Differential Gene Expression
|
|
|
- Winfred Horton
- 6 years ago
- Views:
Transcription
1 IBS 8102 Cell, Molecular, and Developmental Biology Differential Gene Expression January 22, 2008
2 Differential Gene Expression Chromatin structure Gene anatomy Gene sequences Control of gene transcription enhancers transcription factors Epigenetic processes (modification of structure of DNA rather than sequence) methylation imprinting epigenetics RNA processing
3 Chromatin Structure ~140 bp chromatin ~60 bp Histone lysines are protonated = net positive charge DNA PO 4 deprotonated = net negative charge electrostatic attraction ensures tight binding DNA inaccessible to polymerases, etc
4 Nucleosome Tails Core DNA bound to an octameric complex composed of two each of histones H2A, H2B, H3, and H4. From Luger et al, Nature 389: Histone H1 associates nucleosomes into higher order folded complex
5 Chromatin Acetylation Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( ) state of histones controls DNA binding C O acetylated histone (histone acetyltransferase; HAT) destabilizes = transcription allowed e.g. acetylation of K14 and K9 lysines on tail of histone H3 correlated with transcriptional competence deacetylated histones (histone deacetylases; HDAC) stabilizes = transcription repressed 3. Histone methylation (CH 3 ) transcriptional repression methylation of K9 lysine on H3 associated with heterochromatin Multiple histone regulatory modifications may occur at once; work together to change the behavior of the chromatin dynamic systematic = Histone Code reproducible
6 Chromatin Configuration John H. Frenster Heterochromatin remains tightly condensed throughout most of the cell cycle replicates later Euchromatin active chromatin
7 Anatomy of the Gene core promoter transcription initiation site region transcribed region 5 TATAT 3 +1 upstream downstream Transcription produces heterogenous nuclear RNA (hnrna)
8 hnrna Processing promoter exon 1 intron 1 exon 2 intron 2 termination region 7 methyl Gppp cap primary transcript gene transcription, 5 capping 3 cleavage, polyadenylation cleavage factor cleavage signal AAUAA poly(a)polymerase 7 methyl Gppp AAAAAAA(n) splicing 7 methyl Gppp exon 1 exon 2 AAAAAAA(n) = mature mrna, ready to be transported to the cytoplasm and translation
9 Human β globin Sequence
10 Production of β globin
11 Promoters and Enhancers RNA polymerase II binds to the promoter region at the TATA box however, Pol II cannot initiate transcription alone various proteins bind to regulatory sequences upstream and downstream of transcription initiation site promoter region binds Basal Transcription Factors promoter region 5 3 basal transcription factors facilitate Pol II binding and activity
12 Eukaryotic Transcription Initiation Complex (AKA Preinitiation Complex) Basal (General) Transcription Factors: small nuclear proteins bind to promoter region constitutive, ubiquitous bind sequentially binding of individual TFs mediated by small proteins TBP associated factors (TAFs) mediator complex (~25 proteins)
13 Eukaryotic Transcription Initiation Complex 1. TFIID complex binds to the TATA box through its TATA Binding Protein (TBP) subunit 2. TFIID is stabilized by TFIIA 3. TFIIB and TFIIH join the complex on the TATA box; TFIIE and TFIIF associate with RNA polymerase II
14 Eukaryotic Transcription Initiation Complex 4. RNA polymerase II is positioned by TFIIB, and its carboxy terminal domain (CTD) is bound by TFIID 5. The CTD is phosphorylated by TFIIH and is released by TFIID; RNA polymerase II can now transcribe mrna
15 Stabilization of Transcriptional Initiation Complex by TAFs TBP TATA binding protein TAF(s) TBP associated factors SP1, NTF 1 = Transcription factors
16 TAF Histone Acetyltransferase Activity TBP TATA binding protein TAF(s) TBP associated factors
17 Enhancers Enhancers are cis acting regulatory elements cis (same or same side); elements that reside on the same DNA strand; e.g. DNA sequences trans (other side); elements that originate from another DNA strand, e.g. regulatory proteins Enhancers: DNA sequences that regulate gene expression by affecting the transcription initiation complex on the promoter. Function bind specific regulatory proteins (i.e. specific Transcription Factors) TFs generally interact with mediator complex proteins recruit RNA polii Location highly variable with respect to the transcribed portion of the gene. upstream (5 ), downstream (3 ), or within transcribed region close proximity or as many as 10 6 bp away
18 Enhancers Enhancers differ from promoters: 1) need a promoter to work 2) can work at a distance 3) can work in reverse orientation
19 Enhancer Generalizations 1. Most gene transcription requires enhancers. 2. Enhancers are the major determinants of differential transcription in cell types and through developmental stages. 3. There can be multiple signals (e.g. multiple enhancer sites) for a given gene, and each enhancer can be bound by more than one transcription factor (not at the same time). 4. Transcription is regulated by the interaction of transcription factors bound to enhancers and the transcription initiation complex assembled at the promoter. 5. Enhancers are combinatorial. Various DNA sequences regulate temporal and spatial gene expression; these can be mixed and matched. 6. Enhancers are modular. A gene can have several enhancer elements, each of which may turn it on in different sets of cells. 7. Enhancers generally activate transcription by remodeling chromatin to expose the promoter, or by facilitating the binding of RNA polymerase to the promoter by stabilizing TAFs. 8. Enhancers can also inhibit transcription (aka Silencers).
