Conformational Transitions of the Three Recombinant Domains of Human Serum Albumin Depending on ph*
|
|
|
- Veronica Phelps
- 6 years ago
- Views:
Transcription
1 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 275, No. 5, Issue of February 4, pp , by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Conformational Transitions of the Three Recombinant Domains of Human Serum Albumin Depending on ph* (Received for publication, October 1, 1999, and in revised form, November 4, 1999) Michael Dockal, Daniel C. Carter, and Florian Rüker From the Institute of Applied Microbiology, University of Agricultural Sciences, Muthgasse 18, A-1190 Vienna, Austria and New Century Pharmaceuticals Inc., Huntsville, Alabama Human serum albumin (HSA) is a protein of 66.5 kda that is composed of three homologous domains, each of which displays specific structural and functional characteristics. HSA is known to undergo different ph-dependent structural transitions, the N-F and F-E transitions in the acid ph region and the N-B transition at slightly alkaline ph. In order to elucidate the structural behavior of the recombinant HSA domains as standalone proteins and to investigate the molecular and structural origins of the ph-induced conformational changes of the intact molecule, we have employed fluorescence and circular dichroic methods. Here we provide evidence that the loosening of the HSA structure in the N-F transition takes place primarily in HSA-DOM III and that HSA-DOM I undergoes a structural rearrangement with only minor changes in secondary structure, whereas HSA-DOM II transforms to a molten globulelike state as the ph is reduced. In the ph region of the N-B transition of HSA, HSA-DOM I and HSA-DOM II experience a tertiary structural isomerization, whereas with HSA-DOM III no alterations in tertiary structure are observed, as judged from near-uv CD and fluorescence measurements. The albumin molecule is composed of three homologous, predominantly helical evolutionarily related domains. Each domain is made up of two subdomains, which share a common helical motif. Specific characteristics, for example the location of the various ligand-binding sites, were assigned to some of the domains or subdomains in the past (1, 2). We recently described the production of the three domains of HSA 1 as recombinant proteins, and we were able to show that they have structural features as well as ligand binding properties correlating to their specific functionality in the context of the native protein (3). Clear evidence could be provided that the primary binding site for hemin is located on HSA-DOM I and that for diazepam on HSA-DOM III. Human serum albumin undergoes several transitions in dependence of ph, the N-F transition between ph 5.0 and 3.5, the F-E transition or acid expansion below ph 3.5, and the N-B * This work was supported in part by the Austrian Fonds zur Förderung der Wissenschaftlichen Forschung Grant P11280-MED and Jubiläumsfonds der Oesterreichischen Nationalbank Grant P7277-MED. The costs of publication of this article were defrayed in part by the payment of page charges. This article must therefore be hereby marked advertisement in accordance with 18 U.S.C. Section 1734 solely to indicate this fact. Supported in part by New Century Pharmaceuticals, Inc. To whom correspondence should be addressed: Institute of Applied Microbiology, University of Agricultural Sciences, Muthgasse 18, A-1190 Vienna, Austria. Tel.: ; Fax: ; ruker@mail.boku.ac.at. 1 The abbreviations used are: HSA, human serum albumin; BSA, bovine serum albumin; DOM, domain transition between ph 7.0 and 9.0 (1, 2). The acid-induced structural changes of bovine and human serum albumin have been studied by a wide range of methods (1, 2) and are characterized by changes in secondary as well as tertiary structure. An indication for the former is a significant decrease in helical content observed by far-uv CD measurements which was interpreted by Era et al. (4, 5) to represent a helix 3 and a helix 3 coil transition in the case of the N-F transition. As the ph of an albumin solution is reduced below 3.5, further unfolding occurs until about ph 2.5, when the molecule appears to be expanded to the full extent that is allowed by its disulfide bonding structure. Under slightly alkaline conditions, between ph 7.0 and 9.0, HSA and BSA undergo another conformational change, known as the N-B transition. It is more subtle and more gradual in onset and has been proposed to have physiological importance, similar to the acid-induced transformations (1, 2). The B isomerization is supposed to be a structural fluctuation, a loosening of the molecule with loss of rigidity, particularly affecting the N-terminal region and thereby impacting ligand binding (6 9). One way to explore the molecular origins of these transitions is by using albumin fragments. Attempts have been made in the past to assign specific roles and contributions of the albumin domains to the characteristic structural transitions of both HSA and, more frequently, its closely related bovine homolog, BSA. For many of these studies, fragments have been used, which had been produced by enzymatic or chemical cleavage (8 17). An F-like transformation was observed with a tryptic fragment of bovine serum albumin (BSA-T ), shedding some light on the nucleation points for the intramolecular changes that occur during this transition (13). It was shown that this fragment unfolds and abruptly loses structure at ph 4, whereas fragment BSA-P does not do so until ph 3.5 (12), as determined by difference spectroscopic methods. Domain III of BSA was consequently proposed to have a less constrained conformation than the rest of the molecule and to expand through separation of its subunits in the course of the N-F transition. Bos et al. (17) concluded from their work with two large fragments of HSA that the conformational change between ph 6.0 and 9.0 of the P-46 fragment (residues of HSA) clearly resembles the N-B transition of albumin, whereas the T-45 fragment (residues of HSA) does not display such a conformational change. However, the choice of fragments used in these studies was dependent on the availability of suitable cleavage sites and did not necessarily take into account the natural borders of the domains as they can be defined based on amino acid sequence and atomic structure. Based on our previous results on the structure and ligand binding properties of the three recombinant HSA domains, we report here in a systematic way on the conformational behavior of these new proteins in the acidic and slightly alkaline ph region and on their contributions to the origins of the structural This paper is available on line at
2 Conformational Transitions of Recombinant Albumin Domains 3043 transitions of the native molecule. For this purpose, fluorescence and CD spectroscopic measurements were performed depending on ph, and the results are discussed in the context of the atomic coordinates of HSA. EXPERIMENTAL PROCEDURES Materials HSA (essentially free of fatty acids and globulin) was from Sigma (catalog number A3782). This material was further purified by size exclusion chromatography over Superdex 200 prep grad (Amersham Pharmacia Biotech) and delipidated by passing over a Lipidx column (Canberra-Packard). The three domains of HSA were cloned, expressed, and purified as has been described before in detail (3). The domains encompassed the following amino acid residues of HSA: HSA-DOM I, 1 197; HSA-DOM II, ; HSA-DOM III, Due to the restriction site used for cloning, all domains had Glu-Phe as the N-terminal amino acids. All other reagents were of analytical grade. Protein Solutions Protein stock solutions were prepared by dissolving the delipidated proteins in aqua bidest. Protein concentrations were measured by their absorbance at 278 nm on a Hewlett-Packard HP % spectrophotometer. For HSA, an absorption coefficient (A 278 nm,1 cm )of 0.58 was used (8). For HSA-DOM I, HSA-DOM II, and HSA-DOM III, the absorption coefficients were found to be 0.50, 0.79, and 0.30, respectively (3). ph-dependent CD Measurements Measurements were made using a Jasco J-600 spectropolarimeter using a 1-mm cell at 25 C in a thermostated cell holder at a concentration of 4.5 M in the far-uv region. Scans were made from 250 to 190 nm. The slit width was programmed for a half-bandwidth of 1 nm, and the dynode voltage never exceeded 0.6 kv. 10 l of a 100 times concentrated protein stock solution (in aqua bidest.) were mixed with 990 l of the corresponding buffer (ph 9.0 to 8.5, 10 mm boric acid/borax; ph 8.0 to 5.0, 10 mm sodium phosphate; ph 4.5 to 3.5, 10 mm acetic acid/sodium acetate; ph 3.0 and less was adjusted with HCl). All spectra were recorded at least twice. After each measurement, the ph was confirmed with a standard ph meter. The data were expressed as mean residue ellipticity ([ ] MRW, degree cm 2 dmol 1 ), using the mean residue weights of g mol 1 for the intact molecule, g mol 1 for HSA-DOM I, g mol 1 for HSA-DOM II, and g mol 1 for HSA-DOM III. The fractional content of the secondary structure elements of the proteins was calculated from the far-uv CD spectra using the procedure of Provencher and co-workers (18, 19) (CONTIN) with a set of 16 reference proteins. In the near-uv region, measurements were made using a 1-cm cell at 25 C in a thermostated cell-holder at a protein concentration of 20 M. Scans were made from 340 to 250 nm. The slit width was programmed for a half-bandwidth of 1 nm, and the dynode voltage never exceeded 0.4 kv. As above, a 100 times concentrated protein stock solution was diluted with the desired buffer (ph 9.0, 0.1 M boric acid/borax; ph 7.4, 0.1 M sodium phosphate; ph 4.0, 0.1 M acetic acid/sodium acetate; ph 2.0 was adjusted with HCl) to reach the final protein concentration of 20 M. The results are expressed as molar ellipticity ([ ], degree cm 2 dmol 1 ). ph-dependent Fluorescence Spectroscopy Fluorescence measurements were made with a Hitachi F-4500 fluorescence spectrometer equipped with a thermostated cell holder at 25 C. All spectra were recorded in a cm stirred cell (0.5 cm at emission and 1 cm at excitation side) with the excitation and the emission slit width set to 5 nm. For HSA and HSA-DOM II, which contain only a single tryptophanyl residue (Trp-214), an excitation wavelength of 295 nm was routinely used, to ensure that the light was absorbed almost entirely by the lone tryptophanyl residue. HSA-DOM I and HSA-DOM III were excited at 280 nm to record the tyrosyl emission. ph-dependent experiments were carried out as described above with a protein concentration of 4.5 M, and all spectra were recorded at least twice. Each spectrum was corrected for buffer base-line fluorescence. After each measurement the ph was controlled with a standard ph meter. RESULTS Far-UV CD In order to get information about the secondary structure of the proteins used in this study, far-uv CD measurements were performed between 250 and 190 nm depending on the ph. A spectrum for each of the proteins in the native (ph 7.4) and denatured state (9.5 M urea) is shown in Fig. 1A, demonstrating full disruption of the secondary structure of all proteins at 9.5 M urea. FIG. 1. A, far-uv CD spectra of HSA ( ), HSA-DOM I (- - -), HSA- DOM II (- -), and HSA-DOM III (- -) at ph 7.4 and 9.5 M urea, 25 C, protein concentration 4.5 M. B, effect of ph on the far-uv CD at 222 nm of HSA (E), HSA-DOM I ( ), HSA-DOM II (Œ), and HSA-DOM III ( ) at 25 C, protein concentration 4.5 M. The regions of the transitions of HSA are indicated. Fig. 1B presents the ellipticity at 222 nm in the ph range from 9.0 to 1.5 for the four proteins. Alterations in the ellipticity at this wavelength are a useful probe for visualizing varying -helical content. The results of the secondary structure resolved analysis are represented in Table I for ph 9.0, 7.4, 4.0 and 2.5. In the acid ph region from ph 5.0 to 3.5 and further down to ph 2.5, the ellipticity observed with HSA shows a two-step sigmoidal increase, and calculated helix content (f ) decreases, whereas sheet structures (f ) increase. These changes in secondary structure are in accordance with the concept of the N-F transition followed by the acid expansion (1, 2, 4, 5). A similar observation is made with HSA-DOM III, which also displays a characteristic two-step transition correlated with a decrease in -helical content and a gain in -sheet structure. However, the two transitions are clearly split. The onset of the first sigmoidal
