Four different active promoter genes were chosen, ATXN7L2, PSRC1, CELSR2 and
|
|
|
- Norman Charles
- 5 years ago
- Views:
Transcription
1 SUPPLEMENTARY MATERIALS AND METHODS Chromatin Immunoprecipitation for qpcr analysis Four different active promoter genes were chosen, ATXN7L2, PSRC1, CELSR2 and IL24, all located on chromosome 1. Primer sequences are shown in the table below. Gene name Sequence forward primer Sequence reverse primer ATXN7L2 CCTCCCTGCACCTACCCTAT TCCACTCTCCCAAGCTCACT PSRC1 ACCGTGATTGGTGAATCCTC CTCTGCTCCCATCTGCCTAC CELSR2 TGCAACTGTGATGTCAGCAA CACAAGGCCCTCAGCTCTAC IL24 AGCCAGTGGTAGAGCCTGAA AGGAAAAAGAGGGAGGTGGA Non-transcribed region CCCATCTCAACCTCCACACT CTTGTCCAGATTCGCTGTGA Chromatin Immunoprecipitation Chromatin was prepared as described in Fousteri et al. (Fousteri et al, 2006) from untreated XPCS2_SV40 cells or 30 min after global UVC irradiation (16 J/m 2 ). Sonication was performed on ice in a buffer containing 10 mm Tris-HCl [ph 8.0], 140 mm Nacl, 1% Triton X- 100, 0.1% Na-deoxycholate, 0.1% SDS (buffer S). The sonicator was a Diagenode (Liège, Belgium) Bioruptor set at power 5 and cycles were: 10 s pulses followed by 20 s of cooling during 15 min and repeated 3 times. After spinning down (13200 rpm, 15 min) the samples, the supernatant containing the crosslinked chromatin was stored at -80 C. In each ChIP assay, 400 g of protein from crosslinked chromatin was used to immunoprecipitate with 1 g of antibody rabbit IgG polyclonal anti-xpb (S-19) in buffer S. Immunoprecipitation lasted was performed at 4 C for 2 hr. The immunocomplexes were collected by adsorption to protein G Sepharose beads (Upstate-Millipore) overnight at 4 C. The G Sepharose beads were washed with 5 vol of buffer S and resuspended in 1x Laemli SDS buffer. To decrosslink the proteins prior electrophoresis, samples were incubated at 95 C for 90 min. ChIP complexes were resolved by SDS-PAGE and 1
2 western-blotted with the following NER antibodies: IgG polyclonal anti-xpb (S-19), mouse IgG monoclonal anti-p62 (G-10), goat IgG polyclonal anti-p34 (C-19), rabbit IgG polyclonal anti- XPA (FL-273, Santa Cruz Biotechnology), mouse IgG monoclonal anti-xpf (Ab-1, clone 219, Thermoscientific). Measurements of transcription activity after inhibition by -amanitin Cells (MRC5SV, XPCS2SV, XP12ROSV, XP6BESV, XPCS1ROSV) were grown on 24 mm coverslips and treated 6h or 16h with -amanitin with concentrations ranging from 0 to 25 µg/ml at 37 C. RNA production was detected using the Click-iT RNA Alexa Fluor Imaging kit (Invitrogen). Briefly, cells were incubated for 2 hours with 5-ethynyl uridine, fixed with paraformaldehyde 4% and permeabilized with Triton 0.5%. After this treatment, cells were incubated for 30 min with the Click-iT reaction cocktail containing Alexa Fluor Azide 488. After washing, coverslips were mounted on slides with Vectashield (Vector). Imaging was performed on a Zeiss LSM 710 confocal laser scanning microscope (Zeiss), using a 40x/1.3 objective. SUPPLEMENTARY REFERENCES Fousteri M, Vermeulen W, van Zeeland AA, Mullenders LH (2006) Cockayne syndrome A and B proteins differentially regulate recruitment of chromatin remodeling and repair factors to stalled RNA polymerase II in vivo. Mol Cell 23:
3 SUPPLEMENTARY FIGURE LEGENDS Figure S1: Recruitment of the DNA damage response machinery is not pan-nuclear in XP- D/CS cells. (A) Local H2AX phosphorylation in XPCS2 (XP-D/CS) cells after UV exposure. XP-D/CS cells were treated with 60 J/m 2 local UV-irradiation. 45 minutes later the cells were fixed and immunostained with antibodies directed against H2AX (Ser139) (in red) and the XPB protein (in green). (B) Local H2A ubiquitination (in red) and the XPB protein (in green) in XP-D/CS cells 45 minutes after local UV-irradiation. All scale bars, 5 m. Figure S2: Effect of UVC-irradiation on TFIIH binding at promoters in WT and XP-D/CS cells Bar graphs of the qpcr data from the Chromatin Immunoprecipitation (ChIP) experiments on the promoters of 4 different expressed genes in both MRC5 (WT) and XPCS2 (XP-D/CS) cells, prior to (-UV) and 1/2h after exposure to 20 J/m 2 of UVC (+UV). The Y-axis represents the fold enrichment of TFIIH binding to the promoter region of each active gene. Figure S3: Transcription inhibition by alpha-amanitin treatment Confocal images of WT, XP-D/CS, XP-G/CS, XP-A and XP-D cells treated with different concentrations (from 5 µg/ml to 20 µg/ml) of -amanitin for 6h. RNA production is stained in green using the EU Click it kit. All scale bars, 25 m. 3
4 Figure S4: UVC-irradiation does not induce DNA breaks in XP-D/CS cells at early time points. (A) Confocal images of XPB (red), Ku80 (green) and Poly(ADP-ribose) (green) immunostaining in XP-D/CS cells fixed 15 min (top row) and 1 hour (bottom row) after local UV-irradiation (60 J/m 2 ). (B) Confocal images of XPB (red), Ku80 (green) and Poly(ADP-ribose) (green) immunostaining in XP-D/CS cells fixed 15 min after laser induced (multi-photon at 800nm) DNA damage. All scale bars, 5 m. Figure S5: H2AX phosphorylation in primary cells after UV-irradiation. (A) Immunostaining with DAPI (cyan) and Ki67 (green) in confluent XP-D/CS human primary cells. Less than 10% of the cells were Ki67 positive (counting 200 nuclei). Scale bars, 25 m. (B) Confocal images of H2AX (green) and XPB (red) immunofluorescence in confluent primary human cells (C5RO, XPCS2, XPCS1RO and XP12RO) 1h after local UV-irradiation (60 J/m 2 ). Scale bars, 5 m. Figure S6: RPA accumulation in UVC-exposed areas in primary cells and XPF recruitment into repair complexes (A) Confocal images of RPA (green) and XPB (red) immunofluorescence in confluent primary human cells (C5RO, XPCS2 and XPCS1RO) 1h after local UV-irradiation (60 J/m 2 ). Scale bars, 5 m. (B) Western-blot analysis of XPB-specific ChIP using chromatin of crosslinked UVirradiated (lane 2) and untreated (lane 1) TFIIH XP/CS cells. ChIP complexes were resolved by SDS-PAGE and western-blotted with NER antibodies (indicated by arrows). The thick-bands (indicated by *) are heavy and light chain IgG bands, intrinsic to the IP. 4
5 Figure S7: ssdna labeling after high local UV-irradiation. Confocal images of XPB (red) and BrdU (green) immunofluorescence in MRC5 (WT) and XPCS2 (XP-D/CS) cells 1h after local UV-irradiation (100 J/m 2 ). Scale bars, 5 m. Figure S8: XPF depletion 48 h after sirna -XPF treatment. Confocal images of XPB (red) and XPF (green) immunofluorescence in XP-D/CS cells, 1h after UV local exposure with 60 J/m 2. Cells were transfected with 40 nm sirna -XPF, or not (Mock), 48 h prior the experiment. Scale bars, 20 m. Figure S9: Fen1 accumulation on UV-lesion sites only in XP/CS cells. Confocal images of Fen1 (red) and RPA (green) immunostained cells (WT, XP-D/CS, XP-G/CS and XP-A) 1 h after local UV-irradiation (60 J/m 2 ). Scale bars, 5 m. 5
