Antibiotic resistance patterns of Escherichia coli isolates from different aquatic environmental sources in León, Nicaragua
|
|
|
- Bertha Hunter
- 5 years ago
- Views:
Transcription
1 ORIGINAL ARTICLE BACTERIOLOGY Antibiotic resistance patterns of Escherichia coli isolates from different aquatic environmental sources in León, Nicaragua E. Amaya,2 *, D. Reyes 2,3 *, M. Paniagua 2, S. Calderón 2, M.-U. Rashid, P. Colque 3,I.Kühn 3,R.Möllby 3, A. Weintraub and C. E. Nord ) Division of Clinical Microbiology, Department of Laboratory Medicine, Karolinska Institutet, Karolinska University Hospital, Huddinge, Stockholm, Sweden, 2) Department of Microbiology, Faculty of Medical Sciences, National Autonomous University of Nicaragua (UNAN), León, Nicaragua and 3) Department of Microbiology, Tumor and Cell Biology (MTC), Karolinska Institutet, Solna, Stockholm, Sweden Abstract Antibiotic-resistant bacteria have emerged due to the selective pressure of antimicrobial use in humans and animals. Water plays an important role in dissemination of these organisms among humans, animals and the environment. We studied the antibiotic resistance patterns among 43 Escherichia coli isolates from different aquatic environmental sources collected from October 2008 to May 200 in León, Nicaragua. High levels of antibiotic resistance were found in E. coli isolates in hospital sewage water and in eight of 87 well-water samples. Among the resistant isolates from the hospital sewage, ampicillin, chloramphenicol, ciprofloxacin, nalidixic acid, trimethoprimsulphamethoxazole was the most common multi-resistance profile. Among the resistant isolates from the wells, % were resistant to ampicillin, ceftazidime, ceftriaxone, cefotaxime, chloramphenicol, ciprofloxacin, gentamicin, nalidixic acid and trimethoprim-sulphamethoxazole. E. coli producing ESBL and harbouring bla CTX-M genes were detected in one of the hospital sewage samples and in 26% of the resistant isolates from the well-water samples. The bla CTX-M- group was more prevalent in E. coli isolates from the hospital sewage samples and the bla CTX-M- group was more prevalent in the well-water samples. Keywords: Antibiotics, extended-spectrum b-lactamase, hospital, resistance, sewage, water, well Original Submission: 8 February 202; Revised Submission: 20 April 202; Accepted: 20 May 202 Editor: R. Cantón Article published online: 6 June 202 Clin Microbiol Infect 202; 8: E347 E354 0./j x Corresponding author: C.E. Nord, Division of Clinical Microbiology, Karolinska Institutet, Karolinska University Hospital Huddinge, SE-4 86 Stockholm, Sweden carl.erik.nord@ki.se *The first two authors contributed equally to the manuscript. Introduction The emergence of antimicrobial-resistant bacteria presents a major threat to public health because it reduces the effectiveness of antimicrobial treatment, leading to increased morbidity, mortality and healthcare expenditure [ 3]. The main factor driving this process is the selective pressure of antimicrobial use in human and animal medicine, as well as in aquaculture and agriculture [,3,4]. Consequently, this has led to the dissemination of antibiotic-resistant bacteria throughout the environment [5,6]. Residues of human and veterinary drugs are introduced into the environment via a number of pathways, but primarily from discharges of wastewater treatment plants or land application of sewage sludge and animal manure [7,8]. Due to the concern described above, some measures have been taken to address this problem; for example, the World Health Organization has ranked antimicrobials according to their importance in human medicine as a critical step in developing risk management strategies for the use of antimicrobials in animals used for food production []. In the European Union and some other countries such as Switzerland, the use of antibiotics as growth promoters in animal farming has been banned for the last several years. In developing countries, control of the use of antibiotics in animal farming is not being implemented and there is no information regarading the use of antibiotics for such purposes. ª202 The Authors Clinical Microbiology and Infection ª202 European Society of Clinical Microbiology and Infectious Diseases
2 E348 Clinical Microbiology and Infection, Volume 8 Number, September 202 CMI The importance of determining the prevalence of antibiotic-resistant bacteria in an environmental reservoir such as water is that the environmental source is not only a way of dissemination of antibiotic-resistant microorganisms among human and animal populations, but also the route by which resistance genes are introduced into natural bacterial ecosystems. Kümmerer [], in a review about antibiotics in an aquatic environment, showed that bacteria that become resistant through the use of antibiotics during medical treatment, are an important source of resistance found in hospital effluents, municipal sewage and sewage treatment plants. Kim and Aga [7], in their review on the impact of antibiotic and antibiotic-resistant bacteria from wastewater treatment plants, showed that these sites provide favourable conditions for the proliferation of antibiotic-resistant bacteria and spread of resistance genes to non-resistant bacteria. Thus, tracking the spread of antibiotic-resistant bacteria in water samples, such as sewage, tap and well water, is a useful source of information that can be used by policy makers in order to create risk management strategies for water environments [5]. The aim of this study was to determine the antibiotic resistance patterns of Escherichia coli isolates from different aquatic environmental sources in León, Nicaragua. The total number of coliforms was recorded, and E. colilike morphology colonies (n = 240, up to 32 bacterial colonies from each water sample were collected, when possible) were subcultured on MacConkey agar plates following an overnight incubation at 37 C and classified as E. coli using the PhP-RE microplates of the PhenePlate system (PhPlate, Stockholm, Sweden; as previously described [0,]. Isolates with correlations higher than 0.75 to each other were assigned to the same biochemical phenotype (BPT). BPTs with identical isolates were called common (C-BPT) and those with one isolate were called single (S-BPT). All S-BPTs and at least one isolate representing each C-BPT from each sample were subcultured onto a fresh McConkey agar plate at 37 C and stored in Brain Heart Infusion broth containing 5% (v/v) glycerol at )70 C for further characterization. Screening for diarrhoeagenic E. coli (DEC) by PCR assays After thawing, the frozen isolates were subcultured on a MacConkey agar plate and incubated at 37 C for 8 h. A smear of each bacterial culture was used as a template for PCR analyses. Screening for DEC was performed by multiplex-pcr as described [2]. Materials and Methods Sample collection and identification of E. coli strains Sampling was carried out from October 2008 to May 200 from different localities in León, Nicaragua. Samples were collected once from (i) household drinking water (n = 20); (ii) well water used for consumption (n = 87); (iii) sewage water from two municipal sedimentation treatment plants (two samples of each influent and effluent, n = 8); and (iv) sewage effluents of the main hospital in the city (n = 3). All samples were collected in sterile 500-mL glass or polyethylene bottles without preservatives and transported at 4 C to the Department of Microbiology at National Autonomous University of Nicaragua, León, where primary isolation of E. coli was performed. Samples from tap water were membrane filtered directly through 0.45-lm pore size filters (Millipore Corporation, Bedford, MA, USA), while all other water samples were subjected to serial ten-fold dilutions with phosphate-buffered saline before filtration. Membrane filters were placed on Chromogenic Selective Agar (Oxoid, Malmö, Sweden) and Difco m-fc Agar (Becton Dickinson AB, Stockholm, Sweden) plates and incubated aerobically for 24 h at 37 and 44.5 C, respectively. Selection of the E. coli isolates for antibiotic susceptibility testing For screening of antibiotic resistance, initially we selected all of the S-BPTs and at least one isolate representing each C-BPT from the results of the biochemical fingerprints of each of the 48 water samples with E. coli growth. Then, if the E. coli isolate (S-BPT or C-BPT) showed antibiotic resistance to at least one of the tested antimicrobial agents, the rest of the E. coli isolates from the corresponding water sample were tested for antibiotic susceptibility (when possible, up to 32 E. coli isolates from each water sample where a S-BPT or C-BPT was shown to be antibiotic resistant were tested) (see Table ). Antibiotic susceptibility testing Minimal inhibitory concentrations (MICs) were determined by the agar dilution method for the antibiotics ampicillin (AMP; AstraZeneca, Södertälje, Sweden), amoxicillin-clavulanic acid (AMC; Sigma, Steinheim, Germany and SmithKline Beecham, Washington, DC, USA), cefotaxime (CTX; Sigma), ceftazidime (CAZ; Sigma), ceftriaxone (CRO; Sigma), ciprofloxacin (CIP; Sigma), chloramphenicol (CHL; Sigma), gentamicin (GEN; Sigma), nalidixic acid (NAL; Sigma) and trimethoprim-sulphamethoxazole (STX; Sigma). Phenotypic detection of extended-spectrum b-lactamase (ESBL), by using ª202 The Authors Clinical Microbiology and Infection ª202 European Society of Clinical Microbiology and Infectious Diseases, CMI, 8, E347 E354
3 CMI Amaya et al. Resistant E. coli in environmental water from Nicaragua E34 TABLE. Total E. coli isolates from the water samples Water source Sample collected (n) Samples where E. coli was isolated (n) Samples where antibiotic-resistant E. coli a was detected E. coli isolates analysed for antibiotic resistance Tap water 20 Well water Sewage treatment plant Sewage (hospital) Total a Antibiotic-resistant S-BPT or C-BPT E. coli isolate. the Etest Ò system (Biomérieux, Solna, Sweden), was performed in those E. coli isolates that showed resistance to any of the third-generation cephalosporins tested. All of these analyses were carried out according to the Clinical and Laboratory Standards Institute guidelines [3]. E. coli ATCC 2522 and Enterococcus faecalis ATCC 222 strains were used as control strains. The data from the antibiotic susceptibility testing were analysed using WHONET 5.4. bla genes group detection and RAPD analysis All ESBL-positive E. coli strains were screened for the resistance genes encoding bla SHV, bla TEM, bla CTX-M and bla OXA by a multiplex PCR assay using universal primers following the procedure described by Fang et al. [4]. Further detection of CTX-M groups, 2,, 8 and 25 was performed using a multiplex PCR assay as described by Dallenne et al. [5] and a single PCR assay as described by Pitout et al. [6]. PCR amplification was carried out on a DNA thermal cycler GeneAmpÒ PCR system 700 (Applied Biosystems Division, Foster City, CA, USA). The epidemiological relationships between E. coli isolates producing ESBL from the hospital sewage water samples and from well-water samples were analysed by RAPD as described by Touati et al. [7] with some modifications. This analysis was carried out in 22 E. coli isolates from the wellwater samples and 7 from the hospital sewage water samples. E. coli isolates showing the same antibiotic resistance and bla genes profile were selected. Total DNA was prepared with the QIAamp Ò DNA mini Kit (Qiagen, Solna, Sweden) and used for RAPD typing, which was performed using puretaq Ready-To-Go PCR beads (GE Healthcare, Little Chalfont, Buckinghamshire, UK) together with the following primers: primer 4 (5 -AAGAGCCCGT-3 ) and primer 5 (5 -AACGCGCAAC-3 ) (Thermo Fisher Scientific, Limburg, Germany). PCR amplification was carried out as follows: one cycle at 4 C for 5 min, 35 (primer 4) and 3 (primer 5) cycles at 4 C for 5 s, 42 C for 30 s and 72 C for min, with a final extension period at 72 C for 5 min. After amplification, the banding pattern of randomly amplified DNA was visualized and analysed on.5% agarose gel in Tris-acetate buffer. A negative control was included in each PCR run with no target DNA. Reproducibility of the amplification results was evaluated in parallel experiments by the repetition of the PCR reactions three times. Electrophoresed agarose gels were analysed using the BioNumerics Ò version 6 software (Applied Maths, Sint-Martens-Latem, Belgium). Dendrograms based on the Jaccard coefficient and unweighted pair group method using arithmetic averages (UPGMA) were generated. Sequencing analysis To identify the specific bla genes detected in the PCR assays for bla SHV, bla TEM, bla OXA and bla CTX-M, DNA sequence analyses of the amplicons were performed. Based on the RAPD analysis, representative E. coli isolates (mainly those E. coli isolates harbouring bla CTX-M plus another type of bla gene, i.e. bla SHV ) from each clonal group were selected for sequencing analysis. For bla SHV, bla TEM and bla OXA, sequencing primers described by Fang et al. [4] were used. For the bla CTX-M- and bla CTX-M- groups sequencing primers described by Pitout et al.