Supplementary information. Interplay between Penicillin-binding proteins and SEDS proteins promotes. bacterial cell wall synthesis
|
|
|
- Noah Jacobs
- 5 years ago
- Views:
Transcription
1 Supplementary information Interplay between Penicillin-binding proteins and SEDS proteins promotes bacterial cell wall synthesis Sophie Leclercq 1, Adeline Derouaux 1, Samir Olatunji 1, Claudine Fraipont 1, Alexander J. F. Egan 2, Waldemar Vollmer 2, Eefjan Breukink 3 and Mohammed Terrak 1 1 Centre d Ingénierie des Protéines, University of Liège, B6a, Quartier Agora, allée du six Août 11, 4000 Liège 1, Belgium 2 Institute for Cell and Molecular Biosciences, The Centre for Bacterial Cell Biology, Newcastle University, Richardson Road, Newcastle upon Tyne, NE2 4AX, UK 3 Membrane Biochemistry and Biophysics, Department of Chemistry, Faculty of Science, Utrecht University, Padualaan 8, 3584 CH Utrecht, The Netherlands To whom correspondence should be addressed: Mohammed Terrak, Centre d Ingénierie des Protéines, University of Liège, B6a, Quartier Agora, Allée du six Août 11, 4000 Liège 1, Belgium, Tel.: ; mterrak@ulg.ac.be 1
2 Supplementary methods Plasmid constructions pdml924 (S510A). Mutation of the active serine S510A in the transpeptidase domain of PBP1b was generated using the QuikChange Site-Directed Mutagenesis method (Agilent Technologies), oligonucleotides S510A1 and S510A2 (Table S1) and plasmid pdml924 1 as template. pdml2040. The ftsi gene was amplified by PCR from pdml using the IW1 and IW2 primers (Table S1) and inserted in the MCS1 of the petduet-1 vector (Novagen-EMD Millipore) between the BamHI and HindIII sites. The ftsw gene was amplified by PCR from pdml using the IW3 and IW4 primers (Table S1) and inserted in the MCS2 of the petduet-1 vector between the BglII and XhoI sites. This construct allows the production of Histagged PBP3 and untagged FtsW. pdml2041. The ftsw gene was amplified by PCR from pdml2400 using the WI1 and WI2 primers (Table S1) and inserted in the MCS1 of the petduet-1 vector between the BamHI and HindIII sites. The ftsi gene was amplified by PCR from pdml using the WI3 and WI4 primers (Table S1) and inserted in the MCS2 of the petduet-1 vector between the BglII and XhoI sites. This construct allows the production of His-tag FtsW and untagged PBP3. pdml2043. Plasmid pdml contains the ftsw gene with a fragment encoding the antigenic HA peptide (YPYDVPDYA), derived from Human influenza hemagglutinin (HA), inserted between codons for E293 and A294 of the large periplasmic loop. The fragment of the ftsw HA gene between SnaBI-SalI sites and containing the sequence coding for HA peptide was recovered by digestion from the plasmid pdml2422 and inserted at the same sites of plasmid 2
3 pdml2040 yielding pdml2043. This construct allows the production of His-tagged PBP3 and untagged FtsW HA. pdml2045. The ponb gene was amplified by PCR from phk using the SN1b1 and SN1b2 primers (Table S1) and inserted in the MCS1 of the pacycduet-1 (Novagen-EMD Millipore) between the NcoI and BamHI sites. The ftsn gene was obtained by digestion of pdml2032 by NdeI and XhoI and inserted at the same sites in the MCS2 of pacycduet-1. This construct allows the production of untagged PBP1b and FtsN-S. tag. pdml2032. The ftsn gene was amplified by PCR from pdml using the Nhis1 and Nhis2 primers (Table S1), digested by NdeI and XhoI and introduced in pet22b between the same sites. This construct allows the production of FtsN-His-tag. pcip1000. The ponb gene was amplified by PCR using the W1b1 and W1b2 primers (Table S1). The resulting fragment was digested by NdeI/XhoI and used to replace the ftsi gene of plasmid pdml2041 digested with the same enzymes. This construct allows the production of His-tagged FtsW and untagged PBP1b. pcip1083. The ponb gene was amplified from plasmid pcip1000 using the W1b1 and W1b2 primers (Table S1) digested by NdeI/XhoI and cloned into the same sites of the MCS2 of an empty petduet-1 vector to generate pcip1061 (petduet-ponb). Then, the sequence coding for FtsW HA was amplified by PCR from pdml2043 using the WHA1 and WHA2 primers. The PCR product was digested by BamHI/EcoRI and inserted into MCS1 of pcip1061 between the corresponding sites to give rise to pcip1083. This construct allows the production of His-tagged FtsW HA and untagged PBP1b. 3
4 pet28mhl-w Kp and pet28mhl-w Se. The genes coding for FtsW of Klebsiella pneumoniae and Salmonella enterica were amplified by PCR using the genomic DNAs of the respective strains as templates and the primers pairs (WKp1 and WKp2) and (WSe1 and WSe2) (Table S1), digested with NdeI/XhoI and cloned into the same sites of pet28mhl. These constructs allow the production of His-tagged FtsW kp and His-tagged FtsW Se respectively. pcip1051. The DNA encoding the full-length E. coli murj gene was amplified by PCR from E. coli K12 genomic DNA using the MJ1 and MJ2 primers and inserted into the BamHI/HindIII sites of petduet-1 to give rise to pcip1051. This construct allows the production of His-tagged MurJ. 4
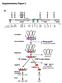
5 Supplementary figures Figure S1 Figure S1. Effect of FtsW and the FtsW-PBP3 complex on the activity of PBP1b in vitro. HPLC chromatograms show the muropeptide profile from in vitro PG assays: PBP1b or PBP1b- LpoB was incubated with lipid II in the presence or absence of FtsW or FtsW-PBP3. The resulting PG was digested with cellosyl, resulting muropetides were reduced with sodium borohydride and separated by HPLC. Peak 1, monophosphorylated disaccharide pentapeptide; peak 2, disaccharide tetrapeptide; peak 3, disaccharide pentapeptide; peak 4, bis-disaccharide tetratetrapeptide; peak 5, bis-disaccharide tetrapentapeptide; peak 6, tris-disaccharide tetratetrapeptide. 5
