Transcription Activators in Eukaryotes. Chapter 12
|
|
|
- Madison Quinn
- 5 years ago
- Views:
Transcription
1 Transcription Activators in Eukaryotes Chapter 12
2 Techniques Run-off transcription Primer extension Quantitative S1 mapping
3
4 Modular Protein domain is an independingly folding region. DA binding domain Transcription-activation domain Dimerization domain most do not bind without dimerization homodimers, heterodimers homodimers, heterodimers, tetramers (rare) some bind small molecules (eg. steroid hormones).
5 C Gal4 Regulation of gal regulon in yeast C GC4 Regulation of amino acid biosynthesis in yeast C C GR SP1 glucocorticoid receptor: binds hormone and then binds DA to alter gene expression Activation domain DA binding domain From Molecular Cell Biology 3rd edition, Lodish et al Scientific American Books 1995 general upstream activator of pol II genes binds the GC box. protein domain independently folds It has been estimated that 8-10% of the proteins coded in the genome are specifically for regulating the transcription of other proteins, yielding more than 1000 in Drosophila alone (Tupler et al., 2001; Adams et al., 2000). (Reference: Moon Draper Thesis, University of Texas at Austin 2005).
6 Activation domains acidic domains - acidic Gal4 Glutamine rich - Sp1 ~25% glutamine Proline-rich - CTF is >20% proline CTF/F-I binding sites are present in viral and cellular promoters and in the origin of DA replication of Adenovirus type 2.
7 Review. Most TF use major groove.
8 O O C 3 T A Major Groove Minor Groove O O C 3 To 1' position of deoxyribose To 1' position of deoxyribose T A Major Groove Minor Groove O O To 1' position of deoxyribose To 1' position of deoxyribose C G Major Groove Minor Groove O O G C Major Groove Minor Groove Most TF use major groove.
9 DA binding domains Zinc containing domains <- eukaryotic omeodomains <- eukaryotic bzip bl <- eukaryotic
10 Zinc Fingers
11 C22 Zinc Finger turn 2 anti-parallel β-sheets β-sheet β-sheet alpha helix EXCEPTIO Don t need to dimerize. All are in the major groove Examples are TFIIIA & zif268 Basic residues on ʻoutsideʼ of alpha helix. These are used to interact/read/bind the DA. B-strands interact with DA backbone and position the alpha helix so that it can read the DA. Abt 23 AA. Characterized by the sequence CX2-4C...X2-4. <-- consensus may be incorrect. eed to verify. Fig 12.2 and 12.3 Zinc Fingers
12 C6 Zinc containing TF 2 zinc atoms Dimer alpha helix dimerization domain -terminus DA binding domain with two Zns consensus sequence CX2CX6CX5-6CX2CX6C example Gal4 Zinc Fingers consensus sequence CX2CX6CX5-6CX2CX6C Top pictures are from your textbook. Bottom one is from
13 Gal4
14 uclear Receptor C4 Zn finger uclear receptors ormones diffuse through the cell membrane enter the nucleus and bind these receptors. The enhancers that they bind are sometimes called ormone Response Elements. Each one binds different small molecules eg. Androgens, estrogens, progesterone, glucocorticoids, vitamin D, thyroid hormone, retinoic acid. Ligand binding domain is separate from DA binding domain. C4 zinc finger: Its consensus sequence is CX2CX13CX2CX14-15CX5CX9CX2C. The first four cysteine residues bind to a zinc ion and the last four cysteine residues bind to another zinc ion (Figure 4-F-2). Reference is DA and ligand binding domains are separate. Zinc Fingers
15 uclear Receptor C4 Zn finger All are ligand-dependent activators of transcription. ow do the activate transcription? Bind a co-activator complex. Includes CBP/p300 Acetylates histones. Some info from Genes VIII Lewin pg 646. C4 zinc finger: Its consensus sequence is CX2CX13CX2CX14-15CX5CX9CX2C. The first four cysteine residues bind to a zinc ion and the last four cysteine residues bind to another zinc ion (Figure 4-F-2). Reference is DA and ligand binding domains are separate. Zinc Fingers
16 uclear Receptors Three general types
17 Type I ormonal glucocorticoid receptor androgen receptor mineralcorticoid receptor progesterone receptor Binds as a dimer. Zinc Fingers Book describes as Type I. Move from cytoplasm to nucleus. The GRE is 2 short half sites. One Zn finger does DA binding the other does dimerization! The dimerization determines what the spacing between the half-sites should be. omodimers recognize palindromic sites. (Genes VIII, Lewin pg 646) The major hormones secreted by the adrenal cortex are cortisol, aldosterone and dehydroepandrosterone (13)...Aldosterone, the major mineralocorticoid of the body, stimulates sodium reabsorption and potassium excretion by the kidney. (Ref: Spinal Cord Medicine 2003).
