Supplemental Figure 1
|
|
|
- Brian Goodman
- 5 years ago
- Views:
Transcription
1 Supplemental Figure 1 A gta2-1 gta2-2 1kb AT4G08350 B Col-0 gta2-1 LP+RP+LBb1.3 C Col-0 gta2-2 LP+RP+LB1 D Col-0 gta2-2 gta2-1 GTA2 TUBLIN E Col0 gta2-1 gta2-2 Supplemental Figure 1. Phenotypic analysis of gta2 mutants. (A) Gene structure of GTA2. Exons are shown as boxes and introns are indicated as lines. (B) and (C) Genotypic analysis of gta2 mutants with left genomic primer (LP), right genomic primer (RP), and the vector primer (LBb1.3 or LB1) (D) The transcripts of GTA2 were tested in Col-0 and gta2 mutants. (E) The flowering time of Col-0 and gta2 mutants. 1
2 Supplemental Figure 2 A At SPT5 NGN KOW KOW KOW CTR KOW 989 aa At GTA2 NGN KOW KOW KOW KOW CTR KOW 1041 aa Spt5N NGN Hs SPT5 KOW KOW KOW KOW CTR KOW 1087 aa Spt5N NGN KOW KOW KOW KOW CTR KOW Dm SPT aa Sc SPT5 Spt5N NGN KOW KOW KOW KOW CTR 1063 aa B GTA2 A. thaliana SPT5 A.thaliana SPT5 H.sapiens SPT5 D.melanogaster SPT5 S.cerevisiae 0.1 Supplemental Figure 2. Phylogenetic analysis of SPT5 proteins from Arabidopsis, humans, Drosophila, and yeast. (A) Diagram of the domain structures of SPT5 proteins from Arabidopsis, humans, Drosophila, and yeast. Proteins all contain NGN (yellow boxes), KOW (blue boxes), and CTR (black boxes) domains. aa = amino acids. (B) The SPT5 proteins from Arabidopsis, human, Drosophila, and yeast were aligned with ClustalW. The relationships of those sequences were examined with MEGA5. The evolutional scar bar was indicated in bottom. 2
3 mrna fold enrichment Supplemental Data. Lu et al. (2017). Plant Cell /tpc Supplemental Figure 3 A Col-0 spt5 T1 T14 B Col-0 spt5 T1 T14 Supplemental Figure 3. Complementation of spt5 with ProSPT5::FLAG-SPT5. (A) Phenotypic analysis of flowering time in 4-week-old plants. Over 40 independent transgenic lines were generated, and two representative transgenic plants are shown. (B) Transcript analysis of SPT5 in the Col-0, spt5 mutant, and two transgenic lines (T1 and T14). Experiments were repeated at least three times, and each experiment included three replicates, the representative experiments shown indicate the mean ±SE, n = 3 replicates. 3
4 Supplemental Figure 4 A CDKC;1 CDKC;1 CDKC;2 CDKC;1+CYCT1;3 CDKC;2+CYCT1;3 CDKC;2 SPT5N CDKC;1 CDKC;2 CDKC;1+CYCT1;3 CDKC;2+CYCT1;3 SPT5N CycH1;3 Coomassie staining Autoradiography B CDKC;1 CDKC;1+CYCT1;3 CDKC;2 CDKC;2+CYCT1;3 CDKC;1 CDKC;1+CYCT1;3 CDKC;2 CDKC;2+CYCT1;3 CDKC;1 CDKC;2 CycH1;3 SPT5C SPT5C Coomassie staining Autoradiography C CDKD;1 CDKD;2 CDKD;3 CDKF:1 CDKD;1 CDKD;2 CDKD;3 CDKF:1 CDKF1 CDKD;2 SPT5N SPT5N Coomassie staining Autoradiography Supplemental Figure 4. The phosphorylation activity of CDKC and CDK proteins (A) and (B) CDKC;1 and CDKC;2 were tested for their ability to phosphorylate the N-terminus of SPT5 (SPT5N) or C-terminus of SPT5 (SPT5C). Cyclin T1;3 was used to enhance the CDKC;1 and CDKC;2 phosphorylation activity. The left panel shows the Coomassie blue-stained gel. The positions of different proteins are indicated on the right. Autoradiography in the right panel shows the activity and specificity of the kinase. The positions of the proteins are indicated by arrows. (C) CDKD;1, CDKD;2, CDKD;3, and CDKF;1 were tested for their ability to phosphorylate the N-terminus of SPT5 (SPTN). The left panel shows the Coomassie blue-stained gel. The positions of different CDK proteins and SPT5 are indicated on the right. Autoradiography in the right panel shows the activity and specificity of the kinase. 4
5 Supplemental Figure 5 A T1 T14 B A13 A15 D4 D21 Anti-FLAG Anti-H3 C Col-0 spt5 A13 A15 D4 D21 Col-0 A9 A41 D11 D43 D Relative expreesion level Col-0 A9 A41 D11 D43 E Total leaf number Col-0 spt5 A9 A13 A15 A41 D4 D11 D21 D43 Supplemental Figure 5. Complementation of spt5 with mutated SPT5 (A) SPT5 accumulation in different transgenic plants. The spt5 mutant was complemented by genomic SPT5 with mutations that encoded nonphosphorylatable (SPT5 TA ; transgenic lines A13 and A15) or phospho-mimic (SPT TD ; transgenic lines D4 and D21) forms of SPT5. T1 and T14 are indicated in Supplemental Figure 3. The amount of SPT5 was determined by immunoblot analysis with a FLAG antibody (anti-flag). H3 was used as the internal control (anti-h3). (B) The flowering time of Col-0, spt5, and transgenic plants (transgenic lines A13, A15, D4 and D21) in a long-day photoperiod. (C) The flowering time of Col-0 and nonphosphorylatable (transgenic lines A9 and A41) or phospho-mimic (transgenic lines D11 and D43) forms of SPT5 in a long-day photoperiod. (D) The transcripts of SPT5 in transgenic plants (A9, A41, D11, and D43). Experiments were repeated at least three times, and each experiment included three replicates, and the representative experiments shown indicate the mean ±SE, n = 3 replicates. (E) Flowering time was assessed by counting the total leaf number at bolting under long-day photoperiod. 5
6 Supplemental Figure 6 A Hs RTF1 At VIP5 VHDFDVKIDLQVPSSESKALA-ITSKAPPAKDGAPRRSLNL--EDYKKRRGLI LQKYGGPQGVQKAFMARKQLTEATVGCRVAENDGKRHGLTLTVSDYKRRRGLL + + +Q K L T A++ R L L DYK+RRGL+ B C His-VIP5 Anti-VIP5 D 100kd Col-0 vip5 70kd Anti-VIP5 50kd Anti-H3 Supplemental Figure 6. Commercial antibody to mammalian RTF1 recognizes Arabidopsis VIP5. (A) The amino acid sequences of human RTF1 ( amino acids) and Arabidopsis VIP5 ( amino acids) were aligned by ClustalW2, with the sequence of the human antigen peptide used to make antibodies shown in red. (B) The cdna for the C-terminus of VIP5 was expressed in E. coli as a N-terminal fusion to His. The His-purified fusion proteins (His-VIP5C) produced were visualized on a Coomassie-stained protein gel. (C) The same purified samples were analyzed by immunoblot using a commercially available antibody to human RTF1 (Abcam ab52887). The antibody is specific to the C-terminus of human RTF1 and recognizes Arabidopsis VIP5. (D) The cell extracts from wild-type and vip5 plants were blotted with anti-rtf1 antibody (top panel) using H3 as a control (anti-h3; bottom panel). 6
