS For Research Use Only PureSpin DNA Clean Up Kit, Micro Columns w/out Caps (Kit Size) OD2080 (50 Preps.) OD2080-2 (200 Preps.) Storage
PureSpin DNA Clean Up Kit Micro Columns INSTRUCTION MANUAL KIT COMPONENTS For Research Use Only PureSpin DNA Clean Up Kit, Micro Columns w/out Caps (Kit Size) OD2080 (50 Preps.) OD2080-2 (200 Preps.) Storage
Genetics Lecture Notes Lectures 17 19
 Genetics Lecture Notes 7.03 2005 Lectures 17 19 Lecture 17 Gene Regulation We are now going to look at ways that genetics can be used to study gene regulation. The issue is how cells adjust the expression
Genetics Lecture Notes 7.03 2005 Lectures 17 19 Lecture 17 Gene Regulation We are now going to look at ways that genetics can be used to study gene regulation. The issue is how cells adjust the expression
Lecture 9 Controlling gene expression
 Lecture 9 Controlling gene expression BIOLOGY Campbell, Reece and Mitchell Chapter 18 334- (352-356) Every cell in your body contains the same number of genes approximately 35, 000 DNA is wound around
Lecture 9 Controlling gene expression BIOLOGY Campbell, Reece and Mitchell Chapter 18 334- (352-356) Every cell in your body contains the same number of genes approximately 35, 000 DNA is wound around
3.1.4 DNA Microarray Technology
 3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
AGRO/ANSC/BIO/GENE/HORT 305 Fall, 2016 Overview of Genetics Lecture outline (Chpt 1, Genetics by Brooker) #1
 AGRO/ANSC/BIO/GENE/HORT 305 Fall, 2016 Overview of Genetics Lecture outline (Chpt 1, Genetics by Brooker) #1 - Genetics: Progress from Mendel to DNA: Gregor Mendel, in the mid 19 th century provided the
AGRO/ANSC/BIO/GENE/HORT 305 Fall, 2016 Overview of Genetics Lecture outline (Chpt 1, Genetics by Brooker) #1 - Genetics: Progress from Mendel to DNA: Gregor Mendel, in the mid 19 th century provided the
CHAPTER 9 DNA Technologies
 CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
Computational Biology I LSM5191
 Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype)
 Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype) Question#1: One-Gene, One-Polypeptide The figure below shows the results of feeding trials with one auxotroph strain of Neurospora
Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype) Question#1: One-Gene, One-Polypeptide The figure below shows the results of feeding trials with one auxotroph strain of Neurospora
INTRODUCTION TO REVERSE TRANSCRIPTION PCR (RT-PCR) ABCF 2016 BecA-ILRI Hub, Nairobi 21 st September 2016 Roger Pelle Principal Scientist
 INTRODUCTION TO REVERSE TRANSCRIPTION PCR (RT-PCR) ABCF 2016 BecA-ILRI Hub, Nairobi 21 st September 2016 Roger Pelle Principal Scientist Objective of PCR To provide a solution to one of the most pressing
INTRODUCTION TO REVERSE TRANSCRIPTION PCR (RT-PCR) ABCF 2016 BecA-ILRI Hub, Nairobi 21 st September 2016 Roger Pelle Principal Scientist Objective of PCR To provide a solution to one of the most pressing
Technical Review. Real time PCR
 Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017
 Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Recent technology allow production of microarrays composed of 70-mers (essentially a hybrid of the two techniques)
 Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Chapter 18: Regulation of Gene Expression. 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer
 Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Use of Quantitative Scanning Electron Microscopy of S. aureus Biofilm Formation in vitro to Identify Strain and Implant Material Specificities
 Use of Quantitative Scanning Electron Microscopy of S. aureus Biofilm Formation in vitro to Identify Strain and Implant Material Specificities Werasak Sutipornpalangkul, MD, PhD 1, Kohei Nishitani, MD,
Use of Quantitative Scanning Electron Microscopy of S. aureus Biofilm Formation in vitro to Identify Strain and Implant Material Specificities Werasak Sutipornpalangkul, MD, PhD 1, Kohei Nishitani, MD,
Higher Human Biology Unit 1: Human Cells Pupils Learning Outcomes
 Higher Human Biology Unit 1: Human Cells Pupils Learning Outcomes 1.1 Division and Differentiation in Human Cells I can state that cellular differentiation is the process by which a cell develops more