3 3044 Conformational Transitions of Recombinant Albumin Domains TABLE I Fraction of secondary structural elements of HSA, HSA-DOM I, HSA- DOM II, and HSA-DOM III estimated by the procedure of Provencher and Glöckner (CONTIN) (18, 19) The abbreviations used are: f, fraction of -helix content; f, fraction of -sheet content; f R, fraction of remainder structure elements. ph f f f R HSA HSA-DOM I HSA-DOM II HSA-DOM III transition is shifted to higher ph as compared with HSA, followed by a second change in the ph region between 3.0 and 2.0. These findings clearly prove that the N-F transition originates in a structural loosening at the C-terminal end of the albumin molecule, probably followed by a separation of domains and subdomains in the course of the acid expansion, as has been suggested before by several authors (1, 2, 13, 15, 16). For HSA-DOM I, a slight increase in [ ] 222 is observed in the ph region from 5.5 to 3.5, which indicates only a minor change in secondary structure, whereas below ph 3.5 this domain shows an acid expansion similar to HSA and HSA-DOM III, which is further evidence for the disruption of intradomain structure of HSA in this ph region. By diminishing the ph of a HSA-DOM II solution from 6.0 to 4.0, unexpectedly a sigmoidal decrease in [ ] 222, is observed, which is correlated with a slight increase in -helix content accompanied by a reduction of -sheet structure. HSA-DOM II shows, in contrast to HSA- DOM I and HSA-DOM III, only a slight increment in ellipticity in the acid expansion region of HSA. Below ph 2.0, the ellipticity of all four proteins investigated is decreased. This can be interpreted as a gain in secondary structure, as was observed previously for many proteins in the absence of salt by Goto et al. (20) and Fink et al. (21). HSA shows a small increase in the ellipticity at 222 nm in the ph range from 7.4 to 9.0, indicating a slight reduction of -helical content and an increase in -structural elements, in accordance with the concept of the well known N-B transition in the absence of added salt (2, 22). Neither of the recombinant domains exhibits significant fluctuation in ellipticity between ph 6.0 and 9.0. In this ph region, only minor changes in secondary structure can therefore be expected, which is further supported by the data from the secondary structure resolved analysis. Tryptophanyl Fluorescence HSA has a single tryptophanyl residue, Trp-214, located in domain II. To examine the conformational variations around this residue in HSA and HSA-DOM II, fluorescence was excited at 295 nm, which provides no excitation of tyrosine residues and therefore neither emission nor energy transfer to the lone indole side chain. The ph profiles of the fluorescence intensity and the wavelength at the respective emission maximum are displayed in Fig. 2A for HSA and HSA-DOM II. The fluorescence of HSA in the acid ph region exhibited a two-step decrease, the first from ph 6.0 to 4.5 and the second, more pronounced, from ph 3.5 to 2.5, representing the N-F and F-E transitions (16, 23, 24). The FIG. 2.A, effect of ph on the Trp-214 fluorescence intensity at the emission maximum (left-hand scale) and emission maximum wavelength (right-hand scale) of HSA (E) and HSA-DOM II (Œ) at25 C, excitation 295 nm, protein concentration 4.5 M. The regions of the transitions of HSA are indicated. B, effect of ph on the Tyr fluorescence intensity at the emission maximum (left-hand scale) and emission maximum wavelength (right-hand scale) of HSA-DOM I ( ) and HSA-DOM III ( ) at 25 C, excitation 280 nm, protein concentration 4.5 M. The regions of the transitions of HSA are indicated. wavelength maximum of the tryptophanyl fluorescence shifted from 340 nm at the high ph end of the N-F transition (ph 5.0) to approximately 332 nm at ph 3.5. By raising the ph of an HSA solution from 7.4 to 9.0, we observed a slight decrease of max and fluorescence intensity (Fig. 2A), which was reported previously for BSA in the absence of salt (22). At ph 7.4, the fluorescence quantum yield of the tryptophanyl residue in HSA-DOM II is 2.6-fold lower compared with Trp-214 in the native molecule, and the emission maximum is shifted from 340 (HSA) to 332 nm (HSA-DOM II). As the ph is reduced, HSA-DOM II shows a sigmoidal rise in fluorescence intensity, reaching a maximum increase of 2.4-fold at ph 4.0. At the onset of this intensity enhancement no change of the emission maximum is observable, whereas from ph 5.5 to ph 4.0 a slight decrease of 1 nm can be detected. The pronounced rise in fluorescence intensity of HSA-DOM II between ph 7.4 and 4.0 coincides with the secondary structural rearrangement
4 Conformational Transitions of Recombinant Albumin Domains 3045 observed by far-uv CD in the same ph region. At ph 4.0, the tryptophanyl residues of both HSA-DOM II and HSA have superimposable fluorescence spectra (data not shown), suggesting that at this ph the tryptophanyl residue in both proteins has a similar environment. By further reduction of the ph, the fluorescence intensity and emission maximum of HSA-DOM II and HSA run nearly parallel, but the diminution in fluorescence intensity below ph 3.0 followed by the increase below ph 2.0 is not as pronounced in HSA-DOM II as it is in HSA. Even at ph 2.0, the tryptophanyl residues are not in a solvent-exposed environment, as indicated by denaturing both proteins with 9.5 M urea, which leads to a red shift of the fluorescence maximum (349 nm for both proteins) and a reduction of the fluorescence intensity (data not shown). In the slightly alkaline ph region HSA-DOM II shows a gain in tryptophanyl fluorescence intensity, which is accompanied by an increase of the emission maximum. Tyrosyl Fluorescence Since HSA harbors only a single tryptophanyl residue, which is located in domain II, we investigated the tyrosyl fluorescence of HSA-DOM I and HSA-DOM III in order to obtain additional insight into the structural characteristics of these two proteins. The ph profiles of the fluorescence intensity and the wavelength at the emission maximum are visualized in Fig. 2B. In the acidic ph region, HSA-DOM I shows a two-step alteration in fluorescence intensity. At first, by lowering the ph to 3.5, the fluorescence increases, which is aligned with a slight, but significant, decrease in the emission maximum. By further reducing the ph, the fluorescence intensity rises to a maximum value at ph 2.0; however, no change in the position of the emission maximum is observed. The spectra of HSA-DOM I in 9.5 M urea and at ph 2.0 (data not shown) are nearly superimposable, indicating that the seven tyrosyl residues in this protein have the same environment at ph 2.0 as in the fully denaturated state at 9.5 M urea. In the neutral and slightly alkaline ph range the tyrosyl fluorescence and the position of the emission maximum of HSA-DOM I show no significant alterations, which indicates no significant changes in the microenvironments of the seven tyrosyl residues. As the ph of the HSA-DOM III solution is reduced to ph 4.0, there is a sharp sigmoidal decrease in the tyrosyl fluorescence intensity and a slight increase in the emission maximum, however, by further lessening of the ph of the solution there is a smooth and continuous increase in fluorescence intensity. The tyrosyl fluorescence intensity of HSA-DOM III in the neutral to slightly alkaline ph region decreases, whereas the emission maximum increases slightly but reproducibly from 304 nm at ph 6.0 to 305 nm at ph 9.0. By raising the ph, a shoulder near 350 nm becomes obvious, which indicates ionization of some tyrosyl side chains (data not shown) (25). Near-UV CD To study the structural alterations in more detail, CD measurements in the near-uv region, which infer to tertiary structure, were performed at specific ph values, and the results are shown in Fig. 3, A D, for HSA and each of the three recombinant domains. The near-uv CD spectra of HSA are shown in Fig. 3A.AtpH 7.4, minima at 262 and 268 nm are observed as well as two shoulders near 276 and 283 nm, which is in accordance with previous findings (26 28). By reducing the ph to 4.0 and further to 2.0 there is an increase in the ellipticity at 262 and 268 nm and a decrease between 290 and 300 nm, denoting loss of tertiary structure in both the N-F transition and the acid expansion, in agreement with the alterations in secondary structure and tryptophanyl fluorescence. These findings agree with results published previously (29, 30). However at ph 2.0 there are still significant CD signals left, compared with the spectrum at 9.5 M urea, suggesting remaining tertiary structure even at this low ph. By changing the ph of the HSA solution from 7.4 to 9.0, we observe a slight gain in the CD signal at 262 and 268 nm, which might reflect perturbations around the numerous disulfide bridges (22, 30). In addition, a small reduction in ellipticity in the region between 280 and 300 nm is seen, which was ascribed to changes in the tryptophanyl environment (22, 30) and which is corroborated here by the changes in tryptophanyl fluorescence that we observed in this ph region. Fig. 3B presents the near-uv CD spectra of HSA-DOM I. Like HSA at ph 7.4, two minima at 262 and 268 nm and a shoulder at 276 nm are seen, and an additional shoulder near 255 nm is evident. Upon reduction of the ph of the solution to 4.0 the spectrum of HSA-DOM I exhibits an increase of [ ] 262 and [ ] 268, a flattening of the shoulder at 255 nm, and a decrease in the region above 280 nm with loss in fine structure. This is a clear indication for a structural rearrangement of HSA-DOM I in the ph region of the N-F transition of HSA, which is further emphasized by the pronounced increase in tyrosyl fluorescence intensity and the shifting emission maximum. By a further decrease of the ph to 2.0, an overall increase in ellipticity compared with the spectrum at ph 4.0 is perceivable, indicating further loss of tertiary structure elements of this domain in the ph region of the F-E transition of HSA. At 9.5 M urea, a dramatic increase in ellipticity was observed. The difference between the spectrum at ph 2.0 and 9.5 M urea denotes that even at ph 2.0 some tertiary structural elements are left. By raising the ph of the HSA-DOM I solution to 9.0, the CD signal at 262 and 268 nm increases slightly and the shoulder near 255 nm flattens, whereas there is a decrease in the region above 285 nm, highlighting the contribution of HSA- DOM I to the N-B transition of HSA. The near-uv CD spectra of HSA-DOM II at different ph values and at 9.5 M urea are presented in Fig. 3C. Similar to HSA the spectrum at ph 7.4 has minima at 262 and 268 nm, shoulders at 276 and 283, and an additional shoulder at 287 nm. As the ph is reduced to 4.0, there is an overall loss in ellipticity below 300 nm, and additional minima at 282 and 291 nm arise. These alterations in the near-uv CD spectra of HSA-DOM II provide evidence for major fluctuations in the tertiary structure and may reflect the changes in the environment of the tryptophanyl residue, which are documented by the 2.4-fold increase in fluorescence intensity as well as the secondary structural changes observed by far-uv CD in the same ph region. At ph 2.0 the ellipticities at 283 and 295 nm are only slightly increased, whereas the changes at 262 and 268 nm are more pronounced, indicating further loss in asymmetry around disulfide bridges and/or aromatic residues. The spectrum of HSA-DOM II in 9.5 M urea is different in shape compared with that at ph 2.0; however, the signal strength is similar, denoting that only minor tertiary structure elements are left at ph 2.0, in contrast to HSA and the other two domains. By increasing the ph of the solution to 9.0, a slight enhancement of the ellipticity at 262 and 268 nm is evident. The shoulder at 287 nm is transformed to a minimum and an additional shoulder at 295 nm arises. These findings and the changes in the tryptophanyl fluorescence intensity and emission maximum suggest that the tertiary structure is changing in the alkaline ph region, whereas there are only small fluctuations in secondary structure (Fig. 1B). In Fig. 3D the near-uv CD spectra of HSA-DOM III are shown. Similar to HSA, minima at 262 and 268 nm and shoulders near 276 and and near 283 nm are observed at ph 7.4. By reduction of the ph from 7.4 to 4.0, the ellipticity below 295 nm increases, and a loss in fine structure is detectable in the
5 3046 Conformational Transitions of Recombinant Albumin Domains FIG. 3.Near-UV CD spectra of HSA (A), HSA-DOM I (B), HSA-DOM II (C), and HSA-DOM III (D) at ph 7.4 ( ), 9.0 (- -), 4.0 (- -), 2.0 (- -), and 9.5 M urea ( ), 25 C, protein concentration 20 M. wavelength region between 295 and 270 nm. As the ph is diminished to 2.0, ellipticities below 310 nm are further enhanced, however less pronounced than at 9.5 M urea, providing evidence that even at ph 2.0 some tertiary structure elements are left. Increasing the ph from 7.4 to 9.0 leads only to modest changes, which indicates only minor perturbations of tertiary structure in the ph region of the N-B transition of HSA. This clearly demonstrates that the major structural rearrangement in the N-B transition of HSA is not likely to happen in domain III of the native albumin molecule. DISCUSSION We described previously the production and characterization of the three recombinant domains of HAS, and we discussed in depth their potential as models for the albumin molecule with an emphasis on studying their basic structural features and ligand binding activities (3). The present study highlights the structural integrity and the behavior under varying ph conditions of these new proteins in comparison with intact albumin. Measurements with HSA were used throughout our experiments as a reference in order to compare all observations made
