Human Merkel Cell Polyomavirus Small T Antigen Is an Oncoprotein
|
|
|
- Jessica Richard
- 6 years ago
- Views:
Transcription
1 Human Merkel Cell Polyomavirus Small T Antigen Is an Oncoprotein Targeting the 4EBP1 Translation Regulator Masahiro Shuda, Hyun Jin Kwun, Huichen Feng, Yuan Chang and Patrick S. Moore Supplemental data 1) Supplemental figures and legends Supplemental Figure 1, related to Figure 1 and Figure 2; Supplemental Figure 2, related to Figure 1; Supplemental Figure 3, related to Figure 3; Supplemental Figure 4, related to Figure 5 and Figure 6; 2) Supplemental methods 3) Supplemental references
2 Supplemental figures and legends mab Epitope MCV 350 Sequence LT Start CM8E6 196 Exon1 Intron1 Exon2 CM5E1 CM2B shrna pan T Frame1 Frame3 Supplemental Figure 1, related to Figure 2 and Figure 3. Mapping of MCV antibodies and shrna sequences. Schematic diagram of the MCV T antigen gene transcript, highlighting sequences targeted for antibodies and shrnas (black bars) used in this study. The overlapping LT and transcripts are encoded by exon1 (frame1) but differential gene expression occurs due to alternative splicing at the exon1intron1 junction (1, 2). CM5E1 mab (red) recognizes a unique peptide sequence (EEYGTLKDYMQSGYNAR) encoded by only. Antibody CM8E6 (yellow) recognizes the Nterminus peptide and detects all isoforms of T antigen encoded by exon1 (3), while CM2B4 (dark green) (4) targets a peptide in exon2 (light green) and does not recognize. Black bars indicate the targeting sequences for pant1 (nucleotide ) and 1 (nucleotide ) shrna, respectively. The pan T1 shrna targets all T antigen isoforms (e.g., both LT and ), while the 1 shrna only targets expression.
3 A CM5E1 () CM2B4 (LT) CM8E6 (pant) B C (kda) Empty vector +LT (genomic) Empty vector +LT (genomic) Empty vector +LT (genomic) LT 57kT 20 αtubulin CM5E1 UISO MS1 MKL2 MKL1 (kda) αtubulin Merged DAPI CM5E1 Supplemental Figure 2, related to Figure 2. CM5E1 antibody specifically detects MCV antigen expression. (A) CM5E1 detects a specific 18 kda MCV antigen protein. The open arrowhead indicates, whereas closed arrowheads indicate LT and 57kT spliced isoforms of MCV LT antigen. 293 cells were transfected with either MCV genomic T antigen (2) or MCV expression constructs and harvested at 48 hours after transfection. Protein expression was examined by immunoblotting using CM5E1 (), CM2B4 (LT/57kT) (4), and CM8E6 (pant) (3) mabs. αtubulin was used as loading control. (B) CM5E1 detection of MCV antigen in MCVinfected MCC cell lines. MCVinfected (MS1, MKL2, and MKL1 (5)) and MCVnegative (UISO (2)) MCC cell lines were lysed and subjected to immunoblot with CM5E1 antibody. (C) MCV is expressed in both nuclear and cytoplasmic compartments. 293 cells expressing MCV were fixed, permeabilized, and stained with CM5E1 to determine the localization of MCV by immunofluorescence. Original magnification, x1000.
4 A Rat1 B Empty vector LT.339 LT.350 C NIH3T3 Empty vector * LT Empty vector LT.339 αtubulin LT.350 D.R7A LT.339 +ST Fold increase R7A.L142A LT Empty.L142A.D44N LT.350 +ST Day Supplemental Figure 3, related to Figure 4. MCV, but not LT, induces transformation of rodent fibroblasts. (A) Transformation assays for Rat1 cells expressing MCV and/or tumorderived LTs (LT.339, LT.350). No increase in colony formation by was noted when tumorderived LTs were coexpressed. Two representative fields are shown for each condition. Original magnification, x100. (B) Expression of and LT proteins in Rat1 cells used in the soft agar assays. LT and proteins were detected with CM2B4 and CM5E1, respectively. Open arrowheads indicate truncated LT bands. Asterisk indicates nonspecific reactivity. Coexpression of LT and was represented by minus/plus (, single gene expression; +, coexpression). (C) MCV induces NIH3T3 colony formation in soft agar. induced soft agar colony formation was unaffected by either PP2Abinding (.R7A and.l142a) or Hsc70binding (.D44N) mutations. Two representative fields are shown for each condition. Original magnification, x100. (D) The effect of and LT on Rat1 cell proliferation. Rat1 cells stably expressing wild type, PP2Abinding mutants of (.R7A,.L142A) or LT were analyzed using a cell proliferation assay in 5% FCS.
