SUPPLEMENTARY INFORMATION
|
|
|
- Kelley York
- 5 years ago
- Views:
Transcription
1 DOI: /ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation status. (a) GBM cell lines were treated with TNF-α (10ng/ml) for 1h and analyzed for TERT, IL-8 and IκBα expression. Plots depict relative fold change in mrna expression. Data shown represent the mean of 2 independent experiments. (b) Relative Fn14 expression in GBM cell lines carrying either C250T or C228T TERT mutation. Data shown are an average of 2 independent experiments per cell line. Plots depict relative mrna levels. All raw data are shown in Supplementary Table 2. (c) Cytoplasmic and nuclear fractions of untreated or TWEAK-stimulated C250T and C228T GBM cell lines were analyzed for p52 and RelB activation by western blotting with the indicated antibodies. Data shown is representative of two independent experiments. Unprocessed original scans of blots are found in Supplementary figure
2 Supplementary Figure 2 Canonical NF-κB activation does not induce p65 or p52 recruitment to TERT promoter in GBM cells. (a) ChIP analysis of control (Ctrl) or TWEAK treated GBM cell lines depicting enrichment of BLC promoter with indicated antibodies. n=3 independent ChIP assays performed per cell line. Error bars represent S.D. (b) Western blot analysis of untreated or TWEAK-treated C250T-mutant GBM cells transduced with shrnas targeting p52, RelB, NIK or vector control. Data shown is representative of two independent experiments. (c-d) ChIP was performed in control or TNF-α stimulated GBM cell lines. Enrichment of TERT promoter (c) and NF-κB1A promoter (d) DNA fragments in ChIP DNA were measured by quantitative real-time PCR (ChIP-qPCR) and normalized to DNA input. n=3 independent ChIP experiments performed for each cell line and error bars represent S.D. *P < 0.05; **P < 0.01; Student s t-test, two-tailed. All raw data are shown in Supplementary Table 2. Unprocessed original scans of blots are found in Supplementary figure
3 Supplementary Figure 3 Lymphotoxin β receptor (LtβR)-mediated activation of non-canonical NF-κB pathway induces recruitment of NF-κB2 p52 and Pol II to C250T TERT promoter, resulting in enhanced TERT transcription and telomerase function. (a) Cells were treated with agonistic human LTβR antibody for 24h and total cell extracts were analyzed by western blotting with indicated antibodies. Data shown is representative of three independent experiments. Unprocessed original scans of blots are found in Supplementary figure 7. (b) Relative TERT expression of control (Ctrl) or anti-ltβr-treated T98G and U251 cells. Data from one experiment are shown which is representative of 2 independent experiments. (c-d) ChIP was performed in control (Ctrl) or anti- LTβR-treated T98G and U251 cells using p52, p65 or Pol II-specific antibodies and IgG as a negative control. Enrichment of TERT promoter DNA (c) and BLC promoter DNA (d) fragments in ChIP DNA was normalized to DNA input. n=3 independent ChIP experiments per treatment group and cell line. Error bars represent S.D. (e) Proliferation assay of control (Ctrl) or anti-ltβr-treated T98G and U251 cells. Data shown are from 3 independent experiments for each cell line. Error bars represent S.E.M. (f) Relative telomerase activity of T98G and U251 cells that were untreated (Ctrl) or stimulated with LTβR antibody for 1-4 days. Plots represent mean ± S.E.M. Data shown are from 3 independent experiments for each cell line. (g) Relative TERT expression of T98G and U251 cells treated with si-control (Ctrl), si-nf-κb2 or si-relb. Data shown represent the mean of 2 independent experiments. All statistical analyses were performed using Student s t-test (two-tailed): *P < 0.05; **P < 0.01; ***P < For raw data, refer to Supplementary Table
4 Supplementary Figure 4 Purified p52 protein binds C250T TERT promoter through its Rel homology domain. (a) Recombinant GST-tagged p52 and GST proteins were analyzed for binding to HIV-κB and C250T TERT promoter DNA-labeled probes with EMSA (right panel). Coomassiestained SDS-PAGE of recombinant p52 protein (left panel). EMSA shown is representative of three independent experiments. (b) EMSA analysis of recombinant wild-type (WT) and mutant p52 (carrying mutation in 2 amino acid residues of Rel-homology domain) proteins binding to HIV-κB and C250T TERT promoter DNA-labeled probes (right panel). Data shown is representative of two independent experiments. Coomassie-stained SDS-PAGE showing amount of WT and mutant p52 proteins used for EMSA analysis (left panel). 4
