Methods and Algorithms for Gene Prediction
|
|
|
- Walter Charles
- 6 years ago
- Views:
Transcription
1 Methods and Algorithms for Gene Prediction Chaochun Wei 韦朝春 Sc.D. Shanghai Jiao Tong University Shanghai Center for Bioinformation Technology 5/12/2011 K-J-C Bioinformatics Training Course 1
2 Outline 1. Introduction 2. Gene prediction methods Gene prediction methods HMM TWINSCAN and N-SCAN Using ESTs for gene prediction Resources Latest progress 3. Gene Prediction FAQs 2
3 1. Introduction: DNA DNA contains genetic information. DNA can be expressed as a sequence of letters A,C,G and T. Eg: ACGTTTCGAGGT 3
4 DNARNAProtein & processing 4
5 RNA Processing Primary mrna 5 5
6 Introduction: Gene Structure Initial Exon Intron Internal Exon Intron Intron Terminal Exon 5 UTR ATG GT AG TAA TAG TGA 3 UTR A gene is a highly structured region of DNA, it is a functional unit of inheritance. 6
7 Patterns in Splice Sites Donor Sites Acceptor Sites Josep F. Abril et al. Genome Res. 2005; 15: Sequence data from RefSeq of human, mouse, rat and chicken. 7
8 A Typical Human Gene Structure 8
9 Genes in a Genome 9
10 In a Mammalian Genome Finding all the genes is hard Mammalian genomes are large 8,000 km of 10pt type Only about 1% protein coding 10
11 DNA, mrna, cdna and EST DNA Primary mrna mrna UTR Coding Region UTR 5 3 Transcription 5 3 RNA Processing 5 3 Reverse Transcription 5 Truncated cdna cdna Full-length cdna EST 3 EST 5 EST 3 EST EST: Short (~650bs) High error rate (~1-5%) Contains only UTRs or coding regions 11
12 The Challenge and Opportunity ~3000 genomes 222 animals 93 plants Genome numbers Better Gene Structure Annotation Numbers from 12
13 2. Gene Prediction Method History Gener ation 1 st Early 1980s 2 nd Early 1990s 3 rd Mid- Date Feature Systems Information or Methods used 1990s Approximate ends of protein coding regions and non-coding regions A complete single gene in a short sequence Multiple complete or partial genes in a long sequence 4 th 2000s Complete gene structures in whole genomes TestCode, Fickett 1982 GRAIL, Uberbacher and Mural 1991 GeneID,Guigo et al GeneParser, Snyder and Stormo 1993 FGENSH, Solovyev et al Genscan, Burge and Karlin, 1997 Twinscan, Korf, et al.2001 N-SCAN, Brown,et al Twinscan_EST, N-SCAN_EST, Wei and Brent, 2006 splice sites promoters Codon usage bias Neuro-network methods + Translation start sites Stop sites Method:HMM UTR Method: Generalized HMM Multiple-genomes Transcript products Method:Generalized HMM Bayesian approach 13
14 Gene Prediction Methods (1) Categorization: by input information 1. Ab initio methods Only need genomic sequences as input GENSCAN (Burge 1997; Burge and Karlin 1997) GeneFinder (Green, unpublished) Fgenesh (Solovyev and Salamov 1997) Can predict novel genes 2. Transcript-alignment-based methods Use cdna, mrna or Protein similarity as major clues ENSEMBL (Birney, Clamp, et. al. 2004) Highly accurate Can only find genes with transcript evidences cdna coverage 50-60% + EST coverage up to 85% 14
15 Gene Prediction Methods (2) Categorization: by input information 3. Hybrid Methods Integrate cdna, mrna, protein and EST alignments into ab initio methods Genie (Kulp, Haussler et al. 1996) Fgenesh+ (Solovyev and Salamov 1997) Genomescan (Yeh, Lim et al. 2001) GAZE (Howe, Chothiea et al. 2002) AUGUSTUS+ (Stanke, Schoffmann et al. 2006) 15
16 Gene Prediction Methods(3) Comparative-Genomics-Based Methods TWINSCAN and N-SCAN De novo Assumption: Coding regions are more conserved. No transcript similarity information (like EST, cdna, mrna, or protein) is used TWINSCAN-EST and N-SCAN_EST Hybrid Use EST to improve prediction accuracy 16
17 TWINSCAN: A Novel Gene Prediction System Using Dual Genomes Conservation sequences represent the conservation patterns between two genomes No transcript similarity information (like EST, cdna, mrna, and protein) is used 17
18 Hidden Markov Model: Model behind gene predictors HMM for two biased coins flipping End e1 ( H) 0.8, e1 ( T) 0.2, e2( H) 0.3, e2( T) 0.7 TTHHTTHTTTTTHTHHHHHTHTH arg max Observed sequence x Hidden state sequence P( x, ) 18
19 states Most Probable Path and Viterbi Algorithm i-1 i L-1 L max Let f Recursion (i=1 L) Time complexity l ( i) max (Pr( x0,..., xi 1, 0,..., i1, i fl ( i) e ptr ( l) i { 0,..., i1} l ( xi ) max( f k arg max( f k k k ( i 1) a ( i 1) a kl kl ); ). O ( N 2 L) space complexity O(NL) l)) 19
20 states Probability of All the Possible Paths and Forward Algorithm i-1 i L-1 L + Let f l ( i) Pr( x,..., x, l) 0 i1 i Recursion (i=1 L) Probability of all the probable paths P ( x) f k ( L) P( x, ) f ( i) e ( x ) ( f ( i 1) a ) l l i k k k kl 20
21 states Posterior Probability and Forward and Backward Algorithm i-1 i L-1 L + Posterior Probability P( i k x) P( i k, P( x) x) 21
22 Posterior Probability and Forward and Backward Algorithm l l l l b x e a x P x P ) (1 ) ( ), ( ) ( 1 0 l l i l kl k i b x e a i b ) 1 ( ) ( ) ( 1 ) ( ) ( ) ( ) ( ), ( ) ( x P i b i f x P x k P x k P k k i i 22 states Recursion (i=l-1,,1) Posterior Probability ),..., Pr( ) ( 1 k x x L b i L i k Let i-1 i L-1 L
23 TWINSCAN Model Generalized HMM Each feature in a gene structure corresponds to one state. State-specific length models. State-specific sequence models Use Conservation information 23
24 Conservation Sequence Generated by projecting local alignments to the target sequence human mouse CTAGAGATGCAAAAGAAACAGGTACCGCAGTGC---CCC : :: :... : CTAGAG AGACAGGTACCATAGGGCTCTCCT Pair each nucleotide of the target with if it is aligned and identical : if it is aligned to mismatch. if it is unaligned 24
25 N-SCAN: A Novel Gene Prediction System Using Multiple Genomes Uses Bayesian model to include phylogenetic tree information Predicts introns in 5 UTR Has Conserved non-coding regions (Brown, Gross and Brent, Genome Res. 2005) 25
26 Using ESTs for Gene Prediction: TWINSCAN_EST Integrating EST alignment information into TWINSCAN to improve its accuracy where EST evidence exits and not to compromise its ability to predict novel genes. 26
27 Sequence Representation of EST Alignments 1. Use EST-to-genome alignment programs BLAT (Kent 2002) 2. Project the top alignment for each EST to the target genomic sequence EST Alignments ESTseq NNN EEE III EEE NNN III NNN EEE NNN 27
