CGTAAGAGTACGTCCAGCATCGGF ATCATTATCTACATCXXXXXTACCATTCATTCAGATCTCACTATCGCATTCTCATGCAGGTCGTAGCCXS
|
|
|
- Abraham Sullivan
- 6 years ago
- Views:
Transcription
1 5 CGTAAGAGTACGTCCAGCATCGGF ATCATTATCTACATCXXXXXTACCATTCATTCAGATCTCACTATCGCATTCTCATGCAGGTCGTAGCCXS 29 Supplementary Figure 1 Sequence of the DNA primer/template pair used in Figure 1bc. A 23nt primer is 6FAM labeled (F) on its 5 terminus and annealed proximal to the 3 end of a 70mer DNA template strand. A blocking oligomer (not shown) is annealed to a nt segment of the template (underlined). The template has 5 abasic residues at position to 29 from the position that act as a reporter for template displacement through the nanopore. An abasic monomer (X) linked to a three carbon spacer (S) protects the template 3 terminus from phi29 DNAPcatalyzed exonucleolysis (see Supplementary Fig. 2).
2 a abasic (X) = P H 2 C H b C3 spacer (S) = P H 3 c d i. FCCATTCATTCAGATCTCACTATCGCATTCTCATGCAGGTCGTAGCC3 ii. FCCATTCATTCAGATCTCACTATCGCATTCTCATGCAGGTCGTAGCCXS3 3 SXCGTAAGAGTACGTCCAGCATCGG iii. FCCATTCATTCAGATCTCACTATCGCATTCTCATGCAGGTCGTAGCC3 3 SXCGTAAGAGTACGTCCAGCATCGG iv. FCCATTCATTCAGATCTCACTATCGCATTCTCATGCAGGTCGTAGCCXS3 DNA substrate i i i ii ii iii iii iv iv wt phi29 DNAP Mg 2 46nt 23nt Supplementary Figure 2 Protection of DNA template 3 termini against exonucleolysis. Phi29 DNAP contains an exonuclease domain that hydrolyzes nucleotides from the 3 termini of ssdna and dsdna. In the nanopore experiments, the 3 termini of DNA primer strands were protected by blocking oligomers, but the 3 termini of the DNA templates also required protection. To this end, we appended a protecting group to the template strands composed of (a) an abasic (1, 2 H) residue (X), and (b) a three carbon spacer (S). (c) DNA strands used to test efficacy of this protection strategy. i) 6FAM (F) labeled 46nt template unprotected at its 3 terminus. ii) Same as (i) but protected at its 3 terminus (XS). iii) Template as in (i) annealed to a 23nt DNA strand protected at its 3 terminus (SX). iv) Template as in (ii) annealed to a 23nt DNA strand protected at its 3 terminus (SX). (d) DNA substrates iiv were incubated for 5 hours in nanopore buffer at 23 C then analyzed by denaturing PAGE (17% gel). MgCl 2 (10 mm) or phi29 DNAP (1 µm) were added as noted. We found that 3 protected DNA templates were not degraded either as singlestranded substrates (lane 5) or as components of doublestranded substrates (lane 9). By comparison, the respective controls absent 3 protection were degraded (lanes 3 & 7).
3 a ii v i iii iv 5 s b SXX X X XXXATGGTAACTAAGTATAGAGTGATA CGTAAGAGTACGTCCAGCATCGG 5 CTCACCTATCCTTCCACTCATACTATCATTATCTACATCXXXXXTACCATTGATTCATATCTCACTATCGCATTCTCATGCAGGTCGTAGCCXS c 29 dgmp d 1 s 1 s Supplementary Figure 3 The causes of the first and second ionic current peaks. In the main text, we hypothesized that traversal of the first ionic current peak in Figure 2b was due to mechanical unzipping of the blocking oligomer, and that traversal of the second ionic current peak was dependent upon primer elongation catalyzed by phi29 DNAP. This hypothesis predicts that traversal of the first peak, but not the second, would be observed absent the Mg 2 ions required for phi29 DNAP catalytic function. This was tested in an experiment in which phi29 DNAP and the DNA substrate shown in Figure 2a of the main text were added to the nanopore cis chamber absent MgCl 2. A representative currrent trace (a) from this experiment shows i) nanopore capture of a phi29 DNAPDNA complex with a blocking oligomer bound, yielding a 2324 ionic current; ii) traversal of a current peak as the abasic insert moves through the major pore constriction due to mechanical unzipping of the blocking oligomer; iii) a long stall at, as the 3 H terminus of the DNA primer reaches the polymerase active site; followed by iv) eventual voltagepromoted dissociation of the complex to restore the v) open
4 channel current. The second peak was not observed in this experiment. The hypothesis also predicts that DNA synthesis would stall if a dntp required for replication was omitted from the nanopore cis chamber in the presence of all other required components. The data presented below are the result of this test. (b) DNA substrate. The template strand is a 94mer protected against exonucleolysis at its 3 end (XS, see Supplementary Fig. 2). It bears a fiveabasic reporter at positions to 29 downstream from the n = 0 position and a unique dgmp residue (red arrow). The DNA template is annealed to a 23nt primer wherein the 3 end is protected by a blocking oligomer (red line; see Fig. 1b, iii for details). (c) Stalling of replication at the dgmp residue of the template strand in the absence of dctp. DNA substrate and phi29 DNAP were added to the nanopore cis chamber containing all constituents necessary for replication (see Fig. 2) except dctp. Under these conditions, capture of the phi29 DNAPDNA complex at 180 mv resulted in traversal of the first ionic current peak due to mechanical unzipping of the blocking oligomer as in Figure 2. However, after removal of the blocking oligomer, the ionic current rose to the second peak then stalled and oscillated between 31 and (red arrow) in of 34 events analyzed. Stalling at this position was anticipated absent dctp (Lieberman, et al., 10, J. Am. Chem. Soc., 132, ) given the distance between the abasic reporter and the unique dgmp of the template strand. (d) Addition of dctp results in progression through the second peak. Conditions of the experiment are the same as panel (c) except for the addition of 100 μm dctp to the cis chamber. Replication proceeds past the dgmp template residue resulting in rapid progression of the ionic current trace from 31 to on the shoulder of the second peak (red arrow). This pattern was observed in 26 of 26 events. These results are consistent with the hypothesis that the second ionic current peak is caused by phi29 DNAPdependent replication, drawing the DNA template strand through the nanopore toward the cis chamber.
