Supplementary Figure 2 Diagram of the expression vector pet28a with an individual gene of interest. Nature Methods: doi: /nmeth.
|
|
|
- Amber McDonald
- 6 years ago
- Views:
Transcription
1 Supplementary Figure 1 Microfluidic chip overview. (a) Photograph of a microfluidic chip bonded to a glass slide with a US dime for scale. Control channels are filled with food dye for better visualization. (b) Pattern of a typical DNA array, consisting of repeats of rows with four different genes and one row with nothing spotted as negative control. (c) Photograph of a bonded PDMS chip onto the glass slide with DNA spots in the back chamber. The orange highlighted frame shows a zoom in of the bottom left corner. (d) Typical fluorescence collage assembled from 640 single fluorescence micrographs of each protein chamber on one single chip shows pattern of expressed protein (assembly not to scale). Fluorescence signal of TagRFP reveals expression levels and Dockerin specificity. Here, low passivation of the protein chamber facilitates visualization. (e) Three of 640 adjacent dumbbell-shaped chambers, one with sfgfp DNA spotted (left), one with Xylanase DNA (center) and one negative control without DNA (right). Control channels are visualized with food dye: neck valve (green), sandwich valve (red), and button valve (blue). (f) Fluorescence images showing GFP signal (top) from expressed and immobilized ybbr-sfgfpdockerin (left), ybbr-xylanase-dockerin (center) with negative control lacking the spotted DNA (right). The bottom row shows the signal from the TagRFP detection construct, which specifically bound to the Dockerin tag via the Cohesin domain.
2 Supplementary Figure 2 Diagram of the expression vector pet28a with an individual gene of interest.
3 Supplementary Figure 3 Schematic of the fibronectin tetramer gene cassette.
4 Supplementary Figure 4 Schematic of the sfgfp dimer gene cassette.
5 Supplementary Figure 5 Schematic of the spectrin dimer gene cassette.
6 Supplementary Figure 6 Schematic of the xylanase gene cassette.
7 Supplementary Figure 7 Exemplary force traces Example curves showing (a) uninterpretable interaction, (b) non-specific interaction of cantilever with surface, (c) no interaction, and (d) a specific Xylanase-Dockerin unfolding and unbinding trace. Curves similar to those shown in a-c were excluded from the analysis.
8 Supplementary Discussion Typically in SMFS experiments, rupture force loading rate plots are used to characterize k off and Δx, the unbinding (or unfolding) probability per time unit and the distance to the transition state along the reaction coordinate, respectively, providing direct information about the energy landscape governing protein folding 1. SMFS experiments are also complemented by all-atom simulations of such systems in silico. Recently, it was shown that high speed SMFS experiments could be performed at speeds achievable in molecular dynamics simulations 2, overcoming a long standing discrepancy between experiment and simulation. In analyzing single-molecule unfolding curves (i.e., Fig. 2), we note that the spotted DNA at the measured array addresses correctly corresponded to the domain of interest encoded by the corresponding spotted DNA at that position. For example, the fibronectin tetramer was measured at array position (237), the spectrin dimer at position (239), the xylanase monomer at position (196), and the sfgfp monomer at position (238), corresponding to the correct genes deposited into the expression chambers at those array positions (Fig. 2). Typically immobilization chambers per microarray were measured. Typically several thousand force curves were acquired giving rise to dozens of interpretable single-molecule interaction curves. Upper force limit Here we extend the discussion regarding the upper force limit for the SMFS-MITOMI system. In all force-distance data traces, the last rupture events represent unbinding of the Coh-Doc complex, not unfolding of a domain. This rupture force of the Coh-Doc complex represents an upper limit in force for the entire construct, since the Doc is used as a handle sequence grabbed by the Coh-modified cantilver. The system we described can therefore interrogate domains with mechanical rupture forces that lie below that of Coh-Doc (~125 pn at 10 nn/s). If proteins with larger unfolding forces should be investigated, other Coh-Doc domains that show even higher complex rupture forces can be used. The Coh-Doc pair from R. flavefaciens, for example (PDB 4IU3) exhibits rupture forces over 600 pn at these loading rates (unpublished data). This could alternatively be used as a handle sequence to interrogate mechanically more stable domains of interest. Computerized image analysis can be used to automate cantilever positioning above the fluorescent rings and subsequent acquisition of unfolding traces at each array address in combination with online force curve analysis to further increase throughput. Additionally, well-characterized reference proteins on the same chip may serve as calibration standards further minimizing uncertainty in absolute force values. It is possible to operate the MITOMI device in a simplified way without the need for microspotting template DNA and chip alignment. This manual option should encourage the interested community to apply the suggested method to their single molecule force spectroscopy experiments. MITOMI enables the experimenter to prepare up to 16 different constructs in one column with 40 repeats each by flow-loading the DNA. Since
