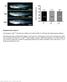
Supplemental Figure 1. Immunoblot analysis of wild-type and mutant forms of JAZ3
|
|
|
- Damon Bell
- 6 years ago
- Views:
Transcription
1 Supplemental Data. Chung and Howe (2009). A Critical Role for the TIFY Motif in Repression of Jasmonate Signaling by a Stabilized Splice Variant of the JASMONATE ZIM-domain protein JAZ10 in Arabidopsis. Supplemental Figure 1. Immunoblot analysis of wild-type and mutant forms of JAZ3 and JAZ10.1. Protein was extracted from yeast strains used for the yeast two-hybrid assays shown in Figures 2E and F. Expression of wild-type (WT) JAZ3-AD and JAZ10.1- AD fusion proteins, as well as mutant derivatives harboring the indicated point mutation in the TIFY motif, was detected with an anti-ha antibody. 1
2 Supplemental Figure 2. Yeast two-hybrid analysis of homo- and heteromeric interactions between Arabidopsis ZIM, ZIM-like1 (ZML1), and ZIM-like2 (ZML2) proteins. Yeast strains co-transformed with empty or ZIM-, ZML1-, and ZML2-containing BD (pgilda) and AD (pb42ad) vectors were plated on media containing X-gal. The photograph was taken after 48 h of incubation of the plate at 30 C. Blue color formation is indicative of a positive protein-protein interaction. 2
3 Supplemental Figure 3. Detection of transcripts encoding JAZ10.3 and JAZ10.4 using transcript-specific primers. (A) Schematic diagram of transcripts encoding JAZ10.3 and JAZ10.4. Left-pointing arrows denote the location of oligonucleotide primers that are specific for each transcript. Primer sequences are listed in Supplemental Table 1. The gene-specific primer for JAZ10.4 hybridizes to a sequence created from splicing of the alternate donor site in exon 3 to the acceptor site in exon 4. (B) Result of an RT-PCR experiment using total RNA isolated from wounded (6 hr post wounding) Arabidopsis leaves and the transcript-specific primers indicated in (A). PCR products were visualized by agarose gel electrophoresis and ethidium bromide staining. 3
4 Supplemental Figure 4. Yeast two-hybrid analysis of the effect of I107A and G111A TIFY mutations on the interaction of JAZ10.4 with other members of the JAZ family. Yeast strains were co-transformed with plasmids encoding an AD-fusion of JAZ10.4 (or the JAZ10.4 I->A and JAZ10.4 G->A mutants; left column) and a BD-fusion of one of the indicated (top row) twelve JAZ family members. Co-transformed strains were plated on medium containing X-gal. The photograph was taken after 48 h of incubation of the plate at 30 C. Blue color formation is indicative of a positive protein-protein interaction. 4
5 Supplemental Table 1. List of constructs and primers used in this study. Clone Name(s) Forward Primers Reverse Primers Vector Cloning site Y2H analysis AD-JAZ1 and BD-JAZ1 a pb42ad or pgilda AD-JAZ2 and BD-JAZ2 5'-GAATTCATGTCGAGTTTTTCTGCCGAGTGT-3' 5'-CTCGAGTTACCGTGAACTGAGCCAAGCTGG-3' pb42ad or pgilda EcoR1-Xho1 AD-JAZ3 and BD-JAZ3 5'-GAATTCATGGAGAGAGATTTCCTCGGGTTG-3' 5'-GAATTCGGTTGCAGAGCTGAGAGAAGAACT-3' pb42ad or pgilda EcoR1-Xho1 AD-JAZ3ΔN 5'-GAATTCTTGGTAGGAGTTCCTATCATG-3' 5'-GAATTCGGTTGCAGAGCTGAGAGAAGAACT-3' pb42ad or pgilda EcoR1-Xho1 AD-JAZ3Jas 5'-GGATTCATGCCTCAAGTCTTTTCG-3' 5'-GAATTCGGTTGCAGAGCTGAGAGAAGAACT-3' pb42ad or pgilda EcoR1-Xho1 AD-JAZ3ΔJas 5'-GAATTCTTGGTAGGAGTTCCTATCATG-3' 5'-CTCGAGCTAAGCCATTACATTGGTAGA-3' pb42ad AD-JAZ3 T180A 5'-TCA CCT GCG CAG TTG GCG ATC TTT TAT GCC-3' b 5'-GGCATAAAAGATCGCCAACTGCGCAGGTGA-3' b pb42ad AD-JAZ3 I181A 5'-CCTGCGCAGTTGACGGCCTTTTATGCCGGTTC-3' b 5'-GAACCGGCATAAAAGGCCGTCAACTGCGCAGG-3' b pb42ad AD-JAZ3 F182A 5'-GCGCAGTTGACGATCGCTTATGCCGGTTCAGT-3' b 5'-ACTGAACCGGCATAAGCGATCGTCAACTGCGC-3' b pb42ad AD-JAZ3 Y183A 5'-CAGTTGACGATCTTTGCTGCCGGTTCAGTTTG-3' b 5'-CAAACTGAACCGGCAGCAAAGATCGTCAACTG-3' b pb42ad AD-JAZ3 G185A 5'-ACGATCTTTTATGCCGCTTCAGTTTGTGTTTA-3' b 5'-TAAACACAAACTGAAGCGGCATAAAAGATCGT-3' b pb42ad AD-JAZ4 and BD-JAZ4 5'-GAATTCATGGAGAGAGATTTTCTCGGGCTG-3' 5'-GAATTCTCATGTCACAATGGGGCTGGTTTC-3' pb42ad or pgilda EcoR1-EcoR1 AD-JAZ5 and BD-JAZ5 5'-GAATTCATGTCGTCGAGCAATGAAAATGCT-3' 5'-CTCGAGCTATAGCCTTAGATCGAGATCTTT-3' pb42ad or pgilda EcoR1-Xho1 AD-JAZ6 and BD-JAZ6 5'-GAATTCATGTCAACGGGACAAGCGCCGGAG-3' 5'-CTCGAGCTAAAGCTTGAGTTCAAGGTTTTT-3' pb42ad or pgilda EcoR1-Xho1 AD-JAZ7 and BD-JAZ7 5'-GAATTCATGATCATCATCATCAAAAACTGC-3' 5'-CTCGAGCTATCGGTAACGGTGGTAAGGGGA-3' pb42ad or pgilda EcoR1-EcoR1 AD-JAZ8 and