[6] were used. Amplified PCR products were purified using the QIAquick PCR Purification Kit (Qiagen) and bidirectional sequencing was performed. Each sequence was then compared with the known bla genes sequences ( by multiplesequence alignment using the BLAST program. Results Screening of diarrhoeagenic E. colis trains None of the samples was positive for any of the tested diarrhoeagenic virulence markers. Identification and selection of the E. coli isolates for antibiotic susceptibility testing Antibiotic-resistant E. coli isolates (S-BPTs or C-BPT from the biochemical fingerprints) were detected in eight of 87 of the well-water samples, seven of eight sewage water samples from the municipal sedimentation treatment plants and in all ª202 The Authors Clinical Microbiology and Infection ª202 European Society of Clinical Microbiology and Infectious Diseases, CMI, 8, E347 E354
4 E350 Clinical Microbiology and Infection, Volume 8 Number, September 202 CMI three hospital sewage water samples. No E. coli isolates were detected in any of the 20 tap water samples. A total of 43 E. coli isolates were included in the study (Table ), all from the water samples with an antibiotic resistant S-BPT or C- BPT E. coli. Antibiotic susceptibilities in the selected E. coli isolates High levels of antibiotic resistance were mainly found in the E. coli isolates from the three samples of hospital sewage water (Table 2). All of the E. coli isolates from samples HB and HC showed resistance to ampicillin, nalidixic acid, ciprofloxacin, trimethoprim-sulphamethoxazole and chloramphenicol but were sensitive to amoxicillin-clavulanic acid, ceftazidime, ceftriaxone, cefotaxime and gentamicin. This finding indicates a very homogenous E. coli population in these two samples, probably one multi-resistant strain, a result that was supported by the phenotyping analyses (not shown). In contrast, E. coli isolates from sample HA showed resistance levels not only to trimethoprim-sulphamethoxazol, chloramphenicol, nalidixic acid and ciprofloxacin but also to the b-lactam drugs (except for amoxicillin-clavulanic acid) and gentamicin. Likewise, multi-resistance patterns were mostly detected in E. coli isolates from the hospital sewage, with resistance to ampicillin, chloramphenicol, ciprofloxacin, nalidixic acid and trimethoprim-sulphamethoxazole as the most common multi-resistance profile in these E. coli isolates (67%). Antibiotic resistance levels in the E. coli isolates from the remaining water samples were low or infrequent, except for the well-water sample 5, from which all of the E. coli isolates were resistant to all tested antimicrobial agents except for amoxicillin-clavulanic acid. ESBL-producing E. coli isolates were detected in 33% of the resistant isolates from the hospital sewage water and in 26% of the resistant isolates from the well-water samples. bla genes group detection and RAPD analysis The gene encoding bla SHV was more commonly detected in ESBL-producing E. coli isolates from the hospital sewage water (53%) than from the well-water samples (22%). The gene encoding for bla TEM was only detected in ESBL-producing E. coli isolates from the hospital sewage water samples (4%). In contrast, the gene encoding for bla OXA was only detected in ESBL-producing E. coli isolates from the wellwater samples (57%). PCR amplification bla genes showed that the bla CTX-M- group was more prevalent in ESBL-producing E. coli isolates from the hospital sewage water samples (65%) than in wellwater samples (26%). In contrast, the bla CTX-M- group was more prevalent in ESBL-producing E. coli isolates from the TABLE 2. Percentage of resistance of E. coli isolates from distinct water sources to different antibiotics Resistance (%) CHL (S 8R 32) SXT (S 2R 4) CIP (S R 4) NAL (S 6 R 32) GEN (S 4R 6) CTX (S R 4) CRO (S R 4) CAZ (S 4R 6) AMC (S 8R 32) AMP (S 8R 32) No. E. coli isolates tested Sample code HA HB HC P P P P P P P SIA a SIA2 b 32 6 SIB a SIB2 b SUA a SUA2 b SUB2 b H, hospital sewage water samples; P, well-water samples; S, sewage water samples from the municipal sedimentation treatment plants: influent a and effluent b. ª202 The Authors Clinical Microbiology and Infection ª202 European Society of Clinical Microbiology and Infectious Diseases, CMI, 8, E347 E354
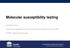
5 CMI Amaya et al. Resistant E. coli in environmental water from Nicaragua E Strain code P04- P04-4 P04-6 P04- P-02 P3-2 P7-3 P7-0 P P0-3 P0-08 P0-4 P0-6 P0-8 P0-20 P0-24 P0-0 P0-06 P0-07 PO-08 ESBL type CTX-M RAPD-clone TEM, SiP4 P P P P P P P2 P2 P2 SiP3 FIG.. Clonal similarity of E. coli from well water by RAPD-PCR. Arrows indicate E. coli isolates selected for sequencing analysis. well-water samples (74%) than in hospital sewage water (34%). The genes encoding for the bla CTX-M-2, bla CTX-M-8 and bla CTX-M-25 groups were not detected in any of the E. coli isolates studied. Twenty-two E. coli isolates from the well-water samples (codes P04, P08, P0, P0, P0, P3, P7 and 5) and 7 from the hospital sewage water sample (code HA) were selected for RAPD analysis. The analysis revealed that the isolates from the well-water samples could be separated into five clones (Fig. ). Among these clones, P and encompass most of the isolates (6 and E. coli isolates, respectively). Interestingly, all of the ESBL-producing E. coli isolates from clone P harboured the gene encoding for the bla CTX- M- group and most of the combination of genes encoding for bla SVH and bla CTX-M. In contrast, all of the ESBL-producing E. coli isolates from clone harboured the gene encoding for the bla CTX-M- group and most of the combination of genes encoding for bla TEM or bla OXA plus bla CTX-M. Regarding hospital sewage water samples, the RAPD analysis revealed that 7 resistant E. coli isolates from the hospital sewage water sample HA could be separated into clones (Fig. 2). Among them, clone encompassed the major number of E. coli isolates. All of them harboured the gene encoding for the bla CTX-M- group and most of the combination of genes encoding for bla SVH and bla CTX-M. The bla CTX-M- group was found mostly in ESBL-producing E. coli isolates that belonged to clone H. The RAPD analysis did not show any clonal similarity between the ESBL-producing E. coli isolates from the wells and the hospital sewage water samples. Sequencing analysis Based on the RAPD analysis, representative isolates from each clonal group were selected for further analysis by sequencing (mainly those E. coli isolates harbouring bla CTX-M plus another type of bla gene, i.e. bla SHV ). The selected E. coli isolates from the well-water and the hospital sewage water samples are marked by arrows (Figs and 2). After sequencing, it was found that bla SHV-/-2, bla TEM- and bla OXA-/-30 were present in the E. coli isolates positive in the PCR assay for bla SHV, bla TEM or bla OXA. For the bla CTX-M groups, it was found that the bla CTX-M-5 gene and bla CTX-M- gene were harboured in the E. coli isolates that were positive in the PCR assay for the bla CTX-M- and bla CTX-M- groups. Discussion Many studies have shown the presence of antibiotic-resistant bacteria or genes conferring resistance in the aquatic environment [6,8,]. Interestingly, this is found even in countries with a high control of the use of antibiotics (e.g. the presence of genes encoding resistance to aminoglycosides, b-lactams and tetracyclines as well as the presence of methicillin-resistant Staphylococcus aureus have been found in waste ª202 The Authors Clinical Microbiology and Infection ª202 European Society of Clinical Microbiology and Infectious Diseases, CMI, 8, E347 E354
6 E352 Clinical Microbiology and Infection, Volume 8 Number, September 202 CMI Strain code ESBL type CTX-M RAPD-clone HA-0 H HA-04 HA- HA-2 HA-0 HA-3 HA-03 HA-05 HA-0 HA-3 HA-28 HA-22 HA-4 HA-25 HA-6 HA-23 HA-32 SHV, SHV, SHV,, H H SiH2 SiH3 SiH4 SiH6 H7 H7 H8 H8 SiH FIG. 2. Clonal similarity of E. coli from the hospital sewage water by RAPD-PCR. Arrows indicate E. coli isolates selected for sequencing analysis. water environments from Sweden) [20]. In our study, the presence of E. coli resistant to at least one of the tested antibiotics was found in 8 of 8 of the environmental water samples collected in León, Nicaragua. Although many studies have shown the relationship of DEC to diarrhoea in Nicaragua [2,2], we could not detect the presence of DEC in the environmental water samples. However, the presence of antibiotic-resistant E. coli strains was frequent. Perhaps a long-term environmental water study covering the diarrhoea season in Nicaragua could show the role of contaminated water in the diarrhoea disease burden in Nicaragua. Ram et al. [22 24], in their studies of surface water samples (drinking water, for irrigation, or other purposes) from the Indian rivers, have demonstrated the presence of antibiotic-resistant shiga toxin and enterotoxin producing E. coli. The authors considered that such a finding could be an important health concern due to the risk of developing waterborne outbreaks. In many developing countries the unregulated sale and dispensing of antibiotics is very common [25,26]. Thus, it is important not only to consider the contribution of hospital effluents but also the contribution of the general community to the input of antibiotic-resistant bacteria to the aquatic environment []. Our results show that among all of the E. coli isolates included in this study, those from the hospital sewage water had higher antibiotic resistance levels to ampicillin (00%), nalidixic acid (70%), ciprofloxacin (6%), chloramphenicol (6%) and trimethoprim-sulphamethoxazole (00%) compared with E. coli isolates from other aquatic samples. The similarities in the frequency of antibiotic resistance found in the E. coli isolates from hospital sewage samples HB and HC, probably as a result of a dominating strain in both samples, could be due to the fact that those sewage systems collect the water effluent from related hospital units. Among the well-water samples that represent the contribution of the community to the input of antibiotic-resistant bacteria to the aquatic environment, E. coli isolates from well-water sample 5 were resistant to the tested antibiotics, which indicated a high contribution to the spread of multi-antibiotic-resistant bacteria, perhaps due to a high use of antibiotics in those settings. Kümmerer [] showed that there is a surprisingly high incidence of antibiotic-resistant E. coli in rural groundwater, perhaps due to run-off from farms or leakage from septic tanks. It has been shown that improper sanitation (e.g. improper excreta management) can lead to the spread of infectious diseases such as diarrhoea. In Nicaraguan rural areas, as in many developing countries, the use of a household latrine is very common and perhaps the presence of antibiotic-resistant bacteria in well-water samples could be due to improper construction of the latrines and hence leakage to the well water. ª202 The Authors Clinical Microbiology and Infection ª202 European Society of Clinical Microbiology and Infectious Diseases, CMI, 8, E347 E354