9 the valves are pressure sensitive it is also possible to operate these manually. This way it is possible to make use of the parallelized method without having the automation tools.supplementary Materials & Methods DNA Sequences Supplementary Table 1. Overview of primers Name Sequence 1 FW-w/o C-Tags MCS TAACTCGAGTAAGATCCGGCTGC 2 REV-N-Tags MCS GCTAGCACTAGTCCATGGGTG 3 FW-DocI GA AAAGTGGTACCTGGTACTCC 4 REV-XylDocI-GA CGGATCTTACTCGAGTTAGTTCTTGTACGGCAATGTATC 5 FW 10FNIII GA CGCACCGGCTCTGGCTCTGGCTCTGTTAGTGATGTTCCGCGTG 6 REV 10 FNIII GA GGAGTACCAGGTACCACTTTGGTGCG 7 REV 10FNIII (auf GS Li) GA ACTAACAGAGCCAGAGCCAGAGCCGGTGCGATAATTGATTGAAATC 8 FW sfgfp (auf MCS) GA CACCCATGGACTAGTGCTAGCAGCAAAGGTGAAGAACTGTTTAC 9 REV sfgfp (auf DocI) GA GGAGTACCAGGTACCACTTTCTTATACAGCTCATCCATACCATG Supplementary Table 2. Overview of DNA plasmids available at Addgene database Addgene ID Construct pet28a-ybbr-his-sfgfp-doci pet28a-ybbr-his-cbm-cohi pet28a-strepii-tagrfp-cohi pet28a-ybbr-his-xyl-doci pet28a-ybbr-his-10fniii(x4)-doci pet28a-ybbr-his-spec(x2)-doci
10 Multiple cloning site for the protein of interest: N terminal region T7 promoter lac operator RBS ATG ybbr Tag HRV 3C protease site HIS Tag (x6) TAATACGACTCACTATAGG GGAATTGTGAGCGGATAACAATTCC CCTGTAGAAATAATTTTGT TTAACTTTAAG AAGGA GATATACAT ATG GGTACC GACTCTCTGGAATTCATCGCTTCTAA ACTGGCT CTGGAAGTTCTGTTCCAGGGTCCG CTGCAG CACCACCACCACCACCAC CCATGG ACTAGTGCTAGC C terminal region Dockerin Type I T7 terminator AAAGTGGTACCTGGTACTCCTTCTACTAAATTATACGGCGACGTCAATGATGACGGAAAAGTTAA CTCAACTGACGCTGTAGCATTGAAGAGATATGTTTTGAGATCAGGTATAAGCATCAACACTGACA ATGCCGATTTGAATGAAGACGGCAGAGTTAATTCAACTGACTTAGGAATTTTGAAGAGATATATT CTCAAAGAAATAGATACATTGCCGTACAAGAAC TAA CTCGAGTAAGATCCGGCTGCTAACAAA GCCCGAAAGGAAGCTGAGTTGGCTGCTGCCACCGCTGAGCAATAA CTAGCATAACCCCTTGGGG CCTCTAAACGGGTCTTGAGGGGTTTTTT 10 FibronectinIII (4x): Glycin-Serin Linker (x6) GTTAGTGATGTTCCGCGTGATCTGGAAGTTGTTGCAGCAACCCCGACCAGCCTGCTGATTAGCTG GGATGCACCGGCAGTTACCGTTCGTTATTATCGTATTACCTATGGTGAAACCGGTGGTAATAGTC CGGTTCAAGAATTTACCGTTCCGGGTAGCAAAAGCACCGCAACCATTAGCGGTCTGAAACCGGGT GTTGATTACACCATTACCGTTTATGCCGTTACCGGTCGTGGTGATTCACCGGCAAGCAGCAAACC GATTAGCATTAACTATCGTACCGGTAGCGGTAGTGGTAGCGTTTCAGATGTGCCTCGCGACCTGG AAGTGGTGGCTGCCACACCGACCTCACTGCTGATCTCATGGGATGCCCCTGCCGTGACCGTGCGC TATTATCGCATCACATATGGCGAGACAGGTGGCAATTCACCTGTGCAAGAATTCACAGTTCCTGG TTCAAAAAGTACCGCCACAATTTCTGGCCTGAAACCTGGCGTGGATTACACAATCACAGTGTATG CAGTGACAGGTCGCGGTGATAGTCCGGCAAGTTCAAAACCGATTTCAATCAATTATCGCACCGGC TCTGGCTCTGGCTCTGTTAGTGATGTTCCGCGTGATCTGGAAGTTGTTGCAGCAACCCCGACCAG CCTGCTGATTAGCTGGGATGCACCGGCAGTTACCGTTCGTTATTATCGTATTACCTATGGTGAAA CCGGTGGTAATAGTCCGGTTCAAGAATTTACCGTTCCGGGTAGCAAAAGCACCGCAACCATTAGC GGTCTGAAACCGGGTGTTGATTACACCATTACCGTTTATGCCGTTACCGGTCGTGGTGATTCACC GGCAAGCAGCAAACCGATTAGCATTAACTATCGTACCGGTAGCGGTAGTGGTAGCGTTTCAGATG TGCCTCGCGACCTGGAAGTGGTGGCTGCCACACCGACCTCACTGCTGATCTCATGGGATGCCCCT GCCGTGACCGTGCGCTATTATCGCATCACATATGGCGAGACAGGTGGCAATTCACCTGTGCAAGA ATTCACAGTTCCTGGTTCAAAAAGTACCGCCACAATTTCTGGCCTGAAACCTGGCGTGGATTACA CAATCACAGTGTATGCAGTGACAGGTCGCGGTGATAGTCCGGCAAGTTCAAAACCGATTTCAATC AAttaTCGCACC
11 sfgfp: AGCAAAGGTGAAGAACTGTTTACCGGTGTTGTTCCGATTCTGGTTGAACTGGATGGTGATGTTAA TGGCCACAAATTTTCAGTTCGTGGTGAAGGCGAAGGTGATGCAACCATTGGTAAACTGACCCTGA AATTTATCTGTACCACCGGCAAACTGCCGGTTCCGTGGCCGACCCTGGTTACCACCCTGACCTAT GGTGTTCAGTGTTTTAGCCGTTATCCGGATCATATGAAACGCCACGATTTTTTCAAAAGCGCAAT GCCGGAAGGTTATGTTCAAGAACGTACCATCTCCTTTAAAGACGACGGTAAATACAAAACCCGTG CCGTTGTTAAATTTGAAGGTGATACCCTGGTGAATCGCATTGAACTGAAAGGCACCGATTTTAAA GAGGATGGTAATATCCTGGGCCACAAACTGGAATATAATTTCAATAGCCACAACGTGTATATCAC CGCAGACAAACAGAAAAATGGCATCAAAGCCAATTTTACCGTGCGCCATAATGTTGAAGATGGTA GCGTGCAGCTGGCAGATCATTATCAGCAGAATACCCCGATTGGTGATGGTCCGGTTCTGCTGCCG GATAATCATTATCTGAGCACCCAGACCGTTCTGAGCAAAGATCCGAATGAAAAACGTGATCATAT GGTGCTGCATGAGTATGTTAATGCAGCAGGTATTACCCATGGTATGGATGAGCTGTATAAG alpha-spectrin repeat 16 (chicken brain) (x2): Glycin-Serine Linker (x6) CGTGCTAAACTGAACGAATCTCACCGTCTGCACCAGTTCTTCCGTGACATGGACGACGAAGAATC TTGGATCAAAGAAAAAAAACTGCTGGTTTCTTCTGAAGACTACGGTCGTGACCTGACCGGTGTTC AGAACCTGCGTAAAAAACACAAACGTCTGGAAGCTGAACTGGCTGCTCACGAACCGGCTATCCAG GGTGTTCTGGACACCGGTAAAAAACTGTCTGACGACAACACCATCGGTAAAGAAGAAATCCAGCA GCGTCTGGCTCAGTTCGTTGACCACTGGAAAGAACTGAAACAGCTGGCTGCTGCTCGTGGTCAGC GTCTGGAAGAATCTCTGGAATACGGTAGCGGTAGCGGTTCACGTGCTAAACTGAACGAATCTCAC CGTCTGCACCAGTTCTTCCGTGACATGGACGACGAAGAATCTTGGATCAAAGAAAAAAAACTGCT GGTTTCTTCTGAAGACTACGGTCGTGACCTGACCGGTGTTCAGAACCTGCGTAAAAAACACAAAC GTCTGGAAGCTGAACTGGCTGCTCACGAACCGGCTATCCAGGGTGTTCTGGACACCGGTAAAAAA CTGTCTGACGACAACACCATCGGTAAAGAAGAAATCCAGCAGCGTCTGGCTCAGTTCGTTGACCA CTGGAAAGAACTGAAACAGCTGGCTGCTGCTCGTGGTCAGCGTCTGGAAGAATCTCTGGAATAt Xylanase: AAGAATGCAGATTCCTATGCGAAAAAACCTCACATCAGCGCATTGAATGCCCCACAATTGGATCA ACGCTACAAAAACGAGTTCACGATTGGTGCGGCAGTAGAACCTTATCAACTACAAAATGAAAAAG ACGTACAAATGCTAAAGCGCCACTTCAACAGCATTGTTGCCGAGAACGTAATGAAACCGATCAGC