Gene Prediction Final Presentation
|
|
|
- Peter Pope
- 6 years ago
- Views:
Transcription
1 Gene Prediction Final Presentation
2 Final Proposed Pipeline Assembled Genome Protein - coding Gene Prediction Ab Initio Prodigal Glimmer GeneMarkS RNA Gene Prediction ncrna Specific trnascanse (trna) RNAmmer (rrna) Infernal Validation BLAST (RefSeq) DEG Merge Validation Final Output
3 Methods for Merging Results Ab initio Tools: GeneMarkS and Prodigal Overlap Criteria: -If there was total overlap, keep the gene -If overlap is not an exact match, then validate via homology 1.Overlap between each pair and between both tools 2.Find overlap between GeneMarkS and Prodigal. Predicted genes with partial or no overlap will then be checked against homology
4 Merging Results from Ab-initio Tools Genes detected by both tools Exact Match Stop Codon Match >60 bp Overlap Genes detected by only one tool Putative Gene List
5 Correcting Previous Results: Reference Genome 2 Tool Exact Match Stop Overlap Extra (FP) Missed (FN) Sensitivity GeneMark- S % 96.7% Prodigal % 98.0% PPV GLIMMER % 92.6% FGenesB % 97.6% Homology % 93.2% Complete Union of Prodigal and GeneMarkS Overlap Union of Prodigal and GeneMarkS % 91.5% % 94.1%
6 Average number of genes predicted by Ab-initio Prodigal tools GMS Prodigal Only* 130 Overlap > 60bp 326 Exact Matches 4257 GMS Only 103
7 Ab initio results
8 Ab initio results
9 Blast Validation Run BLASTx on putative gene list to aid in finding true genes Filtered output Query coverage >95% and Percent identity >95% E-val threshold 1e-10 Putative Genes List: Genes predicted by ab initio programs True positives: Genes verified (by BLASTx) Novel genes: Genes predicted by all ab inito programs but no BLAST hits False positives: Genes predicted by 1 tool but no BLAST hits
10 PREDICTION STATISTICS Sample ID True Positive: Homology G+P NOVEL: Homology G+P False Positive G (or) P OB OB OB OB OB OB OB OB OB OB OB OB Sample ID True Positive: Homology G+P NOVEL: Homology G+P False Positive G (or) P OB OB OB OB OB OB OB OB OB OB OB OB
11 Database for Essential Genes Database contains minimal gene set needed to support cellular life Blast against Database to identify homologs in our predicted genes Salmonella essential genes ~635
12 Sample Predicted Genes Sample Predicted Genes Sample Predicted Genes OB OB OB OB OB OB OB OB OB OB OB OB OB OB OB OB OB OB OB OB OB OB OB OB
13 trnascan-se Prediction Results Command: trnascan-se -P../assembly/OB0001.plasmid.scaffolds.fasta awk -f ~/awks/converttrnascantogtf > OB0001.plasmid.scaffolds.gtf Evaluation according to the NCBI Prokaryotic Genome Annotation Pipeline No trnascan-se scores below of 23 assemblies had a trna for each of 21 amino acids plus selenocysteine OB0005 missing Glu OB0006 missing Glu OB0009 missing Glu, His, Trp, and Ile Average number of predicted trnas was 80.6, the minimum was 74, and the maximum was 90
14 RNAmmer Prediction result Command:./rnammer -S bac -multi -gff output.gff < input.fasta -S kingdom bac -multi runs all molecules and both strands in parallel -gff/ -f: Output formats
15 Rfam with Infernal Prediction of all RNAs Command: cmscan --tblout <cmdb> <seqfile> Input: FASTA Output: Infernal specific table format gff Format File {if (substr($0, 0, 1)!= "#") printf("%- 30s infernal%- 20s %- 10s %- 10s %- 8s %- 4s\n", $3, $1, $8, $9, $15, $10); } Command: awk -f File infernal-table-file > output-file.gff Data is merged based on the scoring system of Infernal
16 Infernal Prediction Result
17 Merging rrna, trna, and CDS Predictions CDS Overlap rrna? No >30bp Overlap trna? No Keep CDS Yes Yes Discard CDS Pseudo or atypical trna Not hypothetical CDS? No Hypothetical CDS trna neither pseudo or atypical? No Yes Ye s Keep Both Keep CDS only Keep trna only
18 ncrna CDS Merging Results Rules: If a protein coding gene overlaps with an rrna gene, it is discarded. If a protein coding gene overlaps by 30 bases or more with a trna gene either may be discarded according to the following: Results: If the protein coding gene is homologous to a CDS gene we consider it a confident prediction. If the trna gene is not a pseudogene we consider it a confident prediction. If either prediction is not a confident prediction, it is discarded, unless both aren t, in which case both are kept 2 protein coding genes from OB0006 removed due to trna conflict 1 protein coding gene from OB0013 removed due to trna conflict
19 References 1. Lukashin A, Borodovsky M: GeneMark.hmm: new solutions for gene finding. Nucleic Acids Res 1998, 26(4): /nar/ Delcher A, Bratke K, Powers E, Salzberg S: Identifying bacterial genes and endosymbiont DNA with Glimmer. Bioinformatics 2007, 23(6): /bioinformatics/btm Lomsadze, A., Tang, S., Gemayel, K., & Borodovsky, M. GeneMarkS-2: Raising Standards of Accuracy in Gene Recognition. 4. Besemer, J., Lomsadze, A., & Borodovsky, M. (2001). GeneMarkS: a self-training method for prediction of gene starts in microbial genomes. Implications for finding sequence motifs in regulatory regions. Nucleic acids research, 29(12), Bocs S, Cruveiller S, Vallenet D, Nuel G, Médigue C. AMIGene: Annotation of MIcrobial Genes. Nucleic Acids Research. 2003;31(13): Oliynyk et al., (2007) Complete genome sequence of the erythromycin-producing bacterium Saccharopolyspora erythraea NRRL23338 Nature Biotechnology 25, John E. Karro, Yangpan Yan, Deyou Zheng, Zhaolei Zhang, Nicholas Carriero, Philip Cayting, Paul Harrrison, Mark Gerstein; Pseudogene.org: a comprehensive database and comparison platform for pseudogene annotation. Nucleic Acids Res 2007; 35 (suppl_1): D55-D60. doi: /nar/gkl Hori H. Methylated nucleosides in trna and trna methyltransferases. Frontiers in Genetics. 2014;5:144. doi: /fgene Gong H, Vu G-P, Bai Y, et al. A Salmonella Small Non-Coding RNA Facilitates Bacterial Invasion and Intracellular Replication by Modulating the Expression of Virulence Factors. Monack DM, ed. PLoS Pathogens. 2011;7(9):e doi: /journal.ppat Sweeney B.A., Roy P., Leontis N.B. (2015) An introduction to recurrent nucleotide interactions in RNA. Wiley Interdisciplinary Reviews: RNA, 6, Harris KA, Lünse CE, Li S, Brewer KI, Breaker RR. Biochemical analysis of pistol self-cleaving ribozymes. RNA. 2015;21(11): doi: /rna Tjaden B, Goodwin SS, Opdyke JA, et al. Target prediction for small, noncoding RNAs in bacteria. Nucleic Acids Research. 2006;34(9): doi: /nar/gkl Lowe TM, Eddy SR. trnascan-se: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Research. 1997;25(5): Lasleett D, Canback B. ARAGORN, a program to detect trna genes and tmrna gnes in nucleotide sequences. Nucleic Acids Research. 2004;32(1): doi: /nar/gkh Pearson, W. R. (2013). An Introduction to Sequence Similarity ( Homology ) Searching. Current Protocols in Bioinformatics / Editoral Board, Andreas D. Baxevanis... [et Al.], 0 3, / bi0301s42.
