Gene Prediction. Mario Stanke. Institut für Mikrobiologie und Genetik Abteilung Bioinformatik. Gene Prediction p.
|
|
|
- Maximillian Parsons
- 5 years ago
- Views:
Transcription
1 Gene Prediction Mario Stanke Institut für Mikrobiologie und Genetik Abteilung Bioinformatik Gene Prediction p.1/23
2 Why Predict Genes with a Computer? tons of data 39/250 eukaryotic/prokaryotic genomes sequenced 547/804 additional sequencing projects started no experimental evicence available: gene not expressed in sample selective experimental verification of predicions Gene Prediction p.2/23
3 What is Predicted? a eukaryotic gene DNA sequence transcription + splicing messenger RNA sequence translation amino acid sequence Legend: not transcibed or transcribed and spliced out transcribed, not spliced out but not translated translated Gene Prediction p.3/23
4 Example of a Human Gene cctcacctctgagaaaacctctttgccaccaataccatgaagctctgcgtgactgtcctgtctctc ctcgtgctagtagctgccttctgctctctagcactctcagcaccaagtaagtctacttttgcagct gctatttcgagtcaaggtgtaggcagagtccttttttctagtcatggctggcaaacagtgggatct ggggatgggacaaaaggcagctaggaagattgccatgtagtctgctgctaaatgtagagtctagta gatattcagtaacattcaagttcctattttcttaagaattagcaaccagcagaggaaaacgatggg ctggaagtcagactgttgaattggctctgcctttaattatttgttcaagcaagcccctgtccctct ctgtgccttggtttccccatctgtcatatgaagggagtgcgatgtgttctgagactgaatccagtt ccaatcttctagatttctttctcgttcttctctgaagatccactattcagaataagactcctgctc atgttaggtgggaatggatacaagggaccatatttggggttctggtagctccacagggatgctcaa tgaagatgcaaaattagaagtcaaaataaacagctcccatgggcagtgttgatctcaccctggcct ttcctttcagtgggctcagaccctcccaccgcctgctgcttttcttacaccgcgaggaagcttcct cgcaactttgtggtagattactatgagaccagcagcctctgctcccagccagctgtggtgtgagta tcaacccctggctgccctgggaggcaagggtgagggctggatttttaaagggggcctgttttgggg agggggtgatgagcgctggggaggcagctctcagggctgaagccttccctgacagcagtgaggtca caggtcatgaactcacttttcaagtgctgaaggcggctgagtggcagccgagacagaagggggttc ctggggaggaagttattcagaggacagggaagcaggggaaggcagacaggtcccatgagatatgga ccaattccttaaaccatgctagaaaaacatgtggaaaagtcactaccaggctggcagggaatgggg caatctattcatactgattgcaatgcccactggttcctaatctgggcaacccctggggcccacagc taaatccagtgagtggaagttacagggagtctgcttccagtgctgctcgaggaaggatcccatcca ccagagctgccccacatggaccatggtcaggcagaggaagatgcctaccacaggcaagggataaag ccagatgacctcaaaggtcccatgggattctaatctgtctgctccttgttctacagattccaaacc aaaagaggcaagcaagtctgcgctgaccccagtgagtcctgggtccaggagtacgtgtatgacctg gaactgaactgagctgctcagagacaggaagtcttc Gene Prediction p.4/23
5 Example of a Human Gene cctcacctctgagaaaacctctttgccaccaataccatgaagctctgcgtgactgtcctgtctctc ctcgtgctagtagctgccttctgctctctagcactctcagcaccaagtaagtctacttttgcagct gctatttcgagtcaaggtgtaggcagagtccttttttctagtcatggctggcaaacagtgggatct ggggatgggacaaaaggcagctaggaagattgccatgtagtctgctgctaaatgtagagtctagta gatattcagtaacattcaagttcctattttcttaagaattagcaaccagcagaggaaaacgatggg ctggaagtcagactgttgaattggctctgcctttaattatttgttcaagcaagcccctgtccctct ctgtgccttggtttccccatctgtcatatgaagggagtgcgatgtgttctgagactgaatccagtt ccaatcttctagatttctttctcgttcttctctgaagatccactattcagaataagactcctgctc atgttaggtgggaatggatacaagggaccatatttggggttctggtagctccacagggatgctcaa tgaagatgcaaaattagaagtcaaaataaacagctcccatgggcagtgttgatctcaccctggcct ttcctttcagtgggctcagaccctcccaccgcctgctgcttttcttacaccgcgaggaagcttcct cgcaactttgtggtagattactatgagaccagcagcctctgctcccagccagctgtggtgtgagta tcaacccctggctgccctgggaggcaagggtgagggctggatttttaaagggggcctgttttgggg agggggtgatgagcgctggggaggcagctctcagggctgaagccttccctgacagcagtgaggtca caggtcatgaactcacttttcaagtgctgaaggcggctgagtggcagccgagacagaagggggttc ctggggaggaagttattcagaggacagggaagcaggggaaggcagacaggtcccatgagatatgga ccaattccttaaaccatgctagaaaaacatgtggaaaagtcactaccaggctggcagggaatgggg caatctattcatactgattgcaatgcccactggttcctaatctgggcaacccctggggcccacagc taaatccagtgagtggaagttacagggagtctgcttccagtgctgctcgaggaaggatcccatcca ccagagctgccccacatggaccatggtcaggcagaggaagatgcctaccacaggcaagggataaag ccagatgacctcaaaggtcccatgggattctaatctgtctgctccttgttctacagattccaaacc aaaagaggcaagcaagtctgcgctgaccccagtgagtcctgggtccaggagtacgtgtatgacctg gaactgaactgagctgctcagagacaggaagtcttc Gene Prediction p.4/23
6 Methods of Gene Prediction signal sensors content sensors alignment with cdna alignment with ESTs protein homology cross-species DNA comparison Gene Prediction p.5/23
7 Methods of Gene Prediction signal sensors content sensors alignment with cdna alignment with ESTs protein homology cross-species DNA comparison } ab initio methods extrinsic methods Gene Prediction p.5/23
8 Signal Sensors promoter translation initation site (TIS) stop codon poly A DNA sequence donor splice site (dss) acceptor splice site (ass) Signals are short sequence segments of the DNA, that control translation or transcription. Gene Prediction p.6/23