18 Type I ormonal hsp90 GRE Binds as a dimer. Zinc Fingers Book describes as Type I. Move from cytoplasm to nucleus. The GRE is 2 short half sites. One Zn finger does DA binding the other does dimerization! The dimerization determines what the spacing between the half-sites should be. omodimers recognize palindromic sites. (Genes VIII, Lewin pg 646) GR can activate or repress: Furthermore, we show that GR acts both as a direct inhibitor of CREB binding protein (CBP)-associated AT activity and also by recruiting DAC2 to the p65- CBP AT complex. Glucocorticoid Receptor Recruitment of istone Deacetylase 2 Inhibits Interleukin-1-Induced istone 4 Acetylation on Lysines 8 and 12 Kazuhiro Ito, Peter J. Barnes, and Ian M. Adcock*
19 retinoic acid receptor X Type II ormonal Thyroid hormone receptor heterodimer is active Core istones are the target. GRE can be several kb up or downstream of the promoter. Deacetylase Without thyroid hormone it is a repressor istone acetyl transferases thyroid hormone receptors & relatives. Book describes as Type II. Stay in nucleus. Retinoic Acid (Vitamen A) important for development in chicks of anterior posterior axis in embryogenesis. These act as receptors in the same way that the laci repressor is a receptor. Zinc Fingers
20 Type III Orphan Receptors - one whose ligand has not been found. Zinc Fingers An orphan receptor is one whose ligand has not been found.
21 omeotic Genes in Drosophila These are transcription factors that act to determine the identify of specific tissue or body parts. Mis-expression can cause, during development, the incorrect body part or tissue to be produced in a particular position. omeodomain Some appear to bind as monomers, some as homodimers, and some as heterodimers. Dimerization domains such as leucine zippers might be found with some of these.
22 omeotic Genes in Drosophila Found in almost all eukaryotes The term homeodomain refers a 60 AA motif which is actually a specialized version of the T motif. 3 helixes. #2 & #3 like are related to the bacterial T motif ydrophobic interactionss hold each helix in position relative to its neighbor. Usually found near the C-terminus terminal part inserts into minor groove Some bind as monomers, some dimers, some both Some are repressors, some are activators omeodomain These are transcription factors that act to determine the identify of specific tissue or body parts. Mis-expression can cause, during development, the incorrect body part or tissue to be produced in a particular position. The term homeodomain refers a 60 AA motif which is actually a specialized version of the T motif. Three protein helixes are involved here. elix #2 and #3 represent the T part of the protein. ydrophobic interactshold each helix in position relative to its neihbor. omeodomain proteins bind as a monomer (Prokaryotic T binds as dimers)
23 Basic Zipper bzip 2 proteins each with a half zipper amphipathic alpha helix Binding and dimerization functions are combined in the same domain. Examples: CREB, Jun, Fos Targets for homodimer binding are inverted repeats with no separation. Examples are CREB ATF/CREB family of basic leucine zippers binds a DA motif called the camp response element which is abbreviated CRE. jun & fos jun forms homo or heterodimers fos forms only heterodimers together called AP1 and bind AP1 sites 10X better than jun homodimers. Lewin Genes VIII pg 652
24 Basic elix-loop-helix bl Two amphipathic helices causes dimerization Binding and dimerization functions are combined in the same domain. elix 1 also contains basic amino acids omodimers & heterodimers form MyoD, Myc and Max are examples MyoD is an example Figure is from page 351 of Weaver, 3rd edition. Amphipathic helix has one hydrophilic face and one hydrophobic face. It has been estimated that 8-10% of the proteins coded in the genome are specifically for regulating the transcription of other proteins, yielding more than 1000 in Drosophila alone (Tupler et al., 2001; Adams et al., 2000). (Reference: Moon Draper Thesis, University of Texas at Austin 2005).