20 Transcription Factors Proteins that bind to enhancer or promoter regions activate or repress transcription Most bind to specific DNA sequences (e.g. enhancers) Transcription factors are grouped together in families, based on structural similarities families share common framework in DNA binding sites slight differences in binding sites cause differences in recognition Arabidopsis currently 41 TF families listed eukaryotes > 75
21
22 TF/DNA Interactions estrogen receptor zinc finger helix loop helix leucine zipper
23 TF/DNA Interactions homeodomain
24 T box Transcription Factor
25 NFkB/Rel Transcription Factor
26 Transcription Factor Domains Three major domains: 1. DNA binding recognizes particular DNA sequence transcription factor engrailed
27 Transcription Factor Domains Three major domains: 1. DNA binding recognizes particular DNA sequence 2. Trans activation activates or represses transcription often involved with transcription coregulator proteins involved in binding RNA polymerase II; e.g. TFIIB, TFIIE often involved with enzymes that modify histones
28 Transcription Factor Domains Three major domains: 1. DNA binding recognizes particular DNA sequence 2. Trans activation activates or represses transcription often involved with transcription coregulator proteins involved in binding RNA polymerase II; e.g. TFIIB, TFIIE often involved with enzymes that modify histones 3. protein protein interaction domain promotes dimerization allows it to be modulated by TAFs or other transcription factors TBP TATA binding protein TAF(s) TBP associated factors NOTE alternate categories combine 2 and 3; also add signal sensing domain for some TFs
29 Transcription Factor Domains Transcription factor MITF (microphthalmia) active in ear and pigment forming cells of eye and skin; osteoclasts pigment cell specific tyrosinases expressed MITF mutation = deafness, multicolored irises, white forelock Structural Family basic helix loop helix leucine zipper Functional protein is a homodimer dimerization common among TFs The transactivating domain is located near the amino terminal when bound to a promoter or enhancer, the protein is able to bind TAF p300/cbp TAF p300/cbp is a histone acetyltransferase
30 Functional Classification of TFs I. Constitutively active e.g. SP1, NF1 CCAAT II. Conditionally active require activation II.A. Developmental (cell specific) expression tightly controlled, but once expressed requires no additional activation; e.g. GATA, HNF, PIT 1, MyoD, MNyf5, Hox, Winged Helix II.B. Signal dependent requires external signal for activation II.B1. Extracellular ligand dependent nuclear receptors II.B2. Intracellular ligand dependent activated by small intracellular molecules; e.g. SREBP, p53, orphan nuclear receptors II.B3. Cell membrane receptor dependent second messenger signaling cascades resulting in the phosphorylation of the transcription factor II.B.3.a. Resident nuclear factors reside in the nucleus regardless of activation state; e.g. CRDEB, AP 1, Mef2 II.B.3.b. Latent cytoplasmic factors inactive form reside in the cytoplasm, but when activated are translocated into the nucleus; e.g. STAT, R0SMAD, NF kb, Notch, TUBBY, NFAT A. H. Brivanlou et al., Science 295, (2002)
31 Functional Classification of TFs A. H. Brivanlou et al., Science 295, (2002)
32 Enhancer Modules Enhancers are modular: e.g. Pax6 gene expressed in the eye, pancreas, nervous system. also expressed in different genes within these tissues Within these modules, transcription factors work in a combinatorial fashion. Transcription factors operate in cascades: one stimulates the production of several others.
33 Combinatorial Enhancers combinatorial transcription factors NOTE enhancers can act as repressors/silencers Kaltoff, 2001
34 Transcription Factor Competition Kaltoff, 2001
35 Epigenetic Mechanisms Development is an epigenetic, not a genetic, process. Epigenetics epi above, etc. control of gene expression not involving sequence Mechanisms histone methylation DNA methylation imprinting sirna alternative RNA splicing translational control
36 Histone Modification Acetylation/deacetylation acetylation destabilizes nucleosome targets: lysines on histone tails Methylation stabilized nucleosome (transcription repressed) hypermethylation of H3 lysine 9 induces ectopic heterochromatin methylated histones recruit DNA methylation enzymes further stabilizes nucleosome demethylated by specific demethylating enzymes Ubiquitination vertebrates, yeast; core histones H2A, H2B, H3 destabilizes nucleosome Phosphorylation histone serine phosphorylation correlated with gene activation Others?
37 DNA Methylation DNA methylation stabilizes nucleosomes; prevents transcription found in all vertebrates; not Drosophila, nematodes, inverts degree of methylation is proportional to degree of transcription in adults tissues cytosine methylation occurs only in CpG sequences 1 5% of CpG in adult somatic cells are methylated in embryonic tissues and stem cells non CpGs nucleotides (CpA, CpT) are also methylated tumor suppressor genes silenced by methylation in many cancers
38 Embryonic DNA Methylation Methylation patterns differ in stage and tissue specific manner methylation correlates inversely with expression methylation pattern changes during development Methylation patterns are established and maintained throughout cell division by DNA (cytosine 5) methyltransferases humans: DNMT3a, DNMT3b act as de novo methyltransferases DNMT1 maintains pattern during DNA replication
39 Genomic Imprinting Special case of DNA methylation: alleles from maternal and paternal genome are differentially methylated methylation patterns can be distinguished based on resulting phenotypes No mammalian parthenogenesis because of imprinting; in mice: gynogenetic (two female pronuclei) embryos showed normal embryonic development, but poor placental development androgenetic (two male pronuclei) embryos showed poor embryonic development, but normal placental development Approximately 80 imprinted genes in humans; many involved with early developmental processes Imprinting seen in mammals, insects, flowering plants in insects, imprinting silences paternal genome produces males (functionally haploid)
40 Genomic Imprinting Imprinted alleles from maternal and paternal genomes are differentially methylated. both methylation patterns are necessary for proper development Angelman and Prader Willi syndromes: Chromosome 15 differentially methylated in males and females Deletions in either parental chromosome 15 result in predominance of reciprocal pattern Female deletion = Angelman syndrome intellectual and developmental delay speech impediment replicated by mutation of Ube3a gene (ubiquitin pathway) Male deletion = Prader Willi syndrome extreme or insatiable appetite small stature learning disabilities imprinted genes: necdin & SNRPN small nuclear ribonucleoprotein polypeptide N
41 Dosage Compensation Mammals, Drosophila, nematodes XX = female, XY = male; diploid XXs have double the X gene products as diploid XYs XX cell Drosophila transcription rate of male X is doubled C. elegans both Xs partially repressed ( = hermaphrodite) Mammals Inactivation of a single X chromosome in mammalian XX cells early embryo both active later embryo only one active Barr bodies