9 Signal Sensors promoter: marks the begin of transcription splice sites: 5 (donor) and 3 (acceptor) end of an intron TIS: Contains the start codon (usually atg) and marks the begin of translation stop codon: marks the end of translation (usually tga, taa or tag) poly-a signal: triggers end of transcription. Gene Prediction p.7/23
10 Donor Splice Site These signals contain typical sequence motifs, but these motifs are not characteristic: The motifs occurr also at positions where actually no signal is. Example donor splice sites: (Almost) every intron begins with the dinucleotide gt, but that is not sufficiently specific and does not suffice for locating donor splice sites. Gene Prediction p.8/23
11 Donor Splice Site cctcacctctgagaaaacctctttgccaccaataccatgaagctctgcgtgactgtcctgtctctc ctcgtgctagtagctgccttctgctctctagcactctcagcaccaagtaagtctacttttgcagct gctatttcgagtcaaggtgtaggcagagtccttttttctagtcatggctggcaaacagtgggatct ggggatgggacaaaaggcagctaggaagattgccatgtagtctgctgctaaatgtagagtctagta gatattcagtaacattcaagttcctattttcttaagaattagcaaccagcagaggaaaacgatggg ctggaagtcagactgttgaattggctctgcctttaattatttgttcaagcaagcccctgtccctct ctgtgccttggtttccccatctgtcatatgaagggagtgcgatgtgttctgagactgaatccagtt ccaatcttctagatttctttctcgttcttctctgaagatccactattcagaataagactcctgctc atgttaggtgggaatggatacaagggaccatatttggggttctggtagctccacagggatgctcaa tgaagatgcaaaattagaagtcaaaataaacagctcccatgggcagtgttgatctcaccctggcct ttcctttcagtgggctcagaccctcccaccgcctgctgcttttcttacaccgcgaggaagcttcct cgcaactttgtggtagattactatgagaccagcagcctctgctcccagccagctgtggtgtgagta tcaacccctggctgccctgggaggcaagggtgagggctggatttttaaagggggcctgttttgggg agggggtgatgagcgctggggaggcagctctcagggctgaagccttccctgacagcagtgaggtca caggtcatgaactcacttttcaagtgctgaaggcggctgagtggcagccgagacagaagggggttc ctggggaggaagttattcagaggacagggaagcaggggaaggcagacaggtcccatgagatatgga ccaattccttaaaccatgctagaaaaacatgtggaaaagtcactaccaggctggcagggaatgggg caatctattcatactgattgcaatgcccactggttcctaatctgggcaacccctggggcccacagc taaatccagtgagtggaagttacagggagtctgcttccagtgctgctcgaggaaggatcccatcca ccagagctgccccacatggaccatggtcaggcagaggaagatgcctaccacaggcaagggataaag ccagatgacctcaaaggtcccatgggattctaatctgtctgctccttgttctacagattccaaacc aaaagaggcaagcaagtctgcgctgaccccagtgagtcctgggtccaggagtacgtgtatgacctg gaactgaactgagctgctcagagacaggaagtctt Gene Prediction p.9/23
12 Donor Splice Site Exon...agagcaaggtacgc......tctttcatgtgagt......actgtcaggtatgt......gacaaaaggtacgt......tccaaaaggcaggg......tttcctaggtaacg......agaagatggtagga......tcctttgggtgagt......gatcctgggtcagt......cacgctgggtacgc......atcacctcgtgagt......cttccagggtgaga......acctcacggtgaga......aacaggaggtacca......gtcggaccgtgagt......aaactgcggtgagt......ccaggaaggtaggg......gcactgtggtgagc......gaggacaggtgagc......cgagggcggtgagc......cctgtcaggtgagt......acgtggaggtgagg......atcagtatgtgagt......tacatcgggtcagt......gccaccaggtaggg......gatctgtggtgaga......tctggccggtttgt......cttccaaggtaggg......gcgccaaggttggc... Intron Gene Prediction p.10/23
13 Donor Splice Site C GTACTAGGT G A T AGC AGC TAC GAC GAC T TGT TC TCA GCTGATCAGA CGT A pictogram of the region -8 to 6 relative to the boundary between exon and intron. The size of the letters of the bases is proportional to the relative frequency of the base at this position. Gene Prediction p.11/23
14 Content Sensors Coding sequences and non-coding sequences (introns, intergenic region) also typically have different base compositions. For example coding: bases g and c slightly more common non-coding: bases a and t slightly more common Gene Prediction p.12/23
15 Content Sensors cctcacctctgagaaaacctctttgccaccaataccatgaagctctgcgtgactgtcctgtctctc ctcgtgctagtagctgccttctgctctctagcactctcagcaccaagtaagtctacttttgcagct gctatttcgagtcaaggtgtaggcagagtccttttttctagtcatggctggcaaacagtgggatct ggggatgggacaaaaggcagctaggaagattgccatgtagtctgctgctaaatgtagagtctagta gatattcagtaacattcaagttcctattttcttaagaattagcaaccagcagaggaaaacgatggg ctggaagtcagactgttgaattggctctgcctttaattatttgttcaagcaagcccctgtccctct ctgtgccttggtttccccatctgtcatatgaagggagtgcgatgtgttctgagactgaatccagtt ccaatcttctagatttctttctcgttcttctctgaagatccactattcagaataagactcctgctc atgttaggtgggaatggatacaagggaccatatttggggttctggtagctccacagggatgctcaa tgaagatgcaaaattagaagtcaaaataaacagctcccatgggcagtgttgatctcaccctggcct ttcctttcagtgggctcagaccctcccaccgcctgctgcttttcttacaccgcgaggaagcttcct cgcaactttgtggtagattactatgagaccagcagcctctgctcccagccagctgtggtgtgagta tcaacccctggctgccctgggaggcaagggtgagggctggatttttaaagggggcctgttttgggg agggggtgatgagcgctggggaggcagctctcagggctgaagccttccctgacagcagtgaggtca caggtcatgaactcacttttcaagtgctgaaggcggctgagtggcagccgagacagaagggggttc ctggggaggaagttattcagaggacagggaagcaggggaaggcagacaggtcccatgagatatgga ccaattccttaaaccatgctagaaaaacatgtggaaaagtcactaccaggctggcagggaatgggg caatctattcatactgattgcaatgcccactggttcctaatctgggcaacccctggggcccacagc taaatccagtgagtggaagttacagggagtctgcttccagtgctgctcgaggaaggatcccatcca ccagagctgccccacatggaccatggtcaggcagaggaagatgcctaccacaggcaagggataaag ccagatgacctcaaaggtcccatgggattctaatctgtctgctccttgttctacagattccaaacc aaaagaggcaagcaagtctgcgctgaccccagtgagtcctgggtccaggagtacgtgtatgacctg gaactgaactgagctgctcagagacaggaagtcttc Gene Prediction p.13/23