BD-JAZ8 5'-CTCGAGATGAAGCTACAGCAAAATTGTGAC-3' 5'-CTCGAGTTATCGTCGTGAATGGTACGGTGA-3' pb42ad or pgilda Xho1-Xho1 AD-JAZ9 and BD-JAZ9 a pb42ad or pgilda AD-JAZ10.1 T106A 5'-GGAACTGTTCCTATGGCGATTTTCTACAATG-3' b 5'-CATTGTAGAAAATCGCCATAGGAACAGTTC-3' b pb42ad AD-JAZ10.1 I107A 5'-ACTGTTCCTATGACGGCTTTCTACAATGGAAG-3' b 5'-CTTCCATTGTAGAAGCCGTCATAGGAACAGT-3' b pb42ad AD-JAZ10.1 F108A 5'-GTTCCTATGACGATTGCCTACAATGGAAGTGT-3' b 5'-ACACTTCCATTGTAGGCAATCGTCATAGGAAC-3' b pb42ad AD-JAZ10.1 Y109A 5'-CCTATGACGATTTTCGCCAATGGAAGTGTTT-3' b 5'-AAACACTTCCATTGGCGAAAATCGTCATAGG-3' b pb42ad AD-JAZ10.1 G111A 5'-ACGATTTTCTACAATGCAAGTGTTTCAGTTTT-3' b 5'-AAAACTGAAACACTTGCATTGTAGAAAATCGT-3' b pb42ad AD-JAZ10.3 and BD-JAZ10.3 5'-GAATTCATGTCGAAAGCTACCATAGAACTC-3' 5'-GAATTCTTACTCTCCTTGCGCTTCTCGAG-3' pb42ad or pgilda EcoR1-EcoR1 AD-JAZ10.4 and BD-JAZ10.4 5'-GAATTCATGTCGAAAGCTACCATAGAACTC-3' 5'-GAATTCCTAATCTCTCCTTGCGCTTCTTGA-3' pb42ad or / pgilda EcoR1-EcoR1 AD-JAZ10.4 I->A 5'-ACTGTTCCTATGACGGCTTTCTACAATGGAAG-3' b 5'-CTTCCATTGTAGAAGCCGTCATAGGAACAGT-3' b pb42ad AD-JAZ10.4 G->A 5'-GTTCCTATGACGATTGCCTACAATGGAAGTGT-3' b 5'-ACACTTCCATTGTAGGCAATCGTCATAGGAAC-3' b pb42ad AD-JAZ11 and BD-JAZ11 5'-GAATTCATGGATTGGTCATTCTCAAGCAAA-3' 5'-GAATTCTTAGTGCAGATGATGAGACTGGAGG-3' pb42ad or pgilda EcoR1-EcoR1 AD-JAZ12 and BD-JAZ12 5'-CCCGGGAATGACTAAGGTGAAGATGA-3' 5'-CCCGGGCTAAGCAGTTGGAAATTCCTC-3' pb42ad or pgilda Sma1-Sma1 AD-ZIM and BD-ZIM 5'-GAATTCATGTTGGTCGCCATTCGATT-3' 5'-CTCGAGTTAGTGATCACCTAACAGAT-3' pb42ad or pgilda EcoR1-Xho1 AD-ZML1 and BD-ZML1 5'-GAATTCATGGATGATCTTCATGGAAG-3' 5'-CTCGAGTCACTGTGTGTTGCTAATGTC-3' pb42ad or pgilda EcoR1-Xho1 AD-ZML2 and BDZML2 5'-GAATTCATGGATGACCTACATGGAAGC-3' 5'-CTCGAGTCACTGTGAGTTGCTTATGTC-3' pb42ad or pgilda EcoR1-Xho1 BD-COI1 c pgilda Stable transgenic lines 35S-JAZ10.1 5'-GGTACCATGTCGAAAGCTACCATAGAACTC-3' 5'-GAGCTCTTAGGCCGATGTCGGATAGTAAGG-3' pbi-ts Kpn1-Sac1 35S-JAZ10.3 5'-GGTACCATGTCGAAAGCTACCATAGAACTC-3' 5'-GAGCTCTTACTCTCCTTGCGCTTCTCGAG-3' pbi-ts Kpn1-Sac1 35S-JAZ10.4 5'-GGTACCATGTCGAAAGCTACCATAGAACTC-3' 5'-GAGCTCCTAATCTCTCCTTGCGCTTCTTGA-3' pbi-ts Kpn1-Sac1 35S-JAZ10.4 I->A 5'-ACTGTTCCTATGACGGCTTTCTACAATGGAAG-3' b 5'-CTTCCATTGTAGAAGCCGTCATAGGAACAGT-3' b pbi-ts Kpn1-Sac1 35S-JAZ10.4 G->A 5'-ACGATTTTCTACAATGCAAGTGTTTCAGTTTT-3' b 5'-AAAACTGAAACACTTGCATTGTAGAAAATCGT-3' b pbi-ts Kpn1-Sac1 35S-JAZ10.4-GUS 5'-ACGATTTTCTACAATGCAAGTGTTTCAGTTTT-3' 5'-AAAACTGAAACACTTGCATTGTAGAAAATCGT-3' pbi121 BamH1/ BamH1 35S-JAZ10.4 I->A -GUS 5'-ACTGTTCCTATGACGGCTTTCTACAATGGAAG-3' b 5'-CTTCCATTGTAGAAGCCGTCATAGGAACAGT-3' b pbi121 BamH1/ BamH1 35S-JAZ10.1-eYFP 5'-GGATCCATGTCGAAAGCTACCATAGAA-3' 5'-GGATCCGGCCGATGTCGGATAGTAAGG-3' pbi-eyfp BamH1-BamH1 35S-JAZ10.3-eYFP 5'-GGATCCATGTCGAAAGCTACCATAGAA-3' 5'-GGATCCCCTCTCCTTGCGCTTCTCGAG-3' pbi-eyfp BamH1-BamH1 35S-JAZ10.4-eYFP 5'-GGATCCATGTCGAAAGCTACCATAGAA-3' 5'-GGATCCATCTCTCCTTGCGCTTCTTGA-3' pbi-eyfp BamH1-BamH1 Transient expression 35S-eYFP 5'-GGTACCATGGTGAGCAAGGGCGAGGAG-3' 5'-GAGCTCTCACTTGTACAGCTCGTCCA-3' pbi-ts Kpn1-Sac1 35S-JAZ10.1-eYFP 5'-GGATCCATGTCGAAAGCTACCATAGAA-3' 5'-GGATCCGGCCGATGTCGGATAGTAAGG-3' pbi-eyfp BamH1-BamH1 35S-JAZ10.3-eYFP 5'-GGATCCATGTCGAAAGCTACCATAGAA-3' 5'-GGATCCCCTCTCCTTGCGCTTCTCGAG-3' pbi-eyfp BamH1-BamH1 35S-JAZ10.4-eYFP 5'-GGATCCATGTCGAAAGCTACCATAGAA-3' 5'-GGATCCATCTCTCCTTGCGCTTCTTGA-3' pbi-eyfp BamH1-BamH1 35S-JAZ10.4 I->A -eyfp 5'-ACTGTTCCTATGACGGCTTTCTACAATGGAAG-3' b 5'-CTTCCATTGTAGAAGCCGTCATAGGAACAGT-3' b pbi-eyfp BamH1-BamH1 JAZ3-nYFP 5'-GTCGACAATGGAGAGAGATTTTCTC-3' 5'-GCGGCCGCGTGGTTGCAGAGCTGAGAGA-3' psy728 Sal1-Not 1 JAZ3-cYFP 5'-GTCGACAATGGAGAGAGATTTTCTC-3' 5'-GCGGCCGCGTGGTTGCAGAGCTGAGAGA-3' psy738 Sal1-Not 1 nyfp-jaz3 5'-GTCGACAATGGAGAGAGATTTTCTC-3' 5'-GAGCTCTTAGGTTGCAGAGCTGAG-3' psy735 Sal1-Sac1 cyfp-jaz3 5'-GTCGACAATGGAGAGAGATTTTCTC-3' 5'-GAGCTCTTAGGTTGCAGAGCTGAG-3' psy736 Sal1-Sac1 Transcript-specific RT-PCR JAZ10.3 5'-GGTACCATGTCGAAAGCTACCATAGAACTC-3' 5'-GGATTGTTGAAGAATCATTAC-3' JAZ10.4 5'-GGTACCATGTCGAAAGCTACCATAGAACTC-3' 5'-ATGGGAAGATCGAAAGATCT-3' a Described in Melotto et al (2008) Plant J 55: b Primers for site-directed mutagensis c Described in Thines et al (2007) Nature 488:
S156AT168AY175A (AAA) were purified as GST-fusion proteins and incubated with GSTfused
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 Supplemental Materials Supplemental Figure S1 (a) Phenotype of the wild type and grik1-2 grik2-1 plants after 8 days in darkness.