25 Independence Without camp With camp CREB A B Gal4 A B ybrid A B CREB activation domain Gal4 DA binding domain A B C R E G A L O TRASCRIPTIO! O TRASCRIPTIO! A B C R E G A L TRASCRIPTIO! O TRASCRIPTIO! Without camp With camp A A B G A L TRASCRIPTIO! B G A L TRASCRIPTIO! Without camp With camp A B G A L C R E O TRASCRIPTIO! O TRASCRIPTIO! A B G A L C R E TRASCRIPTIO! O TRASCRIPTIO! Weaver gives a different but comparable example in Fig pg 331 4th edition. Weaver gives a different example in Fig pg 352 3rd edition.
26 An activation domain can recruit general transcription factors TFIID A TATA Inr B TFIID A Pol II F BTFIID B E A DAB complex Pol II TFIID F B A mediator Pol II F Pol II E B F Pol II E BTFIID F A Complete mediator oloenzyme mediator
27 Step by step recruitment or holoenzyme recruitment? pg Subtitle: Evidence of recruitment of the holoenzyme as a unit. First they describe exp that suggest it is a unit. Then they describe exp that suggest it is not. Other experiments indicate that it can happen both ways.
28 Figure ow does one quantify the components? TAP tagged RA polymerase II & Mediator Dot blot cell extract TAP has part of protein A IgG against peroxidase then add peroxidase and do a peroxidase enzymatic assay.
29 An activation domain can recruit general transcription factors TFIID TFIIB TFIIF TFII probably all of them are targets
30 Figure Stringer et al. TFIID VP16 activation domain fused to protein A. Protein A binds IgG. IgG column to immobilize ela nuclear extracts - enrietta Lackʼs cervical cancer cells Read about them here: <-very interesting. One is heated in a way that breaks TFIID Run off transcription of adenovirus late promoter IgG-VP16 removed essential ingredient ature 345: 784-? 1990 Take home acidic VP16 binds TFIID In the paper, we see that yeast TFIID can complement the depletion! Kill TFIID in an extract. Supplement with stuff from column. Specificity lies in what heat damages! VP16 activation domain fused to protein A. protein A binds IgG. IgG column to immobilize ela nuclear extracts One is heated in a way that breaks TFIID Run off transcription of adenovirus late promoter IgG-VP16 removed essential ingredient ature 345: 784-? 1990 Take home acidic VP16 binds TFIID
31 Run-off transcription
32 VP16 activation domain fused to protein A. Protein A binds IgG. IgG column to immobilize Run off transcription of adenovirus late promoter is used to measure transcription ela nuclear extracts - enrietta Lack s cervical cancer cells Read about them here: <-very interesting. uclear extract is passed through a IgG-VP16 column and some protein binds. Another sample is heated in a way that breaks TFIID. o transcription occurs. The stuff that bound the column complements the heat induced problem Take home acidic VP16 binds TFIID ature 345: 784-? In the paper, we see that yeast TFIID can complement the depletion!