5 a b 100 nm dctp, 100 μm d(a,t,g)tp 1 s GGCTACGACCTGCATGAGAATGC SXCCGATGCTGGACGTACTCTTACGCTATCACTCTAGACTTACTTACCATXXXXX...CTA5 n11 78 n11 c 100 nm dttp, 100 μm d(a,c,g)tp n2 n n10 n12 89 n n n s d n2 n5 n10n12 n16 n n23 GGCTACGACCTGCATGAGAATGC SXCCGATGCTGGACGTACTCTTACGCTATCACTCTAGACTTACTTACCATXXXXX...CTA5 e 100 nm dgtp, 100 μm d(a,t,c)tp n0 n s n6 34 n8 45 n n21 14 n n22 f n0 n4n6n8 n13 n17 n21 n22 GGCTACGACCTGCATGAGAATGC SXCCGATGCTGGACGTACTCTTACGCTATCACTCTAGACTTACTTACCATXXXXX...CTA5 g 0 nm datp, 100 μm d(g,t,c)tp n14 n 10 n9 01 n n3 5 s 4 n n1 n n19 n h n1 n3 n7 n9 n14 n n18 n24 n19 GGCTACGACCTGCATGAGAATGC SXCCGATGCTGGACGTACTCTTACGCTATCACTCTAGACTTACTTACCATXXXXX...CTA5
6 Supplementary Figure 4 Independent confirmation of discrete ionic current states during phi29 DNAPcontrolled template movement. In Figure 3 of the main text, we highlighted 16 discrete ionic current states observed during both forward and reverse ratcheting of a DNA substrate through the nanopore. We aimed to use these 16 states to determine the rate of template movement through the nanopore controlled by phi29 DNAP. Therefore we needed a method to confirm that the observed ionic current states corresponded to translocation of template nucleotides. To this end, we repeated the experiment presented in Figure 2 using the same DNA template. However, rather than using 100 μm of each dntp in the reaction mixture, we ran four independent experiments where, in each case, one of the four dntp substrates was present at 1000 nm and all others were at 100 μm (panels a, c, e, g). In these panels, the first peak is traversed due to voltagedriven unzipping of the blocking oligomer until the 3 hydroxyl of the primer strand is situated in the polymerase catalytic domain (arrow, ). The ionic current trace to the right of in each panel is due to reverse template movement driven by DNA replication. Depending upon the identity of the dilute dntp for each panel, pauses in the current trace > 0 ms were observed at various ionic current states during replication. These pauses occurred as the template base complementary to the dilute dntp arrived at the polymerase active site. For example, in panel (a) (dilute dctp), one discrete pause in the ionic current trace was observed as a template dgmp residue reached the active site. This one pause was predicted from the single dgmp residue at position n11 in the template sequence (panel b). The number of pauses in panels (c), (e), and (g) were likewise predicted from the template sequence (panels d, f, h). Each pause in the current trace is marked by template position (n0 to n24), and by numbers that correspond to ionic current states 016 in Figure 3a,b. Two numbers, e.g. 78 in panel (a), denotes a pause where ionic current fluctuations occurred between two states, while a single number, e.g. 3 in panel (c), denotes a pause with only one ionic current state. In the case where a pause fluctuated between two states, the template position was assigned to the second state. The current traces corresponding to template positions n21n22 (blue) in panel (e) and n1 (green), n18, n19, and n24 (blue) in panel (g) are expanded for clarity. Scale bars in these expansions equal 0ms. In panel (g), pauses at two neighboring template positions (n and n16) gave the same ionic current state (see Supplementary Figure 6). Discontinuities are present in panel (g) because pauses at template positions n2, n16, and n19 lasted 10s of seconds. By combining the data summarized in these four experiments, all nucleotides that could be replicated in the model DNA template were accounted for in sequential order within the 16 ionic current states in Figure 3b.
7 a 5 S X X X X X X X AGTCTAGAGTGATAGCGTAAGAGTACGTCCAGCATCGGF ATCATTATCTACATCXXXXXTACCATTCATTCAGATCTCACTATCGCATTCTCATGCAGGTCGTAGCCXS 29 b blocking oligomer dntps wt phi29 DNAP Mg 2 nt extension 23nt primers degradation products Supplementary Figure 5 A blocking oligomer optimized for increased DNA template throughput on the nanopore. (a) ptimized blocking oligomer (red line) annealed to a DNA substrate. The segment of the blocking oligomer complementary to the template strand was reduced from to nucleotides to facilitate faster removal by the applied voltage at the nanopore. The primer strand is a 23mer bearing a FAM group (F) at its 5 terminus. (b) PAGE gel (17%) demonstrating protection of DNA using the shortened blocking oligomer. DNA substrate (1 μm) was incubated for 5 hr in nanopore buffer at 23 C with or without phi29 DNAP (0.75 μm), dntp (100 μm each), and Mg2 (10 mm) as indicated. Lanes 14 are reactions run absent blocking oligomer; lanes 57 are reactions run in the presence of blocking oligomers preannealed to the DNA substrate. In the presence of phi29 DNAP and Mg 2, but absent blocking oligomer, exonucleolysis by the enzyme resulted in disappearance of the 23nt primer band (lane 3); by comparison the same reaction run present the blocking oligomer showed no loss of the 23nt primer (lane 6). When the reaction was run present all components necessary for replication including dntp but absent blocking oligomer (lane 4) most of the primers were extend by nt to full length products; by comparison the same reaction run present the blocking oligomer showed no appearance of longer products and no loss of the 23nt primer (lane 7). We conclude that this blocking oligomer optimized for removal at the nanopore remains effective at primer protection in bulk phase.
8 a i. ii. iii. iv. v. vi. vii. viii. n0 n10 GGCTACGACCTGCATGAGAATGCGATAGTGAG SXCCGATGCTGGACGTACTCTTACGCTATCACTCTAGACTTACTTACCATXXXXXCTACATCTATTACTA5 n11 GGCTACGACCTGCATGAGAATGCGATAGTGAGA SXCCGATGCTGGACGTACTCTTACGCTATCACTCTAGACTTACTTACCATXXXXXCTACATCTATTACTA5 n12 GGCTACGACCTGCATGAGAATGCGATAGTGAGAT SXCCGATGCTGGACGTACTCTTACGCTATCACTCTAGACTTACTTACCATXXXXXCTACATCTATTACTA5 n13 GGCTACGACCTGCATGAGAATGCGATAGTGAGATC SXCCGATGCTGGACGTACTCTTACGCTATCACTCTAGACTTACTTACCATXXXXXCTACATCTATTACTA5 n14 GGCTACGACCTGCATGAGAATGCGATAGTGAGATCT SXCCGATGCTGGACGTACTCTTACGCTATCACTCTAGACTTACTTACCATXXXXXCTACATCTATTACTA5 n GGCTACGACCTGCATGAGAATGCGATAGTGAGATCTG SXCCGATGCTGGACGTACTCTTACGCTATCACTCTAGACTTACTTACCATXXXXXCTACATCTATTACTA5 n16 GGCTACGACCTGCATGAGAATGCGATAGTGAGATCTGA SXCCGATGCTGGACGTACTCTTACGCTATCACTCTAGACTTACTTACCATXXXXXCTACATCTATTACTA5 n17 GGCTACGACCTGCATGAGAATGCGATAGTGAGATCTGAA SXCCGATGCTGGACGTACTCTTACGCTATCACTCTAGACTTACTTACCATXXXXXCTACATCTATTACTA5 b n13 n12 n14 n n16 n11 28 n10 n Ionic current state Supplementary Figure 6 Verification of ionic current state transitions in Figure 3b that represent a singlenucleotide displacement of the template strand in the nanopore. We aimed to estimate the frequency of singlenucleotide indel (insertion and deletion) errors reported in the ionic current trace during phi29 DNAPcontrolled movement of the template through the nanopore (Fig. 4 of main text). Therefore, we needed to verify specific transitions between ionic current states that represent a single nucleotide displacement of the template strand in the nanopore. Missing or repeating these transitions would be equal to inserting or deleting a single nucleotide read. To this end, we used a series of eight primer/template substrates (panel a). Prior to capture of each substrate on the nanopore, phi29 DNAPcatalyzed incorporation of the appropriate complementary ddntp was allowed to occur in the nanopore bath. After ddntp incorporation, these substrates spanned the ionic current states 712 in Figure 3b. Complexes were captured in
9 the presence of the next complementary dntp in order to form a stable ternary (DNADNAPdNTP) complex at the indicated position along the template (arrow). For example, 1 μm DNA substrate (i) was added to the nanopore chamber in the presence of 0.75 μm phi29 DNAP, 10 mm MgCl 2, 600 μm ddatp, and 50 μm dttp. Two minutes were allowed for phi29 DNAPcatalyzed addition of the ddntp to the primer 3 terminus, after which capture of each ternary complex resulted in an ionic current state corresponding to the n10 templating base residing in enzyme active site. This experiment was repeated for each DNA substrate, and the results are summarized in panel (b). Red circles superimposed on the ionic current states from Figure 3b (black connected circles) indicate the residual ionic current measured when each ternary complex was captured on the nanopore. Each point is the calculated median of at least 10 measured events. These data indicate that transitions from ionic current state 7 to 8, 8 to 9, 10 to 11, and 11 to 12 each represent a single nucleotide displacement of the template strand in the nanopore.
