Epigenetic genome control by RNAi and transposon-derived proteins
|
|
|
- Francis Patterson
- 5 years ago
- Views:
Transcription
1 Epigenetic genome control by RNAi and transposon-derived proteins Shiv Grewal Center for Cancer Research, NCI National Institutes of Health, Bethesda
2 Transposons and their remnants constitute a significant fraction of eukaryotic genomes Organism Yeast Worm Drosophila Mouse Human Transposon content (% of genome) 3.1% 6.% -12% 38% 48% Heterochromatin / RNAi Transposon-derived proteins
3 Distinct patterns of histone modifications specify discrete downstream regulatory events Heterochromatin Euchromatin Adapted from Zhang and Reinberg 2001
4 Heterochromatin coats extended domains associated with a specific classes of repeat elements in S. pombe Centromere Subtelomeric region pgp1 dg dh cnt dg dh cwf20 dh repeat LTRs ChIP fold enrichment H3K9me Swi6 H3K4me ChIP fold enrichment H3K9me Swi6 H3K4me mat1 mat locus IR-L mat2p cenh mat3m IR-R 2. kb ChIP fold enrichment H3K9me Swi6 H3K4me Grewal and Klar Cell 1996
5 Nucleation and spreading of heterochromatin Nucleation (S phase) Spreading H3K9me Clr4 Dcr1 RdRP Repeat elements Rdp1 Chp1 Ago1 Tas3 RITS Clr4 Rik1 H3K9 HMTase Pol II Clr4 Chp1 Ac STOP sirna = Swi6, Chp2 Boundary element Spreading of heterochromatin requires Clr4 chromodomain binding to methylated H3K9 Zhang et al 2008 NSMB 1:381
6 Heterochromatin serves as a versatile platform for targeting effectors across extended domains Grewal and Jia Nat Rev Genet 8:3-46
7 Post-transcriptional and transcriptional heterochromatic silencing Clr6-C1 Dcr1 sirna Factory SHREC H3K9me Clr6-C2 Repeat elements RDRC Rdp1 Chp1 Ago1 Tas3 Rik1 RITS ClrC sirna PTGS Clr4 Pol II Clr4 Clr4 = Swi6, Chp2 Chp1 SHREC TGS Ac STOP Boundary element Ago1 Clr6-C1 Clr6-C2 SHREC RISC Clr6 Pst1 (Sin3) Pst2 (Sin3) Cph2 Prw1 Clr6 (HDAC) Cph1 Alp13 (MRG1) Png2 TGS effector complexes Clr3 Mit1 (Snf2 ATPase) Clr1 Ccq1 Clr3 (HDAC) Clr2 Sds3 Nicolas et al 2007 NSMB Sugiyama et al 2007 Cell
8 Probe SHREC activities facilitate proper positioning of nucleosomes required for higher-order chromatin assembly Micrococcal nuclease digestion patterns Sugiyama et al 2007 Cell 128:491
9 Heterochromatin mediates recruitment of cohesin that is essential for maintenance of genomic integrity Transcriptional and Recombinational Block Euchromatin Heterochromatin Euchromatin mat1 IR-L mat2p cenh mat3m IR-R 2. kb Swi6 Cohesin K. Noma and S. Grewal
10 WT swi (fold enrichment) Swi Complex M cells (fold enrichment) Swi Complex P cells Heterochromatin mediates cell-type specific spreading of a protein complex essential for mating-type switching mat1-p mat1-m WT swi α-swi WCE α-swi WCE mat2-p cenh mat3-m Enhancer Element 2. kb WT swi6 - α-swi WCE α-swi WCE mat act1 mat act1 mat act1 mat act1 WT swi6 - mat act1 mat act1 mat act1 mat act1
11 Boundaries of a heterochromatic domain Transcriptional and Recombinational Block Euchromatin Heterochromatin Euchromatin 2. kb mat1 IR-L mat2p cenh mat3m IR-R Swi6 Fold Enrichment H3 Lys9-methyl H3 Lys4-methyl
12 TFIIIC binding to the boundary elements blocks spreading of heterochromatin heterochromatin spreading mat3 IR-R kan + ura4 + 1 kb wt IR-R IR-R #1 IR-R #2 IR-R #3 IR-R #4 B-box - FOA + FOA mat3m B-box cons. B-box seq#1 B-box seq#2 B-box seq#3 B-box seq#4 B-box seq# 1 kb IR-R...GTTCGANTC... ***** * * AGCTGTTCGTAGCAGCA ******* GCTCTTTCGATTATAGA ********* CCCGGTTCGAGTCCTAT ******** AGCGGTTCGATTTGCGA **** *** TAAGGTTCTACTTTATC mat3m IR-R B-box ura kb H3 Lys9-methyl (fold enrichment) 20 wt B-box
13 TFIIIC, but not Pol III, localizes to heterochromatin domain boundaries ChIP-chip
14 TFIIIC bind to trna gene clusters at centromere heterochromatin domain boundaries ChIP-chip analyses
15 TFIIIC, but not Pol III, peaks at sites are predominantly associated with divergent gene promoter regions COC3 SPBC336.0c sfc3 fold enrichment TFIIIc Pol III B-box COC4 SPBC139.0 SPBC139.07c fold enrichment COC SPCC63.02c SPCC63.03 fold enrichment 40 20
16 TFIIIC bound loci cluster near the nuclear periphery: implications for genome organization and function TFIIIC/DAPI TFIIIC body
17 Host genome surveillance for retrotransposons and repeats by transposon-derived proteins CENP-Bs are conserved proteins that contain DNA binding and dimerization domains CENP-Bs are derived from transposases of POGO DNA transposons S. pombe CENP-B ABP1 CBH1 CBH2 DNA binding domain DDE Dimerization domain S. pombe genome encodes three CENP-Bs that have redundant roles in centromere heterochromatin formation Mammalian CENP-B POGOK transposase
18 CENP-Bs localize to retrotransposons and their remnants in the S. pombe genome Chromosome I Chromosome II 4.6 Mb Fold enrichment.7 Mb tel1l 14 Chromosome III cen1 tel1r tel2l 3. Mb cen2 mat tel2r Tf2-4 Tf Tf2-3 Tf2- Tf2-2 Tf2-6 Tf2-1 LTRs LTR 8 12 Tf2-11 Tf2-9 LTR IG rip1 IG 8 Tf2- LTRs 12 LTRs Fold enrichment 6 1,000 2,000 3,000 4,000, spo20 IG 1 LTR SPAP32A8.02 Tf2-4 Tf2-3 Tf2-2 Tf2-1 LTRs 1,000 Tf2-6 Tf2-3, ,000 2,000 3,000 4,000,000 Chromosome Position (Kb) 00 rip1 IG Tf F.03 IG LTR Tf IG Tf2- cwf IG LTRs 2 LTRs 3,000 4,000 gut2 IG 1,00 2,000 gut2 IG 20 Tf ,000 1, c IG 1,000 srk1 IG rpc34 IG Tf2-13 LTR rpc34 IGIG rpc34 tel3r ars02 wtf6 4,000 2 Tf ,000 LTR 8 6 Cbh1 rdna Tf cen3 rdna Abp1 tel3l srk1 IG LTR Tf2-13 wtf6 00 1,000 1,00 2,000
19 CENP-Bs displaying distinct peaks at LTRs repress Tf2 retrotransposons Relative fold expression to WT Tf2-LTR Tf2-orf RT-PCR abp1δ cbh1δ cbh2δ abp1δ cbh1δ abp1δ cbh2δ cbh1δ cbh2δ abp1δ cbh1δ cbh2δ
20 CENP-Bs silence LTR-associated genes SPCC11E.07c Relative fold enrichment Abp1 Cbh1 1,462 1,466 1,470 1,474 Chromosome III Position (Kb) 27D7.09c LTR rik1 Tandem LTRs 27D7.11c Relative fold expression to WT E.07c abp1 cbh1 cbh2 abp1 cbh1 abp1 cbh2 cbh1 cbh2 abp1 cbh1 cbh2 Relative fold enrichment Abp1 Cbh1 27D7.c 4,21 4,2 4,29 4,33 Chromosome I Position (Kb) Relative fold expression to WT D7.c abp1 cbh1 cbh2 abp1 cbh1 abp1 cbh2 cbh1 cbh2 abp1 cbh1 cbh2
