Supporting Online Material for
|
|
|
- Louisa Edwards
- 6 years ago
- Views:
Transcription
1 Supporting Online Material for Local Positive Feedback Regulation Determines Cell Shape in Root Hair Cells Seiji Takeda, Catherine Gapper, Hidetaka Kaya, Elizabeth Bell, Kazuyuki Kuchitsu, Liam Dolan* *To whom correspondence should be addressed. This PDF file includes: Materials and Methods Figs. S1 to S4 Tables S1 and S2 References Published 29 February 2008, Science 319, 1241 (2008) DOI: /science
2 Supporting online material (SOM) Materials and Methods Plant growth conditions and strains Arabidopsis seeds were sterilized in 5% sodium hypochlorite, washed in water and sown on Murashige and Skoog (Duchefa, Haarlem, The Netherlands) medium (ph 5.8) containing 1% sucrose and 0.8% Phytagel. New rhd2 mutants used in this study were GABI-Kat lines, 841E09 (rhd2-5) and 203A02 (rhd2-6) in which the T-DNA is inserted in the fifth exon and ninth intron, respectively (Fig. S1A). Both mutants have severe short root hair phenotype (Fig. S1C). Fluorescent protein fusion constructs GFP was amplified using GFPattB1F and GFPwosattB2R for GFP without stop codon, or GFPattB1F and GFPstopattB2R with stop codon, and introduced to pdonr207 by BP reaction (Gateway technology, Invitrogen) to generate pentrgfpwos and pentrgfpstop, respectively. For the construction of GFP:genomic fusion genes, the Gateway multisite technique (Invitrogen) was used. RHD2 promoter or promoter+gene fragments were amplified by PCR and cloned into pdonrp4p1r by BP reaction to generate pentrp4p1r clones. 3 region or gene+3 region were amplified and cloned into pdonrp2rp3 to generate pentrp2p3r clones. To generate N-terminal fusion, the promoter in pentrp4p1r, pentrgfpwos, and the gene+3 region in pentrp2rp3 were mixed with pgwbmultisite, which had been modified by M. Tomlinson (John Innes Centre, UK) from pgwb1 (a gift from T. Nakagawa, Shimane University, Japan) and used for LR reaction. For C-terminal fusion, promoter+gene in pentrp4p1r, pentrgfpstop and 3 region in pentrp2rp3 were used in the same way. Point mutated RHD2 genomic fragment was generated
3 by a PCR-based site direct mutagenesis method. The gene structure is shown in Fig. S1B. Primer sequences are listed in Table S2. Plant transformation and screening of T1 plants Agrobacterium tumefaciens strain LBA4404.pBBR1MCS virgn54d was used for plant transformation by the floral dipping method as described (S1). T1 plants were screened on MS and agar plates containing 50 µg/ml hygromycin and 50 µg/ml kanamycin. Imaging and microscopy 4-day grown plants were mounted in liquid MS media (LM: 1 x MS and 3% sucrose, ph 5.8) for imaging. GFP was imaged with Leica TCS SP system attached to Leica DMRE confocal microscope using 488 nm of an Ar/Kr laser or a CCD Hamamatsu TSU ORCA-ER digital camera attached to Nikon Eclipse E600 was used with Metamorph software. For FM4-64 treatment, 4-day old seedlings were incubated in 25 µm FM4-64 solution in LM made from 16.5 mm stock solution in DMSO. For BFA treatment 4-day seedlings were incubated in 25 µm BFA in LM made from 100 mm stock solution in DMSO for 30 min. Images were processed on ImageJ ( and Adobe photoshop CS (Adobe). Recombinant Protein Expression A cdna fragment of RHD2 corresponding to the amino acids was amplified by PCR and cloned into pgex-kg using NcoI and XhoI sites and GST added at the N-terminal end. Amino acid substitution was introduced by PCR. To express theses proteins, the plasmids were transformed into the E. coli strain Rosetta DE3 plyss (Novagen). After growth IPTG was added to the culture to a final concentration of 0.1 mm followed by a further incubation at 30 C for 3 hours. Cell pellets were resuspended in 1/10 culture volume of homogenisation buffer (50 mm Tris ph 8, 250 mm
