Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon Song, Richard M. Amasino, Bosl Noh, and Yoo-Sun Noh
|
|
|
- Lesley Kennedy
- 6 years ago
- Views:
Transcription
1 Developmental Cell, Volume 22 Supplemental Information Control of Seed Germination by Light-Induced Histone Arginine Demethylation Activity Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon Song, Richard M. Amasino, Bosl Noh, and Yoo-Sun Noh Inventory of Supplemental Information Figure S1, related to Figure 1. Sequence Comparison of the JmjC Domains and the Morphological Phenotypes of jmj20, jmj22 Single, and jmj20 jmj22 Double Mutants Figure S2, related to Figure 2. The Expression of JMJ20, JMJ21, and JMJ22 mrnas upon PHYB Activation Figure S3, related to Figure 5. The H3R2me2 and H3K27me3 Levels at the GA3ox1 and GA3ox2 Loci Figure S4, related to Figure 6. The Binding of SOM to JMJ20/JMJ22 Chromatin and the PHYB-dependent Binding of JMJ20/JMJ22 to GA3ox1/GA3ox2 Chromatin Supplemental Experimental Procedures 1
2 Figure S1, related to Figure 1. Sequence Comparison of the JmjC Domains and the Morphological Phenotypes of jmj20, jmj22 Single, and jmj20 jmj22 Double Mutants (A) A multiple amino acid sequence alignment of the JmjC domains of JMJ20, JMJ21, JMJ22, human JMJD6, and human JMJD4. Asterisks and black dots are conserved catalytic residues required for the binding of the Fe (II) and 2-oxoglutarate cofactors, respectively. (B) Phylogenetic analysis of the JmjC domains of JMJ20, JMJ21, JMJ22, JMJD6, and JMJD4. The amino acid sequence alignment (A) and phylogenetic analysis (B) were performed using CLUSTAL W. (C) Expression of JMJ20 and JMJ22 transcripts in the seedlings of jmj22-1, jmj20-1, and jmj20-1 jmj22-1 as assayed by RT-PCR. The gene structures and T-DNA insertion sites are as explained in Figure 1A. The primers used for RT-PCR assays are marked as arrowheads on the diagram, and their sequences are shown in Supplemental Experimental Procedures. (D) Long-day (LD) grown adult plants of WT, jmj22-1, jmj20-1, and jmj20-1 jmj22-1. (E) Seeds harvested from WT, jmj22-1, jmj20-1, and jmj20-1 jmj
3 Figure S2, related to Figure 2. The Expression of JMJ20, JMJ21, and JMJ22 mrnas upon PHYB Activation Seeds were imbibed for 1 h in the dark and were treated with FR for 5 min with a subsequent 5 min of R treatment. Then, the seeds were incubated in the dark and were harvested at 6 h, 12 h, 24 h, or 36 h for RNA extraction. The transcript levels of GA3ox1, GA3ox2, SOM, JMJ20, JMJ21, and JMJ22 were analyzed by RT-qPCR using gene-specific primers (Supplemental Experimental Procedures). The levels at either 0 h (GA3ox1, GA3ox2, JMJ20, JMJ21, and JMJ22) or 36 h (SOM) were set to 1 after normalization by ACT2. Error bars represent sd. 3
4 Figure S3, related to Figure 5. The H3R2me2 and H3K27me3 Levels at the GA3ox1 and GA3ox2 Loci ChIP-PCR assays using light-treated seeds. 1, FR-treated WT; 2, FR-treated jmj20-1 jmj22-1; 3, R-treated WT; 4, R-treated jmj20-1 jmj22-1. The light treatment regime was as described in Figure 3, and the regions tested for PCR are as described in Figure 4A. INPUT indicates chromatin before immunoprecipitation. MOCK refers to control samples lacking antibody. (A) H3R2me2 levels. (B) H3K27me3 levels. 4
5 Figure S4, related to Figure 6. The Binding of SOM to JMJ20/JMJ22 Chromatin and the PHYB-dependent Binding of JMJ20/JMJ22 to GA3ox1/GA3ox2 Chromatin (A) Germination assay of phyb-9 jmj20-1 jmj22-1 seeds exposed to 5 min of R (left) or pil5-1 jmj20-1 jmj22-1 seeds exposed to 5 min of FR (right). Seeds were imbibed for 1 h before light treatment, and germinated seeds were counted every 24 h. Error bars represent sd (A, B, and D-H). (B) The ChIP-qPCR assay with anti-gfp antibody using WT and 35S::SOM:GFP transgenic seeds. The regions tested are shown in Figure 6F. Experimental conditions and data presentations are as described in Figure 6G. 5
6 (C) Yeast One-Hybrid assay. The promoter regions of JMJ20 and JMJ22 used for reporter constructs (A to D; upper left) and the full-length and truncated SOM proteins used for activation-domain (AD) constructs (lower left) are shown. Black boxes represent zinc-finger (ZnF) domains. Yeast cells co-transformed with each combination of reporter and AD constructs were tested for β-galactosidase assay (right). (D and E) Expression of JMJ20, JMJ22, GA3ox1, GA3ox2 transcripts in 35S::JMJ20:GFP and 35S::JMJ22:GFP transgenic seeds as measured by RT-qPCR using gene-specific primers (Supplemental Experimental Procedures). Seeds were imbibed for 1 h, treated with 5 min of FR, and incubated in the dark for 12 h before RNA extraction. (F) Germination of 35S::JMJ20:GFP and 35S::JMJ22:GFP transgenic seeds. Seeds were exposed to various fluence rate of R for 30 seconds after 5 min of FR pulse, and germinated seeds were scored after 5 d. (G) H4R3me2s levels in 35S::JMJ20:GFP and 35S::JMJ22:GFP transgenic seeds as measured by ChIP-qPCR. Seeds were treated as in (D). Normalization was to AtMu1, and the levels in WT were set to 1. The regions tested are shown in Figure 4A (G and H). (H) ChIP-qPCR analyses for the binding of JMJ20:GFP and JMJ22:GFP to GA3ox1 and GA3ox2. Seeds were imbibed for 1 h, treated with 5 min of FR with (R) or without (F) subsequent 5 min of R, and incubated in the dark for 12 h before harvesting. Normalization was to input, and the levels in F Col were set to 1. 6