7 CMI Amaya et al. Resistant E. coli in environmental water from Nicaragua E353 Resistance to ampicillin, nalidixic acid, ciprofloxacin and trimethoprim-sulphamethoxazole, though lower, was also found in some of the E. coli isolates from the sewage water samples from the municipal sedimentation treatment plants (Table 2). Doung et al. [27] showed similar findings in their study of the occurrence of quinolone agents and the number of E. coli resistant to quinolones in hospital wastewater in Hanoi, Vietnam. They found higher levels of antibiotic-resistant bacteria in wastewater from hospitals as compared with wastewater treatment plants. It has been reported that resistant bacteria are eliminated quite well in the sewage treatment plants, which could explain the low level of resistance found in our study []. However, it is important to consider that there are factors that could have influenced our results, such as dilution effects and the viability of antibiotic-resistant bacteria in the environment. In previous studies we have reported on the emergence of bacteria producing ESBL causing infections in Nicaraguan children [8,28,2]. In those isolates, the gene encoding for CTX- M was the most commonly detected. In the present study it is shown that 00% of the ESBL-producing E. coli isolates from both hospital sewage and the well water of the community encode the genes for bla CTX-M. Genes encoding for bla TEM and bla SHV were only found in the hospital samples (44% and 53%, respectively) and the gene encoding for bla OXA was only detected in the well-water samples (57%). After sequencing of selected E. coli isolates (Figs and 2), it was found that bla SHV- /-2, bla TEM- and bla OXA-/-30 were present in the E. coli isolates positive in the PCR assay for bla SHV, bla TEM or bla OXA. CTX-M has become one of the main public health concerns due to its ability to be involved in nosocomial and community-acquired infections. E. coli is most often responsible for producing CTX-M and seems to be a true community ESBL-producing pathogen [30,3]. In the present study, it was found that the bla CTX-M-5 gene and bla CTX-M- gene were detected in the ESBL-producing E. coli isolates that were positive in the PCR assay for the bla CTX-M- group (more prevalent in E. coli isolates from the well-water samples) and bla CTX-M- group (more prevalent in E. coli isolates from the hospital sewage water samples). RAPD analysis did not show any clonal similarities between the ESBL-producing E. coli isolated from the hospital and well-water samples. However, we did find some dominant clones within samples (i.e. clone encompassed most of the ESBL-producing E. coli from well-water samples and clone most of the ESBL-producing E. coli in the hospital samples). In addition, the carriage of the gene encoding for the bla CTX-M groups was more common in some clones (i.e. all of the E. coli isolates from clone harboured the gene encoding genes for the bla CTX-M- group). Even though we did not perform a longitudinal study of environmental water samples, our results suggest that multiresistant ESBL-producing E. coli were widely spread in hospital sewage water and community water samples. Acknowledgements The authors thank Patricia Blandón Roiz for her excellent technical laboratory assistance and Elisa Huete, Martha Mairena, Azucena Laguna and Antonia Obando for their valuable fieldwork activities in sample collection and transportation. We also thank Dr Gilberto Moreno, Harmodio Paredes and Rosa Emelina Alonso from MINSA, SILAIS, León, for their valuable input into this study. Transparency Declaration This study was supported by a grant from the Swedish International Development Cooperation Agency (Ref. No and ) and the National Autonomous University of Nicaragua, León. The authors have no conflict of interest to declare. References. Collignon P, Powers JH, Chiller TM et al. World Health Organization ranking of antimicrobials according to their importance in human medicine: a critical step for developing risk management strategies for the use of antimicrobials in food production animals. Clin Infect Dis 200; 4: Hawkey PM, Jones AM. The changing epidemiology of resistance. J Antimicrob Chemother 200; 64 (suppl ): i Smith RD, Coast J. Antimicrobial resistance: a global response. Bull World Health Organ 2002; 80: Kümmerer K. Antibiotics in the aquatic environment a review part I. Chemosphere 200; 75: Baquero F, Martinez JL, Canton R. Antibiotics and antibiotic resistance in water environments. Curr Opin Biotechnol 2008; : Martinez JL. Environmental pollution by antibiotics and by antibiotic resistance determinants. Environ Pollut 200; 57: Kim S, Aga DS. Potential ecological and human health impacts of antibiotics and antibiotic-resistant bacteria from wastewater treatment plants. J Toxicol Environ Health B Crit Rev 2007; 0: Kümmerer K. Resistance in the environment. J Antimicrob Chemother 2004; 54: Kümmerer K. Antibiotics in the aquatic environment a review part II. Chemosphere 200; 75: Ahmed W, Neller R, Katouli M. Host species-specific metabolic fingerprint database for enterococci and Escherichia coli and its application to identify sources of fecal contamination in surface waters. Appl Environ Microbiol 2005; 7: Landgren M, Oden H, Kuhn I et al. Diversity among 248 Escherichia coli from women with community-acquired lower urinary tract ª202 The Authors Clinical Microbiology and Infection ª202 European Society of Clinical Microbiology and Infectious Diseases, CMI, 8, E347 E354
8 E354 Clinical Microbiology and Infection, Volume 8 Number, September 202 CMI infections in 7 countries. J Antimicrob Chemother 2005; 55: Vilchez S, Reyes D, Paniagua M et al. Prevalence of diarrhoeagenic Escherichia coli in children from León, Nicaragua. J Med Microbiol 200; 58: CLSI. Performance standard for antimicrobial susceptibility testing; Twentieth Informational Supplement. CLSI document M00-S20. Wayne, PA: Clinical and Laboratory Standards Institute, Fang H, Ataker F, Hedin G et al. Molecular epidemiology of extended-spectrum beta-lactamases among Escherichia coli isolates collected in a Swedish hospital and its associated health care facilities from 200 to J Clin Microbiol 2008; 46: Dallenne C, Da Costa A, Decre D et al. Development of a set of multiplex PCR assays for the detection of genes encoding important beta-lactamases in Enterobacteriaceae. J Antimicrob Chemother 200; 65: Pitout JD, Hossain A, Hanson ND. Phenotypic and molecular detection of CTX-M-beta-lactamases produced by Escherichia coli and Klebsiella spp. J Clin Microbiol 2004; 42: Touati A, Benallaoua S, Djoudi F et al. Characterization of CTX-M- 5-producing Klebsiella pneumoniae and Escherichia coli strains isolated from hospital environments in Algeria. Microb Drug Resist 2007; 3: Amaya E, Caceres M, Fang H et al. Extended-spectrum betalactamase-producing Klebsiella pneumoniae in a neonatal intensive care unit in León, Nicaragua. Int J Antimicrob Agents 200; 33: Martinez JL. Antibiotics and antibiotic resistance genes in natural environments. Science 2008; 32: Börjesson S, Dienues O, Jarnheimer PA et al. Quantification of genes encoding resistance to aminoglycosides, beta-lactams and tetracyclines in wastewater environments by real-time PCR. Int J Environ Health Res 200; : Paniagua M, Espinoza F, Ringman M et al. Analysis of incidence of infection with enterotoxigenic Escherichia coli in a prospective cohort study of infant diarrhea in Nicaragua. J Clin Microbiol 7; 35: Ram S, Vajpayee P, Shanker R. Prevalence of multi-antimicrobial-agent resistant, shiga toxin and enterotoxin producing Escherichia coli in surface waters of river Ganga. Environ Sci Technol 2007; 4: Ram S, Vajpayee P, Singh RL et al. Surface water of a perennial river exhibits multi-antimicrobial resistant shiga toxin and enterotoxin producing Escherichia coli. Ecotoxicol Environ Saf 200; 72: Ram S, Vajpayee P, Tripathi U et al.determination of antimicrobial resistance and virulence gene signatures in surface water isolates of Escherichia coli. J Appl Microbiol 2008; 05: Byarugaba DK. Antimicrobial resistance in developing countries and responsible risk factors. Int J Antimicrob Agents 2004; 24: Okeke IN, Lamikanra A, Edelman R. Socioeconomic and behavioral factors leading to acquired bacterial resistance to antibiotics in developing countries. Emerg Infect Dis ; 5: Duong HA, Pham NH, Nguyen HT et al. Occurrence, fate and antibiotic resistance of fluoroquinolone antibacterials in hospital wastewaters in Hanoi, Vietnam. Chemosphere 2008; 72: Amaya E, Caceres M, Fang H et al. Antibiotic resistance patterns in gram-negative and gram-positive bacteria causing septicemia in newborns in León, Nicaragua: correlation with environmental samples. J Chemother 200; 22: Amaya E, Reyes D, Vilchez S et al. Antibiotic resistance patterns of intestinal Escherichia coli isolates from Nicaraguan children. J Med Microbiol 20; 60: Pitout JD, Laupland KB. Extended-spectrum beta-lactamase-producing Enterobacteriaceae: an emerging public-health concern. Lancet Infect Dis 2008; 8: Pitout JD, Nordmann P, Laupland KB et al. Emergence of Enterobacteriaceae producing extended-spectrum beta-lactamases (ESBLs) in the community. J Antimicrob Chemother 2005; 56: ª202 The Authors Clinical Microbiology and Infection ª202 European Society of Clinical Microbiology and Infectious Diseases, CMI, 8, E347 E354
SUMMARY. Key words: antibioticresistance, Enterobacteriaceae, ESBL, CTX-M,
 SUMMARY Key words: antibioticresistance, Enterobacteriaceae, ESBL, CTX-M, The doctoral thesis entitled Prevalence of Enterobacteriaceae producing extendedspectrum beta-lactamases (ESBL) isolated from broilers
SUMMARY Key words: antibioticresistance, Enterobacteriaceae, ESBL, CTX-M, The doctoral thesis entitled Prevalence of Enterobacteriaceae producing extendedspectrum beta-lactamases (ESBL) isolated from broilers
Inside the Burch Lab: E. Coli and Triclosan Resistance. By: Pamela Lammonds
 Inside the Burch Lab: E. Coli and Triclosan Resistance By: Pamela Lammonds Purpose and Goals of Research Concerns over infectious disease have risen in the past few years. In response to this concern,
Inside the Burch Lab: E. Coli and Triclosan Resistance By: Pamela Lammonds Purpose and Goals of Research Concerns over infectious disease have risen in the past few years. In response to this concern,
Discussion Items. Microbial Indicators of Water Quality
 Discussion Items! Announcements! Group Project topics! Discuss previous lab (Microbes in food) Results Lab report Isolates streak isolate by Thursday.! Microbial analysis of water (Tuesday) MPN and MF!
Discussion Items! Announcements! Group Project topics! Discuss previous lab (Microbes in food) Results Lab report Isolates streak isolate by Thursday.! Microbial analysis of water (Tuesday) MPN and MF!