4 NaCl, 0.5% Tween-20, 0.1% β-mercaptoethanol, Complete Protease Inhibitor cocktail (Roche Diagnostics). Cell lysis was performed by sonication on ice. The cell debris was removed by centrifugation at 4 C for 15 minutes at 10,000 g. 3 ml of glutathione agarose (Sigma, prepared as per manufacturers instructions) was added to the supernatant and this mixture was incubated with gentle agitation at 4 C for a 1 hour. The glutathione agarose was recovered by centrifugation at 4 C for 15 minutes at 3,000 rpm. The resin was applied to an empty gravity column and washed with 2 bed volumes of wash buffer (50 mm Tris ph 8, 250 mm NaCl, 0.5% Tween-20). The bound proteins were then eluted with elution buffer (50 mm Tris ph 8, 250 mm NaCl, 20mM Glutathione). Concentration was determined by Bradford Assay. Concentration was adjusted to 1 mg/ml and proteins were stored in 20% glycerol at -20 C. Protein extraction Plant roots were harvested from 14 day old Col-0 seedlings. Roots were excised from just below the hypocotyl and immediately frozen in liquid nitrogen. The frozen plant material was ground under liquid nitrogen using a mortar and pestle. 10 ml of extraction buffer (50mM HEPES ph 7.5, 5 mm EDTA, 1% w/v PVP, 1 mm β-mercaptoethanol, Roche complete protease inhibitor cocktail) was added and vortexed briefly. Samples were then centrifuged for 30 minutes at 4 C at 20,000 g. The supernatant was sterile filtered using a 22 µm filter. Protein concentration was determined by Bradford Assay. Proteins contained in the filtered supernatant were either used directly or rebuffered with an appropriate starting buffer using PD-10 columns (BioRad) and analyzed by SDS-PAGE. In vitro kinase assay For in vitro kinase assays, 5 µl of total plant protein extract or eluted proteins were incubated with 18 µl kinase assay buffer (50 mm HEPES/KOH ph 7.5, 20 mm MgCl 2, 1 mm EGTA (for Ca 2+ free
5 assays) or 10 µm CaCl 2 (for Ca 2+ assays). To this 5 µl of GST-tagged fusion proteins were added. To start the kinase reaction 2 µl of ATP mix (0.1 µl γ-32-p-atp, 0.5 µl 2mM ATP, 1.4 µl water) was added and the reaction allowed to proceed for 30 minutes at room temperature. The reaction was stopped by adding 5 µl loading buffer and incubating at 95 C for 3 minutes. The samples were then separated by SDS-PAGE. Following stain and destain, the gel was vacuum dried and exposed to a PhosphoImager screen to detect radioactive signal. ROS production assay in heterologous expression system Full length of RHD2 cdna was amplified by RT-PCR and cloned into pdonr207 by Gateway technology (Invitrogen). RHD2 carrying point mutation in phosphorylation residues and EF-hand motifs were produced by PCR. The RHD2 fragments were subcloned into pcdna3.1 with FLAG tag at its amino-terminal. Transient transfection of HEK293T cells, immune blotting and ROS measurements were carried out as described before (S2). Production of ROS was determined via chemiluminescence and expressed as relative luminescence units per second (RLU/s, means ± S. E., N=3).
6
7
8
9
10 Supplemental tables Table S1. Number of measured roots and root hairs Number of roots Number of root hairs Col rhd GFP:RHD GFP:RHD GFP:RHD E250A E250A E250A Table S2. Primer sequences used for this work Primer name Sequence (5 to 3 ) GFPattB1F GFPwosattB2R GFPstopattB2R GGGG ACA AGT TTG TAC AAA AAA GCA GGC TCA ATG AGT AAA GGA GAA GAA CTT TTC GGGG AC CAC TTT GTA CAA GAA AGC TGG GTA TTT GTA TAG TTC ATC CAT GCC GGGG AC CAC TTT GTA CAA GAA AGC TGG GTG TCA TTT GTA TAG TTC ATC CAT GCC RHD2PattB4F RHD2PattB1R RHD2GattB1R RHD2GattB2F RHD2UattB2F RHD2UattB3R GGGG ACA ACT TTG TAT AGA AAA GTT GTT AAG CTT CCT CCT GTT TCG AAT GGGG AC TGC TTT TTT GTA CAA ACT TGC TTT TTA ACA CAC TCT ACC TGA GGGG AC TGC TTT TTT GTA CAA ACT TGC GAA ATT CTC TTT GTG GAA GGA GGGG ACA GCT TTC TTG TAC AAA GTG GAA ATG TCT AGA GTG AGT TTT GAA GGGG ACA GCT TTC TTG TAC AAA GTG GAA AGG ACA CAC AGA GTT AAA AGG GGGG AC AAC TTT GTA TAA TAA AGT TGC AGA TCT ATA AGA TCA TTT CCA Supplemental references S1. S. J. Clough, A. F. Bent, Plant J 16, 735 (1998). S2. Y. Ogasawara et al., J. Biol. Chem /jbc.M (2008).
