SUPPORTING INFORMATION. Optical Control of CRISPR/Cas9 Gene Editing
|
|
|
- Norma Stevenson
- 6 years ago
- Views:
Transcription
1 SUPPORTING INFORMATION Optical Control of CRISPR/ Gene Editing James Hemphill 1, Erin K. Borchardt 2, Kalyn Brown 1, Aravind Asokan 2, and Alexander Deiters 1 * 1 University of Pittsburgh, Department of Chemistry, Pittsburgh, PA The University of North Carolina at Chapel Hill, Departments of Genetics, Biochemistry, and Biophysics, Chapel Hill, NC METHODS Plasmid constructs. The CMV-driven gene was PCR amplified from the h expression vector 1 (Addgene 41815) with primers shown in Supporting Table 1 to introduce both NheI and MfeI restriction sites as well as an HA tag on the C-terminus. The ~4.9 kb gene insert was digested, purified, and cloned between the NheI and MfeI restriction sites of the ppckrs plasmid 2 with the Quick Ligation Kit (New England Biolabs). Alanine mutations and amber stop codons were introduced into at five sites through site-directed mutagenesis with primers shown in Supporting Tables 2 & 3. U6-promoted grnas targeting sequences upstream and downstream of the DsRed-terminator cassette, as shown in Supporting Table 1, were ordered as gblock gene fragments (IDT). The gblocks were PCR amplified to introduce SgrAI and AgeI restriction sites. Each amplified gblock was individually cloned into a plasmid backbone using these restriction sites to generate the plasmids U6-gRNA-Upstream and U6- grna-downstream. Following sequence validation, the U6-gRNA-Upstream cassette was PCR amplified to introduce flanking NotI restriction sites. The PCR product was digested and ligated into the plasmid U6-gRNA-Downstream plasmid. The resulting plasmid, grna2, contains both upstream and downstream U6-gRNA cassettes. The U6-promoted CD71 grnas, as shown in Supporting Table 1, was also ordered as gblocks, PCR amplified with primers to introduce Bsu36I and PacI restriction sites, and then cloned between the Bsu36I and PacI restriction sites of the ppylt plasmid. 2 Exon-based grna sequences were identified for target sites with minimal off-target effects. Plasmid feature maps are shown in Supporting Figure 8. Mammalian cell expression of alanine mutants and caged proteins. HEK293T cells were seeded at ~100,000 cell per well into 6-well plates (BD Falcon) and incubated overnight in DMEM growth media supplemented with 10% FBS and 2% penicillin/streptomycin at 37 C in 5% CO 2. Transfections were performed with 2 µg of each plasmid using 10 µl lipofectamine transfection reagent (Invitrogen) in 2 ml Opti-Mem media (Invitrogen) at 37 C for 4 hours. After 4 hours the Opti-Mem transfection mixtures were removed from the cells and replaced with DMEM growth media supplemented with PCK (2 mm) for a 48 hour incubation. Western blots. Total protein was extracted from ~1 million cells using 250 µl mammalian cell lysis buffer (Sigma-Aldrich) and size separated on a 10% SDS-PAGE gel, then transferred to a PVDF membrane. Standard western blot conditions were followed using a mouse-anti-ha primary antibody in addition to a mouse-anti-gadph control (Santa Cruz Biotechnology). The primary antibodies were detected with a goat-anti-mouse-hrp fluorescent secondary antibody for chemiluminescent analysis with the VisiGlo kit (Amresco) and analyzed on a ChemiDoc imager (Bio-Rad). S1
2 Optical activation of reporter gene editing. HEK293T cells were seeded at ~10,000 cell per well into 96-well plates (BD Falcon) and incubated overnight in DMEM growth media supplemented with 10% FBS and 2% penicillin/streptomycin at 37 C in 5% CO 2. Transfections were performed with 200 ng of each plasmid using 2 µl branched polyethylenimine (bpei, Sigma-Aldrich) in 200 µl Opti-Mem/DMEM media (Invitrogen) overnight at 37 C following the standard protocol. The transfection mixtures were supplemented with PCK (2 mm) and removed after overnight incubations followed by exposure to 365 nm using a UV transilluminator (25 W). Fluorescent imaging of the dual reporter was performed after 48 hour incubations. Media was replaced with clear DMEM-high modified growth media (Thermo Scientific) for microscopy imaging on a Zeiss Observer Z1 microscope (10X objective, NA 0.8 plan-apochromat) with (38 HE; ex: BP470/40; em: BP525/50) and DsRed (43 HE; ex: BP550/25; em: BP605/70) filter cubes, then processed in Zen Pro 2011 imaging software. Fluorescent cell counting was performed in ImageJ software (NIH - settings: threshold 5-10%, size >200 pixels 2, and circularity 0-1). Error bars represent the standard deviations of three replicates. For the spatial control experiments, UV irradiations were performed through a tin foil mask to only expose a subset of cells to 365 nm light. Microscopy imaging was then performed on a Nikon A1 confocal microscope (20X objective) in a tiled grid and stitched using Elements software. DNA cleavage assays. HA-tag purification was performed on total protein lysate (see above) with 20 µl of mouse-anti-ha antibody (Santa Cruz Biotechnology) and 100 µl of Protein A Sepharose 4B suspension (Life Technologies) overnight at 4 C. The sepharose beads were then washed three times with 300 µl PBS and resuspended to 100 µl total volume. Synthetic grnas were produced via PCR templates for in vitro T7RNAp transcription. The single stranded templates (Supporting Table 4) were PCR amplified and used in transcription reactions with the MEGAscript T7 transcription kit (Life Technologies) according to the manufacturer s protocol. The DNA cleavage assays were performed by incubating 40 µl of the HApurifications, 2 µl of synthetic grna purification (~6000 ng pre-annealed in TAE/Mg 2+ buffer [0.04 M tris-acetate, 1 mm EDTA, and 12.5 mm magnesium acetate]), and 200 ng of the dual reporter plasmid in activity buffer 3 [20mM HEPES, 150 mm KCl, 0.5 mm DTT, 0.1 mm EDTA, 10 mm MgCl 2, ph 7.4] overnight at 37 C. The DNA products were then heated to 72 C for 20 min, loaded onto a 0.8% agarose gel, and stained with ethidium bromide. Photochemical regulation of endogenous CD71. HeLa cells were seeded at ~10,000 cell per well into 96-well plates (BD Falcon) and incubated overnight in DMEM growth media supplemented with 10% FBS and 2% penicillin/streptomycin at 37 C in 5% CO 2. Transfections were performed with 200 ng of