21 WT CENP-Bs recruit Clr3 and Clr6 histone deacetylases to repress Tf2 retroelements Co-Immunoprecipitation αflag IP Input Clr3-myc Abp1-FLAG IB: αmyc IB: αflag Clr3 ChIP abp Tf2-12 control Clr6 αflag IP - + Input - + Abp1-FLAG IB: αclr6 Input IB: αflag Clr3 = SHREC Clr6 = Clr6 HDAC complexes
22 CENP-Bs and heterochromatin recruit same repressor complexes Clr6 and SHREC repressors 1 CENP-B LTR retrotransposon Repeats Heterochromatic domain Gene
23 CENP-B mechanism silences and immobilizes an extinct retroelement that becomes sequestered into Tf bodies Tf1-neo Tf2 DAPI 7D4.08 DAPI Merged neo - Tf1 7D4.08 Abp1 ChIP Tf1 - + Cbh1 ChIP Tf D4.08 control + Tf1 Tf body Tf LTR CENP-B HDAC?
24 Summary and Implications Pol II transcription of repeats during S-phase facilitates RNAimediated targeting of heterochromatin Heterochromatin is dynamically regulated and serves as a recruiting platform for targeting variety of factors Host genome surveillance for retrotransposons by transposonderived CENP-B proteins - maintenance of genomic integrity - retrotransposon-mediated genome organization CENP-B Tf Bodies retrotransposon surveillance genome organization gene regulatory networks
Heterochromatin Silencing
 Heterochromatin Silencing Heterochromatin silencing Most DNA in eukaryotes consists of repetitive DNA, including retrotransposons, transposable elements. Packaged into a condensed form: Heterochromatin:
Heterochromatin Silencing Heterochromatin silencing Most DNA in eukaryotes consists of repetitive DNA, including retrotransposons, transposable elements. Packaged into a condensed form: Heterochromatin:
Investigating the Role of RNA Polymerase II in RNAi-dependent Heterochromatin Assembly at Centromeric Repeats
 Investigating the Role of RNA Polymerase II in RNAi-dependent Heterochromatin Assembly at Centromeric Repeats Abstract In Schizosaccharomyces pombe, a fission yeast, large domains of heterochromatin are
Investigating the Role of RNA Polymerase II in RNAi-dependent Heterochromatin Assembly at Centromeric Repeats Abstract In Schizosaccharomyces pombe, a fission yeast, large domains of heterochromatin are
Heterochromatin revisited
 Heterochromatin revisited Shiv I. S. Grewal and Songtao Jia Abstract The formation of heterochromatin, which requires methylation of histone H3 at lysine 9 and the subsequent recruitment of chromodomain
Heterochromatin revisited Shiv I. S. Grewal and Songtao Jia Abstract The formation of heterochromatin, which requires methylation of histone H3 at lysine 9 and the subsequent recruitment of chromodomain
Cbh1. Abp1 WCE WCE. Tf2-7. act1. Abp1. Cbh1 WCE WCE. Tf2-8. act WCE. Tf2-fragment1. act1 WCE
 a SPCC0.07c ars00 ars00 SPCC0.09 alg ars00 Centromere IRCL otrl imrr otrr imrl IRCR lys dh dg cnt dg dh azr c d 0 Chromosome III Position (K) LTR Tf7 LTR Tf LTR Tffragment LTRs LTR. 5. 5. 5,9 5,9 5,97
a SPCC0.07c ars00 ars00 SPCC0.09 alg ars00 Centromere IRCL otrl imrr otrr imrl IRCR lys dh dg cnt dg dh azr c d 0 Chromosome III Position (K) LTR Tf7 LTR Tf LTR Tffragment LTRs LTR. 5. 5. 5,9 5,9 5,97
Mediator Directs Co-transcriptional Heterochromatin Assembly by RNA Interference-Dependent and -Independent Pathways
 Mediator Directs Co-transcriptional Heterochromatin Assembly by RNA Interference-Dependent and -Independent Pathways Eriko Oya 1, Hiroaki Kato 2,3, Yuji Chikashige 4, Chihiro Tsutsumi 4, Yasushi Hiraoka
Mediator Directs Co-transcriptional Heterochromatin Assembly by RNA Interference-Dependent and -Independent Pathways Eriko Oya 1, Hiroaki Kato 2,3, Yuji Chikashige 4, Chihiro Tsutsumi 4, Yasushi Hiraoka
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/science.65/dc Supporting Online Material for RNA Elimination Machinery Targeting Meiotic mrnas Promotes Facultative Heterochromatin Formation Martin Zofall, Soichiro
www.sciencemag.org/cgi/content/full/science.65/dc Supporting Online Material for RNA Elimination Machinery Targeting Meiotic mrnas Promotes Facultative Heterochromatin Formation Martin Zofall, Soichiro
RNAi and Heterochromatin in Plants and Fission Yeast Prof. Rob Martienssen
 RNAi AND HETEROCHROMATIN IN PLANTS AND FISSION YEAST Rob Martienssen, PhD 1 Heterochromatic silencing (PEV) resembles transposon silencing B. McClintock, 1954 2 Transposons regulate genes via DNA methylation
RNAi AND HETEROCHROMATIN IN PLANTS AND FISSION YEAST Rob Martienssen, PhD 1 Heterochromatic silencing (PEV) resembles transposon silencing B. McClintock, 1954 2 Transposons regulate genes via DNA methylation
RITS acts in cis to promote RNA interference mediated transcriptional and post-transcriptional silencing
 00 Nature Publishing Group http://www.nature.com/naturegenetics RITS acts in cis to promote RNA interference mediated transcriptional and post-transcriptional silencing Ken-ichi Noma, Tomoyasu Sugiyama,
00 Nature Publishing Group http://www.nature.com/naturegenetics RITS acts in cis to promote RNA interference mediated transcriptional and post-transcriptional silencing Ken-ichi Noma, Tomoyasu Sugiyama,
SHREC, an Effector Complex for Heterochromatic Transcriptional Silencing
 SHREC, an Effector Complex for Heterochromatic Transcriptional Silencing Tomoyasu Sugiyama, 1 Hugh P. Cam, 1 Rie Sugiyama, 1 Ken-ichi Noma, 1 Martin Zofall, 1 Ryuji Kobayashi, 2 and Shiv I.S. Grewal 1,
SHREC, an Effector Complex for Heterochromatic Transcriptional Silencing Tomoyasu Sugiyama, 1 Hugh P. Cam, 1 Rie Sugiyama, 1 Ken-ichi Noma, 1 Martin Zofall, 1 Ryuji Kobayashi, 2 and Shiv I.S. Grewal 1,
Biologia Cellulare Molecolare Avanzata 2.2. Functional & regulatory genomics module
 Biologia Cellulare Molecolare Avanzata 2.2 Functional & regulatory genomics module 1. Complexity of eukaryotic genomes. 2. Basic concepts of gene transcription and regulation 3. Transcriptomes 4. Coding,
Biologia Cellulare Molecolare Avanzata 2.2 Functional & regulatory genomics module 1. Complexity of eukaryotic genomes. 2. Basic concepts of gene transcription and regulation 3. Transcriptomes 4. Coding,
Barrales_SuppFig_S1 N/S 5-FOA. no reporter WT N/S 5-FOA. imr::ura4 + lem2. RT-qPCR: subtelomeric genes. lem2. nur1 clr4. WT + empty vector.