42 X Chromosome Inactivation Lyon hypothesis: 1. Both X chromosomes active in very early female development. 2. One X is inactivated in each cell 3. Inactivation is random 4. The process is irreversible. All progeny cells will retain the same inactivation pattern calico cat: heterozygous for coat color genes contained on early late X chromosome X chromosome inactivation
43 Differential RNA Processing Nuclear RNA selection more genes transcribed than processed into mrna unprocessed genes degraded in nucleus Alternate splicing ~75% of all human genes may be alternatively spliced (splice isoforms; splice variants) = one gene one protein family nrna splicing mediated through spliceosomes Spliceosomes constructed for each new hnrna small nuclear RNAs (snrna) splicing factor proteins Introns contain recognition sequences GU at 5 end splice site, AG at 3 end splice site (typical) also variable length polypyrimidine polypyrimidine tract (PPT) recruit spliceosome factors to site Cell specific splicing factors differ in ability to recognize intron sequences exon in one cells type might be recognized as an intron in another = alternative splicing
44 α Tropomyosin Alternative Splicing striated muscle striated muscle myoblast smooth muscle neuroblast/muscle hepatoma brain
45 Translational Control Differential mrna longevity stability dependent on length of poly(a) tail poly(a) length dependent on 3 UTR Selective inhibition of message translation (e.g. stored maternal messages) messages lacking 5 cap or 3 poly(a) tail are not translated (may be degraded) mrnas circularize mediated by proteins bound to 5 cap and poly(a) tail no 5 cap = no protein binding proximity of 5 and 3 ends important for ribosomal recognition, initiation stored mrnas often lack methylated cap at fertilization, methyltransferase completes cap mrna 5 methylated guanosine cap poly(a) binding protein eukaryotic initiation factors Rhoads, R. E. et al. Mechanism and regulation of translation in C. elegans (January 28, 2006), WormBook, ed. The C. elegans Research Community, WormBook, doi/ /wormbook , ribosomal 40 S subunit
46 MicroRNAs mirnas complementary to part of one or many mrnas animals 3 UTR; plants coding region) inhibits translation or promotes degradation stem loop 70 nt NOTE Drosha/Dicer/RISC mechanism also used to process sirna
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Differential Gene Expression
 Biology 4361 - Developmental Biology Differential Gene Expression June 18, 2009 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Biology 4361 - Developmental Biology Differential Gene Expression June 18, 2009 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Transcription in Eukaryotes
 Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Epigenetics. Medical studies in English, Lecture # 12,
 Epigenetics Medical studies in English, 2018. Lecture # 12, Epigenetics Regulation of gene activity in eukaryotes Correlation of chromatin structure with transcription stably heritable phenotype resulting
Epigenetics Medical studies in English, 2018. Lecture # 12, Epigenetics Regulation of gene activity in eukaryotes Correlation of chromatin structure with transcription stably heritable phenotype resulting
Chapter 18: Regulation of Gene Expression. 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer
 Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
Chapter 18: Regulation of Gene Expression 1. Gene Regulation in Bacteria 2. Gene Regulation in Eukaryotes 3. Gene Regulation & Cancer Gene Regulation Gene regulation refers to all aspects of controlling
BIOLOGY. Chapter 16 GenesExpression
 BIOLOGY Chapter 16 GenesExpression CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 18 Gene Expression 2014 Pearson Education, Inc. Figure 16.1 Differential Gene Expression results
BIOLOGY Chapter 16 GenesExpression CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 18 Gene Expression 2014 Pearson Education, Inc. Figure 16.1 Differential Gene Expression results
Transcription Eukaryotic Cells
 Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
Transcription Eukaryotic Cells Packet #20 1 Introduction Transcription is the process in which genetic information, stored in a strand of DNA (gene), is copied into a strand of RNA. Protein-encoding genes
Chapter 13. The Nucleus. The nucleus is the hallmark of eukaryotic cells; the very term eukaryotic means having a "true nucleus".
 Chapter 13 The Nucleus The nucleus is the hallmark of eukaryotic cells; the very term eukaryotic means having a "true nucleus". Fig.13.1. The EM of the Nucleus of a Eukaryotic Cell 13.1. The Nuclear Envelope
Chapter 13 The Nucleus The nucleus is the hallmark of eukaryotic cells; the very term eukaryotic means having a "true nucleus". Fig.13.1. The EM of the Nucleus of a Eukaryotic Cell 13.1. The Nuclear Envelope
TRANSCRIPTION AND PROCESSING OF RNA
 TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
Division Ave. High School AP Biology
 Control of Eukaryotic Genes 2007-2008 The BIG Questions n How are genes turned on & off in eukaryotes? n How do cells with the same genes differentiate to perform completely different, specialized functions?
Control of Eukaryotic Genes 2007-2008 The BIG Questions n How are genes turned on & off in eukaryotes? n How do cells with the same genes differentiate to perform completely different, specialized functions?
Gene Expression: Transcription
 Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
Structure/function relationship in DNA-binding proteins
 PHRM 836 September 22, 2015 Structure/function relationship in DNA-binding proteins Devlin Chapter 8.8-9 u General description of transcription factors (TFs) u Sequence-specific interactions between DNA
PHRM 836 September 22, 2015 Structure/function relationship in DNA-binding proteins Devlin Chapter 8.8-9 u General description of transcription factors (TFs) u Sequence-specific interactions between DNA
DNA Transcription. Dr Aliwaini
 DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
Eukaryotic Transcription
 Eukaryotic Transcription I. Differences between eukaryotic versus prokaryotic transcription. II. (core vs holoenzyme): RNA polymerase II - Promotor elements. - General Pol II transcription factors (GTF).
Eukaryotic Transcription I. Differences between eukaryotic versus prokaryotic transcription. II. (core vs holoenzyme): RNA polymerase II - Promotor elements. - General Pol II transcription factors (GTF).
Themes: RNA and RNA Processing. Messenger RNA (mrna) What is a gene? RNA is very versatile! RNA-RNA interactions are very important!
 Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat
 SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat TRANSCRIPTION: AN OVERVIEW Transcription: the synthesis of a single-stranded RNA from a doublestranded DNA template.
SIBC504: TRANSCRIPTION & RNA PROCESSING Assistant Professor Dr. Chatchawan Srisawat TRANSCRIPTION: AN OVERVIEW Transcription: the synthesis of a single-stranded RNA from a doublestranded DNA template.
Make the protein through the genetic dogma process.
 Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Year III Pharm.D Dr. V. Chitra
 Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Chromatin Structure and its Effects on Transcription
 Chromatin Structure and its Effects on Transcription Epigenetics 2014 by Nigel Atkinson The University of Texas at Austin From Weaver 4th edition and Armstrong 1st edition What is the point? DNA is not
Chromatin Structure and its Effects on Transcription Epigenetics 2014 by Nigel Atkinson The University of Texas at Austin From Weaver 4th edition and Armstrong 1st edition What is the point? DNA is not
Eukaryotic & Prokaryotic Transcription. RNA polymerases
 Eukaryotic & Prokaryotic Transcription RNA polymerases RNA Polymerases A. E. coli RNA polymerase 1. core enzyme = ββ'(α)2 has catalytic activity but cannot recognize start site of transcription ~500,000
Eukaryotic & Prokaryotic Transcription RNA polymerases RNA Polymerases A. E. coli RNA polymerase 1. core enzyme = ββ'(α)2 has catalytic activity but cannot recognize start site of transcription ~500,000
Lecture 21: Epigenetics Nurture or Nature? Chromatin DNA methylation Histone Code Twin study X-chromosome inactivation Environemnt and epigenetics
 Lecture 21: Epigenetics Nurture or Nature? Chromatin DNA methylation Histone Code Twin study X-chromosome inactivation Environemnt and epigenetics Epigenetics represents the science for the studying heritable
Lecture 21: Epigenetics Nurture or Nature? Chromatin DNA methylation Histone Code Twin study X-chromosome inactivation Environemnt and epigenetics Epigenetics represents the science for the studying heritable
CHAPTER 18 LECTURE NOTES: CONTROL OF GENE EXPRESSION PART B: CONTROL IN EUKARYOTES
 CHAPTER 18 LECTURE NOTES: CONTROL OF GENE EXPRESSION PART B: CONTROL IN EUKARYOTES I. Introduction A. No operon structures in eukaryotes B. Regulation of gene expression is frequently tissue specific.
CHAPTER 18 LECTURE NOTES: CONTROL OF GENE EXPRESSION PART B: CONTROL IN EUKARYOTES I. Introduction A. No operon structures in eukaryotes B. Regulation of gene expression is frequently tissue specific.
Regulation of Gene Expression
 Slide 1 Chapter 18 Regulation of Gene Expression PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions
Slide 1 Chapter 18 Regulation of Gene Expression PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions
DNA makes RNA makes Proteins. The Central Dogma
 DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
CHAPTER 13 LECTURE SLIDES
 CHAPTER 13 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
CHAPTER 13 LECTURE SLIDES Prepared by Brenda Leady University of Toledo To run the animations you must be in Slideshow View. Use the buttons on the animation to play, pause, and turn audio/text on or off.
Regulation of Gene Expression in Eukaryotes
 12 Regulation of Gene Expression in Eukaryotes WORKING WITH THE FIGURES 1. In Figure 12-4, certain mutations decrease the relative transcription rate of the -globin gene. Where are these mutations located,
12 Regulation of Gene Expression in Eukaryotes WORKING WITH THE FIGURES 1. In Figure 12-4, certain mutations decrease the relative transcription rate of the -globin gene. Where are these mutations located,
Chapter 24: Promoters and Enhancers
 Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
TRANSCRIPTION COMPARISON OF DNA & RNA TRANSCRIPTION. Umm AL Qura University. Sugar Ribose Deoxyribose. Bases AUCG ATCG. Strand length Short Long
 Umm AL Qura University TRANSCRIPTION Dr Neda Bogari TRANSCRIPTION COMPARISON OF DNA & RNA RNA DNA Sugar Ribose Deoxyribose Bases AUCG ATCG Strand length Short Long No. strands One Two Helix Single Double
Umm AL Qura University TRANSCRIPTION Dr Neda Bogari TRANSCRIPTION COMPARISON OF DNA & RNA RNA DNA Sugar Ribose Deoxyribose Bases AUCG ATCG Strand length Short Long No. strands One Two Helix Single Double
GENE REGULATION slide shows by Kim Foglia modified Slides with blue edges are Kim s
 GENE REGULATION slide shows by Kim Foglia modified Slides with blue edges are Kim s 2007-2008 Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
GENE REGULATION slide shows by Kim Foglia modified Slides with blue edges are Kim s 2007-2008 Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
DO NOT OPEN UNTIL TOLD TO START
 DO NOT OPEN UNTIL TOLD TO START BIO 312, Section 1: Fall 2012 December 4 th, 2012 Exam 3 Name (print neatly) Signature 7 digit student ID INSTRUCTIONS: 1. There are 12 pages to the exam. Make sure you
DO NOT OPEN UNTIL TOLD TO START BIO 312, Section 1: Fall 2012 December 4 th, 2012 Exam 3 Name (print neatly) Signature 7 digit student ID INSTRUCTIONS: 1. There are 12 pages to the exam. Make sure you
Chromatographic Separation of the three forms of RNA Polymerase II.
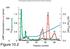 Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Exam 1 ID#: June 29, 2009
 Biology 4361 Name: KEY Exam 1 ID#: June 29, 2009 Multiple choice (one point each; indicate the best answer) 1. According to von Baer s laws, developing embryos a. pass through the adult stages of lower
Biology 4361 Name: KEY Exam 1 ID#: June 29, 2009 Multiple choice (one point each; indicate the best answer) 1. According to von Baer s laws, developing embryos a. pass through the adult stages of lower
Classes of eukaryotic cellular RNAs
 Classes of eukaryotic cellular RNAs ribosomal RNA (rrna) 18S (small subunit) 28S (large subunit) 5.8S (large subunit) 5S (large subunit) transfer RNA (trna) messenger RNA (mrna) heterogeneous nuclear RNA
Classes of eukaryotic cellular RNAs ribosomal RNA (rrna) 18S (small subunit) 28S (large subunit) 5.8S (large subunit) 5S (large subunit) transfer RNA (trna) messenger RNA (mrna) heterogeneous nuclear RNA
Chapter 16: Gene Expression from Biology by OpenStax College is licensed under a Creative Commons Attribution 3.0 Unported license.