Supplemental Data Supplemental Figure 1.
 Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Arabidopsis actin depolymerizing factor AtADF4 mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB
 Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis
 1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH).
 Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were
 1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR
 Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
PCR analysis was performed to show the presence and the integrity of the var1csa and var-
 Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
ΔPDD1 x ΔPDD1. ΔPDD1 x wild type. 70 kd Pdd1. Pdd3
 Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
MacBlunt PCR Cloning Kit Manual
 MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC
 Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supporting Information
 Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
Supplementary. Table 1: Oligonucleotides and Plasmids. complementary to positions from 77 of the SRα '- GCT CTA GAG AAC TTG AAG TAC AGA CTG C
 Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR
 Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
Supplementary Information
 Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
Hes6. PPARα. PPARγ HNF4 CD36
 SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
Supplemental Information. Human Senataxin Resolves RNA/DNA Hybrids. Formed at Transcriptional Pause Sites. to Promote Xrn2-Dependent Termination
 Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells
 Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR
 Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Table S1. Bacterial strains (Related to Results and Experimental Procedures)
 Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Y-chromosomal haplogroup typing Using SBE reaction
 Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Supplemental material
 Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of
 Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Supplementary Figure 1A A404 Cells +/- Retinoic Acid
 Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
RPA-AB RPA-C Supplemental Figure S1: SDS-PAGE stained with Coomassie Blue after protein purification.
 RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
Cat. # Product Size DS130 DynaExpress TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1
 Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Legends for supplementary figures 1-3
 High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
Disease and selection in the human genome 3
 Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system
 Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Supporting information for Biochemistry, 1995, 34(34), , DOI: /bi00034a013
 Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Gene synthesis by circular assembly amplification
 Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
SUPPLEMENTARY MATERIALS AND METHODS. E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1)
 SUPPLEMENTARY MATERIALS AND METHODS E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1) dinb::kan (lab stock) derivative was used as wild-type. MG1655 alka tag dinb (2) is
SUPPLEMENTARY MATERIALS AND METHODS E. coli strains, plasmids, and growth conditions. Escherichia coli strain P90C (1) dinb::kan (lab stock) derivative was used as wild-type. MG1655 alka tag dinb (2) is
Supporting Information
 Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
An engineered tryptophan zipper-type peptide as a molecular recognition scaffold
 SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
SUPPLEMENTARY MATERIAL An engineered tryptophan zipper-type peptide as a molecular recognition scaffold Zihao Cheng and Robert E. Campbell* Supplementary Methods Library construction for FRET-based screening
hcd1tg/hj1tg/ ApoE-/- hcd1tg/hj1tg/ ApoE+/+
 ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
Nongenetic Reprogramming of the Ligand Specificity. of Growth Factor Receptors by Bispecific DNA Aptamers
 Supporting Information For Nongenetic Reprogramming of the Ligand Specificity of Growth Factor Receptors by Bispecific DNA Aptamers Ryosuke Ueki,* Saki Atsuta, Ayaka Ueki and Shinsuke Sando* Department
Supporting Information For Nongenetic Reprogramming of the Ligand Specificity of Growth Factor Receptors by Bispecific DNA Aptamers Ryosuke Ueki,* Saki Atsuta, Ayaka Ueki and Shinsuke Sando* Department
Supplemental Data. Bennett et al. (2010). Plant Cell /tpc
 BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
Supporting Online Information
 Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Supporting Information. Copyright Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, 2006
 Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supplementary Figures
 Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Supplementary Information
 Supplementary Information Load-dependent modulation of non-muscle myosin-2a function by tropomyosin 4.2 Nikolas Hundt 1, Walter Steffen 2, Salma Pathan-Chhatbar 1, Manuel H. Taft 1, Dietmar J. Manstein
Supplementary Information Load-dependent modulation of non-muscle myosin-2a function by tropomyosin 4.2 Nikolas Hundt 1, Walter Steffen 2, Salma Pathan-Chhatbar 1, Manuel H. Taft 1, Dietmar J. Manstein
Supplemental Table 1. Primers used for PCR.
 Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular
![strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular](/thumbs/72/66361212.jpg) Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Luo et al. Supplemental Figures and Materials and Methods
 Luo et al. Supplemental Figures and Materials and Methods The supplemental figures demonstrate that nuclear NFAT is situated at PODs, overexpressed PML does not increase NFAT nuclear localization, and
Luo et al. Supplemental Figures and Materials and Methods The supplemental figures demonstrate that nuclear NFAT is situated at PODs, overexpressed PML does not increase NFAT nuclear localization, and
Dierks Supplementary Fig. S1
 Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
High-throughput cloning and expression in recalcitrant bacteria
 High-throughput cloning and expression in recalcitrant bacteria Eric R Geertsma & Bert Poolman Supplementary text and figures: Supplementary Figure 1 Frequency of SfiI sites yielding identical 3 extensions
High-throughput cloning and expression in recalcitrant bacteria Eric R Geertsma & Bert Poolman Supplementary text and figures: Supplementary Figure 1 Frequency of SfiI sites yielding identical 3 extensions
Multiplexing Genome-scale Engineering
 Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
S4B fluorescence (AU)
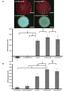 A S4B fluorescence (AU) S4B fluorescence (AU) dsbb csgba csgd dsbb csgba bcsa 5000 * NS NS 4000 * 3000 2000 1000 0 ΔcsgBAΔbcsA ΔcsgDΔdsbBΔbcsA ΔcsgBA ΔdsbBΔcsgBA ΔcsgDΔdsbB B -1000 4000 * * NS 3500 * 3000
A S4B fluorescence (AU) S4B fluorescence (AU) dsbb csgba csgd dsbb csgba bcsa 5000 * NS NS 4000 * 3000 2000 1000 0 ΔcsgBAΔbcsA ΔcsgDΔdsbBΔbcsA ΔcsgBA ΔdsbBΔcsgBA ΔcsgDΔdsbB B -1000 4000 * * NS 3500 * 3000
Amplified Analysis of DNA by the Autonomous Assembly of Polymers Consisting of DNAzyme Wires
 Supporting information for the paper: Amplified Analysis of DNA by the Autonomous Assembly of Polymers Consisting of DNAzyme Wires Fuan Wang, Johann Elbaz, Ron Orbach, Nimrod Magen and Itamar Willner*
Supporting information for the paper: Amplified Analysis of DNA by the Autonomous Assembly of Polymers Consisting of DNAzyme Wires Fuan Wang, Johann Elbaz, Ron Orbach, Nimrod Magen and Itamar Willner*
Glutathione (GSH)-Decorated Magnetic Nanoparticles for Binding Glutathione-S-transferase (GST) Fusion Protein and Manipulating Live Cells
 Glutathione (GSH)-Decorated Magnetic Nanoparticles for Binding Glutathione-S-transferase (GST) Fusion Protein and Manipulating Live Cells Yue Pan, Marcus J. C. Long, Xinming Li, Junfeng Shi, Lizbeth Hedstrom,
Glutathione (GSH)-Decorated Magnetic Nanoparticles for Binding Glutathione-S-transferase (GST) Fusion Protein and Manipulating Live Cells Yue Pan, Marcus J. C. Long, Xinming Li, Junfeng Shi, Lizbeth Hedstrom,
SUPPORTING INFORMATION
 SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
Det matematisk-naturvitenskapelige fakultet
 UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet Exam in: MBV4010 Arbeidsmetoder i molekylærbiologi og biokjemi I MBV4010 Methods in molecular biology and biochemistry I Day of exam: Friday
UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet Exam in: MBV4010 Arbeidsmetoder i molekylærbiologi og biokjemi I MBV4010 Methods in molecular biology and biochemistry I Day of exam: Friday
Overexpression Normal expression Overexpression Normal expression. 26 (21.1%) N (%) P-value a N (%)
 SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
Project 07/111 Final Report October 31, Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines
 Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Project 07/111 Final Report October 31, 2007. Project Title: Cloning and expression of porcine complement C3d for enhanced vaccines Project Leader: Dr Douglas C. Hodgins (519-824-4120 Ex 54758, fax 519-824-5930)
Supporting Information
 upporting Information hiota et al..73/pnas.159218 I Materials and Methods Yeast trains. Yeast strains used in this study are described in Table 1. TOM22FLAG, a yeast haploid strain for expression of C-terminally
upporting Information hiota et al..73/pnas.159218 I Materials and Methods Yeast trains. Yeast strains used in this study are described in Table 1. TOM22FLAG, a yeast haploid strain for expression of C-terminally
Supplementary Information. Construction of Lasso Peptide Fusion Proteins
 Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
2
 1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
1 2 3 4 5 6 7 Supplemental Table 1. Magnaporthe oryzae strains generated in this study. Strain background Genotype Strain name Description Guy-11 H1:RFP H1:RFP Strain expressing Histone H1- encoding gene
Materials Protein synthesis kit. This kit consists of 24 amino acids, 24 transfer RNAs, four messenger RNAs and one ribosome (see below).
 Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURB
 Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURBO DNA-free Kit (Ambion). One µg of total RNA was reverse
Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURBO DNA-free Kit (Ambion). One µg of total RNA was reverse
SUPPLEMENTAL DATA SUPPLEMENTAL FIGURE LEGENDS
 SUPPLEMENTAL DATA SUPPLEMENTAL FIGURE LEGENDS SUPPLEMENTAL FIGURE S1. Identification of BmCREC. (A) Amino acid sequences of BmCREC show the peptides identified in LC-MS/MS analysis (marked by red letters
SUPPLEMENTAL DATA SUPPLEMENTAL FIGURE LEGENDS SUPPLEMENTAL FIGURE S1. Identification of BmCREC. (A) Amino acid sequences of BmCREC show the peptides identified in LC-MS/MS analysis (marked by red letters
A Genetically Encoded Toolbox for Glycocalyx Engineering: Tunable Control of Cell Adhesion,
 TITLE A Genetically Encoded Toolbox for Glycocalyx Engineering: Tunable Control of Cell Adhesion, Survival, and Cancer Cell Behaviors AUTHORS Carolyn R. Shurer *, Marshall J. Colville *, Vivek K. Gupta,
TITLE A Genetically Encoded Toolbox for Glycocalyx Engineering: Tunable Control of Cell Adhesion, Survival, and Cancer Cell Behaviors AUTHORS Carolyn R. Shurer *, Marshall J. Colville *, Vivek K. Gupta,
Supplemental Data. Lin28 Mediates the Terminal Uridylation. of let-7 Precursor MicroRNA. Molecular Cell, Volume 32
 Molecular Cell, Volume 32 Supplemental Data Lin28 Mediates the Terminal Uridylation of let-7 Precursor MicroRNA Inha Heo, Chirlmin Joo, Jun Cho, Minju Ha, Jinju Han, and V. Narry Kim Figure S1. Endogenous
Molecular Cell, Volume 32 Supplemental Data Lin28 Mediates the Terminal Uridylation of let-7 Precursor MicroRNA Inha Heo, Chirlmin Joo, Jun Cho, Minju Ha, Jinju Han, and V. Narry Kim Figure S1. Endogenous
II 0.95 DM2 (RPP1) DM3 (At3g61540) b
 Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
NAME:... MODEL ANSWER... STUDENT NUMBER:... Maximum marks: 50. Internal Examiner: Hugh Murrell, Computer Science, UKZN
 COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
ORFs and genes. Please sit in row K or forward
 ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
Supporting Information
 Supporting Information Figure S1. (A) Schematic representation of PCP GrsA fused N-terminal to MBP with and without linker (yellow) in between PCP and MBP. The numbers denote the site of the fusion and
Supporting Information Figure S1. (A) Schematic representation of PCP GrsA fused N-terminal to MBP with and without linker (yellow) in between PCP and MBP. The numbers denote the site of the fusion and