Higher Human Biology Unit 1: Human Cells Pupils Learning Outcomes 1.1 Division and Differentiation in Human Cells I can state that cellular differentiation is the process by which a cell develops more
Impact of Retinoic acid induced-1 (Rai1) on Regulators of Metabolism and Adipogenesis
 Impact of Retinoic acid induced-1 (Rai1) on Regulators of Metabolism and Adipogenesis The mammalian system undergoes ~24 hour cycles known as circadian rhythms that temporally orchestrate metabolism, behavior,
Impact of Retinoic acid induced-1 (Rai1) on Regulators of Metabolism and Adipogenesis The mammalian system undergoes ~24 hour cycles known as circadian rhythms that temporally orchestrate metabolism, behavior,
Molecular and Cell Biology (MCB)
 Molecular and Cell Biology (MCB) Head of Department: Professor Michael Lynes Department Office: Room 104, Biology/Physics Building For major requirements, see the College of Liberal Arts and Sciences section
Molecular and Cell Biology (MCB) Head of Department: Professor Michael Lynes Department Office: Room 104, Biology/Physics Building For major requirements, see the College of Liberal Arts and Sciences section
Candida albicans ribonuclease P RNA (RPR1) gene. genesig Advanced Kit. 150 tests. Primerdesign Ltd. For general laboratory and research use only
 TM Primerdesign Ltd Candida albicans ribonuclease P RNA (RPR1) gene genesig Advanced Kit 150 tests For general laboratory and research use only 1 Introduction to Candida albicans Candida albicans is a
TM Primerdesign Ltd Candida albicans ribonuclease P RNA (RPR1) gene genesig Advanced Kit 150 tests For general laboratory and research use only 1 Introduction to Candida albicans Candida albicans is a
Cell Growth and DNA Extraction- Technion igem HS
 Growing Cells and DNA Extraction Goals 1. Become familiar with the process of growing bacteria 2. Get to know the DNA extraction process 3. Perform miniprep in the lab Keywords 1. Growth stages 6. Techniques
Growing Cells and DNA Extraction Goals 1. Become familiar with the process of growing bacteria 2. Get to know the DNA extraction process 3. Perform miniprep in the lab Keywords 1. Growth stages 6. Techniques
Microbial Biotechnology agustin krisna wardani
 Microbial Biotechnology agustin krisna wardani 1. The Structure of Microbes Microbes (microorganisms) are tiny organisms that are too small to be seen individually by the naked eye and must be viewed with
Microbial Biotechnology agustin krisna wardani 1. The Structure of Microbes Microbes (microorganisms) are tiny organisms that are too small to be seen individually by the naked eye and must be viewed with
CHAPTER 13 LECTURE SLIDES
 CHAPTER 13 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
CHAPTER 13 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
SureSilencing sirna Array Technology Overview
 SureSilencing sirna Array Technology Overview Pathway-Focused sirna-based RNA Interference Topics to be Covered Who is SuperArray? Brief Introduction to RNA Interference Challenges Facing RNA Interference
SureSilencing sirna Array Technology Overview Pathway-Focused sirna-based RNA Interference Topics to be Covered Who is SuperArray? Brief Introduction to RNA Interference Challenges Facing RNA Interference
Genome Biology and Biotechnology
 Genome Biology and Biotechnology 10. The proteome Prof. M. Zabeau Department of Plant Systems Biology Flanders Interuniversity Institute for Biotechnology (VIB) University of Gent International course
Genome Biology and Biotechnology 10. The proteome Prof. M. Zabeau Department of Plant Systems Biology Flanders Interuniversity Institute for Biotechnology (VIB) University of Gent International course
qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time
 qpcr qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time Differs from endpoint PCR gel on last cycle Used to determines relative amount of template
qpcr qpcr Quantitative PCR or Real-time PCR Gives a measurement of PCR product at end of each cycle real time Differs from endpoint PCR gel on last cycle Used to determines relative amount of template
ReverTra Ace qpcr RT Master Mix