6 SUPPLEMENTARY FIGURES 6
7 7
8 8
9 9
10 10
11 11
12 12
13 13
14 14
Supplementary Material
 Supplementary Material Supplementary Methods Cell synchronization. For synchronized cell growth, thymidine was added to 30% confluent U2OS cells to a final concentration of 2.5mM. Cells were incubated
Supplementary Material Supplementary Methods Cell synchronization. For synchronized cell growth, thymidine was added to 30% confluent U2OS cells to a final concentration of 2.5mM. Cells were incubated
Supplementary Figure Legends
 Supplementary Figure Legends Figure S1 gene targeting strategy for disruption of chicken gene, related to Figure 1 (f)-(i). (a) The locus and the targeting constructs showing HpaI restriction sites. The
Supplementary Figure Legends Figure S1 gene targeting strategy for disruption of chicken gene, related to Figure 1 (f)-(i). (a) The locus and the targeting constructs showing HpaI restriction sites. The
Immunofluorescence images of different core histones and different histone exchange assay.
 Molecular Cell, Volume 51 Supplemental Information Enhanced Chromatin Dynamics by FACT Promotes Transcriptional Restart after UV-Induced DNA Damage Christoffel Dinant, Giannis Ampatziadis-Michailidis,
Molecular Cell, Volume 51 Supplemental Information Enhanced Chromatin Dynamics by FACT Promotes Transcriptional Restart after UV-Induced DNA Damage Christoffel Dinant, Giannis Ampatziadis-Michailidis,
RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the
 Supplementary Methods RT-PCR and real-time PCR analysis RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the manufacturer s protocol and quantified by measuring the
Supplementary Methods RT-PCR and real-time PCR analysis RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the manufacturer s protocol and quantified by measuring the
Confocal immunofluorescence microscopy
 Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
Supplementary Information: Materials and Methods. Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered
 Supplementary Information: Materials and Methods Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered saline (PBS) and lysed in TNN lysis buffer (50mM Tris at ph 8.0, 120mM NaCl
Supplementary Information: Materials and Methods Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered saline (PBS) and lysed in TNN lysis buffer (50mM Tris at ph 8.0, 120mM NaCl
Sarker et al. Supplementary Material. Subcellular Fractionation
 Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Supplementary Information
 Supplementary Information stability is regulated by CK2-dependent interaction with R2TP complex Patrick von Morgen 1,2, Kamila Burdova 1, Thomas G. Flower 3, Nicola J. O'Reilly 4, Simon J. Boulton 5, Stephen
Supplementary Information stability is regulated by CK2-dependent interaction with R2TP complex Patrick von Morgen 1,2, Kamila Burdova 1, Thomas G. Flower 3, Nicola J. O'Reilly 4, Simon J. Boulton 5, Stephen
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Segments of the obstructed intestinal loops were fixed in 4% paraformaldehyde
 Supplementary text Supplementary materials and methods Histopathological examination Segments of the obstructed intestinal loops were fixed in 4% paraformaldehyde (PFA) and embedded in paraffin wax with
Supplementary text Supplementary materials and methods Histopathological examination Segments of the obstructed intestinal loops were fixed in 4% paraformaldehyde (PFA) and embedded in paraffin wax with
Supplemental data. Supplemental Materials and Methods
 Supplemental data Supplemental Materials and Methods Transfection of plasmid. Transfection of plasmids into FRTL5 cells was performed using Lipofectamine LTX with Plus reagent (Invitrogen) according to
Supplemental data Supplemental Materials and Methods Transfection of plasmid. Transfection of plasmids into FRTL5 cells was performed using Lipofectamine LTX with Plus reagent (Invitrogen) according to
Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and
 Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Han et al., http://www.jcb.org/cgi/content/full/jcb.201311007/dc1 Figure S1. SIVA1 interacts with PCNA. (A) HEK293T cells were transiently
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Han et al., http://www.jcb.org/cgi/content/full/jcb.201311007/dc1 Figure S1. SIVA1 interacts with PCNA. (A) HEK293T cells were transiently
SUPPLEMENTARY INFORMATION
 (Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
(Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
Supplemental Materials and Methods
 Supplemental Materials and Methods Co-immunoprecipitation (Co-IP) assay Cells were lysed with NETN buffer (20 mm Tris-HCl, ph 8.0, 0 mm NaCl, 1 mm EDT, 0.5% Nonidet P-40) containing 50 mm β-glycerophosphate,
Supplemental Materials and Methods Co-immunoprecipitation (Co-IP) assay Cells were lysed with NETN buffer (20 mm Tris-HCl, ph 8.0, 0 mm NaCl, 1 mm EDT, 0.5% Nonidet P-40) containing 50 mm β-glycerophosphate,
Quantitative and non-quantitative RT-PCR. cdna was generated from 500ng RNA (iscript;
 Supplemental Methods Quantitative and non-quantitative RT-PCR. cdna was generated from 500ng RNA (iscript; Bio-Rad, Hercules, CA, USA) and standard RT-PCR experiments were carried out using the 2X GoTaq
Supplemental Methods Quantitative and non-quantitative RT-PCR. cdna was generated from 500ng RNA (iscript; Bio-Rad, Hercules, CA, USA) and standard RT-PCR experiments were carried out using the 2X GoTaq
SUPPLEMENTARY INFORMATION
 The Supplementary Information (SI) Methods Cell culture and transfections H1299, U2OS, 293, HeLa cells were maintained in DMEM medium supplemented with 10% fetal bovine serum. H1299 and 293 cells were
The Supplementary Information (SI) Methods Cell culture and transfections H1299, U2OS, 293, HeLa cells were maintained in DMEM medium supplemented with 10% fetal bovine serum. H1299 and 293 cells were
supplementary information
 DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
UV-sensitive syndrome protein UVSSA recruits USP7 to regulate transcription-coupled repair