7 Supplemental Experimental Procedures RT-qPCR Analysis Possible contamination from genomic DNA during the RNA extraction was removed by treating with RNase-free DNase I (Roche) for 1 h at 37ºC before RT. The first cdna strand was synthesized using M-MuLV reverse transcriptase (Fermentas) according to the manufacturer s recommendation using 3 µg of total RNA. qpcr was performed with the Applied Biosystems 7300 real-time PCR system using the SYBR Green I master mix (Kappa) in a volume of 20 µl. A comparative ΔΔC T method was used to calculate the relative amount of DNA. Normalization was to ACT2. All the RT-qPCR results were presented as means + sd of three biological replicates performed in triplicate. For the RT-qPCR experiments in Figures 3 and 6, seeds were imbibed for 1 h in the dark and were treated with FR for 5 min with (R) or without (F) a subsequent 5 min of R treatment. Then, the seeds were incubated in the dark for 12 h and were used for RNA extraction. Primers used for RT-qPCR or RT-PCR assays are as following. Gene name Sequence (5' 3') JMJ20 CATGTCACGGTTCTCTTTGGGG ATCTTCTGCGAACACGAGCG JMJ21 CCATCTGCGACTTGATACC GTAGGTGTTAAGTAGACTCCAAACA JMJ22 CCGGGGAGGTGATGTTTGTAC GAAAGAAAACTTGAAAGTATCAGTT PIL5 GTGGAATGATGCCAATGATG CTCACCCTGCTCGAACTTGG SOM GCTCTTTCGCCTTCCACTCC TCCTAGATCAGGGTCACCAC DAG1 ACACGGCTACGATGGAAACTAG GTATTGTTGTTCTCTATCTTGGGCAT GA3ox1 CCGAAGGTTTCACCATCACT CCCCAAAGGAATGCTACAGA GA3ox2 TAGATCGCATCCCATTCACA 7
8 TGGATAACTGCTTGGGTTCC GA2ox2 AATAACACGGCGGGTCTTCAAATCT TCCTCGATCTCCTTGTATCGGCTAA GAI AGCGTCATGAAACGTTGAGTCAGTG TGCCAACCCAACATGAGACAGC RGA CATTCCCGGAAACGCGATTTATCAG TCACCGTCGTTCCTATGACTCCA ABA1 GATGCAGCCAAATATGGGTCAAGG GCCATTGCATGGATAATAGCGACTC NCED6 ACCGGGTCGGATATAAATTGGGTTG CCCGGGTTGGTTCTCCTGATTC NCED9 AACCGCCGCTATGGTTTTAGACG CCAGTCACCGGAAGGTTATGCAC CYP707A2 ATGGGGTTGCCTTACATCGGAGA TGGCTTGAACAAGTGAGCTTTGCT ABI3 CTTGAAGCAAAGCGACGTGG TGTCTTACTTTAACCCCTCGTAT ACT2 AGGTCCAGGAATCGTTCACAGAAAAT AGAGGGATCAATTCGATCACTCAGAG JMJ20 a ATGGGAATACAGATTATAGGTCAAA JMJ20 b AATAAAATAGATTTGGGAGGAAAAA JMJ20 b GGTTGTATGCGTTGAGCC JMJ22 c ATGCCAAAGTGCAAGAATCTGTTGC JMJ22 d GAAAGAAAACTTGAAAGTATCAGTT ChIP Assay The antibodies used in the ChIP assays of this study were as follows: α-h3k4me3 (Cell Signaling #9727S), α-h3k9me3 (Abcam ab8898), α-h3k27me3 (Upstate ), α- H3R2me2 (Upstate ), α-h4r3me2s (Abcam ab5823), α-gus (Invitrogen A5790), and α-gfp (Invitrogen A6455). The amount of immunoprecipitated DNA was quantified by qpcr using specific primers. In the ChIP assays measuring histone methylation levels, normalizations were to the reference genes specified in each experiment. For ChIP binding assays, the amount of immunoprecipitated DNA was normalized to input DNA. All the ChIPqPCR results were presented as means + sd of three biological replicates performed in 8
9 triplicate. Primers used for ChIP-qPCR assays are as following. Locus GA3ox1 A GA3ox1 B GA3ox1 C GA3ox1 D GA3ox2 A GA3ox2 B GA3ox2 C GA3ox2 D JMJ20 P1 JMJ20 P2 JMJ20 P3 JMJ20 P4 JMJ22 P1 JMJ22 P2 JMJ22 P3 JMJ22 P4 PIL5 P1 ACT2 AtMu1 Sequence (5' 3') GTTATTGATATGGTCGTACGGTAG CAAAAATACTTTGGAAGGGAACG GACAACCCATCATGTGTATGTATC CATATGTTAAAGCACTTGTTTTGGT GTGTTTAGAGGCCATCCCATTCAC GCATGACCGATTTGGTTAGTCGCG TGAAATCTAGTGTAGTAGTGGACC CATATGTTCCTCGTACTCTTCAAC TTGTAACGGTATAAGGCTTGGC GCCTCTCACTTGCTAGTGTTATA TTGTTTTAAGCTGTCTATTTCCAAG CTTTTGGTGGAGAAGAGGAGTG GCATCCCATTCACATCCCACTCTC TGGTGATCTGGAACGCTCCCC CCTTTGGCTACATGACGATTTCTA GTCATGAGGGTCGAGTCTGT CATCACAACACTCGAGTACTTG CTCAAATCATTCAAAATGTAAGGTTC CACAGACCATGGGCCGTAC GGAGTCACGGGACAAATGC AGAAGGAGTACTTGTAGATGG GCAGCAAGTCACTCATGTAC GGCTTTTCCCTAGAGAGATC GATCAAAGTCGCCGGTAAAG TGATGAAGAGCGGAATGCTG TTTTCTCCTCAGCAAACCGC AGGCGATTATGTCGATGAGG AGACGAATCCGTCAATGAGG ATGCAGATACATGAACGGGG GTAATTCACATTATGCGCTGC CGGTGAGGCTACACTAAAAGC GTTTGACGGAGATCAGCC TCCGGTATTCGATCTTCTATC CGGAGATAAAATAATCAGCCTC AGGTCCAGGAATCGTTCACAGAAAAT AGAGGGATCAATTCGATCACTCAGAG CCGAGAACTGGTTGTGGTTT GCTCTTGCTTTGGTGATGGT 9
10 FLC TTGCATCACTCTCGTTTACCC GCGTCACAGAGAACAGAAAGC In Vitro Histone Demethylase Assay 1 or 2 µg of purified JMJ20:6xHis was incubated with 2 µg of calf thymus histone type II-A (Sigma-Aldrich) in demethylation reaction buffer (50 mm HEPES-KOH ph 8.0, 20 µm (NH 4 ) 2 Fe(SO 4 ) 2, 1 mm 2-oxoglutarate, 500 µm ascorbic acid) for 12 h at 37ºC. Histone methylation levels were measured by western blot using the following antibodies: α- H3R2me2 (Upstate ), α-h3k4me3 (Cell signaling #9727), α-h3k9me2 (Upstate ), α-h3k9me3 (Upstate ), α-h3k27me3 (Upstate ), α-h3k36me2 (Upstate ), α-h3k36me3 (Upstate ), α-h3 (Abcam ab1791), α-h4r3me1 (Abcam 17339), α-h4r3me2s (Abcam ab5823), and α-h4 (Abcam ab7311). Yeast One-hybrid Assay For the reporter constructs, 1-kb and 2-kb promoter regions of JMJ20 and JMJ22 were PCR amplified and cloned into placzi (Clonetech) at the EcoRI and SalI sites. AD constructs were generated by cloning the coding regions of SOM into pb42ad (Clonetech) at the XhoI site. As the induction of the full-length SOM protein resulted in the lethality of yeast cells, we also generated the deleted versions of AD:SOM proteins. Both the reporter and AD plasmids were co-transformed into yeast strain EGY48, and the co-transformed cells were selected on SD media lacking L-Uracil and L-Tryptophan. Selected yeast colonies were spotted on galactose and 5-bromo-4-chloro-indolyl-galactopyranoside (X-Gal)-containing media, and tested for their β-galactosidase activity. Sequences of primers used for constructions are available on request. 10
Alternative Cleavage and Polyadenylation of RNA
 Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
Supplemental Data. Lee et al. Plant Cell. (2010) /tpc Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2.
 Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination.
 Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination. Seeds of Col-0 were harvested from plants grown at 16 C, stored for 2 months, imbibed for indicated
Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination. Seeds of Col-0 were harvested from plants grown at 16 C, stored for 2 months, imbibed for indicated
Supplemental Data. Steiner et al. Plant Cell. (2012) /tpc
 Supplemental Figure 1. SPY does not interact with free GST. Invitro pull-down assay using E. coli-expressed MBP-SPY and GST, GST-TCP14 and GST-TCP15. MBP-SPY was used as bait and incubated with equal amount
Supplemental Figure 1. SPY does not interact with free GST. Invitro pull-down assay using E. coli-expressed MBP-SPY and GST, GST-TCP14 and GST-TCP15. MBP-SPY was used as bait and incubated with equal amount
Construction of plant complementation vector and generation of transgenic plants
 MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
Supporting Information. SI Material and Methods
 Supporting Information SI Material and Methods RNA extraction and qrt-pcr RNA extractions were done using the classical phenol-chloroform method. Total RNA samples were treated with Turbo DNA-free (Ambion)
Supporting Information SI Material and Methods RNA extraction and qrt-pcr RNA extractions were done using the classical phenol-chloroform method. Total RNA samples were treated with Turbo DNA-free (Ambion)
supplementary information
 DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
SOP: SYBR Green-based real-time RT-PCR
 SOP: SYBR Green-based real-time RT-PCR By Richard Yu Research fellow Centre for Marine Environmental Research and Innovative Technology (MERIT) Department of Biology and Chemistry City University of Hong
SOP: SYBR Green-based real-time RT-PCR By Richard Yu Research fellow Centre for Marine Environmental Research and Innovative Technology (MERIT) Department of Biology and Chemistry City University of Hong
RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by. 5 AGACACAAACACCAUUGUCACACUCCACAGC; Rand-2 OMe,
 Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
ASPP1 Fw GGTTGGGAATCCACGTGTTG ASPP1 Rv GCCATATCTTGGAGCTCTGAGAG
 Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
sides of the aleurone (Al) but it is excluded from the basal endosperm transfer layer
 Supplemental Data. Gómez et al. (2009). The maize transcription factor MRP-1 (Myb-Related-Protein-1) is a key regulator of the differentiation of transfer cells. Supplemental Figure 1. Expression analyses
Supplemental Data. Gómez et al. (2009). The maize transcription factor MRP-1 (Myb-Related-Protein-1) is a key regulator of the differentiation of transfer cells. Supplemental Figure 1. Expression analyses
Functional Genomics Research Stream. Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update
 Functional Genomics Research Stream Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update Updates Alternate Lab Meeting Fridays 11:30-1:00 WEL 4.224 Welcome to attend either one Lab Log thanks
Functional Genomics Research Stream Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update Updates Alternate Lab Meeting Fridays 11:30-1:00 WEL 4.224 Welcome to attend either one Lab Log thanks
Candidate region (0.74 Mb) ATC TCT GGG ACT CAT GAG CAG GAG GCT AGC ATC TCT GGG ACT CAT TAG CAG GAG GCT AGC
 A idm-3 idm-3 B Physical distance (Mb) 4.6 4.86.6 8.4 C Chr.3 Recom. Rate (%) ATG 3.9.9.9 9.74 Candidate region (.74 Mb) n=4 TAA D idm-3 G3T(E4) G4A(W988) WT idm-3 ATC TCT GGG ACT CAT GAG CAG GAG GCT AGC
A idm-3 idm-3 B Physical distance (Mb) 4.6 4.86.6 8.4 C Chr.3 Recom. Rate (%) ATG 3.9.9.9 9.74 Candidate region (.74 Mb) n=4 TAA D idm-3 G3T(E4) G4A(W988) WT idm-3 ATC TCT GGG ACT CAT GAG CAG GAG GCT AGC
Fatchiyah
 Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
Fatchiyah Email: fatchiya@yahoo.co.id RNAs: mrna trna rrna RNAi DNAs: Protein: genome DNA cdna mikro-makro mono-poly single-multi Analysis: Identification human and animal disease Finger printing Sexing
Reverse transcription-pcr (rt-pcr) Dr. Hani Alhadrami
 Reverse transcription-pcr (rt-pcr) Dr. Hani Alhadrami hanialhadrami@kau.edu.sa www.hanialhadrami.kau.edu.sa Overview Several techniques are available to detect and analyse RNA. Examples of these techniques
Reverse transcription-pcr (rt-pcr) Dr. Hani Alhadrami hanialhadrami@kau.edu.sa www.hanialhadrami.kau.edu.sa Overview Several techniques are available to detect and analyse RNA. Examples of these techniques
Enhancers mutations that make the original mutant phenotype more extreme. Suppressors mutations that make the original mutant phenotype less extreme
 Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
ReverTra Ace qpcr RT Master Mix
 Instruction manual ReverTra Ace qpcr RT Master Mix 1203 F1173K ReverTra Ace qpcr RT Master Mix FSQ-201 200 reactions Store at -20 C Contents [1] Introduction [2] Components [3] Protocol 1. RNA template
Instruction manual ReverTra Ace qpcr RT Master Mix 1203 F1173K ReverTra Ace qpcr RT Master Mix FSQ-201 200 reactions Store at -20 C Contents [1] Introduction [2] Components [3] Protocol 1. RNA template
SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes
 WHITE PAPER SuperScript IV Reverse Transcriptase SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes Abstract Reverse transcriptases (RTs) from avian myeloblastosis virus
WHITE PAPER SuperScript IV Reverse Transcriptase SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes Abstract Reverse transcriptases (RTs) from avian myeloblastosis virus
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr.
 MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
Supporting Online Material, Matsumoto et al.
 Supporting Online Material, Matsumoto et al. Material and Methods Library. Poly(A) + mrna was purified from RAW264.7 cells stimulated with murine IFN-γ (100 units/ml) and bacterial LPS (100 ng/ml) for
Supporting Online Material, Matsumoto et al. Material and Methods Library. Poly(A) + mrna was purified from RAW264.7 cells stimulated with murine IFN-γ (100 units/ml) and bacterial LPS (100 ng/ml) for
Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product.
 Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana.
 SUPPLEMENTARY FIGURES Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana. YFP:, :CFP, HA: and (A) and HA, CFP and YFPtagged and AVR1-CO39 (B) were expressed
SUPPLEMENTARY FIGURES Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana. YFP:, :CFP, HA: and (A) and HA, CFP and YFPtagged and AVR1-CO39 (B) were expressed
Supplementary Information
 Supplementary Information MLL histone methylases regulate expression of HDLR- in presence of estrogen and control plasma cholesterol in vivo Khairul I. Ansari 1, Sahba Kasiri 1, Imran Hussain 1, Samara
Supplementary Information MLL histone methylases regulate expression of HDLR- in presence of estrogen and control plasma cholesterol in vivo Khairul I. Ansari 1, Sahba Kasiri 1, Imran Hussain 1, Samara
USB HotStart-IT. for increased specificity and consistent results. PCR, qpcr and qrt-pcr
 USB HotStart-IT for increased specificity and consistent results PCR, qpcr and qrt-pcr USB PCR Reagents Choose USB HotStart-IT products for increased specificity and consistent results. Long and Accurate
USB HotStart-IT for increased specificity and consistent results PCR, qpcr and qrt-pcr USB PCR Reagents Choose USB HotStart-IT products for increased specificity and consistent results. Long and Accurate
Learning Objectives :
 Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
Technical Review. Real time PCR
 Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
mmu-mir-200a-3p Real-time RT-PCR Detection and U6 Calibration Kit User Manual MyBioSource.com Catalog # MBS826230
 mmu-mir-200a-3p Real-time RT-PCR Detection and U6 Calibration Kit User Manual Catalog # MBS826230 For the detection and quantification of mirnas mmu-mir-200a-3p normalized by U6 snrna using Real-time RT-PCR
mmu-mir-200a-3p Real-time RT-PCR Detection and U6 Calibration Kit User Manual Catalog # MBS826230 For the detection and quantification of mirnas mmu-mir-200a-3p normalized by U6 snrna using Real-time RT-PCR
Student Learning Outcomes (SLOS)
 Student Learning Outcomes (SLOS) KNOWLEDGE AND LEARNING SKILLS USE OF KNOWLEDGE AND LEARNING SKILLS - how to use Annhyb to save and manage sequences - how to use BLAST to compare sequences - how to get
Student Learning Outcomes (SLOS) KNOWLEDGE AND LEARNING SKILLS USE OF KNOWLEDGE AND LEARNING SKILLS - how to use Annhyb to save and manage sequences - how to use BLAST to compare sequences - how to get
Supporting Information
 Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany RNA ligands that distinguish metabolite-induced conformations in the TPP riboswitch Günter Mayer, Marie-Sophie L. Raddatz, Julia D. Grunwald,
Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany RNA ligands that distinguish metabolite-induced conformations in the TPP riboswitch Günter Mayer, Marie-Sophie L. Raddatz, Julia D. Grunwald,
GTTCGGGTTCC TTTTGAGCAG
 Supplementary Figures Splice variants of the SIP1 transcripts play a role in nodule organogenesis in Lotus japonicus. Wang C, Zhu H, Jin L, Chen T, Wang L, Kang H, Hong Z, Zhang Z. 5 UTR CDS 3 UTR TCTCAACCATCCTTTGTCTGCTTCCGCCGCATGGGTGAGGTCATTTTGTCTAGATGACGTGCAATTTACAATGA
Supplementary Figures Splice variants of the SIP1 transcripts play a role in nodule organogenesis in Lotus japonicus. Wang C, Zhu H, Jin L, Chen T, Wang L, Kang H, Hong Z, Zhang Z. 5 UTR CDS 3 UTR TCTCAACCATCCTTTGTCTGCTTCCGCCGCATGGGTGAGGTCATTTTGTCTAGATGACGTGCAATTTACAATGA
Supplementary Information
 Journal : Nature Biotechnology Supplementary Information Targeted genome engineering in human cells with RNA-guided endonucleases Seung Woo Cho, Sojung Kim, Jong Min Kim, and Jin-Soo Kim* National Creative
Journal : Nature Biotechnology Supplementary Information Targeted genome engineering in human cells with RNA-guided endonucleases Seung Woo Cho, Sojung Kim, Jong Min Kim, and Jin-Soo Kim* National Creative
CHAPTER 9 DNA Technologies
 CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
Genetic Engineering & Recombinant DNA
 Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
Genetic Engineering & Recombinant DNA Chapter 10 Copyright The McGraw-Hill Companies, Inc) Permission required for reproduction or display. Applications of Genetic Engineering Basic science vs. Applied
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX
 Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Recombinant DNA Technology. The Role of Recombinant DNA Technology in Biotechnology. yeast. Biotechnology. Recombinant DNA technology.
 PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
PowerPoint Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University C H A P T E R 8 Recombinant DNA Technology The Role of Recombinant DNA Technology in Biotechnology Biotechnology?
SYBR Green Realtime PCR Master Mix
 Instruction manual SYBR Green Realtime PCR Master Mix 0810 F0924K SYBR Green Realtime PCR Master Mix QPK-201T 1 ml x 1 QPK-201 1 ml x 5 Contents [1] Introduction [2] Components [3] Primer design [4] Detection
Instruction manual SYBR Green Realtime PCR Master Mix 0810 F0924K SYBR Green Realtime PCR Master Mix QPK-201T 1 ml x 1 QPK-201 1 ml x 5 Contents [1] Introduction [2] Components [3] Primer design [4] Detection
FOR RESEARCH USE ONLY. NOT FOR HUMAN OR DIAGNOSTIC USE.
 Instruction manual RNA-direct SYBR Green Realtime PCR Master Mix 0810 F0930K RNA-direct SYBR Green Realtime PCR Master Mix Contents QRT-201T QRT-201 0.5mLx2 0.5mLx5 Store at -20 C, protected from light
Instruction manual RNA-direct SYBR Green Realtime PCR Master Mix 0810 F0930K RNA-direct SYBR Green Realtime PCR Master Mix Contents QRT-201T QRT-201 0.5mLx2 0.5mLx5 Store at -20 C, protected from light
Score winning cdna yields with SuperScript III RT
 Score winning cdna yields with RT SuperScript III offers: Higher cdna yields Higher thermal stability Longer half-life than any other RT you could use Announcing SuperScript III Reverse Transcriptase How
Score winning cdna yields with RT SuperScript III offers: Higher cdna yields Higher thermal stability Longer half-life than any other RT you could use Announcing SuperScript III Reverse Transcriptase How
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
sirna Transfection Into Primary Neurons Using Fuse-It-siRNA
 sirna Transfection Into Primary Neurons Using Fuse-It-siRNA This Application Note describes a protocol for sirna transfection into sensitive, primary cortical neurons using Fuse-It-siRNA. This innovative
sirna Transfection Into Primary Neurons Using Fuse-It-siRNA This Application Note describes a protocol for sirna transfection into sensitive, primary cortical neurons using Fuse-It-siRNA. This innovative
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017
 Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
INTRODUCTION TO REVERSE TRANSCRIPTION PCR (RT-PCR) ABCF 2016 BecA-ILRI Hub, Nairobi 21 st September 2016 Roger Pelle Principal Scientist
 INTRODUCTION TO REVERSE TRANSCRIPTION PCR (RT-PCR) ABCF 2016 BecA-ILRI Hub, Nairobi 21 st September 2016 Roger Pelle Principal Scientist Objective of PCR To provide a solution to one of the most pressing
INTRODUCTION TO REVERSE TRANSCRIPTION PCR (RT-PCR) ABCF 2016 BecA-ILRI Hub, Nairobi 21 st September 2016 Roger Pelle Principal Scientist Objective of PCR To provide a solution to one of the most pressing
Mir-X mirna First-Strand Synthesis and SYBR qrt-pcr
 User Manual Mir-X mirna First-Strand Synthesis and SYBR qrt-pcr User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories,
User Manual Mir-X mirna First-Strand Synthesis and SYBR qrt-pcr User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories,
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/323/5910/124/dc1 Supporting Online Material for Regulation of Neuronal Survival Factor MEF2D by Chaperone-Mediated Autophagy Qian Yang, Hua She, Marla Gearing, Emanuela
www.sciencemag.org/cgi/content/full/323/5910/124/dc1 Supporting Online Material for Regulation of Neuronal Survival Factor MEF2D by Chaperone-Mediated Autophagy Qian Yang, Hua She, Marla Gearing, Emanuela
Quantitative Real Time PCR USING SYBR GREEN
 Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/317/5837/477/dc1 Supporting Online Material for AAV Vector Integration Sites in Mouse Hepatocellular Carcinoma Anthony Donsante, Daniel G. Miller, Yi Li, Carole Vogler,
www.sciencemag.org/cgi/content/full/317/5837/477/dc1 Supporting Online Material for AAV Vector Integration Sites in Mouse Hepatocellular Carcinoma Anthony Donsante, Daniel G. Miller, Yi Li, Carole Vogler,
Intracellular receptors specify complex patterns of gene expression that are cell and gene
 SUPPLEMENTAL RESULTS AND DISCUSSION Some HPr-1AR ARE-containing Genes Are Unresponsive to Androgen Intracellular receptors specify complex patterns of gene expression that are cell and gene specific. For
SUPPLEMENTAL RESULTS AND DISCUSSION Some HPr-1AR ARE-containing Genes Are Unresponsive to Androgen Intracellular receptors specify complex patterns of gene expression that are cell and gene specific. For
Chemically defined conditions for human ipsc derivation and culture
 Nature Methods Chemically defined conditions for human ipsc derivation and culture Guokai Chen, Daniel R Gulbranson, Zhonggang Hou, Jennifer M Bolin, Victor Ruotti, Mitchell D Probasco, Kimberly Smuga-Otto,
Nature Methods Chemically defined conditions for human ipsc derivation and culture Guokai Chen, Daniel R Gulbranson, Zhonggang Hou, Jennifer M Bolin, Victor Ruotti, Mitchell D Probasco, Kimberly Smuga-Otto,
mmu-mir-34a Real-time RT-PCR Detection Kit User Manual
 mmu-mir-34a Real-time RT-PCR Detection Kit User Manual Catalog # CPK1272 For the detection and quantification of mirna mmu-mir-34a using Real-Time RT-PCR detection instruments. For research use only. Not
mmu-mir-34a Real-time RT-PCR Detection Kit User Manual Catalog # CPK1272 For the detection and quantification of mirna mmu-mir-34a using Real-Time RT-PCR detection instruments. For research use only. Not
Supplemental Data. LMO4 Controls the Balance between Excitatory. and Inhibitory Spinal V2 Interneurons
 Neuron, Volume 61 Supplemental Data LMO4 Controls the Balance between Excitatory and Inhibitory Spinal V2 Interneurons Kaumudi Joshi, Seunghee Lee, Bora Lee, Jae W. Lee, and Soo-Kyung Lee Supplemental
Neuron, Volume 61 Supplemental Data LMO4 Controls the Balance between Excitatory and Inhibitory Spinal V2 Interneurons Kaumudi Joshi, Seunghee Lee, Bora Lee, Jae W. Lee, and Soo-Kyung Lee Supplemental
PrimeScript RT Master Mix (Perfect Real Time)
 Cat. # RR036A For Research Use PrimeScript RT Master Mix (Perfect Real Time) Product Manual Table of Contents I. Description... 3 II. Kit Components... 3 III. Materials Required but not Provided... 3 IV.