 Instruction manual ReverTra Ace qpcr RT Master Mix 1203 F1173K ReverTra Ace qpcr RT Master Mix FSQ-201 200 reactions Store at -20 C Contents [1] Introduction [2] Components [3] Protocol 1. RNA template
Instruction manual ReverTra Ace qpcr RT Master Mix 1203 F1173K ReverTra Ace qpcr RT Master Mix FSQ-201 200 reactions Store at -20 C Contents [1] Introduction [2] Components [3] Protocol 1. RNA template
Bayesian Networks as framework for data integration
 Bayesian Networks as framework for data integration Jun Zhu, Ph. D. Department of Genomics and Genetic Sciences Icahn Institute of Genomics and Multiscale Biology Icahn Medical School at Mount Sinai New
Bayesian Networks as framework for data integration Jun Zhu, Ph. D. Department of Genomics and Genetic Sciences Icahn Institute of Genomics and Multiscale Biology Icahn Medical School at Mount Sinai New
Lecture 25 (11/15/17)
 Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Reliable and sensitive RT-qPCR analysis of whole-blood RNA samples
 APPLICATION NOTE SuperScript IV Reliable and sensitive RT-qPCR analysis of whole-blood RNA samples Introduction Most commercially available reverse transcription products do not perform well with difficult
APPLICATION NOTE SuperScript IV Reliable and sensitive RT-qPCR analysis of whole-blood RNA samples Introduction Most commercially available reverse transcription products do not perform well with difficult
A Discovery Laboratory Investigating Bacterial Gene Regulation
 Chapter 8 A Discovery Laboratory Investigating Bacterial Gene Regulation Robert Moss Wofford College 429 N. Church Street Spartanburg, SC 29307 mosssre@wofford.edu Bob Moss is an Associate Professor of
Chapter 8 A Discovery Laboratory Investigating Bacterial Gene Regulation Robert Moss Wofford College 429 N. Church Street Spartanburg, SC 29307 mosssre@wofford.edu Bob Moss is an Associate Professor of
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali
 Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
ATIP Avenir Program 2018 Young group leader
 ATIP Avenir Program 2018 Young group leader Important dates - October 17th (4 pm) 2017 : opening of the registrations online - November 23 th 2017: deadline for the online submission, the mailing of the
ATIP Avenir Program 2018 Young group leader Important dates - October 17th (4 pm) 2017 : opening of the registrations online - November 23 th 2017: deadline for the online submission, the mailing of the
Data Sheet. Hippo Pathway TEAD Reporter MCF7 Cell Line Catalog #: 60618
 Data Sheet Hippo Pathway TEAD Reporter MCF7 Cell Line Catalog #: 6618 Background The Hippo pathway regulates cell proliferation and cell death. It is activated by high cell density and cell stress to stop
Data Sheet Hippo Pathway TEAD Reporter MCF7 Cell Line Catalog #: 6618 Background The Hippo pathway regulates cell proliferation and cell death. It is activated by high cell density and cell stress to stop
Storage and Expression of Genetic Information
 Storage and Expression of Genetic Information 29. DNA structure, Replication and Repair ->Ch 25. DNA metabolism 30. RNA Structure, Synthesis and Processing ->Ch 26. RNA metabolism 31. Protein Synthesis
Storage and Expression of Genetic Information 29. DNA structure, Replication and Repair ->Ch 25. DNA metabolism 30. RNA Structure, Synthesis and Processing ->Ch 26. RNA metabolism 31. Protein Synthesis
Virginia Western Community College BIO 205 General Microbiology
 Prerequisites BIO 205 General Microbiology One year of college biology and one year of college chemistry or divisional approval; an ENG 111 placement recommendation, co-enrollment in ENF 3/ENG 111, or
Prerequisites BIO 205 General Microbiology One year of college biology and one year of college chemistry or divisional approval; an ENG 111 placement recommendation, co-enrollment in ENF 3/ENG 111, or
BA, BSc, and MSc Degree Examinations
 Examination Candidate Number: Desk Number: BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Bacterial pathogenesis Time Allowed: 2 hours Marking Scheme: Total marks available
Examination Candidate Number: Desk Number: BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Bacterial pathogenesis Time Allowed: 2 hours Marking Scheme: Total marks available
Blot: a spot or stain, especially of ink on paper.