 ad f + ad e ad + e6 ch + p cid + + cl r4 + ce ntlh dg / + N/S N/S.5 tell.4.3. clr4 5-FOA + empty vector + empty vector + nmt:lem TZ clr4 RT-qPCR: subtelomeric genes.5.4.3..5. 5 5 4 4 3 3 tell transcript
ad f + ad e ad + e6 ch + p cid + + cl r4 + ce ntlh dg / + N/S N/S.5 tell.4.3. clr4 5-FOA + empty vector + empty vector + nmt:lem TZ clr4 RT-qPCR: subtelomeric genes.5.4.3..5. 5 5 4 4 3 3 tell transcript
The fourth chromosome: targeting heterochromatin formation in Drosophila. Drosophila melanogaster chromosomes
 The fourth chromosome: targeting heterochromatin formation in Drosophila Drosophila melanogaster chromosomes modified from TS Painter, 1934, J. Hered 25: 465-476. 1 DNA packaging domains HATs Euchromatin
The fourth chromosome: targeting heterochromatin formation in Drosophila Drosophila melanogaster chromosomes modified from TS Painter, 1934, J. Hered 25: 465-476. 1 DNA packaging domains HATs Euchromatin
Epigenetics in. Saccharomyces cerevisiae. Chapter 4 2/4/14
 Epigenetics in Saccharomyces cerevisiae Chapter 4 2/4/14 The budding yeast - Saccharomyces cerevisiae The fission yeast - Schizosaccharomyces pombe The budding yeast, Saccharomyces cerevisiae and the fission
Epigenetics in Saccharomyces cerevisiae Chapter 4 2/4/14 The budding yeast - Saccharomyces cerevisiae The fission yeast - Schizosaccharomyces pombe The budding yeast, Saccharomyces cerevisiae and the fission
Value Correct Answer Feedback. Student Response. A. Dicer enzyme. complex. C. the Dicer-RISC complex D. none of the above
 1 RNA mediated interference is a post-transcriptional gene silencing mechanism Which component of the RNAi pathway have been implicated in cleavage of the target mrna? A Dicer enzyme B the RISC-siRNA complex
1 RNA mediated interference is a post-transcriptional gene silencing mechanism Which component of the RNAi pathway have been implicated in cleavage of the target mrna? A Dicer enzyme B the RISC-siRNA complex
Control of Eukaryotic Gene Expression (Learning Objectives)
 Control of Eukaryotic Gene Expression (Learning Objectives) 1. Compare and contrast chromatin and chromosome: composition, proteins involved and level of packing. Explain the structure and function of
Control of Eukaryotic Gene Expression (Learning Objectives) 1. Compare and contrast chromatin and chromosome: composition, proteins involved and level of packing. Explain the structure and function of
Supplementary Figure 1. Chromosome 3 is devoid of the telomere-proximal subtelomeric
 Supplementary Figure 1. Chromosome 3 is devoid of the telomere-proximal subtelomeric common sequences. (a) Schematic illustration of the telomere-proximal site of subtelomeres. The restriction map is based
Supplementary Figure 1. Chromosome 3 is devoid of the telomere-proximal subtelomeric common sequences. (a) Schematic illustration of the telomere-proximal site of subtelomeres. The restriction map is based
Toward unraveling biogenesis of Dicerindependent
 Dissertation zur Erlangung des Doktorgrades der Fakultät für Chemie und Pharmazie der Ludwig-Maximilians-Universität München Toward unraveling biogenesis of Dicerindependent prirnas and sirnas in Schizosaccharomyces
Dissertation zur Erlangung des Doktorgrades der Fakultät für Chemie und Pharmazie der Ludwig-Maximilians-Universität München Toward unraveling biogenesis of Dicerindependent prirnas and sirnas in Schizosaccharomyces
RNAi-mediated Heterochromatin Assembly in Fission Yeast
 RNAi-mediated Heterochromatin Assembly in Fission Yeast M. ZOFALL AND S.I.S. GREWAL Laboratory of Molecular Cell Biology, National Cancer Institute, National Institutes of Health, Bethesda, Maryland 20892
RNAi-mediated Heterochromatin Assembly in Fission Yeast M. ZOFALL AND S.I.S. GREWAL Laboratory of Molecular Cell Biology, National Cancer Institute, National Institutes of Health, Bethesda, Maryland 20892
Plant Molecular and Cellular Biology Lecture 9: Nuclear Genome Organization: Chromosome Structure, Chromatin, DNA Packaging, Mitosis Gary Peter
 Plant Molecular and Cellular Biology Lecture 9: Nuclear Genome Organization: Chromosome Structure, Chromatin, DNA Packaging, Mitosis Gary Peter 9/16/2008 1 Learning Objectives 1. List and explain how DNA
Plant Molecular and Cellular Biology Lecture 9: Nuclear Genome Organization: Chromosome Structure, Chromatin, DNA Packaging, Mitosis Gary Peter 9/16/2008 1 Learning Objectives 1. List and explain how DNA
The RNA Polymerase II-associated factor 1 complex represses small-rna-mediated heterochromatin formation and gene silencing
 The RNA Polymerase II-associated factor 1 complex represses small-rna-mediated heterochromatin formation and gene silencing INAUGURALDISSERTATION zur Erlangung der Würde eines Doktors der Philosophie vorgelegt
The RNA Polymerase II-associated factor 1 complex represses small-rna-mediated heterochromatin formation and gene silencing INAUGURALDISSERTATION zur Erlangung der Würde eines Doktors der Philosophie vorgelegt
The Cul4-Ddb1 Cdt2 Ubiquitin Ligase Inhibits Invasion of a Boundary-Associated Antisilencing Factor into Heterochromatin
 The Cul-Ddb Cdt Ubiquitin Ligase Inhibits Invasion of a Boundary-Associated Antisilencing Factor into Heterochromatin Sigurd Braun, Jennifer F. Garcia, Margot Rowley, Mathieu Rougemaille,, Smita Shankar,
The Cul-Ddb Cdt Ubiquitin Ligase Inhibits Invasion of a Boundary-Associated Antisilencing Factor into Heterochromatin Sigurd Braun, Jennifer F. Garcia, Margot Rowley, Mathieu Rougemaille,, Smita Shankar,
The results presented here suggest that the activity Sir2 serves different purposes at
 54 4 DISCUSSION The results presented here suggest that the activity Sir2 serves different purposes at telomeres and the HML locus on the one hand, and at rdna on the other hand: Whereas it controls the
54 4 DISCUSSION The results presented here suggest that the activity Sir2 serves different purposes at telomeres and the HML locus on the one hand, and at rdna on the other hand: Whereas it controls the
In any cell type, a large part of the genes are kept inactive (not expressed) by gene silencing
 Part 1.week 4 8. Chromatin 9. Epigenetics and imprintingi 10. Post-transcriptional control and proteomics 8.1 Nucleosomes 8.2 Covalent modification of histones 8.2 The histone code 8.3 Nucleosome transitions
Part 1.week 4 8. Chromatin 9. Epigenetics and imprintingi 10. Post-transcriptional control and proteomics 8.1 Nucleosomes 8.2 Covalent modification of histones 8.2 The histone code 8.3 Nucleosome transitions
Biology 105 First Exam
 September 26, 2003 Biology 105 First Exam 1. 2. 3. 4. 5. 6. 7. Total 1 1. (15 points) a. Draw the structure of the nucleotide monophosphate 5-Me cytosine. b. Draw the structure of the base analog azacytidine..