6 Figure S2 Figure S2. E. coli C43 was transformed with empty pet-duet plasmid, or pet Duet containing ftsw gene or both FtsW and ftsi genes. The cells were grown in LB medium at 37 C and the optical density (OD) at 600nm of the cell cultures were monitored over time. 6
7 Supplementary table S1. Oligonucleotides used in this study. Primers S510A1: 5 -CGTTCGATTGGTGCGCTTGCAAAACCAG S510A2: 5 -CTGGTTTTGCAAGCGCACCAATCGAACG IW1: 5 -GCCGGATCCGAAAGCAGCGGCGAAAACGCAG IW2: 5 -GCCAAGCTTACGATCTGCCACCTGTCCCCTCG IW3: 5 -GGCAGATCTCCGTTTATCTCTCCCTCGCCTGAAAATG IW4: 5 -CGGCTCGAGTCATCGTGAACCTCGTACAAACGCCTG WI1: 5 -GCCGGATCCGCGTTTATCTCTCCCTCGCCTGAAAATG WI2: 5 -GCCAAGCTTTCATCGTGAACCTCGTACAAACGCCTG WI3: 5 -GCCAGATCTCAAAGCAGCGGCGAAAACGCAG WI4: 5 -GCCCTCGAGTTACGATCTGCCACCTGTCCCCTCG Nhis1: 5 -GCCATATGGCACAACGAGATTATGTACGCC Nhis2: 5 -GGCCTCGAGACCCCCGGCGGCGAGCCG SN1b1: 5 -GAGCCATGGGCATGCCGCGCAAAGGTAAGGGCAAAGG SN1b2: 5 -GACGGATCCTTAATTACTACCAAACATATCCTTGATCCAACCGG W1b1: 5 -TATACATATGGCCGGGAATGACCGCGAG W1b2: 5 -CAGACTCGAGTTAATTACTACCAAACATATCCTTGATCCAAC WHA1: 5 -GCCAGGATCCGCGTTTATCTCTCCCTCGCCTGA WHA2: 5 -CCGCGAATTCTCATCGTGAACCTCGTACAAACGCC WKp1: 5 -AGGTTGCATATGCGTTTCTCTC WKp2: 5 -ACCGCCTCGAGTCATCGCACA WSe1: 5 -CGAAGGAGTTAGGTTGCATATGCGTTTATC WSe2: 5 -GTTGACCACCTCGAGTCATCGAGAA Mj1: 5 -GCCAGGATCCGAATTTATTAAAATCGCTGGCCGCCGTC Mj2: 5 -CCGCAAGCTTTTACACCGTCCGGCGGGCAAATTC Constructs pdml924 (S510A) pdml2040 pdml2041 pdml2032 pdml2045 pcip1000 pcip1083 pet28mhl-w Kp pet28mhl-w Se pcip1051 7
8 Supplementary references 1. Terrak, M. et al. The catalytic, glycosyl transferase and acyl transferase modules of the cell wall peptidoglycan-polymerizing penicillin-binding protein 1b of Escherichia coli. Mol Microbiol 34, (1999). 2. Piette, A. et al. Structural determinants required to target penicillin-binding protein 3 to the septum of Escherichia coli. J Bacteriol 186, (2004). 3. Pastoret, S. et al. Functional analysis of the cell division protein FtsW of Escherichia coli. J Bacteriol 186, (2004). 4. Müller, P. et al. The Essential Cell Division Protein FtsN Interacts with the Murein (Peptidoglycan) Synthase PBP1B in Escherichia coli. J Biol Chem 282, (2007). 8
Supplementary Information
 Supplementary Information Supplementary Figures Figure S1. Study of mgtl translation in vitro. (A) Detection of 5 LR RNA using wild-type and anti-sd (91-95) substituted templates in a transcription-translation
Supplementary Information Supplementary Figures Figure S1. Study of mgtl translation in vitro. (A) Detection of 5 LR RNA using wild-type and anti-sd (91-95) substituted templates in a transcription-translation
Fig. S1. CrgA intracellular levels in M. tuberculosis. Ten and twenty micrograms of
 Supplementary data Fig. S1. CrgA intracellular levels in M. tuberculosis. Ten and twenty micrograms of cell free protein lysates from WT M. tuberculosis (Rv) together with various known concentrations
Supplementary data Fig. S1. CrgA intracellular levels in M. tuberculosis. Ten and twenty micrograms of cell free protein lysates from WT M. tuberculosis (Rv) together with various known concentrations
Supporting information
 Supporting information Construction of strains and plasmids To create ptc67, a PCR product obtained with primers cc2570-162f (gcatgggcaagcttgaggacggcgtcatgt) and cc2570+512f (gaggccgtggtaccatagaggcgggcg),
Supporting information Construction of strains and plasmids To create ptc67, a PCR product obtained with primers cc2570-162f (gcatgggcaagcttgaggacggcgtcatgt) and cc2570+512f (gaggccgtggtaccatagaggcgggcg),
A fail-safe mechanism in the septal ring assembly pathway generated by the sequential recruitment of cell separation amidases and their activators
 Supplemental Information for: A fail-safe mechanism in the septal ring assembly pathway generated by the sequential recruitment of cell separation amidases and their activators Nick T. Peters, Thuy Dinh,
Supplemental Information for: A fail-safe mechanism in the septal ring assembly pathway generated by the sequential recruitment of cell separation amidases and their activators Nick T. Peters, Thuy Dinh,
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL Engineering a pyridoxal '-phosphate supply for cadaverine production by using Escherichia coli whole-cell biocatalysis Weichao Ma 1,,, Weijia Cao 1,, Bowen Zhang 1,, Kequan Chen
SUPPLEMENTARY MATERIAL Engineering a pyridoxal '-phosphate supply for cadaverine production by using Escherichia coli whole-cell biocatalysis Weichao Ma 1,,, Weijia Cao 1,, Bowen Zhang 1,, Kequan Chen
Supporting Information
 Supporting Information Development of a 2,4-Dinitrotoluene-Responsive Synthetic Riboswitch in E. coli cells Molly E. Davidson, Svetlana V. Harbaugh, Yaroslav G. Chushak, Morley O. Stone, Nancy Kelley-
Supporting Information Development of a 2,4-Dinitrotoluene-Responsive Synthetic Riboswitch in E. coli cells Molly E. Davidson, Svetlana V. Harbaugh, Yaroslav G. Chushak, Morley O. Stone, Nancy Kelley-
Molecular Biology Techniques Supporting IBBE
 Molecular Biology Techniques Supporting IBBE Jared Cartwright Protein Production Lab Head Contact Details: email jared.cartwright@york.ac.uk Phone 01904 328797 Presentation Aims Gene synthesis Cloning
Molecular Biology Techniques Supporting IBBE Jared Cartwright Protein Production Lab Head Contact Details: email jared.cartwright@york.ac.uk Phone 01904 328797 Presentation Aims Gene synthesis Cloning
Supporting Information
 Supporting Information Risser and Callahan 10.1073/pnas.0909152106 SI Text Plasmid Construction. Plasmid pdr211 is a mobilizable shuttle vector containing P pete -pats. A fragment containing P pete was
Supporting Information Risser and Callahan 10.1073/pnas.0909152106 SI Text Plasmid Construction. Plasmid pdr211 is a mobilizable shuttle vector containing P pete -pats. A fragment containing P pete was
Human Viperin Causes Radical SAM Dependent Elongation of E. coli Hinting at its Physiological Role