ATTCAACCTGAGGAAGGAAAATTCAATTTTGAACAAGCGGATCGAATTGTGAAGTTCGCTAAGGC AAATGGCATGGATATTCGCTTCCATACACTCGTTTGGCACAGCCAAGTACCTCAATGGTTCTTTC TTGACAAGGAAGGTAAGCCAATGGTTAATGAATGCGATCCAGTGAAACGTGAACAAAATAAACAA CTGCTGTTAAAACGACTTGAAACTCATATTAAAACGATCGTCGAGCGGTACAAAGATGACATTAA GTACTGGGACGTTGTAAATGAGGTTGTGGGGGACGACGGAAAACTGCGCAACTCTCCATGGTATC AAATCGCCGGCATCGATTATATTAAAGTGGCATTCCAAGCAGCTAGAAAATATGGCGGAGACAAC ATTAAGCTTTACATGAATGATTACAATACAGAAGTCGAACCGAAGCGAACCGCTCTTTACAATTT AGTCAAACAACTGAAAGAAGAGGGTGTTCCGATCGACGGCATCGGCCATCAATCCCACATCCAAA TCGGCTGGCCTTCTGAAGCAGAAATCGAGAAAACGATTAACATGTTCGCCGCTCTCGGTTTAGAC AACCAAATCACTGAGCTTGATGTGAGCATGTACGGTTGGCCGCCGCGCGCTTACCCGACGTATGA CGCCATTCCAAAACAAAAGTTTTTGGATCAGGCAGCGCGCTATGATCGTTTGTTCAAACTGTATG AAAAGTTGAGCGATAAAATTAGCAACGTCACCTTCTGGGGCATCGCCGACAATCATACGTGGCTC GACAGCCGTGCGGATGTGTACTATGACGCCAACGGGAATGTTGTGGTTGACCCGAACGCTCCGTA CGCAAAAGTGGAAAAAGGGAAAGGAAAAGATGCGCCGTTCGTTTTTGGACCGGATTACAAAGTCA AACCCGCATATTGGGCTATTATCGACCAC
12 Detection construct RFP-Cohesin: TagRFP-Cohesin: T7 promoter lac operator RBS ATG StrepII Tag TagRFP Linker Cohesin T7 terminator TAATACGACTCACTATAGG GGAATTGTGAGCGGATAACAATTCC CCTGTAGAAATAATTTTGT TTAACTTTAAG AAGGA GATATACAT ATG GGTACC TGGTCTCACCCGCAGTTCGAAAAA G TTTCTAAAGGTGAAGAACTGATCAAAGAAAACATGCACATGAAACTGTACATGGAAGGTACTGTT AACAACCACCACTTCAAATGCACCTCTGAAGGTGAAGGTAAACCGTACGAAGGTACTCAGACCAT GCGTATCAAAGTTGTTGAAGGTGGTCCGCTGCCGTTCGCTTTCGACATCCTGGCTACCTCTTTCA TGTACGGTTCTCGTACCTTCATCAACCACACCCAGGGTATCCCGGACTTCTTCAAACAGTCTTTC CCGGAAGGTTTCACCTGGGAACGTGTTACCACCTACGAAGACGGTGGTGTTCTGACCGCTACCCA GGACACCTCTCTGCAAGACGGTTGCCTGATCTACAACGTTAAAATCCGTGGTGTTAACTTCCCGT CTAACGGTCCGGTTATGCAGAAAAAAACCCTGGGTTGGGAAGCTAACACCGAAATGCTGTACCCG GCTGACGGTGGTCTGGAAGGTCGTTCTGACATGGCTCTGAAACTGGTTGGTGGTGGTCACCTGAT CTGCAACTTCAAAACCACCTACCGTTCTAAAAAACCGGCTAAAAACCTGAAAATGCCGGGTGTTT ACTACGTTGACCACCGTCTGGAACGTATCAAAGAAGCTGACAAAGAAACCTACGTTGAACAGCAC GAAGTTGCTGTTGCTCGTTACTGCGACCTGCCGTCTAAACTGGGTCACAAACTGAAC GGCAGTG TAGTACCATCAACACAGCCTGTAACAACACCACCTGCAACAACAAAACCACCTGCAACAACAATA CCGCCGTCAGATGATCCGAATGCA GGATCCGACGGTGTGGTAGTAGAAATTGGCAAAGTTACGG GATCTGTTGGAACTACAGTTGAAATACCTGTATATTTCAGAGGAGTTCCATCCAAAGGAATAGCA AACTGCGACTTTGTGTTCAGATATGATCCGAATGTATTGGAAATTATAGGGATAGATCCCGGAGA CATAATAGTTGACCCGAATCCTACCAAGAGCTTTGATACTGCAATATATCCTGACAGAAAGATAA TAGTATTCCTGTTTGCGGAAGACAGCGGAACAGGAGCGTATGCAATAACTAAAGACGGAGTATTT GCAAAAATAAGAGCAACTGTAAAATCAAGTGCTCCGGGCTATATTACTTTCGACGAAGTAGGTGG ATTTGCAGATAATGACCTGGTAGAACAGAAGGTATCATTTATAGACGGTGGTGTTAACGTTGGCA ATGCAACA TAA CTCGAGTAAGATCCGGCTGCTAACAAAGCCCGAAAGGAAGCTGAGTTGGCTG CTGCCACCGCTGAGCAATAA CTAGCATAACCCCTTGGGGCCTCTAAACGGGTCTTGAGGGGTTT TTT Molecular weights of synthesized fusion proteins ybbr-(fibronectin) 4 -Dockerin Type I: 53 kda ybbr-(spectrin) 2 -Dockerin Type I: 40 kda ybbr-xylanase-dockerin Type I: 56 kda ybbr-sfgfp-dockerin Type I: 39 kda ybbr-twitchin-dockerin Type I: 52 kda
13 Supplementary Table 3. Yield of interpretable curves Construct Interpretable Curves GFP 25 out of = 0.16 % Fibronectin 27 out of = 0.1 % Xylanase 91 out of 5553 = 1.64 % Spectrin 50 out of = 0.48% References 1. Merkel, R., Nassoy, P., Leung, A., Ritchie, K. & Evans, E. Energy landscapes of receptor ligand bonds explored with dynamic force spectroscopy. Nature 397, (1999). 2. Rico, F., Gonzalez, L., Casuso, I., Puig-Vidal, M. & Scheuring, S. High-Speed Force Spectroscopy Unfolds Titin at the Velocity of Molecular Dynamics Simulations. Science 342, (2013).
From genes to protein mechanics on a chip
 brief communications 2014 Nature America, Inc. All rights reserved. From genes to protein mechanics on a chip Marcus Otten 1,2,4, Wolfgang Ott 1,2,4, Markus A Jobst 1,2,4, Lukas F Milles 1,2, Tobias Verdorfer
brief communications 2014 Nature America, Inc. All rights reserved. From genes to protein mechanics on a chip Marcus Otten 1,2,4, Wolfgang Ott 1,2,4, Markus A Jobst 1,2,4, Lukas F Milles 1,2, Tobias Verdorfer
Supporting Information. Sequence Independent Cloning and Posttranslational. Enzymatic Ligation
 Supporting Information Sequence Independent Cloning and Posttranslational Modification of Repetitive Protein Polymers through Sortase and Sfp-mediated Enzymatic Ligation Wolfgang Ott,,, Thomas Nicolaus,,
Supporting Information Sequence Independent Cloning and Posttranslational Modification of Repetitive Protein Polymers through Sortase and Sfp-mediated Enzymatic Ligation Wolfgang Ott,,, Thomas Nicolaus,,
Enhancers mutations that make the original mutant phenotype more extreme. Suppressors mutations that make the original mutant phenotype less extreme
 Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
CAP BIOINFORMATICS Su-Shing Chen CISE. 10/5/2005 Su-Shing Chen, CISE 1
 CAP 5510-9 BIOINFORMATICS Su-Shing Chen CISE 10/5/2005 Su-Shing Chen, CISE 1 Basic BioTech Processes Hybridization PCR Southern blotting (spot or stain) 10/5/2005 Su-Shing Chen, CISE 2 10/5/2005 Su-Shing
CAP 5510-9 BIOINFORMATICS Su-Shing Chen CISE 10/5/2005 Su-Shing Chen, CISE 1 Basic BioTech Processes Hybridization PCR Southern blotting (spot or stain) 10/5/2005 Su-Shing Chen, CISE 2 10/5/2005 Su-Shing
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr.
 MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
SUPPLEMENTARY INFORMATION
 Nanomechanical mapping of first binding steps of a virus to animal cells David Alsteens, Richard Newton, Rajib Schubert, David Martinez-Martin, Martin Delguste, Botond Roska and Daniel J. Müller This PDF
Nanomechanical mapping of first binding steps of a virus to animal cells David Alsteens, Richard Newton, Rajib Schubert, David Martinez-Martin, Martin Delguste, Botond Roska and Daniel J. Müller This PDF
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
SUPPLEMENTARY INFORMATION
 Results Construct purification and coupling. Two A1-GP1bα ReaLiSM constructs, with and without cysteine residues near the N and C-termini (Fig. S2a), were expressed and purified by Ni affinity chromatography
Results Construct purification and coupling. Two A1-GP1bα ReaLiSM constructs, with and without cysteine residues near the N and C-termini (Fig. S2a), were expressed and purified by Ni affinity chromatography
Supplementary Note DNA Duplex Design and Construction
 Supplementary Note DNA Duplex Design and Construction Double-stranded DNA was amplified from a M13mp18 plasmid (Bayou Biolabs) by Autosticky PCR 1, which in our hands gave higher yields than asymmetric
Supplementary Note DNA Duplex Design and Construction Double-stranded DNA was amplified from a M13mp18 plasmid (Bayou Biolabs) by Autosticky PCR 1, which in our hands gave higher yields than asymmetric
Genome Analysis. Bacterial genome projects
 Genome Analysis Bacterial Genome sequencing does this help us in the investigation of adaptive responses/regulatory systems? Genome Sequencing Projects strategy & methods annotation Comparative genomics
Genome Analysis Bacterial Genome sequencing does this help us in the investigation of adaptive responses/regulatory systems? Genome Sequencing Projects strategy & methods annotation Comparative genomics
Chapter 6 - Molecular Genetic Techniques
 Chapter 6 - Molecular Genetic Techniques Two objects of molecular & genetic technologies For analysis For generation Molecular genetic technologies! For analysis DNA gel electrophoresis Southern blotting
Chapter 6 - Molecular Genetic Techniques Two objects of molecular & genetic technologies For analysis For generation Molecular genetic technologies! For analysis DNA gel electrophoresis Southern blotting
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
An Introduction to printing spotted microarrays
 http://www.flychip.org.uk/printingintroduction.php An Introduction to printing spotted microarrays Overview Microarrays are typically produced by transferring gene or transcript specific PCR amplified
http://www.flychip.org.uk/printingintroduction.php An Introduction to printing spotted microarrays Overview Microarrays are typically produced by transferring gene or transcript specific PCR amplified
Surface Plasmon Resonance Analyzer
 Surface Plasmon Resonance Analyzer 5 6 SPR System Based on Microfluidics Wide Dynamic Range Kinetic Analysis by Detection of Association /Dissociation of Bio-Molecules Measuring of Mass Change below
Surface Plasmon Resonance Analyzer 5 6 SPR System Based on Microfluidics Wide Dynamic Range Kinetic Analysis by Detection of Association /Dissociation of Bio-Molecules Measuring of Mass Change below
Chapter 14 Regulation of Transcription
 Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid
 Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
NPTEL VIDEO COURSE PROTEOMICS PROF. SANJEEVA SRIVASTAVA
 LECTURE-29 Cell-free synthesis based protein microarrays TRANSCRIPT Welcome to the proteomics course. In today s lecture we will talk about protein microarrays based on cell-free synthesis. So cell-free
LECTURE-29 Cell-free synthesis based protein microarrays TRANSCRIPT Welcome to the proteomics course. In today s lecture we will talk about protein microarrays based on cell-free synthesis. So cell-free
Recent technology allow production of microarrays composed of 70-mers (essentially a hybrid of the two techniques)
 Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Microarrays and Transcript Profiling Gene expression patterns are traditionally studied using Northern blots (DNA-RNA hybridization assays). This approach involves separation of total or polya + RNA on
Protocol for cloning SEC-based repair templates using Gibson assembly and ccdb negative selection
 Protocol for cloning SEC-based repair templates using Gibson assembly and ccdb negative selection Written by Dan Dickinson (daniel.dickinson@austin.utexas.edu) and last updated January 2018. A version
Protocol for cloning SEC-based repair templates using Gibson assembly and ccdb negative selection Written by Dan Dickinson (daniel.dickinson@austin.utexas.edu) and last updated January 2018. A version
Nature Structural and Molecular Biology: doi: /nsmb.2937
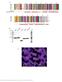 Supplementary Figure 1 Multiple sequence alignment of the CtIP N-terminal domain, purified CtIP protein constructs and details of the 2F o F c electron density map of CtIP-NTD. (a) Multiple sequence alignment,
Supplementary Figure 1 Multiple sequence alignment of the CtIP N-terminal domain, purified CtIP protein constructs and details of the 2F o F c electron density map of CtIP-NTD. (a) Multiple sequence alignment,
CSE : Computational Issues in Molecular Biology. Lecture 19. Spring 2004
 CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
Lecture 22 Eukaryotic Genes and Genomes III
 Lecture 22 Eukaryotic Genes and Genomes III In the last three lectures we have thought a lot about analyzing a regulatory system in S. cerevisiae, namely Gal regulation that involved a hand full of genes.
Lecture 22 Eukaryotic Genes and Genomes III In the last three lectures we have thought a lot about analyzing a regulatory system in S. cerevisiae, namely Gal regulation that involved a hand full of genes.
DNA Microarray Technology
 2 DNA Microarray Technology 2.1 Overview DNA microarrays are assays for quantifying the types and amounts of mrna transcripts present in a collection of cells. The number of mrna molecules derived from
2 DNA Microarray Technology 2.1 Overview DNA microarrays are assays for quantifying the types and amounts of mrna transcripts present in a collection of cells. The number of mrna molecules derived from
Cell-free expression based microarrays
 Dr. Sanjeeva Srivastava IIT Bombay Cell-free expression based microarrays Protein in situ array (PISA) Nucleic Acid Programmable Protein Array (NAPPA) Multiple Spotting Technique (MIST) DNA Array to Protein
Dr. Sanjeeva Srivastava IIT Bombay Cell-free expression based microarrays Protein in situ array (PISA) Nucleic Acid Programmable Protein Array (NAPPA) Multiple Spotting Technique (MIST) DNA Array to Protein
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 February 15, 2013 Multiple choice questions (numbers in brackets indicate the number of correct answers) 1. Which of the following statements are not true Transcriptomes consist of mrnas Proteomes consist
1 February 15, 2013 Multiple choice questions (numbers in brackets indicate the number of correct answers) 1. Which of the following statements are not true Transcriptomes consist of mrnas Proteomes consist
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than. SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with
 Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Chemical Biology In-class discussion: Tethering
 In-class discussion: Tethering This discussion will focus on various Tethering Methods described by workers at Sunesis (for a review, see Erlanson et al. Ann. Rev. Biophys. Biomol. Str. 2004). Regarding