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer
 TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
Lecture 11: Gene Prediction
 Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Supplemental Information. Target-Mediated Protection of Endogenous. MicroRNAs in C. elegans. Inventory of Supplementary Information
 Developmental Cell, Volume 20 Supplemental Information Target-Mediated Protection of Endogenous MicroRNAs in C. elegans Saibal Chatterjee, Monika Fasler, Ingo Büssing, and Helge Großhans Inventory of Supplementary
Developmental Cell, Volume 20 Supplemental Information Target-Mediated Protection of Endogenous MicroRNAs in C. elegans Saibal Chatterjee, Monika Fasler, Ingo Büssing, and Helge Großhans Inventory of Supplementary
SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING
 SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING All of the patients and control subjects were sequenced and genotyped in the same way. Shotgun libraries of approximately 250 bp
SUPPLEMENTAL MATERIAL GENOTYPING WITH MULTIPLEXING TARGETED RESEQUENCING All of the patients and control subjects were sequenced and genotyped in the same way. Shotgun libraries of approximately 250 bp
Supporting Information
 Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Multiplexed Detection of Lung Cancer Cells at the Single-Molecule
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Multiplexed Detection of Lung Cancer Cells at the Single-Molecule
SUPPORTING INFORMATION FILE
 Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
SUPPLEMENTAL TABLE S1. Additional descriptions of plasmid constructions and the oligonucleotides used Plasmid or Oligonucleotide
 SUPPLEMENTAL TABLE S1. Additional descriptions of plasmid constructions and the oligonucleotides used Plasmid or Oligonucleotide former/ working Description a designation Plasmids pes213a b pes213-tn5
SUPPLEMENTAL TABLE S1. Additional descriptions of plasmid constructions and the oligonucleotides used Plasmid or Oligonucleotide former/ working Description a designation Plasmids pes213a b pes213-tn5
Engineering D66N mutant using quick change site directed mutagenesis. Harkewal Singh 09/01/2010
 Engineering D66N mutant using quick change site directed mutagenesis Harkewal Singh 09/01/2010 1 1- What is quick change site directed mutagenesis? 2- An overview of the kit contents. 3- A brief information
Engineering D66N mutant using quick change site directed mutagenesis Harkewal Singh 09/01/2010 1 1- What is quick change site directed mutagenesis? 2- An overview of the kit contents. 3- A brief information
DNA sentences. How are proteins coded for by DNA? Materials. Teacher instructions. Student instructions. Reflection
 DNA sentences How are proteins coded for by DNA? Deoxyribonucleic acid (DNA) is the molecule of life. DNA is one of the most recognizable nucleic acids, a double-stranded helix. The process by which DNA
DNA sentences How are proteins coded for by DNA? Deoxyribonucleic acid (DNA) is the molecule of life. DNA is one of the most recognizable nucleic acids, a double-stranded helix. The process by which DNA
Supplementary Figure 1. The level of pri-mir-8 gradually decreases while those of BR-C and E74 increase during 3rd instar larval development.
 qrt-pcr RT-PCR Relative pri-mir-8 level 1.2 1.0 0.8 0.6 0.4 0.2 0.0 Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) BR-C E74 mtl rrna Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) Supplementary Figure 1.
qrt-pcr RT-PCR Relative pri-mir-8 level 1.2 1.0 0.8 0.6 0.4 0.2 0.0 Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) BR-C E74 mtl rrna Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) Supplementary Figure 1.