16 Content Sensors The frequency of dinucleotides serve even better as a means to distinguish coding from non-coding sequences. Example: The dinucleotide at occurs more often in non-coding sequences than in coding sequences. Gene Prediction p.14/23
17 Content Sensors cctcacctctgagaaaacctctttgccaccaataccatgaagctctgcgtgactgtcctgtctctc ctcgtgctagtagctgccttctgctctctagcactctcagcaccaagtaagtctacttttgcagct gctatttcgagtcaaggtgtaggcagagtccttttttctagtcatggctggcaaacagtgggatct ggggatgggacaaaaggcagctaggaagattgccatgtagtctgctgctaaatgtagagtctagta gatattcagtaacattcaagttcctattttcttaagaattagcaaccagcagaggaaaacgatggg ctggaagtcagactgttgaattggctctgcctttaattatttgttcaagcaagcccctgtccctct ctgtgccttggtttccccatctgtcatatgaagggagtgcgatgtgttctgagactgaatccagtt ccaatcttctagatttctttctcgttcttctctgaagatccactattcagaataagactcctgctc atgttaggtgggaatggatacaagggaccatatttggggttctggtagctccacagggatgctcaa tgaagatgcaaaattagaagtcaaaataaacagctcccatgggcagtgttgatctcaccctggcct ttcctttcagtgggctcagaccctcccaccgcctgctgcttttcttacaccgcgaggaagcttcct cgcaactttgtggtagattactatgagaccagcagcctctgctcccagccagctgtggtgtgagta tcaacccctggctgccctgggaggcaagggtgagggctggatttttaaagggggcctgttttgggg agggggtgatgagcgctggggaggcagctctcagggctgaagccttccctgacagcagtgaggtca caggtcatgaactcacttttcaagtgctgaaggcggctgagtggcagccgagacagaagggggttc ctggggaggaagttattcagaggacagggaagcaggggaaggcagacaggtcccatgagatatgga ccaattccttaaaccatgctagaaaaacatgtggaaaagtcactaccaggctggcagggaatgggg caatctattcatactgattgcaatgcccactggttcctaatctgggcaacccctggggcccacagc taaatccagtgagtggaagttacagggagtctgcttccagtgctgctcgaggaaggatcccatcca ccagagctgccccacatggaccatggtcaggcagaggaagatgcctaccacaggcaagggataaag ccagatgacctcaaaggtcccatgggattctaatctgtctgctccttgttctacagattccaaacc aaaagaggcaagcaagtctgcgctgaccccagtgagtcctgggtccaggagtacgtgtatgacctg gaactgaactgagctgctcagagacaggaagtcttc Gene Prediction p.15/23
18 Content Sensors Reading frame dependent hexamer frequencies is the most commonly used content sensor of current gene prediction programs. Gene Prediction p.16/23
19 Parameters Must Be Species Specific, Example Gene Prediction p.17/23
20 Protein Homology Use local similarity between translated input DNA sequence and amino acid sequence from database to infer evidence about coding regions. Example: input human DNA: gcc atg tcg tcc ggc atc cat gta gcg ctg gtg act gga ggc aac aag ggc atc ggc translated seq.: H V A L V T G G N K G I G A. thaliana homolog: N V A V V T G S N R G I G Programs: GenomeScan AUGUSTUS+ Gene Prediction p.18/23
21 Cross-Species DNA Comparison Consider the DNA sequences of two different species coding for the same (or a similar) protein. Functional parts of the sequence, especially coding regions, tend to be more conserved. Programs: DoubleScan SLAM AUGUSTUS+ AGenDA Gene Prediction p.19/23
22 Cross-Species DNA Comparison Gene Prediction p.20/23
23 The Reliability of Gene Prediction depends on the organism and on the available extrinsic information. Example: humans, ab initio, ENCODE sequences program base exon transcr. gene sn sp sn sp sn sp sn sp Augustus GenemarkHMM Genezilla geneid genscan Gene Prediction p.21/23
24 Problems of Gene Prediction alternative splicing TIS and promoter prediction less reliable exceptions, e.g. overlapping genes non-canonical splice sites (programmed) frameshifts different genetic code Gene Prediction p.22/23
25 Problems of Gene Prediction alternative splicing TIS and promoter prediction less reliable exceptions, e.g. overlapping genes non-canonical splice sites (programmed) frameshifts different genetic code Gene Prediction p.22/23
26 Problems of Gene Prediction alternative splicing TIS and promoter prediction less reliable exceptions, e.g. overlapping genes non-canonical splice sites (programmed) frameshifts different genetic code Gene Prediction p.22/23
27 Problems of Gene Prediction alternative splicing TIS and promoter prediction less reliable exceptions, e.g. overlapping genes non-canonical splice sites (programmed) frameshifts different genetic code Gene Prediction p.22/23
28 Screenshot AUGUSTUS web interface Gene Prediction p.23/23
Lecture 11: Gene Prediction
 Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR
 Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
DNA is normally found in pairs, held together by hydrogen bonds between the bases