each plasmid using 2 µl lipofectamine transfection reagent (Invitrogen) in 200 µl Opti-Mem media (Invitrogen) at 37 C for 4 hours. After 4 hours the Opti- Mem transfection mixtures were removed from the cells and replaced with DMEM growth media supplemented with PCK (2 mm) for a 24 hour incubation. The PCK containing media was removed after the overnight incubation followed by exposure to 365 nm using a UV transilluminator (25 W). After a 48 hour incubation, both quantification of CD71 mrna and fluorescent antibody detection of CD71 protein were performed. Quantification of CD71 mrna was performed by qrt-pcr, in which total RNA was isolated from cells using the Trizol reagent (Invitrogen) and reverse transcribed with the iscript cdna Synthesis kit (Bio-Rad). Quantitative RT-PCR was performed with the SsoFast Evagreen Supermix (Bio-Rad) using primer sets shown in Supporting Table 5. The threshold cycles (Ct) of each sample were normalized to the GAPDH control gene. Protein quantification was performed with anti-human CD71 APC fluorescent antibody (ebioscience) with 0.06 µg for 1 hr at 37 C, then analyzed on a Tecan M1000 plate reader (ex: 635/5; em: 660/10). Error bars represent the standard deviations of three replicates. S2
3 SUPPORTING FIGURES Supporting Figure 1: Structural annotation of critical lysines on. A) Lysines of interest (red) depicted on a surface model of unbound (PDB: 4CMP). B) Lysines of interest depicted on a surface model of bound (PDB: 4OO8), with grna (yellow) and target DNA (blue) shown. K866 is only surface exposed in the grna-target bound structure. C) Detailed view of each lysine of interest in the unbound structure. D) Detailed view of each lysine of interest in the bound structure. Note that K76, K163, K510, and K742 all appear poised to interact with the grna. K866 undergoes a significant conformational change upon binding of the grna, orienting the lysine to become surface exposed. S3
4 A C DsRed B DsRed DsRed ON OFF +reporter wt/grna UV DsRed and grnamediated excision +reporter wt/grna +UV NHEJ repair +reporter +wt/grna UV ON D relative level Supporting Figure 2. Dual reporter CRISPR/ activity assay. A) Depiction of the dual reporter locus. A DsRed gene (red arrow) and an gene (green arrow) are separated by a transcription termination sequence (stop signs). In the absence of, transcription terminates immediately following DsRed, allowing only DsRed expression. B) When functional (blue) and grnas (orange) are present, the complex mediates excision of the DsRed-terminator cassette and NHEJ repair allows expression of. C) Wild-type gene editing of the fluorescent dual reporter construct. HEK293T were transfected with the dual reporter and expression plasmids, then DsRed/ fluorescence was observed at 48 hours. Scale bars indicate 200 µm. D) Analysis of expression by imaging cytometry. Error bars represent standard deviations from three replicates. S4
5 A L WT K76 K163 K510 K742 K866 -HA GAPDH B K76 K163 K510 K742 K866 NT PCK -HA GAPDH Supporting Figure 3. Western blot analysis. The alanine mutant expression plasmids (A) and TAG mutant expression plasmids (B) were transfected and protein was purified for chemiluminescent detection of the C-terminal HA tag. The -HA and GAPDH control bands are annotated, and a horizontal line indicates a cut site on the transfer membrane for antibody staining. S5
6 DsRed K76à Ala K163à Ala K510à Ala K742à Ala K866à Ala Supporting Figure 4. alanine mutant activity scanning. The K A mutant expression plasmids were transfected into cells with the dual reporter system. Scale bars indicate 200 µm. A relative level UV irradiation (min) B relative level UV irradiation (min) Supporting Figure 5. UV irradiation optimization. The wild-type (A) and K866-caged mutant (B) expression plasmids were transfected into cells with the dual reporter system and PCK incorporation constructs, then grown in the presence of PCK (2 mm) for 24 hours. The cells were then UV irradiated for 0-10 minutes and imaged after 48 hour incubation. The images were counted for expressing cells. Error bars represent the standard deviations of three replicates. S6
7 NT WT K163PCK K866PCK min UV nicked linear supercoiled Supporting Figure 6. DNA cleavage assays. The proteins were expressed in HEK293T cells and purified from lysate. purifications were incubated with the dual reporter plasmid and grna overnight at 37 C then analyzed on an agarose gel. A relative CD71 mrna B 1.2 relative fluorescence NT 5 UTR Exon 1 Exon 2 Exon 4 NT 5 UTR Exon 1 Exon 2 Exon 4 Supporting Figure 7. Wild-type controls for the quantification of CD71. HeLa cells were transfected with the and CD71 grna expression constructs, then incubated for 48 hours. Quantification of CD71 mrna with qrt-pcr (A) and CD71 protein with fluorescent antibody detection (B) were performed. Error bars represent the standard deviations of three replicates. S7
8 A TAG ppckrs: PCKRS ppylt PylT (x4) B Dual Reporter DsRed term. grna2 DsRed-4-7 C PylT (x4) grna ppylt: CD71 grna Supporting Figure 8. Plasmid constructs used for the expression of caged (A) and analysis of activity with specific reporter genes (B). The CD71 grna construct is also shown (C). S8
9 SUPPORTING TABLES Supporting Table 1. Sequences of primers used for gene insertion of into ppckrs and grna constructs. Restriction sites bolded, HA tag underlined, and grnas in blue (guide target sequences capitalized). Strand h-nhe1 Forward h-ha-mfe1 Reverse DsRed grna-4 grna-7 CD71 grna-2 5 UTR CD71 grna Exon 1 CD71 grna Exon 2 CD71 grna Exon 4 U6-promoter Sequence (5' 3') TTAAGCTAGCACCATGGACAAGAAGT CGGTGAATTCTTAAGCGTAATCTGGAACATC GTATGGGTACACCTTCCTCTTCTTC TCGACTCTAGAGGATCCACgttttagagctagaaat agcaagttaaaataaggctagtccgttatcaacttgaaaaagtg gcaccgagtcggtgctt TAGCTAGTCTAGGTCGATGCgttttagagctagaaa tagcaagttaaaataaggctagtccgttatcaacttgaaaaagt ggcaccgagtcggtgctt GGACGCGCTAGTGTGAGTGCgttttagagctagaa atagcaagttaaaataaggctagtccgttatcaacttgaaaaag tggcaccgagtcggtgctt GTCATATACCCGGTTCAGCCgttttagagctagaaa tagcaagttaaaataaggctagtccgttatcaacttgaaaaagt ggcaccgagtcggtgctt CTGCAGCACGTCGCTTATATgttttagagctagaaa tagcaagttaaaataaggctagtccgttatcaacttgaaaaagt ggcaccgagtcggtgctt GGGTTATGTGGCGTATAGTAgttttagagctagaaa tagcaagttaaaataaggctagtccgttatcaacttgaaaaagt ggcaccgagtcggtgctt tgtacaaaaaagcaggctttaaaggaaccaattcagtcgactg gatccggtaccaaggtcgggcaggaagagggcctatttcccat gattccttcatatttgcatatacgatacaaggctgttagagagata attagaattaatttgactgtaaacacaaagatattagtacaaaat acgtgacgtagaaagtaataatttcttgggtagtttgcagttttaa aattatgttttaaaatggactatcatatgcttaccgtaacttgaaag tatttcgatttcttggctttatatatcttgtggaaaggacgaaacac cg S9