5 Supplementary Figure 5 Constitutive expression of NF-κB-inducing kinase (NIK) results in transcriptional activation of C250T TERT promoter, which promotes the telomerase activity of GBM cells. (a) T98G and U251 cells were transfected with vector, human NIK wild-type (WT) or kinase-inactive mutant NIK (KK) expression plasmids and total cell lysates were analyzed by western blotting with the indicated antibodies. Data shown is representative of three independent experiments. Original scans of blots are shown in Supplementary figure 7. (b) ChIP was performed in control (Ctrl) or NIK WT-overexpressing T98G and U251 cells using p52, p65 or Pol II-specific antibodies and IgG as a negative control. Enrichment of TERT promoter DNA fragments in ChIP DNA was normalized to input. n= 3 independent ChIP experiments per cell type. Error bars represent S.D. (c) Relative telomerase activity of T98G and U251 cells transfected with vector, NIK WT or NIK KK constructs. Data from one experiment is shown which is representative of 3 and 2 independent experiments for T98G and U251 cells respectively. (d) ChIP analysis of control or NIK WT-overexpressing T98G and U251 cells depicting enrichment of BLC promoter with indicated antibodies. n= 3 independent ChIP experiments per cell type and error bars represent S.D. (e) Luciferase reporter assays were performed in 293T HEK cells that were co-transfected with empty vector or human NIK expression plasmid and the pgl3 basic reporter vector or pgl3 vector containing either the WT TERT promoter region (-340 to -55) or TERT promoter with C250T mutation (-340 to -55). Data shown represent the mean luciferase activity from 2 independent experiments. (f) Luciferase reporter assays were performed in 293T HEK cells that were transfected with pgl3 empty reporter vector or pgl3 vector containing either the WT TERT promoter region or TERT promoter with C250T mutation and subsequently untreated or stimulated with TWEAK (30ng/ml) for 1D. Luciferase assay data shown are from one of two independent experiments. (g-h) Vector- or NIK WT-expressing T98G cells were transfected with si-ctrl, si-relb or si-nf-κb2 and analyzed for relative TERT expression (average fold change ± S.D.; n=3 independent experiments) (g) and western blot analysis (h). Western blot data is representative of two independent experiments. *P < 0.05;***P < 0.001; Student s t-test, two-tailed. All raw data are shown in Supplementary Table 2. Unprocessed original scans of blots are found in Supplementary figure
6 Supplementary Figure 6 Ectopic NIK expression enhances the in vivo tumorigenicity of C250T GBM cells through p52 activation. (a) T98G cells were stably transduced with lentiviral expression constructs for vector, NIK WT and NIK WT in combination with shrna targeting p52 (NIK sh-p52) and total cell lysates were analyzed by western blotting with the indicated antibodies. Representative image from two independent experiments is shown. Unprocessed original scans of blots are found in Supplementary figure 7. (b) 5 NOD-SCID mice were injected subcutaneously with T98G cells expressing vector (red arrows), NIK WT (white arrows) or NIK WT sh-p52 (black arrows) and the tumor growth of these cells in the xenograft mouse model after 10 weeks are depicted. (c) Representative immunohistochemical staining of T98G NIK WT xeonograft tumors with the indicated antibodies. Scale bars, 100μm. (d) Relative telomerase activity of T98G cells expressing vector, NIK WT or NIK sh-p52. Data shown represent n =3 independent experiments here. Error bars represent S.E.M. *P < 0.05; Student s t-test, two-tailed. All raw data are shown in Supplementary Table
7 Vector Flag-p52 ETS1/2 Flag IP Flag ETS1/2 Input Flag Actin Figure 6a Supplementary Figure 7 Unprocessed western blot scans. 7
8 LN444 LN444 D247mg D247mg Cytoplasmic Nuclear Cytoplasmic Nuclear U251 U251 LN229 LN229 Cytoplasmic Nuclear Cytoplasmic Nuclear p100 p RelA RelB GAPDH TRF Supplementary figure 1c Supplementary Figure 7 continued 8
9 Tweak p100 p52 RelB Actin Supplementary figure 2b Supplementary Figure 7 continued 9
10 T98G U251 Ctrl Anti-LtβR 24h Ctrl Anti-LtβR 24h p100 p52 Actin Supplementary figure 3a Supplementary Figure 7 continued 10
11 Left panel NIK WT - NIK KK - T98G Right panel NIK WT NIK KK - - U NIK NIK p100 p52 Actin p100 p52 Actin 40 Supplementary figure 5a Supplementary Figure 7 continued 11
12 NIK 160 p100 p52 RelB RelA Actin Supplementary figure 5h Supplementary Figure 7 continued 12
13 T98G Vector NIK WT NIK sh-p52 NIK p100 p52 Actin Supplementary figure 6a Supplementary Figure 7 continued 13
14 qpcr primer sequences Gene Name Forward primer (5 - - > 3 ) Forward primer (5 - - > 3 ) Actin CCA TCA CGA TGC CAG TGG TA CCA TCA CGA TGC CAG TGG TA TERT CCA AGT TCC TGC ACT GGC TGA TTC CCG ATG CTG CCT GAC NFkB2 GAC AAG GTG CAG AAA GAT GAC AT TCC GGA ACA CAA TGG CAT ACT RelB AAA CAC ATC AGA GCT GCG GA CCC TGC TGA ACA CCA CTG AT IL8 GCCAACACAGAAATTATTGTAAAGCTT CCTCTGCACCCAGTTTTCCTT IkBα CGCACCTCCACTCCATCC AGCCATGGATAGAGGCTAAGTGTA ETS1 AGACCCTCTCCAGACAGACA GGCGATCACAACTATCGTAGCT ETS2 CCCAGTGGAGCAAGGCAAA AACTCCCATCCGTCTCCAGT Fn14 CCAAGCTCCTCCAACCACAA TGGGGCCTAGTGTCAAGTCT ChIP- qpcr primer sequences Gene Promoter Forward primer (5 - - > 3 ) Reverse primer (5 - - > 3 ) TERT AGGGCCTCCACATCATGGCCCC TCCCAGGGTGCAGGGACGCCAG BLC CTGCGTTGCCTTGACAGTTC GTCGAATGGGCCATCTCTCA TERT GTCGAGTGGACACGGTGAT AAGTTTATGCAAACTGGACAGGA CRISPR grna sequences grna2- top grna2-bottom grna8-top grna8-bottom 5 -CACCG AGGGGCTGGGAGGGCCCGGA-3 5 -AAACTCC GGGCCCTCCCAGCCCCT C CACCGAAACTCGCGCCGCGAGGAGA AAACTCTCCTCGCGGCGCGAGTTTC-3 Supplementary Table 1 Primer sequences for qpcr and ChIP, and CRISPR grna sequences. Supplementary Table 2 Statistics source data. 14