Background and Strategy
 Background and Strategy Background Algorithm Ab-initio tools Homology based tools RNA prediction tool Pseudogenes Validation References Gene: DNA sequence that codes for amino acids in a protein Key step
Background and Strategy Background Algorithm Ab-initio tools Homology based tools RNA prediction tool Pseudogenes Validation References Gene: DNA sequence that codes for amino acids in a protein Key step
Gene Prediction: Preliminary Results
 Gene Prediction: Preliminary Results Outline Preliminary Pipeline Programs Program Comparison Tests Metrics Gene Prediction Tools: Usage + Results GeneMarkS Glimmer 3.0 Prodigal BLAST ncrna Prediction
Gene Prediction: Preliminary Results Outline Preliminary Pipeline Programs Program Comparison Tests Metrics Gene Prediction Tools: Usage + Results GeneMarkS Glimmer 3.0 Prodigal BLAST ncrna Prediction
Gene Prediction Group
 Group Ben, Jasreet, Jeff, Jia, Kunal TACCTGAAAAAGCACATAATACTTATGCGTATCCGCCCTAAACACTGCCTTCTTTCTCAA AGAAGATGTCGCCGCTTTTCAACCGAACGATGTGTTCTTCGCCGTTTTCTCGGTAGTGCA TATCGATGATTCACGTTTCGGCAGTGCAGGCACCGGCGCATATTCAGGATACCGGACGCT
Group Ben, Jasreet, Jeff, Jia, Kunal TACCTGAAAAAGCACATAATACTTATGCGTATCCGCCCTAAACACTGCCTTCTTTCTCAA AGAAGATGTCGCCGCTTTTCAACCGAACGATGTGTTCTTCGCCGTTTTCTCGGTAGTGCA TATCGATGATTCACGTTTCGGCAGTGCAGGCACCGGCGCATATTCAGGATACCGGACGCT
Gene Prediction. Lab & Preliminary Results. Faction 2 Saturday, March 11, 2017
 Gene Prediction Lab & Preliminary Results Faction 2 Saturday, March 11, 2017 Group Members: Michelle Kim Khushbu Patel Krithika Xinrui Zhou Chen Lin Sujun Zhao Hannah Hatchell rohini mopuri Jack Cartee
Gene Prediction Lab & Preliminary Results Faction 2 Saturday, March 11, 2017 Group Members: Michelle Kim Khushbu Patel Krithika Xinrui Zhou Chen Lin Sujun Zhao Hannah Hatchell rohini mopuri Jack Cartee
Prokaryotic Annotation Pipeline SOP HGSC, Baylor College of Medicine
 1 Abstract A prokaryotic annotation pipeline was developed to automatically annotate draft and complete bacterial genomes. The protein coding genes in the genomes are predicted by the combination of Glimmer
1 Abstract A prokaryotic annotation pipeline was developed to automatically annotate draft and complete bacterial genomes. The protein coding genes in the genomes are predicted by the combination of Glimmer
Gene Prediction Background & Strategy Faction 2 February 22, 2017
 Gene Prediction Background & Strategy Faction 2 February 22, 2017 Group Members: Michelle Kim Khushbu Patel Krithika Xinrui Zhou Chen Lin Sujun Zhao Hannah Hatchell rohini mopuri Jack Cartee Introduction
Gene Prediction Background & Strategy Faction 2 February 22, 2017 Group Members: Michelle Kim Khushbu Patel Krithika Xinrui Zhou Chen Lin Sujun Zhao Hannah Hatchell rohini mopuri Jack Cartee Introduction
Gene Prediction Background & Strategy. February 24, 2016
 Gene Prediction Background & Strategy February 24, 2016 overview background ab initio prediction tools rna prediction tools homology-based prediction tools combo tools final statements Gene Prediction
Gene Prediction Background & Strategy February 24, 2016 overview background ab initio prediction tools rna prediction tools homology-based prediction tools combo tools final statements Gene Prediction
Bacterial Genome Annotation
 Bacterial Genome Annotation Bacterial Genome Annotation For an annotation you want to predict from the sequence, all of... protein-coding genes their stop-start the resulting protein the function the control
Bacterial Genome Annotation Bacterial Genome Annotation For an annotation you want to predict from the sequence, all of... protein-coding genes their stop-start the resulting protein the function the control
Genome sequence of Acinetobacter baumannii MDR-TJ
 JB Accepts, published online ahead of print on 11 March 2011 J. Bacteriol. doi:10.1128/jb.00226-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
JB Accepts, published online ahead of print on 11 March 2011 J. Bacteriol. doi:10.1128/jb.00226-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
GeneMarkS-2: Raising Standards of Accuracy in Gene Recognition
 GeneMarkS-2: Raising Standards of Accuracy in Gene Recognition Alexandre Lomsadze 1^, Shiyuyun Tang 2^, Karl Gemayel 3^ and Mark Borodovsky 1,2,3 ^ joint first authors 1 Wallace H. Coulter Department of
GeneMarkS-2: Raising Standards of Accuracy in Gene Recognition Alexandre Lomsadze 1^, Shiyuyun Tang 2^, Karl Gemayel 3^ and Mark Borodovsky 1,2,3 ^ joint first authors 1 Wallace H. Coulter Department of
Gene Prediction Chengwei Luo, Amanda McCook, Nadeem Bulsara, Phillip Lee, Neha Gupta, and Divya Anjan Kumar
 Gene Prediction Chengwei Luo, Amanda McCook, Nadeem Bulsara, Phillip Lee, Neha Gupta, and Divya Anjan Kumar Gene Prediction Introduction Protein-coding gene prediction RNA gene prediction Modification
Gene Prediction Chengwei Luo, Amanda McCook, Nadeem Bulsara, Phillip Lee, Neha Gupta, and Divya Anjan Kumar Gene Prediction Introduction Protein-coding gene prediction RNA gene prediction Modification
Leonardo Mariño-Ramírez, PhD NCBI / NLM / NIH. BIOL 7210 A Computational Genomics 2/18/2015
 Leonardo Mariño-Ramírez, PhD NCBI / NLM / NIH BIOL 7210 A Computational Genomics 2/18/2015 The $1,000 genome is here! http://www.illumina.com/systems/hiseq-x-sequencing-system.ilmn Bioinformatics bottleneck
Leonardo Mariño-Ramírez, PhD NCBI / NLM / NIH BIOL 7210 A Computational Genomics 2/18/2015 The $1,000 genome is here! http://www.illumina.com/systems/hiseq-x-sequencing-system.ilmn Bioinformatics bottleneck
Functional Annotation - Faction 2 Background and Strategy
 Functional Annotation - Faction 2 Background and Strategy March 8, 2017 Khushbu Patel Karan Kapuria Angela Mo Harrison Kim David Lu Christian Colon Nolan English Bowen Yang Cong Gao RECAP. WE ARE HERE!!