 Supporting Information Human Viperin Causes Radical SAM Dependent Elongation of E. coli Hinting at its Physiological Role Micah T. Nelp, Anthony P. Young, Branden M. Stepanski, Vahe Bandarian* Department
Supporting Information Human Viperin Causes Radical SAM Dependent Elongation of E. coli Hinting at its Physiological Role Micah T. Nelp, Anthony P. Young, Branden M. Stepanski, Vahe Bandarian* Department
Development of Immunogens to Protect Against Turkey Cellulitis. Douglas. N. Foster and Robyn Gangl. Department of Animal Science
 Development of Immunogens to Protect Against Turkey Cellulitis Douglas. N. Foster and Robyn Gangl Department of Animal Science University of Minnesota St. Paul, MN 55108 Introduction Clostridial dermatitis
Development of Immunogens to Protect Against Turkey Cellulitis Douglas. N. Foster and Robyn Gangl Department of Animal Science University of Minnesota St. Paul, MN 55108 Introduction Clostridial dermatitis
Figure S1. Biosynthetic pathway of GDP-PerNAc. Mass spectrum of purified GDP-PerNAc. Nature Protocols: doi: /nprot
 Synthesis of GDP-PerNAc In E. coli O157, the biosynthesis of GDP- -N-acetyl-D-perosamine (GDP-PerNAc) involves three enzymes (Fig. S1). GDP-D-Mannose is converted by GDP-mannose-4,6-dehydratase (GMD) into
Synthesis of GDP-PerNAc In E. coli O157, the biosynthesis of GDP- -N-acetyl-D-perosamine (GDP-PerNAc) involves three enzymes (Fig. S1). GDP-D-Mannose is converted by GDP-mannose-4,6-dehydratase (GMD) into
Bi 8 Lecture 5. Ellen Rothenberg 19 January 2016
 Bi 8 Lecture 5 MORE ON HOW WE KNOW WHAT WE KNOW and intro to the protein code Ellen Rothenberg 19 January 2016 SIZE AND PURIFICATION BY SYNTHESIS: BASIS OF EARLY SEQUENCING complex mixture of aborted DNA
Bi 8 Lecture 5 MORE ON HOW WE KNOW WHAT WE KNOW and intro to the protein code Ellen Rothenberg 19 January 2016 SIZE AND PURIFICATION BY SYNTHESIS: BASIS OF EARLY SEQUENCING complex mixture of aborted DNA
Antisense RNA Targeting the First Periplasmic Domain of YidC in Escherichia coli Appears to Induce Filamentation but Does Not Affect Cell Viability
 Antisense RNA Targeting the First Periplasmic Domain of YidC in Escherichia coli Appears to Induce Filamentation but Does Not Affect Cell Viability Riaaz Lalani, Nathaniel Susilo, Elisa Xiao, Andrea Xu
Antisense RNA Targeting the First Periplasmic Domain of YidC in Escherichia coli Appears to Induce Filamentation but Does Not Affect Cell Viability Riaaz Lalani, Nathaniel Susilo, Elisa Xiao, Andrea Xu
7.1 Techniques for Producing and Analyzing DNA. SBI4U Ms. Ho-Lau
 7.1 Techniques for Producing and Analyzing DNA SBI4U Ms. Ho-Lau What is Biotechnology? From Merriam-Webster: the manipulation of living organisms or their components to produce useful usually commercial
7.1 Techniques for Producing and Analyzing DNA SBI4U Ms. Ho-Lau What is Biotechnology? From Merriam-Webster: the manipulation of living organisms or their components to produce useful usually commercial
Chapter 8: Recombinant DNA. Ways this technology touches us. Overview. Genetic Engineering
 Chapter 8 Recombinant DNA and Genetic Engineering Genetic manipulation Ways this technology touches us Criminal justice The Justice Project, started by law students to advocate for DNA testing of Death
Chapter 8 Recombinant DNA and Genetic Engineering Genetic manipulation Ways this technology touches us Criminal justice The Justice Project, started by law students to advocate for DNA testing of Death
Figure S1. gfp tola and pal mcherry can complement deletion mutants of tola and pal respectively. (A)When strain LS4522 was grown in the presence of
 Figure S1. gfp tola and pal mcherry can complement deletion mutants of tola and pal respectively. (A)When strain LS4522 was grown in the presence of xylose, inducing expression of gfp-tola, localization
Figure S1. gfp tola and pal mcherry can complement deletion mutants of tola and pal respectively. (A)When strain LS4522 was grown in the presence of xylose, inducing expression of gfp-tola, localization
SUPPLEMENTARY DATA: Supplementary Figures:
 SUPPLEMENTARY DATA: Supplementary Figures: Supplementary Fig. S1: In vivo recombinant Mtb reporter assay: Dependence of mcherry expression as a function of E. coli growth: A. Fluorescent intensity of mcherry
SUPPLEMENTARY DATA: Supplementary Figures: Supplementary Fig. S1: In vivo recombinant Mtb reporter assay: Dependence of mcherry expression as a function of E. coli growth: A. Fluorescent intensity of mcherry
Δsig. ywa. yjbm. rela -re. rela
 A wt A. A, P rela -re la A/Δ yjbm A/Δ ywa C A, P hy-s igd, D (A, P IPTG hy-s igd, ) D D (+ IPTG ) D/Δ rela Figure S1 SigD B. Figure S1. SigD levels and swimming motility in (p)ppgpp synthetase mutants.
A wt A. A, P rela -re la A/Δ yjbm A/Δ ywa C A, P hy-s igd, D (A, P IPTG hy-s igd, ) D D (+ IPTG ) D/Δ rela Figure S1 SigD B. Figure S1. SigD levels and swimming motility in (p)ppgpp synthetase mutants.
Supplemental Information
 Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
Recitation CHAPTER 9 DNA Technologies
 Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
Please sign below if you wish to have your grades posted by the last five digits of your SSN
 BIO 226R EXAM III (Sample) PRINT YOUR NAME Please sign below if you wish to have your grades posted by the last five digits of your Signature BIO 226R Exam III has 8 pages, and 26 questions. There are
BIO 226R EXAM III (Sample) PRINT YOUR NAME Please sign below if you wish to have your grades posted by the last five digits of your Signature BIO 226R Exam III has 8 pages, and 26 questions. There are
The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity
 Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
Certificate of Analysis
 Certificate of Analysis pet6xhn Expression Vector Set Contents Product Information... 1 pet6xhn-n, pet6xhn-c, and pet6xhn-gfpuv Vector Information... 2 Location of Features... 4 Additional Information...