28 Accuracy Measurement Annotated data sets for training/testing RefSeq ( CCDS ( Accuracy in different levels Nucleotide level Exon level Gene level Transcript level Sensitivity and specificity 28
29 Accuracy Measurement (continue) Annotation Prediction Correct Prediction Sensitivity Correct _ Prediction Total _ Annotation Specificit y Correct _ Prediction Total _ Prediction 29
30 percen TWINSCAN_EST and N-SCAN_EST on the Whole Human Genome TWINSCAN2.03 TWINSCAN_EST N-SCAN N-SCAN_EST exact gene sensitivity exact gene specificity exact exon sensitivity exact exon specificity 30
31 An Example of N-SCAN_EST Prediction (Hg17, chr21:33,459,500-33,465,411) 31
32 An Example of N-SCAN_EST Prediction 32
33 Experimental Validation of Predictions Siepel, Genome Research,
34 Experimental Validation of Predictions See The MGC Project Team, The Completion of the Mammalian Gene Collection (MGC), Genome Research, 2009, 19: Wei, C., et al., Closing in on the C.elegans ORFeome by Cloning TWINSCAN predictions, Genome Research, 2005, 15: Tenney, A. E. et al., Gene prediction and verification in a compact genome with numerous small introns, Genome Research,
35 N-SCAN/TWINSCAN Webserver: 35
36 Resources for Gene Prediction Sequence data sets Nucleotide Sequences (NCBI) dbest mrna cdna Annotations RefSeq ( CCDS ( Genome Browser UCSC Genome Browser( 36
37 Latest Progress in Gene Prediction New Methods Conrad: gene prediction using conditional random fields. Decaprio et al., Genome Res Sep;17(9): Not working for vertebrate genomes SVM for splice site Sonneburg et al., BMC Bioinformatics. 2007;8 Suppl 10:S7. CONTRAST: Gross et al., Genome Biology 2007, 8:R269 Best de novo gene predictor for human (gene level accuracy ~50%) Used SVM and conditional random field 37
38 3. Gene Prediction FAQs (from Ian Korf) Algorithms vs. experts Q: are expert biologists better than computer programs? A: Yes and no. Next-generation sequencing Q: Will next-gen transcript sequencing replace gene prediction? A: No. Rare transcripts may require directed experiments to validate. Prediction accuracy Q: Why are gene prediction programs inaccurate? A: We don t always know why. 38
39 Gene Prediction FAQs (continue) Genes in my favorate genome Q: There is no gene predictor for it, what should I do? 39
40 Gene prediction for algal genomes Microalgae Macroalgae 40
41 A Typical Alga Gene Structure A Typical Human Gene Structure 41
42 Gene Prediction FAQs (continue) Genes in my favorate genome Q: There is no gene predictor for it, what should I do? A: Training a gene predictor or use one that is for another organism that is close to this genome. But it may be inaccurate. Difficult genes Q: why some genes are not predicted by any program? A: They are statistical outliers. 42
43 Gene Prediction FAQs (continue) Just coding exons Q: why other parts are not predicted, such as noncoding exons, alternative isoforms, non-canonical splice sites, gene within genes? A: There are trade offs. Pseudogenes Q: why do some gene predictions have tiny introns? A: Retro-pseudo genes often have very strong coding signals, because they are derived from highly expressed genes. 43
44 Gene Prediction FAQs (end) How can I tell a good gene prediction from a bad one? Scores have been assigned to every exon and intron of a gene. People can tell if a gene prediction is good or not by the scores of exons and introns of this gene. You may have to run the program on your own computer to figure them out! 44
45 Acknowledgement Training program organizers Former lab members (WashU) Dr. Michael Brent Dr. Paul Flicek Dr. Ian Korf Samuel Gross 45
Using Expressing Sequence Tags to Improve Gene Structure Annotation
 Washington University in St. Louis Washington University Open Scholarship All Computer Science and Engineering Research Computer Science and Engineering Report Number: WUCS-2006-25 2006-05-01 Using Expressing
Washington University in St. Louis Washington University Open Scholarship All Computer Science and Engineering Research Computer Science and Engineering Report Number: WUCS-2006-25 2006-05-01 Using Expressing
Outline. Introduction to ab initio and evidence-based gene finding. Prokaryotic gene predictions
 Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
UCSC Genome Browser. Introduction to ab initio and evidence-based gene finding
 UCSC Genome Browser Introduction to ab initio and evidence-based gene finding Wilson Leung 06/2006 Outline Introduction to annotation ab initio gene finding Basics of the UCSC Browser Evidence-based gene
UCSC Genome Browser Introduction to ab initio and evidence-based gene finding Wilson Leung 06/2006 Outline Introduction to annotation ab initio gene finding Basics of the UCSC Browser Evidence-based gene
Gene Signal Estimates from Exon Arrays
 Gene Signal Estimates from Exon Arrays I. Introduction: With exon arrays like the GeneChip Human Exon 1.0 ST Array, researchers can examine the transcriptional profile of an entire gene (Figure 1). Being
Gene Signal Estimates from Exon Arrays I. Introduction: With exon arrays like the GeneChip Human Exon 1.0 ST Array, researchers can examine the transcriptional profile of an entire gene (Figure 1). Being
Eukaryotic Gene Prediction. Wei Zhu May 2007
 Eukaryotic Gene Prediction Wei Zhu May 2007 In nature, nothing is perfect... - Alice Walker Gene Structure What is Gene Prediction? Gene prediction is the problem of parsing a sequence into nonoverlapping
Eukaryotic Gene Prediction Wei Zhu May 2007 In nature, nothing is perfect... - Alice Walker Gene Structure What is Gene Prediction? Gene prediction is the problem of parsing a sequence into nonoverlapping
Gene Structure & Gene Finding Part II
 Gene Structure & Gene Finding Part II David Wishart david.wishart@ualberta.ca 30,000 metabolite Gene Finding in Eukaryotes Eukaryotes Complex gene structure Large genomes (0.1 to 10 billion bp) Exons and
Gene Structure & Gene Finding Part II David Wishart david.wishart@ualberta.ca 30,000 metabolite Gene Finding in Eukaryotes Eukaryotes Complex gene structure Large genomes (0.1 to 10 billion bp) Exons and
TIGR THE INSTITUTE FOR GENOMIC RESEARCH
 Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
user s guide Question 3
 Question 3 During a positional cloning project aimed at finding a human disease gene, linkage data have been obtained suggesting that the gene of interest lies between two sequence-tagged site markers.