 Supplementary Information UVsensitive syndrome protein UVSSA recruits to regulate transcriptioncoupled repair Petra Schwertman 1, Anna Lagarou 2, Dick H.W. Dekkers 3, Anja Raams 1, Adriana C. van der Hoek
Supplementary Information UVsensitive syndrome protein UVSSA recruits to regulate transcriptioncoupled repair Petra Schwertman 1, Anna Lagarou 2, Dick H.W. Dekkers 3, Anja Raams 1, Adriana C. van der Hoek
Supplementary Materials and Methods
 Supplementary Materials and Methods sirna sequences used in this study The sequences of Stealth Select RNAi for ALK and FLOT-1 were as follows: ALK sense no.1 (ALK): 5 -AAUACUGACAGCCACAGGCAAUGUC-3 ; ALK
Supplementary Materials and Methods sirna sequences used in this study The sequences of Stealth Select RNAi for ALK and FLOT-1 were as follows: ALK sense no.1 (ALK): 5 -AAUACUGACAGCCACAGGCAAUGUC-3 ; ALK
SUPPLEMENTARY MATERIALS AND METHODS
 SUPPLEMENTARY MATERIALS AND METHODS Chemicals. All chemicals used in supplementary experiments were the same as in the manuscript except the following. Methyl-methanesulfonate (MMS) was from Fluka. NU1025
SUPPLEMENTARY MATERIALS AND METHODS Chemicals. All chemicals used in supplementary experiments were the same as in the manuscript except the following. Methyl-methanesulfonate (MMS) was from Fluka. NU1025
1. Cross-linking and cell harvesting
 ChIP is a powerful tool that allows the specific matching of proteins or histone modifications to regions of the genome. Chromatin is isolated and antibodies to the antigen of interest are used to determine
ChIP is a powerful tool that allows the specific matching of proteins or histone modifications to regions of the genome. Chromatin is isolated and antibodies to the antigen of interest are used to determine
Technical Note Detection of post-immunoprecipitation proteins by Western blot using the Quick Western Kit IRDye 680RD
 Technical Note Detection of post-immunoprecipitation proteins by Western blot using the Quick Western Kit IRDye 680RD Developed for: Aerius, Odyssey Classic, Odyssey CLx and Odyssey Sa Imaging Systems
Technical Note Detection of post-immunoprecipitation proteins by Western blot using the Quick Western Kit IRDye 680RD Developed for: Aerius, Odyssey Classic, Odyssey CLx and Odyssey Sa Imaging Systems
Anti-HB-EGF (Human) mab
 Page 1 For Research Use Only. Not for use in diagnostic procedures. CODE No. D308-3 Anti-HB-EGF (Human) mab CLONALITY CLONE ISOTYPE QUANTITY SOURCE IMMUNOGEN FORMURATION STORAGE Monoclonal 3H4 Mouse IgG1
Page 1 For Research Use Only. Not for use in diagnostic procedures. CODE No. D308-3 Anti-HB-EGF (Human) mab CLONALITY CLONE ISOTYPE QUANTITY SOURCE IMMUNOGEN FORMURATION STORAGE Monoclonal 3H4 Mouse IgG1
Apoptosis assay: Apoptotic cells were identified by Annexin V-Alexa Fluor 488 and Propidium
 Apoptosis assay: Apoptotic cells were identified by Annexin V-Alexa Fluor 488 and Propidium Iodide (Invitrogen, Carlsbad, CA) staining. Briefly, 2x10 5 cells were washed once in cold PBS and resuspended
Apoptosis assay: Apoptotic cells were identified by Annexin V-Alexa Fluor 488 and Propidium Iodide (Invitrogen, Carlsbad, CA) staining. Briefly, 2x10 5 cells were washed once in cold PBS and resuspended
Supplementary methods
 Supplementary methods Cell culture, infection, transfection, and RNA interference HEK293 cells and its derivatives were grown in DMEM supplemented with 10% FBS. Various constructs were introduced into
Supplementary methods Cell culture, infection, transfection, and RNA interference HEK293 cells and its derivatives were grown in DMEM supplemented with 10% FBS. Various constructs were introduced into
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
Supplementary Fig S1 Nutlin-3a treatment does not affect cell cycle progression in the absence
 Supplementary Figure Legends Supplementary Fig S1 Nutlin-3a treatment does not affect cell cycle progression in the absence of p53 or p21. HCT116 cells which were null for either p53 (A) or p21 (B) were
Supplementary Figure Legends Supplementary Fig S1 Nutlin-3a treatment does not affect cell cycle progression in the absence of p53 or p21. HCT116 cells which were null for either p53 (A) or p21 (B) were
Supplementary Information
 Supplementary Information Table of contents: Pages: Supplementary Methods 2-3 Supplementary Figure 1 4 Supplementary Figure 2 5 Supplementary Figure 3 6 Supplementary Figure 4 7 Supplementary Figure 5
Supplementary Information Table of contents: Pages: Supplementary Methods 2-3 Supplementary Figure 1 4 Supplementary Figure 2 5 Supplementary Figure 3 6 Supplementary Figure 4 7 Supplementary Figure 5
For Research Use Only. Not for use in diagnostic procedures. Anti-NRF2 mab
 Page 1 For Research Use Only. Not for use in diagnostic procedures. Anti-NRF2 mab CODE No. M200-3 CLONALITY CLONE ISOTYPE QUANTITY SOURCE IMMUNOGEN FORMURATION STORAGE Monoclonal 1F2 Mouse IgG1 100 L,
Page 1 For Research Use Only. Not for use in diagnostic procedures. Anti-NRF2 mab CODE No. M200-3 CLONALITY CLONE ISOTYPE QUANTITY SOURCE IMMUNOGEN FORMURATION STORAGE Monoclonal 1F2 Mouse IgG1 100 L,
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3209 Supplementary Figure 1 IR induces the association of FH with chromatin. a, U2OS cells synchronized by thymidine double block (2 mm) underwent no release (G1 phase) or release for 2
DOI: 10.1038/ncb3209 Supplementary Figure 1 IR induces the association of FH with chromatin. a, U2OS cells synchronized by thymidine double block (2 mm) underwent no release (G1 phase) or release for 2
Alpha or beta human chorionic gonadotropin knockdown decrease BeWo cell fusion by
 Alpha or beta human chorionic gonadotropin knockdown decrease BeWo cell fusion by down-regulating PKA and CREB activation Sudha Saryu Malhotra 1, Pankaj Suman 2 and Satish Kumar Gupta 1 * 1 Reproductive
Alpha or beta human chorionic gonadotropin knockdown decrease BeWo cell fusion by down-regulating PKA and CREB activation Sudha Saryu Malhotra 1, Pankaj Suman 2 and Satish Kumar Gupta 1 * 1 Reproductive
FIGURE S1. Representative images to illustrate RAD51 foci induction in FANCD2 wild-type (wt) and
 Supplementary Figures FIGURE S1. Representative images to illustrate RAD51 foci induction in FANCD2 wild-type (wt) and mutant (mut) cells. Higher magnification is shown in Figure 1A. Arrows indicate cisplatin-induced
Supplementary Figures FIGURE S1. Representative images to illustrate RAD51 foci induction in FANCD2 wild-type (wt) and mutant (mut) cells. Higher magnification is shown in Figure 1A. Arrows indicate cisplatin-induced