Certificate of Analysis pet6xhn Expression Vector Set Contents Product Information... 1 pet6xhn-n, pet6xhn-c, and pet6xhn-gfpuv Vector Information... 2 Location of Features... 4 Additional Information...
Red Type Indicates Unique Site
 3600 G0605 pscaavmcmvmcsbghpa Plasmid Features: Coordinates Feature 980-1084 AAV2 5 ITR 1144-1666 modified CMV 1667-1761 MCS 1762-1975 BgHpA 1987-2114 AAV2 3 ITR 3031-3891 B-lactamase (Ampicillin) Antibiotic
3600 G0605 pscaavmcmvmcsbghpa Plasmid Features: Coordinates Feature 980-1084 AAV2 5 ITR 1144-1666 modified CMV 1667-1761 MCS 1762-1975 BgHpA 1987-2114 AAV2 3 ITR 3031-3891 B-lactamase (Ampicillin) Antibiotic
CAP BIOINFORMATICS Su-Shing Chen CISE. 10/5/2005 Su-Shing Chen, CISE 1
 CAP 5510-9 BIOINFORMATICS Su-Shing Chen CISE 10/5/2005 Su-Shing Chen, CISE 1 Basic BioTech Processes Hybridization PCR Southern blotting (spot or stain) 10/5/2005 Su-Shing Chen, CISE 2 10/5/2005 Su-Shing
CAP 5510-9 BIOINFORMATICS Su-Shing Chen CISE 10/5/2005 Su-Shing Chen, CISE 1 Basic BioTech Processes Hybridization PCR Southern blotting (spot or stain) 10/5/2005 Su-Shing Chen, CISE 2 10/5/2005 Su-Shing
Rotation Report Sample Version 3. Due Date: August 9, Analysis of the Guanine Nucleotide Exchange Activity of the S. cerevisiae Ats1 Protein
 Rotation Report Sample Version 3 Due Date: August 9, 1998 Analysis of the Guanine Nucleotide Exchange Activity of the S. cerevisiae Ats1 Protein Anita H. Corbett Advisor: Amy Jones Rotation 1 Abstract:
Rotation Report Sample Version 3 Due Date: August 9, 1998 Analysis of the Guanine Nucleotide Exchange Activity of the S. cerevisiae Ats1 Protein Anita H. Corbett Advisor: Amy Jones Rotation 1 Abstract:
Enzyme that uses RNA as a template to synthesize a complementary DNA
 Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Antisense RNA Insert Design for Plasmid Construction to Knockdown Target Gene Expression
 Vol. 1:7-15 Antisense RNA Insert Design for Plasmid Construction to Knockdown Target Gene Expression Ji, Tom, Lu, Aneka, Wu, Kaylee Department of Microbiology and Immunology, University of British Columbia
Vol. 1:7-15 Antisense RNA Insert Design for Plasmid Construction to Knockdown Target Gene Expression Ji, Tom, Lu, Aneka, Wu, Kaylee Department of Microbiology and Immunology, University of British Columbia
Figure 1. Map of cloning vector pgem T-Easy (bacterial plasmid DNA)
 Texas A&M University-Corpus Christi CHEM4402 Biochemistry II Laboratory Laboratory 6: Ligation & Bacterial Transformation (Bring your text and laptop to class if you wish to work on your assignment during
Texas A&M University-Corpus Christi CHEM4402 Biochemistry II Laboratory Laboratory 6: Ligation & Bacterial Transformation (Bring your text and laptop to class if you wish to work on your assignment during
Improved method for assembly of linear yeast expression cassettes using NEBuilder HiFi DNA Assembly Master Mix
 DNA CLONING DNA AMPLIFICATION & PCR Improved method for assembly of linear yeast expression cassettes using NEBuilder HiFi DNA Assembly Master Mix EPIGENETICS RNA ANALYSIS LIBRARY PREP FOR NEXT GEN SEQUENCING
DNA CLONING DNA AMPLIFICATION & PCR Improved method for assembly of linear yeast expression cassettes using NEBuilder HiFi DNA Assembly Master Mix EPIGENETICS RNA ANALYSIS LIBRARY PREP FOR NEXT GEN SEQUENCING
On-chip microbial culture for the specific detection of very low levels of bacteria. Supplementary Information
 On-chip microbial culture for the specific detection of very low levels of bacteria Sihem Bouguelia, Yoann Roupioz, Sami Slimani, Laure Mondani, Guillermina Casabona, Claire Durmort, Thierry Vernet, Roberto
On-chip microbial culture for the specific detection of very low levels of bacteria Sihem Bouguelia, Yoann Roupioz, Sami Slimani, Laure Mondani, Guillermina Casabona, Claire Durmort, Thierry Vernet, Roberto
MCB 102 University of California, Berkeley August 11 13, Problem Set 8
 MCB 102 University of California, Berkeley August 11 13, 2009 Isabelle Philipp Handout Problem Set 8 The answer key will be posted by Tuesday August 11. Try to solve the problem sets always first without
MCB 102 University of California, Berkeley August 11 13, 2009 Isabelle Philipp Handout Problem Set 8 The answer key will be posted by Tuesday August 11. Try to solve the problem sets always first without
Basics of Recombinant DNA Technology Biochemistry 302. March 5, 2004 Bob Kelm
 Basics of Recombinant DNA Technology Biochemistry 302 March 5, 2004 Bob Kelm Applications of recombinant DNA technology Mapping and identifying genes (DNA cloning) Propagating genes (DNA subcloning) Modifying
Basics of Recombinant DNA Technology Biochemistry 302 March 5, 2004 Bob Kelm Applications of recombinant DNA technology Mapping and identifying genes (DNA cloning) Propagating genes (DNA subcloning) Modifying
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
Different Upstream Activators Stimulate Transcription Through Distinct Coactivator Complexes
 Supplemental Material for Different Upstream Activators Stimulate Transcription Through Distinct Coactivator Complexes Dong-ki Lee, Soyoun Kim, and John T. Lis Due to be published as a Research Communication
Supplemental Material for Different Upstream Activators Stimulate Transcription Through Distinct Coactivator Complexes Dong-ki Lee, Soyoun Kim, and John T. Lis Due to be published as a Research Communication
Understanding the Cellular Mechanism of the Excess Microsporocytes I (EMSI) Gene. Andrew ElBardissi, The Pennsylvania State University
 Understanding the Cellular Mechanism of the Excess Microsporocytes I (EMSI) Gene Andrew ElBardissi, The Pennsylvania State University Abstract: Hong Ma, The Pennsylvania State University The Excess Microsporocytes
Understanding the Cellular Mechanism of the Excess Microsporocytes I (EMSI) Gene Andrew ElBardissi, The Pennsylvania State University Abstract: Hong Ma, The Pennsylvania State University The Excess Microsporocytes
Supporting Information
 Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany RNA ligands that distinguish metabolite-induced conformations in the TPP riboswitch Günter Mayer, Marie-Sophie L. Raddatz, Julia D. Grunwald,
Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany RNA ligands that distinguish metabolite-induced conformations in the TPP riboswitch Günter Mayer, Marie-Sophie L. Raddatz, Julia D. Grunwald,
Bi 8 Lecture 4. Ellen Rothenberg 14 January Reading: from Alberts Ch. 8