Yeast BioBrick Assembly (YBA)
 Yeast BioBrick Assembly (YBA) Standardized method for vector assembly of BioBrick devices via homologous recombination in Saccharomyces cerevisiae Martin Schneider, Leonard Fresenborg, Virginia Schadeweg
Yeast BioBrick Assembly (YBA) Standardized method for vector assembly of BioBrick devices via homologous recombination in Saccharomyces cerevisiae Martin Schneider, Leonard Fresenborg, Virginia Schadeweg
TILLING Resources for Japonica and Indica Rice
 TILLING Resources for Japonica and Indica Rice Luca Comai, Steve Henikoff, Tom Tai, Hei Leung TILLING Resources for Japonica and Indica Rice Outline Who: Seattle TILLING Project (Luca Comai, Steve Henikoff)
TILLING Resources for Japonica and Indica Rice Luca Comai, Steve Henikoff, Tom Tai, Hei Leung TILLING Resources for Japonica and Indica Rice Outline Who: Seattle TILLING Project (Luca Comai, Steve Henikoff)
Expression of Recombinant Proteins
 Expression of Recombinant Proteins Uses of Cloned Genes sequencing reagents (eg, probes) protein production insufficient natural quantities modify/mutagenesis library screening Expression Vector Features
Expression of Recombinant Proteins Uses of Cloned Genes sequencing reagents (eg, probes) protein production insufficient natural quantities modify/mutagenesis library screening Expression Vector Features
evaluated with UAS CLB eliciting UAS CIT -N Libraries increase in the
 Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Supporting Online Material
 Supporting Online Material 1. Materials and Methods 2. Supporting online figures 3. Supporting online references 1. Material and Methods Plasmid constructs All AvrPphB and protease inactive AvrPphB() constructs
Supporting Online Material 1. Materials and Methods 2. Supporting online figures 3. Supporting online references 1. Material and Methods Plasmid constructs All AvrPphB and protease inactive AvrPphB() constructs
Table S1. Sequences of mutagenesis primers used to create altered rdpa- and sdpa genes
 Supplementary Table and Figures for Structural Basis for the Enantiospecificities of R- and S-Specific Phenoxypropionate/α-Ketoglutarate Dioxygenases by Tina A. Müller, Maria I. Zavodszky, Michael Feig,
Supplementary Table and Figures for Structural Basis for the Enantiospecificities of R- and S-Specific Phenoxypropionate/α-Ketoglutarate Dioxygenases by Tina A. Müller, Maria I. Zavodszky, Michael Feig,
Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers
 DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
Supporting Information. Trifluoroacetophenone-Linked Nucleotides and DNA for Studying of DNA-protein Interactions by 19 F NMR Spectroscopy
 Supporting Information Trifluoroacetophenone-Linked Nucleotides and DNA for Studying of DNA-protein Interactions by 19 F NMR Spectroscopy Agata Olszewska, Radek Pohl and Michal Hocek # * Institute of Organic
Supporting Information Trifluoroacetophenone-Linked Nucleotides and DNA for Studying of DNA-protein Interactions by 19 F NMR Spectroscopy Agata Olszewska, Radek Pohl and Michal Hocek # * Institute of Organic
Creation of A Caspese-3 Sensing System Using A Combination of Split- GFP and Split-Intein
 Supplementary Information Creation of A Caspese-3 Sensing System Using A Combination of Split- GFP and Split-Intein Seiji Sakamoto,* Mika Terauchi, Anna Hugo, Tanner Kim, Yasuyuki Araki and Takehiko Wada*
Supplementary Information Creation of A Caspese-3 Sensing System Using A Combination of Split- GFP and Split-Intein Seiji Sakamoto,* Mika Terauchi, Anna Hugo, Tanner Kim, Yasuyuki Araki and Takehiko Wada*
PCR-based Markers and Cut Flower Longevity in Carnation
 PCRbased Markers and Cut Flower Longevity in Carnation Laura De Benedetti, Luca Braglia, Simona Bruna, Gianluca Burchi *, Antonio Mercuri and Tito Schiva Istituto Sperimentale per la Floricoltura, Corso
PCRbased Markers and Cut Flower Longevity in Carnation Laura De Benedetti, Luca Braglia, Simona Bruna, Gianluca Burchi *, Antonio Mercuri and Tito Schiva Istituto Sperimentale per la Floricoltura, Corso
Wet Lab Tutorial: Genelet Circuits
 Wet Lab Tutorial: Genelet Circuits DNA 17 This tutorial will introduce the in vitro transcriptional circuits toolkit. The tutorial will focus on the design, synthesis, and experimental testing of a synthetic
Wet Lab Tutorial: Genelet Circuits DNA 17 This tutorial will introduce the in vitro transcriptional circuits toolkit. The tutorial will focus on the design, synthesis, and experimental testing of a synthetic
G+C content. 1 Introduction. 2 Chromosomes Topology & Counts. 3 Genome size. 4 Replichores and gene orientation. 5 Chirochores.