 Bioinformatics Biology Review The genetic code is stored in DNA Deoxyribonucleic acid. DNA molecules are chains of four nucleotide bases Guanine, Thymine, Cytosine, Adenine DNA is normally found in pairs,
Bioinformatics Biology Review The genetic code is stored in DNA Deoxyribonucleic acid. DNA molecules are chains of four nucleotide bases Guanine, Thymine, Cytosine, Adenine DNA is normally found in pairs,
Disease and selection in the human genome 3
 Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
132 Grundlagen der Bioinformatik, SoSe 14, D. Huson, June 22, This exposition is based on the following source, which is recommended reading:
 132 Grundlagen der Bioinformatik, SoSe 14, D. Huson, June 22, 214 1 Gene Prediction Using HMMs This exposition is based on the following source, which is recommended reading: 1. Chris Burge and Samuel
132 Grundlagen der Bioinformatik, SoSe 14, D. Huson, June 22, 214 1 Gene Prediction Using HMMs This exposition is based on the following source, which is recommended reading: 1. Chris Burge and Samuel
Grundlagen der Bioinformatik, SoSe 11, D. Huson, July 4, This exposition is based on the following source, which is recommended reading:
 Grundlagen der Bioinformatik, SoSe 11, D. Huson, July 4, 211 155 12 Gene Prediction Using HMMs This exposition is based on the following source, which is recommended reading: 1. Chris Burge and Samuel
Grundlagen der Bioinformatik, SoSe 11, D. Huson, July 4, 211 155 12 Gene Prediction Using HMMs This exposition is based on the following source, which is recommended reading: 1. Chris Burge and Samuel
Genomics and Gene Recognition Genes and Blue Genes
 Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
Genomics and Gene Recognition Genes and Blue Genes November 1, 2004 Prokaryotic Gene Structure prokaryotes are simplest free-living organisms studying prokaryotes can give us a sense what is the minimum
BioInformatics and Computational Molecular Biology. Course Website
 BioInformatics and Computational Molecular Biology Course Website http://bioinformatics.uchc.edu What is Bioinformatics Bioinformatics upgrades the information content of biological measurements. Discovery
BioInformatics and Computational Molecular Biology Course Website http://bioinformatics.uchc.edu What is Bioinformatics Bioinformatics upgrades the information content of biological measurements. Discovery
Interpretation of sequence results
 Interpretation of sequence results An overview on DNA sequencing: DNA sequencing involves the determination of the sequence of nucleotides in a sample of DNA. It use a modified PCR reaction where both
Interpretation of sequence results An overview on DNA sequencing: DNA sequencing involves the determination of the sequence of nucleotides in a sample of DNA. It use a modified PCR reaction where both
Computational Gene Finding
 Computational Gene Finding Dong Xu Digital Biology Laboratory Computer Science Department Christopher S. Life Sciences Center University of Missouri, Columbia E-mail: xudong@missouri.edu http://digbio.missouri.edu
Computational Gene Finding Dong Xu Digital Biology Laboratory Computer Science Department Christopher S. Life Sciences Center University of Missouri, Columbia E-mail: xudong@missouri.edu http://digbio.missouri.edu
Gene Identification in silico
 Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Computational gene finding
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Eukaryotic Gene Prediction. Wei Zhu May 2007
 Eukaryotic Gene Prediction Wei Zhu May 2007 In nature, nothing is perfect... - Alice Walker Gene Structure What is Gene Prediction? Gene prediction is the problem of parsing a sequence into nonoverlapping
Eukaryotic Gene Prediction Wei Zhu May 2007 In nature, nothing is perfect... - Alice Walker Gene Structure What is Gene Prediction? Gene prediction is the problem of parsing a sequence into nonoverlapping
Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis
 1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 08: Gene finding aatgcatgcggctatgctaatgcatgcggctatgctaagctgggatccgatgacaatgcatgcggctatgctaatgcatgcggc tatgcaagctgggatccgatgactatgctaagctgggatccgatgacaatgcatgcggctatgctaatgaatggtcttgggatt
EECS730: Introduction to Bioinformatics Lecture 08: Gene finding aatgcatgcggctatgctaatgcatgcggctatgctaagctgggatccgatgacaatgcatgcggctatgctaatgcatgcggc tatgcaagctgggatccgatgactatgctaagctgggatccgatgacaatgcatgcggctatgctaatgaatggtcttgggatt
NAME:... MODEL ANSWER... STUDENT NUMBER:... Maximum marks: 50. Internal Examiner: Hugh Murrell, Computer Science, UKZN
 COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
Materials Protein synthesis kit. This kit consists of 24 amino acids, 24 transfer RNAs, four messenger RNAs and one ribosome (see below).
 Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
GenBank Growth. In 2003 ~ 31 million sequences ~ 37 billion base pairs
 Gene Finding GenBank Growth GenBank Growth In 2003 ~ 31 million sequences ~ 37 billion base pairs GenBank: Exponential Growth Growth of GenBank in billions of base pairs from release 3 in April of 1994
Gene Finding GenBank Growth GenBank Growth In 2003 ~ 31 million sequences ~ 37 billion base pairs GenBank: Exponential Growth Growth of GenBank in billions of base pairs from release 3 in April of 1994
PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells
 Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
ORFs and genes. Please sit in row K or forward
 ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
PCR analysis was performed to show the presence and the integrity of the var1csa and var-
 Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Outline. Introduction to ab initio and evidence-based gene finding. Prokaryotic gene predictions
 Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
Genome annotation & EST
 Genome annotation & EST What is genome annotation? The process of taking the raw DNA sequence produced by the genome sequence projects and adding the layers of analysis and interpretation necessary
Genome annotation & EST What is genome annotation? The process of taking the raw DNA sequence produced by the genome sequence projects and adding the layers of analysis and interpretation necessary
Lecture 19A. DNA computing
 Lecture 19A. DNA computing What exactly is DNA (deoxyribonucleic acid)? DNA is the material that contains codes for the many physical characteristics of every living creature. Your cells use different
Lecture 19A. DNA computing What exactly is DNA (deoxyribonucleic acid)? DNA is the material that contains codes for the many physical characteristics of every living creature. Your cells use different
DNA sentences. How are proteins coded for by DNA? Materials. Teacher instructions. Student instructions. Reflection
 DNA sentences How are proteins coded for by DNA? Deoxyribonucleic acid (DNA) is the molecule of life. DNA is one of the most recognizable nucleic acids, a double-stranded helix. The process by which DNA
DNA sentences How are proteins coded for by DNA? Deoxyribonucleic acid (DNA) is the molecule of life. DNA is one of the most recognizable nucleic acids, a double-stranded helix. The process by which DNA
Gene Prediction in Eukaryotes
 Gene Prediction in Eukaryotes Jan-Jaap Wesselink Biomol Informatics, S.L. jjw@biomol-informatics.com June 2010/Madrid jjw@biomol-informatics.com (BI) Gene Prediction June 2010/Madrid 1 / 34 Outline 1 Gene
Gene Prediction in Eukaryotes Jan-Jaap Wesselink Biomol Informatics, S.L. jjw@biomol-informatics.com June 2010/Madrid jjw@biomol-informatics.com (BI) Gene Prediction June 2010/Madrid 1 / 34 Outline 1 Gene
Arabidopsis actin depolymerizing factor AtADF4 mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB
 Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
MODULE 5: TRANSLATION
 MODULE 5: TRANSLATION Lesson Plan: CARINA ENDRES HOWELL, LEOCADIA PALIULIS Title Translation Objectives Determine the codons for specific amino acids and identify reading frames by looking at the Base
MODULE 5: TRANSLATION Lesson Plan: CARINA ENDRES HOWELL, LEOCADIA PALIULIS Title Translation Objectives Determine the codons for specific amino acids and identify reading frames by looking at the Base
The Making of the Fittest: Natural Selection and Adaptation
 INTRODUCTION MOLECULAR GENETICS OF COLOR MUTATIONS IN ROCK POCKET MICE THE ROCK POCKET MOUSE The rock pocket mouse, Chaetodipus intermedius, is a small, nocturnal animal found in the deserts of the southwestern
INTRODUCTION MOLECULAR GENETICS OF COLOR MUTATIONS IN ROCK POCKET MICE THE ROCK POCKET MOUSE The rock pocket mouse, Chaetodipus intermedius, is a small, nocturnal animal found in the deserts of the southwestern
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Supplemental Data Supplemental Figure 1.
 Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Multiplexing Genome-scale Engineering
 Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
UNIT 7. DNA Structure, Replication, and Protein Synthesis
 UNIT 7 DNA Structure, Replication, and Protein Synthesis Section 3 Objectives Describe the difference between DNA and RNA. Define transcription. Define translation. Apply to rules of base pairing to replicate,
UNIT 7 DNA Structure, Replication, and Protein Synthesis Section 3 Objectives Describe the difference between DNA and RNA. Define transcription. Define translation. Apply to rules of base pairing to replicate,
SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer
 TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
Supplementary. Table 1: Oligonucleotides and Plasmids. complementary to positions from 77 of the SRα '- GCT CTA GAG AAC TTG AAG TAC AGA CTG C
 Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Genes and gene finding
 Genes and gene finding Ben Langmead Department of Computer Science You are free to use these slides. If you do, please sign the guestbook (www.langmead-lab.org/teaching-materials), or email me (ben.langmead@gmail.com)
Genes and gene finding Ben Langmead Department of Computer Science You are free to use these slides. If you do, please sign the guestbook (www.langmead-lab.org/teaching-materials), or email me (ben.langmead@gmail.com)
Computational gene finding. Devika Subramanian Comp 470
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) The biological context Lec 1 Lec 2 Lec 3 Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) The biological context Lec 1 Lec 2 Lec 3 Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Genome annotation. Erwin Datema (2011) Sandra Smit (2012, 2013)
 Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
Hes6. PPARα. PPARγ HNF4 CD36
 SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
Computational gene finding
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
TIGR THE INSTITUTE FOR GENOMIC RESEARCH
 Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
Evidence Combination in Hidden Markov Models for Gene Prediction
 Evidence Combination in Hidden Markov Models for Gene Prediction by Bronislava Brejová A thesis presented to the University of Waterloo in fulfilment of the thesis requirement for the degree of Doctor
Evidence Combination in Hidden Markov Models for Gene Prediction by Bronislava Brejová A thesis presented to the University of Waterloo in fulfilment of the thesis requirement for the degree of Doctor
GENETICS and the DNA code NOTES
 GENETICS and the DNA code NOTES BACKGROUND DNA is the hereditary material of most organisms. It is an organic compound made of two strands, twisted around one another to form a double helix. Each strand
GENETICS and the DNA code NOTES BACKGROUND DNA is the hereditary material of most organisms. It is an organic compound made of two strands, twisted around one another to form a double helix. Each strand
Course Introduction. Bioinformatics: Issues and Algorithms. CSE Fall 2007 Lecture 1
 Course Introduction Bioinformatics: Issues and Algorithms CSE 308-408 Fall 2007 Lecture 1-1- Motivation "Biology easily has 500 years of exciting problems to work on." Donald E. Knuth Good news: no prior
Course Introduction Bioinformatics: Issues and Algorithms CSE 308-408 Fall 2007 Lecture 1-1- Motivation "Biology easily has 500 years of exciting problems to work on." Donald E. Knuth Good news: no prior