 Bi 8 Lecture 4 DNA approaches: How we know what we know Ellen Rothenberg 14 January 2016 Reading: from Alberts Ch. 8 Central concept: DNA or RNA polymer length as an identifying feature RNA has intrinsically
Bi 8 Lecture 4 DNA approaches: How we know what we know Ellen Rothenberg 14 January 2016 Reading: from Alberts Ch. 8 Central concept: DNA or RNA polymer length as an identifying feature RNA has intrinsically
Biosynthesis of 3-Methoxy-5-Methyl Naphthoic Acid and Its Incorporation into the Antitumor Antibiotic Azinomycin B
 Biosynthesis of 3-Methoxy-5-Methyl Naphthoic Acid and Its Incorporation into the Antitumor Antibiotic Azinomycin B Wei Ding, a,b Wei Deng, b Mancheng Tang, b Qi Zhang, b Gongli Tang, b Yurong Bi, a and
Biosynthesis of 3-Methoxy-5-Methyl Naphthoic Acid and Its Incorporation into the Antitumor Antibiotic Azinomycin B Wei Ding, a,b Wei Deng, b Mancheng Tang, b Qi Zhang, b Gongli Tang, b Yurong Bi, a and
Fatchiyah
 Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
Genetic Engineering & Recombinant DNA
 Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
Mutating Asn-666 to Glu in the O-helix region of the taq DNA polymerase gene
 Research in Pharmaceutical Sciences, April 2010; 5(1): 15-19 Received: Oct 2009 Accepted: Jan 2010 School of Pharmacy & Pharmaceutical Sciences 15 Isfahan University of Medical Sciences Original Article
Research in Pharmaceutical Sciences, April 2010; 5(1): 15-19 Received: Oct 2009 Accepted: Jan 2010 School of Pharmacy & Pharmaceutical Sciences 15 Isfahan University of Medical Sciences Original Article
Schematic representation of the endogenous PALB2 locus and gene-disruption constructs
 Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Site directed mutagenesis, Insertional and Deletion Mutagenesis. Mitesh Shrestha
 Site directed mutagenesis, Insertional and Deletion Mutagenesis Mitesh Shrestha Mutagenesis Mutagenesis (the creation or formation of a mutation) can be used as a powerful genetic tool. By inducing mutations
Site directed mutagenesis, Insertional and Deletion Mutagenesis Mitesh Shrestha Mutagenesis Mutagenesis (the creation or formation of a mutation) can be used as a powerful genetic tool. By inducing mutations
Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question in Section B and ONE question from Section C.
 UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 2013-2014 MOLECULAR BIOLOGY BIO-2B02 Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question
UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 2013-2014 MOLECULAR BIOLOGY BIO-2B02 Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question
Biotechnology and DNA Technology
 11/27/2017 PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College CHAPTER 9 Biotechnology and DNA Technology Introduction to Biotechnology Learning Objectives Compare
11/27/2017 PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College CHAPTER 9 Biotechnology and DNA Technology Introduction to Biotechnology Learning Objectives Compare
Texas A&M University-Corpus Christi CHEM4402 Biochemistry II Laboratory Laboratory 4 - Polymerase Chain Reaction (PCR)
 Texas A&M University-Corpus Christi CHEM4402 Biochemistry II Laboratory Laboratory 4 - Polymerase Chain Reaction (PCR) Progressing with the sequence of experiments, we are now ready to amplify the green
Texas A&M University-Corpus Christi CHEM4402 Biochemistry II Laboratory Laboratory 4 - Polymerase Chain Reaction (PCR) Progressing with the sequence of experiments, we are now ready to amplify the green
Genome Analysis. Bacterial genome projects
 Genome Analysis Bacterial Genome sequencing does this help us in the investigation of adaptive responses/regulatory systems? Genome Sequencing Projects strategy & methods annotation Comparative genomics
Genome Analysis Bacterial Genome sequencing does this help us in the investigation of adaptive responses/regulatory systems? Genome Sequencing Projects strategy & methods annotation Comparative genomics
SUPPLEMENTARY EXPEMENTAL PROCEDURES
 SUPPLEMENTARY EXPEMENTAL PROCEDURES Plasmids- Total RNAs were extracted from HeLaS3 cells and reverse-transcribed using Superscript III Reverse Transcriptase (Invitrogen) to obtain DNA template for the
SUPPLEMENTARY EXPEMENTAL PROCEDURES Plasmids- Total RNAs were extracted from HeLaS3 cells and reverse-transcribed using Superscript III Reverse Transcriptase (Invitrogen) to obtain DNA template for the
Supporting Information
 Supporting Information Zhang et al. 10.1073/pnas.0809084105 SI Text Cloning of a PKS4 KS MAT Didomain. The KS MAT didomain was cloned as a standalone protein with primer pair pks4-ks-f-ndei and pks4-at-r-noti-s-ecori,
Supporting Information Zhang et al. 10.1073/pnas.0809084105 SI Text Cloning of a PKS4 KS MAT Didomain. The KS MAT didomain was cloned as a standalone protein with primer pair pks4-ks-f-ndei and pks4-at-r-noti-s-ecori,
NAME TA SEC Problem Set 4 FRIDAY October 15, Answers to this problem set must be inserted into the box outside
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel NAME TA SEC 7.012 Problem Set 4 FRIDAY October 15,
Engineering splicing factors with designed specificities
 nature methods Engineering splicing factors with designed specificities Yang Wang, Cheom-Gil Cheong, Traci M Tanaka Hall & Zefeng Wang Supplementary figures and text: Supplementary Figure 1 Supplementary
nature methods Engineering splicing factors with designed specificities Yang Wang, Cheom-Gil Cheong, Traci M Tanaka Hall & Zefeng Wang Supplementary figures and text: Supplementary Figure 1 Supplementary
G0463 pscaavmcsbghpa MCS. Plasmid Features:
 3200 G0463 pscaavmcsbghpa Plasmid Features: Coordinates Feature 980-1085 AAV2 ITR 106bp (mutated ITR) 1110-1226 MCS 1227-1440 BgHpA 1453-1595 AAV2 ITR (143bp) 2496-3356 B-lactamase (Ampicillin) Antibiotic
3200 G0463 pscaavmcsbghpa Plasmid Features: Coordinates Feature 980-1085 AAV2 ITR 106bp (mutated ITR) 1110-1226 MCS 1227-1440 BgHpA 1453-1595 AAV2 ITR (143bp) 2496-3356 B-lactamase (Ampicillin) Antibiotic
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
Deep sequencing reveals global patterns of mrna recruitment
 Supplementary information for: Deep sequencing reveals global patterns of mrna recruitment during translation initiation Rong Gao 1#*, Kai Yu 1#, Ju-Kui Nie 1,Teng-Fei Lian 1, Jian-Shi Jin 1, Anders Liljas
Supplementary information for: Deep sequencing reveals global patterns of mrna recruitment during translation initiation Rong Gao 1#*, Kai Yu 1#, Ju-Kui Nie 1,Teng-Fei Lian 1, Jian-Shi Jin 1, Anders Liljas
Chapter 20 Recombinant DNA Technology. Copyright 2009 Pearson Education, Inc.
 Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
7.012 Problem Set 5. Question 1
 Name Section 7.012 Problem Set 5 Question 1 While studying the problem of infertility, you attempt to isolate a hypothetical rabbit gene that accounts for the prolific reproduction of rabbits. After much
Name Section 7.012 Problem Set 5 Question 1 While studying the problem of infertility, you attempt to isolate a hypothetical rabbit gene that accounts for the prolific reproduction of rabbits. After much
7.013 Practice Quiz
 MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel 7.013 Practice Quiz 2 2004 1 Question 1 A. The primer
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel 7.013 Practice Quiz 2 2004 1 Question 1 A. The primer
TABLE S1. Sequences of the primers used for RT-PCR and PCR
 TABLE S1. Sequences of the primers used for RT-PCR and PCR Primer Sequence (5 to 3 ) Purpose Cloning: posuprtid-1 & posuprtid-2 deletion plasmids DM01 AGCTCCAGCGGGTCGCCGGGGTTGGCC DM02 CCAGCGGGTCGCCGGGGTTGGCC
TABLE S1. Sequences of the primers used for RT-PCR and PCR Primer Sequence (5 to 3 ) Purpose Cloning: posuprtid-1 & posuprtid-2 deletion plasmids DM01 AGCTCCAGCGGGTCGCCGGGGTTGGCC DM02 CCAGCGGGTCGCCGGGGTTGGCC
Biotechnology. Biotechnology is difficult to define but in general it s the use of biological systems to solve problems.
 MITE 2 S Biology Biotechnology Summer 2004 Austin Che Biotechnology is difficult to define but in general it s the use of biological systems to solve problems. Recombinant DNA consists of DNA assembled
MITE 2 S Biology Biotechnology Summer 2004 Austin Che Biotechnology is difficult to define but in general it s the use of biological systems to solve problems. Recombinant DNA consists of DNA assembled
Genetics Lecture 21 Recombinant DNA
 Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
Genetics Lecture 21 Recombinant DNA Recombinant DNA In 1971, a paper published by Kathleen Danna and Daniel Nathans marked the beginning of the recombinant DNA era. The paper described the isolation of
Roche Molecular Biochemicals Technical Note No. LC 12/2000
 Roche Molecular Biochemicals Technical Note No. LC 12/2000 LightCycler Absolute Quantification with External Standards and an Internal Control 1. General Introduction Purpose of this Note Overview of Method
Roche Molecular Biochemicals Technical Note No. LC 12/2000 LightCycler Absolute Quantification with External Standards and an Internal Control 1. General Introduction Purpose of this Note Overview of Method
Regulation of ARE transcript 3 end processing by the. yeast Cth2 mrna decay factor
 Regulation of ARE transcript 3 end processing by the yeast Cth2 mrna decay factor Manoël Prouteau, Marie-Claire Daugeron and Bertrand Séraphin Supplementary Information Material and Methods Plasmid construction
Regulation of ARE transcript 3 end processing by the yeast Cth2 mrna decay factor Manoël Prouteau, Marie-Claire Daugeron and Bertrand Séraphin Supplementary Information Material and Methods Plasmid construction
Genome Sequence Assembly
 Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Supplementary Information for: Engineered synthetic scaffolds for organising proteins within bacterial cytoplasms
 Lee et al. Supplementary Information 1 Supplementary Information for: Engineered synthetic scaffolds for organising proteins within bacterial cytoplasms Authors: Matthew J. Lee, 1 Judith Mantell, 2,3 Lorna
Lee et al. Supplementary Information 1 Supplementary Information for: Engineered synthetic scaffolds for organising proteins within bacterial cytoplasms Authors: Matthew J. Lee, 1 Judith Mantell, 2,3 Lorna
Chapter 6 - Molecular Genetic Techniques
 Chapter 6 - Molecular Genetic Techniques Two objects of molecular & genetic technologies For analysis For generation Molecular genetic technologies! For analysis DNA gel electrophoresis Southern blotting
Chapter 6 - Molecular Genetic Techniques Two objects of molecular & genetic technologies For analysis For generation Molecular genetic technologies! For analysis DNA gel electrophoresis Southern blotting
based on the methyl-sensitivity of MazF RNA endonuclease
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2017 Detection of N 6 -methyladenosine based on the methyl-sensitivity of MazF RNA endonuclease Miki
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2017 Detection of N 6 -methyladenosine based on the methyl-sensitivity of MazF RNA endonuclease Miki
Biology 4100 Minor Assignment 1 January 19, 2007
 Biology 4100 Minor Assignment 1 January 19, 2007 This assignment is due in class on February 6, 2007. It is worth 7.5% of your final mark for this course. Your assignment must be typed double-spaced on
Biology 4100 Minor Assignment 1 January 19, 2007 This assignment is due in class on February 6, 2007. It is worth 7.5% of your final mark for this course. Your assignment must be typed double-spaced on
In vitro mutagenesis
 Core course BMS361N Genetic Engineering In vitro mutagenesis Prof. Narkunaraja Shanmugam Dept. Of Biomedical Science School of Basic Medical Sciences Bharathidasan University In vitro mutagenesis Many
Core course BMS361N Genetic Engineering In vitro mutagenesis Prof. Narkunaraja Shanmugam Dept. Of Biomedical Science School of Basic Medical Sciences Bharathidasan University In vitro mutagenesis Many
Problem Set 8. Answer Key
 MCB 102 University of California, Berkeley August 11, 2009 Isabelle Philipp Online Document Problem Set 8 Answer Key 1. The Genetic Code (a) Are all amino acids encoded by the same number of codons? no
MCB 102 University of California, Berkeley August 11, 2009 Isabelle Philipp Online Document Problem Set 8 Answer Key 1. The Genetic Code (a) Are all amino acids encoded by the same number of codons? no
Supplementary Figure 1 Performance analysis of RedLibs (version 0.2.3). Supplementary Figure 2
 Supplementary Figure 1 Performance analysis of RedLibs (version 0.2.3). The required calculation time for the evaluation of all sub-libraries with a given target size are plotted against the combinatorial
Supplementary Figure 1 Performance analysis of RedLibs (version 0.2.3). The required calculation time for the evaluation of all sub-libraries with a given target size are plotted against the combinatorial
Microbial Biotechnology agustin krisna wardani
 Microbial Biotechnology agustin krisna wardani 1. The Structure of Microbes Microbes (microorganisms) are tiny organisms that are too small to be seen individually by the naked eye and must be viewed with
Microbial Biotechnology agustin krisna wardani 1. The Structure of Microbes Microbes (microorganisms) are tiny organisms that are too small to be seen individually by the naked eye and must be viewed with
A LEUCYL-tRNA SYNTHETASE WITHOUT THE EDITING DOMAIN AND ITS DIRECTED EVOLUTION FOR UNNATURAL PROTEIN ENGINEERING. Emily Fung
 i A LEUCYL-tRNA SYNTHETASE WITHOUT THE EDITING DOMAIN AND ITS DIRECTED EVOLUTION FOR UNNATURAL PROTEIN ENGINEERING By Emily Fung Submitted in partial fulfillment of the Requirements for Departmental Honors
i A LEUCYL-tRNA SYNTHETASE WITHOUT THE EDITING DOMAIN AND ITS DIRECTED EVOLUTION FOR UNNATURAL PROTEIN ENGINEERING By Emily Fung Submitted in partial fulfillment of the Requirements for Departmental Honors
Design. Construction. Characterization
 Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
yellow protein turn off/on label (POOL) , Republic of Korea The PYP gene was subcloned in the pqe80l vector using EZ-Cloning (enzynomics ).
