Nature Structural and Molecular Biology: doi: /nsmb.2937
|
|
|
- Joanna Fox
- 6 years ago
- Views:
Transcription
1
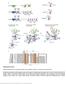
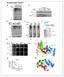
2 Supplementary Figure 1 Multiple sequence alignment of the CtIP N-terminal domain, purified CtIP protein constructs and details of the 2F o F c electron density map of CtIP-NTD. (a) Multiple sequence alignment, including the predicted secondary structure (red rods indicate regions of predicted α-helix) and coiledcoil regions; the probability of coiled-coil formation is indicated by - (<50%), c (50-90%) or C (>90%). Protein sequence analysis was performed with the Jalview suite of bioinformatics tools ( Hs: Homo sapiens, Mm: Mus musculus, Ec: Equus caballus, Oc: Oryctolagus cuniculus, Gg: Gallus gallus, Xl: Xenopus laevis, Dr: Danio rerio. (b) SDS-PAGE analysis of purified recombinant CtIP protein samples: NTD, NTD-DM (C89A, C92A), nntd, cntd, cntd-dm (C89A, C92A) and CTD visualised by Coomassie staining. (c) Details of the 2F o - F c electron density map, contoured at 1.2 rmsd, superimposed on the refined crystallographic model of CtIP-NTD. The figure shows the map in the vicinity of amino acid W24. The map is shown as thin mesh in light purple, the protein atoms are shown in ball-and-stick representation and coloured according to chemical identity. The figure was prepared in Coot
3
4 Supplementary Figure 2 CD analysis of CtIP N-terminal-domain constructs and micropixe analysis of the CtIP N-terminal domain. Circular dichroism (CD) analysis of CtIP N-terminal domain constructs. (a, c, e) Far-UV CD spectra recorded between 260 and 185 nm, in mean residue ellipticity, MRE ([θ]) (( xx ddeegg..ccmm 2.dmol -1.residue -1 ), with α-helical content calculated through deconvolution using the CDSSTR algorithm with normalised root-mean-square deviation values (nrmsd) as shown. (b, d, f) CD thermal denaturation data recorded between 5 and 95 C, in mean residue ellipticity at 222 nm ([θ] 222 ). (a) CD spectra and (b) thermal denaturation of CtIP-NTD (solid line) and CtIP-NTD-DM (dashed line). (c) CD spectra and (d) thermal denaturation of CtIP-cNTD (solid line) and CtIP-cNTD-DM (dashed line). (e) CD spectra and (f) thermal denaturation of CtIP-nNTD. (g) CtIP-NTD was analysed in buffer containing KBr. Rutherford backscattering (RBS) spectrum and (h) PIXE spectrum with raw data shown as black dots and the modelling fitting as red lines. (i) Zinc was detected at a mean atomic ratio to sulphur of 0.095, which given the presence of three methionine and three cysteine residues per chain, corresponds to a CtIP-NTD:Zn 2+ ratio of 1.8:1.
5 Supplementary Figure 3 SEC-MALS analysis of MBP-CtIP-nNTD and F20E CtIP-NTD. (a) MBP-CtIP-nNTD eluted in a single peak corresponding to a molecular weight of 187 kda; its theoretical tetramer size is 197 kda. For comparison, free MBP eluted in a single peak of 44.5 kda, corresponding to its theoretical monomer size of 44.7 kda. The light scattering (LS) as relative Raleigh ratio and the differential Reflective Index (dri) are drawn as solid and dashed lines, respectively. The value for the fitted molecular mass (M r ) of MBP-CtIP-nNTD is shown as diamond shapes across its elution peak. The predicted molecular masses of monomeric MBP-CtIP-nNTD and free MBP are shown in brackets above the respective elution peak. (b) F20E CtIP-NTD forms two distinct species of 62.3 kda and 33.2 kda, corresponding to its theoretical tetramer and dimer sizes of 63.2 kda and 31.6 kda, respectively. The light scattering (LS) as relative Raleigh ratio and the differential Reflective Index (dri) are drawn as solid and dashed lines, respectively. The values for the fitted molecular masses (M r ) of dimeric and tetrameric F20E CtIP-NTD are shown as diamond shapes across the respective elution peaks. The predicted molecular mass of monomeric F20E CtIP-nNTD is shown in brackets above the elution peaks.
6
7 Supplementary Figure 4 Experimental procedure and gating scheme for the TLR system. (a) TLR cells are not fluorescent due to the fact that the TLR cassette contains an in-frame truncated version of the egfp gene and fulllength mcherry is out of frame. When TLR cells are transfected with an I-SceI nuclease encoded in a plasmid also expressing an infrared fluorescent protein (IFP) and an exogenous donor template containing the missing part of egfp that also expresses a blue fluorescent protein (BFP), the different repair outcomes will give different fluorescent cells. If the double-strand break (DSB) is repaired by gene conversion (homologous recombination), egfp is restored and the cells fluoresce in green. If the DSB is repaired by mutagenic end joining causing a +2 frameshift, mcherry gets in frame and its expression yields red fluorescence. The T2A dis-linker between egfp and mcherry allows the downstream-encoded mcherry to escape degradation of the misfolded protein encoded in the +3 reading frame of egfp. (b) Experimental outline of the TLR assay. (c-h) Gating scheme for a TLR assay. (c) Discrimination of cells over debris using a forward scatter vs. side scatter plot (areas). (d) Discrimination of cell singlets over doublets using a forward scatter (area) vs. side scatter (width) plot. (e) Selection of events for quantification. Dot plot showing the intensity of BFP (donor; x axis) vs. intensity of IFP (nuclease; y axis). Cells with negative staining for both BFP and IFP are depicted in (f) to show the absence of egfp or mcherry fluorescence in non-transfected cells, and to establish quantification gates. Cells counted positive for both BFP and IFP (that is, transfected with both the donor and the nuclease) are represented in (g), where quantification gates are the same as the ones established in (f). At least 10,000 events were analysed in the double-positive population. (h) Due to the fact that the amount of donor transfected in the cells affects the outcome of the assay, the intensity of the BFP signal in the double-positive cells (geometric mean of BFP signal in BFP+ IFP+ cells) was calculated for normalisation purposes (see Methods). (i) Mutagenic end-joining (mutej) rates in the TLR-CtIP complementation system. Only expression of the wild-type version of FLAG-CtIP reduced the rate of mutej to values similar to the control (cells transfected with control sirna). EV: empty vector. All quantifications are shown as the average of three independent experiments. Error bars are ± SEM.