In-class discussion: Tethering This discussion will focus on various Tethering Methods described by workers at Sunesis (for a review, see Erlanson et al. Ann. Rev. Biophys. Biomol. Str. 2004). Regarding
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10016 Supplementary discussion on binding site density for protein complexes on the surface: The density of biotin sites on the chip is ~10 3 biotin-peg per µm 2. The biotin sites are
doi:10.1038/nature10016 Supplementary discussion on binding site density for protein complexes on the surface: The density of biotin sites on the chip is ~10 3 biotin-peg per µm 2. The biotin sites are
BBF RFC 44 - Bioscaffold-Linker. Mina Petros & Savery Nigel. 18 October 2009
 BBF RFC 44 Bioscaffold-Linker BBF RFC 44 - Bioscaffold-Linker Mina Petros & Savery Nigel 18 October 2009 1. Purpose Protein fusions are currently not supported by most assembly standards. This assembly
BBF RFC 44 Bioscaffold-Linker BBF RFC 44 - Bioscaffold-Linker Mina Petros & Savery Nigel 18 October 2009 1. Purpose Protein fusions are currently not supported by most assembly standards. This assembly
Exam 2 BIO200, Winter 2012
 Exam 2 BIO200, Winter 2012 Name: Multiple Choice Questions: Circle the one best answer for each question. (2 points each) 1. The 5 cap structure is often described as a backwards G. What makes this nucleotide
Exam 2 BIO200, Winter 2012 Name: Multiple Choice Questions: Circle the one best answer for each question. (2 points each) 1. The 5 cap structure is often described as a backwards G. What makes this nucleotide
Introduction to Microarray Analysis
 Introduction to Microarray Analysis Methods Course: Gene Expression Data Analysis -Day One Rainer Spang Microarrays Highly parallel measurement devices for gene expression levels 1. How does the microarray
Introduction to Microarray Analysis Methods Course: Gene Expression Data Analysis -Day One Rainer Spang Microarrays Highly parallel measurement devices for gene expression levels 1. How does the microarray
Intercalation-based single-molecule fluorescence assay to study DNA supercoil
 Intercalation-based single-molecule fluorescence assay to study DNA supercoil dynamics Mahipal Ganji, Sung Hyun Kim, Jaco van der Torre, Elio Abbondanzieri*, Cees Dekker* Supplementary information Supplementary
Intercalation-based single-molecule fluorescence assay to study DNA supercoil dynamics Mahipal Ganji, Sung Hyun Kim, Jaco van der Torre, Elio Abbondanzieri*, Cees Dekker* Supplementary information Supplementary
DNA is normally found in pairs, held together by hydrogen bonds between the bases
 Bioinformatics Biology Review The genetic code is stored in DNA Deoxyribonucleic acid. DNA molecules are chains of four nucleotide bases Guanine, Thymine, Cytosine, Adenine DNA is normally found in pairs,
Bioinformatics Biology Review The genetic code is stored in DNA Deoxyribonucleic acid. DNA molecules are chains of four nucleotide bases Guanine, Thymine, Cytosine, Adenine DNA is normally found in pairs,
HEK293T. Fig. 1 in the
 Supplementary Information Supplementary Figure 1 Zinc uptake assay of hzip4 and hzip4-δecd transiently expressed in HEK293T cells. The results of one representative e experiment are shown in Fig. 1 in
Supplementary Information Supplementary Figure 1 Zinc uptake assay of hzip4 and hzip4-δecd transiently expressed in HEK293T cells. The results of one representative e experiment are shown in Fig. 1 in
PHYS 498 HW3 Solutions: 1. We have two equations: (1) (2)
 PHYS 498 HW3 Solutions: 1. We have two equations: (1) (2) Where Conc is the initial concentration of [B] or [SA] Since [B] = [SA], the second equation simplifies to: (3) Using equation (1) and (3), we
PHYS 498 HW3 Solutions: 1. We have two equations: (1) (2) Where Conc is the initial concentration of [B] or [SA] Since [B] = [SA], the second equation simplifies to: (3) Using equation (1) and (3), we
7.03, 2006, Lecture 23 Eukaryotic Genes and Genomes IV
 1 Fall 2006 7.03 7.03, 2006, Lecture 23 Eukaryotic Genes and Genomes IV In the last three lectures we have thought a lot about analyzing a regulatory system in S. cerevisiae, namely Gal regulation that
1 Fall 2006 7.03 7.03, 2006, Lecture 23 Eukaryotic Genes and Genomes IV In the last three lectures we have thought a lot about analyzing a regulatory system in S. cerevisiae, namely Gal regulation that
7.05, 2005, Lecture 23 Eukaryotic Genes and Genomes IV
 7.05, 2005, Lecture 23 Eukaryotic Genes and Genomes IV In the last three lectures we have thought a lot about analyzing a regulatory system in S. cerevisiae, namely Gal regulation that involved a hand
7.05, 2005, Lecture 23 Eukaryotic Genes and Genomes IV In the last three lectures we have thought a lot about analyzing a regulatory system in S. cerevisiae, namely Gal regulation that involved a hand
DYNAMICBIOSENSORS. Chip functionalization DNA-encoded addressing and chip configuration. Technology Information #120
 Technology Information #120 Chip functionalization DNA-encoded addressing and chip configuration The switchsense biochip features 24 detection spots, which are arranged in 2 or 4 seperate flow channels.
Technology Information #120 Chip functionalization DNA-encoded addressing and chip configuration The switchsense biochip features 24 detection spots, which are arranged in 2 or 4 seperate flow channels.
Clones. Glass Slide PCR. Purification. Array Printing. Post-process. Hybridize. Scan. Array Fabrication. Sample Preparation/Hybridization.
 Terminologies Reporter: the nucleotide sequence present in a particular location on the array (a.k.a. probe) Feature: the location of a reporter on the array Composite sequence: a set of reporters used
Terminologies Reporter: the nucleotide sequence present in a particular location on the array (a.k.a. probe) Feature: the location of a reporter on the array Composite sequence: a set of reporters used
TECHNICAL BULLETIN. In Vitro Bacterial Split Fluorescent Protein Fold n Glow Solubility Assay Kits
 In Vitro Bacterial Split Fluorescent Protein Fold n Glow Solubility Assay Kits Catalog Numbers APPA001 In Vitro Bacterial Split GFP "Fold 'n' Glow" Solubility Assay Kit (Green) APPA008 In Vitro Bacterial
In Vitro Bacterial Split Fluorescent Protein Fold n Glow Solubility Assay Kits Catalog Numbers APPA001 In Vitro Bacterial Split GFP "Fold 'n' Glow" Solubility Assay Kit (Green) APPA008 In Vitro Bacterial
Methods of Biomaterials Testing Lesson 3-5. Biochemical Methods - Molecular Biology -
 Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Design. Construction. Characterization
 Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Fabricating Microfluidic Devices for High-Density Biological Assays
 Fabricating Microfluidic Devices for High-Density Biological Assays Todd Thorsen Department of Mechanical Engineeering MIT Panamerican Advanced Studies Institute Micro-Electro-Mechanical Systems San Carlos
Fabricating Microfluidic Devices for High-Density Biological Assays Todd Thorsen Department of Mechanical Engineeering MIT Panamerican Advanced Studies Institute Micro-Electro-Mechanical Systems San Carlos
Molecular Cloning. Genomic DNA Library: Contains DNA fragments that represent an entire genome. cdna Library:
 Molecular Cloning Genomic DNA Library: Contains DNA fragments that represent an entire genome. cdna Library: Made from mrna, and represents only protein-coding genes expressed by a cell at a given time.
Molecular Cloning Genomic DNA Library: Contains DNA fragments that represent an entire genome. cdna Library: Made from mrna, and represents only protein-coding genes expressed by a cell at a given time.