7 Supplemental Table 1 GST-CDKC;1 To generate GST-CDKC;1, the CDKC;1 was amplified from an Arabidopsis first strand cdna pool using forward primer (5 - CGGAATTCATGGCGATGGCATCATTCGG-3 ) and reverse primer (5 - GCGTCGACCTGTTGCCATCCGTATTGCT-3 ), and cloned into pgex-6p-1 with EcoRI/ SalI GST-CDKC;2 To generate GST-CDKC2, the CDKC;2 was amplified from an Arabidopsis first strand cdna pool using forward primer (5 - CGGAATTCATGGCGGCTGCGGCTTTTGG-3 ) and reverse primer (5 - GCGTCGACCGGTTGCCATCCATATTGTT-3 ), and cloned into pgex-6p-1 with EcoRI/ SalI GST- CYCT1;3 To generate GST- CYCT1;3, the CYCT1;3 was amplified from an Arabidopsis first strand cdna pool using forward primer (5 - GGAATTCATGGGAGAGGAGCATCCGAGAAAG -3 ) and reverse primer (5 - GACGTCGACCCAGATGCCAGCCTGTCTATAGGA -3 ), and cloned into pgex-6p-1 with EcoRI/ SalI GST-CDKF;1 To generate GST-CDKF;1, the CDKF;1 was amplified from an Arabidopsis first strand cdna pool using forward primer (5 -TCCCCCGGG TTTAAAGGAGATGGATAAACAA-3 ) and reverse primer (5 - ACGCGTCGAC ATGAAAAATAGGGTAAAGAATGGC-3 ), and cloned into pgex-6p-1 with SmaI/ SalI GST-CDKD;1 To generate GST-CDKD;1, the CDKD;1 was amplified from an Arabidopsis first strand cdna pool using forward primer (5 -"CGGGATCC CTAAAAATGTCGATTCCTTCTGTA-3 ) and reverse primer (5 - "CGGAATTCTCAAGAAGAAGCCTGTTACGCGAT-3 ), and cloned into pgex-6p-1 with BamHI/EcoRI GST-CDKD;2 To generate GST-CDKD;2, the CDKD;2 was amplified from an Arabidopsis first strand cdna pool using forward primer (5 - "CGGGATCC GCCCTCGTACTTCCTCGAAGTGTA-3 ) and reverse primer (5 - TCCCCCGGG AATCCTTCAGGACCCATCACTCTC-3 ), and cloned into pgex-6p-1 with BamHI/ SmaI GST-CDKD;3 To generate GST-CDKD;3, the CDKD;3 was amplified from an Arabidopsis first strand cdna pool using forward primer (5 - "CGGGATCC ATTATTGAGTGGGTTCTGAAAAAC-3 ) and reverse primer (5 - ACGCGTCGAC ATAACCAACAGTCTTACTGGAACT-3 ), and cloned into pgex-6p-1 with BamHI/ SalI 7
8 pgex6p-1-spt5 N To generate pgex6p-1-spt5 N, the SPT5 N fragment was amplified from an Arabidopsis first strand cdna pool using forward primer (5 -CGCGGATCCATGCTTTTCAAAGATGGTTTTC-3 ) and reverse primer (5 - CGGAATTCTTGGCTTCCCATATTATACTGCGG -3 ), and cloned into pgex-6p-1 with BamHI/EcoRI. GST-SPT5 C To generate GST-SPT5 C, the C fragment from SPT5 cdna was amplified from an Arabidopsis first strand cdna pool using forward primer (5 - CGGGATCCGTCGCAACCCCGCAGTAT AATAT -3 ) and reverse primer (5 - CGGAATTC TCACTCATGAACTAACTTGGCTAA -3 ), and cloned into pgex-6p-1 with BamHI/EcoR I pgex6p-1-spt5 CTR To generate pgex6p-1-spt5 CTR, the SPT5 CTR fragment was amplified from an Arabidopsis first strand cdna pool using forward primer (5 - CGGGATCC GTCGCAACCCCGCAGTATAATAT-3 ) and reverse primer (5 - CGGAATTC GAGACATGACATCGAGATCGGTTC -3 ), and cloned into pgex-6p-1 with BamHI/EcoRI. PET30a-SPT5 KOW To generate pgbkt7-spt5 KOW6, the KOW6 fragment was amplified from an Arabidopsis first strand cdna pool using forward primer (5 - CGGGATCCAACAAAGTGAGCCTAGTTTGTC-3 ) and reverse primer (5 - CGGAATTC TCACTCATGAACTAACTTGGCTAA-3 ), and cloned into PET30a with BamHI/EcoRI. pgex6p-1-spt5 KOW To generate pgex6p-1-spt5 KOW6, the KOW6 fragment was digested from PET30a-SPT5 KOW6 by BamHI/EcoRI, cloned into pgex6p-1. PET- SPT5 CTR To generate PET- SPT5 CTR4 repeat, the CTR4 repeat was amplified from an Arabidopsis first strand cdna pool using forward primer (5 - CGGGATCCGGAAGCCAAACTCCTATGCATCCTT-3 ) and reverse primer (5 -CGGAATTCGATAGGTGTAGCTCCAGAATGCCGC -3 ), and cloned into PET-30a with BamHI/EcoRI PET- SPT5 CTR SA To generate PET- SPT5 CTR SA, the SPT5 CTR SA fragment was amplified from an Arabidopsis first strand cdna pool using forward primer (5 - CCGGAATTC GAAACACCTATGCATCCTG -3 ) and reverse primer (5 - CCGCTCGAG CCTCATTCCATCATGGATTGGTGTAGCTCCAGC -3 ), and cloned into PET-30a with EcoRI/XhoI PET-SPT5 CTR TA To generate PET-SPT5 CTR TA, the CTR TA fragment was amplified from an Arabidopsis first strand cdna pool using forward primer (5-8
9 CGGGATCCGGAAGCCAAGCTCCTATGCATCCTTCCCGGGCTCCACTT-3 ) and reverse primer (5 - CGGAATTCGATAGGTGCAGCTCCAGAATGCCGCATTGGAGCCATACA -3 ), and cloned into PET-30a with EcoRI/BamHI PET-SPT5 CTR TD To generate PET-SPT5 CTR TD, The CTR TD fragment was amplified from an Arabidopsis first strand cdna pool using forward primer (5 - CATGCCATGG GGAAGCCAAGATCCTATGCATCCTTCCCGGGATCCACTT-3 ) and reverse primer (5 - CGGAATTC GATAGGATCAGCTCCAGAATGCCGCATTGGATCCATACA -3 ), and cloned into PET-30a with NcoI/BamHI PET- SPT5 CTR TE To generate PET- SPT5 CTR TE, the CTR TE fragment was amplified from an Arabidopsis first strand cdna pool using forward primer (5 - CGGGATCC GGAAGCCAAGAACCTATGCATCCTTCCCGGGAACCACTT-3 ) and reverse primer (5 - CGGAATTC GATAGGTTCAGCTCCAGAATGCCGCATTGGTTCCATACA -3 ), and cloned into PET-30a with BamHI/EcoRI ProSPT5::SPT5-1300FLAG/N To generate ProSPT5::SPT5-1300Flag/N. The SPT5 promoter was amplified from an Arabidopsis genomic using forward primer (5 - TGCCTGCAGG AGAGACACTCACCAACAGAGACCT-3 ) and reverse primer (5 -CGGGGTACC TGGTTCGATTACAGAGAAACTAGG-3 ), and cloned into pcambia1300flag/n with SbfI/KpnI. The SPT5 was amplified from an Arabidopsis genomic using forward primer (5 - TCAGC AGTCGAAGAGC ATGTCTCAGTACTCAGACGACGAT-3 ) and reverse primer (5 - TTAGCGTGTGAAGA GC CTCCTGAACTAACTTGGCTAA-3 ), cloned into ProSPT5:: pcambia1300flag/n by Gateway. ProSPT5::SPT5-1300FLAG/N CTR TA To generate ProSPT5::SPT5-1300FLAG/N CTR TA. The CTR T mutant to A was amplified from ProSPT5::SPT5-1300Flag/N using forward primer 1 (5 - GCTCCTATGCATCCTTCCCGGGCTCCACTTCATCCCTGTATGGCTCCAATGCGGCATTCT G-3 ), reverse 1 primer (5 - CCATACAGGGATGAAGTGGAGCCCGGGAAGGATGCATAGGAGCTTGGCTTCCCATAT TA -3 ), forward primer 2 (5 - GCACCTATCCATGATGGAATGAGGGCACCTATGCGTGGt-3 ), reverse primer 2 (5 - TGCCCTCATTCCATCATGGATAGGTGCAGCTGAATTTTT-3 ), forward primer 3 (5 - AGATTGGGGTAGTAGTGCTCCTGGTCGTA-3 ), and reverse primer 3 (5 - AGCACTACTACCCCAATCTGATCCAGGCG-3 ) ProSPT5::SPT5-1300FLAG/N CTR TD To generate ProSPT5::SPT5-1300FLAG/N CTR TD. The CTR T mutant to D was amplified from ProSPT5::SPT5-1300FLAG/N using forward primer 1 (5 - GATCCTATGCATCCTTCCCGGGATCCACTTCATCCCTGTATGGATCCAATGCGGCATTC 9