8
9 Supplementary Figure 5 CtIP expression levels in the different stable cell lines generated and fluorescence-activated cell sorting quantification of GFP-CtIP accumulation on damaged cells. (a) CtIP expression levels in the different U2OS-MMEJ FLAG-CtIP clones. Samples were collected 48 h after sirna transfection. CtIP antibody was a gift from R. Baer. PARP-1 antibody (Cell Signalling) was used as loading control. (b) GFP-CtIP expression levels in the U2OS stable cell lines. RPA34-20 (Merck) antibody was used as loading control. (c) A representative complete dataset for quantification of GFP-CtIP accumulation on damaged cells. Dot plots representing, for each dataset, gating and quantification (in the y axis) of the GFP-positive cells (top panels) and the γh2ax-positive cells (bottom panels) taking into account their DNA content (x axis). Quantification gates were always established using the untreated samples of each dataset. At least 10,000 events were analysed per sample.
10 Supplementary Figure 6 A representative complete dataset for the DNA-end resection assay. Dot plots representing, for each dataset, gating and quantification (in the y axis) of the RPA-positive cells (top panels) and the γh2axpositive cells (bottom panels) taking into account their DNA content (x axis). Quantification gates were always established using the untreated samples of each dataset and were defined as to avoid quantification of RPA chromatinization due to DNA replication. Note that as camptothecin (CPT) only causes DNA double-strand breaks in S-phase cells, both RPA- and γh2ax-positive cells only appear in that cell cycle stage. At least 10,000 events were analyzed per sample.
11 Supplementary Figure 7 CtIP-CTD, but not CtIP-NTD, interacts with double-stranded DNA. Electrophoretic mobility shift assay measuring the ability of CtIP NTD and CTD regions to interact with linear, double-stranded DNA. In the experiment, increasing amounts (0, 0.5, 1.0, 2.0, 5.0 and 10µM) of the CtIP protein were incubated with a 1µM sample of 5 fluorescein-labeled 200bp double-stranded DNA. After incubation, each sample was resolved by electrophoresis on agarose gel and visualised under UV light.
Supplementatry Fig 1. Domain structure, biophysical characterisation and electron microscopy of a TD. (a) XTACC3/Maskin and XMAP215/chTOG domain
 Supplementatry Fig 1. Domain structure, biophysical characterisation and electron microscopy of a TD. (a) XTACC3/Maskin and XMAP215/chTOG domain architecture. Various C-terminal fragments were cloned and
Supplementatry Fig 1. Domain structure, biophysical characterisation and electron microscopy of a TD. (a) XTACC3/Maskin and XMAP215/chTOG domain architecture. Various C-terminal fragments were cloned and
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than. SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with
 Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the
 Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Supplementary Figure 1 PZA inhibits root hair formation as well as cell elongation in the maturation zone of eto1-2 roots. (A) The PI staining of the
 Supplementary Figure 1 PZA inhibits root hair formation as well as cell elongation in the maturation zone of eto1-2 roots. (A) The PI staining of the roots of three-day-old etiolated seedlings of Col-0
Supplementary Figure 1 PZA inhibits root hair formation as well as cell elongation in the maturation zone of eto1-2 roots. (A) The PI staining of the roots of three-day-old etiolated seedlings of Col-0
Electronic Supplementary Materials
 Electronic Supplementary Materials Supplementary Figure S1. Modelling of the coiled-coil based interaction between Mis12 and Nnf1a. 36 alternative structural alignments of Mis12 and Nnf1a proteins on the
Electronic Supplementary Materials Supplementary Figure S1. Modelling of the coiled-coil based interaction between Mis12 and Nnf1a. 36 alternative structural alignments of Mis12 and Nnf1a proteins on the
Supplementary Figure 1. Nature Structural & Molecular Biology: doi: /nsmb.3494
 Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
 DOI: 10.1038/ncb3259 A Ismail et al. Supplementary Figure 1 B 60000 45000 SSC 30000 15000 Live cells 0 0 15000 30000 45000 60000 FSC- PARR 60000 45000 PARR Width 30000 FSC- 15000 Single cells 0 0 15000
DOI: 10.1038/ncb3259 A Ismail et al. Supplementary Figure 1 B 60000 45000 SSC 30000 15000 Live cells 0 0 15000 30000 45000 60000 FSC- PARR 60000 45000 PARR Width 30000 FSC- 15000 Single cells 0 0 15000
Conformation of the Mineralocorticoid Receptor N- terminal Domain: Evidence for Induced and Stable Structure
 ME-10-0005 Conformation of the Mineralocorticoid Receptor N- terminal Domain: Evidence for Induced and Stable Structure Katharina Fischer 1, Sharon M. Kelly 2, Kate Watt 1, Nicholas C. Price 2 and Iain
ME-10-0005 Conformation of the Mineralocorticoid Receptor N- terminal Domain: Evidence for Induced and Stable Structure Katharina Fischer 1, Sharon M. Kelly 2, Kate Watt 1, Nicholas C. Price 2 and Iain
SUPPLEMENTARY INFORMATION FIGURE LEGENDS
 SUPPLEMENTARY INFORMATION FIGURE LEGENDS Fig. S1. Radiation-induced phosphorylation of Rad50 at a specific site. A. Rad50 is an in vitro substrate for ATM. A series of Rad50-GSTs covering the entire molecule
SUPPLEMENTARY INFORMATION FIGURE LEGENDS Fig. S1. Radiation-induced phosphorylation of Rad50 at a specific site. A. Rad50 is an in vitro substrate for ATM. A series of Rad50-GSTs covering the entire molecule
SUPPLEMENTARY INFORMATION. Supplementary Figures 1-8
 SUPPLEMENTARY INFORMATION Supplementary Figures 1-8 Supplementary Figure 1. TFAM residues contacting the DNA minor groove (A) TFAM contacts on nonspecific DNA. Leu58, Ile81, Asn163, Pro178, and Leu182
SUPPLEMENTARY INFORMATION Supplementary Figures 1-8 Supplementary Figure 1. TFAM residues contacting the DNA minor groove (A) TFAM contacts on nonspecific DNA. Leu58, Ile81, Asn163, Pro178, and Leu182
human Cdc45 Figure 1c. (c)
 1 Details of the refined crystallographic model of human Cdc45 and comparison of its active-site region with that of bacterial RecJ. (a) Stereo view of a representative example of the final 2F o -F c electron
1 Details of the refined crystallographic model of human Cdc45 and comparison of its active-site region with that of bacterial RecJ. (a) Stereo view of a representative example of the final 2F o -F c electron
Supplemental Information. Single-Molecule Imaging Reveals How. Mre11-Rad50-Nbs1 Initiates DNA Break Repair
 Molecular Cell, Volume 67 Supplemental Information Single-Molecule Imaging Reveals How Mre11-Rad50-Nbs1 Initiates DNA Break Repair Logan R. Myler, Ignacio F. Gallardo, Michael M. Soniat, Rajashree A. Deshpande,
Molecular Cell, Volume 67 Supplemental Information Single-Molecule Imaging Reveals How Mre11-Rad50-Nbs1 Initiates DNA Break Repair Logan R. Myler, Ignacio F. Gallardo, Michael M. Soniat, Rajashree A. Deshpande,
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Domain architecture and conformational states of the decapping complex, as revealed by structural studies. (a) Domain organization of Schizosaccharomyces pombe (Sp) and Saccharomyces
Supplementary Figure 1 Domain architecture and conformational states of the decapping complex, as revealed by structural studies. (a) Domain organization of Schizosaccharomyces pombe (Sp) and Saccharomyces
Site-specific time-resolved FRET reveals local variations in the unfolding mechanism in an apparently two-state protein unfolding transition
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2017 Supplementary information for Site-specific time-resolved FRET reveals local variations
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2017 Supplementary information for Site-specific time-resolved FRET reveals local variations
RPA-AB RPA-C Supplemental Figure S1: SDS-PAGE stained with Coomassie Blue after protein purification.
 RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid
 Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and
 Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
SUPPLEMENTARY INFORMATION. doi: /nature Human 1 Mouse 1 Xenopus Human 2 Mouse 2 Danio Dros Sulso
 S1 Human 1 Mouse 1 Xenopus Human 2 Mouse 2 Danio Dros Sulso Human 1 Mouse 1 Xenopus Human 2 Mouse 2 Danio Dros Sulso Human 1 Mouse 1 Xenopus Human 2 Mouse 2 Danio Dros Sulso Human 1 Mouse 1 Xenopus Human
S1 Human 1 Mouse 1 Xenopus Human 2 Mouse 2 Danio Dros Sulso Human 1 Mouse 1 Xenopus Human 2 Mouse 2 Danio Dros Sulso Human 1 Mouse 1 Xenopus Human 2 Mouse 2 Danio Dros Sulso Human 1 Mouse 1 Xenopus Human
Supplemental Table 1. List of PCR primers
 Supplemental Table 1. List of PCR primers Primer Sequences (5 to 3 ) 1 TATCCATGGCGCCGGCGGCGAGGGCGGAG 2 TATAAGCTTCTTGGCGTGTCCAGCCCACGGGGCGTAGAACTCGAC 3 ATAAAGCTTGCTCCAGAGTATGAGAAAGCT 4 ATACTCGAGGAGCTCATCCTTGAGAGGCTC
Supplemental Table 1. List of PCR primers Primer Sequences (5 to 3 ) 1 TATCCATGGCGCCGGCGGCGAGGGCGGAG 2 TATAAGCTTCTTGGCGTGTCCAGCCCACGGGGCGTAGAACTCGAC 3 ATAAAGCTTGCTCCAGAGTATGAGAAAGCT 4 ATACTCGAGGAGCTCATCCTTGAGAGGCTC
Figure S1. Sequence alignments of ATRIP and ATR TopBP1 interacting regions.
 A H. sapiens 204 TKLQTS--ERANKLAAPSVSH VSPRKNPSVVIKPEACS-PQFGKTSFPTKESFSANMS LP 259 B. taurus 201 TKLQSS--ERANKLAVPTVSH VSPRKSPSVVIKPEACS-PQFGKPSFPTKESFSANKS LP 257 M. musculus 204 TKSQSN--GRTNKPAAPSVSH
A H. sapiens 204 TKLQTS--ERANKLAAPSVSH VSPRKNPSVVIKPEACS-PQFGKTSFPTKESFSANMS LP 259 B. taurus 201 TKLQSS--ERANKLAVPTVSH VSPRKSPSVVIKPEACS-PQFGKPSFPTKESFSANKS LP 257 M. musculus 204 TKSQSN--GRTNKPAAPSVSH
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2579 Figure S1 Incorporation of heavy isotope-labeled amino acids and enrichment of di-glycine modified peptides. The incorporation of isotopelabeled amino acids in peptides was calculated
DOI: 10.1038/ncb2579 Figure S1 Incorporation of heavy isotope-labeled amino acids and enrichment of di-glycine modified peptides. The incorporation of isotopelabeled amino acids in peptides was calculated
HEK293T. Fig. 1 in the
 Supplementary Information Supplementary Figure 1 Zinc uptake assay of hzip4 and hzip4-δecd transiently expressed in HEK293T cells. The results of one representative e experiment are shown in Fig. 1 in
Supplementary Information Supplementary Figure 1 Zinc uptake assay of hzip4 and hzip4-δecd transiently expressed in HEK293T cells. The results of one representative e experiment are shown in Fig. 1 in
PDIP46 (DNA polymerase δ interacting protein 46) is an activating factor for human DNA polymerase δ
 PDIP46 (DNA polymerase δ interacting protein 46) is an activating factor for human DNA polymerase δ Supplementary Material Figure S1. PDIP46 is associated with Pol isolated by immunoaffinity chromatography.
PDIP46 (DNA polymerase δ interacting protein 46) is an activating factor for human DNA polymerase δ Supplementary Material Figure S1. PDIP46 is associated with Pol isolated by immunoaffinity chromatography.
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature08627 Supplementary Figure 1. DNA sequences used to construct nucleosomes in this work. a, DNA sequences containing the 601 positioning sequence (blue)24 with a PstI restriction site
doi: 10.1038/nature08627 Supplementary Figure 1. DNA sequences used to construct nucleosomes in this work. a, DNA sequences containing the 601 positioning sequence (blue)24 with a PstI restriction site
Supplementary Figure 1
 Supplementary Figure 1 A basic residue of BARD1 promotes Ub-transfer from BRCA1-E2~Ub. A. Scan of external facing BARD1 residues 91-99 for impact on Ub chain formation catalysed by the BRCA1- BARD1 ligase.
Supplementary Figure 1 A basic residue of BARD1 promotes Ub-transfer from BRCA1-E2~Ub. A. Scan of external facing BARD1 residues 91-99 for impact on Ub chain formation catalysed by the BRCA1- BARD1 ligase.
Supplementary Figure S1 Purification of deubiquitinases HEK293 cells were transfected with the indicated DUB-expressing plasmids.