Cat. # RR036A For Research Use PrimeScript RT Master Mix (Perfect Real Time) Product Manual Table of Contents I. Description... 3 II. Kit Components... 3 III. Materials Required but not Provided... 3 IV.
GeNei TM Transformation Teaching Kit Manual
 Teaching Kit Manual Cat No. New Cat No. KT07 107385 KT07A 106220 Revision No.: 00060505 CONTENTS Page No. Objective 3 Principle 3 Kit Description 6 Materials Provided 7 Procedure 9 Observation & Interpretation
Teaching Kit Manual Cat No. New Cat No. KT07 107385 KT07A 106220 Revision No.: 00060505 CONTENTS Page No. Objective 3 Principle 3 Kit Description 6 Materials Provided 7 Procedure 9 Observation & Interpretation
Unit 6: Molecular Genetics & DNA Technology Guided Reading Questions (100 pts total)
 Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 16 The Molecular Basis of Inheritance Unit 6: Molecular Genetics
Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 16 The Molecular Basis of Inheritance Unit 6: Molecular Genetics
A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
Hossain_Supplemental Figure 1
 Hossain_Supplemental Figure 1 GFP-PACT GFP-PACT Motif I GFP-PACT Motif II A. MG132 (1µM) GFP Tubulin GFP-PACT Pericentrin GFP-PACT GFP-PACT Pericentrin Fig. S1. Expression and localization of Orc1 PACT
Hossain_Supplemental Figure 1 GFP-PACT GFP-PACT Motif I GFP-PACT Motif II A. MG132 (1µM) GFP Tubulin GFP-PACT Pericentrin GFP-PACT GFP-PACT Pericentrin Fig. S1. Expression and localization of Orc1 PACT
Nucleic Acid Research Group Real-Time PCR Survey
 Nucleic Acid Research Group Real-Time PCR Survey Association of Biomolecular Resource Facilities (ABRF) INTRODUCTION: This survey is designed to determine the current status of real-time PCR technology
Nucleic Acid Research Group Real-Time PCR Survey Association of Biomolecular Resource Facilities (ABRF) INTRODUCTION: This survey is designed to determine the current status of real-time PCR technology
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
P HENIX. PHENIX PCR Enzyme Guide Tools For Life Science Discovery RESEARCH PRODUCTS
 PHENIX PCR Enzyme Guide PHENIX offers a broad line of premium quality PCR Enzymes. This PCR Enzyme Guide will help simplify your polymerase selection process. Each DNA Polymerase has different characteristics
PHENIX PCR Enzyme Guide PHENIX offers a broad line of premium quality PCR Enzymes. This PCR Enzyme Guide will help simplify your polymerase selection process. Each DNA Polymerase has different characteristics
scgem Workflow Experimental Design Single cell DNA methylation primer design
 scgem Workflow Experimental Design Single cell DNA methylation primer design The scgem DNA methylation assay uses qpcr to measure digestion of target loci by the methylation sensitive restriction endonuclease
scgem Workflow Experimental Design Single cell DNA methylation primer design The scgem DNA methylation assay uses qpcr to measure digestion of target loci by the methylation sensitive restriction endonuclease
Histone Analysis. Histone Analysis
 Histone Analysis Histone Analysis innovative tools designed to help unravel the histone code Histone Antibodies Histone Purification Recombinant Histones Histone Modifying Enzymes Histone Modification
Histone Analysis Histone Analysis innovative tools designed to help unravel the histone code Histone Antibodies Histone Purification Recombinant Histones Histone Modifying Enzymes Histone Modification
1. Cross-linking and cell harvesting
 ChIP is a powerful tool that allows the specific matching of proteins or histone modifications to regions of the genome. Chromatin is isolated and antibodies to the antigen of interest are used to determine
ChIP is a powerful tool that allows the specific matching of proteins or histone modifications to regions of the genome. Chromatin is isolated and antibodies to the antigen of interest are used to determine
Supplementary information, Figure S1
 Supplementary information, Figure S1 (A) Schematic diagram of the sgrna and hspcas9 expression cassettes in a single binary vector designed for Agrobacterium-mediated stable transformation of Arabidopsis
Supplementary information, Figure S1 (A) Schematic diagram of the sgrna and hspcas9 expression cassettes in a single binary vector designed for Agrobacterium-mediated stable transformation of Arabidopsis
Roche Molecular Biochemicals Technical Note No. LC 10/2000
 Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
pgbkt7 Anti- Myc AH109 strain (KDa) 50
 pgbkt7 (KDa) 50 37 Anti- Myc AH109 strain Supplementary Figure 1. Protein expression of CRN and TDR in yeast. To analyse the protein expression of CRNKD and TDRKD, total proteins extracted from yeast culture
pgbkt7 (KDa) 50 37 Anti- Myc AH109 strain Supplementary Figure 1. Protein expression of CRN and TDR in yeast. To analyse the protein expression of CRNKD and TDRKD, total proteins extracted from yeast culture
Microarrays: since we use probes we obviously must know the sequences we are looking at!
 These background are needed: 1. - Basic Molecular Biology & Genetics DNA replication Transcription Post-transcriptional RNA processing Translation Post-translational protein modification Gene expression
These background are needed: 1. - Basic Molecular Biology & Genetics DNA replication Transcription Post-transcriptional RNA processing Translation Post-translational protein modification Gene expression
2054, Chap. 14, page 1
 2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
2054, Chap. 14, page 1 I. Recombinant DNA technology (Chapter 14) A. recombinant DNA technology = collection of methods used to perform genetic engineering 1. genetic engineering = deliberate modification
Biology 201 (Genetics) Exam #3 120 points 20 November Read the question carefully before answering. Think before you write.