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 Supplemental Materials Supplemental Figure S1 (a) Phenotype of the wild type and grik1-2 grik2-1 plants after 8 days in darkness.
SUPPLEMENTAL DATA. Supplementary data
 1 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 SUPPLEMENTAL DATA Supplementary data Figure S1: Verification of pla-iα1 knockdown lines a: Semiquantitative
1 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 SUPPLEMENTAL DATA Supplementary data Figure S1: Verification of pla-iα1 knockdown lines a: Semiquantitative
Supplemental Data. Cui et al. (2012). Plant Cell /tpc a b c d. Stem UBC32 ACTIN
 A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
Supplementary Figures 1-12
 Supplementary Figures 1-12 Supplementary Figure 1. The specificity of anti-abi1 antibody. Total Proteins extracted from the wild type seedlings or abi1-3 null mutant seedlings were used for immunoblotting
Supplementary Figures 1-12 Supplementary Figure 1. The specificity of anti-abi1 antibody. Total Proteins extracted from the wild type seedlings or abi1-3 null mutant seedlings were used for immunoblotting
Department of Energy Plant Research Laboratory, Michigan State University, East Lansing, Michigan b
 The Plant Cell, Vol. 21: 131 145, January 2009, www.plantcell.org ã 2009 American Society of Plant Biologists A Critical Role for the TIFY Motif in Repression of Jasmonate Signaling by a Stabilized Splice
The Plant Cell, Vol. 21: 131 145, January 2009, www.plantcell.org ã 2009 American Society of Plant Biologists A Critical Role for the TIFY Motif in Repression of Jasmonate Signaling by a Stabilized Splice
Supplemental Data. Dai et al. (2013). Plant Cell /tpc Absolute FyPP3. Absolute
 A FyPP1 Absolute B FyPP3 Absolute Dry seeds Imbibed 24 hours Dry seeds Imbibed 24 hours C ABI5 Absolute Dry seeds Imbibed 24 hours Supplemental Figure 1. Expression of FyPP1, FyPP3 and ABI5 during seed
A FyPP1 Absolute B FyPP3 Absolute Dry seeds Imbibed 24 hours Dry seeds Imbibed 24 hours C ABI5 Absolute Dry seeds Imbibed 24 hours Supplemental Figure 1. Expression of FyPP1, FyPP3 and ABI5 during seed
- 1 - Supplemental Data
 - 1-1 Supplemental Data 2 3 4 5 6 7 8 9 Supplemental Figure S1. Differential expression of AtPIP Genes in DC3000-inoculated plants. Gene expression in leaves was analyzed by real-time RT-PCR and expression
- 1-1 Supplemental Data 2 3 4 5 6 7 8 9 Supplemental Figure S1. Differential expression of AtPIP Genes in DC3000-inoculated plants. Gene expression in leaves was analyzed by real-time RT-PCR and expression
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1 shrna mediated knockdown of ZRSR2 in K562 and 293T cells. (a) ZRSR2 transcript levels in stably transduced K562 cells were determined using qrt-pcr. GAPDH was
Supplementary Figure 1 Supplementary Fig. 1 shrna mediated knockdown of ZRSR2 in K562 and 293T cells. (a) ZRSR2 transcript levels in stably transduced K562 cells were determined using qrt-pcr. GAPDH was
Figure S1. DELLA Proteins Act as Positive Regulators to Mediate GA-Regulated Anthocyanin
 Supplemental Information Figure S1. DELLA Proteins Act as Positive Regulators to Mediate GA-Regulated Anthocyanin Biosynthesis. (A) Effect of GA on anthocyanin content in WT and ga1-3 seedlings. Mock,
Supplemental Information Figure S1. DELLA Proteins Act as Positive Regulators to Mediate GA-Regulated Anthocyanin Biosynthesis. (A) Effect of GA on anthocyanin content in WT and ga1-3 seedlings. Mock,
Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana.