 Supplementary Figure S1 Purification of deubiquitinases HEK293 cells were transfected with the indicated DUB-expressing plasmids. The cells were harvested 72 h after transfection. FLAG-tagged deubiquitinases
Supplementary Figure S1 Purification of deubiquitinases HEK293 cells were transfected with the indicated DUB-expressing plasmids. The cells were harvested 72 h after transfection. FLAG-tagged deubiquitinases
Supplementary Online Material. Structural mimicry in transcription regulation of human RNA polymerase II by the. DNA helicase RECQL5
 Supplementary Online Material Structural mimicry in transcription regulation of human RNA polymerase II by the DNA helicase RECQL5 Susanne A. Kassube, Martin Jinek, Jie Fang, Susan Tsutakawa and Eva Nogales
Supplementary Online Material Structural mimicry in transcription regulation of human RNA polymerase II by the DNA helicase RECQL5 Susanne A. Kassube, Martin Jinek, Jie Fang, Susan Tsutakawa and Eva Nogales
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11988 Supplementary Figure 1. Digestion of model DNA substrates. a, Linearized plasmid DNA (pik31- PstI, lanes 1 and 2), supercoiled plasmid (pik31, lanes 3 and 4), singly nicked plasmid
doi:10.1038/nature11988 Supplementary Figure 1. Digestion of model DNA substrates. a, Linearized plasmid DNA (pik31- PstI, lanes 1 and 2), supercoiled plasmid (pik31, lanes 3 and 4), singly nicked plasmid
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11070 Supplementary Figure 1 Purification of FLAG-tagged proteins. a, Purification of FLAG-RNF12 by FLAG-affinity from nuclear extracts of wild-type (WT) and two FLAG- RNF12 transgenic
doi:10.1038/nature11070 Supplementary Figure 1 Purification of FLAG-tagged proteins. a, Purification of FLAG-RNF12 by FLAG-affinity from nuclear extracts of wild-type (WT) and two FLAG- RNF12 transgenic
SUPPLEMENTARY INFORMATION. Reengineering Protein Interfaces Yields Copper-Inducible Ferritin Cage Assembly
 SUPPLEMENTARY INFORMATION Reengineering Protein Interfaces Yields Copper-Inducible Ferritin Cage Assembly Dustin J. E. Huard, Kathleen M. Kane and F. Akif Tezcan* Department of Chemistry and Biochemistry,
SUPPLEMENTARY INFORMATION Reengineering Protein Interfaces Yields Copper-Inducible Ferritin Cage Assembly Dustin J. E. Huard, Kathleen M. Kane and F. Akif Tezcan* Department of Chemistry and Biochemistry,
Near-infrared optogenetic pair for protein regulation and spectral multiplexing
 Supplementary Information Near-infrared optogenetic pair for protein regulation and spectral multiplexing Taras A. Redchuk 1, Evgeniya S. Omelina 1, Konstantin G. Chernov 1 and Vladislav V. Verkhusha 1,2
Supplementary Information Near-infrared optogenetic pair for protein regulation and spectral multiplexing Taras A. Redchuk 1, Evgeniya S. Omelina 1, Konstantin G. Chernov 1 and Vladislav V. Verkhusha 1,2
The Skap-hom Dimerization and PH Domains Comprise
 Molecular Cell, Volume 32 Supplemental Data The Skap-hom Dimerization and PH Domains Comprise a 3 -Phosphoinositide-Gated Molecular Switch Kenneth D. Swanson, Yong Tang, Derek F. Ceccarelli, Florence Poy,
Molecular Cell, Volume 32 Supplemental Data The Skap-hom Dimerization and PH Domains Comprise a 3 -Phosphoinositide-Gated Molecular Switch Kenneth D. Swanson, Yong Tang, Derek F. Ceccarelli, Florence Poy,
Oligonucleotides were purchased from Eurogentec, purified by denaturing gel electrophoresis
 SUPPLEMENRY INFORMION Purification of probes and Oligonucleotides sequence Oligonucleotides were purchased from Eurogentec, purified by denaturing gel electrophoresis and recovered by electroelution. Labelling
SUPPLEMENRY INFORMION Purification of probes and Oligonucleotides sequence Oligonucleotides were purchased from Eurogentec, purified by denaturing gel electrophoresis and recovered by electroelution. Labelling
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity.
 Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
1 24 C63 C β- β-
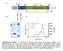 M40 Signal leaved RS1 Domain 59 110 142 Discoidin Domain 223 1 24 63 219 224 + - β- β- M M e e O O H H His 6 -Tag 250 150 100 75 50 37 * 25 20 Supplementary Figure 1 Purification of wild-type retinoschisin.
M40 Signal leaved RS1 Domain 59 110 142 Discoidin Domain 223 1 24 63 219 224 + - β- β- M M e e O O H H His 6 -Tag 250 150 100 75 50 37 * 25 20 Supplementary Figure 1 Purification of wild-type retinoschisin.
Supplementary Information
 4(R/S)-Guanidinylprolyl collagen peptides: On-resin synthesis, complexation with plasmid DNA and the role of peptides in enhancement of transfection Manaswini Nanda, Krishna N. Ganesh,* Chemical Biology
4(R/S)-Guanidinylprolyl collagen peptides: On-resin synthesis, complexation with plasmid DNA and the role of peptides in enhancement of transfection Manaswini Nanda, Krishna N. Ganesh,* Chemical Biology
SUPPLEMENTARY INFORMATION. Tolerance of a knotted near infrared fluorescent protein to random circular permutation
 SUPPLEMENTARY INFORMATION Tolerance of a knotted near infrared fluorescent protein to random circular permutation Naresh Pandey 1,3, Brianna E. Kuypers 2,4, Barbara Nassif 1, Emily E. Thomas 1,3, Razan
SUPPLEMENTARY INFORMATION Tolerance of a knotted near infrared fluorescent protein to random circular permutation Naresh Pandey 1,3, Brianna E. Kuypers 2,4, Barbara Nassif 1, Emily E. Thomas 1,3, Razan
Supplementary Figure 1
 Supplementary Figure 1 Cell cycle distribution (%) 1 8 6 4 2 Cell Cycle G1 S G2 Viability (%) 5 4 3 2 1 Viability Supplementary Figure 1. Cell cycle distribution and viability during DSB repair measurements.
Supplementary Figure 1 Cell cycle distribution (%) 1 8 6 4 2 Cell Cycle G1 S G2 Viability (%) 5 4 3 2 1 Viability Supplementary Figure 1. Cell cycle distribution and viability during DSB repair measurements.
Supplementary Information: Materials and Methods. GST and GST-p53 were purified according to standard protocol after
 Supplementary Information: Materials and Methods Recombinant protein expression and in vitro kinase assay. GST and GST-p53 were purified according to standard protocol after induction with.5mm IPTG for
Supplementary Information: Materials and Methods Recombinant protein expression and in vitro kinase assay. GST and GST-p53 were purified according to standard protocol after induction with.5mm IPTG for
Supplementary Figure S1. Binding of HSA mutants to hfcrn. (a) The levels of titrated amounts of HSA
 Supplementary Figure S1. Binding of HSA mutants to hfcrn. (a) The levels of titrated amounts of HSA variants (5.0-0.002 μg/ml) directly coated in the wells at ph 6.0 were controlled using a horseradish
Supplementary Figure S1. Binding of HSA mutants to hfcrn. (a) The levels of titrated amounts of HSA variants (5.0-0.002 μg/ml) directly coated in the wells at ph 6.0 were controlled using a horseradish
SUPPLEMENTAL DATA SUPPLEMENTAL FIGURE LEGENDS
 SUPPLEMENTAL DATA SUPPLEMENTAL FIGURE LEGENDS SUPPLEMENTAL FIGURE S1. Identification of BmCREC. (A) Amino acid sequences of BmCREC show the peptides identified in LC-MS/MS analysis (marked by red letters
SUPPLEMENTAL DATA SUPPLEMENTAL FIGURE LEGENDS SUPPLEMENTAL FIGURE S1. Identification of BmCREC. (A) Amino acid sequences of BmCREC show the peptides identified in LC-MS/MS analysis (marked by red letters
MT minus end catastrophe
 DOI: 1.138/ncb3241 MT minus end catastrophe frequency [s -1 ].2.15.1.5 control mgfp- TPX2 * * mgfp- TPX2 mini Supplementary Figure 1. TPX2 reduces catastrophes at the microtubule minus ends. Modified box-and-whiskers
DOI: 1.138/ncb3241 MT minus end catastrophe frequency [s -1 ].2.15.1.5 control mgfp- TPX2 * * mgfp- TPX2 mini Supplementary Figure 1. TPX2 reduces catastrophes at the microtubule minus ends. Modified box-and-whiskers
Size-exclusion chromatography TT30 sample was analyzed using a Tosoh Haas TSK Gel G3000SW XL 7.8 mm 30cm column at 1 ml/min and 280 nm detection.