Use of Molecular Assays for Resistance Detection
 Use of Molecular Assays for Resistance Detection Antimicrobial resistance and susceptibility are complex, and current in vitro methods have been developed to predict a microorganism s response to antibacterial
Use of Molecular Assays for Resistance Detection Antimicrobial resistance and susceptibility are complex, and current in vitro methods have been developed to predict a microorganism s response to antibacterial
Detection and characterization of extended spectrum β-lactamase producing Escherichia coli from poultry of eastern India
 Detection and characterization of extended spectrum β-lactamase producing Escherichia coli from poultry of eastern India Dr. Samiran Bandyopadhyay Scientist Indian Veterinary Research Institute Eastern
Detection and characterization of extended spectrum β-lactamase producing Escherichia coli from poultry of eastern India Dr. Samiran Bandyopadhyay Scientist Indian Veterinary Research Institute Eastern
 IOSR Journal of Dental and Medical Sciences (IOSR-JDMS) e-issn: 2279-0853, p-issn: 2279-0861.Volume 16, Issue 10 Ver. VII (Oct. 2017), PP 76-81 www.iosrjournals.org Molecular Characterization of TEM, SHV
IOSR Journal of Dental and Medical Sciences (IOSR-JDMS) e-issn: 2279-0853, p-issn: 2279-0861.Volume 16, Issue 10 Ver. VII (Oct. 2017), PP 76-81 www.iosrjournals.org Molecular Characterization of TEM, SHV
Abstract. Introduction
 ORIGINAL ARTICLE 10.1111/j.1469-0691.2009.02893.x Comparative assessment of inoculum effects on the antimicrobial activity of amoxycillin clavulanate and piperacillin tazobactam with extended-spectrum
ORIGINAL ARTICLE 10.1111/j.1469-0691.2009.02893.x Comparative assessment of inoculum effects on the antimicrobial activity of amoxycillin clavulanate and piperacillin tazobactam with extended-spectrum
and Technology, Fuchu, Tokyo , Japan 3 Research Center for Environmental Technology and Sustainable Development (CETASD),
 Interdisciplinary Studies on Environmental Chemistry Biological Responses to Chemical Pollutants, Eds., Y. Murakami, K. Nakayama, S.-I. Kitamura, H. Iwata and S. Tanabe, pp. 355 359. by TERRAPUB, 2008.
Interdisciplinary Studies on Environmental Chemistry Biological Responses to Chemical Pollutants, Eds., Y. Murakami, K. Nakayama, S.-I. Kitamura, H. Iwata and S. Tanabe, pp. 355 359. by TERRAPUB, 2008.
Cloning and Characterization of E. meningoseptica Beta Lactamase
 Cloning and Characterization of E. meningoseptica Beta Lactamase Authors: Lindsey Purcell, Jessica Matts, Patricia Canaan* Department of Biochemistry and Molecular Biology Abstract Elizabethkingia meningoseptica
Cloning and Characterization of E. meningoseptica Beta Lactamase Authors: Lindsey Purcell, Jessica Matts, Patricia Canaan* Department of Biochemistry and Molecular Biology Abstract Elizabethkingia meningoseptica
1. Procedure for Antibiotic susceptibility test by disc diffusion analysis
 Nanoparticles Functionalized with Ampicillin Destroy Multiple Antibiotic Resistant Isolates of Pseudomonas aeruginosa, Enterobacter aerogenes and Methicillin Resistant Staphylococcus aureus Ashley Brown
Nanoparticles Functionalized with Ampicillin Destroy Multiple Antibiotic Resistant Isolates of Pseudomonas aeruginosa, Enterobacter aerogenes and Methicillin Resistant Staphylococcus aureus Ashley Brown
Virulence factors profile of drug-resistant Escherichia coli isolates from urinary tract infections in Punjab, Pakistan
 DOI 10.1007/s10096-010-1036-6 ARTICLE Virulence factors profile of drug-resistant Escherichia coli isolates from urinary tract infections in Punjab, Pakistan M. Idress & U. Mussarat & Y. Badshah & R. Qamar
DOI 10.1007/s10096-010-1036-6 ARTICLE Virulence factors profile of drug-resistant Escherichia coli isolates from urinary tract infections in Punjab, Pakistan M. Idress & U. Mussarat & Y. Badshah & R. Qamar
Ongoing epidemic of bla VIM-1 -positive Klebsiella pneumoniae in Athens, Greece: a prospective survey
 Journal of Antimicrobial Chemotherapy (2008) 61, 59 63 doi:10.1093/jac/dkm443 Advance Access publication 13 November 2007 Ongoing epidemic of bla VIM-1 -positive Klebsiella pneumoniae in Athens, Greece:
Journal of Antimicrobial Chemotherapy (2008) 61, 59 63 doi:10.1093/jac/dkm443 Advance Access publication 13 November 2007 Ongoing epidemic of bla VIM-1 -positive Klebsiella pneumoniae in Athens, Greece:
VanABC plus. GENSPEED VanABC plus. Molecular multiplex assay for maximum information.
 VanABC plus GENSPEED VanABC plus Molecular multiplex assay for maximum information www.genspeed-biotech.com Vancomycin Resistance? VanABC plus for all relevant targets GENSPEED VanABC plus Molecular multiplex
VanABC plus GENSPEED VanABC plus Molecular multiplex assay for maximum information www.genspeed-biotech.com Vancomycin Resistance? VanABC plus for all relevant targets GENSPEED VanABC plus Molecular multiplex
Curriculum Vitae. Abbas Maleki, Ph.D. Clinical Microbiology Research Center, Ilam University of Medical Sciences, Ilam, Iran
 Curriculum Vitae Abbas Maleki, Ph.D Clinical Microbiology Research Center, Ilam University of Medical Sciences, Ilam, Iran E-mail: abbasmaleki_ilam@yahoo.com maleki-a@medilam.ac.ir Tel: +989187419401 Personal
Curriculum Vitae Abbas Maleki, Ph.D Clinical Microbiology Research Center, Ilam University of Medical Sciences, Ilam, Iran E-mail: abbasmaleki_ilam@yahoo.com maleki-a@medilam.ac.ir Tel: +989187419401 Personal
Ten Minute, Reagent-Free identification of Bacteria Containing Resistance Genes Using a Rapid Intrinsic Fluorescence Method
 548 Ten Minute, Reagent-Free identification of Bacteria Containing Resistance Genes Using a Rapid Intrinsic Fluorescence Method R. Rozen-Sadowsky 1, A. Shinderman 1, D. Gohman 1, D. Shimonov 1, Y. Gluckman
548 Ten Minute, Reagent-Free identification of Bacteria Containing Resistance Genes Using a Rapid Intrinsic Fluorescence Method R. Rozen-Sadowsky 1, A. Shinderman 1, D. Gohman 1, D. Shimonov 1, Y. Gluckman
Prevalence and molecular characterization of clinical isolates of Escherichia coli expressing an AmpC phenotype
 J Antimicrob Chemother 2010; 65: 460 464 doi:10.1093/jac/dkp484 Advance publication 22 January 2010 Prevalence and molecular characterization of clinical isolates of Escherichia coli expressing an AmpC
J Antimicrob Chemother 2010; 65: 460 464 doi:10.1093/jac/dkp484 Advance publication 22 January 2010 Prevalence and molecular characterization of clinical isolates of Escherichia coli expressing an AmpC
Samples across Bangladesh
 Research Article imedpub Journals http://www.imedpub.com/ ARCHIVES OF CLINICAL MICROBIOLOGY Abstract Bacteriological Quality of Drinking Water Samples across Bangladesh A total of 106 (tube well, deep
Research Article imedpub Journals http://www.imedpub.com/ ARCHIVES OF CLINICAL MICROBIOLOGY Abstract Bacteriological Quality of Drinking Water Samples across Bangladesh A total of 106 (tube well, deep
WELCOME. to the CDS WORKSHOP
 WELCOME to the CDS WORKSHOP Sydney 2010 Excel Spreadsheet for Registration Recent Additions to the CDS Doripenem 10mg disc A carbapenem claimed to be more active against Pseudomonas than Meropenem Daptomycin:
WELCOME to the CDS WORKSHOP Sydney 2010 Excel Spreadsheet for Registration Recent Additions to the CDS Doripenem 10mg disc A carbapenem claimed to be more active against Pseudomonas than Meropenem Daptomycin:
Department of Microbiology, University College of Medical Sciences & Guru Tegh Bahadur Hospital & *
 Indian J Med Res 122, October 2005, pp 330-337 Phenotypic characteristics of clinical isolates of Klebsiella pneumoniae & evaluation of available phenotypic techniques for detection of extended spectrum
Indian J Med Res 122, October 2005, pp 330-337 Phenotypic characteristics of clinical isolates of Klebsiella pneumoniae & evaluation of available phenotypic techniques for detection of extended spectrum
NEXT STEP TO STOP THE SPREAD OF ANTIMICROBIAL RESISTANCE. Patricia Ruiz Garbajosa Servicio de Microbiología Hospital Universitario Ramón y Cajal
 NEXT STEP TO STOP THE SPREAD OF ANTIMICROBIAL RESISTANCE Patricia Ruiz Garbajosa Servicio de Microbiología Hospital Universitario Ramón y Cajal ANTIMICROBIAL RESISTANCE AND HEALTHCARE-ASSOCIATED INFECTION
NEXT STEP TO STOP THE SPREAD OF ANTIMICROBIAL RESISTANCE Patricia Ruiz Garbajosa Servicio de Microbiología Hospital Universitario Ramón y Cajal ANTIMICROBIAL RESISTANCE AND HEALTHCARE-ASSOCIATED INFECTION
Detection of aac(6 0 )-Ib-cr in KPC-producing Klebsiella pneumoniae isolates from Tel Aviv, Israel
 Journal of Antimicrobial Chemotherapy (2009) 64, 718 722 doi:10.1093/jac/dkp272 Advance Access publication 4 August 2009 Detection of aac(6 0 )-Ib-cr in KPC-producing Klebsiella pneumoniae isolates from
Journal of Antimicrobial Chemotherapy (2009) 64, 718 722 doi:10.1093/jac/dkp272 Advance Access publication 4 August 2009 Detection of aac(6 0 )-Ib-cr in KPC-producing Klebsiella pneumoniae isolates from
Chapter 9 Antimicrobial Susceptibility Testing (Agar Disk Diffusion Method)
 Chapter 9 Antimicrobial Susceptibility Testing (Agar Disk Diffusion Method) The disk diffusion method presented in this chapter has been carefully standardized by the National Committee for Clinical Laboratory
Chapter 9 Antimicrobial Susceptibility Testing (Agar Disk Diffusion Method) The disk diffusion method presented in this chapter has been carefully standardized by the National Committee for Clinical Laboratory
DEC PCR KIT. SSI Diagnostica
 DEC PCR KIT SSI Diagnostica 2 DEC PCR Kit for PCR Detection of Diarrhoeagenic E. coli (DEC) for in vitro diagnostic use Application The DEC PCR Kit is for in vitro diagnostic PCR detection of diarrhoeagenic
DEC PCR KIT SSI Diagnostica 2 DEC PCR Kit for PCR Detection of Diarrhoeagenic E. coli (DEC) for in vitro diagnostic use Application The DEC PCR Kit is for in vitro diagnostic PCR detection of diarrhoeagenic
Extended-spectrum and CMY-type b-lactamase-producing Escherichia coli in clinical samples and retail meat from Pittsburgh, USA and Seville, Spain
 ORIGINAL ARTICLE 10.1111/j.1469-0691.2009.03001.x Extended-spectrum and CMY-type b-lactamase-producing Escherichia coli in clinical samples and retail meat from, USA and, Spain Y. Doi 1, D. L. Paterson
ORIGINAL ARTICLE 10.1111/j.1469-0691.2009.03001.x Extended-spectrum and CMY-type b-lactamase-producing Escherichia coli in clinical samples and retail meat from, USA and, Spain Y. Doi 1, D. L. Paterson
ORIGINAL ARTICLE /j x
 ORIGINAL ARTICLE 10.1111/j.1469-0691.2007.01807.x Standard and real-time multiplex PCR methods for detection of trimethoprim resistance dfr genes in large collections of bacteria M. Grape 1, A. Motakefi
ORIGINAL ARTICLE 10.1111/j.1469-0691.2007.01807.x Standard and real-time multiplex PCR methods for detection of trimethoprim resistance dfr genes in large collections of bacteria M. Grape 1, A. Motakefi
64 Mukherjee et al. Int. J. Biosci. 2011
 RESEARCH PAPER OPEN ACCESS Detection of blatem and blactx-m genes by multiplex polymerase chain reaction amongst uropathogenic Escherichia coli strains isolated from hospitalized patients in Kolkata, India
RESEARCH PAPER OPEN ACCESS Detection of blatem and blactx-m genes by multiplex polymerase chain reaction amongst uropathogenic Escherichia coli strains isolated from hospitalized patients in Kolkata, India
Curing antibiotic resistance in vivo. Muhammad Kamruzzaman
 Curing antibiotic resistance in vivo Muhammad Kamruzzaman Occurrence of resistance -Some bacteria are naturally resistant to certain antibiotics -Gene mutation -Horizontal transfer of antibiotic resistance
Curing antibiotic resistance in vivo Muhammad Kamruzzaman Occurrence of resistance -Some bacteria are naturally resistant to certain antibiotics -Gene mutation -Horizontal transfer of antibiotic resistance
generated by the usage of antimicrobial drugs is absent. She is the author of 14 publications in international journals.