 SUPPLEMENTARY FIGURES Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana. YFP:, :CFP, HA: and (A) and HA, CFP and YFPtagged and AVR1-CO39 (B) were expressed
SUPPLEMENTARY FIGURES Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana. YFP:, :CFP, HA: and (A) and HA, CFP and YFPtagged and AVR1-CO39 (B) were expressed
Yeast 2-Hybrid Kayla Nygaard
 Yeast 2-Hybrid 2.26.18 Kayla Nygaard Y2H - What is it? A method to screen for protein-protein interactions in yeast Capitalizes on GAL4 system in yeast. GAL4 has 2 domains DNA-Binding Domain (DB) Transcriptional
Yeast 2-Hybrid 2.26.18 Kayla Nygaard Y2H - What is it? A method to screen for protein-protein interactions in yeast Capitalizes on GAL4 system in yeast. GAL4 has 2 domains DNA-Binding Domain (DB) Transcriptional
Supplemental Data. Wu et al. (2). Plant Cell..5/tpc RGLG Hormonal treatment H2O B RGLG µm ABA µm ACC µm GA Time (hours) µm µm MJ µm IA
 Supplemental Data. Wu et al. (2). Plant Cell..5/tpc..4. A B Supplemental Figure. Immunoblot analysis verifies the expression of the AD-PP2C and BD-RGLG proteins in the Y2H assay. Total proteins were extracted
Supplemental Data. Wu et al. (2). Plant Cell..5/tpc..4. A B Supplemental Figure. Immunoblot analysis verifies the expression of the AD-PP2C and BD-RGLG proteins in the Y2H assay. Total proteins were extracted
Enhancers mutations that make the original mutant phenotype more extreme. Suppressors mutations that make the original mutant phenotype less extreme
 Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
SUPPLEMENTARY INFORMATION
 Secondary mutations as a mechanism of cisplatin resistance in BRCA2-mutated cancers Wataru Sakai, Elizabeth M. Swisher, Beth Y. Karlan, Mukesh K. Agarwal, Jake Higgins, Cynthia Friedman, Emily Villegas,
Secondary mutations as a mechanism of cisplatin resistance in BRCA2-mutated cancers Wataru Sakai, Elizabeth M. Swisher, Beth Y. Karlan, Mukesh K. Agarwal, Jake Higgins, Cynthia Friedman, Emily Villegas,
AD-FIL AD-YAB2 -2 BD-JAZ3 AD-YAB3 AD-YAB5 AD-FIL -4 3AT BD-JAZ3 AD-YAB3
 3 4 9 10 11 12 AD-FIL AD-YAB5 2 B AD-YAB3 1 BD-JAZ 5 6 7 8 AD-FIL A AD-YAB2 Supplemental Data. Boter et al. (2015). Plant Cell 10.1105/tpc.15.00220 BD AD-YAB2-2 -2 BD-JAZ3 AD-YAB3 BD AD-YAB5-4 AD-FIL -4
3 4 9 10 11 12 AD-FIL AD-YAB5 2 B AD-YAB3 1 BD-JAZ 5 6 7 8 AD-FIL A AD-YAB2 Supplemental Data. Boter et al. (2015). Plant Cell 10.1105/tpc.15.00220 BD AD-YAB2-2 -2 BD-JAZ3 AD-YAB3 BD AD-YAB5-4 AD-FIL -4
Supplemental Data. Seo et al. (2014). Plant Cell /tpc
 Supplemental Figure 1. Protein alignment of ABD1 from other model organisms. The alignment was performed with H. sapiens DCAF8, M. musculus DCAF8 and O. sativa Os10g0544500. The WD40 domains are underlined.
Supplemental Figure 1. Protein alignment of ABD1 from other model organisms. The alignment was performed with H. sapiens DCAF8, M. musculus DCAF8 and O. sativa Os10g0544500. The WD40 domains are underlined.
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplemental Data. Zhang et al. (2010). Plant Cell /tpc
 Supplemental Figure 1. uvs90 gene cloning The T-DNA insertion in uvs90 was identified using thermal asymmetric interlaced (TAIL)-PCR. Three rounds of amplification were performed; the second (2 nd ) and
Supplemental Figure 1. uvs90 gene cloning The T-DNA insertion in uvs90 was identified using thermal asymmetric interlaced (TAIL)-PCR. Three rounds of amplification were performed; the second (2 nd ) and
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Supplementary Methods
 Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplemental Data. Farmer et al. (2010) Plant Cell /tpc
 Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
AD BD TOC1. Supplementary Figure 1: Yeast two-hybrid assays showing the interaction between
 AD X BD TOC1 AD BD X PIFΔAD PIF TOC1 TOC1 PIFΔAD PIF N TOC1 TOC1 C1 PIFΔAD PIF C1 TOC1 TOC1 C PIFΔAD PIF C TOC1 Supplementary Figure 1: Yeast two-hybrid assays showing the interaction between PIF and TOC1
AD X BD TOC1 AD BD X PIFΔAD PIF TOC1 TOC1 PIFΔAD PIF N TOC1 TOC1 C1 PIFΔAD PIF C1 TOC1 TOC1 C PIFΔAD PIF C TOC1 Supplementary Figure 1: Yeast two-hybrid assays showing the interaction between PIF and TOC1
Supplemental Data. Furlan et al. Plant Cell (2017) /tpc
 Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Figure.
Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Figure.
Electronic Supplementary Materials
 Electronic Supplementary Materials Supplementary Figure S1. Modelling of the coiled-coil based interaction between Mis12 and Nnf1a. 36 alternative structural alignments of Mis12 and Nnf1a proteins on the
Electronic Supplementary Materials Supplementary Figure S1. Modelling of the coiled-coil based interaction between Mis12 and Nnf1a. 36 alternative structural alignments of Mis12 and Nnf1a proteins on the
GTTCGGGTTCC TTTTGAGCAG
 Supplementary Figures Splice variants of the SIP1 transcripts play a role in nodule organogenesis in Lotus japonicus. Wang C, Zhu H, Jin L, Chen T, Wang L, Kang H, Hong Z, Zhang Z. 5 UTR CDS 3 UTR TCTCAACCATCCTTTGTCTGCTTCCGCCGCATGGGTGAGGTCATTTTGTCTAGATGACGTGCAATTTACAATGA
Supplementary Figures Splice variants of the SIP1 transcripts play a role in nodule organogenesis in Lotus japonicus. Wang C, Zhu H, Jin L, Chen T, Wang L, Kang H, Hong Z, Zhang Z. 5 UTR CDS 3 UTR TCTCAACCATCCTTTGTCTGCTTCCGCCGCATGGGTGAGGTCATTTTGTCTAGATGACGTGCAATTTACAATGA
SUPPLEMENTARY INFORMATION
 AS-NMD modulates FLM-dependent thermosensory flowering response in Arabidopsis NATURE PLANTS www.nature.com/natureplants 1 Supplementary Figure 1. Genomic sequence of FLM along with the splice sites. Sequencing
AS-NMD modulates FLM-dependent thermosensory flowering response in Arabidopsis NATURE PLANTS www.nature.com/natureplants 1 Supplementary Figure 1. Genomic sequence of FLM along with the splice sites. Sequencing
Supplemental Figure 1 A
 Supplemental Figure 1 A Supplemental Data. Han et al. (2016). Plant Cell 10.1105/tpc.15.00997 BamHI -1616 bp SINE repeats SalI ATG SacI +1653 bp HindIII LB HYG p35s pfwa LUC Tnos RB pfwa-bamhi-f: CGGGATCCCGCCTTTCTCTTCCTCATCTGC
Supplemental Figure 1 A Supplemental Data. Han et al. (2016). Plant Cell 10.1105/tpc.15.00997 BamHI -1616 bp SINE repeats SalI ATG SacI +1653 bp HindIII LB HYG p35s pfwa LUC Tnos RB pfwa-bamhi-f: CGGGATCCCGCCTTTCTCTTCCTCATCTGC
Supplemental data. Zhao et al. (2009). The Wuschel-related homeobox gene WOX11 is required to activate shoot-borne crown root development in rice.
 Supplemental data. Zhao et al. (2009). The Wuschel-related homeobox gene WOX11 is required to activate shoot-borne crown root development in rice. A B Supplemental Figure 1. Expression of WOX11p-GUS WOX11-GFP
Supplemental data. Zhao et al. (2009). The Wuschel-related homeobox gene WOX11 is required to activate shoot-borne crown root development in rice. A B Supplemental Figure 1. Expression of WOX11p-GUS WOX11-GFP
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
Supplemental Data. Jing et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. Characterization of epp1 Mutants. (A) Cotyledon angles of 5-d-old Col wild-type (gray bars) and epp1-1 (black bars) seedlings under red (R), far-red (FR) and blue (BL) light conditions,
Supplemental Figure 1. Characterization of epp1 Mutants. (A) Cotyledon angles of 5-d-old Col wild-type (gray bars) and epp1-1 (black bars) seedlings under red (R), far-red (FR) and blue (BL) light conditions,
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity.
 Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
Notes of Protein Engineering
 Notes of Protein Engineering Week 2 (01/ 08/ 2016-07/ 08/ 2016) Aug 5th 1. PCR Verify that the E.coli transformed with prs416 linked with parts is well constructed. Fast Taq Polymerase 50μl Protocol; Strategy:
Notes of Protein Engineering Week 2 (01/ 08/ 2016-07/ 08/ 2016) Aug 5th 1. PCR Verify that the E.coli transformed with prs416 linked with parts is well constructed. Fast Taq Polymerase 50μl Protocol; Strategy:
Supplemental Data. Shyu et al. (2012). Plant Cell /tpc
 JAZ11 JAZ12 JAZ6 JAZ5 JAZ10 JAZ9 JAZ2 JAZ1 JAZ3 JAZ4 JAZ7 Supplemental Figure 1. Phylogenetic Tree Constructed from the Jas Motif of Arabidopsis JAZ Proteins. The 27-amino-acid Jas motif in all 12 Arabidopsis
JAZ11 JAZ12 JAZ6 JAZ5 JAZ10 JAZ9 JAZ2 JAZ1 JAZ3 JAZ4 JAZ7 Supplemental Figure 1. Phylogenetic Tree Constructed from the Jas Motif of Arabidopsis JAZ Proteins. The 27-amino-acid Jas motif in all 12 Arabidopsis
ATL1-GFP Marker-mCherry/RFP Merge B C D
 Input IP:α-H EDR1-H stedr1-h EV EDR1-H stedr1-h EV α-h α-mcherry TL1-GFP Marker-mCherry/RFP Merge C D GmMan49-mCherry mcherry-syp21 VH-a1-RFP E F G H I J EDR1-nYFP+TL1-cYFP ra6-mcherry Merge K L M Supplemental
Input IP:α-H EDR1-H stedr1-h EV EDR1-H stedr1-h EV α-h α-mcherry TL1-GFP Marker-mCherry/RFP Merge C D GmMan49-mCherry mcherry-syp21 VH-a1-RFP E F G H I J EDR1-nYFP+TL1-cYFP ra6-mcherry Merge K L M Supplemental
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Supplemental Data. Zhang et al. (2013). Plant Cell /tpc
 SDLTXgal SDLTH SDLTHA ADSV4 BD BDP53 BD BDP53 BD BDP53 ADAt1g1567(aa36358) ADAt1g844(aa9354) ADAt1g844(aa51354) ADAt1g844(full) Prey ADAt1g844(aa11354) ADAt3g5994(aa48418) BD BDPAL2 BD BDPAL2 BD BDPAL2
SDLTXgal SDLTH SDLTHA ADSV4 BD BDP53 BD BDP53 BD BDP53 ADAt1g1567(aa36358) ADAt1g844(aa9354) ADAt1g844(aa51354) ADAt1g844(full) Prey ADAt1g844(aa11354) ADAt3g5994(aa48418) BD BDPAL2 BD BDPAL2 BD BDPAL2
PCR analysis was performed to show the presence and the integrity of the var1csa and var-
 Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplemental Data. Li et al. (2015). Plant Cell /tpc
 Supplemental Data Supplemental Figure 1: Characterization of asr3 T-DNA knockout lines and complementation transgenic lines. (A) The scheme of At2G33550 (ASR3) with gray boxes indicating exons and dash
Supplemental Data Supplemental Figure 1: Characterization of asr3 T-DNA knockout lines and complementation transgenic lines. (A) The scheme of At2G33550 (ASR3) with gray boxes indicating exons and dash
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
Supplemental materials
 Supplemental materials Materials and methods for supplemental figures Yeast two-hybrid assays TAP46-PP2Ac interactions I. The TAP46 was used as the bait and the full-length cdnas of the five C subunits
Supplemental materials Materials and methods for supplemental figures Yeast two-hybrid assays TAP46-PP2Ac interactions I. The TAP46 was used as the bait and the full-length cdnas of the five C subunits
Supplemental Figure 1. Alignment of the NbGAPC amino acid sequences with their Arabidopsis homologues.
 Supplemental Figure 1. Alignment of the NbGAPC amino acid sequences with their Arabidopsis homologues. Homologs from N. benthamiana (NbGAPC1, NbGAPC2, NbGAPC3), Arabidopsis (AtGAPC1, AT3G04120; AtGAPC2,
Supplemental Figure 1. Alignment of the NbGAPC amino acid sequences with their Arabidopsis homologues. Homologs from N. benthamiana (NbGAPC1, NbGAPC2, NbGAPC3), Arabidopsis (AtGAPC1, AT3G04120; AtGAPC2,
Supplementary. Table 1: Oligonucleotides and Plasmids. complementary to positions from 77 of the SRα '- GCT CTA GAG AAC TTG AAG TAC AGA CTG C
 Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Figure S1. Figure S2 RT-PCR. qpcr RT-PCR. Northern. IVSwt ΔIVS IVS IVS IVS. NTC mock IVSwt ΔIVS IVS IVS. mock IVSwt ΔIVS
 Figure S1 40 cycles IVS IVS IVS NTC wt Δ Δ mut pri-mirna163 * Fig. S1 Transcripts generated from the MIR163 gene variants in which splice sites have been mutated are not spliced. products were separated
Figure S1 40 cycles IVS IVS IVS NTC wt Δ Δ mut pri-mirna163 * Fig. S1 Transcripts generated from the MIR163 gene variants in which splice sites have been mutated are not spliced. products were separated
Supplemental Data Supplemental Figure 1.
 Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis
 1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
Supplemental Data. Guo et al. (2015). Plant Cell /tpc
 Supplemental Figure 1. The Mutant exb1-d Displayed Pleiotropic Phenotypes and Produced Branches in the Axils of Cotyledons. (A) Branches were developed in exb1-d but not in wild-type plants. (B) and (C)
Supplemental Figure 1. The Mutant exb1-d Displayed Pleiotropic Phenotypes and Produced Branches in the Axils of Cotyledons. (A) Branches were developed in exb1-d but not in wild-type plants. (B) and (C)
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr.
 MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
MIT Department of Biology 7.01: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel iv) Would Xba I be useful for cloning? Why or why not?