Functional Annotation - Faction 2 Background and Strategy March 8, 2017 Khushbu Patel Karan Kapuria Angela Mo Harrison Kim David Lu Christian Colon Nolan English Bowen Yang Cong Gao RECAP. WE ARE HERE!!
ab initio and Evidence-Based Gene Finding
 ab initio and Evidence-Based Gene Finding A basic introduction to annotation Outline What is annotation? ab initio gene finding Genome databases on the web Basics of the UCSC browser Evidence-based gene
ab initio and Evidence-Based Gene Finding A basic introduction to annotation Outline What is annotation? ab initio gene finding Genome databases on the web Basics of the UCSC browser Evidence-based gene
UCSC Genome Browser. Introduction to ab initio and evidence-based gene finding
 UCSC Genome Browser Introduction to ab initio and evidence-based gene finding Wilson Leung 06/2006 Outline Introduction to annotation ab initio gene finding Basics of the UCSC Browser Evidence-based gene
UCSC Genome Browser Introduction to ab initio and evidence-based gene finding Wilson Leung 06/2006 Outline Introduction to annotation ab initio gene finding Basics of the UCSC Browser Evidence-based gene
Gene Prediction Chengwei Luo, Amanda McCook, Nadeem Bulsara, Phillip Lee, Neha Gupta, and Divya Anjan Kumar
 Gene Prediction Chengwei Luo, Amanda McCook, Nadeem Bulsara, Phillip Lee, Neha Gupta, and Divya Anjan Kumar Gene Prediction Introduction Protein-coding gene prediction RNA gene prediction Modification
Gene Prediction Chengwei Luo, Amanda McCook, Nadeem Bulsara, Phillip Lee, Neha Gupta, and Divya Anjan Kumar Gene Prediction Introduction Protein-coding gene prediction RNA gene prediction Modification
RNA Genomics. BME 110: CompBio Tools Todd Lowe May 14, 2010
 RNA Genomics BME 110: CompBio Tools Todd Lowe May 14, 2010 Admin WebCT quiz on Tuesday cover reading, using Jalview & Pfam Homework #3 assigned today due next Friday (8 days) In Genomes, Two Types of Genes
RNA Genomics BME 110: CompBio Tools Todd Lowe May 14, 2010 Admin WebCT quiz on Tuesday cover reading, using Jalview & Pfam Homework #3 assigned today due next Friday (8 days) In Genomes, Two Types of Genes
A Robust Method for Finding the Automated Best Matched Genes Based on Grouping Similar Fragments of Large-Scale References for Genome Assembly
 S S symmetry Article A Robust Method for Finding the Automated Best Matched Genes Based on Grouping Similar Fragments of Large-Scale References for Genome Assembly Jaehee Jung 1, Jong Im Kim 2, Young-Sik
S S symmetry Article A Robust Method for Finding the Automated Best Matched Genes Based on Grouping Similar Fragments of Large-Scale References for Genome Assembly Jaehee Jung 1, Jong Im Kim 2, Young-Sik
Lecture 7 Motif Databases and Gene Finding
 Introduction to Bioinformatics for Medical Research Gideon Greenspan gdg@cs.technion.ac.il Lecture 7 Motif Databases and Gene Finding Motif Databases & Gene Finding Motifs Recap Motif Databases TRANSFAC
Introduction to Bioinformatics for Medical Research Gideon Greenspan gdg@cs.technion.ac.il Lecture 7 Motif Databases and Gene Finding Motif Databases & Gene Finding Motifs Recap Motif Databases TRANSFAC
Efficient and Accurate Analysis of non coding RNAs with InSyBio ncrnaseq
 Efficient and Accurate Analysis of non coding RNAs with InSyBio ncrnaseq WHITE PAPER By InSyBio Ltd Aigli Korfiati Computer Engineer, MSc, PhD candidate InSyBio Product Development Manager October 2015
Efficient and Accurate Analysis of non coding RNAs with InSyBio ncrnaseq WHITE PAPER By InSyBio Ltd Aigli Korfiati Computer Engineer, MSc, PhD candidate InSyBio Product Development Manager October 2015
Improving Start Codon Prediction Accuracy in Prokaryotic Organisms Using Naïve Bayesian Classification
 Improving Start Codon Prediction Accuracy in Prokaryotic Organisms Using Naïve Bayesian Classification Sean Landman and Imad Rahal Computer Science Department College of St. Benedict / St. John s University
Improving Start Codon Prediction Accuracy in Prokaryotic Organisms Using Naïve Bayesian Classification Sean Landman and Imad Rahal Computer Science Department College of St. Benedict / St. John s University
BME 110 Midterm Examination
 BME 110 Midterm Examination May 10, 2011 Name: (please print) Directions: Please circle one answer for each question, unless the question specifies "circle all correct answers". You can use any resource
BME 110 Midterm Examination May 10, 2011 Name: (please print) Directions: Please circle one answer for each question, unless the question specifies "circle all correct answers". You can use any resource
Complete Genome Sequence of Bifidobacterium longum subsp. longum KACC 91563
 JB Accepts, published online ahead of print on 8 July 2011 J. Bacteriol. doi:10.1128/jb.05620-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
JB Accepts, published online ahead of print on 8 July 2011 J. Bacteriol. doi:10.1128/jb.05620-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
Computational analysis of non-coding RNA. Andrew Uzilov BME110 Tue, Nov 16, 2010
 Computational analysis of non-coding RNA Andrew Uzilov auzilov@ucsc.edu BME110 Tue, Nov 16, 2010 1 Corrected/updated talk slides are here: http://tinyurl.com/uzilovrna redirects to: http://users.soe.ucsc.edu/~auzilov/bme110/fall2010/
Computational analysis of non-coding RNA Andrew Uzilov auzilov@ucsc.edu BME110 Tue, Nov 16, 2010 1 Corrected/updated talk slides are here: http://tinyurl.com/uzilovrna redirects to: http://users.soe.ucsc.edu/~auzilov/bme110/fall2010/
DNA RNA PROTEIN. Professor Andrea Garrison Biology 11 Illustrations 2010 Pearson Education, Inc. unless otherwise noted
 DNA RNA PROTEIN Professor Andrea Garrison Biology 11 Illustrations 2010 Pearson Education, Inc. unless otherwise noted DNA Molecule of heredity Contains all the genetic info our cells inherit Determines
DNA RNA PROTEIN Professor Andrea Garrison Biology 11 Illustrations 2010 Pearson Education, Inc. unless otherwise noted DNA Molecule of heredity Contains all the genetic info our cells inherit Determines