6 Conformational Transitions of Recombinant Albumin Domains 3047 with the three recombinant domains directly to the data obtained with the native protein. In general, good agreement was achieved with published data for HSA. Far-UV CD In the ph region of the N-F transition (ph 5.0 to 3.5), a reduction of helical structure occurs in HSA, whereas the content of sheet structure increases (Fig. 1B and Table I). These findings are in accordance with previously published observations for both HSA and BSA (4, 5, 29 31), where it has been proposed that the N-F transition might represent helix 3 and helix 3 coil transitions. In the acid-expansion region (ph 3.5 to 2.5), we found a further marked reduction in the helix content and an increase in sheet structure which is in agreement with Era et al. (4, 5) and Sogami et al. (30). However, in contrast to these publications we did not find a significant increase in random coil content, and the changes that we observed were not as pronounced as those found by these authors. Furthermore, the onset of the N-F transition occurred about 0.5 ph units earlier in our experiments, which could be ascribed to the different buffer systems that were used in the ph range of the N-F transition. We used acetic acid/sodium acetate in the ph region of 4.5 to 3.5, whereas Era et al. (4, 5) and Sogami et al. (30) adjusted the ph with HCl in 0.1 M NaCl or KCl. Interaction of undissociated acetic acid by hydrogen bonding to carboxylic groups of proteins was demonstrated for BSA and other proteins as revealed by electrophoresis (32 34) and CD (35, 36). Furthermore, Leonard and Foster (37) showed that the ph profile of the N-F transition of BSA in acetic acid/acetate differs from that in chloride. In addition, the earlier onset of the N-F transition of the HSA used in our experiments could be due to differences in the preparation method, which is known to contribute to the structural behavior of albumin (38 40) and/or to the absence of added salt and bound fatty acids, shifting the midpoint of the N-F transition of BSA to higher ph (41). A number of interesting observations was made with the HSA domains in the acid ph range. A high degree of similarity in behavior to the native protein was observed with HSA-DOM III, which displayed a two-step transition with a decrease in helical and an increase in sheet content (Fig. 1B and Table I). However, the two transitions are more distinctly separated from each other than the N-F, and the F-E transitions are in HSA under our conditions; furthermore, the onset of the first structural change occurs earlier and that of the second later than in our control experiments with the native protein. The crystal structure of HSA strongly suggests at least five interdomain salt bridges and six hydrogen bondings for domain III at neutral ph. The fact that HSA-DOM III in its recombinant form is missing these stabilizing interactions may be an explanation for the earlier onset of the first and the delay of the second transition in HSA-DOM III. This notion is further supported by the findings of Sogami et al. (30, 42), who reported for BSA that the initial part of the N-F transition was shifted to higher ph and the acid expansion was moved to lower ph by increasing the salt concentration, especially by using strongly binding anions such as ClO 4, which dissociate hydrophilic and strengthen hydrophobic interfaces. Based on these findings we conclude that the major structural changes in the N-F transition of HSA arise from domain III, as has previously been shown for a C-terminal fragment of BSA (13) underlining that HSA-DOM III as a stand-alone protein clearly reflects the structural behavior of the C terminus of the native molecule. With HSA-DOM I only a minor decrease in helix content is observed in the region of the N-F transition of HSA, whereas a marked structural change occurs in the ph region of the acid expansion of HSA (Fig. 1B and Table I). This is in good agreement with the concept that the albumin molecule is fully extended in the E-form, resulting from a loss of inter-domain and inter-subdomain contacts and a disruption of the structure in the hinge and link regions (1, 2). HSA-DOM II undergoes a rearrangement of secondary structure in the ph region between 6.5 and 4.0 (Fig. 1B), which surprisingly leads to a slight increase in helical content and a reduction in sheet structure (Table I). As we have previously shown (3) the isoelectric point of HSA-DOM II is 5.4, which is almost the midpoint of this transition. It is conceivable that near the isoelectric point, where ionic repulsion between side chains of the same charge is minimized, hydrophobic interactions result in a rearrangement of secondary structure. This suggests that HSA-DOM II has a very flexible structure, which is not surprising since this domain has lost contacts to the two neighboring domains, leading to two previously unexposed surface patches, one at its N terminus and, more importantly, the second in the central part of HSA-DOM II as judged from inspection of the crystal structure. By further lowering the ph, an acid expansion-like transition is observed, which is, however, not as pronounced as compared with HSA-DOM I, HSA- DOM III, and HSA. The partial structural restoration as it is observed with all four proteins at strong acidic ph can be explained by a minimization of the hydrophobic surface area, which is promoted by neutralization of positively charged repulsive forces by chloride anions of HCl, as has been reported by Goto et al. (20) and Fink et al. (21) for many proteins. In the alkaline ph range between ph 7.4 and 9.0, HSA displays a slight reduction in helical content and a small increase in sheet structure, whereas the changes in coil structure are not significant (Fig. 1B and Table I). These findings are in good agreement with the data for BSA reported by Era et al. (22) in the absence of salt. The recombinant domains did not show significant alterations in their secondary structure content in the alkaline ph region (Fig. 1B and Table I). The N-B isomerization is interpreted to be a structural fluctuation, a loosening of the molecule with higher configurational adaptability (2). The B-form in the absence of added salt might be a state where mutual movement of subdomains, connected by flexible hinge regions, takes place, accompanied by fluctuations of the structures of the subdomains themselves relative to each other (22). It is therefore possible that the main losses in secondary structure are affecting the two interdomain helices (h10 DOM I -h1 DOM II, h10 DOM II -h1 DOM III ) of HSA and that the secondary structural integrity of a domain per se is not impaired in the N-B transition as supported by our data presented here. Tryptophanyl Fluorescence To explore further the unexpected structural changes of HSA-DOM II, tryptophanyl fluorescence measurements were performed, since the lone tryptophanyl residue of HSA is located almost centrally in helix 2 of domain II. The position of the energy maximum ( max ) from the arising emission spectrum depends on the properties of the environment of the tryptophanyl residue (43). The fluorescence intensity depends upon the degree of exposure of the tryptophanyl side chain to the polar, aqueous solvent and upon its proximity to specific quenching groups, such as protonated carboxyl, protonated imidazole, deprotonated -amino groups, and tyrosinate anions (44, 45). With HSA, we obtained ph profiles of Fl em max (where Fl is fluorescence intensity) in the acid ph region showing a twostep change (Fig. 2A), one corresponding to the N-F transition and the other to the acid expansion, as reported previously for HSA (16, 23, 24) and for BSA (46, 47). The major changes in Fl em max were complete at ph 4.5, which is approximately at the onset of the N-F transition under conditions where salt is added. It was reported (41) that delipidated BSA and albumin
7 3048 Conformational Transitions of Recombinant Albumin Domains without added salt show a shift of the N-F transition to higher ph regions. The observed shift of the energy maximum to shorter wavelength of HSA in the N-F transition (Fig. 2A) was attributed to sandwiching of the indole side chain in a rigid portion of the protein matrix (24). The ph-dependent decrease of the tryptophanyl fluorescence of HSA at the onset of the N-F transition may be attributed to quenching by protonated imidazole and carboxyl groups, resulting from protonation of such residues in the vicinity of Trp-214. Three amino acids with carboxylate side chains (Glu- 292 of domain II and Glu-450 and Asp-451 of domain III) and one histidine residue (His-242 of domain II) are found within 10 Å distance from Trp-214, based on the atomic coordinates of HSA (Protein Data Bank entries 1UOR (48) and 1AO6 (49)). In addition, Cowgill (50) reported that loss of helical conformation is correlated with a decrease in fluorescence intensity, which is the case in the N-F transition, as discussed in the far-uv CD section. The reduction in fluorescence intensity below ph 3.5 may arise from quenching by protonation of carboxylate groups, whereas the increase in fluorescence at ph values below 2.5 can be attributed to restoration of secondary structure at strong acidic ph without added salt (20, 21), which was reported to trigger fluorescence intensity (50). Consequently, a more hydrophobic environment of Trp-214 may be created, leading to increased tryptophanyl fluorescence intensity as described by Halfman and Nishida (44, 45). The observed decreases of max and fluorescence intensity of HSA in the ph region from 7.4 to 9.0 (Fig. 2A) might either be due to changes in the secondary structure as discussed in the far-uv CD section and/or to quenching of the tryptophanyl fluorophor by deprotonation of groups that are in close proximity to the indole chromophore, such as -amino groups of lysyl residues. Two such residues are found in the crystal structures of HSA, viz. Lys-199 at 3.7 Å and Lys-195 at 7.4 Å distance to Trp-214. Profound differences between the fluorescence behavior of the lone tryptophanyl residue in HSA-DOM II and in HSA at ph 7.4 are highlighted by the 2.6-fold lower quantum yield of HSA-DOM II and the pronounced shift of the emission maximum to shorter wavelength (Fig. 2A), denoting a dramatically changed microenvironment of this residue in HSA-DOM II. As can be deduced from the crystal structures of human serum albumin, Trp-214 acts as an important stabilizer (by hydrophobic packing force) for the interface between subdomain IIA and IIIA (1). This hydrophobic area, which is centered in the crystal structure of domain II, loses its natural counterpart when expressed in the form of a single domain, leading to modified water accessibility of the tryptophanyl residue in the recombinant protein, thereby affecting the alterations in fluorescence behavior of the tryptophanyl residue in HSA-DOM II. The reduced fluorescence intensity may indicate an impairment of the helix harboring the lone tryptophanyl in the native molecule, which is supported by the reduced helical content of HSA-DOM II compared with HSA at ph 7.4 (Table I). By using light meromyosin, a helical muscle protein, Cowgill (50) demonstrated a 3-fold reduction of the tryptophanyl fluorescence intensity by disaggregation of the helical conformation and attributed this observation to the quenching by peptide carbonyl groups in disordered polypeptides. The shifted wavelength maximum of HSA-DOM II compared with HSA may be due to a more hydrophobic environment of the tryptophanyl residue and/or sandwiching of the side chain in the rigid portion of the protein matrix, as has been described Eftink and Ghiron (24) for HSA. In the weak acid ph region, there is a sigmoidal 2.4-fold increase in fluorescence intensity of HSA-DOM II ending near ph 4.0 (Fig. 2A). Interestingly, at ph 4.0 HSA-DOM II and HSA have almost superimposable tryptophanyl fluorescence spectra suggesting similar microenvironments encircling this residue in both proteins. The pronounced increase in fluorescence intensity of HSA-DOM II between ph 7.4 and 4.0 is correlated with the secondary structural rearrangement observed by far-uv CD in the same ph region. It has been reported that loss of helical conformation resulted in decreased fluorescence (see above) (50). Conversely, we detect an increased -helical content with decreasing ph (Fig. 1B and Table I), which therefore may trigger the fluorescence of the tryptophanyl residue in HSA-DOM II. In addition, Steiner and Edelhoch (51) reported that an increase in intensity in the acid ph region is very likely to be connected with a definite conformational change in the protein structure, because all side chains that are known to quench the tryptophanyl fluorescence intensity become protonated in this ph region and should therefore quench the fluorescence (tryptophanyl fluorescence quenching groups: COOH, protonated imidazole, deprotonated -NH 2, SH, S S, tyrosinate anion). The spectra of the tryptophanyl residues of HSA and HSA- DOM II in 9.5 M urea (data not shown) are superimposable and show reduced intensity caused by collisional quenching with the surrounding solvent molecules with an emission maximum red-shifted to approximately 349 nm, which is characteristic for a fully exposed tryptophanyl residue (24, 52). In the weak alkaline ph range, the fluorescence intensity of HSA-DOM II increases, which is accompanied by a shift of the emission maximum to higher wavelength (Fig. 2A). The decrease in fluorescence quenching in this ph region may be due to the deprotonation of imidazole side chains with unusually high pk, which are known to be located in domain II of HSA (17); however, the reduction of the emission maximum suggests a