 Fernando Baquero Doctor in Medicine, Ramón y Cajal Research Professor in Bacterial Evolution at the Biomedical Research Foundation and Department of Microbiology of the Ramón y Cajal Universty Hospital.
Fernando Baquero Doctor in Medicine, Ramón y Cajal Research Professor in Bacterial Evolution at the Biomedical Research Foundation and Department of Microbiology of the Ramón y Cajal Universty Hospital.
Antimicrobial Agents and Chemotherapy New Data Letter
 AAC Accepted Manuscript Posted Online 15 August 2016 Antimicrob. Agents Chemother. doi:10.1128/aac.01519-16 Copyright 2016, American Society for Microbiology. All Rights Reserved. 1 Antimicrobial Agents
AAC Accepted Manuscript Posted Online 15 August 2016 Antimicrob. Agents Chemother. doi:10.1128/aac.01519-16 Copyright 2016, American Society for Microbiology. All Rights Reserved. 1 Antimicrobial Agents
Molecular susceptibility testing
 Molecular susceptibility testing Dr Andrew Ginn Supervising Scientist Antimicrobial Resistance Reference Laboratory ICPMR, Westmead Hospital Resistance genes Gram negatives Transmissible; e.g. ESBLs, MBLs,
Molecular susceptibility testing Dr Andrew Ginn Supervising Scientist Antimicrobial Resistance Reference Laboratory ICPMR, Westmead Hospital Resistance genes Gram negatives Transmissible; e.g. ESBLs, MBLs,
J. Appl. Environ. Biol. Sci., 5(12) , , TextRoad Publication
 J. Appl. Environ. Biol. Sci., 5(2)262-268, 205 205, TextRoad Publication ISSN: 2090-4274 Journal of Applied Environmental and Biological Sciences www.textroad.com Study of routine antibiotic Resistance
J. Appl. Environ. Biol. Sci., 5(2)262-268, 205 205, TextRoad Publication ISSN: 2090-4274 Journal of Applied Environmental and Biological Sciences www.textroad.com Study of routine antibiotic Resistance
CRE Laboratory Testing and CRE Lab Testing Recommendations in-depth recommendations on CRE laboratory detection
 December 2014 Dear Laboratory Director, The Illinois Department of Public Health (IDPH) amended the Control of Communicable Diseases Code (77 Ill. Adm. Code 690) to require reporting of Carbapenem-Resistant
December 2014 Dear Laboratory Director, The Illinois Department of Public Health (IDPH) amended the Control of Communicable Diseases Code (77 Ill. Adm. Code 690) to require reporting of Carbapenem-Resistant
Antibiotic Resistance : Contribution of Hospital Effluents to the
 Antibiotic Resistance : Contribution of Hospital Effluents to the spread of ATBR in environment T. Stalder, O. Barraud, M. Casellas, M. C. Ploy, C. Dagot PILLS final conference 19-20 sept. 2012 1 The emergence
Antibiotic Resistance : Contribution of Hospital Effluents to the spread of ATBR in environment T. Stalder, O. Barraud, M. Casellas, M. C. Ploy, C. Dagot PILLS final conference 19-20 sept. 2012 1 The emergence
Emergence and persistence of integron structures harbouring VIM genes in the Children s Memorial Health Institute, Warsaw, Poland,
 Journal of Antimicrobial Chemotherapy (2009) 63, 269 273 doi:10.1093/jac/dkn512 Advance Access publication 18 December 2008 Emergence and persistence of integron structures harbouring VIM genes in the
Journal of Antimicrobial Chemotherapy (2009) 63, 269 273 doi:10.1093/jac/dkn512 Advance Access publication 18 December 2008 Emergence and persistence of integron structures harbouring VIM genes in the
Extended-spectrum b-lactamases of Escherichia coli and Klebsiella pneumoniae screened by the VITEK 2 system
 Journal of Medical Microbiology (2011), 60, 756 760 DOI 10.1099/jmm.0.024075-0 Extended-spectrum b-lactamases of Escherichia coli and Klebsiella pneumoniae screened by the VITEK 2 system Maria José Espinar,
Journal of Medical Microbiology (2011), 60, 756 760 DOI 10.1099/jmm.0.024075-0 Extended-spectrum b-lactamases of Escherichia coli and Klebsiella pneumoniae screened by the VITEK 2 system Maria José Espinar,
Disk Diffusion Method for Susceptibility Testing of
 JOURNAL OF CLINICAL MICROBIOLOGY, Aug. 1991, p. 1604-1609 0095-1137/91/081604-06$02.00/0 Copyright 1991, American Society for Microbiology Vol. 29, No. 8 Disk Diffusion Method for Susceptibility Testing
JOURNAL OF CLINICAL MICROBIOLOGY, Aug. 1991, p. 1604-1609 0095-1137/91/081604-06$02.00/0 Copyright 1991, American Society for Microbiology Vol. 29, No. 8 Disk Diffusion Method for Susceptibility Testing
Abstract. Introduction
 ORIGINAL ARTICLE BACTERIOLOGY A sensitive and specific phenotypic assay for detection of metallo-blactamases and KPC in Klebsiella pneumoniae with the use of meropenem disks supplemented with aminophenylboronic
ORIGINAL ARTICLE BACTERIOLOGY A sensitive and specific phenotypic assay for detection of metallo-blactamases and KPC in Klebsiella pneumoniae with the use of meropenem disks supplemented with aminophenylboronic
The Cat s Out of the Bag: Microbiological Investigations of Acute Transfusion Reactions.
 The Cat s Out of the Bag: Microbiological Investigations of Acute Transfusion Reactions. Philippe Lagacé-Wiens, MD FRCPC, DTM&H plagacewiens@sharedhealthmb.ca COI declaration I have no conflicts, real
The Cat s Out of the Bag: Microbiological Investigations of Acute Transfusion Reactions. Philippe Lagacé-Wiens, MD FRCPC, DTM&H plagacewiens@sharedhealthmb.ca COI declaration I have no conflicts, real
Methicillin resistant Staphylococcus aureus (MRSA)- Background and analysis
 Methicillin resistant Staphylococcus aureus ()- Background and analysis Outline Staphylococcus aureus Classification Ecology Commensal vs pathogenic S. aureus Definition Resistance determinant and mechanism
Methicillin resistant Staphylococcus aureus ()- Background and analysis Outline Staphylococcus aureus Classification Ecology Commensal vs pathogenic S. aureus Definition Resistance determinant and mechanism
DOI: /IJAPSA S6NDB
 Microbial source tracking through antibiotic resistance indexing of Escherichia coli to identify source of fecal contamination in drinking water Tambekar DH, Dhote SV 1 and Lamkhade KA 2 1,2 PG Dept. of
Microbial source tracking through antibiotic resistance indexing of Escherichia coli to identify source of fecal contamination in drinking water Tambekar DH, Dhote SV 1 and Lamkhade KA 2 1,2 PG Dept. of
Biofilm Protocol Optimization For Pseudomonas aeruginosa. Introduction. Materials and Methods. Culture Media, Incubation Time, and Biofilm Measurement
 Biofilm Protocol Optimization For Pseudomonas aeruginosa Culture Media, Incubation Time, and Biofilm Measurement Introduction In addition to the conventional arsenal of antibiotic resistance mechanisms
Biofilm Protocol Optimization For Pseudomonas aeruginosa Culture Media, Incubation Time, and Biofilm Measurement Introduction In addition to the conventional arsenal of antibiotic resistance mechanisms
Lauren A. Darling1#, Ann M. Evans1, Kathleen A. Stellrecht1,2, Seela M. Nattanmai1,
 JCM Accepted Manuscript Posted Online 20 September 2017 J. Clin. Microbiol. doi:10.1128/jcm.01185-17 Copyright 2017 American Society for Microbiology. All Rights Reserved. 1 JCM Letter to the Editor Submission
JCM Accepted Manuscript Posted Online 20 September 2017 J. Clin. Microbiol. doi:10.1128/jcm.01185-17 Copyright 2017 American Society for Microbiology. All Rights Reserved. 1 JCM Letter to the Editor Submission
Pathogenic bacteria replicate and persevere in ecological niches called reservoirs
 TYPING METHODS WHAT IS TYPING AND WHAT ARE TYPING METHODS? Pathogenic bacteria replicate and persevere in ecological niches called reservoirs Reservoirs may be humans, including (fellow) patients and healthcare
TYPING METHODS WHAT IS TYPING AND WHAT ARE TYPING METHODS? Pathogenic bacteria replicate and persevere in ecological niches called reservoirs Reservoirs may be humans, including (fellow) patients and healthcare
Investigation of Klebsiella pneumoniae Isolates Producing SHV-12 and SHV-11 β-lactamases in Korean Hospitals
 J. Microbiol. Biotechnol. (2009), 19(2), 000 000 doi: 10.4014/jmb.0808.472 First published online 3 June 2009 Investigation of Klebsiella pneumoniae Isolates Producing SHV-12 and SHV-11 β-lactamases in
J. Microbiol. Biotechnol. (2009), 19(2), 000 000 doi: 10.4014/jmb.0808.472 First published online 3 June 2009 Investigation of Klebsiella pneumoniae Isolates Producing SHV-12 and SHV-11 β-lactamases in
H. Wu, B.-G. Liu, J.-H. Liu, Y.-S. Pan, L. Yuan and G.-Z. Hu
 Phenotypic and molecular characterization of CTX-M-14 extended-spectrum β-lactamase and plasmid-mediated ACT-like AmpC β-lactamase produced by Klebsiella pneumoniae isolates from chickens in Henan Province,
Phenotypic and molecular characterization of CTX-M-14 extended-spectrum β-lactamase and plasmid-mediated ACT-like AmpC β-lactamase produced by Klebsiella pneumoniae isolates from chickens in Henan Province,
Detection and molecular characterization of extended spectrum of beta lactamase (ESBL) producing Escherichia coli