10 -3 ), reverse 1 primer (5 - ATCCATACAGGGATGAAGTGGATCCCGGGAAGGATGCATAGGATCTTGGCTTCCCATAt -3 ), forward primer 2 (5 - GATCCTATCCATGATGGAATGAGGGATCCTATGCGTGGt -3 ), reverse primer 2 (5 -ATCCCTCATTCCATCATGGATAGGATCAGCTGAATTTTT-3 ), forward primer 3 (5 - AGATTGGGGTAGTAGTGATCCTGGTCGTA-3 ), and reverse primer 3 (5 - ATCACTACTACCCCAATCTGATCCAGGCG -3 ) pgbkt7-spt5 N To generate pgbkt7-spt5 N, the SPT5 N fragment was amplified from an Arabidopsis first strand cdna pool using forward primer (5 - CATGCCATGGAGGATGAGGACGAAGAAGAC -3 ) and reverse primer (5 - CGGAATTCTAACCCACGAGTCACGGGATAAAT -3 ), and cloned into pgbkt7 with NcoI/EcoR I. pgbkt7-spt5 KOW To generate pgbkt7-spt5 KOW, the KOW fragment was amplified from an Arabidopsis first strand cdna pool using forward primer (5 - CATGCCATGGCTGCTGTTCTTTCTGTCGAG-3 ) and reverse primer (5 - CGGAATTCTTGGCTTCCCATATTATACTGCGG-3 ), and cloned into pgbkt7 with NcoI/EcoR I. PET-SPT5 C To generate PET-SPT5 C, the C fragment from SPT5 cdna was digested from pgex6p-1-spt5 C by BamHI/EcoRI, and cloned into PET30a. pgbkt7-spt5 C To generate pgbkt7-spt5 C, the SPT5 C fragment was digested from PET30a-SPT5 C by NcoI/EcoR I, and cloned into pgbkt7. PET-SPT5 CTR To generate PET-SPT5 CTR, the SPT5 CTR fragment was digested from pgex6p-1-spt5 CTR by BamHI/EcoRI, and cloned into PET30a. pgbkt7-spt5 CTR To generate pgbkt7-spt5 CTR, the CTR fragment was digested from PET3a-SPT5 CTR by NcoI/EcoR I, and cloned into pgbkt7. pgbkt7-spt5 KOW To generate pgbkt7-spt5 KOW6, the KOW6 fragment was digested from PET30a-SPT5 KOW6 by NcoI/EcoR I, and cloned into pgbkt7. PET-VIP5 To generate PET-VIP5, the VIP5 was amplified from an Arabidopsis first strand cdna pool using forward primer (5 - CGCGGATCC ATGGGTGATTTAGAGAACTTGCTT-3 ) and reverse primer (5 - CCGGAATTC AATGCAAATAAATCCGAGAAGA-3 ), and cloned into PET-30a with BamHI/EcoRI 10
11 pgadt7-vip5 To generate pgadt7-vip5, the VIP5 cdna was digested from PET30a-VIP5 by NcoI/ EcoRI, and cloned into pgadt7 pgadt7-vip5 PLUS3 To generate pgadt7-vip5 plus3, the plus3 fragment was amplified from an Arabidopsis first strand cdna pool using forward primer (5 - CATGCCATGGTTGAGTAGTTCCAGCCAAAGTGACA-3 ) and reverse primer (5 - CCGCTCGAGAACATTCATTGGCCTGACTGACGCA-3 ), and cloned into pgadt7 with NcoI/XhoI PUC-spYNE-VIP5 To generate PUC-spYNE-VIP5. The VIP5 cdna was amplified from an Arabidopsis first strand cdna pool using forward primer (5 - CGCGGATCC ATGGGTGATTTAGAGAACTTGCTT-3 ) and reverse primer (5 - TCCCCCGGG AATGCAAATAAATCCGAGAAGA-3 ), and cloned into PUC-spYNE with BamHI/ SmaI PUC-spYNE-CDKD2 To generate PUC-spYNE-CDKD2, the CDKD2 cdna was digested from pgex6p-1-cdkd2 by BamHI/SmaI, and cloned into PUC-spYNE. PUC-spYCE-SPT5 C To generate PUC-spYCE-SPT5 C, the SPT5 C fragment was digested from pgex6p-1-spt5 C by BamHI/Xho1, and cloned into PUC-spYCE. PET-VIP5 C To generate PET-VIP5 C, the VIP5 C fragment was amplified from an Arabidopsis first strand cdna pool using forward primer (5 - CCGGAATTC GAAGCAGCTTTAGAAGCAGCTG-3 ) and reverse primer (5 - TCCCCCGGG AATGCAAATAAATCCGAGAAGA-3 ), and cloned into PET-30a with BamHI/SmaI. Genotyping primers SALK_ Forward primer: GTCTCTTGCCATCGTTCTCTG Reverse primer: ATGTGCCTACGTTTGAGGATG SALKseq_9711 Forward primer: ATTCAACATTCAGCAACCAGC Reverse primer: AGGGTAATATAACGCATGGGG 11
12 SALK_ Forward primer: GATGGTTTGTTTTTGGTGTGG Reverse primer: CTCAGCAAAAATACAGCCTGC SALK_ Forward primer: TTTCCGGATAGCATTTTGTTG Reverse primer: AAGCTGGAATCTGAAGCTTCC SALK_ Forward primer: GGAATTCTGAAACCTCCGAAG Reverse primer: AAGGTCCGTTCCATTATCCAC CS Forward primer: AGGAAATCAAGCCAAACCATG Reverse primer: TGGTGGTAGTTACTCGGATGC flc-3 Forward primer: TATCGCCGGAGGAGAAGC Reverse primer: TAGAAAGAAATAAAGCGAGAAAAGGA FRIGIDA Forward primer: GGGGTACCATGGTGAGCAAGGGCGAGGAGCTGTT Reverse primer: GGGGTACCACTTGTACAGCTCGTCCATGCCGAGA SALK_ Forward primer: GGAATTCTGAAACCTCCGAAG Reverse primer: AAGGTCCGTTCCATTATCCAC CS Forward primer: AGGAAATCAAGCCAAACCATG Reverse primer: TGGTGGTAGTTACTCGGATGC RT-PCR primer SPT5 Forward primer: CGGTGAAAGATGTTGTCAGG Reverse primer: AAGGTTATGCCGGTCATGTAT VIP5 Forward primer: GAAATGAAACCTCGGCGGCT Reverse primer: TCATTGGCCTGACTGACGCA CDKD;2 Forward primer: GATTAGTCTTGTTCTTGATG 12
13 Reverse primer: CTACAGAACTAATCAATTGC FLC Forward primer: CGGTCTCATCGAGAAAGCTC Reverse primer: CCACAAGCTTGCTATCCACA GTA2 Forward primer: GAGTCCTCAATATCAGCCGG Reverse primer: TCACGGTTGCACAAACTTGG UBIQUITIN 10 Forward primer: AGGATGGCAGAACTCTTGCT Reverse primer: TCCCAGTCAACGTCTTAACG Primers for ChIP-PCR FLC Region 1 Forward primer: TGGAGGGAACAACCTAATGC Reverse primer: TCATTGGACCAAACCAAACC Region 2 Forward primer: CGACAAGTCACCTTCTCCAAA Reverse primer: AGGGGGAACAAATGAAAACC Region 3 Forward primer: GGCGGATCTCTTGTTGTTTC Reverse primer: CTTCTTCACGACATTGTTCTTCC Region 4 Forward primer: GGGGCTGCGTTTACATTTTA Reverse primer: GTGATAGCGCTGGCTTTGAT Region 5 Forward primer: CTTTTTCATGGGCAGGATCA Reverse primer: TGACATTTGATCCCACAAGC Region 6 Forward primer: CTTTTTCATGGGCAGGATCA Reverse primer: TGACATTTGATCCCACAAGC Region 6 Forward primer: CGTGTGAGAATTGCATCGAG 13
14 Reverse primer: AAAAACGCGCAGAGAGAGAG UBIQUITIN 10 Forward primer: AGGATGGCAGAACTCTTGCT Reverse primer: TCCCAGTCAACGTCTTAACG 14
S156AT168AY175A (AAA) were purified as GST-fusion proteins and incubated with GSTfused
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 Supplemental Materials Supplemental Figure S1 (a) Phenotype of the wild type and grik1-2 grik2-1 plants after 8 days in darkness.
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 Supplemental Materials Supplemental Figure S1 (a) Phenotype of the wild type and grik1-2 grik2-1 plants after 8 days in darkness.
Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified
 Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified by primers used for mrna expression analysis. Gray
Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified by primers used for mrna expression analysis. Gray
Supplemental Data. Li et al. (2015). Plant Cell /tpc
 Supplemental Data Supplemental Figure 1: Characterization of asr3 T-DNA knockout lines and complementation transgenic lines. (A) The scheme of At2G33550 (ASR3) with gray boxes indicating exons and dash
Supplemental Data Supplemental Figure 1: Characterization of asr3 T-DNA knockout lines and complementation transgenic lines. (A) The scheme of At2G33550 (ASR3) with gray boxes indicating exons and dash
Supplemental Figure 1.
 Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity.
 Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
Supplemental Data. Cui et al. (2012). Plant Cell /tpc a b c d. Stem UBC32 ACTIN
 A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
Supplemental Data. Guo et al. (2015). Plant Cell /tpc
 Supplemental Figure 1. The Mutant exb1-d Displayed Pleiotropic Phenotypes and Produced Branches in the Axils of Cotyledons. (A) Branches were developed in exb1-d but not in wild-type plants. (B) and (C)
Supplemental Figure 1. The Mutant exb1-d Displayed Pleiotropic Phenotypes and Produced Branches in the Axils of Cotyledons. (A) Branches were developed in exb1-d but not in wild-type plants. (B) and (C)
Supplementary Figure 1 Collision-induced dissociation (CID) mass spectra of peptides from PPK1, PPK2, PPK3 and PPK4 respectively.