33 With some promoters the acidic activation domain can act help stabilize TFIIB on the DA.
DNA Binding Domains: Structural Motifs. Effector Domain. Zinc Fingers. Zinc Fingers, continued. Zif268
 DNA Binding Domains: Structural Motifs Studies of known transcription factors have found several motifs of protein design to allow sequence-specific binding of DNA. We will cover only three of these motifs:
DNA Binding Domains: Structural Motifs Studies of known transcription factors have found several motifs of protein design to allow sequence-specific binding of DNA. We will cover only three of these motifs:
Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators
 Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators At least 5 potential gene expression control points Superfamily of Gene Regulators Activation of gene structure Initiation
Chapter 25: Regulating Eukaryotic Transcription The Ligand Responsive Activators At least 5 potential gene expression control points Superfamily of Gene Regulators Activation of gene structure Initiation
Structure/function relationship in DNA-binding proteins
 PHRM 836 September 22, 2015 Structure/function relationship in DNA-binding proteins Devlin Chapter 8.8-9 u General description of transcription factors (TFs) u Sequence-specific interactions between DNA
PHRM 836 September 22, 2015 Structure/function relationship in DNA-binding proteins Devlin Chapter 8.8-9 u General description of transcription factors (TFs) u Sequence-specific interactions between DNA
Molecular Biology (BIOL 4320) Exam #1 March 12, 2002
 Molecular Biology (BIOL 4320) Exam #1 March 12, 2002 Name KEY SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number.
Molecular Biology (BIOL 4320) Exam #1 March 12, 2002 Name KEY SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number.
Enhancers. Activators and repressors of transcription
 Enhancers Can be >50 kb away from the gene they regulate. Can be upstream from a promoter, downstream from a promoter, within an intron, or even downstream of the final exon of a gene. Are often cell type
Enhancers Can be >50 kb away from the gene they regulate. Can be upstream from a promoter, downstream from a promoter, within an intron, or even downstream of the final exon of a gene. Are often cell type
Transcription factors
 Atlas of Genetics and Cytogenetics in Oncology and Haematology Transcription factors I Introduction * II Initiation of transcription III Transcription factors family pdf version I Introduction III.1 Helix-Turn-Helix
Atlas of Genetics and Cytogenetics in Oncology and Haematology Transcription factors I Introduction * II Initiation of transcription III Transcription factors family pdf version I Introduction III.1 Helix-Turn-Helix
Eukaryotic transcription (II)
 Eukaryotic transcription (II) Transcription factors Prokaryote: Sigma factors they are similar to general transcription factor of eukaryotes, they help bring RNA pol to the promoter Transcription factors
Eukaryotic transcription (II) Transcription factors Prokaryote: Sigma factors they are similar to general transcription factor of eukaryotes, they help bring RNA pol to the promoter Transcription factors
Overlap with exam 1 begins
 Exam 2 begins here Overlap with exam 1 begins Another groups asks with what does Gal4- VP16 interact? DNA attached to agarose beads Now add components in steps Ask if the recruitment of a component requires
Exam 2 begins here Overlap with exam 1 begins Another groups asks with what does Gal4- VP16 interact? DNA attached to agarose beads Now add components in steps Ask if the recruitment of a component requires
Another groups asks with what does Gal4- VP16 interact?
 Another groups asks with what does Gal4- VP16 interact? DNA attached to agarose beads Now add components in steps Ask if the recruitment of a component requires that Gal4-VP16 be present. Primer extension
Another groups asks with what does Gal4- VP16 interact? DNA attached to agarose beads Now add components in steps Ask if the recruitment of a component requires that Gal4-VP16 be present. Primer extension
Differential Gene Expression
 Developmental Biology Biology 4361 Differential Gene Expression October 13, 2005 core transcription initiation site 5 promoter 3 TATAT +1 upstream downstream Basal transcription factors (eukaryotes) TFIID
Developmental Biology Biology 4361 Differential Gene Expression October 13, 2005 core transcription initiation site 5 promoter 3 TATAT +1 upstream downstream Basal transcription factors (eukaryotes) TFIID
Synthetic cells: do bacteria need all its genes? No.
 NO NEED TO REFER TO THE SLIDES. بسم هللا الرحمن الرحيم Do we need all the non coding regions of the DNA? Two weeks ago, they discovered that the genome of a plant is very small (recall that plant genome
NO NEED TO REFER TO THE SLIDES. بسم هللا الرحمن الرحيم Do we need all the non coding regions of the DNA? Two weeks ago, they discovered that the genome of a plant is very small (recall that plant genome
Gene expression DNA RNA. Protein DNA. Replication. Initiation Elongation Processing Export. DNA RNA Protein. Transcription. Degradation.