Hes6. PPARα. PPARγ HNF4 CD36
 SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Supplementary Figure 1A A404 Cells +/- Retinoic Acid
 Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of
 Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells
 Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Overexpression Normal expression Overexpression Normal expression. 26 (21.1%) N (%) P-value a N (%)
 SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
Supplementary. Table 1: Oligonucleotides and Plasmids. complementary to positions from 77 of the SRα '- GCT CTA GAG AAC TTG AAG TAC AGA CTG C
 Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supplemental Data Supplemental Figure 1.
 Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Supplementary Figures
 Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Dierks Supplementary Fig. S1
 Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis
 1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR
 Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
SUPPLEMENTARY INFORMATION
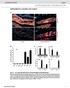 doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
PCR analysis was performed to show the presence and the integrity of the var1csa and var-
 Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR
 Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Lacombe et al. Supplemental material ms INS-BR-TR-2
 Supplemental Figure 1: Rabbit α-gla antibodies specifically recognize carboxylated VKD proteins. (A) HEK293 cells were transfected with plasmids encoding GGCX-FLAG, PT- FLAG or tagged proline rich Gla
Supplemental Figure 1: Rabbit α-gla antibodies specifically recognize carboxylated VKD proteins. (A) HEK293 cells were transfected with plasmids encoding GGCX-FLAG, PT- FLAG or tagged proline rich Gla
ΔPDD1 x ΔPDD1. ΔPDD1 x wild type. 70 kd Pdd1. Pdd3
 Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
RPA-AB RPA-C Supplemental Figure S1: SDS-PAGE stained with Coomassie Blue after protein purification.
 RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC
 Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Arabidopsis actin depolymerizing factor AtADF4 mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB
 Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Supplemental Information. Human Senataxin Resolves RNA/DNA Hybrids. Formed at Transcriptional Pause Sites. to Promote Xrn2-Dependent Termination
 Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
hcd1tg/hj1tg/ ApoE-/- hcd1tg/hj1tg/ ApoE+/+
 ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
Supplementary Figure 1
 Supplementary Figure 1 PTEN promotes virus-induced expression of IFNB1 and its downstream genes. (a) Quantitative RT-PCR analysis of IFNB1 mrna (left) and ELISA of IFN-β (right) in HEK 293 cells (2 10
Supplementary Figure 1 PTEN promotes virus-induced expression of IFNB1 and its downstream genes. (a) Quantitative RT-PCR analysis of IFNB1 mrna (left) and ELISA of IFN-β (right) in HEK 293 cells (2 10
Supporting Information
 Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR
 Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Table S1. Alteration of ZNF322A and FBXW7 protein expression levels in relation to clinicopathological parameters in 135 lung cancer patients.
 SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A and FBXW7 protein expression levels in relation to clinicopathological parameters in 135 lung cancer patients. ZNF322A Normal Overexpression expression
SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A and FBXW7 protein expression levels in relation to clinicopathological parameters in 135 lung cancer patients. ZNF322A Normal Overexpression expression
Table S1. Sequences of mutagenesis primers used to create altered rdpa- and sdpa genes
 Supplementary Table and Figures for Structural Basis for the Enantiospecificities of R- and S-Specific Phenoxypropionate/α-Ketoglutarate Dioxygenases by Tina A. Müller, Maria I. Zavodszky, Michael Feig,
Supplementary Table and Figures for Structural Basis for the Enantiospecificities of R- and S-Specific Phenoxypropionate/α-Ketoglutarate Dioxygenases by Tina A. Müller, Maria I. Zavodszky, Michael Feig,
Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector.
 Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector. (a) Western blotting analysis and (b) qpcr analysis of eif6 expression in HEK293 T cells transfected with either
Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector. (a) Western blotting analysis and (b) qpcr analysis of eif6 expression in HEK293 T cells transfected with either
Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were
 1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
Disease and selection in the human genome 3
 Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Fig. S1: Nkx2.2 is specifically deleted in the intestinal epithelium.