 Name KEY Section Biology 201 (Genetics) Exam #3 120 points 20 November 2006 Read the question carefully before answering. Think before you write. You will have up to 50 minutes to take this exam. After
Name KEY Section Biology 201 (Genetics) Exam #3 120 points 20 November 2006 Read the question carefully before answering. Think before you write. You will have up to 50 minutes to take this exam. After
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or
 Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling
 Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Thyroid peroxidase gene expression is induced by lipopolysaccharide involving Nuclear Factor (NF)-κB p65 subunit phosphorylation
 1 2 3 4 5 SUPPLEMENTAL DATA Thyroid peroxidase gene expression is induced by lipopolysaccharide involving Nuclear Factor (NF)-κB p65 subunit phosphorylation Magalí Nazar, Juan Pablo Nicola, María Laura
1 2 3 4 5 SUPPLEMENTAL DATA Thyroid peroxidase gene expression is induced by lipopolysaccharide involving Nuclear Factor (NF)-κB p65 subunit phosphorylation Magalí Nazar, Juan Pablo Nicola, María Laura
PrimePCR Assay Validation Report
 Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin)
Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin)
Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup
 Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup 1. Introduction The data produced by IHEC is illustrated in Figure 1. Figure 1. The space of epigenomic
Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup 1. Introduction The data produced by IHEC is illustrated in Figure 1. Figure 1. The space of epigenomic
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
Real-Time PCR Validations
 Real-Time PCR Validations Relative gene expression Example experiment: I have 2 samples: untreated and treated. Question: What happens to the expression of gene X when I treat the cells? Answer: Expressed
Real-Time PCR Validations Relative gene expression Example experiment: I have 2 samples: untreated and treated. Question: What happens to the expression of gene X when I treat the cells? Answer: Expressed
M Keramatipour 2. M Keramatipour 1. M Keramatipour 4. M Keramatipour 3. M Keramatipour 5. M Keramatipour
 Molecular Cloning Methods Mohammad Keramatipour MD, PhD keramatipour@tums.ac.ir Outline DNA recombinant technology DNA cloning co Cell based PCR PCR-based Some application of DNA cloning Genomic libraries
Molecular Cloning Methods Mohammad Keramatipour MD, PhD keramatipour@tums.ac.ir Outline DNA recombinant technology DNA cloning co Cell based PCR PCR-based Some application of DNA cloning Genomic libraries
Supplemental Information. PARP1 Represses PAP and Inhibits Polyadenylation during Heat Shock
 Molecular Cell, Volume 49 Supplemental Information PARP1 Represses PAP and Inhibits Polyadenylation during Heat Shock Dafne Campigli Di Giammartino, Yongsheng Shi, and James L. Manley Supplemental Information
Molecular Cell, Volume 49 Supplemental Information PARP1 Represses PAP and Inhibits Polyadenylation during Heat Shock Dafne Campigli Di Giammartino, Yongsheng Shi, and James L. Manley Supplemental Information
% Viability. isw2 ino isw2 ino isw2 ino isw2 ino mM HU 4-NQO CPT
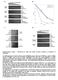 a Drug concentration b 1.3% MMS nhp1 nhp1 8 nhp1 mag1.5% MMS.3% MMS nhp1 nhp1 ino8 9 ino8 9 % Viability 4.5% MMS ino8 9 ino8 9 2.5.1.15 % MMS c d nhp1 nhp1 nhp1 nhp1 nhp1 nhp1 Control (YPD) γ IR (1 gy)
a Drug concentration b 1.3% MMS nhp1 nhp1 8 nhp1 mag1.5% MMS.3% MMS nhp1 nhp1 ino8 9 ino8 9 % Viability 4.5% MMS ino8 9 ino8 9 2.5.1.15 % MMS c d nhp1 nhp1 nhp1 nhp1 nhp1 nhp1 Control (YPD) γ IR (1 gy)
Figure S1. Figure S2. Figure S3 HB Anti-FSP27 (COOH-terminal peptide) Ab. Anti-GST-FSP27(45-127) Ab.
 / 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
/ 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
Quant One Step RT-PCR Kit
 1. Quant One Step RT-PCR Kit For fast and sensitive one-step RT-PCR www.tiangen.com/en RT121221 Quant One Step RT-PCR Kit Kit Contents Cat. no. KR113 Contents Hotmaster Taq Polymerase (2.5 U/μl) Quant
1. Quant One Step RT-PCR Kit For fast and sensitive one-step RT-PCR www.tiangen.com/en RT121221 Quant One Step RT-PCR Kit Kit Contents Cat. no. KR113 Contents Hotmaster Taq Polymerase (2.5 U/μl) Quant
Roche Molecular Biochemicals Technical Note No. LC 9/2000
 Roche Molecular Biochemicals Technical Note No. LC 9/2000 LightCycler Optimization Strategy Introduction Purpose of this Note Table of Contents The LightCycler system provides different detection formats
Roche Molecular Biochemicals Technical Note No. LC 9/2000 LightCycler Optimization Strategy Introduction Purpose of this Note Table of Contents The LightCycler system provides different detection formats
PrimeScript RT reagent Kit (Perfect Real Time)
 Cat. # RR037A For Research Use PrimeScript RT reagent Kit (Perfect Real Time) Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Storage... 3 IV. Features... 4 V. Precautions...
Cat. # RR037A For Research Use PrimeScript RT reagent Kit (Perfect Real Time) Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Storage... 3 IV. Features... 4 V. Precautions...