Cell extracts and western blotting RNA isolation and real-time PCR Chromatin immunoprecipitation (ChIP)
 Cell extracts and western blotting Cells were washed with ice-cold phosphate-buffered saline (PBS) and lysed with lysis buffer. 1 Total cell extracts were separated by SDS-PAGE and transferred to nitrocellulose
Cell extracts and western blotting Cells were washed with ice-cold phosphate-buffered saline (PBS) and lysed with lysis buffer. 1 Total cell extracts were separated by SDS-PAGE and transferred to nitrocellulose
Cdc42 Activation Assay Kit
 A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Cdc42 Activation Assay Kit Catalog Number: 80701 20 assays 1 Table of Content Product Description 3 Assay
A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Cdc42 Activation Assay Kit Catalog Number: 80701 20 assays 1 Table of Content Product Description 3 Assay
Supplementary Material - Methods
 Novel Protein-Protein Interactions in the Schizophrenia interactome Supplementary Material - Methods Experimental validations of predicted interactions Table S1-1: Protein pairs that were validated by
Novel Protein-Protein Interactions in the Schizophrenia interactome Supplementary Material - Methods Experimental validations of predicted interactions Table S1-1: Protein pairs that were validated by
For Research Use Only. Not for use in diagnostic procedures. Anti-NRF2 mab
 Page 1 For Research Use Only. Not for use in diagnostic procedures. Anti-NRF2 mab CODE No. M200-3 CLONALITY CLONE ISOTYPE QUANTITY SOURCE IMMUNOGEN FORMURATION STORAGE Monoclonal 1F2 Mouse IgG1 κ 100 µl,
Page 1 For Research Use Only. Not for use in diagnostic procedures. Anti-NRF2 mab CODE No. M200-3 CLONALITY CLONE ISOTYPE QUANTITY SOURCE IMMUNOGEN FORMURATION STORAGE Monoclonal 1F2 Mouse IgG1 κ 100 µl,
SUPPLEMENTARY INFORMATION. LIN-28 co-transcriptionally binds primary let-7 to regulate mirna maturation in C. elegans
 SUPPLEMENTARY INFORMATION LIN-28 co-transcriptionally binds primary let-7 to regulate mirna maturation in C. elegans Priscilla M. Van Wynsberghe 1, Zoya S. Kai 1, Katlin B. Massirer 2-4, Victoria H. Burton
SUPPLEMENTARY INFORMATION LIN-28 co-transcriptionally binds primary let-7 to regulate mirna maturation in C. elegans Priscilla M. Van Wynsberghe 1, Zoya S. Kai 1, Katlin B. Massirer 2-4, Victoria H. Burton
Supporting Information
 Supporting Information Stavru et al. 0.073/pnas.357840 SI Materials and Methods Immunofluorescence. For immunofluorescence, cells were fixed for 0 min in 4% (wt/vol) paraformaldehyde (Electron Microscopy
Supporting Information Stavru et al. 0.073/pnas.357840 SI Materials and Methods Immunofluorescence. For immunofluorescence, cells were fixed for 0 min in 4% (wt/vol) paraformaldehyde (Electron Microscopy
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling
 Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
CD93 and dystroglycan cooperation in human endothelial cell adhesion and migration
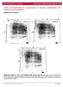 /, Supplementary Advance Publications Materials 2016 CD93 and dystroglycan cooperation in human endothelial cell adhesion and migration Supplementary Materials Supplementary Figure S1: In ECs CD93 silencing
/, Supplementary Advance Publications Materials 2016 CD93 and dystroglycan cooperation in human endothelial cell adhesion and migration Supplementary Materials Supplementary Figure S1: In ECs CD93 silencing
T H E J O U R N A L O F C E L L B I O L O G Y
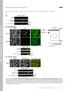 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Nakajima and Tanoue, http://www.jcb.org/cgi/content/full/jcb.201104118/dc1 Figure S1. DLD-1 cells exhibit the characteristic morphology
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Nakajima and Tanoue, http://www.jcb.org/cgi/content/full/jcb.201104118/dc1 Figure S1. DLD-1 cells exhibit the characteristic morphology
Myers Lab ChIP-seq Protocol v Modified January 10, 2014
 Myers Lab ChIP-seq Protocol V011014 1 Contact information: Dr. Florencia Pauli Behn HudsonAlpha Institute for Biotechnology 601 Genome Way Huntsville, AL 35806 Telephone: 256-327-5229 Email: fpauli@hudsonalpha.org
Myers Lab ChIP-seq Protocol V011014 1 Contact information: Dr. Florencia Pauli Behn HudsonAlpha Institute for Biotechnology 601 Genome Way Huntsville, AL 35806 Telephone: 256-327-5229 Email: fpauli@hudsonalpha.org
Supplemental Data. Noncoding Transcription by RNA Polymerase Pol IVb/Pol V Mediates Transcriptional Silencing of Overlapping and Adjacent Genes
 Cell, Volume 135 Supplemental Data Noncoding Transcription by RNA Polymerase Pol IVb/Pol V Mediates Transcriptional Silencing of Overlapping and Adjacent Genes Andrzej T. Wierzbicki, Jeremy R. Haag, and
Cell, Volume 135 Supplemental Data Noncoding Transcription by RNA Polymerase Pol IVb/Pol V Mediates Transcriptional Silencing of Overlapping and Adjacent Genes Andrzej T. Wierzbicki, Jeremy R. Haag, and
ChIP protocol Chromatin fragmentation using the Covaris S2 sonicator by Ethan Ford (version 12/1/11) X- link Cells
 ChIP protocol Chromatin fragmentation using the Covaris S2 sonicator by Ethan Ford (version 12/1/11) X- link Cells 1. Grow six 15 cm plates of HeLa cells to 90% confluency. 2. Remove media and add wash
ChIP protocol Chromatin fragmentation using the Covaris S2 sonicator by Ethan Ford (version 12/1/11) X- link Cells 1. Grow six 15 cm plates of HeLa cells to 90% confluency. 2. Remove media and add wash
A RRM1 H2AX DAPI. RRM1 H2AX DAPI Merge. Cont. sirna RRM1
 A H2AX DAPI H2AX DAPI Merge Cont sirna Figure S1: Accumulation of RRM1 at DNA damage sites (A) HeLa cells were subjected to in situ detergent extraction without IR irradiation, and immunostained with the
A H2AX DAPI H2AX DAPI Merge Cont sirna Figure S1: Accumulation of RRM1 at DNA damage sites (A) HeLa cells were subjected to in situ detergent extraction without IR irradiation, and immunostained with the
Supplementary Figure 1. RAD51 and RAD51 paralogs are enriched spontaneously onto
 Supplementary Figure legends Supplementary Figure 1. and paralogs are enriched spontaneously onto the S-phase chromatin during DN replication. () Chromatin fractionation was carried out as described in
Supplementary Figure legends Supplementary Figure 1. and paralogs are enriched spontaneously onto the S-phase chromatin during DN replication. () Chromatin fractionation was carried out as described in
Supplemental Data Supplementary Figure Legends and Scheme Figure S1.