10 Supporting Table 2. Sequences of primers used in the development of Kà Ala mutants. Mutations introduced capitalized and bolded. Strand K76AlaForward K76ala Reverse K163Ala Forward K163Ala Reverse K510Ala Forward K510Ala Reverse K742Ala Forward K742Ala Reverse K866Ala Forward K866Ala Reverse Sequence (5' 3') gcagagcgaatcggatctgctacctgcaggagatc cgattcgctctgcgggtatatctgcgccgtgctgt tatgatcgcatttcggggacacttcctcatcgagggg cccgaaatgcgatcatatgcgccagcgcgagatagat cttcctgcacactctctgctgtacgagtacttcacagtttataacgagctcaccaa agagtgtgcaggaagcaccttttcgttaggcagatttttatcaaagttagtcatcc accgttgcggtcgtggatgaactcgtcaaagtaa cgaccgcaacggtctgcagtattccctttttgat agaggggcgagtgataacgtcccctcagaag tcactcgcccctctatttttatcggatcttgtcaacac Supporting Table 3. Sequences of primers used in the development of Kà TAG mutants. Mutations introduced capitalized and bolded. Strand K76TAG Forward K76TAG Reverse K163TAG Forward K163TAG Reverse K510TAG Forward K510TAG Reverse K742TAG Forward K742TAG Reverse K866TAG Forward K866TAG Reverse Sequence (5' 3') cgcagatagaatcggatctgctacctgcaggagatc ccgattctatctgcgggtatatctgcgccgtgctgt tatgatctagtttcggggacacttcctcatcgagggg cccgaaactagatcatatgcgccagcgcgagatagat cttccttagcactctctgctgtacgagtacttcacagtttataacgagctcaccaa agagtgctaaggaagcaccttttcgttaggcagatttttatcaaagttagtcatcc accgtttaggtcgtggatgaactcgtcaaagtaa cgacctaaacggtctgcagtattccctttttgat agagggtagagtgataacgtcccctcagaag tcactctaccctctatttttatcggatcttgtcaacac Supporting Table 4. Sequences of templates and primers used for synthetic grna transcription. T7RNAp promoter sequence capitalized. Strand -7 grna template DsRed-4 grna template T7 Forward T7 Reverse Sequence (5' 3') TAATACGACTCACTATAGGGAGAtagct agtctaggtcgatgcgttttagagctagaaatagcaag ttaaaataaggctagtccgttatcaacttgaaaaagtg gcaccgagtcggtgctt TAATACGACTCACTATAGGGAGAtcgact ctagaggatccacgttttagagctagaaatagcaagtt aaaataaggctagtccgttatcaacttgaaaaagtgg caccgagtcggtgctt TAATACGACTCACTATAGGG aaagcaccgactcggtgcca S10
11 Supporting Table 5. Sequences of qrt-pcr primers used. Strand GAPDH Forward GAPDH Reverse CD71 Forward CD71 Reverse Sequence (5' 3') TGCACCACCAACTGCTTAGC GGCATGGACTGTGGTCATGAG AAAATCCGGTGTAGGCACAG GCACTCCAACTGGCAAAGAT REFERENCES 1. Mali, P.; Yang, L.; Esvelt, K. M.; Aach, J.; Guell, M.; DiCarlo, J. E.; Norville, J. E.; Church, G. M., RNA- guided human genome engineering via. Science 2013, 339 (6121), Gautier, A.; Nguyen, D. P.; Lusic, H.; An, W.; Deiters, A.; Chin, J. W., Genetically encoded photocontrol of protein localization in mammalian cells. J Am Chem Soc 2010, 132 (12), Jinek, M.; Chylinski, K.; Fonfara, I.; Hauer, M.; Doudna, J. A.; Charpentier, E., A programmable dual- RNA- guided DNA endonuclease in adaptive bacterial immunity. Science 2012, 337 (6096), S11
XactEdit Cas9 Nuclease with NLS User Manual
 XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX
 Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the
 Supplementary Methods RT-PCR and real-time PCR analysis RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the manufacturer s protocol and quantified by measuring the
Supplementary Methods RT-PCR and real-time PCR analysis RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the manufacturer s protocol and quantified by measuring the
Supplementary Information
 Journal : Nature Biotechnology Supplementary Information Targeted genome engineering in human cells with RNA-guided endonucleases Seung Woo Cho, Sojung Kim, Jong Min Kim, and Jin-Soo Kim* National Creative
Journal : Nature Biotechnology Supplementary Information Targeted genome engineering in human cells with RNA-guided endonucleases Seung Woo Cho, Sojung Kim, Jong Min Kim, and Jin-Soo Kim* National Creative
SUPPORTING INFORMATION. A cleavage-responsive stem-loop hairpin for assaying guide RNA activity
 SUPPORTING INFORMATION A cleavage-responsive stem-loop hairpin for assaying guide RNA activity Tara R. deboer 1, Noreen Wauford 1, Jing-Yi Chung, Miguel Salvador Torres Perez, and Niren Murthy* University
SUPPORTING INFORMATION A cleavage-responsive stem-loop hairpin for assaying guide RNA activity Tara R. deboer 1, Noreen Wauford 1, Jing-Yi Chung, Miguel Salvador Torres Perez, and Niren Murthy* University
RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by. 5 AGACACAAACACCAUUGUCACACUCCACAGC; Rand-2 OMe,
 Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Schematic representation of the endogenous PALB2 locus and gene-disruption constructs
 Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
CRISPR/Cas9 Gene Editing Tools
 CRISPR/Cas9 Gene Editing Tools - Guide-it Products for Successful CRISPR/Cas9 Gene Editing - Why choose Guide-it products? Optimized methods designed for speed and ease of use Complete kits that don t
CRISPR/Cas9 Gene Editing Tools - Guide-it Products for Successful CRISPR/Cas9 Gene Editing - Why choose Guide-it products? Optimized methods designed for speed and ease of use Complete kits that don t
pdsipher and pdsipher -GFP shrna Vector User s Guide
 pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
CRISPR/Cas9 Gene Editing Tools
 CRISPR/Cas9 Gene Editing Tools - Separations Simply Spectacular INDELS Identify indels Determine if one or both copies of your gene have indels The Guide-it Genotype Confirmation Kit: Simple detection
CRISPR/Cas9 Gene Editing Tools - Separations Simply Spectacular INDELS Identify indels Determine if one or both copies of your gene have indels The Guide-it Genotype Confirmation Kit: Simple detection
Ligation Independent Cloning (LIC) Procedure
 Ligation Independent Cloning (LIC) Procedure Ligation Independent Cloning (LIC) LIC cloning allows insertion of DNA fragments without using restriction enzymes into specific vectors containing engineered
Ligation Independent Cloning (LIC) Procedure Ligation Independent Cloning (LIC) LIC cloning allows insertion of DNA fragments without using restriction enzymes into specific vectors containing engineered
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling
 Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
CRISPR/Cas9 Genome Editing: Transfection Methods
 CRISPR/ Genome Editing: Transfection Methods For over 20 years Mirus Bio has developed and manufactured high performance transfection products and technologies. That expertise is now being applied to the
CRISPR/ Genome Editing: Transfection Methods For over 20 years Mirus Bio has developed and manufactured high performance transfection products and technologies. That expertise is now being applied to the
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
Farnham Lab Protocol for CRISPR/Cas 9- mediated enhancer deletion using a puromycin selection vector Created by Yu (Phoebe) Guo
 Farnham Lab Protocol for CRISPR/Cas 9- mediated enhancer deletion using a puromycin selection vector Created by Yu (Phoebe) Guo 20151024 Part A. Design grna oligos 1. Design grnas Go to Optimized CRISPR
Farnham Lab Protocol for CRISPR/Cas 9- mediated enhancer deletion using a puromycin selection vector Created by Yu (Phoebe) Guo 20151024 Part A. Design grna oligos 1. Design grnas Go to Optimized CRISPR
Simple protocol for gene editing using GenCrisprTM Cas9 nuclease
 Simple protocol for gene editing using GenCrisprTM Cas9 nuclease Contents Protocol Step 1: Choose the target DNA sequence Step 2: Design sgrna Step 3: Preparation for sgrna 3.1 In vitro transcription of
Simple protocol for gene editing using GenCrisprTM Cas9 nuclease Contents Protocol Step 1: Choose the target DNA sequence Step 2: Design sgrna Step 3: Preparation for sgrna 3.1 In vitro transcription of
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Guide-it sgrna In Vitro Transcription and Screening Systems User Manual
 Clontech Laboratories, Inc. Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 631438, 631439 & 631440 (042114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra
Clontech Laboratories, Inc. Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 631438, 631439 & 631440 (042114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra
Transfection of neural stem cells with Lipofectamine Stem Transfection Reagent in StemPro medium
 TRANSFECTION PROTOCOL Lipofectamine Stem Transfection Reagent Transfection of neural stem cells with Lipofectamine Stem Transfection Reagent in StemPro medium NSC media, passaging reagents, and complexation
TRANSFECTION PROTOCOL Lipofectamine Stem Transfection Reagent Transfection of neural stem cells with Lipofectamine Stem Transfection Reagent in StemPro medium NSC media, passaging reagents, and complexation
(Supplementary Methods online)
 (Supplementary Methods online) Production and purification of either LC-antisense or control molecules Recombinant phagemids and the phagemid vector were transformed into XL-1 Blue competent bacterial
(Supplementary Methods online) Production and purification of either LC-antisense or control molecules Recombinant phagemids and the phagemid vector were transformed into XL-1 Blue competent bacterial
Protocols for cell lines using CRISPR/CAS
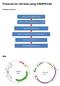 Protocols for cell lines using CRISPR/CAS Procedure overview Map Preparation of CRISPR/CAS plasmids Expression vectors for guide RNA (U6-gRNA) and Cas9 gene (CMV-p-Cas9) are ampicillin-resist ant and stable
Protocols for cell lines using CRISPR/CAS Procedure overview Map Preparation of CRISPR/CAS plasmids Expression vectors for guide RNA (U6-gRNA) and Cas9 gene (CMV-p-Cas9) are ampicillin-resist ant and stable
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
NUCLEIC ACID PURIFICATION KITS FAST SIMPLE QUALITY CONVENIENT REPRODUCIBLE
 NUCLEIC ACID PURIFICATION KITS FAST SIMPLE QUALITY CONVENIENT REPRODUCIBLE COMPANY PROFILE Since its founding in 1998,, Inc. has been at the forefront of nucleic acid purification by offering products
NUCLEIC ACID PURIFICATION KITS FAST SIMPLE QUALITY CONVENIENT REPRODUCIBLE COMPANY PROFILE Since its founding in 1998,, Inc. has been at the forefront of nucleic acid purification by offering products
Gene Forward Primer Reverse Primer GAPDH ATCATCCCTGCCTCTACTGG GTCAGGTCCACCACTGACAC SSB1 AACTTCAGTGAGCCAAACCC GTTCTCAGAGGCTGGAGAGG
 Supplemental Data EXPERIMENTAL PROCEDURES Plasmids and Antibodies- Full length cdna of INT11 or INT12 were cloned into ps- Flag-SBP vector respectively. Anti-RNA pol II (RPB1) was purchased from Santa
Supplemental Data EXPERIMENTAL PROCEDURES Plasmids and Antibodies- Full length cdna of INT11 or INT12 were cloned into ps- Flag-SBP vector respectively. Anti-RNA pol II (RPB1) was purchased from Santa
Guide-it Indel Identification Kit User Manual
 Clontech Laboratories, Inc. Guide-it Indel Identification Kit User Manual Cat. No. 631444 (120114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra Bella Avenue, Mountain View, CA 94043, USA
Clontech Laboratories, Inc. Guide-it Indel Identification Kit User Manual Cat. No. 631444 (120114) Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra Bella Avenue, Mountain View, CA 94043, USA
Confocal immunofluorescence microscopy
 Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
Confocal immunofluorescence microscopy HL-6 and cells were cultured and cytospun onto glass slides. The cells were double immunofluorescence stained for Mt NPM1 and fibrillarin (nucleolar marker). Briefly,
Protocol for Genome Editing via the RNA-guided Cas9 Nuclease in. Zebrafish Embryos 1
 Protocol for Genome Editing via the RNA-guided Cas9 Nuclease in Zebrafish Embryos 1 1. In vitro synthesis of capped Cas9 mrna The full length of humanized Cas9 cdnas with double NLS were cloned into pxt7
Protocol for Genome Editing via the RNA-guided Cas9 Nuclease in Zebrafish Embryos 1 1. In vitro synthesis of capped Cas9 mrna The full length of humanized Cas9 cdnas with double NLS were cloned into pxt7
Supplementary Information
 Supplementary Information Supplementary Figures Figure S1. Study of mgtl translation in vitro. (A) Detection of 5 LR RNA using wild-type and anti-sd (91-95) substituted templates in a transcription-translation
Supplementary Information Supplementary Figures Figure S1. Study of mgtl translation in vitro. (A) Detection of 5 LR RNA using wild-type and anti-sd (91-95) substituted templates in a transcription-translation
For designing necessary primers and oligos for grna, online CRISPR designing tools can be used (e.g.