 Fig. S1: Nkx2.2 is specifically deleted in the intestinal epithelium. PCR of representative tissues with oligonucleotides to identify recombination of the Nkx2.2flox allele. Nkx2.2 is specifically deleted
Fig. S1: Nkx2.2 is specifically deleted in the intestinal epithelium. PCR of representative tissues with oligonucleotides to identify recombination of the Nkx2.2flox allele. Nkx2.2 is specifically deleted
The Acute Phase Protein Orosomucoid Regulates Food Intake and Energy Homeostasis via Leptin Receptor Signaling Pathway. Shanghai , China
 The Acute Phase Protein Orosomucoid Regulates Food Intake and Energy Homeostasis via Leptin Receptor Signaling Pathway Yang Sun, 1 Yili Yang, 2 Zhen Qin, 1 Jinya Cai, 3 Xiuming Guo, 1 Yun Tang, 3 Jingjing
The Acute Phase Protein Orosomucoid Regulates Food Intake and Energy Homeostasis via Leptin Receptor Signaling Pathway Yang Sun, 1 Yili Yang, 2 Zhen Qin, 1 Jinya Cai, 3 Xiuming Guo, 1 Yun Tang, 3 Jingjing
Supplementary Fig. 1. Isolation and in vitro expansion of EpCAM + cholangiocytes. For collagenase perfusion, enzyme solution was injected from the
 Supplementary Fig. 1. Isolation and in vitro expansion of EpCAM + cholangiocytes. For collagenase perfusion, enzyme solution was injected from the portal vein for digesting adult livers, whereas it was
Supplementary Fig. 1. Isolation and in vitro expansion of EpCAM + cholangiocytes. For collagenase perfusion, enzyme solution was injected from the portal vein for digesting adult livers, whereas it was
Supporting Online Information
 Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Supplementary data 1. Prmers and probes used in Taqman real-time PCR.
 Supplementary data 1. Prmers and probes used in Taqman real-time PCR. PIG-T Forward Primer: gatctgcctcacgtgcactgt Reverse Primer: aggttcgggtgaggagattgt Probe: 6FAMTGG CCG TGT GCT ATG GCT CCT TCTAMRA PIG-U
Supplementary data 1. Prmers and probes used in Taqman real-time PCR. PIG-T Forward Primer: gatctgcctcacgtgcactgt Reverse Primer: aggttcgggtgaggagattgt Probe: 6FAMTGG CCG TGT GCT ATG GCT CCT TCTAMRA PIG-U
Supplementary Information. Construction of Lasso Peptide Fusion Proteins
 Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
supplementary information
 DOI: 10.1038/ncb2172 Figure S1 p53 regulates cellular NADPH and lipid levels via inhibition of G6PD. (a) U2OS cells stably expressing p53 shrna or a control shrna were transfected with control sirna or
DOI: 10.1038/ncb2172 Figure S1 p53 regulates cellular NADPH and lipid levels via inhibition of G6PD. (a) U2OS cells stably expressing p53 shrna or a control shrna were transfected with control sirna or
PILRα Is a Herpes Simplex Virus-1 Entry Coreceptor That Associates with Glycoprotein B
 Satoh et al. Page S1 Cell, Volume 132 PILRα Is a Herpes Simplex Virus-1 Entry Coreceptor That Associates with Glycoprotein B Takeshi Satoh, Jun Arii, Tadahiro Suenaga, Jing Wang, Amane Kogure, Junji Uehori,
Satoh et al. Page S1 Cell, Volume 132 PILRα Is a Herpes Simplex Virus-1 Entry Coreceptor That Associates with Glycoprotein B Takeshi Satoh, Jun Arii, Tadahiro Suenaga, Jing Wang, Amane Kogure, Junji Uehori,
Supporting Information
 Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/9/eaat5401/dc1 Supplementary Materials for GLK-IKKβ signaling induces dimerization and translocation of the AhR-RORγt complex in IL-17A induction and autoimmune
advances.sciencemag.org/cgi/content/full/4/9/eaat5401/dc1 Supplementary Materials for GLK-IKKβ signaling induces dimerization and translocation of the AhR-RORγt complex in IL-17A induction and autoimmune
Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and
 Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
Site-Directed Mutagenesis. Mutations in four Smad4 sites of mouse Gat1 promoter
 Supplement Supporting Materials and Methods Site-Directed Mutagenesis. Mutations in four Smad4 sites of mouse Gat1 promoter were independently generated using a two-step PCR method. The Smad4 binding site
Supplement Supporting Materials and Methods Site-Directed Mutagenesis. Mutations in four Smad4 sites of mouse Gat1 promoter were independently generated using a two-step PCR method. The Smad4 binding site
An evolutionarily conserved negative feedback mechanism in the hippo pathway reflects functional difference between LATS1 and LATS2