structural transition in the zone surrounding the tryptophanyl residue in HSA-DOM II. Tyrosyl Fluorescence The fluorescence of proteins originates almost entirely from the tyrosyl and tryptophanyl residues. In proteins that lack tryptophanyl residues (class A, tryptophanyl-free proteins), tyrosyl fluorescence is used as a probe for conformational changes (53). HSA-DOM I harbors seven tyrosyl residues. As indicated by the crystal structures (Protein Data Bank entries: 1UOR (48) and 1AO6 (49)) four of them (residues 30, 138, 140, and 161) are located in helical regions, and all of them are at least minimally exposed to the solvent. We observed a significant increase in tyrosyl fluorescence intensity of HSA-DOM I in the ph region from 6.0 to 3.5 (Fig. 2B). Inspection of the atomic coordinates of HSA shows that two carboxylate side chains, Asp-38 and Asp- 108, are within 5 Å of Tyr-84 and Tyr-148, respectively. As has been demonstrated by Cowgill (54), carboxylate groups serve as acceptors for the phenolic proton in the excited state and thus are among the known quenchers for tyrosyl fluorescence. In addition, it was reported that tyrosyl fluorescence is strongly quenched by hydrogen bonding of the phenolic hydroxyl to carbonyl or amide groups that could be supplied by the protein backbone and/or by side chains of asparagine and glutamine next to tyrosyl residues (53, 55, 56). As a matter of fact, hydrogen bonds between the OH group of Tyr-30 and the amide side chain of Asn-99, as well as between the OH group of Tyr-84 and backbone carbonyl of Gln-33 are strongly indicated by the atomic coordinates of HSA. Therefore, protonation of the aspartate side chains mentioned above accompanied by structural changes which lead to disruption of these hydrogen bondings may explain the observed rise in fluorescence intensity. All this suggests a rearrangement of tertiary rather than of secondary structure of HSA-DOM I in the region of the N-F transition of HSA, which is corroborated by the results of near-uv CD measurements discussed below.
8 Conformational Transitions of Recombinant Albumin Domains 3049 Three out of the four tyrosyl residues of HSA-DOM III are known to be located in helical regions (residues 401, 411, and 452 of HSA) (Protein Data Bank entries 1UOR (48) and 1AO6 (49)). As the ph of a HSA-DOM III solution is reduced to ph 4.0 there is a sharp sigmoidal decrease in the tyrosyl fluorescence intensity and a slight increase in the emission maximum (Fig. 2B). This decrease in fluorescence intensity may be due to a loss of helical structure around the three tyrosyl residues located in -helical regions, in conformity with the pronounced reduction of helical content that we observed in the identical ph range, as described above in the far-uv CD section. Similar observations have been made by Cowgill (54) in the case of helical muscle proteins. In addition, no carboxylate side chains that could potentially trigger the fluorescence by their protonation can be found in 5 Å surrounding the four tyrosyl residues. By further reduction of the ph of the solution, the fluorescence rises continuously, which may be ascribed to the disruption of hydrogen bondings between the hydroxyl groups of tyrosyl residues in HSA-DOM III and carbonyl groups of the peptide backbone (Tyr-452 to Asn-429 and Tyr-497 to Lys-534) (55) emphasized by the loss of secondary structure of HSA- DOM III in the F-E transition. By increasing the ph of a HSA-DOM III solution, the intrinsic tyrosyl fluorescence decreases, whereas a shoulder in the emission spectrum near 350 nm occurs (data not shown). These findings strongly suggest the deprotonation of the phenolic hydroxyl group of some tyrosyl residues in HSA-DOM III, as was suggested previously (25, 54, 57). In addition, Edelhoch et al. (57) reported that significant inhibition of phenol emission occurs by uncharged amino groups. In this ph region some lysyl residues may be deprotonated and lead to quenching of tyrosyl fluorescence. In fact, in the crystal structure of HSA, amino groups of lysyl residues are found in close proximity (5 Å surrounding) to all 4 tyrosyl residues of HSA-DOM III: Tyr-401 to Lys-525, Tyr-411 to Lys-414, Tyr-452 to Lys-432 and Tyr-497 to Lys-534. All these findings and the fact that HSA-DOM III undergoes only minor changes in secondary structure in the slightly alkaline ph region provide evidence that the observed decrease in fluorescence intensity of HSA-DOM III is not correlated with a structural change, which in addition is strongly supported by the near-uv CD results discussed below. Near-UV CD CD spectra in the near-uv region provide information about the asymmetry of the structure around the aromatic amino acid side chains and disulfide bridges and therefore on fluctuations in the tertiary structure of a protein. The main contributions to the ellipticity from tryptophanyl and tyrosyl groups are obvious above 265 nm, with the largest maxima around 279, 284, and 291 nm for tryptophanyl, around 277 nm for tyrosyl, and around 255, 261, and 268 nm for phenylalanine residues. The ellipticity of S S bridges in proteins can be significant from 320 to 250 nm (58 61). The observed alterations in ellipticity of HSA in the acid ph range (Fig. 3A) reflect the tertiary structural alterations in the N-F and F-E transition, in correlation with the loss of secondary structure described above. These findings are in close agreement with previous results obtained with BSA (30). The observed decrease of [ ] in the spectrum at ph 4.0 was ascribed to the immobilization of the tryptophanyl residues of BSA (30). Leonard and Foster (37) reported that approximately four buried tyrosyl residues of BSA appear to become solventexposed during the acid expansion, whereas none do so in the N-F transition. This may, in addition to the environmental perturbations of the disulfide bridges, be an explanation for the overall enhancement of the ellipticity and the loss in fine structure below 280 nm between ph 4.0 and 2.0. The residual near-uv CD signals at ph 2.0, which are significant compared with the spectrum at 9.5 M urea, strongly suggest the existence of tertiary packing, at least around aromatic residues and disulfide bridges even at this low ph. The alterations in the spectra between ph 7.4 and 9.0 demonstrate tertiary structural changes in the N-B transition of HSA (Fig. 3A). This was ascribed to environmental perturbations of the 17 disulfides in BSA, such as variations in the dihedral angles, with some contribution of phenylalanine and tyrosyl residues (29, 30), which were shown by UV difference spectroscopy to gain increased accessibility in this ph range (62). In addition, a slight reduction in ellipticity in the region between 280 and 300 nm emerged, which might be due to an immobilization of the tryptophanyl side chain, as was described in the case of the N-F transition of BSA by Sogami et al. (30) and is supported by the observed changes in the tryptophanyl fluorescence discussed above. The spectra of the three recombinant domains are dominated by the large number of disulfide bridges analogous to HSA, whereas some difference in fine structure is obvious, which may be owing to different relative amounts of aromatic amino acid residues (Fig. 3, B D). The general similarity of the spectra of HSA, HSA-DOM I, HSA-DOM II, and HSA-DOM III at ph 7.4 reflects their structural, evolutionary-based relationship and emphasizes that the recombinant domains as standalone proteins have the ability to adopt folds that are similar to their respective structure in the context of the whole protein. In the ph region of the N-F transition of HSA, all three domains display increases in ellipticity of the minima at 262 and 268 nm (Fig. 3, B D), which may be due to loss of asymmetry around disulfide bridges and/or aromatic residues, indicating disruption of tertiary structure. In addition, major global changes with additional minima in the spectra of HSA- DOM II occur between ph 7.4 and 4.0. This may reflect dramatic changes in the tryptophanyl environment, supported by the observed 2.4-fold increase in fluorescence intensity and the secondary structural changes (Fig. 2A and Fig. 1B). Whereas the ph-dependent changes of the spectra of HSA- DOM II and HSA-DOM III coincide with the observed alterations in secondary structure and tryptophanyl and tyrosyl fluorescence respectively, the most striking result of our CD measurements in the near ultraviolet region is the fact that HSA-DOM I undergoes fluctuations in tertiary structure that are accompanied by only a small loss in secondary structure, which could so far only be deduced from experiments with larger fragments (12). HSA-DOM I shows flattening of the shoulder near 255 nm as the ph is diminished to 4.0 and also as it is increased to 9.0, indicating perturbations of phenylalanine residues. An opening of the crevice harboring Cys-34 has been reported to occur during both the N-F and the N-B transition of BSA (63, 64). The crystal structure of HSA shows that Phe-26 and Phe-37 are located in the same crevice, and it is therefore possible that the observed flattening of the shoulder at 255 nm may partly be attributed to perturbations of these particular residues. By reducing the ph further, all three recombinant proteins lose tertiary structure as indicated by the rise in ellipticity at 262 and 268 nm and the loss of fine structure (Fig. 3, B D), in agreement with the observed decrease of the secondary structure content in the region of the acid expansion of HSA. By inspection of the spectra of HSA-DOM I, HSA-DOM II, and HSA-DOM III at ph 2.0 and 9.5 M urea, which represents the fully denaturated state as indicated by the spectra in the far-uv region (Fig. 1A), it becomes evident that HSA-DOM I and HSA-DOM III, similar to HSA, display significant tertiary structural elements even at ph 2.0. It has been reported for some proteins (20, 21), especially for those that are heavily
UV Fluorescence Polarization as a Means to Investigate Protein Conformational and Mass Change
 A p p l i c a t i o n N o t e UV Fluorescence Polarization as a Means to Investigate Protein Conformational and Mass Change Using Intrinsic Tryptophan Fluorescence in Conjunction with UV-capable Polarizers
A p p l i c a t i o n N o t e UV Fluorescence Polarization as a Means to Investigate Protein Conformational and Mass Change Using Intrinsic Tryptophan Fluorescence in Conjunction with UV-capable Polarizers
DENATURATION OF HEMOGLOBIN IN THE PRESENCE OF TANNIC ACID
 PROCEEDINGS OF THE YEREVAN STATE UNIVERSITY C h e m i s t r y a n d B i o l o g y 2014, 1, p. 23 27 C h emistr y DENATURATION OF HEMOGLOBIN IN THE PRESENCE OF TANNIC ACID K. R. GRIGORYAN, L. S. SARGSYAN
PROCEEDINGS OF THE YEREVAN STATE UNIVERSITY C h e m i s t r y a n d B i o l o g y 2014, 1, p. 23 27 C h emistr y DENATURATION OF HEMOGLOBIN IN THE PRESENCE OF TANNIC ACID K. R. GRIGORYAN, L. S. SARGSYAN
Proteins Amides from Amino Acids
 Chapter 26 and Chapter 28 Proteins Amides from Amino Acids Amino acids contain a basic amino group and an acidic carboxyl group Joined as amides between the ¾NH 2 of one amino acid and the ¾CO 2 H to the
Chapter 26 and Chapter 28 Proteins Amides from Amino Acids Amino acids contain a basic amino group and an acidic carboxyl group Joined as amides between the ¾NH 2 of one amino acid and the ¾CO 2 H to the
The study of protein secondary structure and stability at equilibrium ABSTRACT
 The study of protein secondary structure and stability at equilibrium Michelle Planicka Dept. of Physics, North Georgia College and State University, Dahlonega, GA REU, Dept. of Physics, University of
The study of protein secondary structure and stability at equilibrium Michelle Planicka Dept. of Physics, North Georgia College and State University, Dahlonega, GA REU, Dept. of Physics, University of
The mechanism(s) of protein folding. What is meant by mechanism. Experimental approaches
 The mechanism(s) of protein folding What is meant by mechanism Computational approaches Experimental approaches Questions: What events occur and in what time sequence when a protein folds Is there a specified
The mechanism(s) of protein folding What is meant by mechanism Computational approaches Experimental approaches Questions: What events occur and in what time sequence when a protein folds Is there a specified
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Last modified 29 September 2005
 Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Last modified 9 September 005 Focus concept Purification of a novel seed storage protein allows sequence analysis and
Case 7 A Storage Protein From Seeds of Brassica nigra is a Serine Protease Inhibitor Last modified 9 September 005 Focus concept Purification of a novel seed storage protein allows sequence analysis and
Fundamentals of Protein Structure
 Outline Fundamentals of Protein Structure Yu (Julie) Chen and Thomas Funkhouser Princeton University CS597A, Fall 2005 Protein structure Primary Secondary Tertiary Quaternary Forces and factors Levels
Outline Fundamentals of Protein Structure Yu (Julie) Chen and Thomas Funkhouser Princeton University CS597A, Fall 2005 Protein structure Primary Secondary Tertiary Quaternary Forces and factors Levels
Nanobiotechnology. Place: IOP 1 st Meeting Room Time: 9:30-12:00. Reference: Review Papers. Grade: 50% midterm, 50% final.