 Supplementary Figure 1 lision-induced dissociation (CID) mass spectra of peptides from PPK1, PPK, PPK3 and PPK respectively. % of nuclei with signal / field a 5 c ppif3:gus pppk1:gus 0 35 30 5 0 15 10
Supplementary Figure 1 lision-induced dissociation (CID) mass spectra of peptides from PPK1, PPK, PPK3 and PPK respectively. % of nuclei with signal / field a 5 c ppif3:gus pppk1:gus 0 35 30 5 0 15 10
SUPPLEMENTARY INFORMATION
 AS-NMD modulates FLM-dependent thermosensory flowering response in Arabidopsis NATURE PLANTS www.nature.com/natureplants 1 Supplementary Figure 1. Genomic sequence of FLM along with the splice sites. Sequencing
AS-NMD modulates FLM-dependent thermosensory flowering response in Arabidopsis NATURE PLANTS www.nature.com/natureplants 1 Supplementary Figure 1. Genomic sequence of FLM along with the splice sites. Sequencing
Supplemental Data. Furlan et al. Plant Cell (2017) /tpc
 Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Figure.
Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Data. Furlan et al. Plant Cell (0) 0.0/tpc..00. Supplemental Figure.
Construction of plant complementation vector and generation of transgenic plants
 MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
Nature Genetics: doi: /ng.3556 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE. Supplementary Figure 1
 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
Supplemental Data. Jing et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. Characterization of epp1 Mutants. (A) Cotyledon angles of 5-d-old Col wild-type (gray bars) and epp1-1 (black bars) seedlings under red (R), far-red (FR) and blue (BL) light conditions,
Supplemental Figure 1. Characterization of epp1 Mutants. (A) Cotyledon angles of 5-d-old Col wild-type (gray bars) and epp1-1 (black bars) seedlings under red (R), far-red (FR) and blue (BL) light conditions,
Supplemental Data. Na Xu et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected with the wwp-luc reporter, and FLAG-tagged FHL1,
 Supplementary Data Supplementary Figure Legends Supplementary Figure 1 FHL-mediated TGFβ-responsive reporter transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected
Supplementary Data Supplementary Figure Legends Supplementary Figure 1 FHL-mediated TGFβ-responsive reporter transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected
Supplemental Figure legends Figure S1. (A) (B) (C) (D) Figure S2. Figure S3. (A-E) Figure S4. Figure S5. (A, C, E, G, I) (B, D, F, H, Figure S6.
 Supplemental Figure legends Figure S1. Map-based cloning and complementation testing for ZOP1. (A) ZOP1 was mapped to a ~273-kb interval on Chromosome 1. In the interval, a single-nucleotide G to A substitution
Supplemental Figure legends Figure S1. Map-based cloning and complementation testing for ZOP1. (A) ZOP1 was mapped to a ~273-kb interval on Chromosome 1. In the interval, a single-nucleotide G to A substitution
3 -end. Sau3A. 3 -end TAA TAA. ~9.1kb SalI TAA. ~12.6kb. HindШ ATG 5,860bp TAA. ~12.7kb
 Supplemental Data. Chen et al. (2008). Badh2, encoding betaine aldehyde dehydrogenase, inhibits the biosynthesis of 2-acetyl-1-pyrroline, a major component in rice fragrance. A pcam-cah/cah Sau3A Promoter
Supplemental Data. Chen et al. (2008). Badh2, encoding betaine aldehyde dehydrogenase, inhibits the biosynthesis of 2-acetyl-1-pyrroline, a major component in rice fragrance. A pcam-cah/cah Sau3A Promoter
Alternative Cleavage and Polyadenylation of RNA
 Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
Supplemental Data. Sethi et al. (2014). Plant Cell /tpc
 Supplemental Data Supplemental Figure 1. MYC2 Binds to the E-box but not the E1-box of the MPK6 Promoter. (A) E1-box and E-box (wild type) containing MPK6 promoter fragment. The region shown in red denotes
Supplemental Data Supplemental Figure 1. MYC2 Binds to the E-box but not the E1-box of the MPK6 Promoter. (A) E1-box and E-box (wild type) containing MPK6 promoter fragment. The region shown in red denotes
Nature Genetics: doi: /ng Supplementary Figure 1. ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets.
 Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Sperm cells are passive cargo of the pollen tube in plant fertilization
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 3 ARTICLE NUMBER: 17079 Sperm cells are passive cargo of the pollen tube in plant fertilization Jun Zhang 1, Qingpei
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 3 ARTICLE NUMBER: 17079 Sperm cells are passive cargo of the pollen tube in plant fertilization Jun Zhang 1, Qingpei
Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon Song, Richard M. Amasino, Bosl Noh, and Yoo-Sun Noh
 Developmental Cell, Volume 22 Supplemental Information Control of Seed Germination by Light-Induced Histone Arginine Demethylation Activity Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon
Developmental Cell, Volume 22 Supplemental Information Control of Seed Germination by Light-Induced Histone Arginine Demethylation Activity Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon
Supplemental Data. Zhang et al. (2010). Plant Cell /tpc
 Supplemental Figure 1. uvs90 gene cloning The T-DNA insertion in uvs90 was identified using thermal asymmetric interlaced (TAIL)-PCR. Three rounds of amplification were performed; the second (2 nd ) and
Supplemental Figure 1. uvs90 gene cloning The T-DNA insertion in uvs90 was identified using thermal asymmetric interlaced (TAIL)-PCR. Three rounds of amplification were performed; the second (2 nd ) and
Fig. S1. TPL and TPL N176H protein interactions. (A) Semi-in vivo pull-down assays using recombinant GST N-TPL and GST N-TPL N176H fusions and
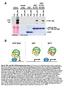 Fig. S1. TPL and TPL N176H protein interactions. (A) Semi-in vivo pull-down assays using recombinant GST N-TPL and GST N-TPL N176H fusions and transgenic Arabidopsis TPL-HA lysates. Immunoblotting of input
Fig. S1. TPL and TPL N176H protein interactions. (A) Semi-in vivo pull-down assays using recombinant GST N-TPL and GST N-TPL N176H fusions and transgenic Arabidopsis TPL-HA lysates. Immunoblotting of input
Supplemental Data. Zhou et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. Confirmation of mutant mapping results. (A) Complementation assay with stably transformed genomic fragments (ComN-N) (2 kb upstream of TSS and 1.5 kb downstream of TES) and CaMV
Supplemental Figure 1. Confirmation of mutant mapping results. (A) Complementation assay with stably transformed genomic fragments (ComN-N) (2 kb upstream of TSS and 1.5 kb downstream of TES) and CaMV
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature10928 Materials and Methods 1. Plant material and growth conditions. All plant lines used were in Col-0 background unless otherwise specified. pif4-101 mutant
SUPPLEMENTARY INFORMATION doi:10.1038/nature10928 Materials and Methods 1. Plant material and growth conditions. All plant lines used were in Col-0 background unless otherwise specified. pif4-101 mutant
Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product.
 Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
Supplementary Information Supplementary Figure S1. Immunodetection of full-length XA21 and the XA21 C-terminal cleavage product. Total protein extracted from Kitaake wild type and rice plants carrying
IDN1 and IDN2: two proteins required for de novo DNA methylation in Arabidopsis thaliana.
 IDN1 and IDN2: two proteins required for de novo DNA methylation in Arabidopsis thaliana. Israel Ausin, Todd C. Mockler, Joanne Chory, and Steven E. Jacobsen Col Ler nrpd1a rdr2 dcl3-1 ago4-1 idn1-1 idn2-1
IDN1 and IDN2: two proteins required for de novo DNA methylation in Arabidopsis thaliana. Israel Ausin, Todd C. Mockler, Joanne Chory, and Steven E. Jacobsen Col Ler nrpd1a rdr2 dcl3-1 ago4-1 idn1-1 idn2-1
GTTCGGGTTCC TTTTGAGCAG
 Supplementary Figures Splice variants of the SIP1 transcripts play a role in nodule organogenesis in Lotus japonicus. Wang C, Zhu H, Jin L, Chen T, Wang L, Kang H, Hong Z, Zhang Z. 5 UTR CDS 3 UTR TCTCAACCATCCTTTGTCTGCTTCCGCCGCATGGGTGAGGTCATTTTGTCTAGATGACGTGCAATTTACAATGA
Supplementary Figures Splice variants of the SIP1 transcripts play a role in nodule organogenesis in Lotus japonicus. Wang C, Zhu H, Jin L, Chen T, Wang L, Kang H, Hong Z, Zhang Z. 5 UTR CDS 3 UTR TCTCAACCATCCTTTGTCTGCTTCCGCCGCATGGGTGAGGTCATTTTGTCTAGATGACGTGCAATTTACAATGA
Supplemental Figure 1 A
 Supplemental Figure 1 A Supplemental Data. Han et al. (2016). Plant Cell 10.1105/tpc.15.00997 BamHI -1616 bp SINE repeats SalI ATG SacI +1653 bp HindIII LB HYG p35s pfwa LUC Tnos RB pfwa-bamhi-f: CGGGATCCCGCCTTTCTCTTCCTCATCTGC
Supplemental Figure 1 A Supplemental Data. Han et al. (2016). Plant Cell 10.1105/tpc.15.00997 BamHI -1616 bp SINE repeats SalI ATG SacI +1653 bp HindIII LB HYG p35s pfwa LUC Tnos RB pfwa-bamhi-f: CGGGATCCCGCCTTTCTCTTCCTCATCTGC
Supplemental Data. Polycomb Silencing of KNOX Genes Confines. Shoot Stem Cell Niches in Arabidopsis Current Biology, Volume 18
 Supplemental Data Polycomb Silencing of KNOX Genes Confines Shoot Stem Cell Niches in Arabidopsis Lin Xu and Wen-Hui Shen - 1 - Figure S1. Phylogram of RING1, BMI1 and RAD18 homologues in several organisms
Supplemental Data Polycomb Silencing of KNOX Genes Confines Shoot Stem Cell Niches in Arabidopsis Lin Xu and Wen-Hui Shen - 1 - Figure S1. Phylogram of RING1, BMI1 and RAD18 homologues in several organisms
Enzyme that uses RNA as a template to synthesize a complementary DNA
 Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
The Arabidopsis Transcription Factor BES1 Is a Direct Substrate of MPK6 and Regulates Immunity
 Supplemental Data The Arabidopsis Transcription Factor BES1 Is a Direct Substrate of MPK6 and Regulates Immunity Authors: Sining Kang, Fan Yang, Lin Li, Huamin Chen, She Chen and Jie Zhang The following
Supplemental Data The Arabidopsis Transcription Factor BES1 Is a Direct Substrate of MPK6 and Regulates Immunity Authors: Sining Kang, Fan Yang, Lin Li, Huamin Chen, She Chen and Jie Zhang The following
SUPPLEMENTARY INFORMATION
 Ca 2+ /calmodulin Regulates Salicylic Acid-mediated Plant Immunity Liqun Du, Gul S. Ali, Kayla A. Simons, Jingguo Hou, Tianbao Yang, A.S.N. Reddy and B. W. Poovaiah * *To whom correspondence should be
Ca 2+ /calmodulin Regulates Salicylic Acid-mediated Plant Immunity Liqun Du, Gul S. Ali, Kayla A. Simons, Jingguo Hou, Tianbao Yang, A.S.N. Reddy and B. W. Poovaiah * *To whom correspondence should be
Supplemental Data. Lee et al. Plant Cell. (2010) /tpc Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2.
 Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Figure S1. DELLA Proteins Act as Positive Regulators to Mediate GA-Regulated Anthocyanin
 Supplemental Information Figure S1. DELLA Proteins Act as Positive Regulators to Mediate GA-Regulated Anthocyanin Biosynthesis. (A) Effect of GA on anthocyanin content in WT and ga1-3 seedlings. Mock,
Supplemental Information Figure S1. DELLA Proteins Act as Positive Regulators to Mediate GA-Regulated Anthocyanin Biosynthesis. (A) Effect of GA on anthocyanin content in WT and ga1-3 seedlings. Mock,
Supplemental Figure 1. Immunoblot analysis of wild-type and mutant forms of JAZ3
 Supplemental Data. Chung and Howe (2009). A Critical Role for the TIFY Motif in Repression of Jasmonate Signaling by a Stabilized Splice Variant of the JASMONATE ZIM-domain protein JAZ10 in Arabidopsis.
Supplemental Data. Chung and Howe (2009). A Critical Role for the TIFY Motif in Repression of Jasmonate Signaling by a Stabilized Splice Variant of the JASMONATE ZIM-domain protein JAZ10 in Arabidopsis.
PIE1 ARP6 SWC6 KU70 ARP6 PIE1. HSA SNF2_N HELICc SANT. pie1-3 A1,A2 K1,K2 K1,K3 K3,LB2 A3, A4 A3,LB1 A1,A2 K1,K2 K1,K3. swc6-1 A3,A4.
 A B N-terminal SWC2 H2A.Z SWC6 ARP6 PIE1 HSA SNF2_N HELICc SANT C pie1-3 D PIE1 ARP6 5 Kb A1 200 bp A3 A2 LB1 arp6-3 A4 E A1,A2 A3, A4 A3,LB1 K1,K2 K1,K3 K3,LB2 SWC6 swc6-1 A1,A2 A3,A4 K1,K2 K1,K3 100
A B N-terminal SWC2 H2A.Z SWC6 ARP6 PIE1 HSA SNF2_N HELICc SANT C pie1-3 D PIE1 ARP6 5 Kb A1 200 bp A3 A2 LB1 arp6-3 A4 E A1,A2 A3, A4 A3,LB1 K1,K2 K1,K3 K3,LB2 SWC6 swc6-1 A1,A2 A3,A4 K1,K2 K1,K3 100
Supplementary Information
 Supplementary Information MED18 interaction with distinct transcription factors regulates plant immunity, flowering time and responses to hormones Supplementary Figure 1. Diagram showing T-DNA insertion
Supplementary Information MED18 interaction with distinct transcription factors regulates plant immunity, flowering time and responses to hormones Supplementary Figure 1. Diagram showing T-DNA insertion
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/5/244/ra72/dc1 Supplementary Materials for An Interaction Between BZR1 and DELLAs Mediates Direct Signaling Crosstalk Between Brassinosteroids and Gibberellins
www.sciencesignaling.org/cgi/content/full/5/244/ra72/dc1 Supplementary Materials for An Interaction Between BZR1 and DELLAs Mediates Direct Signaling Crosstalk Between Brassinosteroids and Gibberellins
Figure S1. Figure S2. Figure S3 HB Anti-FSP27 (COOH-terminal peptide) Ab. Anti-GST-FSP27(45-127) Ab.
 / 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
/ 36B4 mrna ratio Figure S1 * 2. 1.6 1.2.8 *.4 control TNFα BRL49653 Figure S2 Su bw AT p iw Anti- (COOH-terminal peptide) Ab Blot : Anti-GST-(45-127) Ab β-actin Figure S3 HB2 HW AT BA T Figure S4 A TAG
A Nucleus-Encoded Chloroplast Protein YL1 Is Involved in Chloroplast. Development and Efficient Biogenesis of Chloroplast ATP Synthase in Rice
 A Nucleus-Encoded Chloroplast Protein YL1 Is Involved in Chloroplast Development and Efficient Biogenesis of Chloroplast ATP Synthase in Rice Fei Chen 1,*, Guojun Dong 2,*, Limin Wu 1, Fang Wang 3, Xingzheng
A Nucleus-Encoded Chloroplast Protein YL1 Is Involved in Chloroplast Development and Efficient Biogenesis of Chloroplast ATP Synthase in Rice Fei Chen 1,*, Guojun Dong 2,*, Limin Wu 1, Fang Wang 3, Xingzheng
SUPPLEMETARY INFORMATION. Cdk7 mediates RPB1-driven mrna synthesis in Toxoplasma gondii. Abhijit S. Deshmukh 1 *, Pallabi Mitra 2 and Mulaka Maruthi 3
 SUPPLEMETARY INFORMATION Cdk7 mediates RPB1-driven mrna synthesis in Toxoplasma gondii Abhijit S. Deshmukh 1 *, Pallabi Mitra 2 and Mulaka Maruthi 3 1 National Institute of Animal Biotechnology, Hyderabad,
SUPPLEMETARY INFORMATION Cdk7 mediates RPB1-driven mrna synthesis in Toxoplasma gondii Abhijit S. Deshmukh 1 *, Pallabi Mitra 2 and Mulaka Maruthi 3 1 National Institute of Animal Biotechnology, Hyderabad,
supplementary information
 DOI: 10.1038/ncb2116 Figure S1 CDK phosphorylation of EZH2 in cells. (a) Comparison of candidate CDK phosphorylation sites on EZH2 with known CDK substrates by multiple sequence alignments. (b) CDK1 and
DOI: 10.1038/ncb2116 Figure S1 CDK phosphorylation of EZH2 in cells. (a) Comparison of candidate CDK phosphorylation sites on EZH2 with known CDK substrates by multiple sequence alignments. (b) CDK1 and
SUPPLEMENTARY INFORMATION
 R2A MASNSEKNPLL-SDEKPKSTEENKSS-KPESASGSSTSSAMP---GLNFNAFDFSNMASIL 56 R2B MASSSEKTPLIPSDEKNDTKEESKSTTKPESGSGAPPSPS-PTDPGLDFNAFDFSGMAGIL 60 R2A NDPSIREMAEQIAKDPAFNQLAEQLQRSIPNAGQEGGFPNFDPQQYVNTMQQVMHNPEFK
R2A MASNSEKNPLL-SDEKPKSTEENKSS-KPESASGSSTSSAMP---GLNFNAFDFSNMASIL 56 R2B MASSSEKTPLIPSDEKNDTKEESKSTTKPESGSGAPPSPS-PTDPGLDFNAFDFSGMAGIL 60 R2A NDPSIREMAEQIAKDPAFNQLAEQLQRSIPNAGQEGGFPNFDPQQYVNTMQQVMHNPEFK
Biology 105: Introduction to Genetics PRACTICE FINAL EXAM Part I: Definitions. Homology: Reverse transcriptase. Allostery: cdna library
 Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Reverse transcriptase Allostery: cdna library Transformation Part II Short Answer 1. Describe the reasons for
Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Reverse transcriptase Allostery: cdna library Transformation Part II Short Answer 1. Describe the reasons for
Supplemental Data. Seo et al. (2014). Plant Cell /tpc
 Supplemental Figure 1. Protein alignment of ABD1 from other model organisms. The alignment was performed with H. sapiens DCAF8, M. musculus DCAF8 and O. sativa Os10g0544500. The WD40 domains are underlined.