 DESIGN OF grnas For designing necessary primers and oligos for grna, online CRISPR designing tools can be used (e.g. http://crispr.mit.edu/, https://benchling.com). Assembly of grna transcriptional cassettes
DESIGN OF grnas For designing necessary primers and oligos for grna, online CRISPR designing tools can be used (e.g. http://crispr.mit.edu/, https://benchling.com). Assembly of grna transcriptional cassettes
Engineering splicing factors with designed specificities
 nature methods Engineering splicing factors with designed specificities Yang Wang, Cheom-Gil Cheong, Traci M Tanaka Hall & Zefeng Wang Supplementary figures and text: Supplementary Figure 1 Supplementary
nature methods Engineering splicing factors with designed specificities Yang Wang, Cheom-Gil Cheong, Traci M Tanaka Hall & Zefeng Wang Supplementary figures and text: Supplementary Figure 1 Supplementary
Supplementary Information: Materials and Methods. Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered
 Supplementary Information: Materials and Methods Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered saline (PBS) and lysed in TNN lysis buffer (50mM Tris at ph 8.0, 120mM NaCl
Supplementary Information: Materials and Methods Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered saline (PBS) and lysed in TNN lysis buffer (50mM Tris at ph 8.0, 120mM NaCl
Site directed mutagenesis, Insertional and Deletion Mutagenesis. Mitesh Shrestha
 Site directed mutagenesis, Insertional and Deletion Mutagenesis Mitesh Shrestha Mutagenesis Mutagenesis (the creation or formation of a mutation) can be used as a powerful genetic tool. By inducing mutations
Site directed mutagenesis, Insertional and Deletion Mutagenesis Mitesh Shrestha Mutagenesis Mutagenesis (the creation or formation of a mutation) can be used as a powerful genetic tool. By inducing mutations
Recitation CHAPTER 9 DNA Technologies
 Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
Recitation CHAPTER 9 DNA Technologies DNA Cloning: General Scheme A cloning vector and eukaryotic chromosomes are separately cleaved with the same restriction endonuclease. (A single chromosome is shown
Supporting Information
 Supporting Information Development of a 2,4-Dinitrotoluene-Responsive Synthetic Riboswitch in E. coli cells Molly E. Davidson, Svetlana V. Harbaugh, Yaroslav G. Chushak, Morley O. Stone, Nancy Kelley-
Supporting Information Development of a 2,4-Dinitrotoluene-Responsive Synthetic Riboswitch in E. coli cells Molly E. Davidson, Svetlana V. Harbaugh, Yaroslav G. Chushak, Morley O. Stone, Nancy Kelley-
Supplemental Information
 Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
Supplementary Information: Materials and Methods. GST and GST-p53 were purified according to standard protocol after
 Supplementary Information: Materials and Methods Recombinant protein expression and in vitro kinase assay. GST and GST-p53 were purified according to standard protocol after induction with.5mm IPTG for
Supplementary Information: Materials and Methods Recombinant protein expression and in vitro kinase assay. GST and GST-p53 were purified according to standard protocol after induction with.5mm IPTG for
Polymerase Chain Reaction
 Polymerase Chain Reaction Amplify your insert or verify its presence 3H Taq platinum PCR mix primers Ultrapure Water PCR tubes PCR machine A. Insert amplification For insert amplification, use the Taq
Polymerase Chain Reaction Amplify your insert or verify its presence 3H Taq platinum PCR mix primers Ultrapure Water PCR tubes PCR machine A. Insert amplification For insert amplification, use the Taq
Sarker et al. Supplementary Material. Subcellular Fractionation
 Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
supplementary information
 DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
Transfection of pluripotent stem cells with Lipofectamine Stem reagent in mtesr1 Medium on Geltrex matrix
 TRANSFECTION PROTOCOL Lipofectamine Stem Transfection Reagent Transfection of pluripotent stem cells with Lipofectamine Stem reagent in mtesr1 Medium on Geltrex matrix PSC growth medium, passaging reagents,
TRANSFECTION PROTOCOL Lipofectamine Stem Transfection Reagent Transfection of pluripotent stem cells with Lipofectamine Stem reagent in mtesr1 Medium on Geltrex matrix PSC growth medium, passaging reagents,
Antibodies against PCNA were previously described [1]. To deplete PCNA from Xenopus egg
![Antibodies against PCNA were previously described [1]. To deplete PCNA from Xenopus egg Antibodies against PCNA were previously described [1]. To deplete PCNA from Xenopus egg](/thumbs/84/90687567.jpg) Supplementary information Supplementary methods PCNA antibody and immunodepletion Antibodies against PCNA were previously described [1]. To deplete PCNA from Xenopus egg extracts, one volume of protein
Supplementary information Supplementary methods PCNA antibody and immunodepletion Antibodies against PCNA were previously described [1]. To deplete PCNA from Xenopus egg extracts, one volume of protein
Protocols for cloning SEC-based repair templates using SapTrap assembly
 Protocols for cloning SEC-based repair templates using SapTrap assembly Written by Dan Dickinson (ddickins@live.unc.edu) and last updated July 2016. Overview SapTrap (Schwartz and Jorgensen, 2016) is a
Protocols for cloning SEC-based repair templates using SapTrap assembly Written by Dan Dickinson (ddickins@live.unc.edu) and last updated July 2016. Overview SapTrap (Schwartz and Jorgensen, 2016) is a
MeCP2. MeCP2/α-tubulin. GFP mir1-1 mir132
 Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
Guide-it sgrna In Vitro Transcription and Screening Systems User Manual
 Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 632638, 632639, 632635, 632636, 632637 (040618) 1290 Terra Bella Avenue, Mountain View, CA 94043, USA U.S. Technical Support:
Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 632638, 632639, 632635, 632636, 632637 (040618) 1290 Terra Bella Avenue, Mountain View, CA 94043, USA U.S. Technical Support:
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Supplemental Materials and Methods
 Supplemental Materials and Methods In situ hybridization In situ hybridization analysis of HFE2 and genin mrna in rat liver tissues was performed as previously described (1). Briefly, the digoxigenin-labeled
Supplemental Materials and Methods In situ hybridization In situ hybridization analysis of HFE2 and genin mrna in rat liver tissues was performed as previously described (1). Briefly, the digoxigenin-labeled
Functional Genomics Research Stream. Research Meeting: November 8, 2011 cdna Library Construction for RNA-Seq
 Functional Genomics Research Stream Research Meeting: November 8, 2011 cdna Library Construction for RNA-Seq Lab Issues Don t leave out boxes of water + ethidium bromide Minimize tip boxes in phenol waste
Functional Genomics Research Stream Research Meeting: November 8, 2011 cdna Library Construction for RNA-Seq Lab Issues Don t leave out boxes of water + ethidium bromide Minimize tip boxes in phenol waste
Supporting Information
 Supporting Information Deng et al. 10.1073/pnas.1515692112 SI Materials and Methods FPLC. All fusion proteins were expressed and purified through a three-step FPLC purification protocol, as described (20),
Supporting Information Deng et al. 10.1073/pnas.1515692112 SI Materials and Methods FPLC. All fusion proteins were expressed and purified through a three-step FPLC purification protocol, as described (20),
Applications of Cas9 nickases for genome engineering
 application note genome editing Applications of Cas9 nickases for genome engineering Shuqi Yan, Mollie Schubert, Maureen Young, Brian Wang Integrated DNA Technologies, 17 Commercial Park, Coralville, IA,
application note genome editing Applications of Cas9 nickases for genome engineering Shuqi Yan, Mollie Schubert, Maureen Young, Brian Wang Integrated DNA Technologies, 17 Commercial Park, Coralville, IA,
SUPPLEMENTAL MATERIALS. Chromatin immunoprecipitation assays. Single-cell suspensions of pituitary
 SUPPLEMENTAL MATERIALS METHODS Chromatin immunoprecipitation assays. Single-cell suspensions of pituitary and liver cells were prepared from the indicated transgenic mice, as described (Ho et al, 2002).