 ISSN: 2319-7706 Volume 2 Number 8 (2013) pp. 196-205 http://www.ijcmas.com Original Research Article Detection and molecular characterization of extended spectrum of beta lactamase (ESBL) producing Escherichia
ISSN: 2319-7706 Volume 2 Number 8 (2013) pp. 196-205 http://www.ijcmas.com Original Research Article Detection and molecular characterization of extended spectrum of beta lactamase (ESBL) producing Escherichia
EU Reference Laboratory for E. coli Department of Veterinary Public Health and Food Safety Unit of Foodborne Zoonoses Istituto Superiore di Sanità
 Detection of Enterotoxigenic Escherichia coli in food by Real Time PCR amplification of the lt, sth, and stp genes, encoding the heat-labile and heat-stable enterotoxins 1. Aim and field of application
Detection of Enterotoxigenic Escherichia coli in food by Real Time PCR amplification of the lt, sth, and stp genes, encoding the heat-labile and heat-stable enterotoxins 1. Aim and field of application
Drug Susceptibility Pattern of Extraintestinal Pathogenic E.Coli Isolated from Various Clinical Specimens
 International Journal of Research Studies in Microbiology and Biotechnology (IJRSMB) Volume 2, Issue 1, 2016, PP 22-27 ISSN 2454-9428 (Online) www.arcjournals.org Drug Susceptibility Pattern of Extraintestinal
International Journal of Research Studies in Microbiology and Biotechnology (IJRSMB) Volume 2, Issue 1, 2016, PP 22-27 ISSN 2454-9428 (Online) www.arcjournals.org Drug Susceptibility Pattern of Extraintestinal
Occurrence and Detection of AmpC β-lactamases among Enterobacteriaceae in a Tertiary Care Centre in Trivandrum, India
 International Journal of Current Microbiology and Applied Sciences ISSN: 2319-7706 Volume 7 Number 08 (2018) Journal homepage: http://www.ijcmas.com Original Research Article https://doi.org/10.20546/ijcmas.2018.708.023
International Journal of Current Microbiology and Applied Sciences ISSN: 2319-7706 Volume 7 Number 08 (2018) Journal homepage: http://www.ijcmas.com Original Research Article https://doi.org/10.20546/ijcmas.2018.708.023
Prevalence of SHV β-lactamases in Escherichia coli
 African Journal of Microbiology Research Vol. 6(26), pp. 5518-5522, 12 July, 2012 Available online at http://www.academicjournals.org/ajmr DOI: 10.5897/AJMR12.1228 ISSN 1996-0808 2012 Academic Journals
African Journal of Microbiology Research Vol. 6(26), pp. 5518-5522, 12 July, 2012 Available online at http://www.academicjournals.org/ajmr DOI: 10.5897/AJMR12.1228 ISSN 1996-0808 2012 Academic Journals
PERANAN MIKROBIOLOGI DALAM DIAGNOSIS PENYAKIT INFEKSI. dr. Agus Eka Darwinata, Ph.D.
 PERANAN MIKROBIOLOGI DALAM DIAGNOSIS PENYAKIT INFEKSI dr. Agus Eka Darwinata, Ph.D. CLINICAL MICROBIOLOGY Clinical microbiology is the discipline of detection, characterization, and quantification of
PERANAN MIKROBIOLOGI DALAM DIAGNOSIS PENYAKIT INFEKSI dr. Agus Eka Darwinata, Ph.D. CLINICAL MICROBIOLOGY Clinical microbiology is the discipline of detection, characterization, and quantification of
Faecal prevalence of extended-spectrum ß-lactamase (ESBL)- producing coliforms in a geriatric population and among haematology patients
 Malaysian J Pathol 2005; 27(2) : 75 81 FAECAL ESBL-PRODUCING COLIFORMS Faecal prevalence of extended-spectrum ß-lactamase (ESBL)- producing coliforms in a geriatric population and among haematology patients
Malaysian J Pathol 2005; 27(2) : 75 81 FAECAL ESBL-PRODUCING COLIFORMS Faecal prevalence of extended-spectrum ß-lactamase (ESBL)- producing coliforms in a geriatric population and among haematology patients
Resistance, Yonsei University College of Medicine, Seoul, Korea; and 2 Department of
 AAC Accepts, published online ahead of print on March 0 Antimicrob. Agents Chemother. doi:./aac.0- Copyright 0, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
AAC Accepts, published online ahead of print on March 0 Antimicrob. Agents Chemother. doi:./aac.0- Copyright 0, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
M. Ben-David 1, O. Hammer 1, A.Shinderman 1, Y. Gluckman- Yavo 1, M. Fridman 1, D. Gohman 1, G. Ingber 1 and E. Zahavy 2
 437 Fast Antibiotic Susceptibility Testing Utilizing a Unique Spectral Intensity Ratio Analysis via Single Fluorescence Membrane Dye Staining and Flow Cytometry M. Ben-David 1, O. Hammer 1, A.Shinderman
437 Fast Antibiotic Susceptibility Testing Utilizing a Unique Spectral Intensity Ratio Analysis via Single Fluorescence Membrane Dye Staining and Flow Cytometry M. Ben-David 1, O. Hammer 1, A.Shinderman
Genotypic and Phenotypic Correlations of Multidrug Resistant Acinetobacter baumannii-
 JCM Accepts, published online ahead of print on 25 August 2010 J. Clin. Microbiol. doi:10.1128/jcm.01585-10 Copyright 2010, American Society for Microbiology and/or the Listed Authors/Institutions. All
JCM Accepts, published online ahead of print on 25 August 2010 J. Clin. Microbiol. doi:10.1128/jcm.01585-10 Copyright 2010, American Society for Microbiology and/or the Listed Authors/Institutions. All
JOHN DEMPSEY HOSPITAL Farmington, Connecticut ANTIBIOTIC SUSCEPTIBILITY PROFILES for INPATIENT Bacterial Isolates
 JOHN DEMPSEY HOSPITAL Farmington, Connecticut 2017 ANTIBIOTIC SUSCEPTIBILITY PROFILES for INPATIENT Bacterial Isolates **GROUPED BY CULTURE SOURCES** (data from 1/1/17 1/1/18) Prepared by: UCHC/JDH Antimicrobial
JOHN DEMPSEY HOSPITAL Farmington, Connecticut 2017 ANTIBIOTIC SUSCEPTIBILITY PROFILES for INPATIENT Bacterial Isolates **GROUPED BY CULTURE SOURCES** (data from 1/1/17 1/1/18) Prepared by: UCHC/JDH Antimicrobial
UNDERGRADUATE RESEARCH SEMESTER/EXPLORATORY GRANT APPLICATION Budget Worksheet
 UNDERGRADUATE RESEARCH SEMESTER/EXPLORATORY GRANT APPLICATION Budget Worksheet BUDGET ITEM Department or College Funds Outside Agency Funds Persona l Funds Undergrad. Research Funds GRAND TOTAL Materials
UNDERGRADUATE RESEARCH SEMESTER/EXPLORATORY GRANT APPLICATION Budget Worksheet BUDGET ITEM Department or College Funds Outside Agency Funds Persona l Funds Undergrad. Research Funds GRAND TOTAL Materials
LABORATORY PROTOCOL. Isolation of ESBL-, AmpC- and carbapenemase-producing E. coli from caecal samples
 LABORATORY PROTOCOL Isolation of ESBL-, AmpC- and carbapenemase-producing E. coli from caecal samples November 2017 Version 5 Henrik Hasman, Yvonne Agersø, Rene Hendriksen, Lina M. Cavaco (DTU Food) and
LABORATORY PROTOCOL Isolation of ESBL-, AmpC- and carbapenemase-producing E. coli from caecal samples November 2017 Version 5 Henrik Hasman, Yvonne Agersø, Rene Hendriksen, Lina M. Cavaco (DTU Food) and
International Journal of Molecular and Clinical Microbiology
 International Journal of Molecular and Clinical Microbiology 6(2) (2016) 687-692 International Journal of Molecular and Clinical Microbiology Prevalence of extended-spectrum ß-lactamase-producing Escherichia
International Journal of Molecular and Clinical Microbiology 6(2) (2016) 687-692 International Journal of Molecular and Clinical Microbiology Prevalence of extended-spectrum ß-lactamase-producing Escherichia
Maurine A. Leverstein-van Hall,* Hetty E. M. Blok, Armand Paauw, Ad C. Fluit, Annet Troelstra, Ellen M. Mascini, Marc J. M. Bonten, and Jan Verhoef
 JOURNAL OF CLINICAL MICROBIOLOGY, Feb. 2006, p. 518 524 Vol. 44, No. 2 0095-1137/06/$08.00 0 doi:10.1128/jcm.44.2.518 524.2006 Copyright 2006, American Society for Microbiology. All Rights Reserved. Extensive
JOURNAL OF CLINICAL MICROBIOLOGY, Feb. 2006, p. 518 524 Vol. 44, No. 2 0095-1137/06/$08.00 0 doi:10.1128/jcm.44.2.518 524.2006 Copyright 2006, American Society for Microbiology. All Rights Reserved. Extensive
FARM MICROBIOLOGY 2008 PART 7: WATER & WASTEWATER MICROBIOLOGY. B. The water supply and the hydrologic cycle.
 FARM MICROBIOLOGY 2008 PART 7: WATER & WASTEWATER MICROBIOLOGY I. Water General and Microbiology. A. Domestic use of water. Drinking, bathing, cleaning, formulating drugs, mediamaking (for bacteriology
FARM MICROBIOLOGY 2008 PART 7: WATER & WASTEWATER MICROBIOLOGY I. Water General and Microbiology. A. Domestic use of water. Drinking, bathing, cleaning, formulating drugs, mediamaking (for bacteriology
4.1.1 Multidrug resistance: Curing and Mechanism
 4.0 Results and Discussion (Part II) 4.1 Results 4.1.1 Multidrug resistance: Curing and Mechanism Bacterial resistance to antibiotics is one of the most critical problems faced by the physicians all over
4.0 Results and Discussion (Part II) 4.1 Results 4.1.1 Multidrug resistance: Curing and Mechanism Bacterial resistance to antibiotics is one of the most critical problems faced by the physicians all over
International Journal of Pharma and Bio Sciences STUDIES ON EFFECT OF CEFOTAXIME AND TERMINALIA CHEBULA ON ESCHERICHIA COLI ABSTRACT
 Research Article Microbiology International Journal of Pharma and Bio Sciences ISSN 0975-6299 STUDIES ON EFFECT OF CEFOTAXIME AND TERMINALIA CHEBULA ON ESCHERICHIA COLI ALPA RABADIA *, S.D. KAMAT AND D.V.