Supplemental Figure 1. Protein alignment of ABD1 from other model organisms. The alignment was performed with H. sapiens DCAF8, M. musculus DCAF8 and O. sativa Os10g0544500. The WD40 domains are underlined.
Supplemental Data. Hu et al. Plant Cell (2017) /tpc
 1 2 3 4 Supplemental Figure 1. DNA gel blot analysis of homozygous transgenic plants. (Supports Figure 1.) 5 6 7 8 Rice genomic DNA was digested with the restriction enzymes EcoRⅠ and BamHⅠ. Lanes in the
1 2 3 4 Supplemental Figure 1. DNA gel blot analysis of homozygous transgenic plants. (Supports Figure 1.) 5 6 7 8 Rice genomic DNA was digested with the restriction enzymes EcoRⅠ and BamHⅠ. Lanes in the
Schematic representation of the endogenous PALB2 locus and gene-disruption constructs
 Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Problem Set 8. Answer Key
 MCB 102 University of California, Berkeley August 11, 2009 Isabelle Philipp Online Document Problem Set 8 Answer Key 1. The Genetic Code (a) Are all amino acids encoded by the same number of codons? no
MCB 102 University of California, Berkeley August 11, 2009 Isabelle Philipp Online Document Problem Set 8 Answer Key 1. The Genetic Code (a) Are all amino acids encoded by the same number of codons? no
(phosphatase tensin) domain is shown in dark gray, the FH1 domain in black, and the
 Supplemental Figure 1. Predicted Domain Organization of the AFH14 Protein. (A) Schematic representation of the predicted domain organization of AFH14. The PTEN (phosphatase tensin) domain is shown in dark
Supplemental Figure 1. Predicted Domain Organization of the AFH14 Protein. (A) Schematic representation of the predicted domain organization of AFH14. The PTEN (phosphatase tensin) domain is shown in dark
Candidate region (0.74 Mb) ATC TCT GGG ACT CAT GAG CAG GAG GCT AGC ATC TCT GGG ACT CAT TAG CAG GAG GCT AGC
 A idm-3 idm-3 B Physical distance (Mb) 4.6 4.86.6 8.4 C Chr.3 Recom. Rate (%) ATG 3.9.9.9 9.74 Candidate region (.74 Mb) n=4 TAA D idm-3 G3T(E4) G4A(W988) WT idm-3 ATC TCT GGG ACT CAT GAG CAG GAG GCT AGC
A idm-3 idm-3 B Physical distance (Mb) 4.6 4.86.6 8.4 C Chr.3 Recom. Rate (%) ATG 3.9.9.9 9.74 Candidate region (.74 Mb) n=4 TAA D idm-3 G3T(E4) G4A(W988) WT idm-3 ATC TCT GGG ACT CAT GAG CAG GAG GCT AGC
Supplemental Data. Bai et al. Plant Cell. (2012) /tpc A
 A B Unconserved Conserved AIF4 AIF2 AIF3 AIF1 UPB1 IBH1 Supplemental Figure 1. IBH1, UPB1 and AIFs belong to the same HLH family. (A), Part of a phylogenetic tree constructed using conserved domains shows
A B Unconserved Conserved AIF4 AIF2 AIF3 AIF1 UPB1 IBH1 Supplemental Figure 1. IBH1, UPB1 and AIFs belong to the same HLH family. (A), Part of a phylogenetic tree constructed using conserved domains shows
A Repressor Complex Governs the Integration of
 Developmental Cell 15 Supplemental Data A Repressor Complex Governs the Integration of Flowering Signals in Arabidopsis Dan Li, Chang Liu, Lisha Shen, Yang Wu, Hongyan Chen, Masumi Robertson, Chris A.
Developmental Cell 15 Supplemental Data A Repressor Complex Governs the Integration of Flowering Signals in Arabidopsis Dan Li, Chang Liu, Lisha Shen, Yang Wu, Hongyan Chen, Masumi Robertson, Chris A.
Fig. S1. Clustering analysis of expression array and ChIP-PCR assay in the ARF3 locus. (A) Typical examples of the transgenic plants used for
 Fig. S1. Clustering analysis of expression array and ChIP-PCR assay in the ARF3 locus. (A) Typical examples of the transgenic plants used for ChIP-chip and ChIP-PCR assays. The presence of pas1:t7:as1
Fig. S1. Clustering analysis of expression array and ChIP-PCR assay in the ARF3 locus. (A) Typical examples of the transgenic plants used for ChIP-chip and ChIP-PCR assays. The presence of pas1:t7:as1
Reading Lecture 8: Lecture 9: Lecture 8. DNA Libraries. Definition Types Construction
 Lecture 8 Reading Lecture 8: 96-110 Lecture 9: 111-120 DNA Libraries Definition Types Construction 142 DNA Libraries A DNA library is a collection of clones of genomic fragments or cdnas from a certain
Lecture 8 Reading Lecture 8: 96-110 Lecture 9: 111-120 DNA Libraries Definition Types Construction 142 DNA Libraries A DNA library is a collection of clones of genomic fragments or cdnas from a certain
Characterization of a novel gene involved in border cell migration
 Characterization of a novel gene involved in border cell migration Wei-Hao Li, He-Yen Chou and Li-Mei Pai Graduate Institute of Basic Medical Science Chang-Gung University, Tao-Yuan, Taiwan, R.O.C. Abstract
Characterization of a novel gene involved in border cell migration Wei-Hao Li, He-Yen Chou and Li-Mei Pai Graduate Institute of Basic Medical Science Chang-Gung University, Tao-Yuan, Taiwan, R.O.C. Abstract
(b) Genotyping of SALK_ (AT1g51690) and SALK_ (At1g17720) to find homozygous single mutant plants.
 Supplemental Table 1. Primers used for cloning and RT-PCR. The B-type subunits cdna CDS were amplified using high-fidelity PCR amplification with gene specific primers. The cdna CDS sequences were obtained
Supplemental Table 1. Primers used for cloning and RT-PCR. The B-type subunits cdna CDS were amplified using high-fidelity PCR amplification with gene specific primers. The cdna CDS sequences were obtained
Supplemental Data. Steiner et al. Plant Cell. (2012) /tpc
 Supplemental Figure 1. SPY does not interact with free GST. Invitro pull-down assay using E. coli-expressed MBP-SPY and GST, GST-TCP14 and GST-TCP15. MBP-SPY was used as bait and incubated with equal amount
Supplemental Figure 1. SPY does not interact with free GST. Invitro pull-down assay using E. coli-expressed MBP-SPY and GST, GST-TCP14 and GST-TCP15. MBP-SPY was used as bait and incubated with equal amount
7.012 Problem Set 5. Question 1
 Name Section 7.012 Problem Set 5 Question 1 While studying the problem of infertility, you attempt to isolate a hypothetical rabbit gene that accounts for the prolific reproduction of rabbits. After much
Name Section 7.012 Problem Set 5 Question 1 While studying the problem of infertility, you attempt to isolate a hypothetical rabbit gene that accounts for the prolific reproduction of rabbits. After much
Supplementary Figures 1-12
 Supplementary Figures 1-12 Supplementary Figure 1. The specificity of anti-abi1 antibody. Total Proteins extracted from the wild type seedlings or abi1-3 null mutant seedlings were used for immunoblotting
Supplementary Figures 1-12 Supplementary Figure 1. The specificity of anti-abi1 antibody. Total Proteins extracted from the wild type seedlings or abi1-3 null mutant seedlings were used for immunoblotting
Supplemental Data. Farmer et al. (2010) Plant Cell /tpc
 Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
Supplementary Methods
 Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Enhancers mutations that make the original mutant phenotype more extreme. Suppressors mutations that make the original mutant phenotype less extreme
 Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
scrm-d/+ scrm-d Supplemental Fig. 1.