Question 3 During a positional cloning project aimed at finding a human disease gene, linkage data have been obtained suggesting that the gene of interest lies between two sequence-tagged site markers.
Chimp Chunk 3-14 Annotation by Matthew Kwong, Ruth Howe, and Hao Yang
 Chimp Chunk 3-14 Annotation by Matthew Kwong, Ruth Howe, and Hao Yang Ruth Howe Bio 434W April 1, 2010 INTRODUCTION De novo annotation is the process by which a finished genomic sequence is searched for
Chimp Chunk 3-14 Annotation by Matthew Kwong, Ruth Howe, and Hao Yang Ruth Howe Bio 434W April 1, 2010 INTRODUCTION De novo annotation is the process by which a finished genomic sequence is searched for
Genome Annotation. What Does Annotation Describe??? Genome duplications Genes Mobile genetic elements Small repeats Genetic diversity
 Genome Annotation Genome Sequencing Costliest aspect of sequencing the genome o But Devoid of content Genome must be annotated o Annotation definition Analyzing the raw sequence of a genome and describing
Genome Annotation Genome Sequencing Costliest aspect of sequencing the genome o But Devoid of content Genome must be annotated o Annotation definition Analyzing the raw sequence of a genome and describing
Annotating 7G24-63 Justin Richner May 4, Figure 1: Map of my sequence
 Annotating 7G24-63 Justin Richner May 4, 2005 Zfh2 exons Thd1 exons Pur-alpha exons 0 40 kb 8 = 1 kb = LINE, Penelope = DNA/Transib, Transib1 = DINE = Novel Repeat = LTR/PAO, Diver2 I = LTR/Gypsy, Invader
Annotating 7G24-63 Justin Richner May 4, 2005 Zfh2 exons Thd1 exons Pur-alpha exons 0 40 kb 8 = 1 kb = LINE, Penelope = DNA/Transib, Transib1 = DINE = Novel Repeat = LTR/PAO, Diver2 I = LTR/Gypsy, Invader
Baum-Welch and HMM applications. November 16, 2017
 Baum-Welch and HMM applications November 16, 2017 Markov chains 3 states of weather: sunny, cloudy, rainy Observed once a day at the same time All transitions are possible, with some probability Each state
Baum-Welch and HMM applications November 16, 2017 Markov chains 3 states of weather: sunny, cloudy, rainy Observed once a day at the same time All transitions are possible, with some probability Each state
Bioinformatics for Proteomics. Ann Loraine
 Bioinformatics for Proteomics Ann Loraine aloraine@uab.edu What is bioinformatics? The science of collecting, processing, organizing, storing, analyzing, and mining biological information, especially data
Bioinformatics for Proteomics Ann Loraine aloraine@uab.edu What is bioinformatics? The science of collecting, processing, organizing, storing, analyzing, and mining biological information, especially data
Genes and gene finding
 Genes and gene finding Ben Langmead Department of Computer Science You are free to use these slides. If you do, please sign the guestbook (www.langmead-lab.org/teaching-materials), or email me (ben.langmead@gmail.com)
Genes and gene finding Ben Langmead Department of Computer Science You are free to use these slides. If you do, please sign the guestbook (www.langmead-lab.org/teaching-materials), or email me (ben.langmead@gmail.com)
Hands-On Four Investigating Inherited Diseases
 Hands-On Four Investigating Inherited Diseases The purpose of these exercises is to introduce bioinformatics databases and tools. We investigate an important human gene and see how mutations give rise
Hands-On Four Investigating Inherited Diseases The purpose of these exercises is to introduce bioinformatics databases and tools. We investigate an important human gene and see how mutations give rise
MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE?
 MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE? Lesson Plan: Title Introduction to the Genome Browser: what is a gene? JOYCE STAMM Objectives Demonstrate basic skills in using the UCSC Genome
MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE? Lesson Plan: Title Introduction to the Genome Browser: what is a gene? JOYCE STAMM Objectives Demonstrate basic skills in using the UCSC Genome
Model. David Kulp, David Haussler. Baskin Center for Computer Engineering and Computer Science.
 Integrating Database Homology in a Probabilistic Gene Structure Model David Kulp, David Haussler Baskin Center for Computer Engineering and Computer Science University of California, Santa Cruz CA, 95064,
Integrating Database Homology in a Probabilistic Gene Structure Model David Kulp, David Haussler Baskin Center for Computer Engineering and Computer Science University of California, Santa Cruz CA, 95064,
user s guide Question 3
 Question 3 During a positional cloning project aimed at finding a human disease gene, linkage data have been obtained suggesting that the gene of interest lies between two sequence-tagged site markers.
Question 3 During a positional cloning project aimed at finding a human disease gene, linkage data have been obtained suggesting that the gene of interest lies between two sequence-tagged site markers.