Research Article Microbiology International Journal of Pharma and Bio Sciences ISSN 0975-6299 STUDIES ON EFFECT OF CEFOTAXIME AND TERMINALIA CHEBULA ON ESCHERICHIA COLI ALPA RABADIA *, S.D. KAMAT AND D.V.
Prevalence and spread of extendedspectrum b-lactamase-producing Enterobacteriaceae in Ngaoundere, Cameroon. Abstract. Editor: R.
 RESEARCH NOTE BACTERIOLOGY Prevalence and spread of extendedspectrum b-lactamase-producing Enterobacteriaceae in Ngaoundere, Cameroon C. Lonchel Magoue 1, P. Melin 1, J. Gangoue-Pieboji 2, M.-C. Okomo
RESEARCH NOTE BACTERIOLOGY Prevalence and spread of extendedspectrum b-lactamase-producing Enterobacteriaceae in Ngaoundere, Cameroon C. Lonchel Magoue 1, P. Melin 1, J. Gangoue-Pieboji 2, M.-C. Okomo
Occurrence of TEM & SHV gene in extended spectrum b-lactamases (ESBLs) producing Klebsiella sp. isolated from a tertiary care hospital
 Indian J Med Res 125, February 2007, pp 173-178 Occurrence of TEM & SHV gene in extended spectrum b-lactamases (ESBLs) producing Klebsiella sp. isolated from a tertiary care hospital Prabha Lal, Arti Kapil,
Indian J Med Res 125, February 2007, pp 173-178 Occurrence of TEM & SHV gene in extended spectrum b-lactamases (ESBLs) producing Klebsiella sp. isolated from a tertiary care hospital Prabha Lal, Arti Kapil,
Water Quality Testing II:
 The Biotechnology Education Company Revised and Updated PCR-based Testing for Water Bacterial Contaminants Storage: See Page 3 for specific storage instructions Experiment Objective: The objective of this
The Biotechnology Education Company Revised and Updated PCR-based Testing for Water Bacterial Contaminants Storage: See Page 3 for specific storage instructions Experiment Objective: The objective of this
Pr oject Summar y. Escherichia coli O157 indicator organism. Principal Investigator: Terrance M. Arthur and Mohammad Koohmaraie
 Pr oject Summar y Escherichia coli O157 indicator organism Principal Investigator: Terrance M. Arthur and Mohammad Koohmaraie U.S. Department of Agriculture, Agricultural Research Service Study Completed
Pr oject Summar y Escherichia coli O157 indicator organism Principal Investigator: Terrance M. Arthur and Mohammad Koohmaraie U.S. Department of Agriculture, Agricultural Research Service Study Completed
Microbial Source Tracking in Non-Sewered Catchments
 South East Queensland Healthy Waterways Partnership Microbial Source Tracking in Non-Sewered Catchments Report no.2, Septics Study By Warish Ahmed and Mohammad Katouli January 2007 Prepared for: South
South East Queensland Healthy Waterways Partnership Microbial Source Tracking in Non-Sewered Catchments Report no.2, Septics Study By Warish Ahmed and Mohammad Katouli January 2007 Prepared for: South
Supplementary Information for
 Supplementary Information for Direct, rapid antimicrobial susceptibility test from positive blood cultures based on microscopic imaging analysis J. Choi 1, H. Y. Jeong 2,3, G. Y. Lee 2,4, S. Han 1, S.
Supplementary Information for Direct, rapid antimicrobial susceptibility test from positive blood cultures based on microscopic imaging analysis J. Choi 1, H. Y. Jeong 2,3, G. Y. Lee 2,4, S. Han 1, S.
Plasmid Profile of Bacteria
 International Journal of Current Microbiology and Applied Sciences ISSN: 2319-7706 Volume 7 Number 06 (2018) Journal homepage: http://www.ijcmas.com Original Research Article https://doi.org/10.20546/ijcmas.2018.706.021
International Journal of Current Microbiology and Applied Sciences ISSN: 2319-7706 Volume 7 Number 06 (2018) Journal homepage: http://www.ijcmas.com Original Research Article https://doi.org/10.20546/ijcmas.2018.706.021
Susceptibility Tests
 JOURNAL OF CLINICAL MICROBIOLOGY, Aug. 1982, p. 213-217 Vol. 16, No. 2 0095-1137/82/080213-05$02.00/0 In Vitro Studies with Cefotaxime: Disk Diffusion Susceptibility Tests SMITH SHADOMY* AND EDWARD L.
JOURNAL OF CLINICAL MICROBIOLOGY, Aug. 1982, p. 213-217 Vol. 16, No. 2 0095-1137/82/080213-05$02.00/0 In Vitro Studies with Cefotaxime: Disk Diffusion Susceptibility Tests SMITH SHADOMY* AND EDWARD L.
Antimicrobial Susceptibility Testing Disk Diffusion
 Antimicrobial Susceptibility Testing Disk Diffusion Babak Valizadeh,DCLS Babak_Valizadeh@hotmail.com 1390 / 09 / 10 2011.12.01 1 2 3 CLSI - M02-A10 / 2009 4 CLSI M100-S21 / 2011 Antimicrobial Susceptibility
Antimicrobial Susceptibility Testing Disk Diffusion Babak Valizadeh,DCLS Babak_Valizadeh@hotmail.com 1390 / 09 / 10 2011.12.01 1 2 3 CLSI - M02-A10 / 2009 4 CLSI M100-S21 / 2011 Antimicrobial Susceptibility
Providing clear solutions to microbiological challenges TM. cgmp/iso CLIA. Polyphasic Microbial Identification & DNA Fingerprinting
 Providing clear solutions to microbiological challenges TM Cert. No. 2254.01 Polyphasic Microbial Identification & DNA Fingerprinting Microbial Contamination Tracking & Trending cgmp/iso-17025-2005 CLIA
Providing clear solutions to microbiological challenges TM Cert. No. 2254.01 Polyphasic Microbial Identification & DNA Fingerprinting Microbial Contamination Tracking & Trending cgmp/iso-17025-2005 CLIA
ACCEPTED. Variant CTX-M-59 in a Neonatal Intensive Care Unit in. Division of Infectious Diseases, University of Pittsburgh Medical Center, Pittsburgh,
 AAC Accepts, published online ahead of print on 1 March 00 Antimicrob. Agents Chemother. doi:./aac.0-0 Copyright 00, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
AAC Accepts, published online ahead of print on 1 March 00 Antimicrob. Agents Chemother. doi:./aac.0-0 Copyright 00, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
The Prevalence of TEM-1 gene causing resistance to beta-lactam antibiotics in Klebsiella pneumoniae isolates from clinical samples and plasmid curing
 Available online at www.ijmrhs.com ISSN No: 2319-5886 International Journal of Medical Research & Health Sciences, 2016, 5, 11:557-561 The Prevalence of TEM-1 gene causing resistance to beta-lactam antibiotics
Available online at www.ijmrhs.com ISSN No: 2319-5886 International Journal of Medical Research & Health Sciences, 2016, 5, 11:557-561 The Prevalence of TEM-1 gene causing resistance to beta-lactam antibiotics
Extended Spectrum β-lactamases: Critical Tools of Bacterial Resistance
 Review Article Mahidol University Journal of Pharmaceutical Science 2012; 39 (1), 1-8 Extended Spectrum β-lactamases: Critical Tools of Bacterial Resistance Department of Microbiology, Faculty of Pharmacy,
Review Article Mahidol University Journal of Pharmaceutical Science 2012; 39 (1), 1-8 Extended Spectrum β-lactamases: Critical Tools of Bacterial Resistance Department of Microbiology, Faculty of Pharmacy,
INNOVATIVE SCIENCE DRIVES ANTIBIOTIC LEADERSHIP. Corporate Presentation October 2018 Nasdaq: AKAO
 INNOVATIVE SCIENCE DRIVES ANTIBIOTIC LEADERSHIP Corporate Presentation October 2018 Nasdaq: AKAO Forward Looking Statements This presentation contains forward-looking statements. All statements other than
INNOVATIVE SCIENCE DRIVES ANTIBIOTIC LEADERSHIP Corporate Presentation October 2018 Nasdaq: AKAO Forward Looking Statements This presentation contains forward-looking statements. All statements other than
JOURNAL OF INTERNATIONAL ACADEMIC RESEARCH FOR MULTIDISCIPLINARY Impact Factor 1.393, ISSN: , Volume 2, Issue 8, September 2014
 EVALUATION OF CHROMAGAR-CTX FOR THE DETECTION OF CTX-M-ESBL- PRODUCING GRAM NEGATIVE BACTERIAL ISOLATES RASHA H. ELSHERIF* LAMIAA A. MADKOUR** REHAM A. DWEDAR*** *Clinical Pathology Department, Faculty
EVALUATION OF CHROMAGAR-CTX FOR THE DETECTION OF CTX-M-ESBL- PRODUCING GRAM NEGATIVE BACTERIAL ISOLATES RASHA H. ELSHERIF* LAMIAA A. MADKOUR** REHAM A. DWEDAR*** *Clinical Pathology Department, Faculty
Accurate and Timely Identification of Genes Conferring Resistance to Carbapenems Serves as an Important Tool for Infection Control Measures
 International Journal of Current Microbiology and Applied Sciences ISSN: 2319-7706 Volume 7 Number 05 (2018) Journal homepage: http://www.ijcmas.com Original Research Article https://doi.org/10.20546/ijcmas.2018.705.296
International Journal of Current Microbiology and Applied Sciences ISSN: 2319-7706 Volume 7 Number 05 (2018) Journal homepage: http://www.ijcmas.com Original Research Article https://doi.org/10.20546/ijcmas.2018.705.296
Evaluation of Use of a New Chromogenic Agar in Detection of Urinary Tract Pathogens
 JOURNAL OF CLINICAL MICROBIOLOGY, Apr. 1998, p. 990 994 Vol. 36, No. 4 0095-1137/98/$04.00 0 Copyright 1998, American Society for Microbiology Evaluation of Use of a New Chromogenic Agar in Detection of
JOURNAL OF CLINICAL MICROBIOLOGY, Apr. 1998, p. 990 994 Vol. 36, No. 4 0095-1137/98/$04.00 0 Copyright 1998, American Society for Microbiology Evaluation of Use of a New Chromogenic Agar in Detection of
ORIGINAL ARTICLE /j x
 ORIGINAL ARTICLE 10.1111/j.1469-0691.2006.01645.x Development and clinical validation of a molecular diagnostic assay to detect CTX-M-type b-lactamases in Enterobacteriaceae J. D. D. Pitout 1,2,3, N. Hamilton
ORIGINAL ARTICLE 10.1111/j.1469-0691.2006.01645.x Development and clinical validation of a molecular diagnostic assay to detect CTX-M-type b-lactamases in Enterobacteriaceae J. D. D. Pitout 1,2,3, N. Hamilton
Antimicrobial and Antibacterial Agents
 Antimicrobial and Antibacterial Agents Contents Introduction Classification of antimicrobial drugs Special terms Mechanism of action Resistance of antimicrobial agent Introduction Joseph Lister 1867 -
Antimicrobial and Antibacterial Agents Contents Introduction Classification of antimicrobial drugs Special terms Mechanism of action Resistance of antimicrobial agent Introduction Joseph Lister 1867 -
Adaptation of a Bacterial Growth Detection Assay on the VICTOR Nivo Multimode Plate Reader for Measurement of Antibiotic Effects
 APPLICATION NOTE Multimode Detection Authors: Maria Kuzikov Dr. Bernhard Ellinger Fraunhofer IME ScreeningPort Hamburg, Germany Adaptation of a Bacterial Growth Detection Assay on the VICTOR Nivo Multimode
APPLICATION NOTE Multimode Detection Authors: Maria Kuzikov Dr. Bernhard Ellinger Fraunhofer IME ScreeningPort Hamburg, Germany Adaptation of a Bacterial Growth Detection Assay on the VICTOR Nivo Multimode
Prevalence of metallo-β-lactamases in clinical isolates of Pseudomonas aeruginosa from King Abdulaziz University Hospital in Jeddah
 World Journal of Pharmaceutical Sciences ISSN (Print): 2321-3310; ISSN (Online): 2321-3086 Published by Atom and Cell Publishers All Rights Reserved Available online at: http://www.wjpsonline.org/ Original
World Journal of Pharmaceutical Sciences ISSN (Print): 2321-3310; ISSN (Online): 2321-3086 Published by Atom and Cell Publishers All Rights Reserved Available online at: http://www.wjpsonline.org/ Original
Prevalence of Shiga-Toxin Producing Escherichia Coli in Two Cohorts of Beef Cattle is Associated with Diversity of Microflora and Animal Age
 Prevalence of Shiga-Toxin Producing Escherichia Coli in Two Cohorts of Beef Cattle is Associated with Diversity of Microflora and Animal Age Raies A. Mir 1,2, Sarah M. Markland 1,2, Mauricio Elzo 1, Soohyoun
Prevalence of Shiga-Toxin Producing Escherichia Coli in Two Cohorts of Beef Cattle is Associated with Diversity of Microflora and Animal Age Raies A. Mir 1,2, Sarah M. Markland 1,2, Mauricio Elzo 1, Soohyoun
The American University in Cairo
 The American University in Cairo School of Science and Engineering MOLECULAR CHARACTERIZATION OF EXTENDED- SPECTRUM Beta LACTAMASE (ESBL) PRODUCING Klebsiella pneumoniae AND Escherichia coli AMONG HOSPITALIZED
The American University in Cairo School of Science and Engineering MOLECULAR CHARACTERIZATION OF EXTENDED- SPECTRUM Beta LACTAMASE (ESBL) PRODUCING Klebsiella pneumoniae AND Escherichia coli AMONG HOSPITALIZED
ORIGINAL ARTICLE /j x
 ORIGINAL ARTICLE 10.1111/j.1469-0691.2008.02030.x Metallo-b-lactamase-producing Pseudomonas aeruginosa isolated from a large tertiary centre in Kenya J. D. D. Pitout 1,2,3, G. Revathi 4, B. L Chow 1, B.