Lecture 2: Biology Basics Continued. Fall 2018 August 23, 2018
 Lecture 2: Biology Basics Continued Fall 2018 August 23, 2018 Genetic Material for Life Central Dogma DNA: The Code of Life The structure and the four genomic letters code for all living organisms Adenine,
Lecture 2: Biology Basics Continued Fall 2018 August 23, 2018 Genetic Material for Life Central Dogma DNA: The Code of Life The structure and the four genomic letters code for all living organisms Adenine,
BIOSTAT516 Statistical Methods in Genetic Epidemiology Autumn 2005 Handout1, prepared by Kathleen Kerr and Stephanie Monks
 Rationale of Genetic Studies Some goals of genetic studies include: to identify the genetic causes of phenotypic variation develop genetic tests o benefits to individuals and to society are still uncertain
Rationale of Genetic Studies Some goals of genetic studies include: to identify the genetic causes of phenotypic variation develop genetic tests o benefits to individuals and to society are still uncertain
Supplemental Data. Bennett et al. (2010). Plant Cell /tpc
 BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
 www.lessonplansinc.com Topic: Gene Mutations WS Summary: Students will learn about frame shift mutations and base substitution mutations. Goals & Objectives: Students will be able to demonstrate how mutations
www.lessonplansinc.com Topic: Gene Mutations WS Summary: Students will learn about frame shift mutations and base substitution mutations. Goals & Objectives: Students will be able to demonstrate how mutations
Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC
 Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supporting Information. Copyright Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, 2006
 Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
ΔPDD1 x ΔPDD1. ΔPDD1 x wild type. 70 kd Pdd1. Pdd3
 Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
CSE : Computational Issues in Molecular Biology. Lecture 1. Spring 2004
 CSE 397-497: Computational Issues in Molecular Biology Lecture 1 Spring 2004-1- Motivation http://www.ncbi.nlm.nih.gov/genbank/genbankstats.html "Biology easily has 500 years of exciting problems to work
CSE 397-497: Computational Issues in Molecular Biology Lecture 1 Spring 2004-1- Motivation http://www.ncbi.nlm.nih.gov/genbank/genbankstats.html "Biology easily has 500 years of exciting problems to work
The Nature of Genes. The Nature of Genes. Genes and How They Work. Chapter 15/16
 Genes and How They Work Chapter 15/16 The Nature of Genes Beadle and Tatum proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes
Genes and How They Work Chapter 15/16 The Nature of Genes Beadle and Tatum proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes
RPA-AB RPA-C Supplemental Figure S1: SDS-PAGE stained with Coomassie Blue after protein purification.
 RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
Supporting information for Biochemistry, 1995, 34(34), , DOI: /bi00034a013
 Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supplemental material
 Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Eukaryotic Gene Structure
 Eukaryotic Gene Structure Terminology Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Gene Basic physical and
Eukaryotic Gene Structure Terminology Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Gene Basic physical and
Expression of Recombinant Proteins
 Expression of Recombinant Proteins Uses of Cloned Genes sequencing reagents (eg, probes) protein production insufficient natural quantities modify/mutagenesis library screening Expression Vector Features
Expression of Recombinant Proteins Uses of Cloned Genes sequencing reagents (eg, probes) protein production insufficient natural quantities modify/mutagenesis library screening Expression Vector Features
Collect, analyze and synthesize. Annotation. Annotation for D. virilis. Evidence Based Annotation. GEP goals: Evidence for Gene Models 08/22/2017
 Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
DNA Replication and Repair
 DNA Replication and Repair http://hyperphysics.phy-astr.gsu.edu/hbase/organic/imgorg/cendog.gif Overview of DNA Replication SWYK CNs 1, 2, 30 Explain how specific base pairing enables existing DNA strands
DNA Replication and Repair http://hyperphysics.phy-astr.gsu.edu/hbase/organic/imgorg/cendog.gif Overview of DNA Replication SWYK CNs 1, 2, 30 Explain how specific base pairing enables existing DNA strands
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
Additional Table A1. Accession numbers of resource records for all rhodopsin sequences downloaded from NCBI. Species common name
 1 2 3 Additional Table A1. Accession numbers of resource records for all rhodopsin sequences downloaded from NCBI. Species common name Scientific name Accession number Accession number (introns) Codons
1 2 3 Additional Table A1. Accession numbers of resource records for all rhodopsin sequences downloaded from NCBI. Species common name Scientific name Accession number Accession number (introns) Codons
Collect, analyze and synthesize. Annotation. Annotation for D. virilis. GEP goals: Evidence Based Annotation. Evidence for Gene Models 12/26/2018
 Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
Y-chromosomal haplogroup typing Using SBE reaction
 Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Gene Prediction: Statistical Approaches
 Gene Prediction: Statistical Approaches Outline 1. Central Dogma and Codons 2. Discovery of Split Genes 3. Splicing 4. Open Reading Frames 5. Codon Usage 6. Splicing Signals 7. TestCode Section 1: Central
Gene Prediction: Statistical Approaches Outline 1. Central Dogma and Codons 2. Discovery of Split Genes 3. Splicing 4. Open Reading Frames 5. Codon Usage 6. Splicing Signals 7. TestCode Section 1: Central
Codon Bias with PRISM. 2IM24/25, Fall 2007
 Codon Bias with PRISM 2IM24/25, Fall 2007 from RNA to protein mrna vs. trna aminoacid trna anticodon mrna codon codon-anticodon matching Watson-Crick base pairing A U and C G binding first two nucleotide
Codon Bias with PRISM 2IM24/25, Fall 2007 from RNA to protein mrna vs. trna aminoacid trna anticodon mrna codon codon-anticodon matching Watson-Crick base pairing A U and C G binding first two nucleotide
SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING
 SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING All of the patients and control subjects were sequenced and genotyped in the same way. Shotgun libraries of approximately 250 bp
SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING All of the patients and control subjects were sequenced and genotyped in the same way. Shotgun libraries of approximately 250 bp
Supporting Information
 Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
FAST AND ACCURATE GENE PREDICTION BY PROTEIN HOMOLOGY
 FAST AND ACCURATE GENE PREDICTION BY PROTEIN HOMOLOGY by Rong She Master of Science, Simon Fraser University, 2003 Bachelor of Engineering, Shanghai Jiaotong University, 1993 THESIS SUBMITTED IN PARTIAL
FAST AND ACCURATE GENE PREDICTION BY PROTEIN HOMOLOGY by Rong She Master of Science, Simon Fraser University, 2003 Bachelor of Engineering, Shanghai Jiaotong University, 1993 THESIS SUBMITTED IN PARTIAL
Annotation of contig27 in the Muller F Element of D. elegans. Contig27 is a 60,000 bp region located in the Muller F element of the D. elegans.