September 26, 2003 Biology 105 First Exam 1. 2. 3. 4. 5. 6. 7. Total 1 1. (15 points) a. Draw the structure of the nucleotide monophosphate 5-Me cytosine. b. Draw the structure of the base analog azacytidine..
Expanded View Figures
 MO reports Leo1 Paf1 promotes histone turnover Laia Sadeghi et al xpanded View Figures Figure V1. The transposon mutagenesis system. artoon of the Hermes transposon screen. Southern blot of the Hermes
MO reports Leo1 Paf1 promotes histone turnover Laia Sadeghi et al xpanded View Figures Figure V1. The transposon mutagenesis system. artoon of the Hermes transposon screen. Southern blot of the Hermes
Spt6 Is Required for Heterochromatic Silencing in the Fission Yeast Schizosaccharomyces pombe
 MOLECULAR AND CELLULAR BIOLOGY, Oct. 2011, p. 4193 4204 Vol. 31, No. 20 0270-7306/11/$12.00 doi:10.1128/mcb.05568-11 Copyright 2011, American Society for Microbiology. All Rights Reserved. Spt6 Is Required
MOLECULAR AND CELLULAR BIOLOGY, Oct. 2011, p. 4193 4204 Vol. 31, No. 20 0270-7306/11/$12.00 doi:10.1128/mcb.05568-11 Copyright 2011, American Society for Microbiology. All Rights Reserved. Spt6 Is Required
Determinants of Heterochromatic sirna Biogenesis and Function
 Article Determinants of Heterochromatic sirna Biogenesis and Function Ruby Yu, 1 Gloria Jih, 1 Nahid Iglesias, 1 and Danesh Moazed 1, * 1 Department of Cell Biology, Howard Hughes Medical Institute, Harvard
Article Determinants of Heterochromatic sirna Biogenesis and Function Ruby Yu, 1 Gloria Jih, 1 Nahid Iglesias, 1 and Danesh Moazed 1, * 1 Department of Cell Biology, Howard Hughes Medical Institute, Harvard
The heterochromatin protein Swi6/HP1 activates replication origins at the pericentromeric region and silent mating-type locus
 letters The heterochromatin protein Swi6/HP activates replication origins at the pericentromeric region and silent mating-type locus Makoto T. Hayashi, Tatsuro S. Takahashi, Takuro Nakagawa, Jun-ichi Nakayama
letters The heterochromatin protein Swi6/HP activates replication origins at the pericentromeric region and silent mating-type locus Makoto T. Hayashi, Tatsuro S. Takahashi, Takuro Nakagawa, Jun-ichi Nakayama
Asf1/HIRA Facilitate Global Histone Deacetylation and Associate with HP1 to Promote Nucleosome Occupancy at Heterochromatic Loci
 Article Asf1/HIRA Facilitate Global Histone Deacetylation and Associate with HP1 to Promote Nucleosome Occupancy at Heterochromatic Loci Kenichi Yamane, 1 Takeshi Mizuguchi, 1 Bowen Cui, 1 Martin Zofall,
Article Asf1/HIRA Facilitate Global Histone Deacetylation and Associate with HP1 to Promote Nucleosome Occupancy at Heterochromatic Loci Kenichi Yamane, 1 Takeshi Mizuguchi, 1 Bowen Cui, 1 Martin Zofall,
Title. Author(s) 大屋, 恵梨子. Issue Date DOI. Doc URL. Type. File Information. Study on Mediator as a regulator of heterochromatin
 Title Study on Mediator as a regulator of heterochromatin Author(s) 大屋, 恵梨子 Issue Date 2013-09-25 DOI 10.14943/doctoral.k11091 Doc URL http://hdl.handle.net/2115/53917 Type theses (doctoral) File Information
Title Study on Mediator as a regulator of heterochromatin Author(s) 大屋, 恵梨子 Issue Date 2013-09-25 DOI 10.14943/doctoral.k11091 Doc URL http://hdl.handle.net/2115/53917 Type theses (doctoral) File Information
Chromatin Remodeling Factors and DNA Replication
 Chromatin Remodeling Factors and DNA Replication Patrick Varga-Weisz Abstract Chromatin structures have to be precisely duplicated during DNA replication to maintain tissue-specific gene expression patterns
Chromatin Remodeling Factors and DNA Replication Patrick Varga-Weisz Abstract Chromatin structures have to be precisely duplicated during DNA replication to maintain tissue-specific gene expression patterns
Lecture 6. Chromosome Structure and Function in Mitosis
 Lecture 6 Chromosome Structure and Function in Mitosis Outline: Chromosome Organization and Function in Interphase Chromatin effects on replication Effects of nuclear organization on gene expression Chromosomes
Lecture 6 Chromosome Structure and Function in Mitosis Outline: Chromosome Organization and Function in Interphase Chromatin effects on replication Effects of nuclear organization on gene expression Chromosomes
The Mi-2 Homolog Mit1 Actively Positions Nucleosomes Within Heterochromatin to Suppress Transcription
 University of Tennessee Health Science Center UTHSC Digital Commons Theses and Dissertations (ETD) College of Graduate Health Sciences 5-2014 The Mi-2 Homolog Mit1 Actively Positions Nucleosomes Within
University of Tennessee Health Science Center UTHSC Digital Commons Theses and Dissertations (ETD) College of Graduate Health Sciences 5-2014 The Mi-2 Homolog Mit1 Actively Positions Nucleosomes Within
AP Biology. The BIG Questions. Chapter 19. Prokaryote vs. eukaryote genome. Prokaryote vs. eukaryote genome. Why turn genes on & off?