Lecture 11: Gene Prediction
 Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Chimp Sequence Annotation: Region 2_3
 Chimp Sequence Annotation: Region 2_3 Jeff Howenstein March 30, 2007 BIO434W Genomics 1 Introduction We received region 2_3 of the ChimpChunk sequence, and the first step we performed was to run RepeatMasker
Chimp Sequence Annotation: Region 2_3 Jeff Howenstein March 30, 2007 BIO434W Genomics 1 Introduction We received region 2_3 of the ChimpChunk sequence, and the first step we performed was to run RepeatMasker
Gene Prediction: Statistical Approaches
 Gene Prediction: Statistical Approaches Outline Codons Discovery of Split Genes Exons and Introns Splicing Open Reading Frames Codon Usage Splicing Signals TestCode Gene Prediction: Computational Challenge
Gene Prediction: Statistical Approaches Outline Codons Discovery of Split Genes Exons and Introns Splicing Open Reading Frames Codon Usage Splicing Signals TestCode Gene Prediction: Computational Challenge
Agenda. Annotation of Drosophila. Muller element nomenclature. Annotation: Adding labels to a sequence. GEP Drosophila annotation projects 01/03/2018
 Agenda Annotation of Drosophila January 2018 Overview of the GEP annotation project GEP annotation strategy Types of evidence Analysis tools Web databases Annotation of a single isoform (walkthrough) Wilson
Agenda Annotation of Drosophila January 2018 Overview of the GEP annotation project GEP annotation strategy Types of evidence Analysis tools Web databases Annotation of a single isoform (walkthrough) Wilson
RNA-Sequencing analysis
 RNA-Sequencing analysis Markus Kreuz 25. 04. 2012 Institut für Medizinische Informatik, Statistik und Epidemiologie Content: Biological background Overview transcriptomics RNA-Seq RNA-Seq technology Challenges
RNA-Sequencing analysis Markus Kreuz 25. 04. 2012 Institut für Medizinische Informatik, Statistik und Epidemiologie Content: Biological background Overview transcriptomics RNA-Seq RNA-Seq technology Challenges
Genie Gene Finding in Drosophila melanogaster
 Methods Gene Finding in Drosophila melanogaster Martin G. Reese, 1,2,4 David Kulp, 2 Hari Tammana, 2 and David Haussler 2,3 1 Berkeley Drosophila Genome Project, Department of Molecular and Cell Biology,
Methods Gene Finding in Drosophila melanogaster Martin G. Reese, 1,2,4 David Kulp, 2 Hari Tammana, 2 and David Haussler 2,3 1 Berkeley Drosophila Genome Project, Department of Molecular and Cell Biology,
Machine Learning Methods for RNA-seq-based Transcriptome Reconstruction
 Machine Learning Methods for RNA-seq-based Transcriptome Reconstruction Gunnar Rätsch Friedrich Miescher Laboratory Max Planck Society, Tübingen, Germany NGS Bioinformatics Meeting, Paris (March 24, 2010)
Machine Learning Methods for RNA-seq-based Transcriptome Reconstruction Gunnar Rätsch Friedrich Miescher Laboratory Max Planck Society, Tübingen, Germany NGS Bioinformatics Meeting, Paris (March 24, 2010)
Genome Annotation. Stefan Prost 1. May 27th, States of America. Genome Annotation
 Genome Annotation Stefan Prost 1 1 Department of Integrative Biology, University of California, Berkeley, United States of America May 27th, 2015 Outline Genome Annotation 1 Repeat Annotation 2 Repeat
Genome Annotation Stefan Prost 1 1 Department of Integrative Biology, University of California, Berkeley, United States of America May 27th, 2015 Outline Genome Annotation 1 Repeat Annotation 2 Repeat
Outline. Annotation of Drosophila Primer. Gene structure nomenclature. Muller element nomenclature. GEP Drosophila annotation projects 01/04/2018
 Outline Overview of the GEP annotation projects Annotation of Drosophila Primer January 2018 GEP annotation workflow Practice applying the GEP annotation strategy Wilson Leung and Chris Shaffer AAACAACAATCATAAATAGAGGAAGTTTTCGGAATATACGATAAGTGAAATATCGTTCT
Outline Overview of the GEP annotation projects Annotation of Drosophila Primer January 2018 GEP annotation workflow Practice applying the GEP annotation strategy Wilson Leung and Chris Shaffer AAACAACAATCATAAATAGAGGAAGTTTTCGGAATATACGATAAGTGAAATATCGTTCT
BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers
 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html UCSC
BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html UCSC
measuring gene expression December 5, 2017
 measuring gene expression December 5, 2017 transcription a usually short-lived RNA copy of the DNA is created through transcription RNA is exported to the cytoplasm to encode proteins some types of RNA
measuring gene expression December 5, 2017 transcription a usually short-lived RNA copy of the DNA is created through transcription RNA is exported to the cytoplasm to encode proteins some types of RNA
Analysis of Biological Sequences SPH
 Analysis of Biological Sequences SPH 140.638 swheelan@jhmi.edu nuts and bolts meet Tuesdays & Thursdays, 3:30-4:50 no exam; grade derived from 3-4 homework assignments plus a final project (open book,
Analysis of Biological Sequences SPH 140.638 swheelan@jhmi.edu nuts and bolts meet Tuesdays & Thursdays, 3:30-4:50 no exam; grade derived from 3-4 homework assignments plus a final project (open book,
Molecular Biology Primer. CptS 580, Computational Genomics, Spring 09
 Molecular Biology Primer pts 580, omputational enomics, Spring 09 Starting 19 th century What do we know of cellular biology? ell as a fundamental building block 1850s+: ``DNA was discovered by Friedrich
Molecular Biology Primer pts 580, omputational enomics, Spring 09 Starting 19 th century What do we know of cellular biology? ell as a fundamental building block 1850s+: ``DNA was discovered by Friedrich
Open Access. Abstract