 Supplemental Data Supplementary Figure Legends and Scheme Figure S1. UTK1 inhibits the second EGF-induced wave of lamellipodia formation in TT cells. A and B, EGF-induced lamellipodia formation in TT cells,
Supplemental Data Supplementary Figure Legends and Scheme Figure S1. UTK1 inhibits the second EGF-induced wave of lamellipodia formation in TT cells. A and B, EGF-induced lamellipodia formation in TT cells,
SUPPLEMENTAL MATERIAL. Supplemental Methods:
 SUPPLEMENTAL MATERIAL Supplemental Methods: Immunoprecipitation- As we described but with some modifications [22]. As part of another ongoing project, lysate from human umbilical vein endothelial cells
SUPPLEMENTAL MATERIAL Supplemental Methods: Immunoprecipitation- As we described but with some modifications [22]. As part of another ongoing project, lysate from human umbilical vein endothelial cells
Supplementary information. Supplementary Figures
 Supplementary information Supplementary Figures Supplementary Figure 1. A. i. HA-JMY expressing U2OS cells were treated with SAHA (6h). DAPI was used to visualise nuclei. ii. U2OS cells stably expressing
Supplementary information Supplementary Figures Supplementary Figure 1. A. i. HA-JMY expressing U2OS cells were treated with SAHA (6h). DAPI was used to visualise nuclei. ii. U2OS cells stably expressing
IMMUNOPRECIPITATION (IP)
 1 IMMUNOPRECIPITATION (IP) Overview and Technical Tips 2 CONTENTS 3 7 8 9 12 13 17 18 19 20 Introduction Factors Influencing IP General Protocol Modifications Of IP Protocols Troubleshooting Contact Us
1 IMMUNOPRECIPITATION (IP) Overview and Technical Tips 2 CONTENTS 3 7 8 9 12 13 17 18 19 20 Introduction Factors Influencing IP General Protocol Modifications Of IP Protocols Troubleshooting Contact Us
Supplemental Information
 Supplemental Information Intrinsic protein-protein interaction mediated and chaperonin assisted sequential assembly of a stable Bardet Biedl syndome protein complex, the BBSome * Qihong Zhang 1#, Dahai
Supplemental Information Intrinsic protein-protein interaction mediated and chaperonin assisted sequential assembly of a stable Bardet Biedl syndome protein complex, the BBSome * Qihong Zhang 1#, Dahai
Rab5 Activation Assay Kit
 A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Rab5 Activation Assay Kit Catalog Number: 83701 20 assays 24 Whitewoods Lane 1 Table of Content Product
A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Rab5 Activation Assay Kit Catalog Number: 83701 20 assays 24 Whitewoods Lane 1 Table of Content Product
ApoTrack Cytochrome c Apoptosis ICC Antibody
 ab110417 ApoTrack Cytochrome c Apoptosis ICC Antibody Instructions for Use For the Immunocytochemistry analysis of cytochrome c and a mitochondrial marker (Complex Vα) in apoptotic cells and nonapoptotic
ab110417 ApoTrack Cytochrome c Apoptosis ICC Antibody Instructions for Use For the Immunocytochemistry analysis of cytochrome c and a mitochondrial marker (Complex Vα) in apoptotic cells and nonapoptotic
Methods Western blot analysis of plg Quantification of plasminogen accumulation by ELISA Immunohistochemical analysis
 Methods Western blot analysis of plg Wild-type mice first received a standardized burn wound and then were intravenously administered 2 mg of human plg (Omnio AB, Umeå, Sweden). 24 hours after wounding
Methods Western blot analysis of plg Wild-type mice first received a standardized burn wound and then were intravenously administered 2 mg of human plg (Omnio AB, Umeå, Sweden). 24 hours after wounding
Anti-CLOCK (Mouse) mab
 Page 1 For Research Use Only. Not for use in diagnostic procedures. Anti-CLOCK (Mouse) mab CODE No. D349-3 CLONALITY CLONE ISOTYPE QUANTITY SOURCE IMMUNOGEN FORMURATION STORAGE Monoclonal CLSP4 Mouse IgG1
Page 1 For Research Use Only. Not for use in diagnostic procedures. Anti-CLOCK (Mouse) mab CODE No. D349-3 CLONALITY CLONE ISOTYPE QUANTITY SOURCE IMMUNOGEN FORMURATION STORAGE Monoclonal CLSP4 Mouse IgG1
Arf6 Activation Assay Kit
 A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Arf6 Activation Assay Kit Catalog Number: 82401 20 assays NewEast Biosciences 1 Table of Content Product
A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Arf6 Activation Assay Kit Catalog Number: 82401 20 assays NewEast Biosciences 1 Table of Content Product
EPIGENTEK. EpiQuik Chromatin Immunoprecipitation Kit. Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Chromatin Immunoprecipitation Kit Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Chromatin Immunoprecipitation Kit is suitable for combining the specificity of
EpiQuik Chromatin Immunoprecipitation Kit Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Chromatin Immunoprecipitation Kit is suitable for combining the specificity of
ApoTrack Cytochrome c Apoptosis ICC Antibody Kit
 ab110417 ApoTrack Cytochrome c Apoptosis ICC Antibody Kit Instructions for Use For the Immunocytochemistry analysis of cytochrome c and a mitochondrial marker (Complex Vα) in apoptotic cells and non-apoptotic
ab110417 ApoTrack Cytochrome c Apoptosis ICC Antibody Kit Instructions for Use For the Immunocytochemistry analysis of cytochrome c and a mitochondrial marker (Complex Vα) in apoptotic cells and non-apoptotic
SUPPLEMENTAL MATERIAL. Supplemental material contains Supplemental Figure Legends and Supplemental Figures 1 to
 SUPPLEMENTAL MATERIAL Supplemental material contains Supplemental Figure Legends and Supplemental Figures 1 to 6. SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure 1. Overview of the mechanisms by which
SUPPLEMENTAL MATERIAL Supplemental material contains Supplemental Figure Legends and Supplemental Figures 1 to 6. SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure 1. Overview of the mechanisms by which