 Chapter 16: Gene Expression from Biology by OpenStax College is licensed under a Creative Commons Attribution 3.0 Unported license. 2013, Rice University. CHAPTER 16 GENE EXPRESSION 429 16 GENE EXPRESSION
Chapter 16: Gene Expression from Biology by OpenStax College is licensed under a Creative Commons Attribution 3.0 Unported license. 2013, Rice University. CHAPTER 16 GENE EXPRESSION 429 16 GENE EXPRESSION
Fig Ch 17: From Gene to Protein
 Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
Fig. 17-1 Ch 17: From Gene to Protein Basic Principles of Transcription and Translation RNA is the intermediate between genes and the proteins for which they code Transcription is the synthesis of RNA
DNA Binding Domains: Structural Motifs. Effector Domain. Zinc Fingers. Zinc Fingers, continued. Zif268
 DNA Binding Domains: Structural Motifs Studies of known transcription factors have found several motifs of protein design to allow sequence-specific binding of DNA. We will cover only three of these motifs:
DNA Binding Domains: Structural Motifs Studies of known transcription factors have found several motifs of protein design to allow sequence-specific binding of DNA. We will cover only three of these motifs:
CLASS 3.5: 03/29/07 EUKARYOTIC TRANSCRIPTION I: PROMOTERS AND ENHANCERS
 CLASS 3.5: 03/29/07 EUKARYOTIC TRANSCRIPTION I: PROMOTERS AND ENHANCERS A. Promoters and Polymerases (RNA pols): 1. General characteristics - Initiation of transcription requires a. Transcription factors
CLASS 3.5: 03/29/07 EUKARYOTIC TRANSCRIPTION I: PROMOTERS AND ENHANCERS A. Promoters and Polymerases (RNA pols): 1. General characteristics - Initiation of transcription requires a. Transcription factors
DNA Transcription. Visualizing Transcription. The Transcription Process
 DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
Transcription & post transcriptional modification
 Transcription & post transcriptional modification Transcription The synthesis of RNA molecules using DNA strands as the templates so that the genetic information can be transferred from DNA to RNA Similarity
Transcription & post transcriptional modification Transcription The synthesis of RNA molecules using DNA strands as the templates so that the genetic information can be transferred from DNA to RNA Similarity
Higher Human Biology Unit 1: Human Cells Pupils Learning Outcomes
 Higher Human Biology Unit 1: Human Cells Pupils Learning Outcomes 1.1 Division and Differentiation in Human Cells I can state that cellular differentiation is the process by which a cell develops more
Higher Human Biology Unit 1: Human Cells Pupils Learning Outcomes 1.1 Division and Differentiation in Human Cells I can state that cellular differentiation is the process by which a cell develops more
Biochemistry Eukaryotic Transcription
 1 Description of Module Subject Name Paper Name Module Name/Title Dr. Vijaya Khader Dr. MC Varadaraj 2 1. Objectives 1. Understand and have an overview of eucaryotic transcriptional regulation. 2. Explain
1 Description of Module Subject Name Paper Name Module Name/Title Dr. Vijaya Khader Dr. MC Varadaraj 2 1. Objectives 1. Understand and have an overview of eucaryotic transcriptional regulation. 2. Explain
Review of Protein (one or more polypeptide) A polypeptide is a long chain of..
 Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Gene expression Review of Protein (one or more polypeptide) A polypeptide is a long chain of.. In a protein, the sequence of amino acid determines its which determines the protein s A protein with an enzymatic
Name Class Date. Practice Test
 Name Class Date 12 DNA Practice Test Multiple Choice Write the letter that best answers the question or completes the statement on the line provided. 1. What do bacteriophages infect? a. mice. c. viruses.
Name Class Date 12 DNA Practice Test Multiple Choice Write the letter that best answers the question or completes the statement on the line provided. 1. What do bacteriophages infect? a. mice. c. viruses.
Gene Expression and Heritable Phenotype. CBS520 Eric Nabity
 Gene Expression and Heritable Phenotype CBS520 Eric Nabity DNA is Just the Beginning DNA was determined to be the genetic material, and the structure was identified as a (double stranded) double helix.
Gene Expression and Heritable Phenotype CBS520 Eric Nabity DNA is Just the Beginning DNA was determined to be the genetic material, and the structure was identified as a (double stranded) double helix.
Chromatin and Transcription
 Chromatin and Transcription Chromatin Structure Chromatin Represses Transcription Nucleosome Positioning Histone Acetylation Chromatin Remodeling Histone Methylation CHIP Analysis Chromatin and Elongation
Chromatin and Transcription Chromatin Structure Chromatin Represses Transcription Nucleosome Positioning Histone Acetylation Chromatin Remodeling Histone Methylation CHIP Analysis Chromatin and Elongation
RNA : functional role
 RNA : functional role Hamad Yaseen, PhD MLS Department, FAHS Hamad.ali@hsc.edu.kw RNA mrna rrna trna 1 From DNA to Protein -Outline- From DNA to RNA From RNA to Protein From DNA to RNA Transcription: Copying
RNA : functional role Hamad Yaseen, PhD MLS Department, FAHS Hamad.ali@hsc.edu.kw RNA mrna rrna trna 1 From DNA to Protein -Outline- From DNA to RNA From RNA to Protein From DNA to RNA Transcription: Copying
Chapter 8 Lecture Outline. Transcription, Translation, and Bioinformatics
 Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Developmental Biology BY1101 P. Murphy
 Developmental Biology BY1101 P. Murphy Lecture 7 Cellular differentiation and the regulation of gene expression. In this lecture we looked at two main questions: How is gene expression regulated? (revision
Developmental Biology BY1101 P. Murphy Lecture 7 Cellular differentiation and the regulation of gene expression. In this lecture we looked at two main questions: How is gene expression regulated? (revision
WORKING WITH THE FIGURES. 1. In Figure 8-3, why are the arrows for genes 1 and 2 pointing in opposite directions?