 Supporting Information High-throughput instant quantification of protein expression and purity based on photoactive yellow protein turn off/on label (POOL) Youngmin Kim 1,2, Prabhakar Ganesan 1,2 and Hyotcherl
Supporting Information High-throughput instant quantification of protein expression and purity based on photoactive yellow protein turn off/on label (POOL) Youngmin Kim 1,2, Prabhakar Ganesan 1,2 and Hyotcherl
Reading Lecture 8: Lecture 9: Lecture 8. DNA Libraries. Definition Types Construction
 Lecture 8 Reading Lecture 8: 96-110 Lecture 9: 111-120 DNA Libraries Definition Types Construction 142 DNA Libraries A DNA library is a collection of clones of genomic fragments or cdnas from a certain
Lecture 8 Reading Lecture 8: 96-110 Lecture 9: 111-120 DNA Libraries Definition Types Construction 142 DNA Libraries A DNA library is a collection of clones of genomic fragments or cdnas from a certain
Cytochrome c heme lyase can mature a fusion peptide composed of the amino-terminal residues of horse cytochrome c
 Cytochrome c heme lyase can mature a fusion peptide composed of the amino-terminal residues of horse cytochrome c Wesley B. Asher a and Kara L. Bren a * a Department of Chemistry, University of Rochester,
Cytochrome c heme lyase can mature a fusion peptide composed of the amino-terminal residues of horse cytochrome c Wesley B. Asher a and Kara L. Bren a * a Department of Chemistry, University of Rochester,
Chapter 9 Genetic Engineering
 Chapter 9 Genetic Engineering Biotechnology: use of microbes to make a protein product Recombinant DNA Technology: Insertion or modification of genes to produce desired proteins Genetic engineering: manipulation
Chapter 9 Genetic Engineering Biotechnology: use of microbes to make a protein product Recombinant DNA Technology: Insertion or modification of genes to produce desired proteins Genetic engineering: manipulation
Supplemental Data Genetic and Epigenetic Regulation of the FLO Gene Family Generates Cell-Surface Variation in Yeast
 Supplemental Data Genetic and Epigenetic Regulation of the FLO Gene Family Generates Cell-Surface Variation in Yeast S1 Adrian Halme, Stacie Bumgarner, Cora Styles and Gerald R. Fink Yeast Strain Construction
Supplemental Data Genetic and Epigenetic Regulation of the FLO Gene Family Generates Cell-Surface Variation in Yeast S1 Adrian Halme, Stacie Bumgarner, Cora Styles and Gerald R. Fink Yeast Strain Construction
Data Sheet Quick PCR Cloning Kit
 Data Sheet Quick PCR Cloning Kit 6044 Cornerstone Ct. West, Ste. E DESCRIPTION: The Quick PCR Cloning Kit is a simple and highly efficient method to insert any gene or DNA fragment into a vector, without
Data Sheet Quick PCR Cloning Kit 6044 Cornerstone Ct. West, Ste. E DESCRIPTION: The Quick PCR Cloning Kit is a simple and highly efficient method to insert any gene or DNA fragment into a vector, without
Classic cloning with pask-iba and pexpr-iba vectors
 Classic cloning with pask-iba and pexpr-iba vectors General protocol Last date of revision March 2017 Version PR08-0008 For research use only Important licensing information Products featuring CMV promoter,
Classic cloning with pask-iba and pexpr-iba vectors General protocol Last date of revision March 2017 Version PR08-0008 For research use only Important licensing information Products featuring CMV promoter,
Efficient Multi-site-directed Mutagenesis directly from Genomic Template.