 Analytical ultracentrifugation (AUC) and size exclusion chromatography (SEC-HPLC) were used to assess TT30 molecular weight distribution. The sedimentation distribution of TT30 is shown in Fig. S1A. The
Analytical ultracentrifugation (AUC) and size exclusion chromatography (SEC-HPLC) were used to assess TT30 molecular weight distribution. The sedimentation distribution of TT30 is shown in Fig. S1A. The
Supplemental Material: Rev1 promotes replication through UV lesions in conjunction with DNA
 Supplemental Material: Rev1 promotes replication through UV lesions in conjunction with DNA polymerases,, and, but not with DNA polymerase Jung-Hoon Yoon, Jeseong Park, Juan Conde, Maki Wakamiya, Louise
Supplemental Material: Rev1 promotes replication through UV lesions in conjunction with DNA polymerases,, and, but not with DNA polymerase Jung-Hoon Yoon, Jeseong Park, Juan Conde, Maki Wakamiya, Louise
Supplementary Materials for
 www.advances.sciencemag.org/cgi/content/full/1/7/e1500454/dc1 Supplementary Materials for CRISPR-Cas9 delivery to hard-to-transfect cells via membrane deformation Xin Han, Zongbin Liu, Myeong chan Jo,
www.advances.sciencemag.org/cgi/content/full/1/7/e1500454/dc1 Supplementary Materials for CRISPR-Cas9 delivery to hard-to-transfect cells via membrane deformation Xin Han, Zongbin Liu, Myeong chan Jo,
Supplementary Figure 1. Espn-1 knockout characterization. (a) The predicted recombinant Espn-1 -/- allele was detected by PCR of the left (5 )
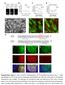 Supplementary Figure 1. Espn-1 knockout characterization. (a) The predicted recombinant Espn-1 -/- allele was detected by PCR of the left (5 ) homologous recombination arm (left) and of the right (3 )
Supplementary Figure 1. Espn-1 knockout characterization. (a) The predicted recombinant Espn-1 -/- allele was detected by PCR of the left (5 ) homologous recombination arm (left) and of the right (3 )
TALEN mediated targeted editing of GM2/GD2-synthase gene modulates anchorage independent growth by reducing anoikis resistance in mouse tumor cells
 Supplementary Information for TALEN mediated targeted editing of GM2/GD2-synthase gene modulates anchorage independent growth by reducing anoikis resistance in mouse tumor cells Barun Mahata 1, Avisek
Supplementary Information for TALEN mediated targeted editing of GM2/GD2-synthase gene modulates anchorage independent growth by reducing anoikis resistance in mouse tumor cells Barun Mahata 1, Avisek
Supplementary Information. Human Antibody-Based Chemically Induced Dimerizers for Cell Therapeutic Applications
 Supplementary Information Human Antibody-Based Chemically Induced Dimerizers for Cell Therapeutic Applications Zachary B Hill 1,4, Alexander J Martinko 1,2,4, Duy P Nguyen 1 & James A Wells *1,3 1 Department
Supplementary Information Human Antibody-Based Chemically Induced Dimerizers for Cell Therapeutic Applications Zachary B Hill 1,4, Alexander J Martinko 1,2,4, Duy P Nguyen 1 & James A Wells *1,3 1 Department
Nature Structural & Molecular Biology: doi: /nsmb.3018
 Supplementary Figure 1 Validation of genetic complementation assay in Bmal1 / Per2 Luc fibroblasts. (a) Only Bmal1, not Bmal2, rescues circadian rhythms from cells. Cells expressing various Bmal constructs
Supplementary Figure 1 Validation of genetic complementation assay in Bmal1 / Per2 Luc fibroblasts. (a) Only Bmal1, not Bmal2, rescues circadian rhythms from cells. Cells expressing various Bmal constructs
Supplemental Materials and Methods
 Supplemental Materials and Methods Co-immunoprecipitation (Co-IP) assay Cells were lysed with NETN buffer (20 mm Tris-HCl, ph 8.0, 0 mm NaCl, 1 mm EDT, 0.5% Nonidet P-40) containing 50 mm β-glycerophosphate,
Supplemental Materials and Methods Co-immunoprecipitation (Co-IP) assay Cells were lysed with NETN buffer (20 mm Tris-HCl, ph 8.0, 0 mm NaCl, 1 mm EDT, 0.5% Nonidet P-40) containing 50 mm β-glycerophosphate,
Stabilization of a virus-like particle and its application as a nanoreactor at physiological conditions
 Supporting Information Stabilization of a virus-like particle and its application as a nanoreactor at physiological conditions Lise Schoonen, b Sjors Maassen, b Roeland J. M. Nolte b and Jan C. M. van
Supporting Information Stabilization of a virus-like particle and its application as a nanoreactor at physiological conditions Lise Schoonen, b Sjors Maassen, b Roeland J. M. Nolte b and Jan C. M. van
SUPPLEMENTAL FIGURES AND TABLES
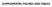 SUPPLEMENTAL FIGURES AND TABLES A B Flag-ALDH1A1 IP: α-ac HEK293T WT 91R 128R 252Q 367R 41/ 419R 435R 495R 412R C Flag-ALDH1A1 NAM IP: HEK293T + + - + D NAM - + + E Relative ALDH1A1 activity 1..8.6.4.2
SUPPLEMENTAL FIGURES AND TABLES A B Flag-ALDH1A1 IP: α-ac HEK293T WT 91R 128R 252Q 367R 41/ 419R 435R 495R 412R C Flag-ALDH1A1 NAM IP: HEK293T + + - + D NAM - + + E Relative ALDH1A1 activity 1..8.6.4.2
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Supplementary information:
 Supplementary information: Distinct amino acids in the C-linker domain of the plant K + subcellular localization and activity at the plasma membrane channel KAT2 determine its Nieves-Cordones et al. Part
Supplementary information: Distinct amino acids in the C-linker domain of the plant K + subcellular localization and activity at the plasma membrane channel KAT2 determine its Nieves-Cordones et al. Part
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling
 Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplementary information
 Supplementary information The E3 ligase RNF8 regulates KU80 removal and NHEJ repair Lin Feng 1, Junjie Chen 1 1 Department of Experimental Radiation Oncology, The University of Texas M. D. Anderson Cancer