Data Retrieval from GenBank
 Data Retrieval from GenBank Peter J. Myler Bioinformatics of Intracellular Pathogens JNU, Feb 7-0, 2009 http://www.ncbi.nlm.nih.gov (January, 2007) http://ncbi.nlm.nih.gov/sitemap/resourceguide.html Accessing
Data Retrieval from GenBank Peter J. Myler Bioinformatics of Intracellular Pathogens JNU, Feb 7-0, 2009 http://www.ncbi.nlm.nih.gov (January, 2007) http://ncbi.nlm.nih.gov/sitemap/resourceguide.html Accessing
Analysis Report. Institution : Macrogen Japan Name : Macrogen Japan Order Number : 1501APB-0004 Sample Name : 8380 Type of Analysis : De novo assembly
 Analysis Report Institution : Macrogen Japan Name : Macrogen Japan Order Number : 1501APB-0004 Sample Name : 8380 Type of Analysis : De novo assembly 1 Table of Contents 1. Result of Whole Genome Assembly
Analysis Report Institution : Macrogen Japan Name : Macrogen Japan Order Number : 1501APB-0004 Sample Name : 8380 Type of Analysis : De novo assembly 1 Table of Contents 1. Result of Whole Genome Assembly
Genome annotation. Erwin Datema (2011) Sandra Smit (2012, 2013)
 Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
A Probabilistic Genome-Wide Gene Reading Frame Sequence Model
 A Probabilistic Genome-Wide Gene Reading Frame Sequence Model Christian Theil Have 1 and Søren Mørk 2 1 Novo Nordisk Foundation Center for Basic Metabolic Resarch, Section of Metabolic Genetics, Copenhagen
A Probabilistic Genome-Wide Gene Reading Frame Sequence Model Christian Theil Have 1 and Søren Mørk 2 1 Novo Nordisk Foundation Center for Basic Metabolic Resarch, Section of Metabolic Genetics, Copenhagen
Computational Genomics Final Presentation. BIOL 7210 Spring 2015
 Computational Genomics Final Presentation BIOL 7210 Spring 2015 Genome Assembly Jillian Walker, Diana Williams, Ke Qi, Xin Wu, Bhanu Gandham, Anuj Gupta, Taylor Griswold, Yuanbo Wang, Sung Im, Maxine Harlemon,
Computational Genomics Final Presentation BIOL 7210 Spring 2015 Genome Assembly Jillian Walker, Diana Williams, Ke Qi, Xin Wu, Bhanu Gandham, Anuj Gupta, Taylor Griswold, Yuanbo Wang, Sung Im, Maxine Harlemon,
Codon usage diversity in city microbiomes
 Codon usage diversity in city microbiomes Haruo Suzuki 1,2 1. Institute for Advanced Biosciences, Keio University, Tsuruoka, Yamagata, Japan 2. Faculty of Environment and Information Studies, Keio University,
Codon usage diversity in city microbiomes Haruo Suzuki 1,2 1. Institute for Advanced Biosciences, Keio University, Tsuruoka, Yamagata, Japan 2. Faculty of Environment and Information Studies, Keio University,
Genome Sequence Assembly
 Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Applied bioinformatics in genomics
 Applied bioinformatics in genomics Productive bioinformatics in a genome sequencing center Heiko Liesegang Warschau 2005 The omics pyramid: 1. 2. 3. 4. 5. Genome sequencing Genome annotation Transcriptomics
Applied bioinformatics in genomics Productive bioinformatics in a genome sequencing center Heiko Liesegang Warschau 2005 The omics pyramid: 1. 2. 3. 4. 5. Genome sequencing Genome annotation Transcriptomics
Module 6 Microbial Genetics. Chapter 8
 Module 6 Microbial Genetics Chapter 8 Structure and function of the genetic material Genetics science of o Study of what genes are, how they determine the characteristics of an organism, how they carry
Module 6 Microbial Genetics Chapter 8 Structure and function of the genetic material Genetics science of o Study of what genes are, how they determine the characteristics of an organism, how they carry
Small Genome Annotation and Data Management at TIGR
 Small Genome Annotation and Data Management at TIGR Michelle Gwinn, William Nelson, Robert Dodson, Steven Salzberg, Owen White Abstract TIGR has developed, and continues to refine, a comprehensive, efficient
Small Genome Annotation and Data Management at TIGR Michelle Gwinn, William Nelson, Robert Dodson, Steven Salzberg, Owen White Abstract TIGR has developed, and continues to refine, a comprehensive, efficient
RNA Genomics II. BME 110: CompBio Tools Todd Lowe & Andrew Uzilov May 17, 2011
 RNA Genomics II BME 110: CompBio Tools Todd Lowe & Andrew Uzilov May 17, 2011 1 TIME Why RNA? An evolutionary perspective The RNA World hypotheses: life arose as self-replicating non-coding RNA (ncrna)
RNA Genomics II BME 110: CompBio Tools Todd Lowe & Andrew Uzilov May 17, 2011 1 TIME Why RNA? An evolutionary perspective The RNA World hypotheses: life arose as self-replicating non-coding RNA (ncrna)
Protein Synthesis: From Gene RNA Protein Trait
 Protein Synthesis: From Gene RNA Protein Trait Human Genome The human genome contains about genes. Each gene is a of DNA (sequence of nitrogen bases) contained within each chromosome. Each chromosome contains
Protein Synthesis: From Gene RNA Protein Trait Human Genome The human genome contains about genes. Each gene is a of DNA (sequence of nitrogen bases) contained within each chromosome. Each chromosome contains
Lecture 10. Ab initio gene finding
 Lecture 10 Ab initio gene finding Uses of probabilistic sequence Segmentation models/hmms Multiple alignment using profile HMMs Prediction of sequence function (gene family models) ** Gene finding ** Review
Lecture 10 Ab initio gene finding Uses of probabilistic sequence Segmentation models/hmms Multiple alignment using profile HMMs Prediction of sequence function (gene family models) ** Gene finding ** Review
GenBank Growth. In 2003 ~ 31 million sequences ~ 37 billion base pairs
 Gene Finding GenBank Growth GenBank Growth In 2003 ~ 31 million sequences ~ 37 billion base pairs GenBank: Exponential Growth Growth of GenBank in billions of base pairs from release 3 in April of 1994
Gene Finding GenBank Growth GenBank Growth In 2003 ~ 31 million sequences ~ 37 billion base pairs GenBank: Exponential Growth Growth of GenBank in billions of base pairs from release 3 in April of 1994