 The BIG Questions Chapter 19. Control of Eukaryotic Genome How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions?
The BIG Questions Chapter 19. Control of Eukaryotic Genome How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions?
ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System
 ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System Liyan Pang, Ph.D. Application Scientist 1 Topics to be Covered Introduction What is ChIP-qPCR? Challenges Facing Biological
ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System Liyan Pang, Ph.D. Application Scientist 1 Topics to be Covered Introduction What is ChIP-qPCR? Challenges Facing Biological
MODES OF TRANSCRIPTIONAL REGULATION
 MODES OF TRANSCRIPTIONAL REULATION RNA-mediated epigenetic regulation of gene expression Daniel Holoch and Danesh Moazed Abstract Diverse classes of RNA, ranging from small to long non-coding RNAs, have
MODES OF TRANSCRIPTIONAL REULATION RNA-mediated epigenetic regulation of gene expression Daniel Holoch and Danesh Moazed Abstract Diverse classes of RNA, ranging from small to long non-coding RNAs, have
Chromatin Structure and its Effects on Transcription
 Chromatin Structure and its Effects on Transcription Epigenetics 2014 by Nigel Atkinson The University of Texas at Austin From Weaver 4th edition and Armstrong 1st edition What is the point? DNA is not
Chromatin Structure and its Effects on Transcription Epigenetics 2014 by Nigel Atkinson The University of Texas at Austin From Weaver 4th edition and Armstrong 1st edition What is the point? DNA is not
HP1 Proteins Form Distinct Complexes and Mediate Heterochromatic Gene Silencing by Nonoverlapping Mechanisms
 Article HP1 Proteins Form Distinct Complexes and Mediate Heterochromatic Gene Silencing by Nonoverlapping Mechanisms Mohammad R. Motamedi, 1,2 Eun-Jin Erica Hong, 1,2 Xue Li, 2,3 Scott Gerber, 2,4 Carilee
Article HP1 Proteins Form Distinct Complexes and Mediate Heterochromatic Gene Silencing by Nonoverlapping Mechanisms Mohammad R. Motamedi, 1,2 Eun-Jin Erica Hong, 1,2 Xue Li, 2,3 Scott Gerber, 2,4 Carilee
Transcription Regulation And Gene Expression in Eukaryotes (Cycle G2# )
 Transcription Regulation And Gene Expression in Eukaryotes (Cycle G2#13709-01) SMALL RNA REGULATORS OF GENE EXPRESSION RG. Clerc May 05, 2010 www.fmi.ch/training/teaching RNAi for RNA interference : discovered
Transcription Regulation And Gene Expression in Eukaryotes (Cycle G2#13709-01) SMALL RNA REGULATORS OF GENE EXPRESSION RG. Clerc May 05, 2010 www.fmi.ch/training/teaching RNAi for RNA interference : discovered
Lecture 9. Eukaryotic gene regulation: DNA METHYLATION
 Lecture 9 Eukaryotic gene regulation: DNA METHYLATION Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation sequences Transcriptional
Lecture 9 Eukaryotic gene regulation: DNA METHYLATION Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation sequences Transcriptional
Measuring Protein-DNA interactions
 Measuring Protein-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Transcription Factors are genetic switches 3 Regulation of Gene Expression by Transcription
Measuring Protein-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Transcription Factors are genetic switches 3 Regulation of Gene Expression by Transcription
NGS Approaches to Epigenomics
 I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
Gene expression analysis. Biosciences 741: Genomics Fall, 2013 Week 5. Gene expression analysis
 Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Chapter 19 Genetic Regulation of the Eukaryotic Genome. A. Bergeron AP Biology PCHS
 Chapter 19 Genetic Regulation of the Eukaryotic Genome A. Bergeron AP Biology PCHS 2 Do Now - Eukaryotic Transcription Regulation The diagram below shows five genes (with their enhancers) from the genome
Chapter 19 Genetic Regulation of the Eukaryotic Genome A. Bergeron AP Biology PCHS 2 Do Now - Eukaryotic Transcription Regulation The diagram below shows five genes (with their enhancers) from the genome
8.1 Nucleosomes 8.2 Covalent modification of histones 8.3 The histone code 8.4 Decoding proteins 8.5 DNA CpG methylation
 8th / 9th Chromatin - silencing epigenetics - imprinting 8.1 Nucleosomes 8.2 Covalent modification of histones 8.3 The histone code 8.4 Decoding proteins 8.5 DNA CpG methylation In any cell type, a large
8th / 9th Chromatin - silencing epigenetics - imprinting 8.1 Nucleosomes 8.2 Covalent modification of histones 8.3 The histone code 8.4 Decoding proteins 8.5 DNA CpG methylation In any cell type, a large
Sir2 is Required for Clr4 to Initiate Centromeric Heterochromatin Assembly in Fission Yeast
 Sacred Heart University DigitalCommons@SHU Chemistry & Physics Faculty Publications Chemistry and Physics 13 Sir is Required for Clr4 to Initiate Centromeric Heterochromatin Assembly in Fission Yeast Benjamin
Sacred Heart University DigitalCommons@SHU Chemistry & Physics Faculty Publications Chemistry and Physics 13 Sir is Required for Clr4 to Initiate Centromeric Heterochromatin Assembly in Fission Yeast Benjamin
sirna-mediated Heterochromatin Establishment Requires HP1 and Is Associated with Antisense Transcription
 Article sirna-mediated Heterochromatin Establishment Requires HP1 and Is Associated with Antisense Transcription Tetsushi Iida, 1 Jun-ichi Nakayama, 2 and Danesh Moazed 1, * 1 Department of Cell Biology,
Article sirna-mediated Heterochromatin Establishment Requires HP1 and Is Associated with Antisense Transcription Tetsushi Iida, 1 Jun-ichi Nakayama, 2 and Danesh Moazed 1, * 1 Department of Cell Biology,
Reprogramming of the Genome by Toxic Injury Ken Ramos, MD, PhD, ATS
 Reprogramming of the Genome by Toxic Injury Ken Ramos, MD, PhD, ATS Professor of dicine, University of Arizona Health Sciences Center & Associate Vice President for Precision Health Sciences Office of
Reprogramming of the Genome by Toxic Injury Ken Ramos, MD, PhD, ATS Professor of dicine, University of Arizona Health Sciences Center & Associate Vice President for Precision Health Sciences Office of
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali
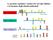 Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
Chapter 13. The Nucleus. The nucleus is the hallmark of eukaryotic cells; the very term eukaryotic means having a "true nucleus".