 /, Supplementary Advance Publications Materials 2015 2016 An evolutionarily conserved negative feedback mechanism in the hippo pathway reflects functional difference between LATS1 and LATS2 Supplementary
/, Supplementary Advance Publications Materials 2015 2016 An evolutionarily conserved negative feedback mechanism in the hippo pathway reflects functional difference between LATS1 and LATS2 Supplementary
SUPPORTING INFORMATION FILE
 Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2271 Supplementary Figure a! WM266.4 mock WM266.4 #7 sirna WM266.4 #10 sirna SKMEL28 mock SKMEL28 #7 sirna SKMEL28 #10 sirna WM1361 mock WM1361 #7 sirna WM1361 #10 sirna 9 WM266. WM136
DOI: 10.1038/ncb2271 Supplementary Figure a! WM266.4 mock WM266.4 #7 sirna WM266.4 #10 sirna SKMEL28 mock SKMEL28 #7 sirna SKMEL28 #10 sirna WM1361 mock WM1361 #7 sirna WM1361 #10 sirna 9 WM266. WM136
Supplemental material
 Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Supporting information for Biochemistry, 1995, 34(34), , DOI: /bi00034a013
 Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Multiplexing Genome-scale Engineering
 Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
Table S1. Bacterial strains (Related to Results and Experimental Procedures)
 Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3562 In the format provided by the authors and unedited. Supplementary Figure 1 Glucose deficiency induced FH-ATF2 interaction. In b-m, immunoblotting or immunoprecipitation analyses were
DOI: 10.1038/ncb3562 In the format provided by the authors and unedited. Supplementary Figure 1 Glucose deficiency induced FH-ATF2 interaction. In b-m, immunoblotting or immunoprecipitation analyses were
Supplementary Figure 1. The level of pri-mir-8 gradually decreases while those of BR-C and E74 increase during 3rd instar larval development.
 qrt-pcr RT-PCR Relative pri-mir-8 level 1.2 1.0 0.8 0.6 0.4 0.2 0.0 Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) BR-C E74 mtl rrna Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) Supplementary Figure 1.
qrt-pcr RT-PCR Relative pri-mir-8 level 1.2 1.0 0.8 0.6 0.4 0.2 0.0 Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) BR-C E74 mtl rrna Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) Supplementary Figure 1.
Materials Protein synthesis kit. This kit consists of 24 amino acids, 24 transfer RNAs, four messenger RNAs and one ribosome (see below).
 Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
SUPPLEMENTAL DATA SUPPLEMENTAL FIGURE LEGENDS
 SUPPLEMENTAL DATA SUPPLEMENTAL FIGURE LEGENDS SUPPLEMENTAL FIGURE S1. Identification of BmCREC. (A) Amino acid sequences of BmCREC show the peptides identified in LC-MS/MS analysis (marked by red letters
SUPPLEMENTAL DATA SUPPLEMENTAL FIGURE LEGENDS SUPPLEMENTAL FIGURE S1. Identification of BmCREC. (A) Amino acid sequences of BmCREC show the peptides identified in LC-MS/MS analysis (marked by red letters
Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan).
 1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
HEAHKFLKVEFPARVRSSQATYEIQFGHLQRPTHYNTSWDWARFEVWAHRWMDLSEHGFG Sbjct: 805 HEAHKFLKVEFPARVRSSQATYEIQFGHLQRPTHYNTSWDWARFEVWAHRWMDLSEHGFG 864
 Supplementary Fig S1 a gi 27923805 sp Q9NTJ4 MAN2C1HUMAN Alpha-mannosidase 2C1 (Alpha-D-mannoside mannohydrolase) (Mannosidase alpha class 2C member 1) (Alpha mannosidase 6A8B) Query: 1 HEAHKFLKVEFPARVRSSQATYEIQFGHLQRPTHYNTSWDWARFEVWAHRWMDLSEHGFG
Supplementary Fig S1 a gi 27923805 sp Q9NTJ4 MAN2C1HUMAN Alpha-mannosidase 2C1 (Alpha-D-mannoside mannohydrolase) (Mannosidase alpha class 2C member 1) (Alpha mannosidase 6A8B) Query: 1 HEAHKFLKVEFPARVRSSQATYEIQFGHLQRPTHYNTSWDWARFEVWAHRWMDLSEHGFG
Supporting Information. Copyright Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, 2006
 Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
(a) Immunoblotting to show the migration position of Flag-tagged MAVS
 Supplementary Figure 1 Characterization of six MAVS isoforms. (a) Immunoblotting to show the migration position of Flag-tagged MAVS isoforms. HEK293T Mavs -/- cells were transfected with constructs expressing
Supplementary Figure 1 Characterization of six MAVS isoforms. (a) Immunoblotting to show the migration position of Flag-tagged MAVS isoforms. HEK293T Mavs -/- cells were transfected with constructs expressing
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
supplementary information