Collect, analyze and synthesize. Annotation. Annotation for D. virilis. GEP goals: Evidence Based Annotation. Evidence for Gene Models 12/26/2018
 Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
Big Idea 3C Basic Review
 Big Idea 3C Basic Review 1. A gene is a. A sequence of DNA that codes for a protein. b. A sequence of amino acids that codes for a protein. c. A sequence of codons that code for nucleic acids. d. The end
Big Idea 3C Basic Review 1. A gene is a. A sequence of DNA that codes for a protein. b. A sequence of amino acids that codes for a protein. c. A sequence of codons that code for nucleic acids. d. The end
Probiotic Strain Isolated from the Vagina of Healthy Women
 JB Accepts, published online ahead of print on 1 April 2011 J. Bacteriol. doi:10.1128/jb.00358-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
JB Accepts, published online ahead of print on 1 April 2011 J. Bacteriol. doi:10.1128/jb.00358-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
BIO4342 Lab Exercise: Detecting and Interpreting Genetic Homology
 BIO4342 Lab Exercise: Detecting and Interpreting Genetic Homology Jeremy Buhler March 15, 2004 In this lab, we ll annotate an interesting piece of the D. melanogaster genome. Along the way, you ll get
BIO4342 Lab Exercise: Detecting and Interpreting Genetic Homology Jeremy Buhler March 15, 2004 In this lab, we ll annotate an interesting piece of the D. melanogaster genome. Along the way, you ll get
Tutorial for Stop codon reassignment in the wild
 Tutorial for Stop codon reassignment in the wild Learning Objectives This tutorial has two learning objectives: 1. Finding evidence of stop codon reassignment on DNA fragments. 2. Detecting and confirming
Tutorial for Stop codon reassignment in the wild Learning Objectives This tutorial has two learning objectives: 1. Finding evidence of stop codon reassignment on DNA fragments. 2. Detecting and confirming
COMPUTER RESOURCES II:
 COMPUTER RESOURCES II: Using the computer to analyze data, using the internet, and accessing online databases Bio 210, Fall 2006 Linda S. Huang, Ph.D. University of Massachusetts Boston In the first computer
COMPUTER RESOURCES II: Using the computer to analyze data, using the internet, and accessing online databases Bio 210, Fall 2006 Linda S. Huang, Ph.D. University of Massachusetts Boston In the first computer
Designing Filters for Fast Protein and RNA Annotation. Yanni Sun Dept. of Computer Science and Engineering Advisor: Jeremy Buhler
 Designing Filters for Fast Protein and RNA Annotation Yanni Sun Dept. of Computer Science and Engineering Advisor: Jeremy Buhler 1 Outline Background on sequence annotation Protein annotation acceleration
Designing Filters for Fast Protein and RNA Annotation Yanni Sun Dept. of Computer Science and Engineering Advisor: Jeremy Buhler 1 Outline Background on sequence annotation Protein annotation acceleration
G4120: Introduction to Computational Biology
 ICB Fall 2009 G4120: Computational Biology Oliver Jovanovic, Ph.D. Columbia University Department of Microbiology & Immunology Copyright 2009 Oliver Jovanovic, All Rights Reserved. Analysis of Protein
ICB Fall 2009 G4120: Computational Biology Oliver Jovanovic, Ph.D. Columbia University Department of Microbiology & Immunology Copyright 2009 Oliver Jovanovic, All Rights Reserved. Analysis of Protein
Transcriptome Assembly, Functional Annotation (and a few other related thoughts)
 Transcriptome Assembly, Functional Annotation (and a few other related thoughts) Monica Britton, Ph.D. Sr. Bioinformatics Analyst June 23, 2017 Differential Gene Expression Generalized Workflow File Types
Transcriptome Assembly, Functional Annotation (and a few other related thoughts) Monica Britton, Ph.D. Sr. Bioinformatics Analyst June 23, 2017 Differential Gene Expression Generalized Workflow File Types
Genome Annotation. Stefan Prost 1. May 27th, States of America. Genome Annotation
 Genome Annotation Stefan Prost 1 1 Department of Integrative Biology, University of California, Berkeley, United States of America May 27th, 2015 Outline Genome Annotation 1 Repeat Annotation 2 Repeat
Genome Annotation Stefan Prost 1 1 Department of Integrative Biology, University of California, Berkeley, United States of America May 27th, 2015 Outline Genome Annotation 1 Repeat Annotation 2 Repeat
Complete genome sequence of Clostridium acetobutylicum. DSM 1731, a solvent producing strain with multi-replicon
 JB Accepts, published online ahead of print on 8 July 2011 J. Bacteriol. doi:10.1128/jb.05596-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
JB Accepts, published online ahead of print on 8 July 2011 J. Bacteriol. doi:10.1128/jb.05596-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
RNA folding & ncrna discovery
 I519 Introduction to Bioinformatics RNA folding & ncrna discovery Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Non-coding RNAs and their functions RNA structures RNA folding
I519 Introduction to Bioinformatics RNA folding & ncrna discovery Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Non-coding RNAs and their functions RNA structures RNA folding
Collect, analyze and synthesize. Annotation. Annotation for D. virilis. Evidence Based Annotation. GEP goals: Evidence for Gene Models 08/22/2017
 Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
Annotation Annotation for D. virilis Chris Shaffer July 2012 l Big Picture of annotation and then one practical example l This technique may not be the best with other projects (e.g. corn, bacteria) l
Chimp Sequence Annotation: Region 2_3
 Chimp Sequence Annotation: Region 2_3 Jeff Howenstein March 30, 2007 BIO434W Genomics 1 Introduction We received region 2_3 of the ChimpChunk sequence, and the first step we performed was to run RepeatMasker
Chimp Sequence Annotation: Region 2_3 Jeff Howenstein March 30, 2007 BIO434W Genomics 1 Introduction We received region 2_3 of the ChimpChunk sequence, and the first step we performed was to run RepeatMasker