5 A B γ β α γ β α γ β α γ β α MKL2 MS1 WaGa sh ctrl sh pant1 sh 1 sh ctrl sh pant1 sh 1 sh ctrl sh pant1 sh 1 p4ebp1 S65 p4ebp1 T70 p4ebp1 T37/T46 Total 4EBP1 LY MK2206 sh ctrl sh pant1 sh 1 pp85 pp70 pp85 pp ps6k T389 Total S6K αtubulin αtubulin * * LT Supplemental Figure 4, related to Figure 7 and Figure 8. (A) Knockdown of in other MCVpositive cell lines reduces 4EBP1 phosphorylation as in MKL1 cells. PanT1 and 1 shrnas reduce 4EBP1 hyperphosphorylation in MCVinfected MKL2, MS1 and WaGa cells (5). Priming site (T37/T46) phosphorylation in hypophosphorylated forms (α and β) of 4EBP1 is relatively preserved while phosphorylation of secondary γ forms (S65 and T70) is markedly reduced by knockdown. (B) and pant antigen knockdown in MKL1 cells reduces S6K phosphorylation at residue T389. Protein samples from Figure 6D were used. Knockdown was confirmed by immunoblotting with CM8E6. Asterisks indicate nonspecific reactivity.
6 Supplemental methods Constructs and transfections. We synthesized codonoptimized (DNA 2.0, Inc) LT and that deleted potential splicing donor and acceptor sites in the sequence. To generate pcdna6 co, codonoptimized sequence was amplified using primers (5 GGG CGG CGA TAT CAC CAT GGA CTT GGT CCT TAA CAG G3 and 5 GGG CCG GCT CGA GTT ATC AGA AGA GAT GCA AGT GAA GCA AGC 3 ) and cloned into pcdna6 vector (Invitrogen) with EcoRV and XhoI restriction sites..r7a,.l142a, and.d44n were generated by sitedirected mutagenesis, with pcdna6 co as a template, using the QuickChange Lightning SiteDirected Mutagenesis kit (Stratagene) and the following primer pairs: co.r7a.f (5 TTG GTC CTT AAC GCG AAA GAA CGA GAG3 )/co.r7a.r (5 CTC TCG TTC TTT CGC GTT AAG GAC CAA3 ), co.l142a.f (5 CAG AAG AAT TGC GCC ACA TGG GGA GAA3 )/co.l142a.r (5 TTC TCC CCA TGT GGC GCA ATT CTT CTG3 ), and.co.d44n.f (5 CTC AAA CAT CAC CCA AAT AAG GGT GGC AAC CC3 )/co.d44n.r (5 GGG TTG CCA CCC TTA TTT GGG TGA TGT TTG AG 3 ). The codonoptimized LT sequence, generated with flanking EcoRV and XhoI sites, was cloned with these sites into a pcdna6 vector to generate pcdna6 LTco. Expression of codonoptimized LT in 293 cells did not produce 57 kt protein, a spliced isoform of LT antigen (2) (data not shown). DNA transfection was carried out using Lipofectamine 2000 reagent (Invitrogen) according to the manufacturer s instructions. Lentivirus construction, production and infection. plvx puro lentiviral vector (Clontech) was modified by replacing the CMV promoter with the elongation factor1 (EF1) promoter using ClaI and AfeI restriction sites to generate plvx EF MCS vector. The EF promoter was amplified by PCR using primers (5 GGG CGG CAT CGA TGG CTC CGG TG CCC GTC AGT G3 and 5 GGG CGG CCC GGG CGC GCC TCG AGC CTG CAG GTG TAC AGC GCT TGT CTA TAT GCA GAA GGA GTT TGC 3 ) from pef1 vector (Invitrogen). Codonoptimized MCV LT and sequences were amplified
7 and cloned into the plvx EF MCS vector described above using AfeI and SmaI sites. Tumorderived LTs, LT.339 and LT.350 were amplified from pcdna6 LTco using forward primer (5 GGG CGG CCG GCC AAC ATG GAT CTG GTA CTG AAT CGC 3 ) and two reverse primers (5 GGG CGG CTT AAT TAA CTA GTC AAG TTC ATA TTC GCA TGC 3 and 5 GGG CGG CTT AAT TAA CTA GTC TGT AAA CTG GGA CGA CG 3 for LT.350 and LT.339, respectively). These two fragments were cloned into the psmpuwhygro lentiviral vector (Cell Biolabs, Inc) using FseI and PacI restriction sites. For lentiviral shrna expression, 6 shrna sequences were designed to target T antigen intron1: 1 (568588; sense strand, 5 CCG GAA GTT GTC TCG CCA GCA TTG TTC AAG AGA CAA TGC TGG CGAGAC AAC TTT TTT TG3 ), 2 (720740; sense strand, 5 CCG GAA CTG ACT ACT GCT TAC TGC ATC AAG AGT GCA GTA AGC AGT AGT CAG TTT TTT TG 3 ), 3 (498520; sense strand, 5 CCG GTA GAT TTT GCA GAG GTC CTG GGT CAC GAG ACC CAG GAC CTC TGC AAA ATC TAT TTT TTT G 3 ), 4 (568590; sense strand, 5 CCG GAA GTT GTC TCG CCA GCA TTG GAG CTC GAG CTC CAA TGC TGG CGA GAC AAC TTT TTT TTT G 3 ), 5, (613635; sense strand, 5 CCG GAA CTG TCT GAC GTG GGG AGG GTG CTC GAG CAC CCT CCC CAC GTC AGA CAG