ASPP1 Fw GGTTGGGAATCCACGTGTTG ASPP1 Rv GCCATATCTTGGAGCTCTGAGAG
 Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
rev. 05/31/12 Box 2 contains the following buffers/reagents:
 Store at RT, 4 C, and 20 C #9002 SimpleChIP Enzymatic Chromatin IP Kit (Agarose Beads) 3 n 1 Kit (30 Immunoprecipitations) For Research Use Only. Not For Use In Diagnostic Procedures. rev. 05/31/12 Orders
Store at RT, 4 C, and 20 C #9002 SimpleChIP Enzymatic Chromatin IP Kit (Agarose Beads) 3 n 1 Kit (30 Immunoprecipitations) For Research Use Only. Not For Use In Diagnostic Procedures. rev. 05/31/12 Orders
Gα i Activation Assay Kit
 A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Gα i Activation Assay Kit Catalog Number 80301 20 assays NewEast Biosciences, Inc 1 Table of Content Product
A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Gα i Activation Assay Kit Catalog Number 80301 20 assays NewEast Biosciences, Inc 1 Table of Content Product
Transcriptional regulation of BRCA1 expression by a metabolic switch: Di, Fernandez, De Siervi, Longo, and Gardner. H3K4Me3
 ChIP H3K4Me3 enrichment.25.2.15.1.5 H3K4Me3 H3K4Me3 ctrl H3K4Me3 + E2 NS + E2 1. kb kb +82 kb Figure S1. Estrogen promotes entry of MCF-7 into the cell cycle but does not significantly change activation-associated
ChIP H3K4Me3 enrichment.25.2.15.1.5 H3K4Me3 H3K4Me3 ctrl H3K4Me3 + E2 NS + E2 1. kb kb +82 kb Figure S1. Estrogen promotes entry of MCF-7 into the cell cycle but does not significantly change activation-associated
Figure S2. Response of mouse ES cells to GSK3 inhibition. Mentioned in discussion
 Stem Cell Reports, Volume 1 Supplemental Information Robust Self-Renewal of Rat Embryonic Stem Cells Requires Fine-Tuning of Glycogen Synthase Kinase-3 Inhibition Yaoyao Chen, Kathryn Blair, and Austin
Stem Cell Reports, Volume 1 Supplemental Information Robust Self-Renewal of Rat Embryonic Stem Cells Requires Fine-Tuning of Glycogen Synthase Kinase-3 Inhibition Yaoyao Chen, Kathryn Blair, and Austin
HPV E6 oncoprotein targets histone methyltransferases for modulating specific. Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu,
 1 HPV E oncoprotein targets histone methyltransferases for modulating specific gene transcription 3 5 Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu, Cheng-Ming Chiang, Sheng-Chung
1 HPV E oncoprotein targets histone methyltransferases for modulating specific gene transcription 3 5 Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu, Cheng-Ming Chiang, Sheng-Chung
ipure kit v2 Magnetic DNA Purification kit for epigenetic applications Cat. No. C (x24) C (x100) Version 2 I 07.15
 ipure kit v2 Magnetic DNA Purification kit for epigenetic applications Cat. No. C03010014 (x24) C03010015 (x100) Version 2 I 07.15 Ordering information diagenode headquarters Diagenode s.a. BELGIUM EUROPE
ipure kit v2 Magnetic DNA Purification kit for epigenetic applications Cat. No. C03010014 (x24) C03010015 (x100) Version 2 I 07.15 Ordering information diagenode headquarters Diagenode s.a. BELGIUM EUROPE
ChIP-seq Protocol for RNA-Binding Proteins
 ChIP-seq: Cells were grown according to the approved ENCODE cell culture protocols. Cells were fixed in 1% formaldehyde and resuspended in lysis buffer. Chromatin was sheared to 100-500 bp using a Branson
ChIP-seq: Cells were grown according to the approved ENCODE cell culture protocols. Cells were fixed in 1% formaldehyde and resuspended in lysis buffer. Chromatin was sheared to 100-500 bp using a Branson
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3363 Supplementary Figure 1 Several WNTs bind to the extracellular domains of PKD1. (a) HEK293T cells were co-transfected with indicated plasmids. Flag-tagged proteins were immunoprecipiated
DOI: 10.1038/ncb3363 Supplementary Figure 1 Several WNTs bind to the extracellular domains of PKD1. (a) HEK293T cells were co-transfected with indicated plasmids. Flag-tagged proteins were immunoprecipiated
H3K36me3 polyclonal antibody
 H3K36me3 polyclonal antibody Cat. No. C15410192 Type: Polyclonal ChIP-grade/ChIP-seq grade Source: Rabbit Lot #: A1845P Size: 50 µg/32 µl Concentration: 1.6 μg/μl Specificity: Human, mouse, Arabidopsis,
H3K36me3 polyclonal antibody Cat. No. C15410192 Type: Polyclonal ChIP-grade/ChIP-seq grade Source: Rabbit Lot #: A1845P Size: 50 µg/32 µl Concentration: 1.6 μg/μl Specificity: Human, mouse, Arabidopsis,
RheB Activation Assay Kit
 A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based RheB Activation Assay Kit Catalog Number: 81201 20 assays NewEast Biosciences 1 FAX: 610-945-2008 Table
A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based RheB Activation Assay Kit Catalog Number: 81201 20 assays NewEast Biosciences 1 FAX: 610-945-2008 Table
Histone H3K27 Methylation Antibody Panel Pack Base Catalog # C Component Size Shipping Temperature
 Histone H3K27 Methylation Antibody Panel Pack Base Catalog # PACK CONTENTS Component Size Shipping Temperature Upon Receipt Checklist 3K27M Histone H3K27me1 (H3K27 Monomethyl) Polyclonal Antibody 25 µg
Histone H3K27 Methylation Antibody Panel Pack Base Catalog # PACK CONTENTS Component Size Shipping Temperature Upon Receipt Checklist 3K27M Histone H3K27me1 (H3K27 Monomethyl) Polyclonal Antibody 25 µg
Gα 13 Activation Assay Kit
 A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Gα 13 Activation Assay Kit Catalog Number: 80401 20 assays NewEast Biosciences 1 Table of Content Product
A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Gα 13 Activation Assay Kit Catalog Number: 80401 20 assays NewEast Biosciences 1 Table of Content Product