Introduction to Bioinformatics CPSC 265. What is bioinformatics? Textbooks
 Introduction to Bioinformatics CPSC 265 Thanks to Jonathan Pevsner, Ph.D. Textbooks Johnathan Pevsner, who I stole most of these slides from (thanks!) has written a textbook, Bioinformatics and Functional
Introduction to Bioinformatics CPSC 265 Thanks to Jonathan Pevsner, Ph.D. Textbooks Johnathan Pevsner, who I stole most of these slides from (thanks!) has written a textbook, Bioinformatics and Functional
Computational gene finding
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Gene Expression. Student:
 Gene Expression Student: 1. A ribozyme is A. a section of the DNA that is expressed in the mrna. B. a self-splicing intron that acts like an enzyme. C. a complex made up of many ribosomes replicating the
Gene Expression Student: 1. A ribozyme is A. a section of the DNA that is expressed in the mrna. B. a self-splicing intron that acts like an enzyme. C. a complex made up of many ribosomes replicating the
Practical Bioinformatics for Life Scientists. Week 14, Lecture 27. István Albert Bioinformatics Consulting Center Penn State
 Practical Bioinformatics for Life Scientists Week 14, Lecture 27 István Albert Bioinformatics Consulting Center Penn State No homework this week Project to be given out next Thursday (Dec 1 st ) Due following
Practical Bioinformatics for Life Scientists Week 14, Lecture 27 István Albert Bioinformatics Consulting Center Penn State No homework this week Project to be given out next Thursday (Dec 1 st ) Due following
Genome Annotation - 2. Qi Sun Bioinformatics Facility Cornell University
 Genome Annotation - 2 Qi Sun Bioinformatics Facility Cornell University Output from Maker GFF file: Annotated gene, transcripts, and CDS FASTA file: Predicted transcript sequences Predicted protein sequences
Genome Annotation - 2 Qi Sun Bioinformatics Facility Cornell University Output from Maker GFF file: Annotated gene, transcripts, and CDS FASTA file: Predicted transcript sequences Predicted protein sequences
Unit 3c. Microbial Gene0cs
 Unit 3c Microbial Gene0cs Microbial Genetics! Gene0cs: the science of heredity Genome: the gene0c informa0on in the cell Genomics: the sequencing and molecular characteriza0on of genomes Gregor Mendel
Unit 3c Microbial Gene0cs Microbial Genetics! Gene0cs: the science of heredity Genome: the gene0c informa0on in the cell Genomics: the sequencing and molecular characteriza0on of genomes Gregor Mendel
MAKER: An easy to use genome annotation pipeline. Carson Holt Yandell Lab Department of Human Genetics University of Utah
 MAKER: An easy to use genome annotation pipeline Carson Holt Yandell Lab Department of Human Genetics University of Utah Introduction to Genome Annotation What annotations are Importance of genome annotations
MAKER: An easy to use genome annotation pipeline Carson Holt Yandell Lab Department of Human Genetics University of Utah Introduction to Genome Annotation What annotations are Importance of genome annotations
Gene Identification in silico
 Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
G4120: Introduction to Computational Biology
 ICB Fall 2004 G4120: Computational Biology Oliver Jovanovic, Ph.D. Columbia University Department of Microbiology Copyright 2004 Oliver Jovanovic, All Rights Reserved. Analysis of Protein Sequences Coding
ICB Fall 2004 G4120: Computational Biology Oliver Jovanovic, Ph.D. Columbia University Department of Microbiology Copyright 2004 Oliver Jovanovic, All Rights Reserved. Analysis of Protein Sequences Coding
Protein Sequence Analysis. BME 110: CompBio Tools Todd Lowe April 19, 2007 (Slide Presentation: Carol Rohl)
 Protein Sequence Analysis BME 110: CompBio Tools Todd Lowe April 19, 2007 (Slide Presentation: Carol Rohl) Linear Sequence Analysis What can you learn from a (single) protein sequence? Calculate it s physical
Protein Sequence Analysis BME 110: CompBio Tools Todd Lowe April 19, 2007 (Slide Presentation: Carol Rohl) Linear Sequence Analysis What can you learn from a (single) protein sequence? Calculate it s physical
Annotation Practice Activity [Based on materials from the GEP Summer 2010 Workshop] Special thanks to Chris Shaffer for document review Parts A-G
![Annotation Practice Activity [Based on materials from the GEP Summer 2010 Workshop] Special thanks to Chris Shaffer for document review Parts A-G Annotation Practice Activity [Based on materials from the GEP Summer 2010 Workshop] Special thanks to Chris Shaffer for document review Parts A-G](/thumbs/95/123686269.jpg) Annotation Practice Activity [Based on materials from the GEP Summer 2010 Workshop] Special thanks to Chris Shaffer for document review Parts A-G Introduction: A genome is the total genetic content of
Annotation Practice Activity [Based on materials from the GEP Summer 2010 Workshop] Special thanks to Chris Shaffer for document review Parts A-G Introduction: A genome is the total genetic content of
Bioinformatics: Sequence Analysis. COMP 571 Luay Nakhleh, Rice University
 Bioinformatics: Sequence Analysis COMP 571 Luay Nakhleh, Rice University Course Information Instructor: Luay Nakhleh (nakhleh@rice.edu); office hours by appointment (office: DH 3119) TA: Leo Elworth (DH
Bioinformatics: Sequence Analysis COMP 571 Luay Nakhleh, Rice University Course Information Instructor: Luay Nakhleh (nakhleh@rice.edu); office hours by appointment (office: DH 3119) TA: Leo Elworth (DH
From Infection to Genbank
 From Infection to Genbank How a pathogenic bacterium gets its genome to NCBI Torsten Seemann VLSCI - Life Sciences Computation Centre - Genomics Theme - Lab Meeting - Friday 27 April 2012 The steps 1.
From Infection to Genbank How a pathogenic bacterium gets its genome to NCBI Torsten Seemann VLSCI - Life Sciences Computation Centre - Genomics Theme - Lab Meeting - Friday 27 April 2012 The steps 1.