 Nanobiotechnology Place: IOP 1 st Meeting Room Time: 9:30-12:00 Reference: Review Papers Grade: 50% midterm, 50% final Midterm: 5/15 History Atom Earth, Air, Water Fire SEM: 20-40 nm Silver 66.2% Gold
Nanobiotechnology Place: IOP 1 st Meeting Room Time: 9:30-12:00 Reference: Review Papers Grade: 50% midterm, 50% final Midterm: 5/15 History Atom Earth, Air, Water Fire SEM: 20-40 nm Silver 66.2% Gold
Fluorescence Quenching of Human Serum Albumin by Caffeine
 CHEM 411L Instrumental Analysis Laboratory Revision 2.1 Fluorescence Quenching of Human Serum Albumin by Caffeine In this laboratory exercise we will examine the fluorescence of Human Serum Albumin (HSA)
CHEM 411L Instrumental Analysis Laboratory Revision 2.1 Fluorescence Quenching of Human Serum Albumin by Caffeine In this laboratory exercise we will examine the fluorescence of Human Serum Albumin (HSA)
Structure formation and association of biomolecules. Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München
 Structure formation and association of biomolecules Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München Motivation Many biomolecules are chemically synthesized
Structure formation and association of biomolecules Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München Motivation Many biomolecules are chemically synthesized
MOLEBIO LAB #3: Electrophoretic Separation of Proteins
 MOLEBIO LAB #3: Electrophoretic Separation of Proteins Introduction: Proteins occupy a central position in the structure and function of all living organisms. Some proteins serve as structural components
MOLEBIO LAB #3: Electrophoretic Separation of Proteins Introduction: Proteins occupy a central position in the structure and function of all living organisms. Some proteins serve as structural components
 Problem: The GC base pairs are more stable than AT base pairs. Why? 5. Triple-stranded DNA was first observed in 1957. Scientists later discovered that the formation of triplestranded DNA involves a type
Problem: The GC base pairs are more stable than AT base pairs. Why? 5. Triple-stranded DNA was first observed in 1957. Scientists later discovered that the formation of triplestranded DNA involves a type
Purification: Step 1. Protein and Peptide Chemistry. Lecture 11. Big Problem: Crude extract is not the natural environment. Cells: Break them open!
 Lecture 11 Protein and Peptide Chemistry Margaret A. Daugherty Fall 2003 Purification: Step 1 Cells: Break them open! Crude Extract Total contents of cell Big Problem: Crude extract is not the natural
Lecture 11 Protein and Peptide Chemistry Margaret A. Daugherty Fall 2003 Purification: Step 1 Cells: Break them open! Crude Extract Total contents of cell Big Problem: Crude extract is not the natural
Purification: Step 1. Lecture 11 Protein and Peptide Chemistry. Cells: Break them open! Crude Extract
 Purification: Step 1 Lecture 11 Protein and Peptide Chemistry Cells: Break them open! Crude Extract Total contents of cell Margaret A. Daugherty Fall 2003 Big Problem: Crude extract is not the natural
Purification: Step 1 Lecture 11 Protein and Peptide Chemistry Cells: Break them open! Crude Extract Total contents of cell Margaret A. Daugherty Fall 2003 Big Problem: Crude extract is not the natural
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL Materials and Methods Circular dichroism (CD) spectroscopy. Far ultraviolet (UV) CD spectra of apo- and holo- CaM and the CaM mutants were recorded on a Jasco J-715 spectropolarimeter
SUPPLEMENTARY MATERIAL Materials and Methods Circular dichroism (CD) spectroscopy. Far ultraviolet (UV) CD spectra of apo- and holo- CaM and the CaM mutants were recorded on a Jasco J-715 spectropolarimeter
Amino Acids. Amino Acid Structure
 Amino Acids Pratt & Cornely Chapter 4 Alpha carbon Sidechain Proteins peptides Amino Acid Structure 1 L amino acids Glycine R/S vs D/L L isoleucine racemization Stereochemisty Common Amino Acids 2 Which
Amino Acids Pratt & Cornely Chapter 4 Alpha carbon Sidechain Proteins peptides Amino Acid Structure 1 L amino acids Glycine R/S vs D/L L isoleucine racemization Stereochemisty Common Amino Acids 2 Which
Quantitative Evaluation of the Ability of Ionic Liquids to Offset the Cold- Induced Unfolding of Proteins
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2014 Supplimentary informations Quantitative Evaluation of the Ability of Ionic Liquids
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2014 Supplimentary informations Quantitative Evaluation of the Ability of Ionic Liquids
Protein Structure and Function! Lecture 4: ph, pka and pi!
 Protein Structure and Function! Lecture 4: ph, pka and pi! Definition of ph and pk a! ph is a measure of the concentration of H +.! + ph = log 10[H ] For a weak acid,! HA #!!"! H + + A!, K a = [H + ][A!
Protein Structure and Function! Lecture 4: ph, pka and pi! Definition of ph and pk a! ph is a measure of the concentration of H +.! + ph = log 10[H ] For a weak acid,! HA #!!"! H + + A!, K a = [H + ][A!
Addison Ault, Department of Chemistry, Cornell College, Mount Vernon, IA. The isoelectric point is a point on the ph scale. It is the ph at which the
 1 Isoelectric Points On the Back of the Envelope Addison Ault, Department of Chemistry, Cornell College, Mount Vernon, IA The isoelectric point is a point on the ph scale. It is the ph at which the average
1 Isoelectric Points On the Back of the Envelope Addison Ault, Department of Chemistry, Cornell College, Mount Vernon, IA The isoelectric point is a point on the ph scale. It is the ph at which the average
BIL 256 Cell and Molecular Biology Lab Spring, 2007 BACKGROUND INFORMATION I. PROTEIN COMPOSITION AND STRUCTURE: A REVIEW OF THE BASICS
 BIL 256 Cell and Molecular Biology Lab Spring, 2007 BACKGROUND INFORMATION I. PROTEIN COMPOSITION AND STRUCTURE: A REVIEW OF THE BASICS Proteins occupy a central position in the structure and function
BIL 256 Cell and Molecular Biology Lab Spring, 2007 BACKGROUND INFORMATION I. PROTEIN COMPOSITION AND STRUCTURE: A REVIEW OF THE BASICS Proteins occupy a central position in the structure and function
CFSSP: Chou and Fasman Secondary Structure Prediction server
 Wide Spectrum, Vol. 1, No. 9, (2013) pp 15-19 CFSSP: Chou and Fasman Secondary Structure Prediction server T. Ashok Kumar Department of Bioinformatics, Noorul Islam College of Arts and Science, Kumaracoil
Wide Spectrum, Vol. 1, No. 9, (2013) pp 15-19 CFSSP: Chou and Fasman Secondary Structure Prediction server T. Ashok Kumar Department of Bioinformatics, Noorul Islam College of Arts and Science, Kumaracoil
Introduction to Proteins
 Introduction to Proteins Lecture 4 Module I: Molecular Structure & Metabolism Molecular Cell Biology Core Course (GSND5200) Matthew Neiditch - Room E450U ICPH matthew.neiditch@umdnj.edu What is a protein?
Introduction to Proteins Lecture 4 Module I: Molecular Structure & Metabolism Molecular Cell Biology Core Course (GSND5200) Matthew Neiditch - Room E450U ICPH matthew.neiditch@umdnj.edu What is a protein?