10 Supplementary Video 1 Forward and reverse ratcheting of DNA in a nanopore controlled by phi29 DNA polymerase. A single αhl pore (gray) is embedded in a lipid bilayer that separates two chambers each containing ~100 μl of a 0.3 M KCl solution buffered at ph 8 (23 C). The solution at the top is supplemented with Mg 2, dntp, and DNA substrate (red backbone) that is protected by a blocking oligomer (yellow backbone). Phi29 DNA polymerase (orange) is prebound to the DNA. Catalysis in bulk phase does not occur because the blocking oligomer prevents enzyme access to the 3 prime hydroxyl of the primer strand. A 180 mv potential is applied across the nanopore resulting in an ionic current (white trace at left). Initially (t = 0 seconds) the channel is open with a current of ~60. Capture of the DNA template (t = 2 s) results in a current drop to ~23.5. Due to the force applied by the electric field, the blocking oligomer is mechanically unzipped from the DNA template complex in single nucleotide increments. During this unzipping process, a five abasic insert within the template strand (green bars with white highlights) passes through the pore causing traversal of a current peak (t = 8 to s) Eventually, the blocking oligomer unzips completely from the DNA template and dissociates exposing the 3 prime hydroxyl of the primer strand (flashing red circle at t = 24 s). Primer elongation now drives the template and the abasic insert back through the nanopore in reverse order, causing the ionic current to traverse the once again (t = 29 to 34 s). Template movement is ~ 10fold faster during the catalytic process compared to the mechanical unzipping process. The animation is not in real time for clarity. Supplementary Video 2 Typical ionic current traces caused by forward and reverse ratcheting of DNA in a nanopore. See Figure 2 for details. Three consecutive events are shown wherein the peak is traversed twice, first due to mechanical unzipping of the blocking oligomer from the template and second due to elongation of the primer strand driving the DNA template back through the pore in reverse order. The peak corresponds to passage of an abasic segment of the template through the limiting constriction of the nanopore. The data were processed with a digital 2 khz Gaussian low pass filter for clarity and presented at 1.5 X real time. The three events begin at 2.7 s, 13.4 s, and 29 s as indicated by the X axis.
Processive Replication of Single DNA Molecules in a Nanopore Catalyzed by phi29 DNA Polymerase
 Processive Replication of Single DNA Molecules in a Nanopore Catalyzed by phi29 DNA Polymerase Kate R. Lieberman, Gerald M. Cherf, Michael J. Doody, Felix Olasagasti, Yvette Kolodji, and Mark Akeson* Department
Processive Replication of Single DNA Molecules in a Nanopore Catalyzed by phi29 DNA Polymerase Kate R. Lieberman, Gerald M. Cherf, Michael J. Doody, Felix Olasagasti, Yvette Kolodji, and Mark Akeson* Department
Additional Activity: Sanger Dideoxy Sequencing: A Simulation Activity
 Student Worksheet Additional Activity: Sanger Dideoxy Sequencing: A Simulation Activity LSM 6.3-7 In 1977, Frederick Sanger developed a method by which the nucleotide sequence of a DNA fragment could be
Student Worksheet Additional Activity: Sanger Dideoxy Sequencing: A Simulation Activity LSM 6.3-7 In 1977, Frederick Sanger developed a method by which the nucleotide sequence of a DNA fragment could be
Enzymatic assembly of DNA molecules up to several hundred kilobases
 nature methods Enzymatic assembly of DNA molecules up to several hundred kilobases Daniel G Gibson, Lei Young, Ray-Yuan Chuang, J Craig Venter, Clyde A Hutchison III & Hamilton O Smith Supplementary figures
nature methods Enzymatic assembly of DNA molecules up to several hundred kilobases Daniel G Gibson, Lei Young, Ray-Yuan Chuang, J Craig Venter, Clyde A Hutchison III & Hamilton O Smith Supplementary figures
Supporting Information
 Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany Rolling-circle Amplification of a DNA Nanojunction Chenxiang Lin, Mingyi Xie, Julian J.L. Chen, Yan Liu and Hao Yan A. RCA replication of the
Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany Rolling-circle Amplification of a DNA Nanojunction Chenxiang Lin, Mingyi Xie, Julian J.L. Chen, Yan Liu and Hao Yan A. RCA replication of the
DNA Replication. DNA Replication. Meselson & Stahl Experiment. Contents
 DNA Replication Contents 1 DNA Replication 1.1 Meselson & Stahl Experiment 1.2 Replication Machinery 2 Polymerase Chain Reaction (PCR) 3 External Resources: DNA Replication Meselson & Stahl Experiment
DNA Replication Contents 1 DNA Replication 1.1 Meselson & Stahl Experiment 1.2 Replication Machinery 2 Polymerase Chain Reaction (PCR) 3 External Resources: DNA Replication Meselson & Stahl Experiment
 The replication of DNA Kornberg 1957 Meselson and Stahl 1958 Cairns 1963 Okazaki 1968 DNA Replication The driving force for DNA synthesis. The addition of a nucleotide to a growing polynucleotide
The replication of DNA Kornberg 1957 Meselson and Stahl 1958 Cairns 1963 Okazaki 1968 DNA Replication The driving force for DNA synthesis. The addition of a nucleotide to a growing polynucleotide
Supplementary Info. Directed Enzymatic Activation of 1-D DNA Tiles
 Supplementary Info Directed Enzymatic Activation of 1-D DNA Tiles Sudhanshu Garg,, Harish Chandran,, Nikhil Gopalkrishnan,, Thomas H. LaBean,, and John Reif Department of Computer Science, Duke University,
Supplementary Info Directed Enzymatic Activation of 1-D DNA Tiles Sudhanshu Garg,, Harish Chandran,, Nikhil Gopalkrishnan,, Thomas H. LaBean,, and John Reif Department of Computer Science, Duke University,
Instructions for Use Life Science Kits & Assays
 Instructions for Use Life Science Kits & Assays Product specifications 1 Product specifications The innumix Standard PCR MasterMix contains all reagents required for routine high throughput PCR amplifications
Instructions for Use Life Science Kits & Assays Product specifications 1 Product specifications The innumix Standard PCR MasterMix contains all reagents required for routine high throughput PCR amplifications
Session 3 Cloning Overview & Polymerase Chain Reaction
 Session 3 Cloning Overview & Polymerase Chain Reaction Learning Objective: In this lab exercise, you will become familiar with the steps of a polymerase chain reaction, the required reagents for a successful
Session 3 Cloning Overview & Polymerase Chain Reaction Learning Objective: In this lab exercise, you will become familiar with the steps of a polymerase chain reaction, the required reagents for a successful
Storage and Expression of Genetic Information
 Storage and Expression of Genetic Information 29. DNA structure, Replication and Repair ->Ch 25. DNA metabolism 30. RNA Structure, Synthesis and Processing ->Ch 26. RNA metabolism 31. Protein Synthesis
Storage and Expression of Genetic Information 29. DNA structure, Replication and Repair ->Ch 25. DNA metabolism 30. RNA Structure, Synthesis and Processing ->Ch 26. RNA metabolism 31. Protein Synthesis
Polymerase Chain Reaction (PCR)
 Laboratory for Environmental Pathogens Research Department of Environmental Sciences University of Toledo Polymerase Chain Reaction (PCR) Background information The polymerase chain reaction (PCR) is an
Laboratory for Environmental Pathogens Research Department of Environmental Sciences University of Toledo Polymerase Chain Reaction (PCR) Background information The polymerase chain reaction (PCR) is an
DNA polymerase proofreading: active site switching catalyzed by the bacteriophage T4 DNA polymerase
 5452 5463 Nucleic Acids Research, 2007, Vol. 35, No. 16 Published online 15 August 2007 doi:10.1093/nar/gkm591 DNA polymerase proofreading: active site switching catalyzed by the bacteriophage T4 DNA polymerase
5452 5463 Nucleic Acids Research, 2007, Vol. 35, No. 16 Published online 15 August 2007 doi:10.1093/nar/gkm591 DNA polymerase proofreading: active site switching catalyzed by the bacteriophage T4 DNA polymerase
Cat. # R006A. For Research Use. TaKaRa Z-Taq DNA Polymerase. Product Manual. v201411da
 Cat. # R006A For Research Use TaKaRa Z-Taq DNA Polymerase Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Specifications... 3 IV. Optimization of Reaction Conditions... 4
Cat. # R006A For Research Use TaKaRa Z-Taq DNA Polymerase Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Specifications... 3 IV. Optimization of Reaction Conditions... 4
DNA Replication semiconservative replication conservative replication dispersive replication DNA polymerase