SUPPLEMENTAL MATERIALS METHODS Chromatin immunoprecipitation assays. Single-cell suspensions of pituitary and liver cells were prepared from the indicated transgenic mice, as described (Ho et al, 2002).
Protocol CRISPR Genome Editing In Cell Lines
 Protocol CRISPR Genome Editing In Cell Lines Protocol 2: HDR donor plasmid applications (gene knockout, gene mutagenesis, gene tagging, Safe Harbor ORF knock-in) Notes: 1. sgrna validation: GeneCopoeia
Protocol CRISPR Genome Editing In Cell Lines Protocol 2: HDR donor plasmid applications (gene knockout, gene mutagenesis, gene tagging, Safe Harbor ORF knock-in) Notes: 1. sgrna validation: GeneCopoeia
Efficient Method for Isolation of High Quality Concentrated Cellular RNA with Extremely Low Levels of Genomic DNA Contamination Application
 Efficient Method for Isolation of High Quality Concentrated Cellular RNA with Extremely Low Levels of Genomic DNA Contamination Application Gene Expression Authors Ilgar Abbaszade, Claudia Robbins, John
Efficient Method for Isolation of High Quality Concentrated Cellular RNA with Extremely Low Levels of Genomic DNA Contamination Application Gene Expression Authors Ilgar Abbaszade, Claudia Robbins, John
Supplementary Materials
 Supplementary Materials Supplementary Methods Supplementary Discussion Supplementary Figure 1 Calculated frequencies of embryo cells bearing bi-allelic alterations. Targeted indel mutations induced by
Supplementary Materials Supplementary Methods Supplementary Discussion Supplementary Figure 1 Calculated frequencies of embryo cells bearing bi-allelic alterations. Targeted indel mutations induced by
Rapid amplification of cdna ends (RACE)
 Rapid amplification of cdna ends (RACE) Rapid amplification of cdna ends (RACE) is a technique used in molecular biology to obtain the full length sequence of an RNA transcript found within a cell. RACE
Rapid amplification of cdna ends (RACE) Rapid amplification of cdna ends (RACE) is a technique used in molecular biology to obtain the full length sequence of an RNA transcript found within a cell. RACE
QUANTITATIVE RT-PCR PROTOCOL (SYBR Green I) (Last Revised: April, 2007)
 QUANTITATIVE RT-PCR PROTOCOL (SYBR Green I) (Last Revised: April, 007) Please contact Center for Plant Genomics (CPG) facility manager Hailing Jin (hljin@iastate.edu) regarding questions or corrections.
QUANTITATIVE RT-PCR PROTOCOL (SYBR Green I) (Last Revised: April, 007) Please contact Center for Plant Genomics (CPG) facility manager Hailing Jin (hljin@iastate.edu) regarding questions or corrections.
Alt-Ring the genome with CRISPR- Cas9
 Alt-Ring the genome with CRISPR- Cas9 Genome editing done easily and fast CHIA Jin Ngee Regional Application Specialist 1 Integrated DNA Technologies The Custom Biology Company 2 IDT was founded by a scientist
Alt-Ring the genome with CRISPR- Cas9 Genome editing done easily and fast CHIA Jin Ngee Regional Application Specialist 1 Integrated DNA Technologies The Custom Biology Company 2 IDT was founded by a scientist
SuperiorScript III cdna Synthesis Kit Instruction Manual
 SuperiorScript III cdna Synthesis Kit Instruction Manual Cat.# EZ405S, EZ405M SuperiorScript III cdna Synthesis Kit Table of Contents I. Description... 3 II. Kit... 4 III. Procedure... 5 IV. Control Experiment
SuperiorScript III cdna Synthesis Kit Instruction Manual Cat.# EZ405S, EZ405M SuperiorScript III cdna Synthesis Kit Table of Contents I. Description... 3 II. Kit... 4 III. Procedure... 5 IV. Control Experiment
Grb2-Mediated Alteration in the Trafficking of AβPP: Insights from Grb2-AICD Interaction
 Journal of Alzheimer s Disease 20 (2010) 1 9 1 IOS Press Supplementary Material Grb2-Mediated Alteration in the Trafficking of AβPP: Insights from Grb2-AICD Interaction Mithu Raychaudhuri and Debashis
Journal of Alzheimer s Disease 20 (2010) 1 9 1 IOS Press Supplementary Material Grb2-Mediated Alteration in the Trafficking of AβPP: Insights from Grb2-AICD Interaction Mithu Raychaudhuri and Debashis
A novel mechanism of post-translational modulation of HMGA functions by the histone chaperone nucleophosmin.
 A novel mechanism of post-translational modulation of HMGA functions by the histone chaperone nucleophosmin. Laura Arnoldo 1,#, Riccardo Sgarra 1,#, Eusebio Chiefari 2, Stefania Iiritano 2, Biagio Arcidiacono
A novel mechanism of post-translational modulation of HMGA functions by the histone chaperone nucleophosmin. Laura Arnoldo 1,#, Riccardo Sgarra 1,#, Eusebio Chiefari 2, Stefania Iiritano 2, Biagio Arcidiacono
Supplementary Methods
 Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary Figure 1 - Characterization of rbag3 binding on macrophages cell surface.
 Supplementary Figure 1 - Characterization of rbag3 binding on macrophages cell surface. (a) Human PDAC cell lines were treated as indicated in Figure 1 panel F. Cells were analyzed for FITC-rBAG3 binding
Supplementary Figure 1 - Characterization of rbag3 binding on macrophages cell surface. (a) Human PDAC cell lines were treated as indicated in Figure 1 panel F. Cells were analyzed for FITC-rBAG3 binding
Design. Construction. Characterization
 Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
SUPPLEMENTARY INFORMATION
 Gene replacements and insertions in rice by intron targeting using CRISPR Cas9 Table of Contents Supplementary Figure 1. sgrna-induced targeted mutations in the OsEPSPS gene in rice protoplasts. Supplementary
Gene replacements and insertions in rice by intron targeting using CRISPR Cas9 Table of Contents Supplementary Figure 1. sgrna-induced targeted mutations in the OsEPSPS gene in rice protoplasts. Supplementary
DNA Visualizer Extraction Kit
 DNA Visualizer Extraction Kit Catalog Number D0006 50 reactions Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 General Information...
DNA Visualizer Extraction Kit Catalog Number D0006 50 reactions Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 General Information...
Revised: RG-RV2 by Fukuhara et al.
 Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
Protocol for cloning SEC-based repair templates using Gibson assembly and ccdb negative selection
 Protocol for cloning SEC-based repair templates using Gibson assembly and ccdb negative selection Written by Dan Dickinson (daniel.dickinson@austin.utexas.edu) and last updated January 2018. A version
Protocol for cloning SEC-based repair templates using Gibson assembly and ccdb negative selection Written by Dan Dickinson (daniel.dickinson@austin.utexas.edu) and last updated January 2018. A version
Data Sheet Quick PCR Cloning Kit
 Data Sheet Quick PCR Cloning Kit 6044 Cornerstone Ct. West, Ste. E DESCRIPTION: The Quick PCR Cloning Kit is a simple and highly efficient method to insert any gene or DNA fragment into a vector, without
Data Sheet Quick PCR Cloning Kit 6044 Cornerstone Ct. West, Ste. E DESCRIPTION: The Quick PCR Cloning Kit is a simple and highly efficient method to insert any gene or DNA fragment into a vector, without
At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in
 Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
HiPer RT-PCR Teaching Kit
 HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
Supplemental methods:
 Supplemental methods: ASC-J9 treatment ASC-J9 was patented by the University of Rochester, the University of North Carolina, and AndroScience Corp., and then licensed to AndroScience Corp. Both the University
Supplemental methods: ASC-J9 treatment ASC-J9 was patented by the University of Rochester, the University of North Carolina, and AndroScience Corp., and then licensed to AndroScience Corp. Both the University
Somatic Primary pirna Biogenesis Driven by cis-acting RNA Elements and Trans-Acting Yb
 Cell Reports Supplemental Information Somatic Primary pirna Biogenesis Driven by cis-acting RNA Elements and Trans-Acting Yb Hirotsugu Ishizu, Yuka W. Iwasaki, Shigeki Hirakata, Haruka Ozaki, Wataru Iwasaki,
Cell Reports Supplemental Information Somatic Primary pirna Biogenesis Driven by cis-acting RNA Elements and Trans-Acting Yb Hirotsugu Ishizu, Yuka W. Iwasaki, Shigeki Hirakata, Haruka Ozaki, Wataru Iwasaki,
B. ADM: C. D. Apoptosis: 1.68% 2.99% 1.31% Figure.S1,Li et al. number. invaded cells. HuH7 BxPC-3 DLD-1.
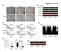 A. - Figure.S1,Li et al. B. : - + - + - + E-cadherin CK19 α-sma vimentin β -actin C. D. Apoptosis: 1.68% 2.99% 1.31% - : - + - + - + Apoptosis: 48.33% 45.32% 44.59% E. invaded cells number 400 300 200
A. - Figure.S1,Li et al. B. : - + - + - + E-cadherin CK19 α-sma vimentin β -actin C. D. Apoptosis: 1.68% 2.99% 1.31% - : - + - + - + Apoptosis: 48.33% 45.32% 44.59% E. invaded cells number 400 300 200
Surrogate reporter-based enrichment of cells containing RNA-guided Cas9 nucleaseinduced
 Supplementary Data Surrogate reporter-based enrichment of cells containing RNA-guided Cas9 nucleaseinduced mutations Suresh Ramakrishna 1, Seung Woo Cho 2, Sojung Kim 2, Myungjae Song 1, Ramu Gopalappa
Supplementary Data Surrogate reporter-based enrichment of cells containing RNA-guided Cas9 nucleaseinduced mutations Suresh Ramakrishna 1, Seung Woo Cho 2, Sojung Kim 2, Myungjae Song 1, Ramu Gopalappa
Supplemental Table 1: Sequences of real time PCR primers. Primers were intronspanning
 Symbol Accession Number Sense-primer (5-3 ) Antisense-primer (5-3 ) T a C ACTB NM_001101.3 CCAGAGGCGTACAGGGATAG CCAACCGCGAGAAGATGA 57 HSD3B2 NM_000198.3 CTTGGACAAGGCCTTCAGAC TCAAGTACAGTCAGCTTGGTCCT 60
Symbol Accession Number Sense-primer (5-3 ) Antisense-primer (5-3 ) T a C ACTB NM_001101.3 CCAGAGGCGTACAGGGATAG CCAACCGCGAGAAGATGA 57 HSD3B2 NM_000198.3 CTTGGACAAGGCCTTCAGAC TCAAGTACAGTCAGCTTGGTCCT 60
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity.
 Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
LINGO-1, A TRANSMEMBRANE SIGNALING PROTEIN, INHIBITS OLIGODENDROCYTE DIFFERENTIATION AND MYELINATION THROUGH INTERCELLULAR SELF- INTERACTIONS.
 Supplemental Data: LINGO-1, A TRANSMEMBRANE SIGNALING PROTEIN, INHIBITS OLIGODENDROCYTE DIFFERENTIATION AND MYELINATION THROUGH INTERCELLULAR SELF- INTERACTIONS. Scott Jepson, Bryan Vought, Christian H.
Supplemental Data: LINGO-1, A TRANSMEMBRANE SIGNALING PROTEIN, INHIBITS OLIGODENDROCYTE DIFFERENTIATION AND MYELINATION THROUGH INTERCELLULAR SELF- INTERACTIONS. Scott Jepson, Bryan Vought, Christian H.
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1 shrna mediated knockdown of ZRSR2 in K562 and 293T cells. (a) ZRSR2 transcript levels in stably transduced K562 cells were determined using qrt-pcr. GAPDH was
Supplementary Figure 1 Supplementary Fig. 1 shrna mediated knockdown of ZRSR2 in K562 and 293T cells. (a) ZRSR2 transcript levels in stably transduced K562 cells were determined using qrt-pcr. GAPDH was
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
Integrated Omics Study Delineates the Dynamics of Lipid Droplets in Rhodococcus Opacus PD630
 School of Natural Sciences and Mathematics 2013-10-22 Integrated Omics Study Delineates the Dynamics of Lipid Droplets in Rhodococcus Opacus PD630 UTD AUTHOR(S): Michael Qiwei Zhang 2013 The Authors This
School of Natural Sciences and Mathematics 2013-10-22 Integrated Omics Study Delineates the Dynamics of Lipid Droplets in Rhodococcus Opacus PD630 UTD AUTHOR(S): Michael Qiwei Zhang 2013 The Authors This
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Document S1. Supplemental Experimental Procedures and Three Figures (see next page)
 Supplemental Data Document S1. Supplemental Experimental Procedures and Three Figures (see next page) Table S1. List of Candidate Genes Identified from the Screen. Candidate genes, corresponding dsrnas
Supplemental Data Document S1. Supplemental Experimental Procedures and Three Figures (see next page) Table S1. List of Candidate Genes Identified from the Screen. Candidate genes, corresponding dsrnas
Protocols. We used a buffer mix with polymerase (either Mango Mix (forhandler) or 2x High Fidelity) and the appropriate primers and template DNA.