Bi 8 Lecture 7. Ellen Rothenberg 26 January Reading: Ch. 3, pp ; panel 3-1
 Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
AnaTag HiLyte Fluor 647 Protein Labeling Kit
 AnaTag HiLyte Fluor 647 Protein Labeling Kit Catalog # 72049 Kit Size 3 Conjugation Reactions This kit is optimized to conjugate HiLyte Fluor 647 SE to proteins (e.g., IgG). It provides ample materials
AnaTag HiLyte Fluor 647 Protein Labeling Kit Catalog # 72049 Kit Size 3 Conjugation Reactions This kit is optimized to conjugate HiLyte Fluor 647 SE to proteins (e.g., IgG). It provides ample materials
Types of chromatography
 Chromatography Physical separation method based on the differential migration of analytes in a mobile phase as they move along a stationary phase. Mechanisms of Separation: Partitioning Adsorption Exclusion
Chromatography Physical separation method based on the differential migration of analytes in a mobile phase as they move along a stationary phase. Mechanisms of Separation: Partitioning Adsorption Exclusion
PROCEDURE FOR USE NICKEL NTA Magnetic Agarose Beads (5%)
 1 AFFINITY HIS-TAG PURIFICATION PROCEDURE FOR USE NICKEL NTA Magnetic Agarose Beads (5%) DESCRIPTION Nickel NTA Magnetic Agarose Beads are products that allow rapid and easy small-scale purification of
1 AFFINITY HIS-TAG PURIFICATION PROCEDURE FOR USE NICKEL NTA Magnetic Agarose Beads (5%) DESCRIPTION Nickel NTA Magnetic Agarose Beads are products that allow rapid and easy small-scale purification of
Proteins Higher Order Structures
 Proteins Higher Order Structures Dr. Mohammad Alsenaidy Department of Pharmaceutics College of Pharmacy King Saud University Office: AA 101 msenaidy@ksu.edu.sa Previously on PHT 426!! Protein Structures
Proteins Higher Order Structures Dr. Mohammad Alsenaidy Department of Pharmaceutics College of Pharmacy King Saud University Office: AA 101 msenaidy@ksu.edu.sa Previously on PHT 426!! Protein Structures
School of Chemistry and Chemical Engineering, Sun Yat-Sen University, Guangzhou , P.R.China
 Sequence-specific recognition of double-stranded DNA with molecular beacon with the aid of Ag + under neutral ph environment Zhiyou Xiao, Xiaoting Guo, Liansheng Ling * School of Chemistry and Chemical
Sequence-specific recognition of double-stranded DNA with molecular beacon with the aid of Ag + under neutral ph environment Zhiyou Xiao, Xiaoting Guo, Liansheng Ling * School of Chemistry and Chemical
1. Bloomsbury BBSRC Centre for Structural Biology, Birkbeck College and University College London.
 Purification/Polishing of His-tagged proteins - Application of Centrifugal Vivapure Ion-exchange Membrane Devices to the Purification/Polishing of Histagged Background Multi-milligram quantities of highly
Purification/Polishing of His-tagged proteins - Application of Centrifugal Vivapure Ion-exchange Membrane Devices to the Purification/Polishing of Histagged Background Multi-milligram quantities of highly
Rapid Kinetics with IR Protein folding examples
 Rapid Kinetics with IR Protein folding examples Time dependent data with FTIR Stop-flow methods - msec limits so far Continuous, micro-flow methods - < 100 µsec Rapid scan FT-IR - msec Multichannel laser
Rapid Kinetics with IR Protein folding examples Time dependent data with FTIR Stop-flow methods - msec limits so far Continuous, micro-flow methods - < 100 µsec Rapid scan FT-IR - msec Multichannel laser
Reducing Non-Specific Binding in Surface Plasmon Resonance Experiments
 1 Reducing Non-Specific Binding in Surface Plasmon Resonance Experiments SUMMARY Reducing non-specific binding (NSB) is essential to generating accurate data with SPR The effect of bovine serum albumin,
1 Reducing Non-Specific Binding in Surface Plasmon Resonance Experiments SUMMARY Reducing non-specific binding (NSB) is essential to generating accurate data with SPR The effect of bovine serum albumin,
Protein Folding Problem I400: Introduction to Bioinformatics
 Protein Folding Problem I400: Introduction to Bioinformatics November 29, 2004 Protein biomolecule, macromolecule more than 50% of the dry weight of cells is proteins polymer of amino acids connected into
Protein Folding Problem I400: Introduction to Bioinformatics November 29, 2004 Protein biomolecule, macromolecule more than 50% of the dry weight of cells is proteins polymer of amino acids connected into
Hmwk # 8 : DNA-Binding Proteins : Part II
 The purpose of this exercise is : Hmwk # 8 : DNA-Binding Proteins : Part II 1). to examine the case of a tandem head-to-tail homodimer binding to DNA 2). to view a Zn finger motif 3). to consider the case
The purpose of this exercise is : Hmwk # 8 : DNA-Binding Proteins : Part II 1). to examine the case of a tandem head-to-tail homodimer binding to DNA 2). to view a Zn finger motif 3). to consider the case
Packing of Secondary Structures
 7.88 Lecture Notes - 5 7.24/7.88J/5.48J The Protein Folding and Human Disease Packing of Secondary Structures Packing of Helices against sheets Packing of sheets against sheets Parallel Orthogonal Table:
7.88 Lecture Notes - 5 7.24/7.88J/5.48J The Protein Folding and Human Disease Packing of Secondary Structures Packing of Helices against sheets Packing of sheets against sheets Parallel Orthogonal Table:
Zool 3200: Cell Biology Exam 3 3/6/15
 Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
Name: Trask Zool 3200: Cell Biology Exam 3 3/6/15 Answer each of the following questions in the space provided; circle the correct answer or answers for each multiple choice question and circle either
Purification of (recombinant) proteins. Pekka Lappalainen, Institute of Biotechnology, University of Helsinki
 Purification of (recombinant) proteins Pekka Lappalainen, Institute of Biotechnology, University of Helsinki Physical properties of proteins that can be applied for purification -size -charge (isoelectric
Purification of (recombinant) proteins Pekka Lappalainen, Institute of Biotechnology, University of Helsinki Physical properties of proteins that can be applied for purification -size -charge (isoelectric
Supporting Information
 Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2014 Supporting Information Integration of Graphene Oxide and DNA as Universal Platform
Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2014 Supporting Information Integration of Graphene Oxide and DNA as Universal Platform
nanodsf 2bind: Your service provider for biophysical characterization of proteins Precisely revealing protein folding and stability
 nanodsf Precisely revealing protein folding and stability 2bind: Your service provider for biophysical characterization of proteins This booklet was written and designed by 2bind 08 2015 Any reproduction
nanodsf Precisely revealing protein folding and stability 2bind: Your service provider for biophysical characterization of proteins This booklet was written and designed by 2bind 08 2015 Any reproduction
RNA does not adopt the classic B-DNA helix conformation when it forms a self-complementary double helix
 Reason: RNA has ribose sugar ring, with a hydroxyl group (OH) If RNA in B-from conformation there would be unfavorable steric contact between the hydroxyl group, base, and phosphate backbone. RNA structure
Reason: RNA has ribose sugar ring, with a hydroxyl group (OH) If RNA in B-from conformation there would be unfavorable steric contact between the hydroxyl group, base, and phosphate backbone. RNA structure
Multi-Volume Based Protein Quantification
 A p p l i c a t i o n G u i d e Multi-Volume Based Methods Why Quantify Proteins? Proteins are central to our understanding of biology. In cells, they are multipurpose: from actin providing structural
A p p l i c a t i o n G u i d e Multi-Volume Based Methods Why Quantify Proteins? Proteins are central to our understanding of biology. In cells, they are multipurpose: from actin providing structural
A nucleic acid-based fluorescent sensor for expeditious detection of pyrophosphate anions at nanomolar concentrations
 Supporting Information for A nucleic acid-based fluorescent sensor for expeditious detection of pyrophosphate anions at nanomolar concentrations Xin Su, Chen Zhang, Xianjin Xiao, Anqin Xu, Zhendong Xu
Supporting Information for A nucleic acid-based fluorescent sensor for expeditious detection of pyrophosphate anions at nanomolar concentrations Xin Su, Chen Zhang, Xianjin Xiao, Anqin Xu, Zhendong Xu
PolyCAT A For Cation-Exchange
 PolyCAT A For Cation-Exchange The PolyCAT A is made though a unique process for attaching Poly(aspartic acid) covalently to silica. Proteins elute from this polypeptide coating in sharp peaks with little
PolyCAT A For Cation-Exchange The PolyCAT A is made though a unique process for attaching Poly(aspartic acid) covalently to silica. Proteins elute from this polypeptide coating in sharp peaks with little
All Rights Reserved. U.S. Patents 6,471,520B1; 5,498,190; 5,916, North Market Street, Suite CC130A, Milwaukee, WI 53202
 Secondary Structure In the previous protein folding activity, you created a hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously folds
Secondary Structure In the previous protein folding activity, you created a hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously folds
Imaging of protein crystals with two photon microscopy
 Supporting Information Imaging of protein crystals with two photon microscopy Pius Padayatti,*, Grazyna Palczewska,*, Wenyu Sun, Krzysztof Palczewski,# and David Salom Polgenix Inc., Cleveland, Ohio 44106,
Supporting Information Imaging of protein crystals with two photon microscopy Pius Padayatti,*, Grazyna Palczewska,*, Wenyu Sun, Krzysztof Palczewski,# and David Salom Polgenix Inc., Cleveland, Ohio 44106,
BETA STRAND Prof. Alejandro Hochkoeppler Department of Pharmaceutical Sciences and Biotechnology University of Bologna
 Prof. Alejandro Hochkoeppler Department of Pharmaceutical Sciences and Biotechnology University of Bologna E-mail: a.hochkoeppler@unibo.it C-ter NH and CO groups: right, left, right (plane of the slide)
Prof. Alejandro Hochkoeppler Department of Pharmaceutical Sciences and Biotechnology University of Bologna E-mail: a.hochkoeppler@unibo.it C-ter NH and CO groups: right, left, right (plane of the slide)
Proteins. Amino Acids (APK) Peptides (APK) 5/23/2012. Peptide bond. Acid. Amino
 Proteins Amino Acids (APK) Acid Amino Image courtesy of Biotech (biotech.chem.indiana.edu/pages/ protein_intro.html) Peptides (APK) Peptide bond 1 Proteins (polypeptides) Segment of a protein Peptide bonds
Proteins Amino Acids (APK) Acid Amino Image courtesy of Biotech (biotech.chem.indiana.edu/pages/ protein_intro.html) Peptides (APK) Peptide bond 1 Proteins (polypeptides) Segment of a protein Peptide bonds
R R Innovation Way P/N SECKIT-7830 Newark, DE 19711, USA Tel: Fax: Website: Published in November 2013
 5-100 Innovation Way Newark, DE 19711, USA Tel:302-3661101 Fax:302-3661151 Website: www.sepax-tech.com Published in November 2013 P/N SECKIT-7830 These Phases are developed based on innovative surface
5-100 Innovation Way Newark, DE 19711, USA Tel:302-3661101 Fax:302-3661151 Website: www.sepax-tech.com Published in November 2013 P/N SECKIT-7830 These Phases are developed based on innovative surface
NPTEL VIDEO COURSE PROTEOMICS PROF. SANJEEVA SRIVASTAVA
 LECTURE-06 PROTEIN PURIFICATION AND PEPTIDE ISOLATION USING CHROMATOGRAPHY TRANSCRIPT Welcome to the proteomics course. Today, we will talk about protein purification and peptide isolation using chromatography
LECTURE-06 PROTEIN PURIFICATION AND PEPTIDE ISOLATION USING CHROMATOGRAPHY TRANSCRIPT Welcome to the proteomics course. Today, we will talk about protein purification and peptide isolation using chromatography
AFFINITY HIS-TAG PURIFICATION
 DESCRIPTION Nickel NTA Agarose Cartridges 5ml are used for purification of histidine-tagged proteins in native or denaturing conditions. This cartridge can be used with an automated chromatography system,
DESCRIPTION Nickel NTA Agarose Cartridges 5ml are used for purification of histidine-tagged proteins in native or denaturing conditions. This cartridge can be used with an automated chromatography system,
Overview. Secondary Structure. Tertiary Structure
 Protein Structure Disclaimer: All information and images were taken from outside sources and the author claims no legal ownership of any material. Sources for images are linked on each slide and the information
Protein Structure Disclaimer: All information and images were taken from outside sources and the author claims no legal ownership of any material. Sources for images are linked on each slide and the information
Effect of Hydrogen Ion Concentration and Buffer Composition on Ligand Binding Characteristics and Polymerization of Cow s Milk Folate Binding Protein
 Bioscience Reports, Vol. 21, No. 6, December 2001 ( 2002) Effect of Hydrogen Ion Concentration and Buffer Composition on Ligand Binding Characteristics and Polymerization of Cow s Milk Folate Binding Protein
Bioscience Reports, Vol. 21, No. 6, December 2001 ( 2002) Effect of Hydrogen Ion Concentration and Buffer Composition on Ligand Binding Characteristics and Polymerization of Cow s Milk Folate Binding Protein
Structural bioinformatics
 Structural bioinformatics Why structures? The representation of the molecules in 3D is more informative New properties of the molecules are revealed, which can not be detected by sequences Eran Eyal Plant