 DOI: 10.1038/ncb2116 Figure S1 CDK phosphorylation of EZH2 in cells. (a) Comparison of candidate CDK phosphorylation sites on EZH2 with known CDK substrates by multiple sequence alignments. (b) CDK1 and
DOI: 10.1038/ncb2116 Figure S1 CDK phosphorylation of EZH2 in cells. (a) Comparison of candidate CDK phosphorylation sites on EZH2 with known CDK substrates by multiple sequence alignments. (b) CDK1 and
Supplemental Data. Bennett et al. (2010). Plant Cell /tpc
 BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
SUPPLEMENTAL FIGURES AND TABLES
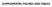 SUPPLEMENTAL FIGURES AND TABLES A B Flag-ALDH1A1 IP: α-ac HEK293T WT 91R 128R 252Q 367R 41/ 419R 435R 495R 412R C Flag-ALDH1A1 NAM IP: HEK293T + + - + D NAM - + + E Relative ALDH1A1 activity 1..8.6.4.2
SUPPLEMENTAL FIGURES AND TABLES A B Flag-ALDH1A1 IP: α-ac HEK293T WT 91R 128R 252Q 367R 41/ 419R 435R 495R 412R C Flag-ALDH1A1 NAM IP: HEK293T + + - + D NAM - + + E Relative ALDH1A1 activity 1..8.6.4.2
SUPPLEMENTARY EXPEMENTAL PROCEDURES
 SUPPLEMENTARY EXPEMENTAL PROCEDURES Plasmids- Total RNAs were extracted from HeLaS3 cells and reverse-transcribed using Superscript III Reverse Transcriptase (Invitrogen) to obtain DNA template for the
SUPPLEMENTARY EXPEMENTAL PROCEDURES Plasmids- Total RNAs were extracted from HeLaS3 cells and reverse-transcribed using Superscript III Reverse Transcriptase (Invitrogen) to obtain DNA template for the
Supplementary Figure 1. TRIM9 does not affect AP-1, NF-AT or ISRE activity. (a,b) At 24h post-transfection with TRIM9 or vector and indicated
 Supplementary Figure 1. TRIM9 does not affect AP-1, NF-AT or ISRE activity. (a,b) At 24h post-transfection with TRIM9 or vector and indicated reporter luciferase constructs, HEK293T cells were stimulated
Supplementary Figure 1. TRIM9 does not affect AP-1, NF-AT or ISRE activity. (a,b) At 24h post-transfection with TRIM9 or vector and indicated reporter luciferase constructs, HEK293T cells were stimulated
Regulation of transcription by the MLL2 complex and MLL complex-associated AKAP95
 Supplementary Information Regulation of transcription by the complex and MLL complex-associated Hao Jiang, Xiangdong Lu, Miho Shimada, Yali Dou, Zhanyun Tang, and Robert G. Roeder Input HeLa NE IP lot:
Supplementary Information Regulation of transcription by the complex and MLL complex-associated Hao Jiang, Xiangdong Lu, Miho Shimada, Yali Dou, Zhanyun Tang, and Robert G. Roeder Input HeLa NE IP lot:
Supplementary Fig. 1 Kinetics of appearence of the faster migrating form of Bcl-10.
 α-cd3 + α-cd28: Time (min): + + + + + + + + + 0 5 15 30 60 120 180 240 300 360 360 n.s. Supplementary Fig. 1 Kinetics of appearence of the faster migrating form of. Immunoblot of lysates from Jurkat cells
α-cd3 + α-cd28: Time (min): + + + + + + + + + 0 5 15 30 60 120 180 240 300 360 360 n.s. Supplementary Fig. 1 Kinetics of appearence of the faster migrating form of. Immunoblot of lysates from Jurkat cells
Supplementary Figure 1. GST pull-down analysis of the interaction of GST-cIAP1 (A, B), GSTcIAP1
 Legends Supplementary Figure 1. GST pull-down analysis of the interaction of GST- (A, B), GST mutants (B) or GST- (C) with indicated proteins. A, B, Cell lysate from untransfected HeLa cells were loaded
Legends Supplementary Figure 1. GST pull-down analysis of the interaction of GST- (A, B), GST mutants (B) or GST- (C) with indicated proteins. A, B, Cell lysate from untransfected HeLa cells were loaded
-15 diopter negative lenses in wild-type and homozygous CHRM2-deleted mice, and
 Supplementary Materials Supplementary Figure 1: Myopia induction was performed using uniocular -10 and -15 diopter negative lenses in wild-type and homozygous CHRM2-deleted mice, and results at 2, 4 and
Supplementary Materials Supplementary Figure 1: Myopia induction was performed using uniocular -10 and -15 diopter negative lenses in wild-type and homozygous CHRM2-deleted mice, and results at 2, 4 and
SUPPLEMENTARY INFORMATION
 doi:1.138/nature11496 Cl. 8 Cl. E93 Rag1 -/- 3H9 + BM Rag1 -/- BM CD CD c-kit c-kit c-kit wt Spleen c-kit B22 B22 IgM IgM IgM Supplementary Figure 1. FACS analysis of single-cell-derived pre-b cell clones.
doi:1.138/nature11496 Cl. 8 Cl. E93 Rag1 -/- 3H9 + BM Rag1 -/- BM CD CD c-kit c-kit c-kit wt Spleen c-kit B22 B22 IgM IgM IgM Supplementary Figure 1. FACS analysis of single-cell-derived pre-b cell clones.