 1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
CHE-3H84: Protein Engineering Past Exam Papers
 CHE-3H84: Protein Engineering Past Exam Papers Sorted by Topic then Year Knowledge-Based Engineering of Proteins, Large Scale Production of Recombinant Proteins, and Protein Purification Dr. Hemmings 2006/7
CHE-3H84: Protein Engineering Past Exam Papers Sorted by Topic then Year Knowledge-Based Engineering of Proteins, Large Scale Production of Recombinant Proteins, and Protein Purification Dr. Hemmings 2006/7
Supplementary Information
 Supplementary Information Microbead-based biomimetic synthetic neighbors enhance survival and function of rat pancreatic β-cells Wei Li, a Samuel Lee, b Minglin Ma, a, f Soo Min Kim, b Patrick Guye, c
Supplementary Information Microbead-based biomimetic synthetic neighbors enhance survival and function of rat pancreatic β-cells Wei Li, a Samuel Lee, b Minglin Ma, a, f Soo Min Kim, b Patrick Guye, c
Supplementary Figure 1 Tmod3 expression and phosphorylation. (a) Expression of Tmod3 in 3T3-L1 preadipocytes and differentiated adipocytes.
 1 2 3 1 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 Supplementary Figure 1 Tmod3 expression and phosphorylation. (a) Expression of Tmod3 in 3T3-L1 preadipocytes and differentiated
1 2 3 1 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 Supplementary Figure 1 Tmod3 expression and phosphorylation. (a) Expression of Tmod3 in 3T3-L1 preadipocytes and differentiated
Efficient genome replication of hepatitis B virus using adenovirus vector: a compact pregenomic RNA-expression unit
 Efficient genome replication of hepatitis B virus using adenovirus vector: a compact pregenomic RNA-expression unit Mariko Suzuki 1, Saki Kondo 1, Manabu Yamasaki 2, Norie Matsuda 2, Akio Nomoto 2, Tetsuro
Efficient genome replication of hepatitis B virus using adenovirus vector: a compact pregenomic RNA-expression unit Mariko Suzuki 1, Saki Kondo 1, Manabu Yamasaki 2, Norie Matsuda 2, Akio Nomoto 2, Tetsuro
Supplementary information
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2017 Supplementary information Structural insights into the EthR-DNA interaction from native mass spectrometry
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2017 Supplementary information Structural insights into the EthR-DNA interaction from native mass spectrometry
Supplemental Material S1, Zhang et al. (2009)
 Supplemental Material S1, Zhang et al. (2009) Cloning of PS small RNA fragments. PS RNA labeled with [γ 32 P]-ATP (NEN) of a length from 30 to 90 nucleotides were isolated from the gel after denaturing
Supplemental Material S1, Zhang et al. (2009) Cloning of PS small RNA fragments. PS RNA labeled with [γ 32 P]-ATP (NEN) of a length from 30 to 90 nucleotides were isolated from the gel after denaturing
Primer Design Workshop. École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria
 Primer Design Workshop École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria Scenario You have discovered the presence of a novel endophy5c organism living inside the cells
Primer Design Workshop École d'été en géné-que des champignons 2012 Dr. Will Hintz University of Victoria Scenario You have discovered the presence of a novel endophy5c organism living inside the cells
Event-specific Method for the Quantification of Soybean SYHT0H2 by Real-time PCR. Validated Method
 EUROPEAN COMMISSION JOINT RESEARCH CENTRE Institute for Health and Consumer Protection Molecular Biology and Genomics Unit Event-specific Method for the Quantification of Soybean SYHT0H2 by Real-time PCR
EUROPEAN COMMISSION JOINT RESEARCH CENTRE Institute for Health and Consumer Protection Molecular Biology and Genomics Unit Event-specific Method for the Quantification of Soybean SYHT0H2 by Real-time PCR