3.1.4 DNA Microarray Technology
 3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
3.1.4 DNA Microarray Technology Scientists have discovered that one of the differences between healthy and cancer is which genes are turned on in each. Scientists can compare the gene expression patterns
Introduction to Bioinformatics
 Introduction to Bioinformatics If the 19 th century was the century of chemistry and 20 th century was the century of physic, the 21 st century promises to be the century of biology...professor Dr. Satoru
Introduction to Bioinformatics If the 19 th century was the century of chemistry and 20 th century was the century of physic, the 21 st century promises to be the century of biology...professor Dr. Satoru
Genetic Parts in Bacterial Gene Expression
 Genetic Parts in Bacterial Gene Expression The Focus Promoters Operators Transcription Factors Transcriptional Terminators Ribosome Binding Sites An Abstract Annotation We ll Pay Particular Attention To:
Genetic Parts in Bacterial Gene Expression The Focus Promoters Operators Transcription Factors Transcriptional Terminators Ribosome Binding Sites An Abstract Annotation We ll Pay Particular Attention To:
Supplementary Figures 1-6
 1 Supplementary Figures 1-6 2 3 4 5 6 7 Supplementary Fig. 1: GFP-KlpA forms a homodimer. a, Hydrodynamic analysis of the purified full-length GFP-KlpA protein. Fractions from size exclusion chromatography
1 Supplementary Figures 1-6 2 3 4 5 6 7 Supplementary Fig. 1: GFP-KlpA forms a homodimer. a, Hydrodynamic analysis of the purified full-length GFP-KlpA protein. Fractions from size exclusion chromatography
11/22/13. Proteomics, functional genomics, and systems biology. Biosciences 741: Genomics Fall, 2013 Week 11
 Proteomics, functional genomics, and systems biology Biosciences 741: Genomics Fall, 2013 Week 11 1 Figure 6.1 The future of genomics Functional Genomics The field of functional genomics represents the
Proteomics, functional genomics, and systems biology Biosciences 741: Genomics Fall, 2013 Week 11 1 Figure 6.1 The future of genomics Functional Genomics The field of functional genomics represents the
Genome Sequence Assembly
 Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Genome Sequence Assembly Learning Goals: Introduce the field of bioinformatics Familiarize the student with performing sequence alignments Understand the assembly process in genome sequencing Introduction:
Interpretation of sequence results
 Interpretation of sequence results An overview on DNA sequencing: DNA sequencing involves the determination of the sequence of nucleotides in a sample of DNA. It use a modified PCR reaction where both
Interpretation of sequence results An overview on DNA sequencing: DNA sequencing involves the determination of the sequence of nucleotides in a sample of DNA. It use a modified PCR reaction where both
Functional Genomics Research Stream. Research Meetings: November 2 & 3, 2009 Next Generation Sequencing
 Functional Genomics Research Stream Research Meetings: November 2 & 3, 2009 Next Generation Sequencing Current Issues Research Meetings: Meet with me this Thursday or Friday. (bring laboratory notebook
Functional Genomics Research Stream Research Meetings: November 2 & 3, 2009 Next Generation Sequencing Current Issues Research Meetings: Meet with me this Thursday or Friday. (bring laboratory notebook
MICROARRAYS: CHIPPING AWAY AT THE MYSTERIES OF SCIENCE AND MEDICINE
 MICROARRAYS: CHIPPING AWAY AT THE MYSTERIES OF SCIENCE AND MEDICINE National Center for Biotechnology Information With only a few exceptions, every
MICROARRAYS: CHIPPING AWAY AT THE MYSTERIES OF SCIENCE AND MEDICINE National Center for Biotechnology Information With only a few exceptions, every
Personal Exam Code: 1. 16p
 Exam, Methods in Molecular Biology, BIOR47, 2012-04-26 Please hand in answers to each question on its paper. Use additional papers if needed. Mark all papers with your code. Your answers should be logical
Exam, Methods in Molecular Biology, BIOR47, 2012-04-26 Please hand in answers to each question on its paper. Use additional papers if needed. Mark all papers with your code. Your answers should be logical
Genome Biology and Biotechnology
 Genome Biology and Biotechnology Functional Genomics Prof. M. Zabeau Department of Plant Systems Biology Flanders Interuniversity Institute for Biotechnology (VIB) University of Gent International course
Genome Biology and Biotechnology Functional Genomics Prof. M. Zabeau Department of Plant Systems Biology Flanders Interuniversity Institute for Biotechnology (VIB) University of Gent International course
(a) Overview of the 2-helix bundle (2HB) nanospring design used in this study. The
 1 Supplementary Figure 1 Design of the DNA origami spring (nanospring). (a) Overview of the 2-helix bundle (2HB) nanospring design used in this study. The scheme was produced by cadnano software 1. Scaffold,
1 Supplementary Figure 1 Design of the DNA origami spring (nanospring). (a) Overview of the 2-helix bundle (2HB) nanospring design used in this study. The scheme was produced by cadnano software 1. Scaffold,
Nature Methods: doi: /nmeth Supplementary Figure 1. Accuracy of microsphere tracking in x, y and z.
 Supplementary Figure 1 Accuracy of microsphere tracking in x, y and z. Accuracy of tracking at 50 Hz in x, y, and z of a 4.5 µm diameter polystyrene microsphere (a) and a 1.5 µm diameter silica microsphere
Supplementary Figure 1 Accuracy of microsphere tracking in x, y and z. Accuracy of tracking at 50 Hz in x, y, and z of a 4.5 µm diameter polystyrene microsphere (a) and a 1.5 µm diameter silica microsphere
The Structure of Proteins The Structure of Proteins. How Proteins are Made: Genetic Transcription, Translation, and Regulation
 How Proteins are Made: Genetic, Translation, and Regulation PLAY The Structure of Proteins 14.1 The Structure of Proteins Proteins - polymer amino acids - monomers Linked together with peptide bonds A
How Proteins are Made: Genetic, Translation, and Regulation PLAY The Structure of Proteins 14.1 The Structure of Proteins Proteins - polymer amino acids - monomers Linked together with peptide bonds A
Biotechnology Project Lab
 Only for teaching purposes - not for reproduction or sale Advanced Cell Biology & Biotechnology Biotechnology Project Lab Giovanna Gambarotta COMPETENCES THAT YOU WILL ACQUIRE - compare DNA sequences -
Only for teaching purposes - not for reproduction or sale Advanced Cell Biology & Biotechnology Biotechnology Project Lab Giovanna Gambarotta COMPETENCES THAT YOU WILL ACQUIRE - compare DNA sequences -
Recitation CHAPTER 9 DNA Technologies
 Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
Microarrays: since we use probes we obviously must know the sequences we are looking at!
 These background are needed: 1. - Basic Molecular Biology & Genetics DNA replication Transcription Post-transcriptional RNA processing Translation Post-translational protein modification Gene expression
These background are needed: 1. - Basic Molecular Biology & Genetics DNA replication Transcription Post-transcriptional RNA processing Translation Post-translational protein modification Gene expression
Restriction Enzymes (Site-Specific Endonuclease) Enzymes that recognize and cleave dsdna in a highly sequence specific manner.