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system
 Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Chapter 13 Chromatin Structure and its Effects on Transcription
 Chapter 13 Chromatin Structure and its Effects on Transcription Students must be positive that they understand standard PCR. There is a resource on the web for this purpose. Warn them before this class.
Chapter 13 Chromatin Structure and its Effects on Transcription Students must be positive that they understand standard PCR. There is a resource on the web for this purpose. Warn them before this class.
Y-chromosomal haplogroup typing Using SBE reaction
 Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency.
 Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency. (a) Transfection of different concentration of GAS5-overexpressing
Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency. (a) Transfection of different concentration of GAS5-overexpressing
Legends for supplementary figures 1-3
 High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
Project 07/111 Final Report October 31, Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines
 Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Lecture 11: Gene Prediction
 Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature09732 Supplementary Figure 1: Depletion of Fbw7 results in elevated Mcl-1 abundance. a, Total thymocytes from 8-wk-old Lck-Cre/Fbw7 +/fl (Control) or Lck-Cre/Fbw7 fl/fl (Fbw7 KO) mice
doi:10.1038/nature09732 Supplementary Figure 1: Depletion of Fbw7 results in elevated Mcl-1 abundance. a, Total thymocytes from 8-wk-old Lck-Cre/Fbw7 +/fl (Control) or Lck-Cre/Fbw7 fl/fl (Fbw7 KO) mice
Cat. # Product Size DS130 DynaExpress TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1
 Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
NAME:... MODEL ANSWER... STUDENT NUMBER:... Maximum marks: 50. Internal Examiner: Hugh Murrell, Computer Science, UKZN
 COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal antibody was examined in
 Supplementary information Supplementary figures Supplementary Figure 1 Determination of the s pecificity of in-house anti-rhbdd1 mouse monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal
Supplementary information Supplementary figures Supplementary Figure 1 Determination of the s pecificity of in-house anti-rhbdd1 mouse monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal
Supplementary Figure 1 PARP1 is involved in regulating the stability of mrnas from pro-inflammatory cytokine/chemokine mediators.
 Supplementary Figure 1 PARP1 is involved in regulating the stability of mrnas from pro-inflammatory cytokine/chemokine mediators. (a) A graphic depiction of the approach to determining the stability of
Supplementary Figure 1 PARP1 is involved in regulating the stability of mrnas from pro-inflammatory cytokine/chemokine mediators. (a) A graphic depiction of the approach to determining the stability of
Nature Immunology: doi: /ni Supplementary Figure 1. Zranb1 gene targeting.
 Supplementary Figure 1 Zranb1 gene targeting. (a) Schematic picture of Zranb1 gene targeting using an FRT-LoxP vector, showing the first 6 exons of Zranb1 gene (exons 7-9 are not shown). Targeted mice
Supplementary Figure 1 Zranb1 gene targeting. (a) Schematic picture of Zranb1 gene targeting using an FRT-LoxP vector, showing the first 6 exons of Zranb1 gene (exons 7-9 are not shown). Targeted mice
WB: anti-flag (Orai1) WB: anti-stim1
 [Ca 2+ ] i (µm) 1.5 1..5 2Ca Ca 2 mm Ca 1 µm TG HeLa µm 2-APB HeLa O+S KDa DSP (mm).12..5 1. 2.5 3 6 9 12 2 Complex Monomer KDa 2 1 DSP (mm).12..5 1. 2.5 Complex Monomer (Orai1) WB: anti-stim1 Figure S1
[Ca 2+ ] i (µm) 1.5 1..5 2Ca Ca 2 mm Ca 1 µm TG HeLa µm 2-APB HeLa O+S KDa DSP (mm).12..5 1. 2.5 3 6 9 12 2 Complex Monomer KDa 2 1 DSP (mm).12..5 1. 2.5 Complex Monomer (Orai1) WB: anti-stim1 Figure S1
Partial list of differentially expressed genes from cdna microarray, comparing MUC18-
 Supplemental Figure legends Table-1 Partial list of differentially expressed genes from cdna microarray, comparing MUC18- silenced and NT-transduced A375SM cells. Supplemental Figure 1 Effect of MUC-18
Supplemental Figure legends Table-1 Partial list of differentially expressed genes from cdna microarray, comparing MUC18- silenced and NT-transduced A375SM cells. Supplemental Figure 1 Effect of MUC-18
Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURB
 Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURBO DNA-free Kit (Ambion). One µg of total RNA was reverse
Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURBO DNA-free Kit (Ambion). One µg of total RNA was reverse