 David Wang Bio 434W 4/27/15 Annotation of contig27 in the Muller F Element of D. elegans Abstract Contig27 is a 60,000 bp region located in the Muller F element of the D. elegans. Genscan predicted six
David Wang Bio 434W 4/27/15 Annotation of contig27 in the Muller F Element of D. elegans Abstract Contig27 is a 60,000 bp region located in the Muller F element of the D. elegans. Genscan predicted six
Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers
 DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
Table S1. Sequences of mutagenesis primers used to create altered rdpa- and sdpa genes
 Supplementary Table and Figures for Structural Basis for the Enantiospecificities of R- and S-Specific Phenoxypropionate/α-Ketoglutarate Dioxygenases by Tina A. Müller, Maria I. Zavodszky, Michael Feig,
Supplementary Table and Figures for Structural Basis for the Enantiospecificities of R- and S-Specific Phenoxypropionate/α-Ketoglutarate Dioxygenases by Tina A. Müller, Maria I. Zavodszky, Michael Feig,
Gene Prediction: Statistical Approaches
 Gene Prediction: Statistical Approaches Outline Codons Discovery of Split Genes Exons and Introns Splicing Open Reading Frames Codon Usage Splicing Signals TestCode Gene Prediction: Computational Challenge
Gene Prediction: Statistical Approaches Outline Codons Discovery of Split Genes Exons and Introns Splicing Open Reading Frames Codon Usage Splicing Signals TestCode Gene Prediction: Computational Challenge
Outline. Gene Finding Questions. Recap: Prokaryotic gene finding Eukaryotic gene finding The human gene complement Regulation
 Tues, Nov 29: Gene Finding 1 Online FCE s: Thru Dec 12 Thurs, Dec 1: Gene Finding 2 Tues, Dec 6: PS5 due Project presentations 1 (see course web site for schedule) Thurs, Dec 8 Final papers due Project
Tues, Nov 29: Gene Finding 1 Online FCE s: Thru Dec 12 Thurs, Dec 1: Gene Finding 2 Tues, Dec 6: PS5 due Project presentations 1 (see course web site for schedule) Thurs, Dec 8 Final papers due Project
Improved Splice Site Detection in Genie
 Improved Splice Site Detection in Genie Martin Reese Informatics Group Human Genome Center Lawrence Berkeley National Laboratory MGReese@lbl.gov http://www-hgc.lbl.gov/inf Santa Fe, 1/23/97 Database Homologies
Improved Splice Site Detection in Genie Martin Reese Informatics Group Human Genome Center Lawrence Berkeley National Laboratory MGReese@lbl.gov http://www-hgc.lbl.gov/inf Santa Fe, 1/23/97 Database Homologies
Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH).
 Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Protein Structure Analysis
 BINF 731 Protein Structure Analysis http://binf.gmu.edu/vaisman/binf731/ Iosif Vaisman COMPUTATIONAL BIOLOGY COMPUTATIONAL STRUCTURAL BIOLOGY COMPUTATIONAL MOLECULAR BIOLOGY BIOINFORMATICS STRUCTURAL BIOINFORMATICS
BINF 731 Protein Structure Analysis http://binf.gmu.edu/vaisman/binf731/ Iosif Vaisman COMPUTATIONAL BIOLOGY COMPUTATIONAL STRUCTURAL BIOLOGY COMPUTATIONAL MOLECULAR BIOLOGY BIOINFORMATICS STRUCTURAL BIOINFORMATICS
Supplementary Figure 1A A404 Cells +/- Retinoic Acid
 Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
SUPPLEMENTARY INFORMATION
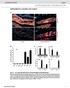 doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
Gene Prediction: Statistical Approaches
 Gene Prediction: Statistical Approaches Outline Codons Discovery of Split Genes Exons and Introns Splicing Open Reading Frames Codon Usage Splicing Signals TestCode Gene Prediction: Computational Challenge
Gene Prediction: Statistical Approaches Outline Codons Discovery of Split Genes Exons and Introns Splicing Open Reading Frames Codon Usage Splicing Signals TestCode Gene Prediction: Computational Challenge
2
 1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
ProGen: GPHMM for prokaryotic genomes
 ProGen: GPHMM for prokaryotic genomes Sharad Akshar Punuganti May 10, 2011 Abstract ProGen is an implementation of a Generalized Pair Hidden Markov Model (GPHMM), a model which can be used to perform both
ProGen: GPHMM for prokaryotic genomes Sharad Akshar Punuganti May 10, 2011 Abstract ProGen is an implementation of a Generalized Pair Hidden Markov Model (GPHMM), a model which can be used to perform both
Gene Finding Genome Annotation
 Gene Finding Genome Annotation Gene finding is a cornerstone of genomic analysis Genome content and organization Differential expression analysis Epigenomics Population biology & evolution Medical genomics
Gene Finding Genome Annotation Gene finding is a cornerstone of genomic analysis Genome content and organization Differential expression analysis Epigenomics Population biology & evolution Medical genomics
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Coding and noncoding transcription at the vg locus. (a,b) qpcr analysis of developmental and tissue-specific vg mrna (a) and PRE/TRE (b) transcription, shown as percentage of TBP
Supplementary Figure 1 Coding and noncoding transcription at the vg locus. (a,b) qpcr analysis of developmental and tissue-specific vg mrna (a) and PRE/TRE (b) transcription, shown as percentage of TBP