Supplementary information The E3 ligase RNF8 regulates KU80 removal and NHEJ repair Lin Feng 1, Junjie Chen 1 1 Department of Experimental Radiation Oncology, The University of Texas M. D. Anderson Cancer
Supplementary Figure Legends
 Supplementary Figure Legends Figure S1 gene targeting strategy for disruption of chicken gene, related to Figure 1 (f)-(i). (a) The locus and the targeting constructs showing HpaI restriction sites. The
Supplementary Figure Legends Figure S1 gene targeting strategy for disruption of chicken gene, related to Figure 1 (f)-(i). (a) The locus and the targeting constructs showing HpaI restriction sites. The
Supplementary Figure 1 Structural modeling and purification of V. cholerae ABH. (a) The migration of the purified rabh and catalytically inactive
 Supplementary Figure 1 Structural modeling and purification of V. cholerae ABH. (a) The migration of the purified rabh and catalytically inactive variants rabhs, rabhd, and rabhh are shown on 12% SDS-PAGE
Supplementary Figure 1 Structural modeling and purification of V. cholerae ABH. (a) The migration of the purified rabh and catalytically inactive variants rabhs, rabhd, and rabhh are shown on 12% SDS-PAGE
Hoffmann et al., http ://www.jcb.org /cgi /content /full /jcb /DC1
 Supplemental material JCB Hoffmann et al., http ://www.jcb.org /cgi /content /full /jcb.201506071 /DC1 THE JOU RNAL OF CELL BIO LOGY Figure S1. TRA IP harbors a PCNA-binding PIP box required for its recruitment
Supplemental material JCB Hoffmann et al., http ://www.jcb.org /cgi /content /full /jcb.201506071 /DC1 THE JOU RNAL OF CELL BIO LOGY Figure S1. TRA IP harbors a PCNA-binding PIP box required for its recruitment
Figure S1, related to Figure 1. Characterization of biosensor behavior in vivo.
 Developmental Cell, Volume 23 Supplemental Information Separase Biosensor Reveals that Cohesin Cleavage Timing Depends on Phosphatase PP2A Cdc55 Regulation Gilad Yaakov, Kurt Thorn, and David O. Morgan
Developmental Cell, Volume 23 Supplemental Information Separase Biosensor Reveals that Cohesin Cleavage Timing Depends on Phosphatase PP2A Cdc55 Regulation Gilad Yaakov, Kurt Thorn, and David O. Morgan
Coleman et al., Supplementary Figure 1
 Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
supplementary information
 DOI: 10.1038/ncb2172 Figure S1 p53 regulates cellular NADPH and lipid levels via inhibition of G6PD. (a) U2OS cells stably expressing p53 shrna or a control shrna were transfected with control sirna or
DOI: 10.1038/ncb2172 Figure S1 p53 regulates cellular NADPH and lipid levels via inhibition of G6PD. (a) U2OS cells stably expressing p53 shrna or a control shrna were transfected with control sirna or
Supplementary
 Supplementary information Supplementary Material and Methods Plasmid construction The transposable element vectors for inducible expression of RFP-FUS wt and EGFP-FUS R521C and EGFP-FUS P525L were derived
Supplementary information Supplementary Material and Methods Plasmid construction The transposable element vectors for inducible expression of RFP-FUS wt and EGFP-FUS R521C and EGFP-FUS P525L were derived
SUPPLEMENTARY INFORMATION
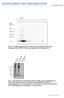 doi:10.1038/nature10324 Fig. S1: Two dimensional IEF/SDS-PAGE/Western blot analysis of RBC lysate crosslinked with 4 mm BS 3. The blot was probed with αsyn antibody C20. Fig. S2: SDS-PAGE/silver stain
doi:10.1038/nature10324 Fig. S1: Two dimensional IEF/SDS-PAGE/Western blot analysis of RBC lysate crosslinked with 4 mm BS 3. The blot was probed with αsyn antibody C20. Fig. S2: SDS-PAGE/silver stain
Bootcamp: Molecular Biology Techniques and Interpretation
 Bootcamp: Molecular Biology Techniques and Interpretation Bi8 Winter 2016 Today s outline Detecting and quantifying nucleic acids and proteins: Basic nucleic acid properties Hybridization PCR and Designing
Bootcamp: Molecular Biology Techniques and Interpretation Bi8 Winter 2016 Today s outline Detecting and quantifying nucleic acids and proteins: Basic nucleic acid properties Hybridization PCR and Designing
Chapter 14 Regulation of Transcription
 Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Modulating the Cascade architecture of a minimal Type I-F CRISPR-Cas system
 SUPPLEMENTARY DATA Modulating the Cascade architecture of a minimal Type I-F CRISPR-Cas system Daniel Gleditzsch 1, Hanna Müller-Esparza 1, Patrick Pausch 2,3, Kundan Sharma 4, Srivatsa Dwarakanath 1,
SUPPLEMENTARY DATA Modulating the Cascade architecture of a minimal Type I-F CRISPR-Cas system Daniel Gleditzsch 1, Hanna Müller-Esparza 1, Patrick Pausch 2,3, Kundan Sharma 4, Srivatsa Dwarakanath 1,
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or
 Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Supplementary Information Crystal structure, biochemical and cellular activities demonstrate separate functions of MTH1 and MTH2
 Supplementary Information Crystal structure, biochemical and cellular activities demonstrate separate functions of MTH1 and MTH2 Supplementary Figure 1 Comparison of activities of NUDIX proteins with nucleotide
Supplementary Information Crystal structure, biochemical and cellular activities demonstrate separate functions of MTH1 and MTH2 Supplementary Figure 1 Comparison of activities of NUDIX proteins with nucleotide
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature10258 Supplementary Figure 1 Reconstitution of the CENP-A nucleosome with recombinant human histones H2A, H2B, H4, and CENP-A. a, Purified recombinant human
SUPPLEMENTARY INFORMATION doi:10.1038/nature10258 Supplementary Figure 1 Reconstitution of the CENP-A nucleosome with recombinant human histones H2A, H2B, H4, and CENP-A. a, Purified recombinant human
Structural and functional analysis of Hikeshi, a new nuclear transport receptor of Hsp70
 Supporting information Volume 71 (2015) Supporting information for article: Structural and functional analysis of Hikeshi, a new nuclear transport receptor of Hsp70 Jinsue Song, Shingo Kose, Ai Watanabe,