 Software ProSplicer: a database of putative alternative splicing information derived from protein, mrna and expressed sequence tag sequence data Hsien-Da Huang*, Jorng-Tzong Horng*, Chau-Chin Lee and Baw-Jhiune
Software ProSplicer: a database of putative alternative splicing information derived from protein, mrna and expressed sequence tag sequence data Hsien-Da Huang*, Jorng-Tzong Horng*, Chau-Chin Lee and Baw-Jhiune
Guided tour to Ensembl
 Guided tour to Ensembl Introduction Introduction to the Ensembl project Walk-through of the browser Variations and Functional Genomics Comparative Genomics BioMart Ensembl Genome browser http://www.ensembl.org
Guided tour to Ensembl Introduction Introduction to the Ensembl project Walk-through of the browser Variations and Functional Genomics Comparative Genomics BioMart Ensembl Genome browser http://www.ensembl.org
Gene Prediction: Statistical Approaches
 Gene Prediction: Statistical Approaches Outline Codons Discovery of Split Genes Exons and Introns Splicing Open Reading Frames Codon Usage Splicing Signals TestCode Gene Prediction: Computational Challenge
Gene Prediction: Statistical Approaches Outline Codons Discovery of Split Genes Exons and Introns Splicing Open Reading Frames Codon Usage Splicing Signals TestCode Gene Prediction: Computational Challenge
RNA-Seq with the Tuxedo Suite
 RNA-Seq with the Tuxedo Suite Monica Britton, Ph.D. Sr. Bioinformatics Analyst September 2015 Workshop The Basic Tuxedo Suite References Trapnell C, et al. 2009 TopHat: discovering splice junctions with
RNA-Seq with the Tuxedo Suite Monica Britton, Ph.D. Sr. Bioinformatics Analyst September 2015 Workshop The Basic Tuxedo Suite References Trapnell C, et al. 2009 TopHat: discovering splice junctions with
Bio 101 Sample questions: Chapter 10
 Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
Research AceView: a comprehensive cdna-supported gene and transcripts annotation Danielle Thierry-Mieg and Jean Thierry-Mieg
 Research AceView: a comprehensive cdna-supported gene and transcripts annotation Danielle Thierry-Mieg and Jean Thierry-Mieg Address: National Center for Biotechnology Information, National Library of
Research AceView: a comprehensive cdna-supported gene and transcripts annotation Danielle Thierry-Mieg and Jean Thierry-Mieg Address: National Center for Biotechnology Information, National Library of
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene-centered resources at NCBI
 COURSE OF BIOINFORMATICS a.a. 2014-2015 Gene-centered resources at NCBI We searched Accession Number: M60495 AT NCBI Nucleotide Gene has been implemented at NCBI to organize information about genes, serving
COURSE OF BIOINFORMATICS a.a. 2014-2015 Gene-centered resources at NCBI We searched Accession Number: M60495 AT NCBI Nucleotide Gene has been implemented at NCBI to organize information about genes, serving
Introduction to Microarray Data Analysis and Gene Networks. Alvis Brazma European Bioinformatics Institute
 Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
Introduction to Microarray Data Analysis and Gene Networks Alvis Brazma European Bioinformatics Institute A brief outline of this course What is gene expression, why it s important Microarrays and how
ELE4120 Bioinformatics. Tutorial 5
 ELE4120 Bioinformatics Tutorial 5 1 1. Database Content GenBank RefSeq TPA UniProt 2. Database Searches 2 Databases A common situation for alignment is to search through a database to retrieve the similar
ELE4120 Bioinformatics Tutorial 5 1 1. Database Content GenBank RefSeq TPA UniProt 2. Database Searches 2 Databases A common situation for alignment is to search through a database to retrieve the similar
MATH 5610, Computational Biology
 MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
Biotechnology Explorer
 Biotechnology Explorer C. elegans Behavior Kit Bioinformatics Supplement explorer.bio-rad.com Catalog #166-5120EDU This kit contains temperature-sensitive reagents. Open immediately and see individual
Biotechnology Explorer C. elegans Behavior Kit Bioinformatics Supplement explorer.bio-rad.com Catalog #166-5120EDU This kit contains temperature-sensitive reagents. Open immediately and see individual
7.2 Protein Synthesis. From DNA to Protein Animation
 7.2 Protein Synthesis From DNA to Protein Animation Proteins Why are proteins so important? They break down your food They build up muscles They send signals through your brain that control your body They
7.2 Protein Synthesis From DNA to Protein Animation Proteins Why are proteins so important? They break down your food They build up muscles They send signals through your brain that control your body They
Computational aspects of ncrna research. Mihaela Zavolan Biozentrum, Basel Swiss Institute of Bioinformatics
 Computational aspects of ncrna research Mihaela Zavolan Biozentrum, Basel Swiss Institute of Bioinformatics Computational aspects on ncrna Bacterial ncrnas research Gene discovery Target discovery Discovery
Computational aspects of ncrna research Mihaela Zavolan Biozentrum, Basel Swiss Institute of Bioinformatics Computational aspects on ncrna Bacterial ncrnas research Gene discovery Target discovery Discovery
RNA-Seq Software, Tools, and Workflows
 RNA-Seq Software, Tools, and Workflows Monica Britton, Ph.D. Sr. Bioinformatics Analyst September 1, 2016 Some mrna-seq Applications Differential gene expression analysis Transcriptional profiling Assumption:
RNA-Seq Software, Tools, and Workflows Monica Britton, Ph.D. Sr. Bioinformatics Analyst September 1, 2016 Some mrna-seq Applications Differential gene expression analysis Transcriptional profiling Assumption:
Training materials.