 Blotting technique Blot: a spot or stain, especially of ink on paper. 2/27 In molecular biology and genetics, a blot is a method of transferring proteins, DNA or RNA, onto a carrier (for example, a nitrocellulose,pvdf
Blotting technique Blot: a spot or stain, especially of ink on paper. 2/27 In molecular biology and genetics, a blot is a method of transferring proteins, DNA or RNA, onto a carrier (for example, a nitrocellulose,pvdf
Evolutionary models of conflict and cooperation. May 13, 2010 Joao Xavier
 Evolutionary models of conflict and cooperation May 13, 2010 Joao Xavier Evolutionary cooperation is central to all life Why evolutionary cooperation is important Simple definitions: Evolution Fitness
Evolutionary models of conflict and cooperation May 13, 2010 Joao Xavier Evolutionary cooperation is central to all life Why evolutionary cooperation is important Simple definitions: Evolution Fitness
YeaStar Genomic DNA Kit Catalog No. D2002
 Instructions YeaStar Genomic DNA Kit Catalog No. D2002 Table of Contents General Information...... 1 Package Contents Ordering Information General Description...... 2 Protocol I.......... 2 Protocol II.
Instructions YeaStar Genomic DNA Kit Catalog No. D2002 Table of Contents General Information...... 1 Package Contents Ordering Information General Description...... 2 Protocol I.......... 2 Protocol II.
Lecture 3 Mutagens and Mutagenesis. 1. Mutagens A. Physical and Chemical mutagens B. Transposons and retrotransposons C. T-DNA
 Lecture 3 Mutagens and Mutagenesis 1. Mutagens A. Physical and Chemical mutagens B. Transposons and retrotransposons C. T-DNA 2. Mutagenesis A. Screen B. Selection C. Lethal mutations Read: 508-514 Figs:
Lecture 3 Mutagens and Mutagenesis 1. Mutagens A. Physical and Chemical mutagens B. Transposons and retrotransposons C. T-DNA 2. Mutagenesis A. Screen B. Selection C. Lethal mutations Read: 508-514 Figs:
Proteomics And Cancer Biomarker Discovery. Dr. Zahid Khan Institute of chemical Sciences (ICS) University of Peshawar. Overview. Cancer.