 Chapter 13 The Nucleus The nucleus is the hallmark of eukaryotic cells; the very term eukaryotic means having a "true nucleus". Fig.13.1. The EM of the Nucleus of a Eukaryotic Cell 13.1. The Nuclear Envelope
Chapter 13 The Nucleus The nucleus is the hallmark of eukaryotic cells; the very term eukaryotic means having a "true nucleus". Fig.13.1. The EM of the Nucleus of a Eukaryotic Cell 13.1. The Nuclear Envelope
Supplementary Materials
 Supplementary Materials Construction of Synthetic Nucleoli in Human Cells Reveals How a Major Functional Nuclear Domain is Formed and Propagated Through Cell Divisision Authors: Alice Grob, Christine Colleran
Supplementary Materials Construction of Synthetic Nucleoli in Human Cells Reveals How a Major Functional Nuclear Domain is Formed and Propagated Through Cell Divisision Authors: Alice Grob, Christine Colleran
Einführung in die Genetik
 Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
H3K9me-Independent Gene Silencing in Fission Yeast Heterochromatin by Clr5 and Histone Deacetylases
 H3K9me-Independent Gene Silencing in Fission Yeast Heterochromatin by Clr5 and Histone Deacetylases Klavs R. Hansen 1,2, Idit Hazan 3, Sreenath Shanker 4, Stephen Watt 5,6, Janne Verhein-Hansen 1,Jürg
H3K9me-Independent Gene Silencing in Fission Yeast Heterochromatin by Clr5 and Histone Deacetylases Klavs R. Hansen 1,2, Idit Hazan 3, Sreenath Shanker 4, Stephen Watt 5,6, Janne Verhein-Hansen 1,Jürg
Einführung in die Genetik
 Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Twitter: @PlantDevTUM, #genetiktum FB: Plant Development TUM Prof. Dr. Claus Schwechheimer
Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Twitter: @PlantDevTUM, #genetiktum FB: Plant Development TUM Prof. Dr. Claus Schwechheimer
Control of heterochromatin localization and silencing by the nuclear membrane protein Lem2
 Control of heterochromatin localization and silencing by the nuclear membrane protein Lem Ramón Ramos Barrales, Marta Forn,,3 Paula Raluca Georgescu,,3 Zsuzsa Sarkadi, and Sigurd Braun, Department of Physiological
Control of heterochromatin localization and silencing by the nuclear membrane protein Lem Ramón Ramos Barrales, Marta Forn,,3 Paula Raluca Georgescu,,3 Zsuzsa Sarkadi, and Sigurd Braun, Department of Physiological
Chapter 17 Lecture. Concepts of Genetics. Tenth Edition. Regulation of Gene Expression in Eukaryotes
 Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Nature Genetics: doi: /ng Supplementary Figure 1. ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets.
 Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
2/10/17. Contents. Applications of HMMs in Epigenomics
 2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
Gene Regulation in Eukaryotes. Dr. Syahril Abdullah Medical Genetics Laboratory
 Gene Regulation in Eukaryotes Dr. Syahril Abdullah Medical Genetics Laboratory syahril@medic.upm.edu.my Lecture Outline 1. The Genome 2. Overview of Gene Control 3. Cellular Differentiation in Higher Eukaryotes
Gene Regulation in Eukaryotes Dr. Syahril Abdullah Medical Genetics Laboratory syahril@medic.upm.edu.my Lecture Outline 1. The Genome 2. Overview of Gene Control 3. Cellular Differentiation in Higher Eukaryotes
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation
 Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation Mario Halic 1 and Danesh Moazed 1,2, * 1 Department of ell Biology 2 Howard Hughes Medical Institute Harvard Medical School, Boston,
Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation Mario Halic 1 and Danesh Moazed 1,2, * 1 Department of ell Biology 2 Howard Hughes Medical Institute Harvard Medical School, Boston,
Chromatin and Transcription
 Chromatin and Transcription Chromatin Structure Chromatin Represses Transcription Nucleosome Positioning Histone Acetylation Chromatin Remodeling Histone Methylation CHIP Analysis Chromatin and Elongation
Chromatin and Transcription Chromatin Structure Chromatin Represses Transcription Nucleosome Positioning Histone Acetylation Chromatin Remodeling Histone Methylation CHIP Analysis Chromatin and Elongation
Supplemental Figure legends Figure S1. (A) (B) (C) (D) Figure S2. Figure S3. (A-E) Figure S4. Figure S5. (A, C, E, G, I) (B, D, F, H, Figure S6.
 Supplemental Figure legends Figure S1. Map-based cloning and complementation testing for ZOP1. (A) ZOP1 was mapped to a ~273-kb interval on Chromosome 1. In the interval, a single-nucleotide G to A substitution
Supplemental Figure legends Figure S1. Map-based cloning and complementation testing for ZOP1. (A) ZOP1 was mapped to a ~273-kb interval on Chromosome 1. In the interval, a single-nucleotide G to A substitution
Transcription and RNAi in heterochromatic gene silencing
 CHROMATIN REVIEW Transcription and RNAi in heterochromatic gene silencing Marc Bühler & Danesh Moazed Recent findings have challenged the longstanding belief that heterochromatin is an inert and transcriptionally
CHROMATIN REVIEW Transcription and RNAi in heterochromatic gene silencing Marc Bühler & Danesh Moazed Recent findings have challenged the longstanding belief that heterochromatin is an inert and transcriptionally
Lecture 21: Epigenetics Nurture or Nature? Chromatin DNA methylation Histone Code Twin study X-chromosome inactivation Environemnt and epigenetics
 Lecture 21: Epigenetics Nurture or Nature? Chromatin DNA methylation Histone Code Twin study X-chromosome inactivation Environemnt and epigenetics Epigenetics represents the science for the studying heritable
Lecture 21: Epigenetics Nurture or Nature? Chromatin DNA methylation Histone Code Twin study X-chromosome inactivation Environemnt and epigenetics Epigenetics represents the science for the studying heritable
H3K36me3 polyclonal antibody
 H3K36me3 polyclonal antibody Cat. No. C15410192 Type: Polyclonal ChIP-grade/ChIP-seq grade Source: Rabbit Lot #: A1845P Size: 50 µg/32 µl Concentration: 1.6 μg/μl Specificity: Human, mouse, Arabidopsis,
H3K36me3 polyclonal antibody Cat. No. C15410192 Type: Polyclonal ChIP-grade/ChIP-seq grade Source: Rabbit Lot #: A1845P Size: 50 µg/32 µl Concentration: 1.6 μg/μl Specificity: Human, mouse, Arabidopsis,
3.1 The Role of the Enzymatic Activity of Sir2 for Efficient. The Sir proteins are recruited to DNA by site-specific DNA-binding proteins and
 35 3 RESULTS 3.1 The Role of the Enzymatic Activity of Sir2 for Efficient Association of the SIR Complex with DNA The Sir proteins are recruited to DNA by site-specific DNA-binding proteins and subsequently
35 3 RESULTS 3.1 The Role of the Enzymatic Activity of Sir2 for Efficient Association of the SIR Complex with DNA The Sir proteins are recruited to DNA by site-specific DNA-binding proteins and subsequently
Bioinformatics of Transcriptional Regulation
 Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Control of Gene Expression
 Control of Gene Expression Different cell types of a organism contain the same DNA but the DNA is expressed differently. External signals can cause a cell to change the expression of its genes. DNA elements
Control of Gene Expression Different cell types of a organism contain the same DNA but the DNA is expressed differently. External signals can cause a cell to change the expression of its genes. DNA elements
CELL BIOLOGY - CLUTCH CH. 7 - GENE EXPRESSION.