 DNA Replication DNA Strands are templates for DNA synthesis: Watson and Crick suggested that the existing strands of DNA served as a template for the producing of new strands, with bases being added to
DNA Replication DNA Strands are templates for DNA synthesis: Watson and Crick suggested that the existing strands of DNA served as a template for the producing of new strands, with bases being added to
Guidelines for Preventing Contamination of PCR Reference Guidelines for Primer Design Estimation of Primer Melting Temperature
 Guidelines for Preventing Contamination of PCR During PCR more than 10 million copies of a template DNA are generated. Therefore, care must be taken to avoid contamination with other templates and amplicons
Guidelines for Preventing Contamination of PCR During PCR more than 10 million copies of a template DNA are generated. Therefore, care must be taken to avoid contamination with other templates and amplicons
Masayoshi Honda, Jeehae Park, Robert A. Pugh, Taekjip Ha, and Maria Spies
 Molecular Cell, Volume 35 Supplemental Data Single-Molecule Analysis Reveals Differential Effect of ssdna-binding Proteins on DNA Translocation by XPD Helicase Masayoshi Honda, Jeehae Park, Robert A. Pugh,
Molecular Cell, Volume 35 Supplemental Data Single-Molecule Analysis Reveals Differential Effect of ssdna-binding Proteins on DNA Translocation by XPD Helicase Masayoshi Honda, Jeehae Park, Robert A. Pugh,
Recombinant DNA Technology
 History of recombinant DNA technology Recombinant DNA Technology (DNA cloning) Majid Mojarrad Recombinant DNA technology is one of the recent advances in biotechnology, which was developed by two scientists
History of recombinant DNA technology Recombinant DNA Technology (DNA cloning) Majid Mojarrad Recombinant DNA technology is one of the recent advances in biotechnology, which was developed by two scientists
Current ( pa) Current (pa) Voltage (mv) Voltage ( mv)
 Current ( pa) 3000 2000 1000 0-1000 -2000-3000 a -400-200 0 200 400 Voltage (mv) P1 P2 P3 P4 P5 P6 P7 P8 P9 P10 P11 P12 P13 P14 P15 P16 P17 P18 Average Current (pa) 4500 3000 1500 0-1500 -3000-4500 b -400-200
Current ( pa) 3000 2000 1000 0-1000 -2000-3000 a -400-200 0 200 400 Voltage (mv) P1 P2 P3 P4 P5 P6 P7 P8 P9 P10 P11 P12 P13 P14 P15 P16 P17 P18 Average Current (pa) 4500 3000 1500 0-1500 -3000-4500 b -400-200
Wang Tian, Ying T. Hwang, and Charles B. C. Hwang*
 JOURNAL OF VIROLOGY, Sept. 2008, p. 8937 8941 Vol. 82, No. 17 0022-538X/08/$08.00 0 doi:10.1128/jvi.00911-08 Copyright 2008, American Society for Microbiology. All Rights Reserved. The Enhanced DNA Replication
JOURNAL OF VIROLOGY, Sept. 2008, p. 8937 8941 Vol. 82, No. 17 0022-538X/08/$08.00 0 doi:10.1128/jvi.00911-08 Copyright 2008, American Society for Microbiology. All Rights Reserved. The Enhanced DNA Replication
Polymerase Chain Reaction (PCR)
 Polymerase Chain Reaction (PCR) PCR protocols Polymerase Chain Reaction (PCR) A technique for the in vitro amplification of specific DNA sequences by the simultaneous primer extension of complementary
Polymerase Chain Reaction (PCR) PCR protocols Polymerase Chain Reaction (PCR) A technique for the in vitro amplification of specific DNA sequences by the simultaneous primer extension of complementary
TaKaRa PCR Amplification Kit
 Cat. # R011 For Research Use TaKaRa PCR Amplification Kit Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Storage... 4 IV. Materials Required but not Provided... 4 V. Principle...
Cat. # R011 For Research Use TaKaRa PCR Amplification Kit Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Storage... 4 IV. Materials Required but not Provided... 4 V. Principle...
Cat. # For Research Use. BcaBEST Labeling Kit. Product Manual. v201701da
 Cat. # 6046 For Research Use BcaBEST Labeling Kit Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Storage... 3 IV. Principles... 4 V. Protocol... 5 VI. Effect of Template
Cat. # 6046 For Research Use BcaBEST Labeling Kit Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Storage... 3 IV. Principles... 4 V. Protocol... 5 VI. Effect of Template
Replication. Obaidur Rahman
 Replication Obaidur Rahman DIRCTION OF DNA SYNTHESIS How many reactions can a DNA polymerase catalyze? So how many reactions can it catalyze? So 4 is one answer, right, 1 for each nucleotide. But what
Replication Obaidur Rahman DIRCTION OF DNA SYNTHESIS How many reactions can a DNA polymerase catalyze? So how many reactions can it catalyze? So 4 is one answer, right, 1 for each nucleotide. But what
TaKaRa Taq HS PCR Kit, UNG plus
 Cat. # R013S/A For Research Use TaKaRa Taq HS PCR Kit, UNG plus Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Storage... 4 IV. Materials Required but not Provided... 4 V.
Cat. # R013S/A For Research Use TaKaRa Taq HS PCR Kit, UNG plus Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Storage... 4 IV. Materials Required but not Provided... 4 V.
CircLigase ssdna Ligase
 Cat. Nos. CL4111K and CL4115K Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook (facebook.com/epicentrebio), and Twitter (@EpicentreBio). www.epicentre.com Lit. # 222 10/2012 1 EPILIT222
Cat. Nos. CL4111K and CL4115K Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook (facebook.com/epicentrebio), and Twitter (@EpicentreBio). www.epicentre.com Lit. # 222 10/2012 1 EPILIT222
Biotechnology. Explorer Program. Serious About Science Education 5/17/09 1
 Biotechnology Explorer Program Serious About Science Education 5/17/09 1 Chromosome 8: PCR TM PCR Workshop Kirk Brown,, Tracy High School; Tracy, Ca Stan Hitomi,, Monte Vista High School; Danville, CA
Biotechnology Explorer Program Serious About Science Education 5/17/09 1 Chromosome 8: PCR TM PCR Workshop Kirk Brown,, Tracy High School; Tracy, Ca Stan Hitomi,, Monte Vista High School; Danville, CA
Polymerase Chain Reaction (PCR) and Its Applications
 Polymerase Chain Reaction (PCR) and Its Applications What is PCR? PCR is an exponentially progressing synthesis of the defined target DNA sequences in vitro. It was invented in 1983 by Dr. Kary Mullis,
Polymerase Chain Reaction (PCR) and Its Applications What is PCR? PCR is an exponentially progressing synthesis of the defined target DNA sequences in vitro. It was invented in 1983 by Dr. Kary Mullis,
Instructions for Use Life Science Kits & Assays
 Instructions for Use Life Science Kits & Assays Content Content 1 Product and order number... I 2 Storage conditions... I 3 Description... II 3.1 Quality data... II 3.2 Unit definition... II 4 Delivered
Instructions for Use Life Science Kits & Assays Content Content 1 Product and order number... I 2 Storage conditions... I 3 Description... II 3.1 Quality data... II 3.2 Unit definition... II 4 Delivered
Supporting Information
 Supporting Information Velocity of DNA during translocation through a solid state nanopore Calin Plesa, Nick van Loo, Philip Ketterer, Hendrik Dietz, Cees Dekker Department of Bionanoscience, Kavli Institute
Supporting Information Velocity of DNA during translocation through a solid state nanopore Calin Plesa, Nick van Loo, Philip Ketterer, Hendrik Dietz, Cees Dekker Department of Bionanoscience, Kavli Institute
The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity
 Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
Promega Notes Magazine Number 62, 1997, p. 02 The GeneEditor TM in vitro Mutagenesis System: Site- Directed Mutagenesis Using Altered Beta-Lactamase Specificity By Christine Andrews and Scott Lesley Promega
CHEM 4420 Exam I Spring 2013 Page 1 of 6
 CHEM 4420 Exam I Spring 2013 Page 1 of 6 Name Use complete sentences when requested. There are 100 possible points on this exam. The multiple choice questions are worth 2 points each. All other questions
CHEM 4420 Exam I Spring 2013 Page 1 of 6 Name Use complete sentences when requested. There are 100 possible points on this exam. The multiple choice questions are worth 2 points each. All other questions
Color Figures for: DNA Microarray Experiments: Biological and Technological Aspects
 Color Figures for: DNA Microarray Experiments: Biological and Technological Aspects Danh V. Nguyen 1, A. Bulak Arpat 2, Naisyin Wang 1, and Raymond J. Carroll 1 1 Department of Statistics, Texas A&M University,
Color Figures for: DNA Microarray Experiments: Biological and Technological Aspects Danh V. Nguyen 1, A. Bulak Arpat 2, Naisyin Wang 1, and Raymond J. Carroll 1 1 Department of Statistics, Texas A&M University,
Mechanism of Transcription Termination by RNA Polymerase III Utilizes a Non-template Strand Sequence-Specific Signal Element
 Molecular Cell Supplemental Information Mechanism of ranscription ermination by RNA Polymerase III Utilizes a Non-template Strand Sequence-Specific Signal Element Aneeshkumar G. Arimbasseri and Richard