Supporting information Volume 71 (2015) Supporting information for article: Structural and functional analysis of Hikeshi, a new nuclear transport receptor of Hsp70 Jinsue Song, Shingo Kose, Ai Watanabe,
Supplemental Information
 Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
Supplementary Information for. Regulation of Rev1 by the Fanconi Anemia Core Complex
 Supplementary Information for Regulation of Rev1 by the Fanconi Anemia Core Complex Hyungjin Kim, Kailin Yang, Donniphat Dejsuphong, Alan D. D Andrea* *Corresponding Author: Alan D. D Andrea, M.D. Alan_dandrea@dfci.harvard.edu
Supplementary Information for Regulation of Rev1 by the Fanconi Anemia Core Complex Hyungjin Kim, Kailin Yang, Donniphat Dejsuphong, Alan D. D Andrea* *Corresponding Author: Alan D. D Andrea, M.D. Alan_dandrea@dfci.harvard.edu
SUPPLEMENTAL MATERIAL. Supplemental material contains Supplemental Figure Legends and Supplemental Figures 1 to
 SUPPLEMENTAL MATERIAL Supplemental material contains Supplemental Figure Legends and Supplemental Figures 1 to 6. SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure 1. Overview of the mechanisms by which
SUPPLEMENTAL MATERIAL Supplemental material contains Supplemental Figure Legends and Supplemental Figures 1 to 6. SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure 1. Overview of the mechanisms by which
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/NCHEM.1805 Visualization and Selective Chemical Targeting of RNA G-quadruplex Structures in the Cytoplasm of Human Cells Giulia Biffi 1, Marco Di Antonio 2, David Tannahill 1 and Shankar Balasubramanian
DOI: 10.1038/NCHEM.1805 Visualization and Selective Chemical Targeting of RNA G-quadruplex Structures in the Cytoplasm of Human Cells Giulia Biffi 1, Marco Di Antonio 2, David Tannahill 1 and Shankar Balasubramanian
SUPPLEMENTARY INFORMATION. Design and Characterization of Bivalent BET Inhibitors
 SUPPLEMENTARY INFORMATION Design and Characterization of Bivalent BET Inhibitors Minoru Tanaka 1,2,#, Justin M. Roberts 1,#, Hyuk-Soo Seo 3, Amanda Souza 1, Joshiawa Paulk 1, Thomas G. Scott 1, Stephen
SUPPLEMENTARY INFORMATION Design and Characterization of Bivalent BET Inhibitors Minoru Tanaka 1,2,#, Justin M. Roberts 1,#, Hyuk-Soo Seo 3, Amanda Souza 1, Joshiawa Paulk 1, Thomas G. Scott 1, Stephen
Supplementary Note 1. Enzymatic properties of the purified Syn BVR
 Supplementary Note 1. Enzymatic properties of the purified Syn BVR The expression vector pet15b-syn bvr allowed us to routinely prepare 15 mg of electrophoretically homogenous Syn BVR from 2.5 L of TB-medium
Supplementary Note 1. Enzymatic properties of the purified Syn BVR The expression vector pet15b-syn bvr allowed us to routinely prepare 15 mg of electrophoretically homogenous Syn BVR from 2.5 L of TB-medium
The molecular basis of lysine 48 ubiquitin chain synthesis by Ube2K
 Supplementary Information The molecular basis of lysine 48 ubiquitin chain synthesis by Adam J. Middleton, Catherine L. Day* Department of Biochemistry, Otago School of Medical Sciences, University of
Supplementary Information The molecular basis of lysine 48 ubiquitin chain synthesis by Adam J. Middleton, Catherine L. Day* Department of Biochemistry, Otago School of Medical Sciences, University of
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06721 SUPPLEMENTARY INFORMATION. Supplemental Figure Legends Supplemental Figure 1 The distribution of hatx-1[82q] in Cos7 cells. Cos7 cells are co-transfected with hatx-1[82q]-gfp (green)
doi: 10.1038/nature06721 SUPPLEMENTARY INFORMATION. Supplemental Figure Legends Supplemental Figure 1 The distribution of hatx-1[82q] in Cos7 cells. Cos7 cells are co-transfected with hatx-1[82q]-gfp (green)
Supplementary Information
 Supplementary Information Self-assembled Messenger RNA Nanoparticles (mrna-nps) for Efficient Gene Expression Hyejin Kim 1, Yongkuk Park 1 and Jong Bum Lee 1, * 1 Department of Chemical Engineering, University
Supplementary Information Self-assembled Messenger RNA Nanoparticles (mrna-nps) for Efficient Gene Expression Hyejin Kim 1, Yongkuk Park 1 and Jong Bum Lee 1, * 1 Department of Chemical Engineering, University
Gene Forward Primer Reverse Primer GAPDH ATCATCCCTGCCTCTACTGG GTCAGGTCCACCACTGACAC SSB1 AACTTCAGTGAGCCAAACCC GTTCTCAGAGGCTGGAGAGG
 Supplemental Data EXPERIMENTAL PROCEDURES Plasmids and Antibodies- Full length cdna of INT11 or INT12 were cloned into ps- Flag-SBP vector respectively. Anti-RNA pol II (RPB1) was purchased from Santa
Supplemental Data EXPERIMENTAL PROCEDURES Plasmids and Antibodies- Full length cdna of INT11 or INT12 were cloned into ps- Flag-SBP vector respectively. Anti-RNA pol II (RPB1) was purchased from Santa
Nature Methods: doi: /nmeth Supplementary Figure 1. Comparison of REC2 and BCL-xL domains and modeling domain replacement.
 Supplementary Figure 1 Comparison of REC2 and BCL-xL domains and modeling domain replacement. (a) The REC2 domain of Cas9 and (b) BCL-xL display similar globular structures and volumes. (c, d) Modeling
Supplementary Figure 1 Comparison of REC2 and BCL-xL domains and modeling domain replacement. (a) The REC2 domain of Cas9 and (b) BCL-xL display similar globular structures and volumes. (c, d) Modeling
SUPPORTING INFORMATION.
 SUPPORTING INFORMATION. Structural and dynamic changes associated with beneficial engineered single amino acid deletion mutations in enhanced Green Fluorescent Protein James A. J. Arpino 1, Pierre J. Rizkallah
SUPPORTING INFORMATION. Structural and dynamic changes associated with beneficial engineered single amino acid deletion mutations in enhanced Green Fluorescent Protein James A. J. Arpino 1, Pierre J. Rizkallah
Supplementary Information. Structural basis for duplex RNA recognition and cleavage by A.