Structural bioinformatics Why structures? The representation of the molecules in 3D is more informative New properties of the molecules are revealed, which can not be detected by sequences Eran Eyal Plant
Protein Structure Databases, cont. 11/09/05
 11/9/05 Protein Structure Databases (continued) Prediction & Modeling Bioinformatics Seminars Nov 10 Thurs 3:40 Com S Seminar in 223 Atanasoff Computational Epidemiology Armin R. Mikler, Univ. North Texas
11/9/05 Protein Structure Databases (continued) Prediction & Modeling Bioinformatics Seminars Nov 10 Thurs 3:40 Com S Seminar in 223 Atanasoff Computational Epidemiology Armin R. Mikler, Univ. North Texas
Solution Structure of the DNA-binding Domain of GAL4 from Saccharomyces cerevisiae
 Vol. 14, No. 1-4 175 Solution Structure of the DNA-binding Domain of GAL4 from Saccharomyces cerevisiae James D. Baleja, V. Thanabal, Ted Mau, and Gerhard Wagner Department of Biological Chemistry and
Vol. 14, No. 1-4 175 Solution Structure of the DNA-binding Domain of GAL4 from Saccharomyces cerevisiae James D. Baleja, V. Thanabal, Ted Mau, and Gerhard Wagner Department of Biological Chemistry and
7.88J Protein Folding Problem Fall 2007
 MIT OpenCourseWare http://ocw.mit.edu 7.88J Protein Folding Problem Fall 2007 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms. 7.88 Lecture Notes - 8 7.24/7.88J/5.48J
MIT OpenCourseWare http://ocw.mit.edu 7.88J Protein Folding Problem Fall 2007 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms. 7.88 Lecture Notes - 8 7.24/7.88J/5.48J
ProSEC 300S. Protein Characterization columns
 ProSEC 300S Protein Characterization columns Agilent s ProSEC 300S is a silica-based material specifically designed for the analysis of proteins by aqueous size exclusion chromatography. With a proprietary
ProSEC 300S Protein Characterization columns Agilent s ProSEC 300S is a silica-based material specifically designed for the analysis of proteins by aqueous size exclusion chromatography. With a proprietary
In Vitro Monitoring of the Formation of Pentamers from the Monomer of GST Fused HPV 16 L1
 This journal is The Royal Society of Chemistry 213 In Vitro Monitoring of the Formation of Pentamers from the Monomer of GST Fused HPV 16 L1 Dong-Dong Zheng, a Dong Pan, a Xiao Zha, ac Yuqing Wu,* a Chunlai
This journal is The Royal Society of Chemistry 213 In Vitro Monitoring of the Formation of Pentamers from the Monomer of GST Fused HPV 16 L1 Dong-Dong Zheng, a Dong Pan, a Xiao Zha, ac Yuqing Wu,* a Chunlai
Ligand immobilization using thiol-disulphide exchange
 A P P L I C A T I T E 9 Ligand immobilization using thiol-disulphide exchange Abstract This Application ote describes an immobilization procedure based on thioldisulphide exchange, providing a valuable
A P P L I C A T I T E 9 Ligand immobilization using thiol-disulphide exchange Abstract This Application ote describes an immobilization procedure based on thioldisulphide exchange, providing a valuable
Randomly arrayed G-rich DNA sequence for label-free and realtime. assay of enzyme activity
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Randomly arrayed G-rich DNA sequence for label-free and realtime assay of enzyme activity Zhuoliang
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Randomly arrayed G-rich DNA sequence for label-free and realtime assay of enzyme activity Zhuoliang
Application of Quantum Mechanics to Biology
 Application of Quantum Mechanics to Biology How can we apply quantum mechanics to biology? Polymers of nucleotides and amino acids - millions of atoms bounded into a large molecule Visual System Must turn
Application of Quantum Mechanics to Biology How can we apply quantum mechanics to biology? Polymers of nucleotides and amino acids - millions of atoms bounded into a large molecule Visual System Must turn
Stoichiometry and Affinity of Human Serum Albumin Alzheimer s A Peptide Interactions
 Biophysical Journal, Volume 100 Supporting Material Stoichiometry and Affinity of Human Serum Albumin Alzheimer s A Peptide Interactions Julijana Milojevic and Giuseppe Melacini S1 Supplementary Material
Biophysical Journal, Volume 100 Supporting Material Stoichiometry and Affinity of Human Serum Albumin Alzheimer s A Peptide Interactions Julijana Milojevic and Giuseppe Melacini S1 Supplementary Material
Chapter 3 Nucleic Acids, Proteins, and Enzymes
 3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
Protocol S1: Supporting Information
 Protocol S1: Supporting Information Basis for the specificity of the kinase domain of Abl for peptide substrates The crystal structures reported in this work were obtained using two different ATP analog-peptide
Protocol S1: Supporting Information Basis for the specificity of the kinase domain of Abl for peptide substrates The crystal structures reported in this work were obtained using two different ATP analog-peptide
Kinetics Review. Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets)
 Quiz 1 Kinetics Review Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets) I will post the problems with solutions on Toolkit for those that can t make
Quiz 1 Kinetics Review Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets) I will post the problems with solutions on Toolkit for those that can t make
Your Name: MID TERM ANSWER SHEET SIN: ( )
 MIDTERM EXAMINATION (October 23, 2008) BIOE150. Introduction to Bio-Nanoscience & Bio-Nanotechnology Professor Seung-Wuk Lee Fall Semester, 2008 0. Write down your name and the last digit of your SIN in
MIDTERM EXAMINATION (October 23, 2008) BIOE150. Introduction to Bio-Nanoscience & Bio-Nanotechnology Professor Seung-Wuk Lee Fall Semester, 2008 0. Write down your name and the last digit of your SIN in
STRUCTURAL BIOLOGY. α/β structures Closed barrels Open twisted sheets Horseshoe folds
 STRUCTURAL BIOLOGY α/β structures Closed barrels Open twisted sheets Horseshoe folds The α/β domains Most frequent domain structures are α/β domains: A central parallel or mixed β sheet Surrounded by α
STRUCTURAL BIOLOGY α/β structures Closed barrels Open twisted sheets Horseshoe folds The α/β domains Most frequent domain structures are α/β domains: A central parallel or mixed β sheet Surrounded by α
Protein Structure Prediction by Constraint Logic Programming
 MPRI C2-19 Protein Structure Prediction by Constraint Logic Programming François Fages, Constraint Programming Group, INRIA Rocquencourt mailto:francois.fages@inria.fr http://contraintes.inria.fr/ Molecules
MPRI C2-19 Protein Structure Prediction by Constraint Logic Programming François Fages, Constraint Programming Group, INRIA Rocquencourt mailto:francois.fages@inria.fr http://contraintes.inria.fr/ Molecules
AFFINITY HIS-TAG PURIFICATION
 DESCRIPTION Resins are products that allow batch or column purifications. This product is supplied as a suspension in 50% aqueous suspension containing 30 vol % ethanol. INSTRUCTIONS The resins are adapted
DESCRIPTION Resins are products that allow batch or column purifications. This product is supplied as a suspension in 50% aqueous suspension containing 30 vol % ethanol. INSTRUCTIONS The resins are adapted
Zwitterion Chromatography ZIC
 Zwitterion Chromatography ZIC A novel technique, with unique selectivity, suitable for preparative scale separations? PhD Einar Pontén What is Zwitterion Chromatography? Our definition: Liquid chromatography
Zwitterion Chromatography ZIC A novel technique, with unique selectivity, suitable for preparative scale separations? PhD Einar Pontén What is Zwitterion Chromatography? Our definition: Liquid chromatography
Purification of DNA from living cells
 Purification of DNA from living cells Total cell DNA & Plasmid DNA Grow and harvest bacterial culture Prepare cell extract Purify DNA from a cell extract Concentrate DNA samples Measure DNA concentration
Purification of DNA from living cells Total cell DNA & Plasmid DNA Grow and harvest bacterial culture Prepare cell extract Purify DNA from a cell extract Concentrate DNA samples Measure DNA concentration
Proteins the primary biological macromolecules of living organisms
 Proteins the primary biological macromolecules of living organisms Protein structure and folding Primary Secondary Tertiary Quaternary structure of proteins Structure of Proteins Protein molecules adopt
Proteins the primary biological macromolecules of living organisms Protein structure and folding Primary Secondary Tertiary Quaternary structure of proteins Structure of Proteins Protein molecules adopt
Nucleic Acids, Proteins, and Enzymes
 3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
3 Nucleic Acids, Proteins, and Enzymes Chapter 3 Nucleic Acids, Proteins, and Enzymes Key Concepts 3.1 Nucleic Acids Are Informational Macromolecules 3.2 Proteins Are Polymers with Important Structural
Analysis and Purification of Polypeptides by Reversed-Phase HPLC
 Analysis and Purification of Polypeptides by Reversed-Phase HPLC Reversed-phase HPLC is a valuable tool for the analysis and purification of proteins and peptides. It is effective in separating peptide
Analysis and Purification of Polypeptides by Reversed-Phase HPLC Reversed-phase HPLC is a valuable tool for the analysis and purification of proteins and peptides. It is effective in separating peptide
TECHNICAL BULLETIN. In Vitro Bacterial Split Fluorescent Protein Fold n Glow Solubility Assay Kits
 In Vitro Bacterial Split Fluorescent Protein Fold n Glow Solubility Assay Kits Catalog Numbers APPA001 In Vitro Bacterial Split GFP "Fold 'n' Glow" Solubility Assay Kit (Green) APPA008 In Vitro Bacterial
In Vitro Bacterial Split Fluorescent Protein Fold n Glow Solubility Assay Kits Catalog Numbers APPA001 In Vitro Bacterial Split GFP "Fold 'n' Glow" Solubility Assay Kit (Green) APPA008 In Vitro Bacterial
The Cys 2. His 2. Research Article 451
 Research Article 451 High-resolution structures of variant Zif268 DNA complexes: implications for understanding zinc finger DNA recognition Monicia Elrod-Erickson 1, Timothy E Benson 1 and Carl O Pabo
Research Article 451 High-resolution structures of variant Zif268 DNA complexes: implications for understanding zinc finger DNA recognition Monicia Elrod-Erickson 1, Timothy E Benson 1 and Carl O Pabo
CSE : Computational Issues in Molecular Biology. Lecture 19. Spring 2004
 CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
BIRKBECK COLLEGE (University of London)
 BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
Biochemistry study of the molecular basis of life
 Biochemistry : An Introduction Biochemistry study of the molecular basis of life n Study of the chemistry of living organisms Studies organic molecules & organic reactions in living organisms n Living
Biochemistry : An Introduction Biochemistry study of the molecular basis of life n Study of the chemistry of living organisms Studies organic molecules & organic reactions in living organisms n Living
Nickel-NTA Agarose Suspension
 Nickel-NTA Agarose Suspension Agarose beads for purification of His-tagged proteins Product No. A9735 Description Nickel-NTA Agarose Suspension is an agarose-based affinity chromatography resin allowing
Nickel-NTA Agarose Suspension Agarose beads for purification of His-tagged proteins Product No. A9735 Description Nickel-NTA Agarose Suspension is an agarose-based affinity chromatography resin allowing
Bivalirudin Purification:
 Bivalirudin Purification: Sorbent Screening and Overload Experiments Marc Jacob, Joshua Heng, and Tivadar Farkas Phenomenex, Inc., 411 Madrid Ave., Torrance, CA 90501 USA PO94190412_W Abstract In this
Bivalirudin Purification: Sorbent Screening and Overload Experiments Marc Jacob, Joshua Heng, and Tivadar Farkas Phenomenex, Inc., 411 Madrid Ave., Torrance, CA 90501 USA PO94190412_W Abstract In this
Sequence Analysis '17 -- lecture Secondary structure 3. Sequence similarity and homology 2. Secondary structure prediction
 Sequence Analysis '17 -- lecture 16 1. Secondary structure 3. Sequence similarity and homology 2. Secondary structure prediction Alpha helix Right-handed helix. H-bond is from the oxygen at i to the nitrogen
Sequence Analysis '17 -- lecture 16 1. Secondary structure 3. Sequence similarity and homology 2. Secondary structure prediction Alpha helix Right-handed helix. H-bond is from the oxygen at i to the nitrogen
0E.03. profos AG. creative bioscience solutions. product information. creative bioscience solutions
 0E.03 profos AG profos AG is a reliable and steadily growing biotechnology company. The combination of science, creativity and versatility under the paradigm of efficiency are our philosophy. Our products
0E.03 profos AG profos AG is a reliable and steadily growing biotechnology company. The combination of science, creativity and versatility under the paradigm of efficiency are our philosophy. Our products
LIGAND BINDING LABORATORIES
 LIGAND BINDING LABORATORIES Laboratory Page Aims of laboratory 1 Introduction to the ligand binding studies 1 Direct methods of measuring ligand binding 3 EXPERIMENT 1 An Equilibrium Dialysis Study of
LIGAND BINDING LABORATORIES Laboratory Page Aims of laboratory 1 Introduction to the ligand binding studies 1 Direct methods of measuring ligand binding 3 EXPERIMENT 1 An Equilibrium Dialysis Study of
Alpha-helices, beta-sheets and U-turns within a protein are stabilized by (hint: two words).