 Protocols PCRs Colony PCRs We used a buffer mix with polymerase (either Mango Mix (forhandler) or 2x High Fidelity) and the appropriate primers and template DNA. For a 10 µl reaction 2x Mango Mix/2x High
Protocols PCRs Colony PCRs We used a buffer mix with polymerase (either Mango Mix (forhandler) or 2x High Fidelity) and the appropriate primers and template DNA. For a 10 µl reaction 2x Mango Mix/2x High
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Selected Techniques Part I
 1 Selected Techniques Part I Gel Electrophoresis Can be both qualitative and quantitative Qualitative About what size is the fragment? How many fragments are present? Is there in insert or not? Quantitative
1 Selected Techniques Part I Gel Electrophoresis Can be both qualitative and quantitative Qualitative About what size is the fragment? How many fragments are present? Is there in insert or not? Quantitative
NEBNext RNase III RNA Fragmentation Module
 SAMPLE PREPARATION NEBNext RNase III RNA Fragmentation Module Instruction Manual NEB #E6146S 100 reactions NEBNext RNase III RNA Fragmentation Module Table of Contents: Description....2 Applications....2
SAMPLE PREPARATION NEBNext RNase III RNA Fragmentation Module Instruction Manual NEB #E6146S 100 reactions NEBNext RNase III RNA Fragmentation Module Table of Contents: Description....2 Applications....2
Lipofection of Cas9/synthetic RNA ribonucleoprotein (RNP) complexes for CRISPR/Cas9 genome editing (Thermo CRISPRMAX Kit)
 Lipofection of Cas9/synthetic RNA ribonucleoprotein (RNP) complexes for CRISPR/Cas9 genome editing (Thermo CRISPRMAX Kit) BACKGROUND This protocol describes how to transfect cultured cells with ribonucleoprotein
Lipofection of Cas9/synthetic RNA ribonucleoprotein (RNP) complexes for CRISPR/Cas9 genome editing (Thermo CRISPRMAX Kit) BACKGROUND This protocol describes how to transfect cultured cells with ribonucleoprotein
sirna Transfection Into Primary Neurons Using Fuse-It-siRNA
 sirna Transfection Into Primary Neurons Using Fuse-It-siRNA This Application Note describes a protocol for sirna transfection into sensitive, primary cortical neurons using Fuse-It-siRNA. This innovative
sirna Transfection Into Primary Neurons Using Fuse-It-siRNA This Application Note describes a protocol for sirna transfection into sensitive, primary cortical neurons using Fuse-It-siRNA. This innovative
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the
 Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
CRISPR RNA-guided activation of endogenous human genes
 CRISPR RNA-guided activation of endogenous human genes Morgan L Maeder, Samantha J Linder, Vincent M Cascio, Yanfang Fu, Quan H Ho, J Keith Joung Supplementary Figure 1 Comparison of VEGF activation induced
CRISPR RNA-guided activation of endogenous human genes Morgan L Maeder, Samantha J Linder, Vincent M Cascio, Yanfang Fu, Quan H Ho, J Keith Joung Supplementary Figure 1 Comparison of VEGF activation induced
used at a final concentration of 5 ng/ml. Rabbit anti-bim and mouse anti-mkp2 antibodies were
 1 Supplemental Methods Reagents and chemicals: TGFβ was a generous gift from Genzyme Inc. (Cambridge, MA) and was used at a final concentration of 5 ng/ml. Rabbit anti-bim and mouse anti-mkp2 antibodies
1 Supplemental Methods Reagents and chemicals: TGFβ was a generous gift from Genzyme Inc. (Cambridge, MA) and was used at a final concentration of 5 ng/ml. Rabbit anti-bim and mouse anti-mkp2 antibodies
Supplementary Methods pcfd5 cloning protocol
 Supplementary Methods cloning protocol vermilion trna grna trna grna U6:3 Terminator AmpR attb is a vector for expressing one or multiple trna-flanked Cas9 grnas under the control of the strong, ubiquitous
Supplementary Methods cloning protocol vermilion trna grna trna grna U6:3 Terminator AmpR attb is a vector for expressing one or multiple trna-flanked Cas9 grnas under the control of the strong, ubiquitous
Protocols for cloning SEC-based repair templates using SapTrap assembly
 Protocols for cloning SEC-based repair templates using SapTrap assembly Written by Dan Dickinson (daniel.dickinson@austin.utexas.edu) and last updated January 2018. Overview SapTrap (Schwartz and Jorgensen,
Protocols for cloning SEC-based repair templates using SapTrap assembly Written by Dan Dickinson (daniel.dickinson@austin.utexas.edu) and last updated January 2018. Overview SapTrap (Schwartz and Jorgensen,
TALEN mediated targeted editing of GM2/GD2-synthase gene modulates anchorage independent growth by reducing anoikis resistance in mouse tumor cells
 Supplementary Information for TALEN mediated targeted editing of GM2/GD2-synthase gene modulates anchorage independent growth by reducing anoikis resistance in mouse tumor cells Barun Mahata 1, Avisek
Supplementary Information for TALEN mediated targeted editing of GM2/GD2-synthase gene modulates anchorage independent growth by reducing anoikis resistance in mouse tumor cells Barun Mahata 1, Avisek
SUPPLEMENTARY INFORMATION
 (Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
(Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
BS 50 Genetics and Genomics Week of Oct 24
 BS 50 Genetics and Genomics Week of Oct 24 Additional Practice Problems for Section Question 1: The following table contains a list of statements that apply to replication, transcription, both, or neither.
BS 50 Genetics and Genomics Week of Oct 24 Additional Practice Problems for Section Question 1: The following table contains a list of statements that apply to replication, transcription, both, or neither.
Fatchiyah
 Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
CELLTECHGEN For Research Only. Construction of all-in-one vector for Lenti-virus system (Example: Lenti-EF1 -Cas9-EGFP-U6 sgrna vector)
 Construction of all-in-one vector for Lenti-virus system (Example: Lenti-EF1 -Cas9-EGFP-U6 sgrna vector) Catalog number: CTG-CAS9-18 Introduction The vector Lenti-EF1 -Cas9-EGFP-U6 sgrna is designed for
Construction of all-in-one vector for Lenti-virus system (Example: Lenti-EF1 -Cas9-EGFP-U6 sgrna vector) Catalog number: CTG-CAS9-18 Introduction The vector Lenti-EF1 -Cas9-EGFP-U6 sgrna is designed for
CRISPR Ribonucleoprotein (RNP) User Manual
 CRISPR Ribonucleoprotein (RNP) User Manual Description This user manual describes how to use GenScript s CRISPR Ribonucleoprotein (RNP) products for targeted genome editing. The RNP system is comprised
CRISPR Ribonucleoprotein (RNP) User Manual Description This user manual describes how to use GenScript s CRISPR Ribonucleoprotein (RNP) products for targeted genome editing. The RNP system is comprised
Fig. S1 TGF RI inhibitor SB effectively blocks phosphorylation of Smad2 induced by TGF. FET cells were treated with TGF in the presence of
 Fig. S1 TGF RI inhibitor SB525334 effectively blocks phosphorylation of Smad2 induced by TGF. FET cells were treated with TGF in the presence of different concentrations of SB525334. Cells were lysed and
Fig. S1 TGF RI inhibitor SB525334 effectively blocks phosphorylation of Smad2 induced by TGF. FET cells were treated with TGF in the presence of different concentrations of SB525334. Cells were lysed and