Molecular Cell Supplemental Information Mechanism of ranscription ermination by RNA Polymerase III Utilizes a Non-template Strand Sequence-Specific Signal Element Aneeshkumar G. Arimbasseri and Richard
Amplification and Labeling of DNA for Microarray Hybridization, using the Round A/B/C Random DNA Amplification Protocol
 Amplification and Labeling of DNA for Microarray Hybridization, using the Round A/B/C Random DNA Amplification Protocol Goal: To amplify and label DNA prior to hybridization to a microarray, in a relatively
Amplification and Labeling of DNA for Microarray Hybridization, using the Round A/B/C Random DNA Amplification Protocol Goal: To amplify and label DNA prior to hybridization to a microarray, in a relatively
DNA Replication II Biochemistry 302. January 25, 2006
 DNA Replication II Biochemistry 302 January 25, 2006 Following in Dad s footsteps Original A. Kornberg E. coli DNA Pol I is a lousy replicative enzyme. 400 molecules/cell but ~2 replication forks/cell
DNA Replication II Biochemistry 302 January 25, 2006 Following in Dad s footsteps Original A. Kornberg E. coli DNA Pol I is a lousy replicative enzyme. 400 molecules/cell but ~2 replication forks/cell
Application guide for rhpcr
 Application guide for rhpcr rhpcr Primers Description Terminal bases* Synthesis scale Purification rhprimer GEN1 rddddmx 100 nmol rhprimer GEN2 rdxxdm 100 nmol Standard desalting RNase H2 Enzyme and Buffer
Application guide for rhpcr rhpcr Primers Description Terminal bases* Synthesis scale Purification rhprimer GEN1 rddddmx 100 nmol rhprimer GEN2 rdxxdm 100 nmol Standard desalting RNase H2 Enzyme and Buffer
Lecture Four. Molecular Approaches I: Nucleic Acids
 Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
Lecture Four. Molecular Approaches I: Nucleic Acids I. Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single
DNA and Its Role in Heredity. DNA and Its Role in Heredity. A. DNA: The Genetic Material. A. DNA: The Genetic Material.
 DNA and Its Role in Heredity A. DNA: The Genetic Material Lecture Series 8 DNA and Its Role in Heredity B. The Structure of DNA C. DNA E. DNA Proofreading and Repair F. Practical Applications of DNA A.
DNA and Its Role in Heredity A. DNA: The Genetic Material Lecture Series 8 DNA and Its Role in Heredity B. The Structure of DNA C. DNA E. DNA Proofreading and Repair F. Practical Applications of DNA A.
High Throughput Sequencing Technologies. J Fass UCD Genome Center Bioinformatics Core Tuesday December 16, 2014
 High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Tuesday December 16, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
High Throughput Sequencing Technologies J Fass UCD Genome Center Bioinformatics Core Tuesday December 16, 2014 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion
DNA damage alters DNA polymerase d to a form that exhibits increased discrimination against modified template bases and mismatched primers
 Published online 11 December 2008 Nucleic Acids Research, 2009, Vol. 37, No. 2 647 657 doi:10.1093/nar/gkn1000 DNA damage alters DNA polymerase d to a form that exhibits increased discrimination against
Published online 11 December 2008 Nucleic Acids Research, 2009, Vol. 37, No. 2 647 657 doi:10.1093/nar/gkn1000 DNA damage alters DNA polymerase d to a form that exhibits increased discrimination against
DNA REPLICATION & REPAIR
 DNA REPLICATION & REPAIR Table of contents 1. DNA Replication Model 2. DNA Replication Mechanism 3. DNA Repair: Proofreading 1. DNA Replication Model Replication in the cell cycle 3 models of DNA replication
DNA REPLICATION & REPAIR Table of contents 1. DNA Replication Model 2. DNA Replication Mechanism 3. DNA Repair: Proofreading 1. DNA Replication Model Replication in the cell cycle 3 models of DNA replication
Roche Molecular Biochemicals Technical Note No. LC 9/2000
 Roche Molecular Biochemicals Technical Note No. LC 9/2000 LightCycler Optimization Strategy Introduction Purpose of this Note Table of Contents The LightCycler system provides different detection formats
Roche Molecular Biochemicals Technical Note No. LC 9/2000 LightCycler Optimization Strategy Introduction Purpose of this Note Table of Contents The LightCycler system provides different detection formats
Product Name : Simple mirna Detection Kit
 Product Name : Simple mirna Detection Kit Code No. : DS700 This product is for research use only Kit Contents This kit provides sufficient reagents to perform 20 reactions for detecting microrna. Components
Product Name : Simple mirna Detection Kit Code No. : DS700 This product is for research use only Kit Contents This kit provides sufficient reagents to perform 20 reactions for detecting microrna. Components
DNA Replication I Biochemistry 302. Bob Kelm January 24, 2005
 DNA Replication I Biochemistry 302 Bob Kelm January 24, 2005 Watson Crick prediction: Each stand of parent DNA serves as a template for synthesis of a new complementary daughter strand Fig. 4.12 Proof
DNA Replication I Biochemistry 302 Bob Kelm January 24, 2005 Watson Crick prediction: Each stand of parent DNA serves as a template for synthesis of a new complementary daughter strand Fig. 4.12 Proof
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than. SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with
 Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Proposed Models of DNA Replication. Conservative Model. Semi-Conservative Model. Dispersive model
 5.2 DNA Replication Cell Cycle Life cycle of a cell Cells can reproduce Daughter cells receive an exact copy of DNA from parent cell DNA replication happens during the S phase Proposed Models of DNA Replication
5.2 DNA Replication Cell Cycle Life cycle of a cell Cells can reproduce Daughter cells receive an exact copy of DNA from parent cell DNA replication happens during the S phase Proposed Models of DNA Replication
Supplementary Information
 Supplementary Information Minimal mechanistic model of sirna-dependent target RNA slicing by recombinant human Argonaute2 protein Andrea Deerberg*, Sarah Willkomm* and Tobias Restle Institute of Molecular
Supplementary Information Minimal mechanistic model of sirna-dependent target RNA slicing by recombinant human Argonaute2 protein Andrea Deerberg*, Sarah Willkomm* and Tobias Restle Institute of Molecular
ø29 DNA Polymerase Residue Ser 122, a Single-stranded DNA Ligand for 3-5 Exonucleolysis, Is Required to Interact with the Terminal Protein*
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 273, No. 44, Issue of October 30, pp. 28966 28977, 1998 1998 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. ø29 DNA Polymerase
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 273, No. 44, Issue of October 30, pp. 28966 28977, 1998 1998 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. ø29 DNA Polymerase
The Polymerase Chain Reaction. Chapter 6: Background
 The Polymerase Chain Reaction Chapter 6: Background Invention of PCR Kary Mullis Mile marker 46.58 in April of 1983 Pulled off the road and outlined a way to conduct DNA replication in a tube Worked for
The Polymerase Chain Reaction Chapter 6: Background Invention of PCR Kary Mullis Mile marker 46.58 in April of 1983 Pulled off the road and outlined a way to conduct DNA replication in a tube Worked for
Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a
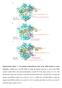 Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a HNH-state 1 in PDB 4OO8, in which the distance from the C atom of the HNH catalytic residue 840 to the
Supplementary Figure 1 Two distinct conformational states of the HNH domain in crystal structures. a HNH-state 1 in PDB 4OO8, in which the distance from the C atom of the HNH catalytic residue 840 to the
Computational Biology I LSM5191
 Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
Computational Biology I LSM5191 Lecture 5 Notes: Genetic manipulation & Molecular Biology techniques Broad Overview of: Enzymatic tools in Molecular Biology Gel electrophoresis Restriction mapping DNA
CircLigase II ssdna Ligase
 Cat. Nos. CL9021K and CL9025K Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook (facebook.com/epicentrebio), and Twitter (@EpicentreBio). www.epicentre.com Lit. # 298 12/2012 1 EPILIT298
Cat. Nos. CL9021K and CL9025K Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook (facebook.com/epicentrebio), and Twitter (@EpicentreBio). www.epicentre.com Lit. # 298 12/2012 1 EPILIT298
Genetic Fingerprinting
 Genetic Fingerprinting Introduction DA fingerprinting In the R & D sector: -involved mostly in helping to identify inherited disorders. In forensics: -identification of possible suspects involved in offences.