 1 Quiz1 Q1 2011 Alpha-helices, beta-sheets and U-turns within a protein are stabilized by (hint: two words) Value Correct Answer 1 noncovalent interactions 100% Equals hydrogen bonds (100%) Equals H-bonds
1 Quiz1 Q1 2011 Alpha-helices, beta-sheets and U-turns within a protein are stabilized by (hint: two words) Value Correct Answer 1 noncovalent interactions 100% Equals hydrogen bonds (100%) Equals H-bonds
Protein Techniques 1 APPENDIX TO CHAPTER 5
 Protein Techniques 1 APPENDIX T CHAPTER 5 Dialysis and Ultrafiltration If a solution of protein is separated from a bathing solution by a semipermeable membrane, small molecules and ions can pass through
Protein Techniques 1 APPENDIX T CHAPTER 5 Dialysis and Ultrafiltration If a solution of protein is separated from a bathing solution by a semipermeable membrane, small molecules and ions can pass through
Time-resolved Measurements Using the Agilent Cary Eclipse Fluorescence Spectrophotometer A Versatile Instrument for Accurate Measurements
 Time-resolved Measurements Using the Agilent Cary Eclipse Fluorescence Spectrophotometer A Versatile Instrument for Accurate Measurements Technical Overview Authors Dr. Fabian Zieschang, Katherine MacNamara,
Time-resolved Measurements Using the Agilent Cary Eclipse Fluorescence Spectrophotometer A Versatile Instrument for Accurate Measurements Technical Overview Authors Dr. Fabian Zieschang, Katherine MacNamara,
Overview. Things to do Prior to Protein Work. Preparation of Protein for Crystallization. Crystallization. Strategies when Crystal is Absent
 Sum Chan on Nov 16, 2007 Overview Things to do Prior to Protein Work Preparation of Protein for Crystallization Crystallization Strategies when Crystal is Absent Strategies when Crystal is Present Before
Sum Chan on Nov 16, 2007 Overview Things to do Prior to Protein Work Preparation of Protein for Crystallization Crystallization Strategies when Crystal is Absent Strategies when Crystal is Present Before
PROTEINS & NUCLEIC ACIDS
 Chapter 3 Part 2 The Molecules of Cells PROTEINS & NUCLEIC ACIDS Lecture by Dr. Fernando Prince 3.11 Nucleic Acids are the blueprints of life Proteins are the machines of life We have already learned that
Chapter 3 Part 2 The Molecules of Cells PROTEINS & NUCLEIC ACIDS Lecture by Dr. Fernando Prince 3.11 Nucleic Acids are the blueprints of life Proteins are the machines of life We have already learned that
Microfluidic Modulation Spectroscopy (MMS) For Protein Therapeutic Drug Analysis. Application Note #2017-1
 Microfluidic Modulation Spectroscopy (MMS) For Protein Therapeutic Drug Analysis Application Note #2017-1 April 2017 Jeff Zonderman RedShift Bioanalytics 131 Middlesex Turnpike Burlington, MA 01803 jzonderman@redshiftbio.com
Microfluidic Modulation Spectroscopy (MMS) For Protein Therapeutic Drug Analysis Application Note #2017-1 April 2017 Jeff Zonderman RedShift Bioanalytics 131 Middlesex Turnpike Burlington, MA 01803 jzonderman@redshiftbio.com
Ion exchange chromatography
 Ion exchange chromatography Objectives: 1- The objective of this experiment is to learn the principles of ion exchange chromatography by separating the charged molecules using buffer and salt. 2- A practical
Ion exchange chromatography Objectives: 1- The objective of this experiment is to learn the principles of ion exchange chromatography by separating the charged molecules using buffer and salt. 2- A practical
X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction
 X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction Tomohisa Shimasaki 1, Hiromi Yoshida 2, Shigehiro Kamitori 2 & Koji
X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction Tomohisa Shimasaki 1, Hiromi Yoshida 2, Shigehiro Kamitori 2 & Koji
Analysis of strongly absorbing chromophores by UV-visible ATR spectroscopy
 Technical Note: AN 915 Rev. B Analysis of strongly absorbing chromophores by UV-visible ATR spectroscopy Walter M. Doyle and Lani Tran This paper illustrates the potential of the attenuated total reflectance
Technical Note: AN 915 Rev. B Analysis of strongly absorbing chromophores by UV-visible ATR spectroscopy Walter M. Doyle and Lani Tran This paper illustrates the potential of the attenuated total reflectance
Vectors for Gene Cloning: Plasmids and Bacteriophages
 Vectors for Gene Cloning: Plasmids and Bacteriophages DNA molecule must be able to replicate within the host cell to be able to act as a vector for gene cloning, so that numerous copies of the recombinant
Vectors for Gene Cloning: Plasmids and Bacteriophages DNA molecule must be able to replicate within the host cell to be able to act as a vector for gene cloning, so that numerous copies of the recombinant
GENETICS الفريق الطبي االكاديمي. DNA Genes & Chromosomes. DONE BY : Buthaina Al-masaeed & Yousef Qandeel. Page 0
 GENETICS ومن أحياها DNA Genes & Chromosomes الفريق الطبي االكاديمي DNA Genes & Chromosomes DONE BY : Buthaina Al-masaeed & Yousef Qandeel Page 0 T(0:44 min) In the pre lecture we take about the back bone
GENETICS ومن أحياها DNA Genes & Chromosomes الفريق الطبي االكاديمي DNA Genes & Chromosomes DONE BY : Buthaina Al-masaeed & Yousef Qandeel Page 0 T(0:44 min) In the pre lecture we take about the back bone
Virtual bond representation
 Today s subjects: Virtual bond representation Coordination number Contact maps Sidechain packing: is it an instrumental way of selecting and consolidating a fold? ASA of proteins Interatomic distances
Today s subjects: Virtual bond representation Coordination number Contact maps Sidechain packing: is it an instrumental way of selecting and consolidating a fold? ASA of proteins Interatomic distances
Homology Modelling. Thomas Holberg Blicher NNF Center for Protein Research University of Copenhagen
 Homology Modelling Thomas Holberg Blicher NNF Center for Protein Research University of Copenhagen Why are Protein Structures so Interesting? They provide a detailed picture of interesting biological features,
Homology Modelling Thomas Holberg Blicher NNF Center for Protein Research University of Copenhagen Why are Protein Structures so Interesting? They provide a detailed picture of interesting biological features,
11 questions for a total of 120 points
 Your Name: BYS 201, Final Exam, May 3, 2010 11 questions for a total of 120 points 1. 25 points Take a close look at these tables of amino acids. Some of them are hydrophilic, some hydrophobic, some positive
Your Name: BYS 201, Final Exam, May 3, 2010 11 questions for a total of 120 points 1. 25 points Take a close look at these tables of amino acids. Some of them are hydrophilic, some hydrophobic, some positive
encouraged to use the Version of Record that, when published, will replace this version. The most /BCJ BIOCHEMICAL JOURNAL
 Biochemical Journal: this is an Accepted Manuscript, not the final Version of Record. You are encouraged to use the Version of Record that, when published, will replace this version. The most up-to-date
Biochemical Journal: this is an Accepted Manuscript, not the final Version of Record. You are encouraged to use the Version of Record that, when published, will replace this version. The most up-to-date
Chapter 4 Fluorescence Resonance Energy Transfer (FRET) by Minor Groove-Associated Cyanine-Polyamide Conjugates
 Chapter 4 Fluorescence Resonance Energy Transfer (FRET) by Minor Groove-Associated Cyanine-Polyamide Conjugates The work described in this chapter was accomplished in collaboration with V. Rucker (Dervan
Chapter 4 Fluorescence Resonance Energy Transfer (FRET) by Minor Groove-Associated Cyanine-Polyamide Conjugates The work described in this chapter was accomplished in collaboration with V. Rucker (Dervan
SPARK. Southern Illinois University Edwardsville. Drake Jensen Southern Illinois University Edwardsville,
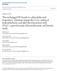 Southern Illinois University Edwardsville SPARK Chemistry Faculty Research, Scholarship, and Creative Activity Chemistry Spring 2-15-2015 The exchanged EF-hands in calmodulin and troponin C chimeras impair
Southern Illinois University Edwardsville SPARK Chemistry Faculty Research, Scholarship, and Creative Activity Chemistry Spring 2-15-2015 The exchanged EF-hands in calmodulin and troponin C chimeras impair