 8 RNA: Transcription and Processing WORKING WITH THE FIGURES 1. In Figure 8-3, why are the arrows for genes 1 and 2 pointing in opposite directions? The arrows for genes 1 and 2 indicate the direction
8 RNA: Transcription and Processing WORKING WITH THE FIGURES 1. In Figure 8-3, why are the arrows for genes 1 and 2 pointing in opposite directions? The arrows for genes 1 and 2 indicate the direction
Einführung in die Genetik
 Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
NUCLEUS. Fig. 2. Various stages in the condensation of chromatin
 NUCLEUS Animal cells contain DNA in nucleus (contains ~ 98% of cell DNA) and mitochondrion. Both compartments are surrounded by an envelope (double membrane). Nuclear DNA represents some linear molecules
NUCLEUS Animal cells contain DNA in nucleus (contains ~ 98% of cell DNA) and mitochondrion. Both compartments are surrounded by an envelope (double membrane). Nuclear DNA represents some linear molecules
Einführung in die Genetik
 Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Twitter: @PlantDevTUM, #genetiktum FB: Plant Development TUM Prof. Dr. Claus Schwechheimer
Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Twitter: @PlantDevTUM, #genetiktum FB: Plant Development TUM Prof. Dr. Claus Schwechheimer
AP Biology Gene Expression/Biotechnology REVIEW
 AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
Protein Synthesis Notes
 Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
Protein Synthesis Notes Protein Synthesis: Overview Transcription: synthesis of mrna under the direction of DNA. Translation: actual synthesis of a polypeptide under the direction of mrna. Transcription
Chapter 14. How many genes? Control of Eukaryotic Genome. Repetitive DNA. What about the rest of the DNA? Fragile X Syndrome
 Chapter 14 Control of Eukaryotic Genome How many genes? Genes only ~3% of human genome protein-coding sequences 1% of human genome non-protein coding genes 2% of human genome trna ribosomal RNAs sirnas
Chapter 14 Control of Eukaryotic Genome How many genes? Genes only ~3% of human genome protein-coding sequences 1% of human genome non-protein coding genes 2% of human genome trna ribosomal RNAs sirnas
8/21/2014. From Gene to Protein
 From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
Epigenetics, Environment and Human Health
 Epigenetics, Environment and Human Health A. Karim Ahmed National Council for Science and the Environment Washington, DC May, 2015 Epigenetics A New Biological Paradigm A Question about Cells: All cells
Epigenetics, Environment and Human Health A. Karim Ahmed National Council for Science and the Environment Washington, DC May, 2015 Epigenetics A New Biological Paradigm A Question about Cells: All cells
REGULATION OF PROTEIN SYNTHESIS. II. Eukaryotes
 REGULATION OF PROTEIN SYNTHESIS II. Eukaryotes Complexities of eukaryotic gene expression! Several steps needed for synthesis of mrna! Separation in space of transcription and translation! Compartmentation
REGULATION OF PROTEIN SYNTHESIS II. Eukaryotes Complexities of eukaryotic gene expression! Several steps needed for synthesis of mrna! Separation in space of transcription and translation! Compartmentation
ALSO: look at figure 5-11 showing exonintron structure of the beta globin gene
 S08 Biology 205 6/4/08 Reading Assignment Chapter 7: From DNA to Protein: How cells read the genome pg 237-243 on exons and introns (you are not responsible for the biochemistry of splicing: figures 7-15,16
S08 Biology 205 6/4/08 Reading Assignment Chapter 7: From DNA to Protein: How cells read the genome pg 237-243 on exons and introns (you are not responsible for the biochemistry of splicing: figures 7-15,16
Regulatory Dynamics in Engineered Gene Networks
 Regulatory Dynamics in Engineered Gene Networks The Physico-chemical Foundation of Transcriptional Regulation with Applications to Systems Biology Mads Kærn Boston University Center for BioDynamics Center
Regulatory Dynamics in Engineered Gene Networks The Physico-chemical Foundation of Transcriptional Regulation with Applications to Systems Biology Mads Kærn Boston University Center for BioDynamics Center
DNA RNA PROTEIN SYNTHESIS -NOTES-
 DNA RNA PROTEIN SYNTHESIS -NOTES- THE COMPONENTS AND STRUCTURE OF DNA DNA is made up of units called nucleotides. Nucleotides are made up of three basic components:, called deoxyribose in DNA In DNA, there
DNA RNA PROTEIN SYNTHESIS -NOTES- THE COMPONENTS AND STRUCTURE OF DNA DNA is made up of units called nucleotides. Nucleotides are made up of three basic components:, called deoxyribose in DNA In DNA, there
Chapter 13. From DNA to Protein
 Chapter 13 From DNA to Protein Proteins All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequenceof a gene The Path From Genes to
Chapter 13 From DNA to Protein Proteins All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequenceof a gene The Path From Genes to
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION
 M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
Protein Synthesis
 HEBISD Student Expectations: Identify that RNA Is a nucleic acid with a single strand of nucleotides Contains the 5-carbon sugar ribose Contains the nitrogen bases A, G, C and U instead of T. The U is
HEBISD Student Expectations: Identify that RNA Is a nucleic acid with a single strand of nucleotides Contains the 5-carbon sugar ribose Contains the nitrogen bases A, G, C and U instead of T. The U is
Nucleic acids and protein synthesis
 THE FUNCTIONS OF DNA Nucleic acids and protein synthesis The full name of DNA is deoxyribonucleic acid. Every nucleotide has the same sugar molecule and phosphate group, but each nucleotide contains one
THE FUNCTIONS OF DNA Nucleic acids and protein synthesis The full name of DNA is deoxyribonucleic acid. Every nucleotide has the same sugar molecule and phosphate group, but each nucleotide contains one
Chapter 5 DNA and Chromosomes
 Chapter 5 DNA and Chromosomes DNA as the genetic material Heat-killed bacteria can transform living cells S Smooth R Rough Fred Griffith, 1920 DNA is the genetic material Oswald Avery Colin MacLeod Maclyn
Chapter 5 DNA and Chromosomes DNA as the genetic material Heat-killed bacteria can transform living cells S Smooth R Rough Fred Griffith, 1920 DNA is the genetic material Oswald Avery Colin MacLeod Maclyn
Eukaryotic Gene Structure
 Eukaryotic Gene Structure Terminology Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Gene Basic physical and
Eukaryotic Gene Structure Terminology Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Gene Basic physical and
Summary 12 1 DNA RNA and Protein Synthesis Chromosomes and DNA Replication. Name Class Date
 Chapter 12 Summary DNA and RNA 12 1 DNA To understand genetics, biologists had to learn the chemical structure of the gene. Frederick Griffith first learned that some factor from dead, disease-causing
Chapter 12 Summary DNA and RNA 12 1 DNA To understand genetics, biologists had to learn the chemical structure of the gene. Frederick Griffith first learned that some factor from dead, disease-causing
Bio11 Announcements. Ch 21: DNA Biology and Technology. DNA Functions. DNA and RNA Structure. How do DNA and RNA differ? What are genes?
 Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
AP BIOLOGY RNA, DNA, & Proteins Chapters 16 & 17 Review
 AP BIOLOGY RNA, DNA, & Proteins Chapters 16 & 17 Review Enzyme that adds nucleotide subunits to an RNA primer during replication DNA polymerase III Another name for protein synthesis translation Sugar
AP BIOLOGY RNA, DNA, & Proteins Chapters 16 & 17 Review Enzyme that adds nucleotide subunits to an RNA primer during replication DNA polymerase III Another name for protein synthesis translation Sugar
Biological information flow
 BCMB 3100 Chapters 36-38 Transcription & RNA Processing Biological information flow Definition of gene RNA Polymerase Gene coding vs template strand Promoter Transcription in E. coli Transcription factors
BCMB 3100 Chapters 36-38 Transcription & RNA Processing Biological information flow Definition of gene RNA Polymerase Gene coding vs template strand Promoter Transcription in E. coli Transcription factors
Chromatin. Structure and modification of chromatin. Chromatin domains
 Chromatin Structure and modification of chromatin Chromatin domains 2 DNA consensus 5 3 3 DNA DNA 4 RNA 5 ss RNA forms secondary structures with ds hairpins ds forms 6 of nucleic acids Form coiling bp/turn
Chromatin Structure and modification of chromatin Chromatin domains 2 DNA consensus 5 3 3 DNA DNA 4 RNA 5 ss RNA forms secondary structures with ds hairpins ds forms 6 of nucleic acids Form coiling bp/turn
Biological information flow
 BCMB 3100 Chapters 36-38 Transcription & RNA Processing Biological information flow Definition of gene RNA Polymerase Gene coding vs template strand Promoter Transcription in E. coli Transcription factors
BCMB 3100 Chapters 36-38 Transcription & RNA Processing Biological information flow Definition of gene RNA Polymerase Gene coding vs template strand Promoter Transcription in E. coli Transcription factors
EUKARYOTIC REGULATION C H A P T E R 1 3
 EUKARYOTIC REGULATION C H A P T E R 1 3 EUKARYOTIC REGULATION Every cell in an organism contains a complete set of DNA. But it doesn t use all of the DNA it receives Each cell chooses different DNA sequences
EUKARYOTIC REGULATION C H A P T E R 1 3 EUKARYOTIC REGULATION Every cell in an organism contains a complete set of DNA. But it doesn t use all of the DNA it receives Each cell chooses different DNA sequences
Matakuliah Genetika (BIO612206) Jurusan Biologi FMIPA Universitas Lampung. Priyambodo, M.Sc. staff.unila.ac.id/priyambodo
 Matakuliah Genetika (BIO612206) Jurusan Biologi FMIPA Universitas Lampung Priyambodo, M.Sc. staff.unila.ac.id/priyambodo Prokariotik Eukariotik staff.unila.ac.id/priyambodo Regulasi ekspresi gen pada
Matakuliah Genetika (BIO612206) Jurusan Biologi FMIPA Universitas Lampung Priyambodo, M.Sc. staff.unila.ac.id/priyambodo Prokariotik Eukariotik staff.unila.ac.id/priyambodo Regulasi ekspresi gen pada
Chapter 31. Transcription and RNA processing
 Chapter 31 Transcription and RNA processing RNA polymerase (RNAP) E. coli promoters Components of E. coli RNA Polymerase Holoenzyme (α 2 ββ'ωσ) Structure of prokaryotic RNAP The closed and open state of
Chapter 31 Transcription and RNA processing RNA polymerase (RNAP) E. coli promoters Components of E. coli RNA Polymerase Holoenzyme (α 2 ββ'ωσ) Structure of prokaryotic RNAP The closed and open state of
Trasposable elements: Uses of P elements Problem set B at the end
 Trasposable elements: Uses of P elements Problem set B at the end P-elements have revolutionized the way Drosophila geneticists conduct their research. Here, we will discuss just a few of the approaches
Trasposable elements: Uses of P elements Problem set B at the end P-elements have revolutionized the way Drosophila geneticists conduct their research. Here, we will discuss just a few of the approaches
Regulation of gene expression. (Lehninger pg )
 Regulation of gene expression (Lehninger pg. 1072-1085) Today s lecture Gene expression Constitutive, inducible, repressible genes Specificity factors, activators, repressors Negative and positive gene
Regulation of gene expression (Lehninger pg. 1072-1085) Today s lecture Gene expression Constitutive, inducible, repressible genes Specificity factors, activators, repressors Negative and positive gene
Genes - DNA - Chromosome. Chutima Talabnin Ph.D. School of Biochemistry,Institute of Science, Suranaree University of Technology
 Genes - DNA - Chromosome Chutima Talabnin Ph.D. School of Biochemistry,Institute of Science, Suranaree University of Technology DNA Cellular DNA contains genes and intragenic regions both of which may
Genes - DNA - Chromosome Chutima Talabnin Ph.D. School of Biochemistry,Institute of Science, Suranaree University of Technology DNA Cellular DNA contains genes and intragenic regions both of which may
Chapter 8: DNA and RNA
 Chapter 8: DNA and RNA Lecture Outline Enger, E. D., Ross, F. C., & Bailey, D. B. (2012). Concepts in biology (14th ed.). New York: McGraw- Hill. 1 8-1 DNA and the Importance of Proteins Proteins play
Chapter 8: DNA and RNA Lecture Outline Enger, E. D., Ross, F. C., & Bailey, D. B. (2012). Concepts in biology (14th ed.). New York: McGraw- Hill. 1 8-1 DNA and the Importance of Proteins Proteins play
PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein
 PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein This is also known as: The central dogma of molecular biology Protein Proteins are made
PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein This is also known as: The central dogma of molecular biology Protein Proteins are made
Mechanism of action-1
 Mechanism of action-1 receptors: mediators of hormone action, membrane associated vs. intracellular receptors: measurements of receptor - ligand interactions, regulation mechanisms surface - receptors:
Mechanism of action-1 receptors: mediators of hormone action, membrane associated vs. intracellular receptors: measurements of receptor - ligand interactions, regulation mechanisms surface - receptors:
BEADLE & TATUM EXPERIMENT
 FROM DNA TO PROTEINS: gene expression Chapter 14 LECTURE OBJECTIVES What Is the Evidence that Genes Code for Proteins? How Does Information Flow from Genes to Proteins? How Is the Information Content in
FROM DNA TO PROTEINS: gene expression Chapter 14 LECTURE OBJECTIVES What Is the Evidence that Genes Code for Proteins? How Does Information Flow from Genes to Proteins? How Is the Information Content in
Biological information flow
 BCMB 3100 Chapters 36-38 Transcription & RNA Processing Definition of gene RNA Polymerase Gene coding vs template strand Promoter Transcription in E. coli Transcription factors mrna processing Biological
BCMB 3100 Chapters 36-38 Transcription & RNA Processing Definition of gene RNA Polymerase Gene coding vs template strand Promoter Transcription in E. coli Transcription factors mrna processing Biological
RNA metabolism. DNA dependent synthesis of RNA RNA processing RNA dependent synthesis of RNA and DNA.