Genetic Fingerprinting Introduction DA fingerprinting In the R & D sector: -involved mostly in helping to identify inherited disorders. In forensics: -identification of possible suspects involved in offences.
Supporting Information
 Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany RNA ligands that distinguish metabolite-induced conformations in the TPP riboswitch Günter Mayer, Marie-Sophie L. Raddatz, Julia D. Grunwald,
Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany RNA ligands that distinguish metabolite-induced conformations in the TPP riboswitch Günter Mayer, Marie-Sophie L. Raddatz, Julia D. Grunwald,
Ch 10 Molecular Biology of the Gene
 Ch 10 Molecular Biology of the Gene For Next Week Lab -Hand in questions from 4 and 5 by TUES in my mailbox (Biology Office) -Do questions for Lab 6 for next week -Lab practical next week Lecture Read
Ch 10 Molecular Biology of the Gene For Next Week Lab -Hand in questions from 4 and 5 by TUES in my mailbox (Biology Office) -Do questions for Lab 6 for next week -Lab practical next week Lecture Read
Methods of Biomaterials Testing Lesson 3-5. Biochemical Methods - Molecular Biology -
 Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Bio Rad PCR Song Lyrics
 Bio Rad PCR Song Lyrics There was a time when to amplify DNA, You had to grow tons and tons of tiny cells. (Oooh) Then along came a guy named Dr. Kary Mullis, Said you can amplify in vitro just as well.
Bio Rad PCR Song Lyrics There was a time when to amplify DNA, You had to grow tons and tons of tiny cells. (Oooh) Then along came a guy named Dr. Kary Mullis, Said you can amplify in vitro just as well.
Laboratory #7 PCR PCR
 1 Laboratory #7 Polymerase chain reaction () is DNA replication in a test tube. In vitro enzymatic amplification of a specific segment of DNA. Many Applications. direct cloning from DNA or cdna. Mutagenesis
1 Laboratory #7 Polymerase chain reaction () is DNA replication in a test tube. In vitro enzymatic amplification of a specific segment of DNA. Many Applications. direct cloning from DNA or cdna. Mutagenesis
CHAPTER 17 FROM GENE TO PROTEIN. Section C: The Synthesis of Protein
 CHAPTER 17 FROM GENE TO PROTEIN Section C: The Synthesis of Protein 1. Translation is the RNA-directed synthesis of a polypeptide: a closer look 2. Signal peptides target some eukaryotic polypeptides to
CHAPTER 17 FROM GENE TO PROTEIN Section C: The Synthesis of Protein 1. Translation is the RNA-directed synthesis of a polypeptide: a closer look 2. Signal peptides target some eukaryotic polypeptides to
DNA Hybridization and Detection
 Chapter 6 DNA Hybridization and Detection Fluorescence Polarization Detection of DNA Hybridization........................................................ 6-2 Introduction.............................................................................................................
Chapter 6 DNA Hybridization and Detection Fluorescence Polarization Detection of DNA Hybridization........................................................ 6-2 Introduction.............................................................................................................
RNA/cDNA Quality Assay User Manual
 .Clontech Laboratories, Inc. RNA/cDNA Quality Assay User Manual (PR0X3688) Cat. No. 636841 Published 1/19/2011 Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra Bella Avenue, Mountain View, CA
.Clontech Laboratories, Inc. RNA/cDNA Quality Assay User Manual (PR0X3688) Cat. No. 636841 Published 1/19/2011 Clontech Laboratories, Inc. A Takara Bio Company 1290 Terra Bella Avenue, Mountain View, CA
RNA is a single strand molecule composed of subunits called nucleotides joined by phosphodiester bonds.
 The Versatility of RNA Primary structure of RNA RNA is a single strand molecule composed of subunits called nucleotides joined by phosphodiester bonds. Each nucleotide subunit is composed of a ribose sugar,
The Versatility of RNA Primary structure of RNA RNA is a single strand molecule composed of subunits called nucleotides joined by phosphodiester bonds. Each nucleotide subunit is composed of a ribose sugar,
NIH Public Access Author Manuscript Nat Nanotechnol. Author manuscript; available in PMC 2008 August 12.
 NIH Public Access Author Manuscript Published in final edited form as: Nat Nanotechnol. 2007 November ; 2(11): 718 724. Sequence-specific detection of individual DNA polymerase complexes in real time using
NIH Public Access Author Manuscript Published in final edited form as: Nat Nanotechnol. 2007 November ; 2(11): 718 724. Sequence-specific detection of individual DNA polymerase complexes in real time using
SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes
 WHITE PAPER SuperScript IV Reverse Transcriptase SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes Abstract Reverse transcriptases (RTs) from avian myeloblastosis virus
WHITE PAPER SuperScript IV Reverse Transcriptase SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes Abstract Reverse transcriptases (RTs) from avian myeloblastosis virus
1. A brief overview of sequencing biochemistry
 Supplementary reading materials on Genome sequencing (optional) The materials are from Mark Blaxter s lecture notes on Sequencing strategies and Primary Analysis 1. A brief overview of sequencing biochemistry
Supplementary reading materials on Genome sequencing (optional) The materials are from Mark Blaxter s lecture notes on Sequencing strategies and Primary Analysis 1. A brief overview of sequencing biochemistry
Finishing Drosophila Ananassae Fosmid 2728G16
 Finishing Drosophila Ananassae Fosmid 2728G16 Kyle Jung March 8, 2013 Bio434W Professor Elgin Page 1 Abstract For my finishing project, I chose to finish fosmid 2728G16. This fosmid carries a segment of
Finishing Drosophila Ananassae Fosmid 2728G16 Kyle Jung March 8, 2013 Bio434W Professor Elgin Page 1 Abstract For my finishing project, I chose to finish fosmid 2728G16. This fosmid carries a segment of
High Throughput Sequencing Technologies. UCD Genome Center Bioinformatics Core Monday 15 June 2015
 High Throughput Sequencing Technologies UCD Genome Center Bioinformatics Core Monday 15 June 2015 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion 2011 PacBio
High Throughput Sequencing Technologies UCD Genome Center Bioinformatics Core Monday 15 June 2015 Sequencing Explosion www.genome.gov/sequencingcosts http://t.co/ka5cvghdqo Sequencing Explosion 2011 PacBio
Independent Study Guide The Blueprint of Life, from DNA to Protein (Chapter 7)
 Independent Study Guide The Blueprint of Life, from DNA to Protein (Chapter 7) I. General Principles (Chapter 7 introduction) a. Morse code distinct series of dots and dashes encode the 26 letters of the
Independent Study Guide The Blueprint of Life, from DNA to Protein (Chapter 7) I. General Principles (Chapter 7 introduction) a. Morse code distinct series of dots and dashes encode the 26 letters of the
Premix Ex Taq (Probe qpcr)
 For Research Use Premix Ex Taq (Probe qpcr) Product Manual Table of Contents I. Description... 3 II. Principle... 4 III. Components... 5 IV. Materials Required but not Provided... 5 V. Storage... 5 VI.