 !! www.clutchprep.com CONCEPT: CONTROL OF GENE EXPRESSION BASICS Gene expression is the process through which cells selectively to express some genes and not others Every cell in an organism is a clone
!! www.clutchprep.com CONCEPT: CONTROL OF GENE EXPRESSION BASICS Gene expression is the process through which cells selectively to express some genes and not others Every cell in an organism is a clone
Section C: The Control of Gene Expression
 Section C: The Control of Gene Expression 1. Each cell of a multicellular eukaryote expresses only a small fraction of its genes 2. The control of gene expression can occur at any step in the pathway from
Section C: The Control of Gene Expression 1. Each cell of a multicellular eukaryote expresses only a small fraction of its genes 2. The control of gene expression can occur at any step in the pathway from
Differential Gene Expression
 Biology 4361 - Developmental Biology Differential Gene Expression June 18, 2009 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Biology 4361 - Developmental Biology Differential Gene Expression June 18, 2009 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Tethering RITS to a Nascent Transcript Initiates RNAi- and Heterochromatin- Dependent Gene Silencing
 Tethering RITS to a Nascent Transcript Initiates RNAi- and Heterochromatin- Dependent Gene Silencing Marc Bühler, 1 André Verdel, 1 and Danesh Moazed 1, * 1 Department of Cell Biology, Harvard Medical
Tethering RITS to a Nascent Transcript Initiates RNAi- and Heterochromatin- Dependent Gene Silencing Marc Bühler, 1 André Verdel, 1 and Danesh Moazed 1, * 1 Department of Cell Biology, Harvard Medical
Control of Eukaryotic Genes. AP Biology
 Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Gene Regulation Biology
 Gene Regulation Biology Potential and Limitations of Cell Re-programming in Cancer Research Eric Blanc KCL April 13, 2010 Eric Blanc (KCL) Gene Regulation Biology April 13, 2010 1 / 21 Outline 1 The Central
Gene Regulation Biology Potential and Limitations of Cell Re-programming in Cancer Research Eric Blanc KCL April 13, 2010 Eric Blanc (KCL) Gene Regulation Biology April 13, 2010 1 / 21 Outline 1 The Central
A Genetic Screen to Identify Mammalian Chromatin Modifiers In Vivo.
 Gus Frangou, Stephanie Palmer & Mark Groudine Fred Hutchinson Cancer Research Center, Seattle- USA A Genetic Screen to Identify Mammalian Chromatin Modifiers In Vivo. During mammalian development and differentiation
Gus Frangou, Stephanie Palmer & Mark Groudine Fred Hutchinson Cancer Research Center, Seattle- USA A Genetic Screen to Identify Mammalian Chromatin Modifiers In Vivo. During mammalian development and differentiation
Local and Genomic Determinants of sirna-mediated Heterochromatin Formation and Silencing in Fission Yeast
 Local and Genomic Determinants of sirna-mediated Heterochromatin Formation and Silencing in Fission Yeast The Harvard community has made this article openly available. Please share how this access benefits
Local and Genomic Determinants of sirna-mediated Heterochromatin Formation and Silencing in Fission Yeast The Harvard community has made this article openly available. Please share how this access benefits
V23: the histone code
 V23: the histone code The DNA of eukaryotic organisms is packaged into chromatin, whose basic repeating unit is the nucleosome. A nucleosome is formed by wrapping 147 base pairs of DNA twice around an
V23: the histone code The DNA of eukaryotic organisms is packaged into chromatin, whose basic repeating unit is the nucleosome. A nucleosome is formed by wrapping 147 base pairs of DNA twice around an
DNA:CHROMATIN INTERACTIONS
 DNA:CHROMATIN INTERACTIONS Exploring transcription factor binding and the epigenomic landscape Chris Seward Introductions Cell and Developmental Biology PhD Candidate in Dr. Lisa Stubbs Laboratory Currently
DNA:CHROMATIN INTERACTIONS Exploring transcription factor binding and the epigenomic landscape Chris Seward Introductions Cell and Developmental Biology PhD Candidate in Dr. Lisa Stubbs Laboratory Currently
elements surrounding centromeres, and with the silent mating-type loci (1). Assembly of heterochromatin at these loci involves an orchestrated
 Fig. 4. Mutations in components of the RNAi system result in a loss of histone H3-mK9, and a delocalization of heterochromatin proteins HP1 and HP2. Polytene chromosomes (prepared as in Fig. 3) were treated
Fig. 4. Mutations in components of the RNAi system result in a loss of histone H3-mK9, and a delocalization of heterochromatin proteins HP1 and HP2. Polytene chromosomes (prepared as in Fig. 3) were treated
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3209 Supplementary Figure 1 IR induces the association of FH with chromatin. a, U2OS cells synchronized by thymidine double block (2 mm) underwent no release (G1 phase) or release for 2
DOI: 10.1038/ncb3209 Supplementary Figure 1 IR induces the association of FH with chromatin. a, U2OS cells synchronized by thymidine double block (2 mm) underwent no release (G1 phase) or release for 2
RNA and epigenetic silencing: Insight from fission yeast
 The Japanese Society of Developmental Biologists Develop. Growth Differ. (2012) 54, 129 141 doi: 10.1111/j.1440-169X.2011.01310.x Review Article RNA and epigenetic silencing: Insight from fission yeast
The Japanese Society of Developmental Biologists Develop. Growth Differ. (2012) 54, 129 141 doi: 10.1111/j.1440-169X.2011.01310.x Review Article RNA and epigenetic silencing: Insight from fission yeast
RNA folding and its importance. Mitesh Shrestha
 RNA folding and its importance Mitesh Shrestha Diseases Caused due to Protein Misfolding Alzheimer s Disease Parkinson s Disease Cataracts Sickle Cell Disease Prion Diseases Cystic Fibrosis Ribozymes Ribonucleic
RNA folding and its importance Mitesh Shrestha Diseases Caused due to Protein Misfolding Alzheimer s Disease Parkinson s Disease Cataracts Sickle Cell Disease Prion Diseases Cystic Fibrosis Ribozymes Ribonucleic
Controlling chromatin structure
 Chapter 23 1 Controlling chromatin structure 23.1 Introduction W hen transcription is treated in terms of interactions involving DNA and individual transcription factors and RNA polymerases, we get an
Chapter 23 1 Controlling chromatin structure 23.1 Introduction W hen transcription is treated in terms of interactions involving DNA and individual transcription factors and RNA polymerases, we get an
The contradictory definitions of heterochromatin: transcription and silencing
 Chromosoma (2006) 115: 110 122 DOI 10.1007/s00412-006-0052-x MINI-REVIEW Kathryn L. Huisinga. Brent Brower-Toland. Sarah C. R. Elgin The contradictory definitions of heterochromatin: transcription and
Chromosoma (2006) 115: 110 122 DOI 10.1007/s00412-006-0052-x MINI-REVIEW Kathryn L. Huisinga. Brent Brower-Toland. Sarah C. R. Elgin The contradictory definitions of heterochromatin: transcription and
The Plant Cell, March 2013, American Society of Plant Biologists. All rights reserved.