ORIGINAL ARTICLE 10.1111/j.1469-0691.2008.02030.x Metallo-b-lactamase-producing Pseudomonas aeruginosa isolated from a large tertiary centre in Kenya J. D. D. Pitout 1,2,3, G. Revathi 4, B. L Chow 1, B.
MIC & Etest. Dr. M. Talebi Ph.D of Bacteriology Tehran University of Medical Sciences
 MIC & Etest Dr. M. Talebi Ph.D of Bacteriology Tehran University of Medical Sciences MIC The minimum inhibitory concentration (MIC) is defined as the lowest concentration of the antimicrobial agent required
MIC & Etest Dr. M. Talebi Ph.D of Bacteriology Tehran University of Medical Sciences MIC The minimum inhibitory concentration (MIC) is defined as the lowest concentration of the antimicrobial agent required
Stability of Antibiotics and Chemotherapeutics in
 APPUED MICROBIOLOGY, Sept. 1970, p. 447-451 Copyright 1970 American Society for Microbiology Vol. 20, No. 3 Printed in U.S.A. Stability of Antibiotics and Chemotherapeutics in Agar Plates KENNETH J. RYAN,
APPUED MICROBIOLOGY, Sept. 1970, p. 447-451 Copyright 1970 American Society for Microbiology Vol. 20, No. 3 Printed in U.S.A. Stability of Antibiotics and Chemotherapeutics in Agar Plates KENNETH J. RYAN,
Search and you will find: detecting extendedspectrum β-lactamase-producing Klebsiella pneumoniae from a patient's immediate environment.
 Royal College of Surgeons in Ireland e-publications@rcsi Clinical Microbiology Articles Department of Clinical Microbiology 1-5-2013 Search and you will find: detecting extendedspectrum β-lactamase-producing
Royal College of Surgeons in Ireland e-publications@rcsi Clinical Microbiology Articles Department of Clinical Microbiology 1-5-2013 Search and you will find: detecting extendedspectrum β-lactamase-producing
Supplementary appendix
 Supplementary appendix This appendix formed part of the original submission and has been peer reviewed. We post it as supplied by the authors. Supplement to: Quan J, Li X, Chen Y, et al. Prevalence of
Supplementary appendix This appendix formed part of the original submission and has been peer reviewed. We post it as supplied by the authors. Supplement to: Quan J, Li X, Chen Y, et al. Prevalence of
Title: Description of the First Escherichia coli Clinical Isolate Harboring Colistin-
 AAC Accepted Manuscript Posted Online 24 October 2016 Antimicrob. Agents Chemother. doi:10.1128/aac.01945-16 Copyright 2016, American Society for Microbiology. All Rights Reserved. 1 2 Title: Description
AAC Accepted Manuscript Posted Online 24 October 2016 Antimicrob. Agents Chemother. doi:10.1128/aac.01945-16 Copyright 2016, American Society for Microbiology. All Rights Reserved. 1 2 Title: Description
DETERMINATION OF THE ID50 VALUES OF ANTIBACTERIAL AGENTS IN AGAR. TAKAKO KATO, SATONORI KURASHIGE, Y. A. CHABBERT* and SUSUMU MITSUHASHI
 1299 DETERMINATION OF THE ID50 VALUES OF ANTIBACTERIAL AGENTS IN AGAR TAKAKO KATO, SATONORI KURASHIGE, Y. A. CHABBERT* and SUSUMU MITSUHASHI Department of Microbiology, School of Medicine, Gunma University,
1299 DETERMINATION OF THE ID50 VALUES OF ANTIBACTERIAL AGENTS IN AGAR TAKAKO KATO, SATONORI KURASHIGE, Y. A. CHABBERT* and SUSUMU MITSUHASHI Department of Microbiology, School of Medicine, Gunma University,
Biochemical Fingerprinting Compared with Ribotyping and Pulsed-Field Gel Electrophoresis of DNA for Epidemiological Typing of Enterococci
 JOURNAL OF CLINICAL MICROBIOLOGY, Nov. 1995, p. 2812 2817 Vol. 33, No. 11 0095-1137/95/$04.00 0 Copyright 1995, American Society for Microbiology Biochemical Fingerprinting Compared with Ribotyping and
JOURNAL OF CLINICAL MICROBIOLOGY, Nov. 1995, p. 2812 2817 Vol. 33, No. 11 0095-1137/95/$04.00 0 Copyright 1995, American Society for Microbiology Biochemical Fingerprinting Compared with Ribotyping and
Rapid Detection of Bacterial Growth in Blood Cultures by Bioluminescent Assay of Bacterial ATP
 JOURNAL OF CLINICAL MICROBIOLOGY, Sept. 1983, p. 521-525 0095-1137/83/090521-05$02.00/O Copyright C 1983, American Society for Microbiology Vol. 18, No. 3 Rapid Detection of Bacterial Growth in Blood Cultures
JOURNAL OF CLINICAL MICROBIOLOGY, Sept. 1983, p. 521-525 0095-1137/83/090521-05$02.00/O Copyright C 1983, American Society for Microbiology Vol. 18, No. 3 Rapid Detection of Bacterial Growth in Blood Cultures
Mochamad Nurcholis. Food Technology Department Agricuktural Technology Faculty Brawijaya University 2013
 Mochamad Nurcholis Food Technology Department Agricuktural Technology Faculty Brawijaya University 2013 Microbial Identification Type Conventional Identification DNA Hybridization PCR Amplification DNA
Mochamad Nurcholis Food Technology Department Agricuktural Technology Faculty Brawijaya University 2013 Microbial Identification Type Conventional Identification DNA Hybridization PCR Amplification DNA
GENOMERA TM MRSA/SA PRODUCTS A NEW ERA IN DIRECT MRSA DNA TESTING. Bringing genetic MRSA results in less than 1 hour!
 GENOMERA TM MRSA/SA PRODUCTS A NEW ERA IN DIRECT MRSA DNA TESTING Bringing genetic MRSA results in less than 1 hour! IMPROVING PATIENT CARE THROUGH EARLIER DETECTION OF MRSA AND SA FAST MRSA AND SA DETECTION
GENOMERA TM MRSA/SA PRODUCTS A NEW ERA IN DIRECT MRSA DNA TESTING Bringing genetic MRSA results in less than 1 hour! IMPROVING PATIENT CARE THROUGH EARLIER DETECTION OF MRSA AND SA FAST MRSA AND SA DETECTION
Rapid Purification of DNA with High PCR Efficiency from Mastitis Bacteria in Milk Using Silicon Carbide
 Rapid Purification of DNA with High PCR Efficiency from Mastitis Bacteria in Milk Using Silicon Carbide T. Richardson 1, B. Lam 2 and Y. Haj-Ahmad 1,2. 1 Brock University, St. Catharines, ON, CANADA, 2
Rapid Purification of DNA with High PCR Efficiency from Mastitis Bacteria in Milk Using Silicon Carbide T. Richardson 1, B. Lam 2 and Y. Haj-Ahmad 1,2. 1 Brock University, St. Catharines, ON, CANADA, 2
Analysis of Shigella strains by Pulsed Field Gel Electrophoresis
 Analysis of Shigella strains by Pulsed Field Gel Electrophoresis Kim N. Pham Public Health Internship Program School of Biological Sciences The University of Texas at Austin Mentor: Ana Maria Valle-Rivera,
Analysis of Shigella strains by Pulsed Field Gel Electrophoresis Kim N. Pham Public Health Internship Program School of Biological Sciences The University of Texas at Austin Mentor: Ana Maria Valle-Rivera,
CHAPTER 24. Immunology
 CHAPTER 24 Diagnostic i Microbiology and Immunology Growth-Dependent Diagnostic Methods Isolation of Pathogens from Clinical Specimens Proper sampling and culture of a suspected pathogen is the most reliable
CHAPTER 24 Diagnostic i Microbiology and Immunology Growth-Dependent Diagnostic Methods Isolation of Pathogens from Clinical Specimens Proper sampling and culture of a suspected pathogen is the most reliable