TTT TTT TTT G 3 ) and 6 (660682; sense strand, 5 CCG GTT GGT TTG GAT TTC CTC CTG CTT CTC GAG AAG CAG GAG GAA ATC CAA ACC AAT TTT TTT G 3 ). These were cloned at AgeI and EcoRI sites of plko.1 lentiviral vector to generate 1, 2, 3, 4, 5, and 6 shrnas. Of them, only 1 knocked down MCV expression efficiently in MCVinfected cell lines. An shrna sequence to target raptor (sense strand, 5 CCG GGA CAA CGG CCA CAA GTA CTT CTC GAG AAG TAC TTG TGG CCG TTG TCC TTT TTG 3 ) was also cloned into plko.1. The control plko.1 plasmid, possessing a control nontargeting shorthairpin RNA sequence (sh ctrl), was obtained from Addgene. We previously described two puromycinselectable lentiviral shrnas (sht1.puro and sht2.puro) that target the common T antigen
8 exon1 sequence (5). In the current study, sht1.puro is used and is renamed pant1 for the sake of clarity since it targets all T antigen isoforms. For Lentivirus production, 293FT cells (Invitrogen) were transfected with lentiviral constructs and packaging plasmids, pspax2 and pmd2.g (Addgene) by Lipofectamine 2000 (Invitrogen). One day after transfection, medium containing plasmids and transfection reagent was replaced with fresh 10% FBS DMEM medium. 72 hours after transfection, culture supernatant was collected as infectious lentivirus, aliquoted, and stored at 80 C. Rat1 or NIH3T3 cells seeded in 60 mm dishes were infected with lentivirus in the presence of 1 µg/ml polybrene. Immunoblot analysis. Cells transfected with expression constructs or lentiviruses were lysed in buffer [10 mm TrisHCl (ph 8.0), 0.6% SDS] containing protease inhibitor cocktail (Roche) and phosphatase inhibitors (2 mm NaF and 2 mm NaVO 3 ). Total protein was quantified using BioRad Protein Assay kit (BioRad). The lysates were electrophoresed in SDSPAGE and transferred to nitrocellulose membranes (Amersham). Blots were blocked in 5% (w/v) nonfat dry milk in PBST (PBS with 0.05% Tween 20) for 1 hour. Primary antibodies were incubated overnight at 4 C, followed by antimouse IgGHRP (Amersham) or antirabbit IgGHRP (Amersham) for 1 hour at room temperature. Peroxidase activity was detected using Western Lightning plusecl reagent (Perkin Elmer). The following primary antibodies were used: CM5E1 (1:1000), CM2B4 (1:2500) (4), CM8E6 (1:250) (3), 4EBP1, phospho 4EBP1 T37/T46, phospho 4EBP1 T70, phospho 4EBP1 S65, S6K, phospho S6K T389 (1:1000, Cell Signaling). A monoclonal antibody to αtubulin (1:2500, Sigma) was used for the protein loading control. Cell proliferation Assay. Rat1 cells expressing T antigens were seeded in a 96 well plate (2.0x10 3 cells per 96 well), and cell proliferation was monitored the following day by measuring optical densities.
9 To normalize, OD values were divided by the value of day 1. Foldincrease was calculated to evaluate cell proliferation. Immunofluorescence. 293 cells were spotted on glass slides by Cytospin3 (Shandon), fixed with 10% buffered formalin for 20 min and permeabilized with phosphatebuffered saline (PBS) with 0.1% Triton X100. After blocking, cells were incubated with CM5E1 (1:500 dilution) followed by secondary antibody (Alexa Fluor 568conjugated antimouse, 1:1000, Invitrogen) for 1 hour at room temperature. Stained cells were mounted in aqueous medium containing DAPI (Vector Laboratories, CA). Cells were analyzed using an Olympus AX70 epifluorescence microscope equipped with a Spot RT digital camera. Soft agar colony formation assay. Rat1 or NIH3T3 cells stably expressing MCV T antigens were trypsinized to single cells and counted, suspended in complete medium containing 0.3% agarose (Noble agar; Sigma), and seeded on a 0.6% agar layer in 60 mm dishes ( cells) or 6 well ( cells) plate. After 3 weeks, colonies were stained with crystal violet (0.025% in PBS) and plates were photographed. Stable Rat1 cell lines expressing truncated LT alone (LT.339 and LT.350) (1) or both and truncated LT (LT.339+ and LT.350 +) were established by dual infection, followed by selection with two different selection drugs (puromycin or/and hygromycin).