Supplementary information, Figure S1
 Supplementary information, Figure S1 (A) Schematic diagram of the sgrna and hspcas9 expression cassettes in a single binary vector designed for Agrobacterium-mediated stable transformation of Arabidopsis
Supplementary information, Figure S1 (A) Schematic diagram of the sgrna and hspcas9 expression cassettes in a single binary vector designed for Agrobacterium-mediated stable transformation of Arabidopsis
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Figure S2. Verification of ihog and boi
 Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
Supplemental Figure 1
 Supplemental Figure 1 A gta2-1 gta2-2 1kb AT4G08350 B Col-0 gta2-1 LP+RP+LBb1.3 C Col-0 gta2-2 LP+RP+LB1 D Col-0 gta2-2 gta2-1 GTA2 TUBLIN E Col0 gta2-1 gta2-2 Supplemental Figure 1. Phenotypic analysis
Supplemental Figure 1 A gta2-1 gta2-2 1kb AT4G08350 B Col-0 gta2-1 LP+RP+LBb1.3 C Col-0 gta2-2 LP+RP+LB1 D Col-0 gta2-2 gta2-1 GTA2 TUBLIN E Col0 gta2-1 gta2-2 Supplemental Figure 1. Phenotypic analysis
scrm-d/+ scrm-d Supplemental Fig. 1.
 Kanaoka, Pillitteri, Fujii, Yoshida, Bogenschutz, Takabayashi, Zhu, and Torii (2008) SCRM/ICE1 and SCRM2 specify three cell-state transitional steps leading to Arabidopsis stomatal differentiation WT scrm-d/+
Kanaoka, Pillitteri, Fujii, Yoshida, Bogenschutz, Takabayashi, Zhu, and Torii (2008) SCRM/ICE1 and SCRM2 specify three cell-state transitional steps leading to Arabidopsis stomatal differentiation WT scrm-d/+
FMF NIRCA PROTOCOL STEP 1.
 FMF NIRCA PROTOCOL STEP 1. After you have isolated patient s DNA and DNA from a healthy donor (wild type), you perform a nested PCR. The primers used to amplify exon 2 and exon 10 of the mefv gene are
FMF NIRCA PROTOCOL STEP 1. After you have isolated patient s DNA and DNA from a healthy donor (wild type), you perform a nested PCR. The primers used to amplify exon 2 and exon 10 of the mefv gene are
Supplemental Figure 1.
 Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Problem Set 4
 7.016- Problem Set 4 Question 1 Arginine biosynthesis is an example of multi-step biochemical pathway where each step is catalyzed by a specific enzyme (E1, E2 and E3) as is outlined below. E1 E2 E3 A
7.016- Problem Set 4 Question 1 Arginine biosynthesis is an example of multi-step biochemical pathway where each step is catalyzed by a specific enzyme (E1, E2 and E3) as is outlined below. E1 E2 E3 A
Arabidopsis actin depolymerizing factor AtADF4 mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB
 Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Alternative Cleavage and Polyadenylation of RNA
 Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
SUPPLEMENTARY INFORMATION
 R2A MASNSEKNPLL-SDEKPKSTEENKSS-KPESASGSSTSSAMP---GLNFNAFDFSNMASIL 56 R2B MASSSEKTPLIPSDEKNDTKEESKSTTKPESGSGAPPSPS-PTDPGLDFNAFDFSGMAGIL 60 R2A NDPSIREMAEQIAKDPAFNQLAEQLQRSIPNAGQEGGFPNFDPQQYVNTMQQVMHNPEFK
R2A MASNSEKNPLL-SDEKPKSTEENKSS-KPESASGSSTSSAMP---GLNFNAFDFSNMASIL 56 R2B MASSSEKTPLIPSDEKNDTKEESKSTTKPESGSGAPPSPS-PTDPGLDFNAFDFSGMAGIL 60 R2A NDPSIREMAEQIAKDPAFNQLAEQLQRSIPNAGQEGGFPNFDPQQYVNTMQQVMHNPEFK
Supplemental Data. Meng et al. (2011). Plant Cell /tpc B73 CML311 CML436. Gaspé Flint
 Gaspé Flint B73 CML311 CML436 A B C D 10 th leaf 10 th leaf 10 th leaf Supplemental Figure 1. Whole plant images of the four varieties used in this study (A) Extreme early flowering temperate line Gaspé
Gaspé Flint B73 CML311 CML436 A B C D 10 th leaf 10 th leaf 10 th leaf Supplemental Figure 1. Whole plant images of the four varieties used in this study (A) Extreme early flowering temperate line Gaspé
Construction of plant complementation vector and generation of transgenic plants
 MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
B. Transgenic plants with strong phenotype (%)
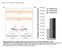 A. TCTAGTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT AAATATGGTCTAAAGAAGAAGAATATGGTCTAAAGAAGAAGAATATGG 2XP35S STTM165 5 GGGGGATGAAGctaCCTGGTCCGA3 3 CCCCCUACUUC---GGACCAGGCU5 mir165 HindIII mir165 96 nt GTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT
A. TCTAGTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT AAATATGGTCTAAAGAAGAAGAATATGGTCTAAAGAAGAAGAATATGG 2XP35S STTM165 5 GGGGGATGAAGctaCCTGGTCCGA3 3 CCCCCUACUUC---GGACCAGGCU5 mir165 HindIII mir165 96 nt GTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT
Box 7905, Engineering Building I, North Carolina State University, Raleigh, NC 27695, Phone: Fax:
 Inefficient ribosomal skipping enables simultaneous secretion and display of proteins in Saccharomyces cerevisiae Carlos A. Cruz-Teran 1, Karthik Tiruthani 1, Adam Mischler, and Balaji M. Rao * Department
Inefficient ribosomal skipping enables simultaneous secretion and display of proteins in Saccharomyces cerevisiae Carlos A. Cruz-Teran 1, Karthik Tiruthani 1, Adam Mischler, and Balaji M. Rao * Department
Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan).
 1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
sides of the aleurone (Al) but it is excluded from the basal endosperm transfer layer
 Supplemental Data. Gómez et al. (2009). The maize transcription factor MRP-1 (Myb-Related-Protein-1) is a key regulator of the differentiation of transfer cells. Supplemental Figure 1. Expression analyses
Supplemental Data. Gómez et al. (2009). The maize transcription factor MRP-1 (Myb-Related-Protein-1) is a key regulator of the differentiation of transfer cells. Supplemental Figure 1. Expression analyses
Supplemental Data. Na Xu et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product.
 Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Regulation of transcription by the MLL2 complex and MLL complex-associated AKAP95
 Supplementary Information Regulation of transcription by the complex and MLL complex-associated Hao Jiang, Xiangdong Lu, Miho Shimada, Yali Dou, Zhanyun Tang, and Robert G. Roeder Input HeLa NE IP lot:
Supplementary Information Regulation of transcription by the complex and MLL complex-associated Hao Jiang, Xiangdong Lu, Miho Shimada, Yali Dou, Zhanyun Tang, and Robert G. Roeder Input HeLa NE IP lot:
Supplemental Information
 Supplemental Information Itemized List Materials and Methods, Related to Supplemental Figures S5A-C and S6. Supplemental Figure S1, Related to Figures 1 and 2. Supplemental Figure S2, Related to Figure
Supplemental Information Itemized List Materials and Methods, Related to Supplemental Figures S5A-C and S6. Supplemental Figure S1, Related to Figures 1 and 2. Supplemental Figure S2, Related to Figure
Supplemental Figure 1 Human REEP family of proteins can be divided into two distinct subfamilies. Residues (single letter amino acid code) identical
 Supplemental Figure Human REEP family of proteins can be divided into two distinct subfamilies. Residues (single letter amino acid code) identical in all six REEPs are highlighted in green. Additional
Supplemental Figure Human REEP family of proteins can be divided into two distinct subfamilies. Residues (single letter amino acid code) identical in all six REEPs are highlighted in green. Additional
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
NAME TA SEC Problem Set 3 FRIDAY March 5, Problem sets will NOT be accepted late.