 RNA metabolism DNA dependent synthesis of RNA RNA processing RNA dependent synthesis of RNA and DNA http://www.youtube.com/watch?v=ovc8nxobxmq DNA dependent synthesis of RNA : production of an RNA molecule
RNA metabolism DNA dependent synthesis of RNA RNA processing RNA dependent synthesis of RNA and DNA http://www.youtube.com/watch?v=ovc8nxobxmq DNA dependent synthesis of RNA : production of an RNA molecule
Lecture 9 Controlling gene expression
 Lecture 9 Controlling gene expression BIOLOGY Campbell, Reece and Mitchell Chapter 18 334- (352-356) Every cell in your body contains the same number of genes approximately 35, 000 DNA is wound around
Lecture 9 Controlling gene expression BIOLOGY Campbell, Reece and Mitchell Chapter 18 334- (352-356) Every cell in your body contains the same number of genes approximately 35, 000 DNA is wound around
Bundle 5 Test Review
 Bundle 5 Test Review DNA vs. RNA DNA Replication Gene Mutations- Protein Synthesis 1. Label the different components and complete the complimentary base pairing. What is this molecule called? _Nucleic
Bundle 5 Test Review DNA vs. RNA DNA Replication Gene Mutations- Protein Synthesis 1. Label the different components and complete the complimentary base pairing. What is this molecule called? _Nucleic
3. How Genomes Function
 3. How Genomes Function In order for the cell to utilize the biological information contained within its genome, groups of genes, each gene representing a single unit of information, have to be expressed
3. How Genomes Function In order for the cell to utilize the biological information contained within its genome, groups of genes, each gene representing a single unit of information, have to be expressed
DNA and RNA. Chapter 12
 DNA and RNA Chapter 12 Warm Up Exercise Test Corrections Make sure to indicate your new answer and provide an explanation for why this is the correct answer. Do this with a red pen in the margins of your
DNA and RNA Chapter 12 Warm Up Exercise Test Corrections Make sure to indicate your new answer and provide an explanation for why this is the correct answer. Do this with a red pen in the margins of your
Bio 101 Sample questions: Chapter 10
 Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
Bio 101 Sample questions: Chapter 10 1. Which of the following is NOT needed for DNA replication? A. nucleotides B. ribosomes C. Enzymes (like polymerases) D. DNA E. all of the above are needed 2 The information
Repression of trx - introductory remarks
 Repression MBV4230 Odd S. Gabrielsen Repression of trx - introductory remarks Repression - confusing language Cis-elements termed silencers, extinguishers, operators, negatively acting sequences, URS etc.
Repression MBV4230 Odd S. Gabrielsen Repression of trx - introductory remarks Repression - confusing language Cis-elements termed silencers, extinguishers, operators, negatively acting sequences, URS etc.
RNA and PROTEIN SYNTHESIS. Chapter 13
 RNA and PROTEIN SYNTHESIS Chapter 13 DNA Double stranded Thymine Sugar is RNA Single stranded Uracil Sugar is Ribose Deoxyribose Types of RNA 1. Messenger RNA (mrna) Carries copies of instructions from
RNA and PROTEIN SYNTHESIS Chapter 13 DNA Double stranded Thymine Sugar is RNA Single stranded Uracil Sugar is Ribose Deoxyribose Types of RNA 1. Messenger RNA (mrna) Carries copies of instructions from
13.1 RNA. Lesson Objectives. Lesson Summary
 13.1 RNA Lesson Objectives Contrast RNA and DNA. Explain the process of transcription. Lesson Summary The Role of RNA RNA (ribonucleic acid) is a nucleic acid like DNA. It consists of a long chain of nucleotides.
13.1 RNA Lesson Objectives Contrast RNA and DNA. Explain the process of transcription. Lesson Summary The Role of RNA RNA (ribonucleic acid) is a nucleic acid like DNA. It consists of a long chain of nucleotides.
Nucleic acids deoxyribonucleic acid (DNA) ribonucleic acid (RNA) nucleotide
 Nucleic Acids Nucleic acids are molecules that store information for cellular growth and reproduction There are two types of nucleic acids: - deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) These
Nucleic Acids Nucleic acids are molecules that store information for cellular growth and reproduction There are two types of nucleic acids: - deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) These
Transcription Regulation And Gene Expression in Eukaryotes FS 2016 Graduate Course G2 P Matthias and RG Clerc Pharmazentrum Hörsaal 2 16h15-18h00
 Transcription Regulation And Gene Expression in Eukaryotes FS 2016 Graduate Course G2 P Matthias and RG Clerc Pharmazentrum Hörsaal 2 16h15-18h00 The general problem RG Clerc March 2, 2016 RNA Transcription
Transcription Regulation And Gene Expression in Eukaryotes FS 2016 Graduate Course G2 P Matthias and RG Clerc Pharmazentrum Hörsaal 2 16h15-18h00 The general problem RG Clerc March 2, 2016 RNA Transcription
Protein Synthesis & Gene Expression
 DNA provides the instructions for how to build proteins Each gene dictates how to build a single protein in prokaryotes The sequence of nucleotides (AGCT) in DNA dictates the order of amino acids that
DNA provides the instructions for how to build proteins Each gene dictates how to build a single protein in prokaryotes The sequence of nucleotides (AGCT) in DNA dictates the order of amino acids that