Supplemental Information. Human Senataxin Resolves RNA/DNA Hybrids. Formed at Transcriptional Pause Sites. to Promote Xrn2-Dependent Termination
 Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Gene synthesis by circular assembly amplification
 Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
PRINCIPLES OF BIOINFORMATICS
 PRINCIPLES OF BIOINFORMATICS BIO540/STA569/CSI660, Fall 2010 Lecture 3 (Sep-13-2010) Primer on Molecular Biology/Genomics Igor Kuznetsov Department of Epidemiology & Biostatistics Cancer Research Center
PRINCIPLES OF BIOINFORMATICS BIO540/STA569/CSI660, Fall 2010 Lecture 3 (Sep-13-2010) Primer on Molecular Biology/Genomics Igor Kuznetsov Department of Epidemiology & Biostatistics Cancer Research Center
Bacterial Genome Annotation
 Bacterial Genome Annotation Bacterial Genome Annotation For an annotation you want to predict from the sequence, all of... protein-coding genes their stop-start the resulting protein the function the control
Bacterial Genome Annotation Bacterial Genome Annotation For an annotation you want to predict from the sequence, all of... protein-coding genes their stop-start the resulting protein the function the control
Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were
 1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
G+C content. 1 Introduction. 2 Chromosomes Topology & Counts. 3 Genome size. 4 Replichores and gene orientation. 5 Chirochores.
 1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
Unit 1: DNA and the Genome. Sub-Topic (1.3) Gene Expression
 Unit 1: DNA and the Genome Sub-Topic (1.3) Gene Expression Unit 1: DNA and the Genome Sub-Topic (1.3) Gene Expression On completion of this subtopic I will be able to State the meanings of the terms genotype,
Unit 1: DNA and the Genome Sub-Topic (1.3) Gene Expression Unit 1: DNA and the Genome Sub-Topic (1.3) Gene Expression On completion of this subtopic I will be able to State the meanings of the terms genotype,
#26 - Gene Prediction 10/22/07
 BCB 444/544 Required Reading (before lecture) Lecture 26 Mon Oct 22 - Lecture 26 Gene Prediction Chp 8 - pp 97-112 Gene Prediction Wed Oct 24 - Lecture 27 (will not be covered on Exam 2) Regulatory Element
BCB 444/544 Required Reading (before lecture) Lecture 26 Mon Oct 22 - Lecture 26 Gene Prediction Chp 8 - pp 97-112 Gene Prediction Wed Oct 24 - Lecture 27 (will not be covered on Exam 2) Regulatory Element
May 16. Gene Finding
 Gene Finding j T[j,k] k i Q is a set of states T is a matrix of transition probabilities T[j,k]: probability of moving from state j to state k Σ is a set of symbols e j (S) is the probability of emitting
Gene Finding j T[j,k] k i Q is a set of states T is a matrix of transition probabilities T[j,k]: probability of moving from state j to state k Σ is a set of symbols e j (S) is the probability of emitting
strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular
![strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular](/thumbs/72/66361212.jpg) Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
PROTEIN SYNTHESIS Study Guide
 PART A. Read the following: PROTEIN SYNTHESIS Study Guide Protein synthesis is the process used by the body to make proteins. The first step of protein synthesis is called Transcription. It occurs in the
PART A. Read the following: PROTEIN SYNTHESIS Study Guide Protein synthesis is the process used by the body to make proteins. The first step of protein synthesis is called Transcription. It occurs in the
Genome 373: Gene Predic/on I. Doug Fowler
 Genome 373: Gene Predic/on I Doug Fowler Outline Review of gene structure Scale of the problem Solu;ons Empirical methods Ab ini&o predic;on What is a gene? A locatable region of genomic sequence, corresponding
Genome 373: Gene Predic/on I Doug Fowler Outline Review of gene structure Scale of the problem Solu;ons Empirical methods Ab ini&o predic;on What is a gene? A locatable region of genomic sequence, corresponding
Genes and How They Work. Chapter 15
 Genes and How They Work Chapter 15 The Nature of Genes They proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes The central
Genes and How They Work Chapter 15 The Nature of Genes They proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes The central
Replication, Transcription, and Translation
 Replication, Transcription, and Translation Information Flow from DNA to Protein The Central Dogma of Molecular Biology Replication is the copying of DNA in the course of cell division. Transcription is
Replication, Transcription, and Translation Information Flow from DNA to Protein The Central Dogma of Molecular Biology Replication is the copying of DNA in the course of cell division. Transcription is
Four different segments of a DNA molecule are represented below.
 Four different segments of a DNA molecule are represented below. There is an error in the DNA in which molecule? A. segment 1 only B. segment 3 only C. segment 2 and 3 D. segment 2 and 4 Explain the basic
Four different segments of a DNA molecule are represented below. There is an error in the DNA in which molecule? A. segment 1 only B. segment 3 only C. segment 2 and 3 D. segment 2 and 4 Explain the basic