B. ADM: C. D. Apoptosis: 1.68% 2.99% 1.31% Figure.S1,Li et al. number. invaded cells. HuH7 BxPC-3 DLD-1.
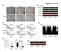 A. - Figure.S1,Li et al. B. : - + - + - + E-cadherin CK19 α-sma vimentin β -actin C. D. Apoptosis: 1.68% 2.99% 1.31% - : - + - + - + Apoptosis: 48.33% 45.32% 44.59% E. invaded cells number 400 300 200
A. - Figure.S1,Li et al. B. : - + - + - + E-cadherin CK19 α-sma vimentin β -actin C. D. Apoptosis: 1.68% 2.99% 1.31% - : - + - + - + Apoptosis: 48.33% 45.32% 44.59% E. invaded cells number 400 300 200
ab Ran Activation Assay Kit
 ab173247 Ran Activation Assay Kit Instructions for Use For the simple and fast measurement of Ran activation. This product is for research use only and is not intended for diagnostic use. Version 1 Last
ab173247 Ran Activation Assay Kit Instructions for Use For the simple and fast measurement of Ran activation. This product is for research use only and is not intended for diagnostic use. Version 1 Last
Description of supplementary material file
 Description of supplementary material file In the supplementary results we show that the VHL-fibronectin interaction is indirect, mediated by fibronectin binding to COL4A2. This provides additional information
Description of supplementary material file In the supplementary results we show that the VHL-fibronectin interaction is indirect, mediated by fibronectin binding to COL4A2. This provides additional information
Mechanism of Induction and Suppression of Antiviral Immunity Directed by Virus-Derived Small RNAs in Drosophila
 Cell Host & Microbe, Volume 4 Supplemental Data Mechanism of Induction and Suppression of Antiviral Immunity Directed by Virus-Derived Small RNAs in Drosophila Roghiyh Aliyari, Qingfa Wu, Hong-Wei Li,
Cell Host & Microbe, Volume 4 Supplemental Data Mechanism of Induction and Suppression of Antiviral Immunity Directed by Virus-Derived Small RNAs in Drosophila Roghiyh Aliyari, Qingfa Wu, Hong-Wei Li,
Beta3 integrin promotes long-lasting activation and polarization of Vascular Endothelial Growth Factor Receptor 2 by immobilized ligand
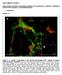 SUPPLEMENTAL FIGURES Beta3 integrin promotes long-lasting activation and polarization of Vascular Endothelial Growth Factor Receptor 2 by immobilized ligand C. Ravelli et al. FIGURE S. I Figure S. I: Gremlin
SUPPLEMENTAL FIGURES Beta3 integrin promotes long-lasting activation and polarization of Vascular Endothelial Growth Factor Receptor 2 by immobilized ligand C. Ravelli et al. FIGURE S. I Figure S. I: Gremlin
Supplementary Fig. 1
 a FL (1-2266) NL (1-1190) CL (1191-2266) HA-ICE1: - HA-ICE1: - - - FLAG-ICE2: + + + + FLAG-ELL: + + + + + + IP: anti-ha FLAG-ICE2 HA-ICE1-FL HA-ICE1-NL HA-ICE1-CL FLAG-ICE2 b IP: anti-ha FL (1-2266) NL
a FL (1-2266) NL (1-1190) CL (1191-2266) HA-ICE1: - HA-ICE1: - - - FLAG-ICE2: + + + + FLAG-ELL: + + + + + + IP: anti-ha FLAG-ICE2 HA-ICE1-FL HA-ICE1-NL HA-ICE1-CL FLAG-ICE2 b IP: anti-ha FL (1-2266) NL
Coleman et al., Supplementary Figure 1
 Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Antibodies against PCNA were previously described [1]. To deplete PCNA from Xenopus egg
![Antibodies against PCNA were previously described [1]. To deplete PCNA from Xenopus egg Antibodies against PCNA were previously described [1]. To deplete PCNA from Xenopus egg](/thumbs/84/90687567.jpg) Supplementary information Supplementary methods PCNA antibody and immunodepletion Antibodies against PCNA were previously described [1]. To deplete PCNA from Xenopus egg extracts, one volume of protein
Supplementary information Supplementary methods PCNA antibody and immunodepletion Antibodies against PCNA were previously described [1]. To deplete PCNA from Xenopus egg extracts, one volume of protein
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Histone H3 Methylation Antibody Panel Pack I - Active Genes Base Catalog # C Component Size Shipping Temperature
 Histone H3 Methylation Antibody Panel Pack I - Active Genes Base Catalog # PACK CONTENTS Component Size Shipping Temperature Upon Receipt Checklist 3K4D Histone H3K4me2 (H3K4 Dimethyl) Polyclonal Antibody
Histone H3 Methylation Antibody Panel Pack I - Active Genes Base Catalog # PACK CONTENTS Component Size Shipping Temperature Upon Receipt Checklist 3K4D Histone H3K4me2 (H3K4 Dimethyl) Polyclonal Antibody
Like use other ChIP kits, before handle ChIP assay please choose a good antibody suitable for precipitation the crosslinked protein / DNA complexes.
 ChIP Assay Kit Cat:RK20100 Like use other ChIP kits, before handle ChIP assay please choose a good antibody suitable for precipitation the crosslinked protein / DNA complexes. Manufactured by Global Headquarters
ChIP Assay Kit Cat:RK20100 Like use other ChIP kits, before handle ChIP assay please choose a good antibody suitable for precipitation the crosslinked protein / DNA complexes. Manufactured by Global Headquarters
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or
 Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
A549-luc mock cells, A549-luc/shCon cells and A549-luc/shYY1 cells were seeded onto 35 mm
 Supplementary Material and Methods Real-time luciferase gene expression in A549-luc cells A549-luc mock cells, A549-luc/shCon cells and A549-luc/shYY1 cells were seeded onto 35 mm dishes (100,000 cells/dish),
Supplementary Material and Methods Real-time luciferase gene expression in A549-luc cells A549-luc mock cells, A549-luc/shCon cells and A549-luc/shYY1 cells were seeded onto 35 mm dishes (100,000 cells/dish),
ApoTrack Cytochrome c Apoptosis ICC Antibody Kit: 2 color immunocytochemistry of cytochrome c and mitochondria.