Outline. Gene Finding Questions. Recap: Prokaryotic gene finding Eukaryotic gene finding The human gene complement Regulation
 Tues, Nov 29: Gene Finding 1 Online FCE s: Thru Dec 12 Thurs, Dec 1: Gene Finding 2 Tues, Dec 6: PS5 due Project presentations 1 (see course web site for schedule) Thurs, Dec 8 Final papers due Project
Tues, Nov 29: Gene Finding 1 Online FCE s: Thru Dec 12 Thurs, Dec 1: Gene Finding 2 Tues, Dec 6: PS5 due Project presentations 1 (see course web site for schedule) Thurs, Dec 8 Final papers due Project
Proteogenomics Workflow for Neoantigen Discovery
 Proteogenomics Workflow for Neoantigen Discovery XIE Lu, Shanghai Center for Bioinformation Technology August 29-31, 2018. The 16th KJC Bioinformatics Symposium, Hayama, Japan Proteogenomics 目录 CONTENTS
Proteogenomics Workflow for Neoantigen Discovery XIE Lu, Shanghai Center for Bioinformation Technology August 29-31, 2018. The 16th KJC Bioinformatics Symposium, Hayama, Japan Proteogenomics 目录 CONTENTS
Challenging algorithms in bioinformatics
 Challenging algorithms in bioinformatics 11 October 2018 Torbjørn Rognes Department of Informatics, UiO torognes@ifi.uio.no What is bioinformatics? Definition: Bioinformatics is the development and use
Challenging algorithms in bioinformatics 11 October 2018 Torbjørn Rognes Department of Informatics, UiO torognes@ifi.uio.no What is bioinformatics? Definition: Bioinformatics is the development and use
ELE4120 Bioinformatics. Tutorial 5
 ELE4120 Bioinformatics Tutorial 5 1 1. Database Content GenBank RefSeq TPA UniProt 2. Database Searches 2 Databases A common situation for alignment is to search through a database to retrieve the similar
ELE4120 Bioinformatics Tutorial 5 1 1. Database Content GenBank RefSeq TPA UniProt 2. Database Searches 2 Databases A common situation for alignment is to search through a database to retrieve the similar
Functional Annotation: Preliminary Results
 Functional Annotation: Preliminary Results Vani Rajan Gena Tang Neha Varghese Kevin Lee Gabriel Mitchell Tripp Jones Robert Petit Shaupu Qin Outline Motivation Naming scheme Preliminary Program Results
Functional Annotation: Preliminary Results Vani Rajan Gena Tang Neha Varghese Kevin Lee Gabriel Mitchell Tripp Jones Robert Petit Shaupu Qin Outline Motivation Naming scheme Preliminary Program Results
Gene Prediction 10/21/05
 Gene Prediction 1/21/5 1/21/5 Gene Prediction Announcements Eam 2 - net Friday Posted online: Eam 2 Study Guide 544 Reading Assignment (2 papers) (formerly Gene Prediction - ) 1/21/5 D Dobbs ISU - BCB
Gene Prediction 1/21/5 1/21/5 Gene Prediction Announcements Eam 2 - net Friday Posted online: Eam 2 Study Guide 544 Reading Assignment (2 papers) (formerly Gene Prediction - ) 1/21/5 D Dobbs ISU - BCB
Annotation of contig27 in the Muller F Element of D. elegans. Contig27 is a 60,000 bp region located in the Muller F element of the D. elegans.
 David Wang Bio 434W 4/27/15 Annotation of contig27 in the Muller F Element of D. elegans Abstract Contig27 is a 60,000 bp region located in the Muller F element of the D. elegans. Genscan predicted six
David Wang Bio 434W 4/27/15 Annotation of contig27 in the Muller F Element of D. elegans Abstract Contig27 is a 60,000 bp region located in the Muller F element of the D. elegans. Genscan predicted six
Genes and gene finding
 Genes and gene finding Ben Langmead Department of Computer Science You are free to use these slides. If you do, please sign the guestbook (www.langmead-lab.org/teaching-materials), or email me (ben.langmead@gmail.com)
Genes and gene finding Ben Langmead Department of Computer Science You are free to use these slides. If you do, please sign the guestbook (www.langmead-lab.org/teaching-materials), or email me (ben.langmead@gmail.com)
Computational gene finding
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Community-assisted genome annotation: The Pseudomonas example. Geoff Winsor, Simon Fraser University Burnaby (greater Vancouver), Canada
 Community-assisted genome annotation: The Pseudomonas example Geoff Winsor, Simon Fraser University Burnaby (greater Vancouver), Canada Overview Pseudomonas Community Annotation Project (PseudoCAP) Past
Community-assisted genome annotation: The Pseudomonas example Geoff Winsor, Simon Fraser University Burnaby (greater Vancouver), Canada Overview Pseudomonas Community Annotation Project (PseudoCAP) Past
Introduction to Cellular Biology and Bioinformatics. Farzaneh Salari
 Introduction to Cellular Biology and Bioinformatics Farzaneh Salari Outline Bioinformatics Cellular Biology A Bioinformatics Problem What is bioinformatics? Computer Science Statistics Bioinformatics Mathematics...
Introduction to Cellular Biology and Bioinformatics Farzaneh Salari Outline Bioinformatics Cellular Biology A Bioinformatics Problem What is bioinformatics? Computer Science Statistics Bioinformatics Mathematics...
Annotating the Genome (H)
 Annotating the Genome (H) Annotation principles (H1) What is annotation? In general: annotation = explanatory note* What could be useful as an annotation of a DNA sequence? an amino acid sequence? What
Annotating the Genome (H) Annotation principles (H1) What is annotation? In general: annotation = explanatory note* What could be useful as an annotation of a DNA sequence? an amino acid sequence? What
Videos. Lesson Overview. Fermentation
 Lesson Overview Fermentation Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast
Lesson Overview Fermentation Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast
CSE 549: RNA-Seq aided gene finding
 CSE 549: RNA-Seq aided gene finding Finding Genes We ll break gene finding methods into 3 main categories. ab initio latin from the beginning w/o experimental evidence comparative make use of knowledge
CSE 549: RNA-Seq aided gene finding Finding Genes We ll break gene finding methods into 3 main categories. ab initio latin from the beginning w/o experimental evidence comparative make use of knowledge
Outline. Introduction to ab initio and evidence-based gene finding. Prokaryotic gene predictions
 Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
Computational gene finding. Devika Subramanian Comp 470
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) The biological context Lec 1 Lec 2 Lec 3 Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) The biological context Lec 1 Lec 2 Lec 3 Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Basic Bioinformatics: Homology, Sequence Alignment,
 Basic Bioinformatics: Homology, Sequence Alignment, and BLAST William S. Sanders Institute for Genomics, Biocomputing, and Biotechnology (IGBB) High Performance Computing Collaboratory (HPC 2 ) Mississippi
Basic Bioinformatics: Homology, Sequence Alignment, and BLAST William S. Sanders Institute for Genomics, Biocomputing, and Biotechnology (IGBB) High Performance Computing Collaboratory (HPC 2 ) Mississippi
An Investigation of Palindromic Sequences in the Pseudomonas fluorescens SBW25 Genome Bachelor of Science Honors Thesis
 An Investigation of Palindromic Sequences in the Pseudomonas fluorescens SBW25 Genome Bachelor of Science Honors Thesis Lina L. Faller Department of Computer Science University of New Hampshire June 2008