An engineered tryptophan zipper-type peptide as a molecular recognition scaffold
 SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2743 Figure S1 stabilizes cellular protein level, post-transcriptionally. (a, b) and DDR1 were RNAi-depleted from HEK.293.-CBG cells. Western blots with indicated antibodies (a). RT-PCRs
DOI: 10.1038/ncb2743 Figure S1 stabilizes cellular protein level, post-transcriptionally. (a, b) and DDR1 were RNAi-depleted from HEK.293.-CBG cells. Western blots with indicated antibodies (a). RT-PCRs
Supplemental Information. Target-Mediated Protection of Endogenous. MicroRNAs in C. elegans. Inventory of Supplementary Information
 Developmental Cell, Volume 20 Supplemental Information Target-Mediated Protection of Endogenous MicroRNAs in C. elegans Saibal Chatterjee, Monika Fasler, Ingo Büssing, and Helge Großhans Inventory of Supplementary
Developmental Cell, Volume 20 Supplemental Information Target-Mediated Protection of Endogenous MicroRNAs in C. elegans Saibal Chatterjee, Monika Fasler, Ingo Büssing, and Helge Großhans Inventory of Supplementary
mir-155 regulates differentiation of brown and beige adipocytes via a bistable circuit
 SUPPLEMENTARY INFORMATION mir-155 regulates differentiation of brown and beige adipocytes via a bistable circuit Yong Chen 1,2,, Franziska Siegel 1,, Stefanie Kipschull 1, Bodo Haas 1,3, Holger Fröhlich
SUPPLEMENTARY INFORMATION mir-155 regulates differentiation of brown and beige adipocytes via a bistable circuit Yong Chen 1,2,, Franziska Siegel 1,, Stefanie Kipschull 1, Bodo Haas 1,3, Holger Fröhlich
Supporting Information
 Supporting Information AbuOdeh et al. 10.1073/pnas.1409252111 SI Materials and Methods List of Antibodies. Antibodies are listed: polyclonal anti antibody (1:500; gift from Kay Huebner, Ohio State University,
Supporting Information AbuOdeh et al. 10.1073/pnas.1409252111 SI Materials and Methods List of Antibodies. Antibodies are listed: polyclonal anti antibody (1:500; gift from Kay Huebner, Ohio State University,
supplementary information
 DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
A Genetically Encoded Toolbox for Glycocalyx Engineering: Tunable Control of Cell Adhesion,
 TITLE A Genetically Encoded Toolbox for Glycocalyx Engineering: Tunable Control of Cell Adhesion, Survival, and Cancer Cell Behaviors AUTHORS Carolyn R. Shurer *, Marshall J. Colville *, Vivek K. Gupta,
TITLE A Genetically Encoded Toolbox for Glycocalyx Engineering: Tunable Control of Cell Adhesion, Survival, and Cancer Cell Behaviors AUTHORS Carolyn R. Shurer *, Marshall J. Colville *, Vivek K. Gupta,
Estradiol-Estrogen Receptor α Mediates the Expression of the CXXC5 Gene through the Estrogen Response Element-Dependent Signaling Pathway
 Estradiol-Estrogen Receptor α Mediates the Expression of the CXXC5 Gene through the Estrogen Response Element-Dependent Signaling Pathway Pelin Yaşar, Gamze Ayaz and Mesut Muyan SUPPLEMENTARY INFORMATION
Estradiol-Estrogen Receptor α Mediates the Expression of the CXXC5 Gene through the Estrogen Response Element-Dependent Signaling Pathway Pelin Yaşar, Gamze Ayaz and Mesut Muyan SUPPLEMENTARY INFORMATION
SUPPLEMENTARY INFORMATION
 doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
Supplemental Figure 1
 mrna level (fold induction) Supplemental Figure 1 a-flag pmx-rank WT pmx-rank Mt 0 1 3 6 0 1 3 6 (h) c-fos b-actin 5 4 3 2 1 0 0 1 3 6 0 1 3 6 (h) pmx-rank WT pmx-rank Mt TRAP+ Mononuclear pre-ocs/well
mrna level (fold induction) Supplemental Figure 1 a-flag pmx-rank WT pmx-rank Mt 0 1 3 6 0 1 3 6 (h) c-fos b-actin 5 4 3 2 1 0 0 1 3 6 0 1 3 6 (h) pmx-rank WT pmx-rank Mt TRAP+ Mononuclear pre-ocs/well
b alternative classical none
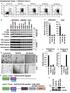 Supplementary Figure. 1: Related to Figure.1 a d e b alternative classical none NIK P-IkBa Total IkBa Tubulin P52 (Lighter) P52 (Darker) RelB (Lighter) RelB (Darker) HDAC1 Control-Sh RelB-Sh NF-kB2-Sh
Supplementary Figure. 1: Related to Figure.1 a d e b alternative classical none NIK P-IkBa Total IkBa Tubulin P52 (Lighter) P52 (Darker) RelB (Lighter) RelB (Darker) HDAC1 Control-Sh RelB-Sh NF-kB2-Sh
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Coding and noncoding transcription at the vg locus. (a,b) qpcr analysis of developmental and tissue-specific vg mrna (a) and PRE/TRE (b) transcription, shown as percentage of TBP
Supplementary Figure 1 Coding and noncoding transcription at the vg locus. (a,b) qpcr analysis of developmental and tissue-specific vg mrna (a) and PRE/TRE (b) transcription, shown as percentage of TBP