 Training materials Ensembl training materials are protected by a CC BY license http://creativecommons.org/licenses/by/4.0/ If you wish to re-use these materials, please credit Ensembl for their creation
Training materials Ensembl training materials are protected by a CC BY license http://creativecommons.org/licenses/by/4.0/ If you wish to re-use these materials, please credit Ensembl for their creation
Review Article A Review of Soft Computing Techniques for Gene Prediction
 ISRN Genomics Volume 2013, Article ID 191206, 8 pages http://dx.doi.org/10.1155/2013/191206 Review Article A Review of Soft Computing Techniques for Gene Prediction Neelam Goel, Shailendra Singh, and Trilok
ISRN Genomics Volume 2013, Article ID 191206, 8 pages http://dx.doi.org/10.1155/2013/191206 Review Article A Review of Soft Computing Techniques for Gene Prediction Neelam Goel, Shailendra Singh, and Trilok
Gene prediction in novel fungal genomes using an ab initio algorithm with unsupervised training
 Methods Gene prediction in novel fungal genomes using an ab initio algorithm with unsupervised training Vardges Ter-Hovhannisyan, 1,4 Alexandre Lomsadze, 2,4 Yury O. Chernoff, 1 and Mark Borodovsky 2,3,5
Methods Gene prediction in novel fungal genomes using an ab initio algorithm with unsupervised training Vardges Ter-Hovhannisyan, 1,4 Alexandre Lomsadze, 2,4 Yury O. Chernoff, 1 and Mark Borodovsky 2,3,5
Eukaryotic Gene Structure
 Eukaryotic Gene Structure Terminology Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Gene Basic physical and
Eukaryotic Gene Structure Terminology Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Gene Basic physical and
 Genome Annotation Genome annotation What is the function of each part of the genome? Where are the genes? What is the mrna sequence (transcription, splicing) What is the protein sequence? What does
Genome Annotation Genome annotation What is the function of each part of the genome? Where are the genes? What is the mrna sequence (transcription, splicing) What is the protein sequence? What does
RNA-Seq Workshop AChemS Sunil K Sukumaran Monell Chemical Senses Center Philadelphia
 RNA-Seq Workshop AChemS 2017 Sunil K Sukumaran Monell Chemical Senses Center Philadelphia Benefits & downsides of RNA-Seq Benefits: High resolution, sensitivity and large dynamic range Independent of prior
RNA-Seq Workshop AChemS 2017 Sunil K Sukumaran Monell Chemical Senses Center Philadelphia Benefits & downsides of RNA-Seq Benefits: High resolution, sensitivity and large dynamic range Independent of prior
Lecture #1. Introduction to microarray technology
 Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
Lecture #1 Introduction to microarray technology Outline General purpose Microarray assay concept Basic microarray experimental process cdna/two channel arrays Oligonucleotide arrays Exon arrays Comparing
From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow
 From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow Technical Overview Import VCF Introduction Next-generation sequencing (NGS) studies have created unanticipated challenges with
From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow Technical Overview Import VCF Introduction Next-generation sequencing (NGS) studies have created unanticipated challenges with
Interpretation of sequence results
 Interpretation of sequence results An overview on DNA sequencing: DNA sequencing involves the determination of the sequence of nucleotides in a sample of DNA. It use a modified PCR reaction where both
Interpretation of sequence results An overview on DNA sequencing: DNA sequencing involves the determination of the sequence of nucleotides in a sample of DNA. It use a modified PCR reaction where both
Bio11 Announcements. Ch 21: DNA Biology and Technology. DNA Functions. DNA and RNA Structure. How do DNA and RNA differ? What are genes?
 Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
Review of Protein (one or more polypeptide) A polypeptide is a long chain of..
 Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Alignment to a database. November 3, 2016
 Alignment to a database November 3, 2016 How do you create a database? 1982 GenBank (at LANL, 2000 sequences) 1988 A way to search GenBank (FASTA) Genome Project 1982 GenBank (at LANL, 2000 sequences)
Alignment to a database November 3, 2016 How do you create a database? 1982 GenBank (at LANL, 2000 sequences) 1988 A way to search GenBank (FASTA) Genome Project 1982 GenBank (at LANL, 2000 sequences)
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Genome and DNA Sequence Databases. BME 110: CompBio Tools Todd Lowe April 5, 2007
 Genome and DNA Sequence Databases BME 110: CompBio Tools Todd Lowe April 5, 2007 Admin Reading: Chapters 2 & 3 Notes available in PDF format on-line (see class calendar page): http://www.soe.ucsc.edu/classes/bme110/spring07/bme110-calendar.html
Genome and DNA Sequence Databases BME 110: CompBio Tools Todd Lowe April 5, 2007 Admin Reading: Chapters 2 & 3 Notes available in PDF format on-line (see class calendar page): http://www.soe.ucsc.edu/classes/bme110/spring07/bme110-calendar.html
Make the protein through the genetic dogma process.
 Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Host : Dr. Nobuyuki Nukina Tutor : Dr. Fumitaka Oyama
 Method to assign the coding regions of ESTs Céline Becquet Summer Program 2002 Structural Neuropathology Lab Molecular Neuropathology Group RIKEN Brain Science Institute Host : Dr. Nobuyuki Nukina Tutor
Method to assign the coding regions of ESTs Céline Becquet Summer Program 2002 Structural Neuropathology Lab Molecular Neuropathology Group RIKEN Brain Science Institute Host : Dr. Nobuyuki Nukina Tutor
Synthetic Biology. Sustainable Energy. Therapeutics Industrial Enzymes. Agriculture. Accelerating Discoveries, Expanding Possibilities. Design.
 Synthetic Biology Accelerating Discoveries, Expanding Possibilities Sustainable Energy Therapeutics Industrial Enzymes Agriculture Design Build Generate Solutions to Advance Synthetic Biology Research
Synthetic Biology Accelerating Discoveries, Expanding Possibilities Sustainable Energy Therapeutics Industrial Enzymes Agriculture Design Build Generate Solutions to Advance Synthetic Biology Research
Measuring transcriptomes with RNA-Seq. BMI/CS 776 Spring 2016 Anthony Gitter
 Measuring transcriptomes with RNA-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2016 Anthony Gitter gitter@biostat.wisc.edu Overview RNA-Seq technology The RNA-Seq quantification problem Generative
Measuring transcriptomes with RNA-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2016 Anthony Gitter gitter@biostat.wisc.edu Overview RNA-Seq technology The RNA-Seq quantification problem Generative
Regulation of eukaryotic transcription:
 Promoter definition by mass genome annotation data: in silico primer extension EMBNET course Bioinformatics of transcriptional regulation Jan 28 2008 Christoph Schmid Regulation of eukaryotic transcription:
Promoter definition by mass genome annotation data: in silico primer extension EMBNET course Bioinformatics of transcriptional regulation Jan 28 2008 Christoph Schmid Regulation of eukaryotic transcription:
Introduction to the UCSC genome browser
 Introduction to the UCSC genome browser Dominik Beck NHMRC Peter Doherty and CINSW ECR Fellow, Senior Lecturer Lowy Cancer Research Centre, UNSW and Centre for Health Technology, UTS SYDNEY NSW AUSTRALIA
Introduction to the UCSC genome browser Dominik Beck NHMRC Peter Doherty and CINSW ECR Fellow, Senior Lecturer Lowy Cancer Research Centre, UNSW and Centre for Health Technology, UTS SYDNEY NSW AUSTRALIA
Analysis of large deletions in human-chimp genomic alignments. Erika Kvikstad BioInformatics I December 14, 2004
 Analysis of large deletions in human-chimp genomic alignments Erika Kvikstad BioInformatics I December 14, 2004 Outline Mutations, mutations, mutations Project overview Strategy: finding, classifying indels
Analysis of large deletions in human-chimp genomic alignments Erika Kvikstad BioInformatics I December 14, 2004 Outline Mutations, mutations, mutations Project overview Strategy: finding, classifying indels
Types of Databases - By Scope
 Biological Databases Bioinformatics Workshop 2009 Chi-Cheng Lin, Ph.D. Department of Computer Science Winona State University clin@winona.edu Biological Databases Data Domains - By Scope - By Level of
Biological Databases Bioinformatics Workshop 2009 Chi-Cheng Lin, Ph.D. Department of Computer Science Winona State University clin@winona.edu Biological Databases Data Domains - By Scope - By Level of
Chapter 15 Gene Technologies and Human Applications
 Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Chapter Outline Chapter 15 Gene Technologies and Human Applications Section 1: The Human Genome KEY IDEAS > Why is the Human Genome Project so important? > How do genomics and gene technologies affect
Mapping strategies for sequence reads
 Mapping strategies for sequence reads Ernest Turro University of Cambridge 21 Oct 2013 Quantification A basic aim in genomics is working out the contents of a biological sample. 1. What distinct elements
Mapping strategies for sequence reads Ernest Turro University of Cambridge 21 Oct 2013 Quantification A basic aim in genomics is working out the contents of a biological sample. 1. What distinct elements
#28 - Promoter Prediction 10/29/07
 BCB 444/544 Required Reading (before lecture) Lecture 28 Mon Oct 29 - Lecture 28 Promoter & Regulatory Element Prediction Chp 9 - pp 113-126 Gene Prediction - finish it Wed Oct 30 - Lecture 29 Phylogenetics
BCB 444/544 Required Reading (before lecture) Lecture 28 Mon Oct 29 - Lecture 28 Promoter & Regulatory Element Prediction Chp 9 - pp 113-126 Gene Prediction - finish it Wed Oct 30 - Lecture 29 Phylogenetics
Measuring transcriptomes with RNA-Seq
 Measuring transcriptomes with RNA-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2017 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC
Measuring transcriptomes with RNA-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2017 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC
Ensembl Tools. EBI is an Outstation of the European Molecular Biology Laboratory.