SUPPLEMENTARY MATERIAL AND METHODS
 SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
Supplemental Information. HEXIM1 and NEAT1 Long Non-coding RNA Form. a Multi-subunit Complex that Regulates. DNA-Mediated Innate Immune Response
 Molecular Cell, Volume 67 Supplemental Information HEXIM1 and NEAT1 Long Non-coding RNA Form a Multi-subunit Complex that Regulates DNA-Mediated Innate Immune Response Mehdi Morchikh, Alexandra Cribier,
Molecular Cell, Volume 67 Supplemental Information HEXIM1 and NEAT1 Long Non-coding RNA Form a Multi-subunit Complex that Regulates DNA-Mediated Innate Immune Response Mehdi Morchikh, Alexandra Cribier,
Methods of Biomaterials Testing Lesson 3-5. Biochemical Methods - Molecular Biology -
 Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
Methods of Biomaterials Testing Lesson 3-5 Biochemical Methods - Molecular Biology - Chromosomes in the Cell Nucleus DNA in the Chromosome Deoxyribonucleic Acid (DNA) DNA has double-helix structure The
The Polymerase Chain Reaction. Chapter 6: Background
 The Polymerase Chain Reaction Chapter 6: Background Invention of PCR Kary Mullis Mile marker 46.58 in April of 1983 Pulled off the road and outlined a way to conduct DNA replication in a tube Worked for
The Polymerase Chain Reaction Chapter 6: Background Invention of PCR Kary Mullis Mile marker 46.58 in April of 1983 Pulled off the road and outlined a way to conduct DNA replication in a tube Worked for
XactEdit Cas9 Nuclease with NLS User Manual
 XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
Supplementary Figure 1. RAD51 and RAD51 paralogs are enriched spontaneously onto
 Supplementary Figure legends Supplementary Figure 1. and paralogs are enriched spontaneously onto the S-phase chromatin during DN replication. () Chromatin fractionation was carried out as described in
Supplementary Figure legends Supplementary Figure 1. and paralogs are enriched spontaneously onto the S-phase chromatin during DN replication. () Chromatin fractionation was carried out as described in
Please purchase PDFcamp Printer on to remove this watermark. DNA microarray
 DNA microarray Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. A DNA microarray is a multiplex technology used in molecular biology. It consists of
DNA microarray Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. A DNA microarray is a multiplex technology used in molecular biology. It consists of
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
Supplementary Figures Montero et al._supplementary Figure 1
 Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
Montero et al_suppl. Info 1 Supplementary Figures Montero et al._supplementary Figure 1 Montero et al_suppl. Info 2 Supplementary Figure 1. Transcripts arising from the structurally conserved subtelomeres
Technical tips Session 5
 Technical tips Session 5 Chromatine Immunoprecipitation (ChIP): This is a powerful in vivo method to quantitate interaction of proteins associated with specific regions of the genome. It involves the immunoprecipitation
Technical tips Session 5 Chromatine Immunoprecipitation (ChIP): This is a powerful in vivo method to quantitate interaction of proteins associated with specific regions of the genome. It involves the immunoprecipitation
ab ChIP Kit Magnetic One-Step
 ab156907 ChIP Kit Magnetic One-Step Instructions for Use For selective enrichment of a chromatin fraction containing specific DNA sequences in a high throughput format using chromatin isolated from various
ab156907 ChIP Kit Magnetic One-Step Instructions for Use For selective enrichment of a chromatin fraction containing specific DNA sequences in a high throughput format using chromatin isolated from various
Chapter 9 Genetic Engineering
 Chapter 9 Genetic Engineering Biotechnology: use of microbes to make a protein product Recombinant DNA Technology: Insertion or modification of genes to produce desired proteins Genetic engineering: manipulation
Chapter 9 Genetic Engineering Biotechnology: use of microbes to make a protein product Recombinant DNA Technology: Insertion or modification of genes to produce desired proteins Genetic engineering: manipulation
Premix Ex Taq (Probe qpcr)
 For Research Use Premix Ex Taq (Probe qpcr) Product Manual Table of Contents I. Description... 3 II. Principle... 4 III. Components... 5 IV. Materials Required but not Provided... 5 V. Storage... 5 VI.
For Research Use Premix Ex Taq (Probe qpcr) Product Manual Table of Contents I. Description... 3 II. Principle... 4 III. Components... 5 IV. Materials Required but not Provided... 5 V. Storage... 5 VI.
Supplemental Information. Boundary Formation through a Direct. Threshold-Based Readout. of Mobile Small RNA Gradients
 Developmental Cell, Volume 43 Supplemental Information Boundary Formation through a Direct Threshold-Based Readout of Mobile Small RNA Gradients Damianos S. Skopelitis, Anna H. Benkovics, Aman Y. Husbands,
Developmental Cell, Volume 43 Supplemental Information Boundary Formation through a Direct Threshold-Based Readout of Mobile Small RNA Gradients Damianos S. Skopelitis, Anna H. Benkovics, Aman Y. Husbands,
PrimePCR Assay Validation Report
 Gene Information Gene Name SRY (sex determining region Y)-box 6 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID SOX6 Human This gene encodes a member of the
Gene Information Gene Name SRY (sex determining region Y)-box 6 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID SOX6 Human This gene encodes a member of the
Table of Contents. PrimeScript TM RT-PCR Kit. I. Kit Contents...2. Storage...3. Principle...4. Features...5. V. Notes...5. Protocol...
 Table of Contents I. Kit Contents...2 II. III. IV. Storage...3 Principle...4 Features...5 V. Notes...5 VI. Protocol...6 VII. PCR Condition...8 VIII. Application...8 IX. Preparation of RNA sample...10 X.
Table of Contents I. Kit Contents...2 II. III. IV. Storage...3 Principle...4 Features...5 V. Notes...5 VI. Protocol...6 VII. PCR Condition...8 VIII. Application...8 IX. Preparation of RNA sample...10 X.
SideStep Lysis and Stabilization Buffer
 SideStep Lysis and Stabilization Buffer INSTRUCTION MANUAL Catalog #400900 Revision B.0 For Research Use Only. Not for use in diagnostic procedures. 400900-12 LIMITED PRODUCT WARRANTY This warranty limits
SideStep Lysis and Stabilization Buffer INSTRUCTION MANUAL Catalog #400900 Revision B.0 For Research Use Only. Not for use in diagnostic procedures. 400900-12 LIMITED PRODUCT WARRANTY This warranty limits
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Description of the observed lymphatic metastases in two different SIX1-induced MCF7 metastasis models (Nude and NOD/SCID). Supplementary Figure 2. MCF7-SIX1
Supplementary Figures Supplementary Figure 1. Description of the observed lymphatic metastases in two different SIX1-induced MCF7 metastasis models (Nude and NOD/SCID). Supplementary Figure 2. MCF7-SIX1
Table S1. Primer sequences
 Table S1. Primer sequences Primers for quantitative PCR Tgf 1 Forward Tgf 1 Reverse Tgf 2Forward Tgf 2Reverse Tgf 3 Forward Tgf 3 Reverse Tgf r1 Forward Tgf r1 Reverse Tgf r2 Forward Tgf r2 Reverse Thbs1
Table S1. Primer sequences Primers for quantitative PCR Tgf 1 Forward Tgf 1 Reverse Tgf 2Forward Tgf 2Reverse Tgf 3 Forward Tgf 3 Reverse Tgf r1 Forward Tgf r1 Reverse Tgf r2 Forward Tgf r2 Reverse Thbs1