10 Supplemental References 1. Feng, H., Shuda, M., Chang, Y., and Moore, P.S. Clonal integration of a polyomavirus in human Merkel cell carcinoma. Science. 2008;319(5866): Shuda, M., Feng, H., Kwun, H.J., Rosen, S.T., Gjoerup, O., Moore, P.S., and Chang, Y. T antigen mutations are a human tumorspecific signature for Merkel cell polyomavirus. Proc Natl Acad Sci U S A. 2008;105(42): Kwun, H.J., Guastafierro, A., Shuda, M., Meinke, G., Bohm, A., Moore, P.S., and Chang, Y. The minimum replication origin of merkel cell polyomavirus has a unique large Tantigen loading architecture and requires small Tantigen expression for optimal replication. J Virol. 2009;83(23): Shuda, M., Arora, R., Kwun, H.J., Feng, H., Sarid, R., FernandezFigueras, M.T., Tolstov, Y., Gjoerup, O., Mansukhani, M.M., Swerdlow, S.H., et al. Human Merkel cell polyomavirus infection I. MCV T antigen expression in Merkel cell carcinoma, lymphoid tissues and lymphoid tumors. Int J Cancer. 2009;125(6): Houben, R., Shuda, M., Weinkam, R., Schrama, D., Feng, H., Chang, Y., Moore, P.S., and Becker, J.C. Merkel cell polyomavirusinfected Merkel cell carcinoma cells require expression of viral T antigens. J Virol. 2010;84(14):
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Supplementary Figure 1A A404 Cells +/- Retinoic Acid
 Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
PCR analysis was performed to show the presence and the integrity of the var1csa and var-
 Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of
 Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis
 1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
Supplemental Data Supplemental Figure 1.
 Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR
 Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR
 Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Supplementary. Table 1: Oligonucleotides and Plasmids. complementary to positions from 77 of the SRα '- GCT CTA GAG AAC TTG AAG TAC AGA CTG C
 Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Arabidopsis actin depolymerizing factor AtADF4 mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB
 Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC
 Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells
 Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Hes6. PPARα. PPARγ HNF4 CD36
 SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
Supplementary Figures
 Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
ΔPDD1 x ΔPDD1. ΔPDD1 x wild type. 70 kd Pdd1. Pdd3
 Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Materials Protein synthesis kit. This kit consists of 24 amino acids, 24 transfer RNAs, four messenger RNAs and one ribosome (see below).
 Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH).
 Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Supporting information for Biochemistry, 1995, 34(34), , DOI: /bi00034a013
 Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supplemental material
 Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Disease and selection in the human genome 3
 Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were
 1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
Supporting Information. Copyright Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, 2006
 Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR
 Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Dierks Supplementary Fig. S1
 Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Cat. # Product Size DS130 DynaExpress TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1
 Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
hcd1tg/hj1tg/ ApoE-/- hcd1tg/hj1tg/ ApoE+/+
 ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
Overexpression Normal expression Overexpression Normal expression. 26 (21.1%) N (%) P-value a N (%)
 SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
Supplemental Information. Human Senataxin Resolves RNA/DNA Hybrids. Formed at Transcriptional Pause Sites. to Promote Xrn2-Dependent Termination
 Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Table S1. Bacterial strains (Related to Results and Experimental Procedures)
 Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Lecture 11: Gene Prediction
 Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
ORFs and genes. Please sit in row K or forward
 ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
Supporting Information
 Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
Supplementary Figure 1. The level of pri-mir-8 gradually decreases while those of BR-C and E74 increase during 3rd instar larval development.
 qrt-pcr RT-PCR Relative pri-mir-8 level 1.2 1.0 0.8 0.6 0.4 0.2 0.0 Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) BR-C E74 mtl rrna Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) Supplementary Figure 1.
qrt-pcr RT-PCR Relative pri-mir-8 level 1.2 1.0 0.8 0.6 0.4 0.2 0.0 Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) BR-C E74 mtl rrna Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) Supplementary Figure 1.
strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular
![strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular](/thumbs/72/66361212.jpg) Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Supplemental Table 1. Primers used for PCR.
 Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
RPA-AB RPA-C Supplemental Figure S1: SDS-PAGE stained with Coomassie Blue after protein purification.
 RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
Supporting Information
 Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
NAME:... MODEL ANSWER... STUDENT NUMBER:... Maximum marks: 50. Internal Examiner: Hugh Murrell, Computer Science, UKZN
 COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
Y-chromosomal haplogroup typing Using SBE reaction
 Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Supporting Online Information
 Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Project 07/111 Final Report October 31, Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines
 Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system
 Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
A Genetically Encoded Toolbox for Glycocalyx Engineering: Tunable Control of Cell Adhesion,
 TITLE A Genetically Encoded Toolbox for Glycocalyx Engineering: Tunable Control of Cell Adhesion, Survival, and Cancer Cell Behaviors AUTHORS Carolyn R. Shurer *, Marshall J. Colville *, Vivek K. Gupta,
TITLE A Genetically Encoded Toolbox for Glycocalyx Engineering: Tunable Control of Cell Adhesion, Survival, and Cancer Cell Behaviors AUTHORS Carolyn R. Shurer *, Marshall J. Colville *, Vivek K. Gupta,
SUPPORTING INFORMATION FILE
 Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
SUPPLEMENTARY INFORMATION
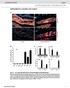 doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
MacBlunt PCR Cloning Kit Manual
 MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
Supporting Information
 Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supplementary Information. Construction of Lasso Peptide Fusion Proteins
 Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Gene synthesis by circular assembly amplification
 Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Supplemental Data. Bennett et al. (2010). Plant Cell /tpc
 BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
Supplementary Information
 Supplementary Information Microbead-based biomimetic synthetic neighbors enhance survival and function of rat pancreatic β-cells Wei Li, a Samuel Lee, b Minglin Ma, a, f Soo Min Kim, b Patrick Guye, c
Supplementary Information Microbead-based biomimetic synthetic neighbors enhance survival and function of rat pancreatic β-cells Wei Li, a Samuel Lee, b Minglin Ma, a, f Soo Min Kim, b Patrick Guye, c
Nongenetic Reprogramming of the Ligand Specificity. of Growth Factor Receptors by Bispecific DNA Aptamers
 Supporting Information For Nongenetic Reprogramming of the Ligand Specificity of Growth Factor Receptors by Bispecific DNA Aptamers Ryosuke Ueki,* Saki Atsuta, Ayaka Ueki and Shinsuke Sando* Department
Supporting Information For Nongenetic Reprogramming of the Ligand Specificity of Growth Factor Receptors by Bispecific DNA Aptamers Ryosuke Ueki,* Saki Atsuta, Ayaka Ueki and Shinsuke Sando* Department
Legends for supplementary figures 1-3
 High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
SUPPORTING INFORMATION
 SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
S4B fluorescence (AU)
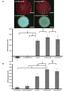 A S4B fluorescence (AU) S4B fluorescence (AU) dsbb csgba csgd dsbb csgba bcsa 5000 * NS NS 4000 * 3000 2000 1000 0 ΔcsgBAΔbcsA ΔcsgDΔdsbBΔbcsA ΔcsgBA ΔdsbBΔcsgBA ΔcsgDΔdsbB B -1000 4000 * * NS 3500 * 3000
A S4B fluorescence (AU) S4B fluorescence (AU) dsbb csgba csgd dsbb csgba bcsa 5000 * NS NS 4000 * 3000 2000 1000 0 ΔcsgBAΔbcsA ΔcsgDΔdsbBΔbcsA ΔcsgBA ΔdsbBΔcsgBA ΔcsgDΔdsbB B -1000 4000 * * NS 3500 * 3000
Lecture 19A. DNA computing
 Lecture 19A. DNA computing What exactly is DNA (deoxyribonucleic acid)? DNA is the material that contains codes for the many physical characteristics of every living creature. Your cells use different
Lecture 19A. DNA computing What exactly is DNA (deoxyribonucleic acid)? DNA is the material that contains codes for the many physical characteristics of every living creature. Your cells use different
Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURB
 Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURBO DNA-free Kit (Ambion). One µg of total RNA was reverse
Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURBO DNA-free Kit (Ambion). One µg of total RNA was reverse
2
 1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
G+C content. 1 Introduction. 2 Chromosomes Topology & Counts. 3 Genome size. 4 Replichores and gene orientation. 5 Chirochores.
 1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer
 TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
Supplementary Figure 1 Tmod3 expression and phosphorylation. (a) Expression of Tmod3 in 3T3-L1 preadipocytes and differentiated adipocytes.
 1 2 3 1 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 Supplementary Figure 1 Tmod3 expression and phosphorylation. (a) Expression of Tmod3 in 3T3-L1 preadipocytes and differentiated
1 2 3 1 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 Supplementary Figure 1 Tmod3 expression and phosphorylation. (a) Expression of Tmod3 in 3T3-L1 preadipocytes and differentiated
Multiplexing Genome-scale Engineering
 Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
SUPPLEMENTAL TABLE S1. Additional descriptions of plasmid constructions and the oligonucleotides used Plasmid or Oligonucleotide
 SUPPLEMENTAL TABLE S1. Additional descriptions of plasmid constructions and the oligonucleotides used Plasmid or Oligonucleotide former/ working Description a designation Plasmids pes213a b pes213-tn5
SUPPLEMENTAL TABLE S1. Additional descriptions of plasmid constructions and the oligonucleotides used Plasmid or Oligonucleotide former/ working Description a designation Plasmids pes213a b pes213-tn5
An engineered tryptophan zipper-type peptide as a molecular recognition scaffold
 SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