 Enzymes Restriction Enzymes (Site-Specific Endonuclease) Enzymes that recognize and cleave dsdna in a highly sequence specific manner. Generally recognize an inverted repeat sequence 4, 6, or 8 base pairs
Enzymes Restriction Enzymes (Site-Specific Endonuclease) Enzymes that recognize and cleave dsdna in a highly sequence specific manner. Generally recognize an inverted repeat sequence 4, 6, or 8 base pairs
Genetics and Biotechnology. Section 1. Applied Genetics
 Section 1 Applied Genetics Selective Breeding! The process by which desired traits of certain plants and animals are selected and passed on to their future generations is called selective breeding. Section
Section 1 Applied Genetics Selective Breeding! The process by which desired traits of certain plants and animals are selected and passed on to their future generations is called selective breeding. Section
A New Biobrick Assembly Strategy Designed for Facile Protein Engineering
 A New Biobrick Assembly Strategy Designed for Facile Protein Engineering Ira E. Phillips and Pamela A. Silver April 18, 2006 Abstract The existing biobrick assembly technique[1] provides a straightforward
A New Biobrick Assembly Strategy Designed for Facile Protein Engineering Ira E. Phillips and Pamela A. Silver April 18, 2006 Abstract The existing biobrick assembly technique[1] provides a straightforward
7/24/2012. DNA Probes. Hybridization and Probes. CLS 420 Immunology & Molecular Diagnostics. Target Sequences. Target Sequences. Nucleic Acid Probes
 Hybridization and Probes CLS 420 Immunology & Molecular Diagnostics Molecular Diagnostics Techniques: Hybridization and Probes Nucleic acid probes: A short, known sequence of DNA or RNA Used to detect
Hybridization and Probes CLS 420 Immunology & Molecular Diagnostics Molecular Diagnostics Techniques: Hybridization and Probes Nucleic acid probes: A short, known sequence of DNA or RNA Used to detect
Bioinformatics & Protein Structural Analysis. Bioinformatics & Protein Structural Analysis. Learning Objective. Proteomics
 The molecular structures of proteins are complex and can be defined at various levels. These structures can also be predicted from their amino-acid sequences. Protein structure prediction is one of the
The molecular structures of proteins are complex and can be defined at various levels. These structures can also be predicted from their amino-acid sequences. Protein structure prediction is one of the
Microarray. Key components Array Probes Detection system. Normalisation. Data-analysis - ratio generation
 Microarray Key components Array Probes Detection system Normalisation Data-analysis - ratio generation MICROARRAY Measures Gene Expression Global - Genome wide scale Why Measure Gene Expression? What information
Microarray Key components Array Probes Detection system Normalisation Data-analysis - ratio generation MICROARRAY Measures Gene Expression Global - Genome wide scale Why Measure Gene Expression? What information
Supplementary Figure 1 Performance analysis of RedLibs (version 0.2.3). Supplementary Figure 2
 Supplementary Figure 1 Performance analysis of RedLibs (version 0.2.3). The required calculation time for the evaluation of all sub-libraries with a given target size are plotted against the combinatorial
Supplementary Figure 1 Performance analysis of RedLibs (version 0.2.3). The required calculation time for the evaluation of all sub-libraries with a given target size are plotted against the combinatorial
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 14: Microarray Some slides were adapted from Dr. Luke Huan (University of Kansas), Dr. Shaojie Zhang (University of Central Florida), and Dr. Dong Xu and
EECS730: Introduction to Bioinformatics Lecture 14: Microarray Some slides were adapted from Dr. Luke Huan (University of Kansas), Dr. Shaojie Zhang (University of Central Florida), and Dr. Dong Xu and
Lecture 25 (11/15/17)
 Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Feature Selection of Gene Expression Data for Cancer Classification: A Review
 Available online at www.sciencedirect.com ScienceDirect Procedia Computer Science 50 (2015 ) 52 57 2nd International Symposium on Big Data and Cloud Computing (ISBCC 15) Feature Selection of Gene Expression
Available online at www.sciencedirect.com ScienceDirect Procedia Computer Science 50 (2015 ) 52 57 2nd International Symposium on Big Data and Cloud Computing (ISBCC 15) Feature Selection of Gene Expression
Supplementary Figure 1: (a) 3D illustration of the mobile phone microscopy attachment, containing a 3D movable sample holder (red piece for z
 Supplementary Figure 1: (a) 3D illustration of the mobile phone microscopy attachment, containing a 3D movable sample holder (red piece for z movement and blue piece for x-y movement). Illumination sources
Supplementary Figure 1: (a) 3D illustration of the mobile phone microscopy attachment, containing a 3D movable sample holder (red piece for z movement and blue piece for x-y movement). Illumination sources
Lecture 2: High-Throughput Biology
 Lecture 2: High-Throughput Biology COMP 465 Fall 2013 Study Chapter 3.8-3.11 8/27/2013 Comp 465 Fall 2013 1 Analyzing DNA Recall DNA is the essential information determining the function of living organisms
Lecture 2: High-Throughput Biology COMP 465 Fall 2013 Study Chapter 3.8-3.11 8/27/2013 Comp 465 Fall 2013 1 Analyzing DNA Recall DNA is the essential information determining the function of living organisms
Nanoscale-Controlled Surface Materials, Bioanalysis, and Commercialization - NANO KOREA
 Nanoscale-Controlled Surface Materials, Bioanalysis, and Commercialization - NAN KREA 2007 - Aug 28, 2007 Joon Won Park NanoSurface Biosciences PSTECH Pohang University of Science and Technology Biomolecules
Nanoscale-Controlled Surface Materials, Bioanalysis, and Commercialization - NAN KREA 2007 - Aug 28, 2007 Joon Won Park NanoSurface Biosciences PSTECH Pohang University of Science and Technology Biomolecules
Expressed genes profiling (Microarrays) Overview Of Gene Expression Control Profiling Of Expressed Genes
 Expressed genes profiling (Microarrays) Overview Of Gene Expression Control Profiling Of Expressed Genes Genes can be regulated at many levels Usually, gene regulation, are referring to transcriptional
Expressed genes profiling (Microarrays) Overview Of Gene Expression Control Profiling Of Expressed Genes Genes can be regulated at many levels Usually, gene regulation, are referring to transcriptional
Supplementary Figure S1. Np95 interacts with de novo methyltransferases Dnmt3a and 3b. (A) Co-immunoprecipitation of endogenous Dnmt3a2 (left and
 Supplementary Figure S1. Np95 interacts with de novo methyltransferases Dnmt3a and 3b. (A) Co-immunoprecipitation of endogenous Dnmt3a2 (left and right), Dnmt3b isoforms (left) and Dnmt1 (right) with GFP-Np95
Supplementary Figure S1. Np95 interacts with de novo methyltransferases Dnmt3a and 3b. (A) Co-immunoprecipitation of endogenous Dnmt3a2 (left and right), Dnmt3b isoforms (left) and Dnmt1 (right) with GFP-Np95
Circular mrna. Gifu University
 Circular mrna 2015 Gifu University What is a circular mrna? Translation stops!! Linear Stop codon Produce protein What is a circular mrna? Infinite translation!! Huge and consecutive protein Circular mrna
Circular mrna 2015 Gifu University What is a circular mrna? Translation stops!! Linear Stop codon Produce protein What is a circular mrna? Infinite translation!! Huge and consecutive protein Circular mrna
Gene Expression Prokaryotes and Viruses. BIT 220 Chapter 23
 Gene Expression Prokaryotes and Viruses BIT 220 Chapter 23 Types of Regulatory Mechanisms Rapid turn-on and turn off of gene expression (responds to some external source Expression of a cascade of gene
Gene Expression Prokaryotes and Viruses BIT 220 Chapter 23 Types of Regulatory Mechanisms Rapid turn-on and turn off of gene expression (responds to some external source Expression of a cascade of gene
Genomes contain all of the information needed for an organism to grow and survive.
 Section 3: Genomes contain all of the information needed for an organism to grow and survive. K What I Know W What I Want to Find Out L What I Learned Essential Questions What are the components of the
Section 3: Genomes contain all of the information needed for an organism to grow and survive. K What I Know W What I Want to Find Out L What I Learned Essential Questions What are the components of the
Lecture 7: Affinity Chromatography-II
 Lecture 7: Affinity Chromatography-II We have studied basics of affinity purification during last lecture. The current lecture is continuation of last lecture and we will cover following: 1. Few specific
Lecture 7: Affinity Chromatography-II We have studied basics of affinity purification during last lecture. The current lecture is continuation of last lecture and we will cover following: 1. Few specific
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Design and sequence of system 1 for targeted demethylation.