 Ensembl Tools EBI is an Outstation of the European Molecular Biology Laboratory. Questions? We ve muted all the mics Ask questions in the Chat box in the webinar interface I will check the Chat box periodically
Ensembl Tools EBI is an Outstation of the European Molecular Biology Laboratory. Questions? We ve muted all the mics Ask questions in the Chat box in the webinar interface I will check the Chat box periodically
Files for this Tutorial: All files needed for this tutorial are compressed into a single archive: [BLAST_Intro.tar.gz]
![Files for this Tutorial: All files needed for this tutorial are compressed into a single archive: [BLAST_Intro.tar.gz] Files for this Tutorial: All files needed for this tutorial are compressed into a single archive: [BLAST_Intro.tar.gz]](/thumbs/75/71877651.jpg) BLAST Exercise: Detecting and Interpreting Genetic Homology Adapted by W. Leung and SCR Elgin from Detecting and Interpreting Genetic Homology by Dr. J. Buhler Prequisites: None Resources: The BLAST web
BLAST Exercise: Detecting and Interpreting Genetic Homology Adapted by W. Leung and SCR Elgin from Detecting and Interpreting Genetic Homology by Dr. J. Buhler Prequisites: None Resources: The BLAST web
TRANSCRIPTION AND TRANSLATION
 TRANSCRIPTION AND TRANSLATION Bell Ringer (5 MINUTES) 1. Have your homework (any missing work) out on your desk and ready to turn in 2. Draw and label a nucleotide. 3. Summarize the steps of DNA replication.
TRANSCRIPTION AND TRANSLATION Bell Ringer (5 MINUTES) 1. Have your homework (any missing work) out on your desk and ready to turn in 2. Draw and label a nucleotide. 3. Summarize the steps of DNA replication.
Comparative Bioinformatics. BSCI348S Fall 2003 Midterm 1
 BSCI348S Fall 2003 Midterm 1 Multiple Choice: select the single best answer to the question or completion of the phrase. (5 points each) 1. The field of bioinformatics a. uses biomimetic algorithms to
BSCI348S Fall 2003 Midterm 1 Multiple Choice: select the single best answer to the question or completion of the phrase. (5 points each) 1. The field of bioinformatics a. uses biomimetic algorithms to
Leonardo Mariño-Ramírez, PhD NCBI / NLM / NIH. BIOL 7210 A Computational Genomics 2/18/2015
 Leonardo Mariño-Ramírez, PhD NCBI / NLM / NIH BIOL 7210 A Computational Genomics 2/18/2015 The $1,000 genome is here! http://www.illumina.com/systems/hiseq-x-sequencing-system.ilmn Bioinformatics bottleneck
Leonardo Mariño-Ramírez, PhD NCBI / NLM / NIH BIOL 7210 A Computational Genomics 2/18/2015 The $1,000 genome is here! http://www.illumina.com/systems/hiseq-x-sequencing-system.ilmn Bioinformatics bottleneck
Contents 1 Introduction Information ow in biology Probabilistic Finite State Machines PFS
 Sequence alignment in bioinformatics Ewan Birney Sanger Centre Wellcome Trust Genome Campus Hinxton, Cambridge CB10 1SA, England. Email: birney@sanger.ac.uk March 28, 2000 Contents 1 Introduction 7 1.1
Sequence alignment in bioinformatics Ewan Birney Sanger Centre Wellcome Trust Genome Campus Hinxton, Cambridge CB10 1SA, England. Email: birney@sanger.ac.uk March 28, 2000 Contents 1 Introduction 7 1.1
Gene Regulation Solutions. Microarrays and Next-Generation Sequencing
 Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
Genomics AGRY Michael Gribskov Hock 331
 Genomics AGRY 60000 Michael Gribskov gribskov@purdue.edu Hock 331 Computing Essentials Resources In this course we will assemble and annotate both genomic and transcriptomic sequence assemblies We will
Genomics AGRY 60000 Michael Gribskov gribskov@purdue.edu Hock 331 Computing Essentials Resources In this course we will assemble and annotate both genomic and transcriptomic sequence assemblies We will
Tree Depth in a Forest
 Tree Depth in a Forest Mark Segal Center for Bioinformatics & Molecular Biostatistics Division of Bioinformatics Department of Epidemiology and Biostatistics UCSF NUS / IMS Workshop on Classification and
Tree Depth in a Forest Mark Segal Center for Bioinformatics & Molecular Biostatistics Division of Bioinformatics Department of Epidemiology and Biostatistics UCSF NUS / IMS Workshop on Classification and
Microarray Gene Expression Analysis at CNIO
 Microarray Gene Expression Analysis at CNIO Orlando Domínguez Genomics Unit Biotechnology Program, CNIO 8 May 2013 Workflow, from samples to Gene Expression data Experimental design user/gu/ubio Samples
Microarray Gene Expression Analysis at CNIO Orlando Domínguez Genomics Unit Biotechnology Program, CNIO 8 May 2013 Workflow, from samples to Gene Expression data Experimental design user/gu/ubio Samples
Fig Ch 17: From Gene to Protein
 Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Recommendations from the BCB Graduate Curriculum Committee 1
 Recommendations from the BCB Graduate Curriculum Committee 1 Vasant Honavar, Volker Brendel, Karin Dorman, Scott Emrich, David Fernandez-Baca, and Steve Willson April 10, 2006 Background The current BCB
Recommendations from the BCB Graduate Curriculum Committee 1 Vasant Honavar, Volker Brendel, Karin Dorman, Scott Emrich, David Fernandez-Baca, and Steve Willson April 10, 2006 Background The current BCB
Hybrid Error Correction and De Novo Assembly with Oxford Nanopore