 PROTOCOL ApoTrack Cytochrome c Apoptosis ICC Antibody Kit 1850 Millrace Drive, Suite 3A Eugene, Oregon 97403 MSA07 Rev.1 DESCRIPTION ApoTrack Cytochrome c Apoptosis ICC Antibody Kit: 2 color immunocytochemistry
PROTOCOL ApoTrack Cytochrome c Apoptosis ICC Antibody Kit 1850 Millrace Drive, Suite 3A Eugene, Oregon 97403 MSA07 Rev.1 DESCRIPTION ApoTrack Cytochrome c Apoptosis ICC Antibody Kit: 2 color immunocytochemistry
supplementary information
 DOI: 10.1038/ncb2172 Figure S1 p53 regulates cellular NADPH and lipid levels via inhibition of G6PD. (a) U2OS cells stably expressing p53 shrna or a control shrna were transfected with control sirna or
DOI: 10.1038/ncb2172 Figure S1 p53 regulates cellular NADPH and lipid levels via inhibition of G6PD. (a) U2OS cells stably expressing p53 shrna or a control shrna were transfected with control sirna or
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2579 Figure S1 Incorporation of heavy isotope-labeled amino acids and enrichment of di-glycine modified peptides. The incorporation of isotopelabeled amino acids in peptides was calculated
DOI: 10.1038/ncb2579 Figure S1 Incorporation of heavy isotope-labeled amino acids and enrichment of di-glycine modified peptides. The incorporation of isotopelabeled amino acids in peptides was calculated
Instruction Manual Version IPure kit. Magnetic DNA Purification kit for epigenetic applications. Cat. No. C (AL )
 Instruction Manual Version 4-12.15 IPure kit Magnetic DNA Purification kit for epigenetic applications Cat. No. C03010011 (AL-100-0024) PAGE 3 Content Introduction 4 Method Overview 5 Kit Content 6 Time
Instruction Manual Version 4-12.15 IPure kit Magnetic DNA Purification kit for epigenetic applications Cat. No. C03010011 (AL-100-0024) PAGE 3 Content Introduction 4 Method Overview 5 Kit Content 6 Time
T H E J O U R N A L O F C E L L B I O L O G Y
 Supplemental material Thompson et al., http://www.jcb.org/cgi/content/full/jcb.200909067/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Modification-specific antibodies do not detect unmodified
Supplemental material Thompson et al., http://www.jcb.org/cgi/content/full/jcb.200909067/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Modification-specific antibodies do not detect unmodified
supplementary information
 DOI: 10.1038/ncb1864 Figure S1 Apak specifically inhibits p53 transcriptional activity. Transcription activity of p53 was measured in U2OS (p53 wild-type) and H1299 (p53 deficient) cells which were transfected
DOI: 10.1038/ncb1864 Figure S1 Apak specifically inhibits p53 transcriptional activity. Transcription activity of p53 was measured in U2OS (p53 wild-type) and H1299 (p53 deficient) cells which were transfected
Supplemental information
 Supplemental information - Control samples (200 subjects) - Immunohistochemistry of rat brain - Immunocytochemistry on neuronal cultures - Immunocompetition assay - Immunoprecipitation - Immunocytochemistry
Supplemental information - Control samples (200 subjects) - Immunohistochemistry of rat brain - Immunocytochemistry on neuronal cultures - Immunocompetition assay - Immunoprecipitation - Immunocytochemistry
Supplementary Information: Materials and Methods. GST and GST-p53 were purified according to standard protocol after
 Supplementary Information: Materials and Methods Recombinant protein expression and in vitro kinase assay. GST and GST-p53 were purified according to standard protocol after induction with.5mm IPTG for
Supplementary Information: Materials and Methods Recombinant protein expression and in vitro kinase assay. GST and GST-p53 were purified according to standard protocol after induction with.5mm IPTG for
Supplementary Figure 1. Intracellular distribution of the EPE peptide. HeLa cells were serum-starved (16 h, 0.1%), and treated with EPE peptide,
 Supplementary Figure 1. Intracellular distribution of the EPE peptide. HeLa cells were serum-starved (16 h, 0.1%), and treated with EPE peptide, conjugated with either TAT or Myristic acid and biotin for
Supplementary Figure 1. Intracellular distribution of the EPE peptide. HeLa cells were serum-starved (16 h, 0.1%), and treated with EPE peptide, conjugated with either TAT or Myristic acid and biotin for
Supplemental Information. PARP1 Represses PAP and Inhibits Polyadenylation during Heat Shock
 Molecular Cell, Volume 49 Supplemental Information PARP1 Represses PAP and Inhibits Polyadenylation during Heat Shock Dafne Campigli Di Giammartino, Yongsheng Shi, and James L. Manley Supplemental Information
Molecular Cell, Volume 49 Supplemental Information PARP1 Represses PAP and Inhibits Polyadenylation during Heat Shock Dafne Campigli Di Giammartino, Yongsheng Shi, and James L. Manley Supplemental Information
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the
 Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Supplemental Material: Rev1 promotes replication through UV lesions in conjunction with DNA
 Supplemental Material: Rev1 promotes replication through UV lesions in conjunction with DNA polymerases,, and, but not with DNA polymerase Jung-Hoon Yoon, Jeseong Park, Juan Conde, Maki Wakamiya, Louise
Supplemental Material: Rev1 promotes replication through UV lesions in conjunction with DNA polymerases,, and, but not with DNA polymerase Jung-Hoon Yoon, Jeseong Park, Juan Conde, Maki Wakamiya, Louise
Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas.
 Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas. Target Gene Non-target / Control BTG2 BRCA1 NFE2L2 Target Sequence ON-TARGET plus Non-targeting sirna # 1 (Cat# D-001810-01-05) sirna1: GAACCGACAUGCUCCCGGA
Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas. Target Gene Non-target / Control BTG2 BRCA1 NFE2L2 Target Sequence ON-TARGET plus Non-targeting sirna # 1 (Cat# D-001810-01-05) sirna1: GAACCGACAUGCUCCCGGA
EpiQuik Methyl-CpG Binding Domain Protein 2 ChIP Kit
 EpiQuik Methyl-CpG Binding Domain Protein 2 ChIP Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Methyl-CpG Binding Domain Protein 2 ChIP Kit is suitable for combining the
EpiQuik Methyl-CpG Binding Domain Protein 2 ChIP Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Methyl-CpG Binding Domain Protein 2 ChIP Kit is suitable for combining the