 Efficient Multi-site-directed Mutagenesis directly from Genomic Template. Fengtao Luo 1, Xiaolan Du 1, Tujun Weng 1, Xuan Wen 1, Junlan Huang 1, Lin Chen 1 Running title: Multi-site-directed Mutagenesis
Efficient Multi-site-directed Mutagenesis directly from Genomic Template. Fengtao Luo 1, Xiaolan Du 1, Tujun Weng 1, Xuan Wen 1, Junlan Huang 1, Lin Chen 1 Running title: Multi-site-directed Mutagenesis
Bioinformatics Support of Genome Sequencing Projects. Seminar in biology
 Bioinformatics Support of Genome Sequencing Projects Seminar in biology Introduction The Big Picture Biology reminder Enzyme for DNA manipulation DNA cloning DNA mapping Sequencing genomes Alignment of
Bioinformatics Support of Genome Sequencing Projects Seminar in biology Introduction The Big Picture Biology reminder Enzyme for DNA manipulation DNA cloning DNA mapping Sequencing genomes Alignment of
A universal chassis plasmid pwh34 was first constructed to facilitate the
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 File name: Supplementary method Plasmids construction A universal chassis plasmid pwh34 was first constructed to facilitate the construction
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 File name: Supplementary method Plasmids construction A universal chassis plasmid pwh34 was first constructed to facilitate the construction
Supplementary Information
 Supplementary Information Experimental procedures Identifying the viable circular permutation sites of E. coli EPSPS Bioinformatics prediction and E. coli complementation assay were performed together
Supplementary Information Experimental procedures Identifying the viable circular permutation sites of E. coli EPSPS Bioinformatics prediction and E. coli complementation assay were performed together
Chapter 10 (Part II) Gene Isolation and Manipulation
 Biology 234 J. G. Doheny Chapter 10 (Part II) Gene Isolation and Manipulation Practice Questions: Answer the following questions with one or two sentences. 1. What does PCR stand for? 2. What does the
Biology 234 J. G. Doheny Chapter 10 (Part II) Gene Isolation and Manipulation Practice Questions: Answer the following questions with one or two sentences. 1. What does PCR stand for? 2. What does the
BS1940 Course Topics Fall 2001 Drs. Hatfull and Arndt
 BS1940 Course Topics Fall 2001 Drs. Hatfull and Arndt Introduction to molecular biology Combining genetics, biochemistry, structural chemistry Information flow in biological systems: The Central Dogma
BS1940 Course Topics Fall 2001 Drs. Hatfull and Arndt Introduction to molecular biology Combining genetics, biochemistry, structural chemistry Information flow in biological systems: The Central Dogma
Journal of Experimental Microbiology and Immunology (JEMI) Vol. 6:20-25 Copyright December 2004, M&I UBC
 Preparing Plasmid Constructs to Investigate the Characteristics of Thiol Reductase and Flavin Reductase With Regard to Solubilizing Insoluble Proteinase Inhibitor 2 in Bacterial Protein Overexpression
Preparing Plasmid Constructs to Investigate the Characteristics of Thiol Reductase and Flavin Reductase With Regard to Solubilizing Insoluble Proteinase Inhibitor 2 in Bacterial Protein Overexpression
pdsipher and pdsipher -GFP shrna Vector User s Guide
 pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
3 Designing Primers for Site-Directed Mutagenesis
 3 Designing Primers for Site-Directed Mutagenesis 3.1 Learning Objectives During the next two labs you will learn the basics of site-directed mutagenesis: you will design primers for the mutants you designed
3 Designing Primers for Site-Directed Mutagenesis 3.1 Learning Objectives During the next two labs you will learn the basics of site-directed mutagenesis: you will design primers for the mutants you designed
CHEM 4420 Exam I Spring 2013 Page 1 of 6
 CHEM 4420 Exam I Spring 2013 Page 1 of 6 Name Use complete sentences when requested. There are 100 possible points on this exam. The multiple choice questions are worth 2 points each. All other questions
CHEM 4420 Exam I Spring 2013 Page 1 of 6 Name Use complete sentences when requested. There are 100 possible points on this exam. The multiple choice questions are worth 2 points each. All other questions
Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question in Section B and ONE question from Section C.
 UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 2015-2016 FUNDAMENTALS OF MOLECULAR BIOLOGY AND GENETICS BIO-4003A Time allowed: 2 hours Answer ALL questions in Section
UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 2015-2016 FUNDAMENTALS OF MOLECULAR BIOLOGY AND GENETICS BIO-4003A Time allowed: 2 hours Answer ALL questions in Section
Molecular Genetics Techniques. BIT 220 Chapter 20
 Molecular Genetics Techniques BIT 220 Chapter 20 What is Cloning? Recombinant DNA technologies 1. Producing Recombinant DNA molecule Incorporate gene of interest into plasmid (cloning vector) 2. Recombinant
Molecular Genetics Techniques BIT 220 Chapter 20 What is Cloning? Recombinant DNA technologies 1. Producing Recombinant DNA molecule Incorporate gene of interest into plasmid (cloning vector) 2. Recombinant
Supplementary Online Material. An Expanded Eukaryotic Genetic Code 2QH, UK. * To whom correspondence should be addressed.
 Supplementary Online Material An Expanded Eukaryotic Genetic Code Jason W. Chin 1, T. Ashton Cropp, J. Christopher Anderson, Mridul Mukherji, Zhiwen Zhang and Peter G. Schultz* Department of Chemistry
Supplementary Online Material An Expanded Eukaryotic Genetic Code Jason W. Chin 1, T. Ashton Cropp, J. Christopher Anderson, Mridul Mukherji, Zhiwen Zhang and Peter G. Schultz* Department of Chemistry
CHAPTER 20 DNA TECHNOLOGY AND GENOMICS. Section A: DNA Cloning
 Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Section A: DNA Cloning 1. DNA technology makes it possible to clone genes for basic research and commercial applications: an overview 2. Restriction enzymes are used to make recombinant DNA 3. Genes can
Supplementary Figure 2
 Supplementary Figure 2 a SBS-C1 SBS-C2 SBS-C3 SBS-C4 SBS-C5 SBS-C6 SBS-C7 SBS-C8 SBS-C9 LCR CNS-1 CNS-2 Il5 Rad50 Il13 Il4 Kif3a Sept8 0 50 100 150 200kb Sau3AI 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17
Supplementary Figure 2 a SBS-C1 SBS-C2 SBS-C3 SBS-C4 SBS-C5 SBS-C6 SBS-C7 SBS-C8 SBS-C9 LCR CNS-1 CNS-2 Il5 Rad50 Il13 Il4 Kif3a Sept8 0 50 100 150 200kb Sau3AI 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17
Biology 105: Introduction to Genetics PRACTICE FINAL EXAM Part I: Definitions. Homology: Reverse transcriptase. Allostery: cdna library
 Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Reverse transcriptase Allostery: cdna library Transformation Part II Short Answer 1. Describe the reasons for
Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Reverse transcriptase Allostery: cdna library Transformation Part II Short Answer 1. Describe the reasons for
Motivation From Protein to Gene
 MOLECULAR BIOLOGY 2003-4 Topic B Recombinant DNA -principles and tools Construct a library - what for, how Major techniques +principles Bioinformatics - in brief Chapter 7 (MCB) 1 Motivation From Protein
MOLECULAR BIOLOGY 2003-4 Topic B Recombinant DNA -principles and tools Construct a library - what for, how Major techniques +principles Bioinformatics - in brief Chapter 7 (MCB) 1 Motivation From Protein
Supporting Information
 S-1 Supporting Information Covalent and Site-Specific Immobilization of Antibody using Photoactivatable Antibody Fc-Binding Proteins Expressed in Escherichia coli Yeolin Lee, Jiyun Jeong, Gabi Lee, Jeong
S-1 Supporting Information Covalent and Site-Specific Immobilization of Antibody using Photoactivatable Antibody Fc-Binding Proteins Expressed in Escherichia coli Yeolin Lee, Jiyun Jeong, Gabi Lee, Jeong