SUPPLEMENTARY MATERIALS AND METHODS. E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1)
 SUPPLEMENTARY MATERIALS AND METHODS E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1) dinb::kan (lab stock) derivative was used as wild-type. MG1655 alka tag dinb (2) is
SUPPLEMENTARY MATERIALS AND METHODS E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1) dinb::kan (lab stock) derivative was used as wild-type. MG1655 alka tag dinb (2) is
SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING
 SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING All of the patients and control subjects were sequenced and genotyped in the same way. Shotgun libraries of approximately 250 bp
SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING All of the patients and control subjects were sequenced and genotyped in the same way. Shotgun libraries of approximately 250 bp
evaluated with UAS CLB eliciting UAS CIT -N Libraries increase in the
 Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Expression of Recombinant Proteins
 Expression of Recombinant Proteins Uses of Cloned Genes sequencing reagents (eg, probes) protein production insufficient natural quantities modify/mutagenesis library screening Expression Vector Features
Expression of Recombinant Proteins Uses of Cloned Genes sequencing reagents (eg, probes) protein production insufficient natural quantities modify/mutagenesis library screening Expression Vector Features
Supplemental Information. Target-Mediated Protection of Endogenous. MicroRNAs in C. elegans. Inventory of Supplementary Information
 Developmental Cell, Volume 20 Supplemental Information Target-Mediated Protection of Endogenous MicroRNAs in C. elegans Saibal Chatterjee, Monika Fasler, Ingo Büssing, and Helge Großhans Inventory of Supplementary
Developmental Cell, Volume 20 Supplemental Information Target-Mediated Protection of Endogenous MicroRNAs in C. elegans Saibal Chatterjee, Monika Fasler, Ingo Büssing, and Helge Großhans Inventory of Supplementary
Supplemental Data. Short Article. Transcriptional Regulation of Adipogenesis by KLF4. Kıvanç Birsoy, Zhu Chen, and Jeffrey Friedman
 Cell Metabolism, Volume 7 Supplemental Data Short Article Transcriptional Regulation of Adipogenesis by KLF4 Kıvanç Birsoy, Zhu Chen, and Jeffrey Friedman Supplemental Experimental Procedures Plasmids
Cell Metabolism, Volume 7 Supplemental Data Short Article Transcriptional Regulation of Adipogenesis by KLF4 Kıvanç Birsoy, Zhu Chen, and Jeffrey Friedman Supplemental Experimental Procedures Plasmids
SUPPLEMENTAL DATA SUPPLEMENTAL FIGURE LEGENDS
 SUPPLEMENTAL DATA SUPPLEMENTAL FIGURE LEGENDS SUPPLEMENTAL FIGURE S1. Identification of BmCREC. (A) Amino acid sequences of BmCREC show the peptides identified in LC-MS/MS analysis (marked by red letters
SUPPLEMENTAL DATA SUPPLEMENTAL FIGURE LEGENDS SUPPLEMENTAL FIGURE S1. Identification of BmCREC. (A) Amino acid sequences of BmCREC show the peptides identified in LC-MS/MS analysis (marked by red letters
Efficient genome replication of hepatitis B virus using adenovirus vector: a compact pregenomic RNA-expression unit
 Efficient genome replication of hepatitis B virus using adenovirus vector: a compact pregenomic RNA-expression unit Mariko Suzuki 1, Saki Kondo 1, Manabu Yamasaki 2, Norie Matsuda 2, Akio Nomoto 2, Tetsuro
Efficient genome replication of hepatitis B virus using adenovirus vector: a compact pregenomic RNA-expression unit Mariko Suzuki 1, Saki Kondo 1, Manabu Yamasaki 2, Norie Matsuda 2, Akio Nomoto 2, Tetsuro
Supporting Information
 upporting Information hiota et al..73/pnas.159218 I Materials and Methods Yeast trains. Yeast strains used in this study are described in Table 1. TOM22FLAG, a yeast haploid strain for expression of C-terminally
upporting Information hiota et al..73/pnas.159218 I Materials and Methods Yeast trains. Yeast strains used in this study are described in Table 1. TOM22FLAG, a yeast haploid strain for expression of C-terminally
Engineering D66N mutant using quick change site directed mutagenesis. Harkewal Singh 09/01/2010
 Engineering D66N mutant using quick change site directed mutagenesis Harkewal Singh 09/01/2010 1 1- What is quick change site directed mutagenesis? 2- An overview of the kit contents. 3- A brief information
Engineering D66N mutant using quick change site directed mutagenesis Harkewal Singh 09/01/2010 1 1- What is quick change site directed mutagenesis? 2- An overview of the kit contents. 3- A brief information
Homework. A bit about the nature of the atoms of interest. Project. The role of electronega<vity
 Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
Table S1. Sequences of mutagenesis primers used to create altered rdpa- and sdpa genes
 Supplementary Table and Figures for Structural Basis for the Enantiospecificities of R- and S-Specific Phenoxypropionate/α-Ketoglutarate Dioxygenases by Tina A. Müller, Maria I. Zavodszky, Michael Feig,
Supplementary Table and Figures for Structural Basis for the Enantiospecificities of R- and S-Specific Phenoxypropionate/α-Ketoglutarate Dioxygenases by Tina A. Müller, Maria I. Zavodszky, Michael Feig,
II 0.95 DM2 (RPP1) DM3 (At3g61540) b
 Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
Det matematisk-naturvitenskapelige fakultet
 UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet Exam in: MBV4010 Arbeidsmetoder i molekylærbiologi og biokjemi I MBV4010 Methods in molecular biology and biochemistry I Day of exam: Friday
UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet Exam in: MBV4010 Arbeidsmetoder i molekylærbiologi og biokjemi I MBV4010 Methods in molecular biology and biochemistry I Day of exam: Friday
MCB421 FALL2005 EXAM#3 ANSWERS Page 1 of 12. ANSWER: Both transposon types form small duplications of adjacent host DNA sequences.