 Kanaoka, Pillitteri, Fujii, Yoshida, Bogenschutz, Takabayashi, Zhu, and Torii (2008) SCRM/ICE1 and SCRM2 specify three cell-state transitional steps leading to Arabidopsis stomatal differentiation WT scrm-d/+
Kanaoka, Pillitteri, Fujii, Yoshida, Bogenschutz, Takabayashi, Zhu, and Torii (2008) SCRM/ICE1 and SCRM2 specify three cell-state transitional steps leading to Arabidopsis stomatal differentiation WT scrm-d/+
Supplemental Table 1. List of PCR primers
 Supplemental Table 1. List of PCR primers Primer Sequences (5 to 3 ) 1 TATCCATGGCGCCGGCGGCGAGGGCGGAG 2 TATAAGCTTCTTGGCGTGTCCAGCCCACGGGGCGTAGAACTCGAC 3 ATAAAGCTTGCTCCAGAGTATGAGAAAGCT 4 ATACTCGAGGAGCTCATCCTTGAGAGGCTC
Supplemental Table 1. List of PCR primers Primer Sequences (5 to 3 ) 1 TATCCATGGCGCCGGCGGCGAGGGCGGAG 2 TATAAGCTTCTTGGCGTGTCCAGCCCACGGGGCGTAGAACTCGAC 3 ATAAAGCTTGCTCCAGAGTATGAGAAAGCT 4 ATACTCGAGGAGCTCATCCTTGAGAGGCTC
Supplemental Data. Huo et al. (2013). Plant Cell /tpc
 Supplemental Data. Huo et al. (2013). Plant Cell 10.1105/tpc.112.108902 1 Supplemental Figure 1. Alignment of NCED4 Amino Acid Sequences from Different Lettuce Varieties. NCED4 amino acid sequences are
Supplemental Data. Huo et al. (2013). Plant Cell 10.1105/tpc.112.108902 1 Supplemental Figure 1. Alignment of NCED4 Amino Acid Sequences from Different Lettuce Varieties. NCED4 amino acid sequences are
Supplemental materials
 Supplemental materials Materials and methods for supplemental figures Yeast two-hybrid assays TAP46-PP2Ac interactions I. The TAP46 was used as the bait and the full-length cdnas of the five C subunits
Supplemental materials Materials and methods for supplemental figures Yeast two-hybrid assays TAP46-PP2Ac interactions I. The TAP46 was used as the bait and the full-length cdnas of the five C subunits
Supplemental Data. Osakabe et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. Phylogenetic analysis of KUP in various species. The amino acid sequences of KUPs from green algae and land plants were identified with a BLAST search and aligned using ClustalW
Supplemental Figure 1. Phylogenetic analysis of KUP in various species. The amino acid sequences of KUPs from green algae and land plants were identified with a BLAST search and aligned using ClustalW
0.5% Sucrose. Supplemental Data. Chao et al. (2011). Plant Cell /tpc Col-0 tsc10a-2. Root length (mm)
 A 0% 0.5% 1% Col-0 tsc10a-2 B Root length (mm) 35 30 25 20 15 10 5 0 0% 0.50% 1% 2% 4% Sucrose concentration 2% 4% Sucrose Col-0 tsc10a-2 Supplemental Figure 1. The effect of sucrose on root growth of
A 0% 0.5% 1% Col-0 tsc10a-2 B Root length (mm) 35 30 25 20 15 10 5 0 0% 0.50% 1% 2% 4% Sucrose concentration 2% 4% Sucrose Col-0 tsc10a-2 Supplemental Figure 1. The effect of sucrose on root growth of
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11070 Supplementary Figure 1 Purification of FLAG-tagged proteins. a, Purification of FLAG-RNF12 by FLAG-affinity from nuclear extracts of wild-type (WT) and two FLAG- RNF12 transgenic
doi:10.1038/nature11070 Supplementary Figure 1 Purification of FLAG-tagged proteins. a, Purification of FLAG-RNF12 by FLAG-affinity from nuclear extracts of wild-type (WT) and two FLAG- RNF12 transgenic
Supplementary Information
 Supplementary Information Supplementary Fig. 1. Seed dormancy and germination responses of RVE1 and PIF1. (a) Diagram of RVE1 and the T-DNA insertion of the rve1-2 mutant (SAIL_326_A01). Black boxes represent
Supplementary Information Supplementary Fig. 1. Seed dormancy and germination responses of RVE1 and PIF1. (a) Diagram of RVE1 and the T-DNA insertion of the rve1-2 mutant (SAIL_326_A01). Black boxes represent
DAY 1 DAY 2 DAY 3. GUS activity. Time h (ZT) Cluster 1. Cluster 2 Cluster 3. Cluster 4. Cluster 5. Cluster 6
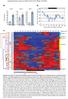 Supplemental Data. Giraud et al. (21). Plant Cell 1.115/tpc.11.74518 A) B) 6 5 4 3 2 1 DAY 1 2 15 1 5 IP SITE II DAY 2 7 6 5 4 3 2 1 IP SITE II DAY 3 IP SITE II ealtive transcript abundance 12 1 75 5 25
Supplemental Data. Giraud et al. (21). Plant Cell 1.115/tpc.11.74518 A) B) 6 5 4 3 2 1 DAY 1 2 15 1 5 IP SITE II DAY 2 7 6 5 4 3 2 1 IP SITE II DAY 3 IP SITE II ealtive transcript abundance 12 1 75 5 25
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Supplemental Data. Dai et al. (2013). Plant Cell /tpc Absolute FyPP3. Absolute
 A FyPP1 Absolute B FyPP3 Absolute Dry seeds Imbibed 24 hours Dry seeds Imbibed 24 hours C ABI5 Absolute Dry seeds Imbibed 24 hours Supplemental Figure 1. Expression of FyPP1, FyPP3 and ABI5 during seed
A FyPP1 Absolute B FyPP3 Absolute Dry seeds Imbibed 24 hours Dry seeds Imbibed 24 hours C ABI5 Absolute Dry seeds Imbibed 24 hours Supplemental Figure 1. Expression of FyPP1, FyPP3 and ABI5 during seed
- PDI5. - Actin A 300-UTR5. PDI5 lines WT. Arabidopsis Chromosome 1. PDI5 (At1g21750) SALK_ SALK_ SALK_015253
 Supplemental Data. Ondzighi et al. (2008). Arabidopsis Protein Disulfide Isomerase-5 Inhibits ysteine Proteases During Trafficking to Vacuoles Prior to Programmed ell Death of the dothelium in Developing
Supplemental Data. Ondzighi et al. (2008). Arabidopsis Protein Disulfide Isomerase-5 Inhibits ysteine Proteases During Trafficking to Vacuoles Prior to Programmed ell Death of the dothelium in Developing
Design. Construction. Characterization
 Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Supplemental Data. Rodrigues et al. (2013). Plant Cell /tpc L-W+A+H -L-W-A-H * * *
 A Bait -L-W+A+H -L-W-A-H Prey Rodrigues_Fig S1 BD-SnRK1.1 BD-SnRK1.1-CD BD-SnRK1.1-RD BD-SnRK1.1 ΔKA1 BD-SnRK1.1 KA1 BD BD AD AD AD AD AD AD-ABI1 AD-PP2CA B MW kda 75 50 37 * * * α MYC 75 50 37 * * * α
A Bait -L-W+A+H -L-W-A-H Prey Rodrigues_Fig S1 BD-SnRK1.1 BD-SnRK1.1-CD BD-SnRK1.1-RD BD-SnRK1.1 ΔKA1 BD-SnRK1.1 KA1 BD BD AD AD AD AD AD AD-ABI1 AD-PP2CA B MW kda 75 50 37 * * * α MYC 75 50 37 * * * α
State Key Laboratory of Plant Physiology and Biochemistry, College of Biological Sciences, China Agricultural University, Beijing , China d
 The Plant Cell, Vol. 22: 2353 2369, July 2010, www.plantcell.org ã 2010 American Society of Plant Biologists Arabidopsis Cockayne Syndrome A-Like Proteins 1A and 1B Form a Complex with CULLIN4 and Damage
The Plant Cell, Vol. 22: 2353 2369, July 2010, www.plantcell.org ã 2010 American Society of Plant Biologists Arabidopsis Cockayne Syndrome A-Like Proteins 1A and 1B Form a Complex with CULLIN4 and Damage
Supplemental Data. Benstein et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. Purification of the heterologously expressed PGDH1, PGDH2 and PGDH3 enzymes by Ni-NTA affinity chromatography. Protein extracts (2 µl) of different fractions (lane 1 = total extract,
Supplemental Figure 1. Purification of the heterologously expressed PGDH1, PGDH2 and PGDH3 enzymes by Ni-NTA affinity chromatography. Protein extracts (2 µl) of different fractions (lane 1 = total extract,
Supplemental Fig. S1. Key to underlines: Key to amino acids:
 AspA-F1 AspA 1 MKQMETKGYGYFRKTKAYGLVCGIT--------------LAGALTLGTTSVSADDVTTLNPATNLTTLQTPPTADQTQLAHQAGQQSGELVSEVSNTEWD 86 SspB 1 MQKREV--FG-FRKSKVAKTLCGAV-LGAALIAIADQQVLADEVTETNSTANVAVTTTGNPATNLPEAQGEATEAASQSQAQAGSKDGALPVEVSADDLN
AspA-F1 AspA 1 MKQMETKGYGYFRKTKAYGLVCGIT--------------LAGALTLGTTSVSADDVTTLNPATNLTTLQTPPTADQTQLAHQAGQQSGELVSEVSNTEWD 86 SspB 1 MQKREV--FG-FRKSKVAKTLCGAV-LGAALIAIADQQVLADEVTETNSTANVAVTTTGNPATNLPEAQGEATEAASQSQAQAGSKDGALPVEVSADDLN
Abcam.com. hutton.ac.uk. Ipmdss.dk. Bo Gong and Eva Chou
 Abcam.com Bo Gong and Eva Chou Ipmdss.dk hutton.ac.uk What is a homeotic gene? A gene which regulates the developmental fate of anatomical structures in an organism Why study them? Understand the underlying
Abcam.com Bo Gong and Eva Chou Ipmdss.dk hutton.ac.uk What is a homeotic gene? A gene which regulates the developmental fate of anatomical structures in an organism Why study them? Understand the underlying