For Research Use Premix Ex Taq (Probe qpcr) Product Manual Table of Contents I. Description... 3 II. Principle... 4 III. Components... 5 IV. Materials Required but not Provided... 5 V. Storage... 5 VI.
Instruction Manual cdna Synthesis System
 Instruction Manual cdna Synthesis System CAT. NO. 18267-013 Table of Contents 1. Notices to Customer........................................1 1.1 Important Information.............................................1
Instruction Manual cdna Synthesis System CAT. NO. 18267-013 Table of Contents 1. Notices to Customer........................................1 1.1 Important Information.............................................1
Prime-It II Random Primer Labeling Kit
 Prime-It II Random Primer Labeling Kit Instruction Manual Catalog #300385 Revision C0 For Research Use Only. Not for use in diagnostic procedures. 300385-12 LIMITED PRODUCT WARRANTY This warranty limits
Prime-It II Random Primer Labeling Kit Instruction Manual Catalog #300385 Revision C0 For Research Use Only. Not for use in diagnostic procedures. 300385-12 LIMITED PRODUCT WARRANTY This warranty limits
Rotor-Gene Multiplex Handbook
 Second Edition July 2011 Rotor-Gene Multiplex Handbook Rotor-Gene Multiplex PCR Kit Rotor-Gene Multiplex RT-PCR Kit For fast multiplex real-time PCR, two-step RT-PCR, and one-step RT-PCR using sequence-specific
Second Edition July 2011 Rotor-Gene Multiplex Handbook Rotor-Gene Multiplex PCR Kit Rotor-Gene Multiplex RT-PCR Kit For fast multiplex real-time PCR, two-step RT-PCR, and one-step RT-PCR using sequence-specific
Lecture for Wednesday. Dr. Prince BIOL 1408
 Lecture for Wednesday Dr. Prince BIOL 1408 THE FLOW OF GENETIC INFORMATION FROM DNA TO RNA TO PROTEIN Copyright 2009 Pearson Education, Inc. Genes are expressed as proteins A gene is a segment of DNA that
Lecture for Wednesday Dr. Prince BIOL 1408 THE FLOW OF GENETIC INFORMATION FROM DNA TO RNA TO PROTEIN Copyright 2009 Pearson Education, Inc. Genes are expressed as proteins A gene is a segment of DNA that
SequaMark DNA Size Marker
 SequaMark DNA Size Marker For sequencing and microsatellite marker sizing Catalog nos. 701001, 701004, 701005, 701006 Version B July 25, 2002 25-0565 www.invitrogen.com tech_service@invitrogen.com 2 Contents
SequaMark DNA Size Marker For sequencing and microsatellite marker sizing Catalog nos. 701001, 701004, 701005, 701006 Version B July 25, 2002 25-0565 www.invitrogen.com tech_service@invitrogen.com 2 Contents
Fig. 16-7a. 5 end Hydrogen bond 3 end. 1 nm. 3.4 nm nm
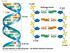 Fig. 16-7a end Hydrogen bond end 1 nm 3.4 nm 0.34 nm (a) Key features of DNA structure end (b) Partial chemical structure end Fig. 16-8 Adenine (A) Thymine (T) Guanine (G) Cytosine (C) Concept 16.2: Many
Fig. 16-7a end Hydrogen bond end 1 nm 3.4 nm 0.34 nm (a) Key features of DNA structure end (b) Partial chemical structure end Fig. 16-8 Adenine (A) Thymine (T) Guanine (G) Cytosine (C) Concept 16.2: Many
Chapter 10. DNA: The Molecule of Heredity. Lectures by Gregory Ahearn. University of North Florida. Copyright 2009 Pearson Education, Inc.
 Chapter 10 DNA: The Molecule of Heredity Lectures by Gregory Ahearn University of North Florida Copyright 2009 Pearson Education, Inc. 10.1 What Is The Structure Of DNA? Deoxyribonucleic acid (DNA) is
Chapter 10 DNA: The Molecule of Heredity Lectures by Gregory Ahearn University of North Florida Copyright 2009 Pearson Education, Inc. 10.1 What Is The Structure Of DNA? Deoxyribonucleic acid (DNA) is
HiPer RT-PCR Teaching Kit
 HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
 PRODUCT INFORMATION Long PCR Enzyme Mix #K0182 500 u Lot Exp. 00.0000 Store at -20 C. CERTIFICATE OF ANALYSIS Long PCR Enzyme Mix is functionally tested in PCR amplification of 47.4 kb fragment from lambda
PRODUCT INFORMATION Long PCR Enzyme Mix #K0182 500 u Lot Exp. 00.0000 Store at -20 C. CERTIFICATE OF ANALYSIS Long PCR Enzyme Mix is functionally tested in PCR amplification of 47.4 kb fragment from lambda
DNA Structure Requirements for the Escherichia coli Complex Clamp Loader and DNA Polymerase III Holoenzyme*
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 275, No. 15, Issue of April 14, pp. 11440 11450, 2000 2000 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. DNA Structure
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 275, No. 15, Issue of April 14, pp. 11440 11450, 2000 2000 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. DNA Structure
QUANTITATIVE RT-PCR PROTOCOL (SYBR Green I) (Last Revised: April, 2007)
 QUANTITATIVE RT-PCR PROTOCOL (SYBR Green I) (Last Revised: April, 007) Please contact Center for Plant Genomics (CPG) facility manager Hailing Jin (hljin@iastate.edu) regarding questions or corrections.
QUANTITATIVE RT-PCR PROTOCOL (SYBR Green I) (Last Revised: April, 007) Please contact Center for Plant Genomics (CPG) facility manager Hailing Jin (hljin@iastate.edu) regarding questions or corrections.
Pre-Steady-State Kinetic Studies of the Fidelity and Mechanism of Polymerization Catalyzed by Truncated Human DNA Polymerase λ
 Biochemistry 2004, 43, 6751-6762 6751 Pre-Steady-State Kinetic Studies of the Fidelity and Mechanism of Polymerization Catalyzed by Truncated Human DNA Polymerase λ Kevin A. Fiala,,, Wissam Abdel-Gawad,,,
Biochemistry 2004, 43, 6751-6762 6751 Pre-Steady-State Kinetic Studies of the Fidelity and Mechanism of Polymerization Catalyzed by Truncated Human DNA Polymerase λ Kevin A. Fiala,,, Wissam Abdel-Gawad,,,
Finishing Fosmid DMAC-27a of the Drosophila mojavensis third chromosome
 Finishing Fosmid DMAC-27a of the Drosophila mojavensis third chromosome Ruth Howe Bio 434W 27 February 2010 Abstract The fourth or dot chromosome of Drosophila species is composed primarily of highly condensed,
Finishing Fosmid DMAC-27a of the Drosophila mojavensis third chromosome Ruth Howe Bio 434W 27 February 2010 Abstract The fourth or dot chromosome of Drosophila species is composed primarily of highly condensed,
SYBR Advantage qpcr Premix. User Manual
 User Manual SYBR Advantage qpcr Premix User Manual United States/Canada 800.66.566 Asia Pacific +.650.99.7300 Europe +33.(0).3904.6880 Japan +8.(0)77.543.66 Clontech Laboratories, Inc. A Takara Bio Company
User Manual SYBR Advantage qpcr Premix User Manual United States/Canada 800.66.566 Asia Pacific +.650.99.7300 Europe +33.(0).3904.6880 Japan +8.(0)77.543.66 Clontech Laboratories, Inc. A Takara Bio Company
INTRODUCTION TO REVERSE TRANSCRIPTION PCR (RT-PCR) ABCF 2016 BecA-ILRI Hub, Nairobi 21 st September 2016 Roger Pelle Principal Scientist
 INTRODUCTION TO REVERSE TRANSCRIPTION PCR (RT-PCR) ABCF 2016 BecA-ILRI Hub, Nairobi 21 st September 2016 Roger Pelle Principal Scientist Objective of PCR To provide a solution to one of the most pressing
INTRODUCTION TO REVERSE TRANSCRIPTION PCR (RT-PCR) ABCF 2016 BecA-ILRI Hub, Nairobi 21 st September 2016 Roger Pelle Principal Scientist Objective of PCR To provide a solution to one of the most pressing
DNA Replication. The Organization of DNA. Recall:
 Recall: The Organization of DNA DNA Replication Chromosomal form appears only during mitosis, and is used in karyotypes. folded back upon itself (chromosomes) coiled around itself (chromatin) wrapped around
Recall: The Organization of DNA DNA Replication Chromosomal form appears only during mitosis, and is used in karyotypes. folded back upon itself (chromosomes) coiled around itself (chromatin) wrapped around
The replication forks Summarising what we know:
 When does replication occur? MBLG1001 lecture 10 Replication the once in a lifetime event! Full blown replication only occurs once, just before cell division BUT the DNA template is constantly being repaired.