 Page 1 of 12 (10pts) 1. There are two mechanisms for transposition used by bacterial transposable elements: replicative (Tn3) and non-replicative (Tn5 and Tn10). Compare and contrast the two mechanisms
Page 1 of 12 (10pts) 1. There are two mechanisms for transposition used by bacterial transposable elements: replicative (Tn3) and non-replicative (Tn5 and Tn10). Compare and contrast the two mechanisms
Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers
 DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
Supplementary Material and Methods
 Supplementary Material and Methods Synaptosomes preparation and RT-PCR analysis. Synaptoneurosome fractions were prepared as previously described in 1. Briefly, rat total brain was homogenized in ice-cold
Supplementary Material and Methods Synaptosomes preparation and RT-PCR analysis. Synaptoneurosome fractions were prepared as previously described in 1. Briefly, rat total brain was homogenized in ice-cold
SUPPLEMENTARY INFORMATION
 doi:1.138/nature11496 Cl. 8 Cl. E93 Rag1 -/- 3H9 + BM Rag1 -/- BM CD CD c-kit c-kit c-kit wt Spleen c-kit B22 B22 IgM IgM IgM Supplementary Figure 1. FACS analysis of single-cell-derived pre-b cell clones.
doi:1.138/nature11496 Cl. 8 Cl. E93 Rag1 -/- 3H9 + BM Rag1 -/- BM CD CD c-kit c-kit c-kit wt Spleen c-kit B22 B22 IgM IgM IgM Supplementary Figure 1. FACS analysis of single-cell-derived pre-b cell clones.
DNA sentences. How are proteins coded for by DNA? Materials. Teacher instructions. Student instructions. Reflection
 DNA sentences How are proteins coded for by DNA? Deoxyribonucleic acid (DNA) is the molecule of life. DNA is one of the most recognizable nucleic acids, a double-stranded helix. The process by which DNA
DNA sentences How are proteins coded for by DNA? Deoxyribonucleic acid (DNA) is the molecule of life. DNA is one of the most recognizable nucleic acids, a double-stranded helix. The process by which DNA
Luo et al. Supplemental Figures and Materials and Methods
 Luo et al. Supplemental Figures and Materials and Methods The supplemental figures demonstrate that nuclear NFAT is situated at PODs, overexpressed PML does not increase NFAT nuclear localization, and
Luo et al. Supplemental Figures and Materials and Methods The supplemental figures demonstrate that nuclear NFAT is situated at PODs, overexpressed PML does not increase NFAT nuclear localization, and
SUPPLEMENTARY INFORMATION. Material and methods
 SUPPLEMENTARY INFORMATION Material and methods Cell culture Human hepatocellular carcinoma (HepG) cells and human embryonic kidney (HEK)93 cells were grown in Dulbecco s modified Eagle s medium (Invitrogen
SUPPLEMENTARY INFORMATION Material and methods Cell culture Human hepatocellular carcinoma (HepG) cells and human embryonic kidney (HEK)93 cells were grown in Dulbecco s modified Eagle s medium (Invitrogen
PCR-based Markers and Cut Flower Longevity in Carnation
 PCRbased Markers and Cut Flower Longevity in Carnation Laura De Benedetti, Luca Braglia, Simona Bruna, Gianluca Burchi *, Antonio Mercuri and Tito Schiva Istituto Sperimentale per la Floricoltura, Corso
PCRbased Markers and Cut Flower Longevity in Carnation Laura De Benedetti, Luca Braglia, Simona Bruna, Gianluca Burchi *, Antonio Mercuri and Tito Schiva Istituto Sperimentale per la Floricoltura, Corso
for Programmed Chemo-enzymatic Synthesis of Antigenic Oligosaccharides
 Supporting Information Design of α-transglucosidases of Controlled Specificity for Programmed Chemo-enzymatic Synthesis of Antigenic Oligosaccharides Elise Champion ±,,,, Isabelle André ±,,, Claire Moulis
Supporting Information Design of α-transglucosidases of Controlled Specificity for Programmed Chemo-enzymatic Synthesis of Antigenic Oligosaccharides Elise Champion ±,,,, Isabelle André ±,,, Claire Moulis
Codon Bias with PRISM. 2IM24/25, Fall 2007
 Codon Bias with PRISM 2IM24/25, Fall 2007 from RNA to protein mrna vs. trna aminoacid trna anticodon mrna codon codon-anticodon matching Watson-Crick base pairing A U and C G binding first two nucleotide
Codon Bias with PRISM 2IM24/25, Fall 2007 from RNA to protein mrna vs. trna aminoacid trna anticodon mrna codon codon-anticodon matching Watson-Crick base pairing A U and C G binding first two nucleotide
PILRα Is a Herpes Simplex Virus-1 Entry Coreceptor That Associates with Glycoprotein B