To investigate the heredity of the WFP gene, we selected plants that were homozygous
 Supplementary information Supplementary Note ST-12 WFP allele is semi-dominant To investigate the heredity of the WFP gene, we selected plants that were homozygous for chromosome 1 of Nipponbare and heterozygous
Supplementary information Supplementary Note ST-12 WFP allele is semi-dominant To investigate the heredity of the WFP gene, we selected plants that were homozygous for chromosome 1 of Nipponbare and heterozygous
Supplemental Data. Borg et al. Plant Cell (2014) /tpc
 Supplementary Figure 1 - Alignment of selected angiosperm DAZ1 and DAZ2 homologs Multiple sequence alignment of selected DAZ1 and DAZ2 homologs. A consensus sequence built using default parameters is shown
Supplementary Figure 1 - Alignment of selected angiosperm DAZ1 and DAZ2 homologs Multiple sequence alignment of selected DAZ1 and DAZ2 homologs. A consensus sequence built using default parameters is shown
Supplemental Figure 1
 Supplemental Figure 1 A LK sls1 lks1-2 F 1 sls1 B LK sls1 lks1-2 F 1 lks1-2 sls1 F 1 lks1-2 sls1 F 2 Col lks1-2 Col+LKS1 Col Col+LKS1 Supplemental Figure 1. Genetic analysis of sls1 mutant. (A) and (B)
Supplemental Figure 1 A LK sls1 lks1-2 F 1 sls1 B LK sls1 lks1-2 F 1 lks1-2 sls1 F 1 lks1-2 sls1 F 2 Col lks1-2 Col+LKS1 Col Col+LKS1 Supplemental Figure 1. Genetic analysis of sls1 mutant. (A) and (B)
Before starting, write your name on the top of each page Make sure you have all pages
 Biology 105: Introduction to Genetics Name Student ID Before starting, write your name on the top of each page Make sure you have all pages You can use the back-side of the pages for scratch, but we will
Biology 105: Introduction to Genetics Name Student ID Before starting, write your name on the top of each page Make sure you have all pages You can use the back-side of the pages for scratch, but we will
Supplemental Data. Wu et al. (2). Plant Cell..5/tpc RGLG Hormonal treatment H2O B RGLG µm ABA µm ACC µm GA Time (hours) µm µm MJ µm IA
 Supplemental Data. Wu et al. (2). Plant Cell..5/tpc..4. A B Supplemental Figure. Immunoblot analysis verifies the expression of the AD-PP2C and BD-RGLG proteins in the Y2H assay. Total proteins were extracted
Supplemental Data. Wu et al. (2). Plant Cell..5/tpc..4. A B Supplemental Figure. Immunoblot analysis verifies the expression of the AD-PP2C and BD-RGLG proteins in the Y2H assay. Total proteins were extracted
Biotechnology and DNA Technology
 11/27/2017 PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College CHAPTER 9 Biotechnology and DNA Technology Introduction to Biotechnology Learning Objectives Compare
11/27/2017 PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College CHAPTER 9 Biotechnology and DNA Technology Introduction to Biotechnology Learning Objectives Compare
PHT1;2-CFP YFP-PHF + PHT1;2-CFP YFP-PHF
 YFP-PHF1 CFP-PHT1;2 PHT1;2-CFP YFP-PHF + PHT1;2-CFP YFP-PHF + CFP-PHT1;2 Negative control!-gfp Supplemental Figure 1: PHT1;2 accumulation is PHF1 dependent. Immunoblot analysis on total protein extract
YFP-PHF1 CFP-PHT1;2 PHT1;2-CFP YFP-PHF + PHT1;2-CFP YFP-PHF + CFP-PHT1;2 Negative control!-gfp Supplemental Figure 1: PHT1;2 accumulation is PHF1 dependent. Immunoblot analysis on total protein extract
BSCI410-Liu/Spring 06 Exam #1 Feb. 23, 06
 Your Name: Your UID# 1. (20 points) Match following mutations with corresponding mutagens (X-RAY, Ds transposon excision, UV, EMS, Proflavin) a) Thymidine dimmers b) Breakage of DNA backbone c) Frameshift
Your Name: Your UID# 1. (20 points) Match following mutations with corresponding mutagens (X-RAY, Ds transposon excision, UV, EMS, Proflavin) a) Thymidine dimmers b) Breakage of DNA backbone c) Frameshift
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1. Validation of CDK9-inhibitor treatment.
 Supplementary Figure 1 Validation of CDK9-inhibitor treatment. (a) Schematic of GAPDH with the middle of the amplicons indicated in base pairs. The transcription start site (TSS) and the terminal polyadenylation
Supplementary Figure 1 Validation of CDK9-inhibitor treatment. (a) Schematic of GAPDH with the middle of the amplicons indicated in base pairs. The transcription start site (TSS) and the terminal polyadenylation
Chapter 20 Recombinant DNA Technology. Copyright 2009 Pearson Education, Inc.
 Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
6/256 1/256 0/256 1/256 2/256 7/256 10/256. At3g06290 (SAC3B)
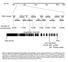 Chr.III M 5M 1M 15M 2M 23M BAC clones F22F7 F1A16 F24F17 F24P17 T8E24 F17A9 F21O3 F17A17 F18C1 F2O1 F28L1 F5E6 F3E22 T1B9 MLP3 Number of recombinants 6/256 1/256 /256 1/256 2/256 7/256 1/256 At3g629 (SAC3B)
Chr.III M 5M 1M 15M 2M 23M BAC clones F22F7 F1A16 F24F17 F24P17 T8E24 F17A9 F21O3 F17A17 F18C1 F2O1 F28L1 F5E6 F3E22 T1B9 MLP3 Number of recombinants 6/256 1/256 /256 1/256 2/256 7/256 1/256 At3g629 (SAC3B)
Intron distance to transcription start (nt)*
 Schwab et al. SOM: Enhanced microrna accumulation through stemloop-adjacent introns 1 Supplemental Table 1. Characteristics of introns used in this study Intron name Length intron (nt) Length cloned fragment
Schwab et al. SOM: Enhanced microrna accumulation through stemloop-adjacent introns 1 Supplemental Table 1. Characteristics of introns used in this study Intron name Length intron (nt) Length cloned fragment
Supplementary Fig 1. The responses of ERF109 to different hormones and stresses. (a to k) The induced expression of ERF109 in 7-day-old Arabidopsis
 Supplementary Fig 1. The responses of ERF109 to different hormones and stresses. (a to k) The induced expression of ERF109 in 7-day-old Arabidopsis seedlings expressing ERF109pro-GUS. The GUS staining
Supplementary Fig 1. The responses of ERF109 to different hormones and stresses. (a to k) The induced expression of ERF109 in 7-day-old Arabidopsis seedlings expressing ERF109pro-GUS. The GUS staining
At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in
 Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
previously. co and fca-9 alleles were isolated from the Col background. Plants were
 Supporting Online Material Materials and Methods Plant Growth Conditions The flc-3, FRI Sf2 -Col (S1), fve-4 (S2), ld-1 (S3) and fpa (S4) lines were described previously. co and fca-9 alleles were isolated
Supporting Online Material Materials and Methods Plant Growth Conditions The flc-3, FRI Sf2 -Col (S1), fve-4 (S2), ld-1 (S3) and fpa (S4) lines were described previously. co and fca-9 alleles were isolated
Expression constructs used in this study: Rtn1pQM-Ntag/A. The expression vector was generated as described previously (Mannan et al., 2006).
 Supplementary file 1 Sequences of primers used: C-mycTag-new-F: 5 CCGAATTCCGTCGGAGCAAGCTTGATTTA3 Ad.C-mycTag-R: 5 TTGCGGCCGCCTTTTGCTCCATGGTGAGGTC3 adpxt1-orf-f: 5 GGTGCGGCCGCTGGTGGCATGCAGCTTAGACACATTGG3
Supplementary file 1 Sequences of primers used: C-mycTag-new-F: 5 CCGAATTCCGTCGGAGCAAGCTTGATTTA3 Ad.C-mycTag-R: 5 TTGCGGCCGCCTTTTGCTCCATGGTGAGGTC3 adpxt1-orf-f: 5 GGTGCGGCCGCTGGTGGCATGCAGCTTAGACACATTGG3
SUPPLEMENTARY EXPEMENTAL PROCEDURES
 SUPPLEMENTARY EXPEMENTAL PROCEDURES Plasmids- Total RNAs were extracted from HeLaS3 cells and reverse-transcribed using Superscript III Reverse Transcriptase (Invitrogen) to obtain DNA template for the
SUPPLEMENTARY EXPEMENTAL PROCEDURES Plasmids- Total RNAs were extracted from HeLaS3 cells and reverse-transcribed using Superscript III Reverse Transcriptase (Invitrogen) to obtain DNA template for the
Learning Objectives :
 Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
Learning Objectives : Understand the basic differences between genomic and cdna libraries Understand how genomic libraries are constructed Understand the purpose for having overlapping DNA fragments in