 MIT Department of Biology 7.013: Introductory Biology - Spring 2004 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. laudette ardel NME T SE 7.013 Problem Set 3 FRIDY March 5, 2004 Problem
MIT Department of Biology 7.013: Introductory Biology - Spring 2004 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. laudette ardel NME T SE 7.013 Problem Set 3 FRIDY March 5, 2004 Problem
Hahn et al. (2011). Plant Cell /tpc
 Supplemental Figure 1. Influence of Hsp70 and Hsp90 on the expression of GUS reporter constructs (A,B) Tomato protoplasts were transformed with the Hsf-dependent promoter GUS reporter plasmid pgmhsp17.3b-ci*:gus
Supplemental Figure 1. Influence of Hsp70 and Hsp90 on the expression of GUS reporter constructs (A,B) Tomato protoplasts were transformed with the Hsf-dependent promoter GUS reporter plasmid pgmhsp17.3b-ci*:gus
Supplemental Data. Aung et al. (2011). Plant Cell /tpc
 35S pro:pmd1-yfp 10 µm Supplemental Figure 1. -terminal YFP fusion of PMD1 (PMD1-YFP, in green) is localized to the cytosol. grobacterium cells harboring 35S pro :PMD1-YFP were infiltrated into tobacco
35S pro:pmd1-yfp 10 µm Supplemental Figure 1. -terminal YFP fusion of PMD1 (PMD1-YFP, in green) is localized to the cytosol. grobacterium cells harboring 35S pro :PMD1-YFP were infiltrated into tobacco
Characterization of a novel gene involved in border cell migration
 Characterization of a novel gene involved in border cell migration Wei-Hao Li, He-Yen Chou and Li-Mei Pai Graduate Institute of Basic Medical Science Chang-Gung University, Tao-Yuan, Taiwan, R.O.C. Abstract
Characterization of a novel gene involved in border cell migration Wei-Hao Li, He-Yen Chou and Li-Mei Pai Graduate Institute of Basic Medical Science Chang-Gung University, Tao-Yuan, Taiwan, R.O.C. Abstract
Supplemental Figure 1. Mutation in NLA Causes Increased Pi Uptake Activity and
 Supplemental Figure 1. Mutation in NLA Causes Increased Pi Uptake Activity and PHT1 Protein Amounts. (A) Shoot morphology of 19-day-old nla mutants under Pi-sufficient conditions. (B) [ 33 P]Pi uptake
Supplemental Figure 1. Mutation in NLA Causes Increased Pi Uptake Activity and PHT1 Protein Amounts. (A) Shoot morphology of 19-day-old nla mutants under Pi-sufficient conditions. (B) [ 33 P]Pi uptake
Supplementary Figure 1. Homozygous rag2 E450fs mutants are healthy and viable similar to wild-type and heterozygous siblings.
 Supplementary Figure 1 Homozygous rag2 E450fs mutants are healthy and viable similar to wild-type and heterozygous siblings. (left) Representative bright-field images of wild type (wt), heterozygous (het)
Supplementary Figure 1 Homozygous rag2 E450fs mutants are healthy and viable similar to wild-type and heterozygous siblings. (left) Representative bright-field images of wild type (wt), heterozygous (het)
Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified
 Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified by primers used for mrna expression analysis. Gray
Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified by primers used for mrna expression analysis. Gray
Supplemental Data. Xie et al. (2014). Plant Cell /tpc
 Supplemental Figure 1. The phylogenetic tree of rice and Arabidopsis Group A/B/C MAPKs that contain the TEY motif in their activation loop. Tobacco SIPK and WIPK, human ERK2 and P38 were also included
Supplemental Figure 1. The phylogenetic tree of rice and Arabidopsis Group A/B/C MAPKs that contain the TEY motif in their activation loop. Tobacco SIPK and WIPK, human ERK2 and P38 were also included
CONTENTS. LIST OF FIGURES... i. LIST OF TABLES... iii. SUMMARY OF THE WORK DONE... vi
 CONTENTS SECTION ~ LIST OF FIGURES... i LIST OF TABLES... iii LIST OF ABBREVIATIONS USED... ~... iv SUMMARY OF THE WORK DONE... vi 1. INTROD UCTION... :.................................... 1 1.1. AN OVERVIEW
CONTENTS SECTION ~ LIST OF FIGURES... i LIST OF TABLES... iii LIST OF ABBREVIATIONS USED... ~... iv SUMMARY OF THE WORK DONE... vi 1. INTROD UCTION... :.................................... 1 1.1. AN OVERVIEW
Supplemental Data. Liu et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. The GFP Tag Does Not Disturb the Physiological Functions of WDL3. (A) RT-PCR analysis of WDL3 expression in wild-type, WDL3-GFP, and WDL3 (without the GFP tag) transgenic seedlings.
Supplemental Figure 1. The GFP Tag Does Not Disturb the Physiological Functions of WDL3. (A) RT-PCR analysis of WDL3 expression in wild-type, WDL3-GFP, and WDL3 (without the GFP tag) transgenic seedlings.
Supplementary Data Supplementary Figures
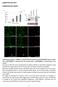 Supplementary Data Supplementary Figures Supplementary Figure 1. Pi04314 is expressed during infection, each GFP-Pi04314 fusion is stable and myr GFP-Pi04314 is removed from the nucleus while NLS GFP-Pi04314
Supplementary Data Supplementary Figures Supplementary Figure 1. Pi04314 is expressed during infection, each GFP-Pi04314 fusion is stable and myr GFP-Pi04314 is removed from the nucleus while NLS GFP-Pi04314
Understanding the Cellular Mechanism of the Excess Microsporocytes I (EMSI) Gene. Andrew ElBardissi, The Pennsylvania State University
 Understanding the Cellular Mechanism of the Excess Microsporocytes I (EMSI) Gene Andrew ElBardissi, The Pennsylvania State University Abstract: Hong Ma, The Pennsylvania State University The Excess Microsporocytes
Understanding the Cellular Mechanism of the Excess Microsporocytes I (EMSI) Gene Andrew ElBardissi, The Pennsylvania State University Abstract: Hong Ma, The Pennsylvania State University The Excess Microsporocytes
(phosphatase tensin) domain is shown in dark gray, the FH1 domain in black, and the
 Supplemental Figure 1. Predicted Domain Organization of the AFH14 Protein. (A) Schematic representation of the predicted domain organization of AFH14. The PTEN (phosphatase tensin) domain is shown in dark
Supplemental Figure 1. Predicted Domain Organization of the AFH14 Protein. (A) Schematic representation of the predicted domain organization of AFH14. The PTEN (phosphatase tensin) domain is shown in dark
Supplemental Data. Lee et al. Plant Cell. (2010) /tpc Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2.
 Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Cassette denotes the ORFs expressed by the expression cassette targeted by the PCR primers used to
 Stanyon, C.A., Limjindaporn, T., and Finley, Jr., R.L. Simultaneous transfer of open reading frames into several different expression vectors. Biotechniques, 35, 520-536, 2003. http://www.biotechniques.com
Stanyon, C.A., Limjindaporn, T., and Finley, Jr., R.L. Simultaneous transfer of open reading frames into several different expression vectors. Biotechniques, 35, 520-536, 2003. http://www.biotechniques.com
Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon Song, Richard M. Amasino, Bosl Noh, and Yoo-Sun Noh
 Developmental Cell, Volume 22 Supplemental Information Control of Seed Germination by Light-Induced Histone Arginine Demethylation Activity Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon
Developmental Cell, Volume 22 Supplemental Information Control of Seed Germination by Light-Induced Histone Arginine Demethylation Activity Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon
Supplemental Data. Bai et al. Plant Cell. (2012) /tpc A
 A B Unconserved Conserved AIF4 AIF2 AIF3 AIF1 UPB1 IBH1 Supplemental Figure 1. IBH1, UPB1 and AIFs belong to the same HLH family. (A), Part of a phylogenetic tree constructed using conserved domains shows
A B Unconserved Conserved AIF4 AIF2 AIF3 AIF1 UPB1 IBH1 Supplemental Figure 1. IBH1, UPB1 and AIFs belong to the same HLH family. (A), Part of a phylogenetic tree constructed using conserved domains shows
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
Intron distance to transcription start (nt)*
 Schwab et al. SOM: Enhanced microrna accumulation through stemloop-adjacent introns 1 Supplemental Table 1. Characteristics of introns used in this study Intron name Length intron (nt) Length cloned fragment
Schwab et al. SOM: Enhanced microrna accumulation through stemloop-adjacent introns 1 Supplemental Table 1. Characteristics of introns used in this study Intron name Length intron (nt) Length cloned fragment
Chinese Academy of Sciences, Datun Road, Chaoyang District, Beijing, 10101, China
 SUPPLEMENTARY INFORMATION Title: Bi-directional processing of pri-mirnas with branched terminal loops by Arabidopsis Dicer-like1 Hongliang Zhu 1,2,3, Yuyi Zhou 1,2,4, Claudia Castillo-González 1,2, Amber
SUPPLEMENTARY INFORMATION Title: Bi-directional processing of pri-mirnas with branched terminal loops by Arabidopsis Dicer-like1 Hongliang Zhu 1,2,3, Yuyi Zhou 1,2,4, Claudia Castillo-González 1,2, Amber
Impact of Nutraceuticals on TERT gene encoded protein
 Impact of Nutraceuticals on TERT gene encoded protein Xu Liu Department of Biological Sciences Fordham University, Bronx, New York, 10458 Abstract Telomerase is a Ribonucleo-protein polymerase that plays
Impact of Nutraceuticals on TERT gene encoded protein Xu Liu Department of Biological Sciences Fordham University, Bronx, New York, 10458 Abstract Telomerase is a Ribonucleo-protein polymerase that plays
Supplementary Figure S1 Complementarity between VR-RNA and cola mrna 5 UTR is important for cola regulation. (A) Mutation sites within VR-RNA-colA
 Supplementary Figure S1 Complementarity between VR-RNA and cola mrna 5 UTR is important for cola regulation. (A) Mutation sites within VR-RNA-colA RNA duplex. Mutation sites in cola mrna 5 UTR or VR-RNA
Supplementary Figure S1 Complementarity between VR-RNA and cola mrna 5 UTR is important for cola regulation. (A) Mutation sites within VR-RNA-colA RNA duplex. Mutation sites in cola mrna 5 UTR or VR-RNA
Problem Set 4
 2006 7.012 Problem Set 4 Due before 5 PM on FRIDAY, October 27, 2006. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. You are studying a specific gene in yeast,
2006 7.012 Problem Set 4 Due before 5 PM on FRIDAY, October 27, 2006. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. You are studying a specific gene in yeast,
Site directed mutagenesis, Insertional and Deletion Mutagenesis. Mitesh Shrestha
 Site directed mutagenesis, Insertional and Deletion Mutagenesis Mitesh Shrestha Mutagenesis Mutagenesis (the creation or formation of a mutation) can be used as a powerful genetic tool. By inducing mutations
Site directed mutagenesis, Insertional and Deletion Mutagenesis Mitesh Shrestha Mutagenesis Mutagenesis (the creation or formation of a mutation) can be used as a powerful genetic tool. By inducing mutations
Supplemental Figure legends Figure S1. (A) (B) (C) (D) Figure S2. Figure S3. (A-E) Figure S4. Figure S5. (A, C, E, G, I) (B, D, F, H, Figure S6.
 Supplemental Figure legends Figure S1. Map-based cloning and complementation testing for ZOP1. (A) ZOP1 was mapped to a ~273-kb interval on Chromosome 1. In the interval, a single-nucleotide G to A substitution
Supplemental Figure legends Figure S1. Map-based cloning and complementation testing for ZOP1. (A) ZOP1 was mapped to a ~273-kb interval on Chromosome 1. In the interval, a single-nucleotide G to A substitution
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the
 Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
The Biotechnology Toolbox
 Chapter 15 The Biotechnology Toolbox Cutting and Pasting DNA Cutting DNA Restriction endonuclease or restriction enzymes Cellular protection mechanism for infected foreign DNA Recognition and cutting specific
Chapter 15 The Biotechnology Toolbox Cutting and Pasting DNA Cutting DNA Restriction endonuclease or restriction enzymes Cellular protection mechanism for infected foreign DNA Recognition and cutting specific
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/5/244/ra72/dc1 Supplementary Materials for An Interaction Between BZR1 and DELLAs Mediates Direct Signaling Crosstalk Between Brassinosteroids and Gibberellins
www.sciencesignaling.org/cgi/content/full/5/244/ra72/dc1 Supplementary Materials for An Interaction Between BZR1 and DELLAs Mediates Direct Signaling Crosstalk Between Brassinosteroids and Gibberellins
SUPPLEMENTAL MATERIALS
 SUPPLEMENTAL MATERIALS SUPPLEMENTAL FIGURES Supplemental Figure S1. Analyses of the CRY1-SPA1 interaction (A) An auxotrophy growth assay (left) and a filter-based -galactosidase colorimetric assay (right)
SUPPLEMENTAL MATERIALS SUPPLEMENTAL FIGURES Supplemental Figure S1. Analyses of the CRY1-SPA1 interaction (A) An auxotrophy growth assay (left) and a filter-based -galactosidase colorimetric assay (right)
Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were
 1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system
 Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