 Satoh et al. Page S1 Cell, Volume 132 PILRα Is a Herpes Simplex Virus-1 Entry Coreceptor That Associates with Glycoprotein B Takeshi Satoh, Jun Arii, Tadahiro Suenaga, Jing Wang, Amane Kogure, Junji Uehori,
Satoh et al. Page S1 Cell, Volume 132 PILRα Is a Herpes Simplex Virus-1 Entry Coreceptor That Associates with Glycoprotein B Takeshi Satoh, Jun Arii, Tadahiro Suenaga, Jing Wang, Amane Kogure, Junji Uehori,
Primer Design Workshop. École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria
 Primer Design Workshop École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria Scenario You have discovered the presence of a novel endophy5c organism living inside the cells
Primer Design Workshop École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria Scenario You have discovered the presence of a novel endophy5c organism living inside the cells
Promoter 1. Promoter 2. Enhancer 2
 Essays 1. An animal normally has two copies of the Xan gene. The protein made by this gene helps the animal clear petroleum-based pollutants from its body. Tine Xan What we know about the Xan gene is shown
Essays 1. An animal normally has two copies of the Xan gene. The protein made by this gene helps the animal clear petroleum-based pollutants from its body. Tine Xan What we know about the Xan gene is shown
-15 diopter negative lenses in wild-type and homozygous CHRM2-deleted mice, and
 Supplementary Materials Supplementary Figure 1: Myopia induction was performed using uniocular -10 and -15 diopter negative lenses in wild-type and homozygous CHRM2-deleted mice, and results at 2, 4 and
Supplementary Materials Supplementary Figure 1: Myopia induction was performed using uniocular -10 and -15 diopter negative lenses in wild-type and homozygous CHRM2-deleted mice, and results at 2, 4 and
Supplemental Materials and Methods
 Supplemental Materials and Methods Antibodies: Anti-SRF (cat# Sc-335) and anti-igf1r (sc-712) (Santa Cruz Biotech), and anti- ADAM-10 (14-6211) were from e-bioscience, anti-ku70 (cat# MS-329-P) (Labvision),
Supplemental Materials and Methods Antibodies: Anti-SRF (cat# Sc-335) and anti-igf1r (sc-712) (Santa Cruz Biotech), and anti- ADAM-10 (14-6211) were from e-bioscience, anti-ku70 (cat# MS-329-P) (Labvision),
Complexity of the Ruminococcus flavefaciens FD-1 cellulosome reflects an expansion of family-related protein-protein interactions
 Complexity of the Ruminococcus flavefaciens FD-1 cellulosome reflects an expansion of family-related protein-protein interactions Vered Israeli-Ruimy 1,*, Pedro Bule 2,*, Sadanari Jindou 3, Bareket Dassa
Complexity of the Ruminococcus flavefaciens FD-1 cellulosome reflects an expansion of family-related protein-protein interactions Vered Israeli-Ruimy 1,*, Pedro Bule 2,*, Sadanari Jindou 3, Bareket Dassa
Dynamic enhancer-gene body contacts during transcription elongation
 Dynamic enhancer-gene body contacts during transcription elongation Kiwon Lee, Chris C.-S. Hsiung,, Peng Huang, Arjun Raj, *, and Gerd A. Blobel Division of Hematology, The Children s Hospital of Philadelphia,
Dynamic enhancer-gene body contacts during transcription elongation Kiwon Lee, Chris C.-S. Hsiung,, Peng Huang, Arjun Raj, *, and Gerd A. Blobel Division of Hematology, The Children s Hospital of Philadelphia,
An evolutionarily conserved negative feedback mechanism in the hippo pathway reflects functional difference between LATS1 and LATS2
 /, Supplementary Advance Publications Materials 2015 2016 An evolutionarily conserved negative feedback mechanism in the hippo pathway reflects functional difference between LATS1 and LATS2 Supplementary
/, Supplementary Advance Publications Materials 2015 2016 An evolutionarily conserved negative feedback mechanism in the hippo pathway reflects functional difference between LATS1 and LATS2 Supplementary
Supplemental Data. Lin28 Mediates the Terminal Uridylation. of let-7 Precursor MicroRNA. Molecular Cell, Volume 32
 Molecular Cell, Volume 32 Supplemental Data Lin28 Mediates the Terminal Uridylation of let-7 Precursor MicroRNA Inha Heo, Chirlmin Joo, Jun Cho, Minju Ha, Jinju Han, and V. Narry Kim Figure S1. Endogenous
Molecular Cell, Volume 32 Supplemental Data Lin28 Mediates the Terminal Uridylation of let-7 Precursor MicroRNA Inha Heo, Chirlmin Joo, Jun Cho, Minju Ha, Jinju Han, and V. Narry Kim Figure S1. Endogenous