An Investigation of Palindromic Sequences in the Pseudomonas fluorescens SBW25 Genome Bachelor of Science Honors Thesis Lina L. Faller Department of Computer Science University of New Hampshire June 2008
Lesson Overview. Fermentation 13.1 RNA
 13.1 RNA The Role of RNA Genes contain coded DNA instructions that tell cells how to build proteins. The first step in decoding these genetic instructions is to copy part of the base sequence from DNA
13.1 RNA The Role of RNA Genes contain coded DNA instructions that tell cells how to build proteins. The first step in decoding these genetic instructions is to copy part of the base sequence from DNA
Algorithms in Bioinformatics
 Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline Central Dogma of Molecular
Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline Central Dogma of Molecular
RNA-Seq with the Tuxedo Suite
 RNA-Seq with the Tuxedo Suite Monica Britton, Ph.D. Sr. Bioinformatics Analyst September 2015 Workshop The Basic Tuxedo Suite References Trapnell C, et al. 2009 TopHat: discovering splice junctions with
RNA-Seq with the Tuxedo Suite Monica Britton, Ph.D. Sr. Bioinformatics Analyst September 2015 Workshop The Basic Tuxedo Suite References Trapnell C, et al. 2009 TopHat: discovering splice junctions with
From assembled genome to annotated genome
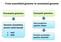 From assembled genome to annotated genome Procaryotic genomes Eucaryotic genomes Genome annotation servers (web based) 1. RAST 2. NCBI Gene prediction pipeline: Maker Function annotation pipeline: Blast2GO
From assembled genome to annotated genome Procaryotic genomes Eucaryotic genomes Genome annotation servers (web based) 1. RAST 2. NCBI Gene prediction pipeline: Maker Function annotation pipeline: Blast2GO
Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype)
 Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype) Question#1: One-Gene, One-Polypeptide The figure below shows the results of feeding trials with one auxotroph strain of Neurospora
Self-test Quiz for Chapter 12 (From DNA to Protein: Genotype to Phenotype) Question#1: One-Gene, One-Polypeptide The figure below shows the results of feeding trials with one auxotroph strain of Neurospora
Introduction to 'Omics and Bioinformatics
 Introduction to 'Omics and Bioinformatics Chris Overall Department of Bioinformatics and Genomics University of North Carolina Charlotte Acquire Store Analyze Visualize Bioinformatics makes many current
Introduction to 'Omics and Bioinformatics Chris Overall Department of Bioinformatics and Genomics University of North Carolina Charlotte Acquire Store Analyze Visualize Bioinformatics makes many current
MCB 102 University of California, Berkeley August 11 13, Problem Set 8
 MCB 102 University of California, Berkeley August 11 13, 2009 Isabelle Philipp Handout Problem Set 8 The answer key will be posted by Tuesday August 11. Try to solve the problem sets always first without
MCB 102 University of California, Berkeley August 11 13, 2009 Isabelle Philipp Handout Problem Set 8 The answer key will be posted by Tuesday August 11. Try to solve the problem sets always first without
Prediction of noncoding RNAs with RNAz
 Prediction of noncoding RNAs with RNAz John Dzmil, III Steve Griesmer Philip Murillo April 4, 2007 What is non-coding RNA (ncrna)? RNA molecules that are not translated into proteins Size range from 20
Prediction of noncoding RNAs with RNAz John Dzmil, III Steve Griesmer Philip Murillo April 4, 2007 What is non-coding RNA (ncrna)? RNA molecules that are not translated into proteins Size range from 20
Gene-centered resources at NCBI
 COURSE OF BIOINFORMATICS a.a. 2014-2015 Gene-centered resources at NCBI We searched Accession Number: M60495 AT NCBI Nucleotide Gene has been implemented at NCBI to organize information about genes, serving
COURSE OF BIOINFORMATICS a.a. 2014-2015 Gene-centered resources at NCBI We searched Accession Number: M60495 AT NCBI Nucleotide Gene has been implemented at NCBI to organize information about genes, serving
ABSTRACT METHODS FOR MICROBIAL GENOMICS. Professor Steven L. Salzberg Department of Computer Science
 ABSTRACT Title of Dissertation: COMPARATIVE AND COMPUTATIONAL METHODS FOR MICROBIAL GENOMICS Derrick Edward Wood, Doctor of Philosophy, 2014 Directed by: Professor Steven L. Salzberg Department of Computer
ABSTRACT Title of Dissertation: COMPARATIVE AND COMPUTATIONAL METHODS FOR MICROBIAL GENOMICS Derrick Edward Wood, Doctor of Philosophy, 2014 Directed by: Professor Steven L. Salzberg Department of Computer
Videos. Bozeman Transcription and Translation: Drawing transcription and translation:
 Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast RNA and DNA. 29b) I can explain
Videos Bozeman Transcription and Translation: https://youtu.be/h3b9arupxzg Drawing transcription and translation: https://youtu.be/6yqplgnjr4q Objectives 29a) I can contrast RNA and DNA. 29b) I can explain
FUNCTIONAL ANNOTATION PRELIMINARY RESULTS. Compgenomics 2010
 FUNCTIONAL ANNOTATION PRELIMINARY RESULTS Compgenomics 2010 E FIRST LEVEL OF ANNOTION F Pathways OPERONS BLASTP BLASTP OPERON_DB KEGG DOORS CONSENSUS SCRIPT SECOD LEVEL OF ANNOTION Operon prediction Introduction
FUNCTIONAL ANNOTATION PRELIMINARY RESULTS Compgenomics 2010 E FIRST LEVEL OF ANNOTION F Pathways OPERONS BLASTP BLASTP OPERON_DB KEGG DOORS CONSENSUS SCRIPT SECOD LEVEL OF ANNOTION Operon prediction Introduction
7.014 Problem Set 4 Answers to this problem set are to be turned in. Problem sets will not be accepted late. Solutions will be posted on the web.
 MIT Department of Biology 7.014 Introductory Biology, Spring 2005 Name: Section : 7.014 Problem Set 4 Answers to this problem set are to be turned in. Problem sets will not be accepted late. Solutions
MIT Department of Biology 7.014 Introductory Biology, Spring 2005 Name: Section : 7.014 Problem Set 4 Answers to this problem set are to be turned in. Problem sets will not be accepted late. Solutions
Sequence Based Function Annotation
 Sequence Based Function Annotation Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Sequence Based Function Annotation 1. Given a sequence, how to predict its biological
Sequence Based Function Annotation Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Sequence Based Function Annotation 1. Given a sequence, how to predict its biological
Two Mark question and Answers
 1. Define Bioinformatics Two Mark question and Answers Bioinformatics is the field of science in which biology, computer science, and information technology merge into a single discipline. There are three
1. Define Bioinformatics Two Mark question and Answers Bioinformatics is the field of science in which biology, computer science, and information technology merge into a single discipline. There are three
Bundle 5 Test Review
 Bundle 5 Test Review DNA vs. RNA DNA Replication Gene Mutations- Protein Synthesis 1. Label the different components and complete the complimentary base pairing. What is this molecule called? _Nucleic
Bundle 5 Test Review DNA vs. RNA DNA Replication Gene Mutations- Protein Synthesis 1. Label the different components and complete the complimentary base pairing. What is this molecule called? _Nucleic