 Supplementary Figure 1 Design and sequence of system 1 for targeted demethylation. (a) Design of system 1 for targeted demethylation. TET1CD was fused to a catalytic inactive Cas9 nuclease (dcas9) and
Supplementary Figure 1 Design and sequence of system 1 for targeted demethylation. (a) Design of system 1 for targeted demethylation. TET1CD was fused to a catalytic inactive Cas9 nuclease (dcas9) and
Fluorescent labeling of the nuclear envelope by localizing GFP on the inner nuclear membrane
 Supporting Information Fluorescent labeling of the nuclear envelope by localizing GFP on the inner nuclear membrane Toshiyuki Taniyama 1, Natsumi Tsuda 1, and Shinji Sueda 1,2,* 1 Department of Bioscience
Supporting Information Fluorescent labeling of the nuclear envelope by localizing GFP on the inner nuclear membrane Toshiyuki Taniyama 1, Natsumi Tsuda 1, and Shinji Sueda 1,2,* 1 Department of Bioscience
beta-galactosidase (A420) ctrl
 0.6 beta-galactosidase (A420) 0.4 0.2 ctrl 2A 22A 35B 36B 58A 64A 68E 75C 86Fa 86Fb 102F Fig. S1. Levels of induced expression at various attp sites. The induced β-galactosidase activity was measured from
0.6 beta-galactosidase (A420) 0.4 0.2 ctrl 2A 22A 35B 36B 58A 64A 68E 75C 86Fa 86Fb 102F Fig. S1. Levels of induced expression at various attp sites. The induced β-galactosidase activity was measured from
Nature Biotechnology: doi: /nbt.4166
 Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
Clamping down on pathogenic bacteria how to shut down a key DNA polymerase complex
 Clamping down on pathogenic bacteria how to shut down a key DNA polymerase complex Bacterial DNA-replication machinery Pathogenic bacteria that are resistant to the current armoury of antibiotics are an
Clamping down on pathogenic bacteria how to shut down a key DNA polymerase complex Bacterial DNA-replication machinery Pathogenic bacteria that are resistant to the current armoury of antibiotics are an
Kinetics Review. Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets)
 Quiz 1 Kinetics Review Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets) I will post the problems with solutions on Toolkit for those that can t make
Quiz 1 Kinetics Review Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets) I will post the problems with solutions on Toolkit for those that can t make
DNA Model Stations. For the following activity, you will use the following DNA sequence.
 Name: DNA Model Stations DNA Replication In this lesson, you will learn how a copy of DNA is replicated for each cell. You will model a 2D representation of DNA replication using the foam nucleotide pieces.
Name: DNA Model Stations DNA Replication In this lesson, you will learn how a copy of DNA is replicated for each cell. You will model a 2D representation of DNA replication using the foam nucleotide pieces.
MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE?
 MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE? Lesson Plan: Title Introduction to the Genome Browser: what is a gene? JOYCE STAMM Objectives Demonstrate basic skills in using the UCSC Genome
MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE? Lesson Plan: Title Introduction to the Genome Browser: what is a gene? JOYCE STAMM Objectives Demonstrate basic skills in using the UCSC Genome
A synthetic recombinase-based feedback loop results in robust expression Supporting Information
 A synthetic recombinase-based feedback loop results in robust expression Supporting Information 1 Fig. S1: Control experiments performed to demonstrate the expected function of our closed-loop feedback
A synthetic recombinase-based feedback loop results in robust expression Supporting Information 1 Fig. S1: Control experiments performed to demonstrate the expected function of our closed-loop feedback
Quantitative Real time PCR. Only for teaching purposes - not for reproduction or sale
 Quantitative Real time PCR PCR reaction conventional versus real time PCR real time PCR principles threshold cycle C T efficiency relative quantification reference genes primers detection chemistry GLP
Quantitative Real time PCR PCR reaction conventional versus real time PCR real time PCR principles threshold cycle C T efficiency relative quantification reference genes primers detection chemistry GLP
Lecture 11 Microarrays and Expression Data
 Introduction to Bioinformatics for Medical Research Gideon Greenspan gdg@cs.technion.ac.il Lecture 11 Microarrays and Expression Data Genetic Expression Data Microarray experiments Applications Expression
Introduction to Bioinformatics for Medical Research Gideon Greenspan gdg@cs.technion.ac.il Lecture 11 Microarrays and Expression Data Genetic Expression Data Microarray experiments Applications Expression
Lectures 9 and 10: Random Walks and the Structure of Macromolecules (contd.)
 Lectures 9 and 10: Random Walks and the Structure of Macromolecules (contd.) Lecturer: Brigita Urbanc Office: 12 909 (E mail: brigita@drexel.edu) Course website: www.physics.drexel.edu/~brigita/courses/biophys_2011
Lectures 9 and 10: Random Walks and the Structure of Macromolecules (contd.) Lecturer: Brigita Urbanc Office: 12 909 (E mail: brigita@drexel.edu) Course website: www.physics.drexel.edu/~brigita/courses/biophys_2011
Single Molecule Force Spectroscopy
 Single Molecule Force Spectroscopy force 0 250 pn rupture force counts cantilever-position x rupture force [pn] E.-L. Florin, V. T. Moy, H. E. Gaub, Science 264, 415 (1994) 1 Streptavidin/Biotin Unbinding
Single Molecule Force Spectroscopy force 0 250 pn rupture force counts cantilever-position x rupture force [pn] E.-L. Florin, V. T. Moy, H. E. Gaub, Science 264, 415 (1994) 1 Streptavidin/Biotin Unbinding
Illumina Sequencing Overview
 Illumina Sequencing Overview Part # 15045845_Rev.C 2013 Illumina, Inc. All rights reserved. Illumina, IlluminaDx, BaseSpace, BeadArray, BeadXpress, cbot, CSPro, DASL, DesignStudio, Eco, GAIIx, Genetic
Illumina Sequencing Overview Part # 15045845_Rev.C 2013 Illumina, Inc. All rights reserved. Illumina, IlluminaDx, BaseSpace, BeadArray, BeadXpress, cbot, CSPro, DASL, DesignStudio, Eco, GAIIx, Genetic
Programmable Sequence-Specific Transcriptional Regulation of Mammalian Genome Using Designer TAL Effectors
 Supplementary Information Programmable Sequence-Specific Transcriptional Regulation of Mammalian Genome Using Designer TAL Effectors Feng Zhang 1,2,3,5,7 *,±, Le Cong 2,3,4 *, Simona Lodato 5,6, Sriram
Supplementary Information Programmable Sequence-Specific Transcriptional Regulation of Mammalian Genome Using Designer TAL Effectors Feng Zhang 1,2,3,5,7 *,±, Le Cong 2,3,4 *, Simona Lodato 5,6, Sriram
TALEN mediated targeted editing of GM2/GD2-synthase gene modulates anchorage independent growth by reducing anoikis resistance in mouse tumor cells
 Supplementary Information for TALEN mediated targeted editing of GM2/GD2-synthase gene modulates anchorage independent growth by reducing anoikis resistance in mouse tumor cells Barun Mahata 1, Avisek
Supplementary Information for TALEN mediated targeted editing of GM2/GD2-synthase gene modulates anchorage independent growth by reducing anoikis resistance in mouse tumor cells Barun Mahata 1, Avisek
Next generation sequencing techniques" Toma Tebaldi Centre for Integrative Biology University of Trento
 Next generation sequencing techniques" Toma Tebaldi Centre for Integrative Biology University of Trento Mattarello September 28, 2009 Sequencing Fundamental task in modern biology read the information
Next generation sequencing techniques" Toma Tebaldi Centre for Integrative Biology University of Trento Mattarello September 28, 2009 Sequencing Fundamental task in modern biology read the information
z-movi High-throughput Label-free Cell Interaction Studies z-movi I Application & Product Brochure
 z-movi High-throughput Label-free Cell Interaction Studies z-movi I Application & Product Brochure Avidity based screening & sorting: z-movi enables direct and label-free screening and sorting of thousands
z-movi High-throughput Label-free Cell Interaction Studies z-movi I Application & Product Brochure Avidity based screening & sorting: z-movi enables direct and label-free screening and sorting of thousands
Supplemental Data. Farmer et al. (2010) Plant Cell /tpc
 Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