 Supplementary Information Structural asis for duplex RNA recognition and cleavage y A. fulgidus C3PO Eneida arizotto 1, Edward D Lowe 1 & James S Parker 1 1 Department of Biochemistry University of Oxford
Supplementary Information Structural asis for duplex RNA recognition and cleavage y A. fulgidus C3PO Eneida arizotto 1, Edward D Lowe 1 & James S Parker 1 1 Department of Biochemistry University of Oxford
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
2.5. Equipment and materials supplied by user PCR based template preparation Influence of temperature on in vitro EGFP synthesis 11
 Manual 15 Reactions LEXSY in vitro Translation Cell-free protein expression kit based on Leishmania tarentolae for PCR-based template generation Cat. No. EGE-2010-15 FOR RESEARCH USE ONLY. NOT INTENDED
Manual 15 Reactions LEXSY in vitro Translation Cell-free protein expression kit based on Leishmania tarentolae for PCR-based template generation Cat. No. EGE-2010-15 FOR RESEARCH USE ONLY. NOT INTENDED
Supporting Information for: DNA-based delivery vehicles: ph-controlled disassembly and cargo release by Jung-Won Keum and Harry Bermudez
 Supporting Information for: DNA-based delivery vehicles: ph-controlled disassembly and cargo release by Jung-Won Keum and Harry Bermudez DNA sequences Strand Sequence 1- GGGTTAGGGTTAGGGTTAGGGAGGGTTAGGGTTAGGGTTAGGGAGGGTTAGGGTTAGGGTTAGGG
Supporting Information for: DNA-based delivery vehicles: ph-controlled disassembly and cargo release by Jung-Won Keum and Harry Bermudez DNA sequences Strand Sequence 1- GGGTTAGGGTTAGGGTTAGGGAGGGTTAGGGTTAGGGTTAGGGAGGGTTAGGGTTAGGGTTAGGG
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Contents: Supplementary Figure 1. Additional structural and binding data for designed tuim peptides. Supplementary Figure 2. Subcellular localization patterns of designed tuim
SUPPLEMENTARY INFORMATION Contents: Supplementary Figure 1. Additional structural and binding data for designed tuim peptides. Supplementary Figure 2. Subcellular localization patterns of designed tuim
In Vitro Monitoring of the Formation of Pentamers from the Monomer of GST Fused HPV 16 L1
 This journal is The Royal Society of Chemistry 213 In Vitro Monitoring of the Formation of Pentamers from the Monomer of GST Fused HPV 16 L1 Dong-Dong Zheng, a Dong Pan, a Xiao Zha, ac Yuqing Wu,* a Chunlai
This journal is The Royal Society of Chemistry 213 In Vitro Monitoring of the Formation of Pentamers from the Monomer of GST Fused HPV 16 L1 Dong-Dong Zheng, a Dong Pan, a Xiao Zha, ac Yuqing Wu,* a Chunlai
Supplementary Figure 1. Schematic for the growth of high-quality uniform
 Supplementary Figure 1. Schematic for the growth of high-quality uniform monolayer WS 2 by ambient-pressure CVD. Supplementary Figure 2. Schematic structures of the initial state (IS) and the final state
Supplementary Figure 1. Schematic for the growth of high-quality uniform monolayer WS 2 by ambient-pressure CVD. Supplementary Figure 2. Schematic structures of the initial state (IS) and the final state
Supplemental Data. Cui et al. (2012). Plant Cell /tpc a b c d. Stem UBC32 ACTIN
 A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
Supplementary information
 Supplementary information Supplementary figures Figure S1 Level of mycdet1 protein in DET1 OE-1, OE-2 and OE-3 transgenic lines. Total protein extract from wild type Col0, det1-1 mutant and DET1 OE lines
Supplementary information Supplementary figures Figure S1 Level of mycdet1 protein in DET1 OE-1, OE-2 and OE-3 transgenic lines. Total protein extract from wild type Col0, det1-1 mutant and DET1 OE lines
A Domain Swapping Study of Nano-Capsule Proteins. Rongli Fan, Aimee L. Boyle, Vee Vee Cheong, See Liang Ng, Brendan P. Orner*
 A Domain Swapping Study of Nano-Capsule Proteins Rongli Fan, Aimee L. Boyle, Vee Vee Cheong, See Liang Ng, Brendan P. Orner* Figure S1. Amino acid sequences of proteins used in this study. Grey shading
A Domain Swapping Study of Nano-Capsule Proteins Rongli Fan, Aimee L. Boyle, Vee Vee Cheong, See Liang Ng, Brendan P. Orner* Figure S1. Amino acid sequences of proteins used in this study. Grey shading
Supplementary Information For. A genetically encoded tool for manipulation of NADP + /NADPH in living cells
 Supplementary Information For A genetically encoded tool for manipulation of NADP + /NADPH in living cells Valentin Cracan 1,2,3, Denis V. Titov 1,2,3, Hongying Shen 1,2,3, Zenon Grabarek 1* and Vamsi
Supplementary Information For A genetically encoded tool for manipulation of NADP + /NADPH in living cells Valentin Cracan 1,2,3, Denis V. Titov 1,2,3, Hongying Shen 1,2,3, Zenon Grabarek 1* and Vamsi
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Multiple sequence alignments of four Swi2/Snf2 subfamily proteins, ScChd1, SsoRad54 and the RNA helicase Vasa. The sequence alignments of the Swi2/Snf2 subfamily proteins, ScChd1
Supplementary Figure 1 Multiple sequence alignments of four Swi2/Snf2 subfamily proteins, ScChd1, SsoRad54 and the RNA helicase Vasa. The sequence alignments of the Swi2/Snf2 subfamily proteins, ScChd1
Supplemental Data. Furlan et al. Plant Cell (2017) /tpc
 Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Figure.
Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Figure.
(phosphatase tensin) domain is shown in dark gray, the FH1 domain in black, and the
 Supplemental Figure 1. Predicted Domain Organization of the AFH14 Protein. (A) Schematic representation of the predicted domain organization of AFH14. The PTEN (phosphatase tensin) domain is shown in dark
Supplemental Figure 1. Predicted Domain Organization of the AFH14 Protein. (A) Schematic representation of the predicted domain organization of AFH14. The PTEN (phosphatase tensin) domain is shown in dark
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature09937 a Name Position Primersets 1a 1b 2 3 4 b2 Phenotype Genotype b Primerset 1a D T C R I E 10000 8000 6000 5000 4000 3000 2500 2000 1500 1000 800 Donor (D)
SUPPLEMENTARY INFORMATION doi:10.1038/nature09937 a Name Position Primersets 1a 1b 2 3 4 b2 Phenotype Genotype b Primerset 1a D T C R I E 10000 8000 6000 5000 4000 3000 2500 2000 1500 1000 800 Donor (D)
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1. Analyses of ECTRs by C-circle and T-circle assays.
 Supplementary Figure 1 Analyses of ECTRs by C-circle and T-circle assays. (a) C-circle and (b) T-circle amplification reactions using genomic DNA from different cell lines in the presence (+) or absence
Supplementary Figure 1 Analyses of ECTRs by C-circle and T-circle assays. (a) C-circle and (b) T-circle amplification reactions using genomic DNA from different cell lines in the presence (+) or absence