 Epigenetics (TTPB4) Teaching Guide Overview Epigenetic modifications of DNA and chromatin affect the activity of genes and transposons. Epigenetic controls affect processes as diverse as time-of-flowering,
Epigenetics (TTPB4) Teaching Guide Overview Epigenetic modifications of DNA and chromatin affect the activity of genes and transposons. Epigenetic controls affect processes as diverse as time-of-flowering,
2. Outline the levels of DNA packing in the eukaryotic nucleus below next to the diagram provided.
 AP Biology Reading Packet 6- Molecular Genetics Part 2 Name Chapter 19: Eukaryotic Genomes 1. Define the following terms: a. Euchromatin b. Heterochromatin c. Nucleosome 2. Outline the levels of DNA packing
AP Biology Reading Packet 6- Molecular Genetics Part 2 Name Chapter 19: Eukaryotic Genomes 1. Define the following terms: a. Euchromatin b. Heterochromatin c. Nucleosome 2. Outline the levels of DNA packing
Molecular Biology (BIOL 4320) Exam #1 March 12, 2002
 Molecular Biology (BIOL 4320) Exam #1 March 12, 2002 Name KEY SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number.
Molecular Biology (BIOL 4320) Exam #1 March 12, 2002 Name KEY SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number.
Epigenetic Regulation of Condensin-Mediated Genome Organization during the Cell Cycle and upon DNA Damage through Histone H3 Lysine 56 Acetylation
 Article Epigenetic Regulation of Condensin-Mediated Genome Organization during the Cell Cycle and upon DNA Damage through Histone H3 Lysine 5 Acetylation Atsunari Tanaka, Hideki Tanizawa, Sira Sriswasdi,
Article Epigenetic Regulation of Condensin-Mediated Genome Organization during the Cell Cycle and upon DNA Damage through Histone H3 Lysine 5 Acetylation Atsunari Tanaka, Hideki Tanizawa, Sira Sriswasdi,
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Gene Regulation. Sonja J. Prohaska WS 2017/18. Computational EvoDevo, University of Leipzig Santa Fe Institute, Santa Fe, NM, U.S.
 Computational EvoDevo, University of Leipzig Santa Fe Institute, Santa Fe, NM, U.S. WS 2017/18 The structural definition: What does that mean? used by the molecular biology community: Chemical modification
Computational EvoDevo, University of Leipzig Santa Fe Institute, Santa Fe, NM, U.S. WS 2017/18 The structural definition: What does that mean? used by the molecular biology community: Chemical modification
RNAi dependent heterochromatin assembly in fission yeast Schizosaccharomyces pombe requires heat shock molecular chaperones Hsp90 and Mas5
 https://doi.org/10.1186/s13072-018-0199-8 Epigenetics & Chromatin RESEARCH Open Access RNAi dependent heterochromatin assembly in fission yeast Schizosaccharomyces pombe requires heat shock molecular chaperones
https://doi.org/10.1186/s13072-018-0199-8 Epigenetics & Chromatin RESEARCH Open Access RNAi dependent heterochromatin assembly in fission yeast Schizosaccharomyces pombe requires heat shock molecular chaperones
Methoden zur Analyse von Transkriptionsfaktoren. Seminar: BCII, Lausen
 Methoden zur Analyse von Transkriptionsfaktoren Seminar: BCII, Lausen Gene expression: from transcription to translation Orphanides G, Reinberg D.Cell. 2002 Feb 22;108(4):439-51. Schematic of a gene regulatory
Methoden zur Analyse von Transkriptionsfaktoren Seminar: BCII, Lausen Gene expression: from transcription to translation Orphanides G, Reinberg D.Cell. 2002 Feb 22;108(4):439-51. Schematic of a gene regulatory
EUKARYOTIC GENE CONTROL
 EUKARYOTIC GENE CONTROL THE BIG QUESTIONS How are genes turned on and off? How do cells with the same DNA/ genes differentiate to perform completely different and specialized functions? GENE EXPRESSION
EUKARYOTIC GENE CONTROL THE BIG QUESTIONS How are genes turned on and off? How do cells with the same DNA/ genes differentiate to perform completely different and specialized functions? GENE EXPRESSION
EMBO. The JmjC domain protein Epe1 prevents unregulated assembly and disassembly of heterochromatin EMBO. open
 The EMBO Journal (2007) 26, 4670 4682 & 2007 European Molecular Biology Organization Some Rights Reserved 0261-4189/07 www.embojournal.org The JmjC domain protein Epe1 prevents unregulated assembly and
The EMBO Journal (2007) 26, 4670 4682 & 2007 European Molecular Biology Organization Some Rights Reserved 0261-4189/07 www.embojournal.org The JmjC domain protein Epe1 prevents unregulated assembly and
THE ANALYSIS OF CHROMATIN CONDENSATION STATE AND TRANSCRIPTIONAL ACTIVITY USING DNA MICROARRAYS 1. INTRODUCTION
 JOURNAL OF MEDICAL INFORMATICS & TECHNOLOGIES Vol.6/2003, ISSN 1642-6037 Piotr WIDŁAK *, Krzysztof FUJAREWICZ ** DNA microarrays, chromatin, transcription THE ANALYSIS OF CHROMATIN CONDENSATION STATE AND
JOURNAL OF MEDICAL INFORMATICS & TECHNOLOGIES Vol.6/2003, ISSN 1642-6037 Piotr WIDŁAK *, Krzysztof FUJAREWICZ ** DNA microarrays, chromatin, transcription THE ANALYSIS OF CHROMATIN CONDENSATION STATE AND
Chapter 18. Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression 2007-2008 Control of Prokaryotic (Bacterial) Genes 2007- Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
Chapter 18 Regulation of Gene Expression 2007-2008 Control of Prokaryotic (Bacterial) Genes 2007- Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06263 SUPPLEMENTARY INFORMATION Table of Contents: Supplementary Results Supplementary Discussion Supplementary References Supplementary Figures S1-S11 Supplementary Table 1-2. Supplementary
doi: 10.1038/nature06263 SUPPLEMENTARY INFORMATION Table of Contents: Supplementary Results Supplementary Discussion Supplementary References Supplementary Figures S1-S11 Supplementary Table 1-2. Supplementary
B. Incorrect! Centromeric DNA is largely heterochromatin, which is inactive DNA.
 MCAT Biology - Problem Drill 06: Molecular Biology of Eukaryotes Question No. 1 of 10 1. Which type of DNA would have the highest level of expression? Question #01 (A) Heterochromatin. (B) Centromeric
MCAT Biology - Problem Drill 06: Molecular Biology of Eukaryotes Question No. 1 of 10 1. Which type of DNA would have the highest level of expression? Question #01 (A) Heterochromatin. (B) Centromeric