 Hybrid Error Correction and De Novo Assembly with Oxford Nanopore Michael Schatz Jan 13, 2015 PAG Bioinformatics @mike_schatz / #PAGXXIII Oxford Nanopore MinION Thumb drive sized sequencer powered over
Hybrid Error Correction and De Novo Assembly with Oxford Nanopore Michael Schatz Jan 13, 2015 PAG Bioinformatics @mike_schatz / #PAGXXIII Oxford Nanopore MinION Thumb drive sized sequencer powered over
PrimePCR Assay Validation Report
 Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin)
Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin)
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017
 Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Gene Expression: Transcription
 Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
DNA makes RNA makes Proteins. The Central Dogma
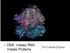 DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
Gene Prediction. Srivani Narra Indian Institute of Technology Kanpur
 Gene Prediction Srivani Narra Indian Institute of Technology Kanpur Email: srivani@iitk.ac.in Supervisor: Prof. Harish Karnick Indian Institute of Technology Kanpur Email: hk@iitk.ac.in Keywords: DNA,
Gene Prediction Srivani Narra Indian Institute of Technology Kanpur Email: srivani@iitk.ac.in Supervisor: Prof. Harish Karnick Indian Institute of Technology Kanpur Email: hk@iitk.ac.in Keywords: DNA,
Assessing De-Novo Transcriptome Assemblies
 Assessing De-Novo Transcriptome Assemblies Shawn T. O Neil Center for Genome Research and Biocomputing Oregon State University Scott J. Emrich University of Notre Dame 100K Contigs, Perfect 1M Contigs,
Assessing De-Novo Transcriptome Assemblies Shawn T. O Neil Center for Genome Research and Biocomputing Oregon State University Scott J. Emrich University of Notre Dame 100K Contigs, Perfect 1M Contigs,
Whole Transcriptome Analysis of Illumina RNA- Seq Data. Ryan Peters Field Application Specialist
 Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Serial Analysis of Gene Expression
 Serial Analysis of Gene Expression Cloning of Tissue-Specific Genes Using SAGE and a Novel Computational Substraction Approach. Genomic (2001) Hung-Jui Shih Outline of Presentation SAGE EST Article TPE
Serial Analysis of Gene Expression Cloning of Tissue-Specific Genes Using SAGE and a Novel Computational Substraction Approach. Genomic (2001) Hung-Jui Shih Outline of Presentation SAGE EST Article TPE
BABELOMICS: Microarray Data Analysis
 BABELOMICS: Microarray Data Analysis Madrid, 21 June 2010 Martina Marbà mmarba@cipf.es Bioinformatics and Genomics Department Centro de Investigación Príncipe Felipe (CIPF) (Valencia, Spain) DNA Microarrays
BABELOMICS: Microarray Data Analysis Madrid, 21 June 2010 Martina Marbà mmarba@cipf.es Bioinformatics and Genomics Department Centro de Investigación Príncipe Felipe (CIPF) (Valencia, Spain) DNA Microarrays
Ensembl gene annotation project
 Ensembl gene annotation project Sus scrofa (Pig) Raw Computes Stage: Searching for sequence patterns, aligning proteins and cdnas to the genome. The annotation process of the Sscrofa10.2 assembly began
Ensembl gene annotation project Sus scrofa (Pig) Raw Computes Stage: Searching for sequence patterns, aligning proteins and cdnas to the genome. The annotation process of the Sscrofa10.2 assembly began
PrimePCR Assay Validation Report
 Gene Information Gene Name collagen, type IV, alpha 1 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID COL4A1 Human This gene encodes the major type IV alpha
Gene Information Gene Name collagen, type IV, alpha 1 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID COL4A1 Human This gene encodes the major type IV alpha
Gene mutation and DNA polymorphism
 Gene mutation and DNA polymorphism Outline of this chapter Gene Mutation DNA Polymorphism Gene Mutation Definition Major Types Definition A gene mutation is a change in the nucleotide sequence that composes
Gene mutation and DNA polymorphism Outline of this chapter Gene Mutation DNA Polymorphism Gene Mutation Definition Major Types Definition A gene mutation is a change in the nucleotide sequence that composes
La traduzione. Il codice genetico
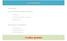 La traduzione Introduction Key-components during translation ØtRNAs ØAminoacil-tRNA sintetasi ØRibosoma The process of translation ØIniziazione ØAllungamento ØTerminazione Il codice genetico IL DOGMA CENTRALE
La traduzione Introduction Key-components during translation ØtRNAs ØAminoacil-tRNA sintetasi ØRibosoma The process of translation ØIniziazione ØAllungamento ØTerminazione Il codice genetico IL DOGMA CENTRALE
Bio-Reagent Services. Custom Gene Services. Gateway to Smooth Molecular Biology! Your Innovation Partner in Drug Discovery!
 Bio-Reagent Services Custom Gene Services Gateway to Smooth Molecular Biology! Gene Synthesis Mutagenesis Mutant Libraries Plasmid Preparation sirna and mirna Services Large-scale DNA Sequencing GenPool
Bio-Reagent Services Custom Gene Services Gateway to Smooth Molecular Biology! Gene Synthesis Mutagenesis Mutant Libraries Plasmid Preparation sirna and mirna Services Large-scale DNA Sequencing GenPool