 Proteomics And Cancer Biomarker Discovery Dr. Zahid Khan Institute of chemical Sciences (ICS) University of Peshawar Overview Proteomics Cancer Aims Tools Data Base search Challenges Summary 1 Overview
Proteomics And Cancer Biomarker Discovery Dr. Zahid Khan Institute of chemical Sciences (ICS) University of Peshawar Overview Proteomics Cancer Aims Tools Data Base search Challenges Summary 1 Overview
Course Descriptions. BIOL: Biology. MICB: Microbiology. [1]
![Course Descriptions. BIOL: Biology. MICB: Microbiology. [1] Course Descriptions. BIOL: Biology. MICB: Microbiology. [1]](/thumbs/72/66855241.jpg) Course Descriptions BIOL: Biology http://www.calendar.ubc.ca/vancouver/courses.cfm?code=biol [1] BIOL 112 (3) Biology of the Cell The principles of cellular and molecular biology using bacterial and eukaryotic
Course Descriptions BIOL: Biology http://www.calendar.ubc.ca/vancouver/courses.cfm?code=biol [1] BIOL 112 (3) Biology of the Cell The principles of cellular and molecular biology using bacterial and eukaryotic
Hsp90 Governs Echinocandin Resistance in the Pathogenic Yeast Candida albicans via Calcineurin
 in the Pathogenic Yeast Candida albicans via Calcineurin Sheena D. Singh 1, Nicole Robbins 1, Aimee K. Zaas 2, Wiley A. Schell 2, John R. Perfect 2,3, Leah E. Cowen 1 * 1 Department of Molecular Genetics,
in the Pathogenic Yeast Candida albicans via Calcineurin Sheena D. Singh 1, Nicole Robbins 1, Aimee K. Zaas 2, Wiley A. Schell 2, John R. Perfect 2,3, Leah E. Cowen 1 * 1 Department of Molecular Genetics,
Scientific Method. Name: NetID: Exam 1 Version 1 September 12, 2017 Dr. A. Pimentel
 Name: NetID: Exam 1 Version 1 September 12, 2017 Dr. A. Pimentel Each question has a value of 4 points and there is a total of 156 points in the exam. However, the maximum score of this exam will be capped
Name: NetID: Exam 1 Version 1 September 12, 2017 Dr. A. Pimentel Each question has a value of 4 points and there is a total of 156 points in the exam. However, the maximum score of this exam will be capped
Catalog No. SA-40002: 50 preparation SA-40001: 100 preparation
 BDtract TM Genomic DNA Isolation Kit Instruction Manual Catalog No. SA-40002: 50 preparation SA-40001: 100 preparation Maxim Biotech, Inc. 780 Dubuque Avenue, So. San Francisco, CA 94080, U.S.A. Tel: (800)
BDtract TM Genomic DNA Isolation Kit Instruction Manual Catalog No. SA-40002: 50 preparation SA-40001: 100 preparation Maxim Biotech, Inc. 780 Dubuque Avenue, So. San Francisco, CA 94080, U.S.A. Tel: (800)
Gut microbial colonization in day old chicks. Henri Woelders DirkJan Schokker Mari A. Smits
 Gut microbial colonization in day old chicks Henri Woelders DirkJan Schokker Mari A. Smits Trends Robustness; Resilience. Ability to keep healthy, or recover from disease (with minimal human intervention).
Gut microbial colonization in day old chicks Henri Woelders DirkJan Schokker Mari A. Smits Trends Robustness; Resilience. Ability to keep healthy, or recover from disease (with minimal human intervention).
Analysis of the expression of homoeologous genes in polyploid cereals using fluorescence cdna-sscp
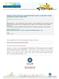 Analysis of the expression of homoeologous genes in polyploid cereals using fluorescence cdna-sscp De Bustos A., Perez R., Jouve N. in Molina-Cano J.L. (ed.), Christou P. (ed.), Graner A. (ed.), Hammer
Analysis of the expression of homoeologous genes in polyploid cereals using fluorescence cdna-sscp De Bustos A., Perez R., Jouve N. in Molina-Cano J.L. (ed.), Christou P. (ed.), Graner A. (ed.), Hammer
PV92 PCR Bio Informatics
 Purpose of PCR Chromosome 16 PV92 PV92 PCR Bio Informatics Alu insert, PV92 locus, chromosome 16 Introduce the polymerase chain reaction (PCR) technique Apply PCR to population genetics Directly measure
Purpose of PCR Chromosome 16 PV92 PV92 PCR Bio Informatics Alu insert, PV92 locus, chromosome 16 Introduce the polymerase chain reaction (PCR) technique Apply PCR to population genetics Directly measure