When does replication occur? MBLG1001 lecture 10 Replication the once in a lifetime event! Full blown replication only occurs once, just before cell division BUT the DNA template is constantly being repaired.
SYBR Premix DimerEraser (Perfect Real Time)
 Cat. # RR091A or Research Use SYBR Premix DimerEraser (Perfect Real Time) Product Manual Table of Contents I. Description... 3 II. Principle... 3 III. Components... 4 IV. Storage... 5 V. eatures... 5 VI.
Cat. # RR091A or Research Use SYBR Premix DimerEraser (Perfect Real Time) Product Manual Table of Contents I. Description... 3 II. Principle... 3 III. Components... 4 IV. Storage... 5 V. eatures... 5 VI.
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Conserved arginines on the rim of Hfq catalyze base pair formation and exchange Subrata Panja and Sarah A. Woodson T.C. Jenkins Department of Biophysics, Johns Hopkins University,
SUPPLEMENTARY INFORMATION Conserved arginines on the rim of Hfq catalyze base pair formation and exchange Subrata Panja and Sarah A. Woodson T.C. Jenkins Department of Biophysics, Johns Hopkins University,
Module1TheBasicsofRealTimePCR Monday, March 19, 2007
 Objectives Slide notes: Page 1 of 41 Module 1: The Basics Of Real Time PCR Slide notes: Module 1: The Basics of real time PCR Page 2 of 41 Polymerase Chain Reaction Slide notes: Here is a review of PCR,
Objectives Slide notes: Page 1 of 41 Module 1: The Basics Of Real Time PCR Slide notes: Module 1: The Basics of real time PCR Page 2 of 41 Polymerase Chain Reaction Slide notes: Here is a review of PCR,
DNA 5 End-Labeling System INSTRUCTIONS FOR USE OF PRODUCT U2010.
 Technical Bulletin DNA 5 End-Labeling System INSTRUCTIONS FOR USE OF PRODUCT U2010. PRINTED IN USA. Revised 12/12 DNA 5 End-Labeling System All technical literature is available on the Internet at: www.promega.com/protocols/
Technical Bulletin DNA 5 End-Labeling System INSTRUCTIONS FOR USE OF PRODUCT U2010. PRINTED IN USA. Revised 12/12 DNA 5 End-Labeling System All technical literature is available on the Internet at: www.promega.com/protocols/
Student Name Biochem. 461 Exam 1 Key, September 23, 2010
 Student Name Biochem. 461 Exam 1 Key, September 23, 2010 (100 POINTS TOTAL) ANSWER ALL QUESTIONS IN THE SPACES PROVIDED [10 pts] We will learn in Chapter 28 that ribonuclosides and deoxyribonucleosides
Student Name Biochem. 461 Exam 1 Key, September 23, 2010 (100 POINTS TOTAL) ANSWER ALL QUESTIONS IN THE SPACES PROVIDED [10 pts] We will learn in Chapter 28 that ribonuclosides and deoxyribonucleosides
Replication Review. 1. What is DNA Replication? 2. Where does DNA Replication take place in eukaryotic cells?
 Replication Review 1. What is DNA Replication? 2. Where does DNA Replication take place in eukaryotic cells? 3. Where does DNA Replication take place in the cell cycle? 4. 4. What guides DNA Replication?
Replication Review 1. What is DNA Replication? 2. Where does DNA Replication take place in eukaryotic cells? 3. Where does DNA Replication take place in the cell cycle? 4. 4. What guides DNA Replication?
DNA/RNA STUDY GUIDE. Match the following scientists with their accomplishments in discovering DNA using the statement in the box below.
 Name: Period: Date: DNA/RNA STUDY GUIDE Part A: DNA History Match the following scientists with their accomplishments in discovering DNA using the statement in the box below. Used a technique called x-ray
Name: Period: Date: DNA/RNA STUDY GUIDE Part A: DNA History Match the following scientists with their accomplishments in discovering DNA using the statement in the box below. Used a technique called x-ray
From Gene to Protein transcription, messenger RNA (mrna) translation, RNA processing triplet code, template strand, codons,
 From Gene to Protein I. Transcription and translation are the two main processes linking gene to protein. A. RNA is chemically similar to DNA, except that it contains ribose as its sugar and substitutes
From Gene to Protein I. Transcription and translation are the two main processes linking gene to protein. A. RNA is chemically similar to DNA, except that it contains ribose as its sugar and substitutes
Randomly arrayed G-rich DNA sequence for label-free and realtime. assay of enzyme activity
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Randomly arrayed G-rich DNA sequence for label-free and realtime assay of enzyme activity Zhuoliang
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Randomly arrayed G-rich DNA sequence for label-free and realtime assay of enzyme activity Zhuoliang
DNA vs. RNA DNA: deoxyribonucleic acid (double stranded) RNA: ribonucleic acid (single stranded) Both found in most bacterial and eukaryotic cells RNA
 DNA Replication DNA vs. RNA DNA: deoxyribonucleic acid (double stranded) RNA: ribonucleic acid (single stranded) Both found in most bacterial and eukaryotic cells RNA molecule can assume different structures
DNA Replication DNA vs. RNA DNA: deoxyribonucleic acid (double stranded) RNA: ribonucleic acid (single stranded) Both found in most bacterial and eukaryotic cells RNA molecule can assume different structures
Pre-Lab: Molecular Biology
 Pre-Lab: Molecular Biology Name 1. What are the three chemical parts of a nucleotide. Draw a simple sketch to show how the three parts are arranged. 2. What are the rules of base pairing? 3. In double
Pre-Lab: Molecular Biology Name 1. What are the three chemical parts of a nucleotide. Draw a simple sketch to show how the three parts are arranged. 2. What are the rules of base pairing? 3. In double
IDTutorial: DNA Replication
 IDTutorial: DNA Replication Introduction In their report announcing the structure of the DNA molecule, Watson and Crick (Nature, 171: 737-738, 1953) observe, It has not escaped our notice that the specific
IDTutorial: DNA Replication Introduction In their report announcing the structure of the DNA molecule, Watson and Crick (Nature, 171: 737-738, 1953) observe, It has not escaped our notice that the specific
Detection of the P35S promoter in transgenic maize via various isothermal amplification techniques: a practical approach
 Analytical and Bioanalytical Chemistry Electronic Supplementary Material Detection of the P35S promoter in transgenic maize via various isothermal amplification techniques: a practical approach C. Zahradnik,
Analytical and Bioanalytical Chemistry Electronic Supplementary Material Detection of the P35S promoter in transgenic maize via various isothermal amplification techniques: a practical approach C. Zahradnik,
P HENIX. PHENIX PCR Enzyme Guide Tools For Life Science Discovery RESEARCH PRODUCTS
 PHENIX PCR Enzyme Guide PHENIX offers a broad line of premium quality PCR Enzymes. This PCR Enzyme Guide will help simplify your polymerase selection process. Each DNA Polymerase has different characteristics
PHENIX PCR Enzyme Guide PHENIX offers a broad line of premium quality PCR Enzymes. This PCR Enzyme Guide will help simplify your polymerase selection process. Each DNA Polymerase has different characteristics