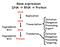 Gene expression DNA RNA Protein DNA DNA Degradation RNA Degradation Protein Replication Transcription Translation Initiation Elongation Processing Export Initiation Elongation Processing Targeting Chapter
Gene expression DNA RNA Protein DNA DNA Degradation RNA Degradation Protein Replication Transcription Translation Initiation Elongation Processing Export Initiation Elongation Processing Targeting Chapter
Regulation of gene expression. (Lehninger pg )
 Regulation of gene expression (Lehninger pg. 1072-1085) Today s lecture Gene expression Constitutive, inducible, repressible genes Specificity factors, activators, repressors Negative and positive gene
Regulation of gene expression (Lehninger pg. 1072-1085) Today s lecture Gene expression Constitutive, inducible, repressible genes Specificity factors, activators, repressors Negative and positive gene
Gene regulation V Biochemistry 302. March 6, 2006
 Gene regulation V Biochemistry 302 March 6, 2006 Common structural motifs associated with transcriptional regulatory proteins Helix-turn-helix Prokaryotic repressors and activators Eukaryotic homeodomain
Gene regulation V Biochemistry 302 March 6, 2006 Common structural motifs associated with transcriptional regulatory proteins Helix-turn-helix Prokaryotic repressors and activators Eukaryotic homeodomain
Chapter 14 Regulation of Transcription
 Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Eukaryotic Gene Expression Prof. P. N. Rangarajan Department of Biochemistry Indian Institute of Science, Bangalore
 Eukaryotic Gene Expression Prof. P. N. Rangarajan Department of Biochemistry Indian Institute of Science, Bangalore Lecture No. # 06 Eukaryotic Transcription Factors: Transcription Activation Domains In
Eukaryotic Gene Expression Prof. P. N. Rangarajan Department of Biochemistry Indian Institute of Science, Bangalore Lecture No. # 06 Eukaryotic Transcription Factors: Transcription Activation Domains In
CHAPTER 18 LECTURE NOTES: CONTROL OF GENE EXPRESSION PART B: CONTROL IN EUKARYOTES
 CHAPTER 18 LECTURE NOTES: CONTROL OF GENE EXPRESSION PART B: CONTROL IN EUKARYOTES I. Introduction A. No operon structures in eukaryotes B. Regulation of gene expression is frequently tissue specific.
CHAPTER 18 LECTURE NOTES: CONTROL OF GENE EXPRESSION PART B: CONTROL IN EUKARYOTES I. Introduction A. No operon structures in eukaryotes B. Regulation of gene expression is frequently tissue specific.
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
Biology 4361 Developmental Biology Differential Gene Expression September 28, 2006 Chromatin Structure ~140 bp ~60 bp Transcriptional Regulation: 1. Packing prevents access CH 3 2. Acetylation ( C O )
CELL BIOLOGY - CLUTCH CH. 7 - GENE EXPRESSION.
 !! www.clutchprep.com CONCEPT: CONTROL OF GENE EXPRESSION BASICS Gene expression is the process through which cells selectively to express some genes and not others Every cell in an organism is a clone
!! www.clutchprep.com CONCEPT: CONTROL OF GENE EXPRESSION BASICS Gene expression is the process through which cells selectively to express some genes and not others Every cell in an organism is a clone
Differential Gene Expression
 Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Biology 4361 Developmental Biology Differential Gene Expression June 19, 2008 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
- all cells express large number of the same genes - housekeeping or common genes - many cells also express cell-type-specific genes
 Developmental Biology - Biology 4361 Lecture 8 - Differential Gene Expression October 13, 2005 The principle of genomic equivalence states that all cells in a developing organism have the same genetic
Developmental Biology - Biology 4361 Lecture 8 - Differential Gene Expression October 13, 2005 The principle of genomic equivalence states that all cells in a developing organism have the same genetic
Eukaryotic Transcription
 Eukaryotic Transcription I. Differences between eukaryotic versus prokaryotic transcription. II. (core vs holoenzyme): RNA polymerase II - Promotor elements. - General Pol II transcription factors (GTF).
Eukaryotic Transcription I. Differences between eukaryotic versus prokaryotic transcription. II. (core vs holoenzyme): RNA polymerase II - Promotor elements. - General Pol II transcription factors (GTF).
Mechanism of action-1
 Mechanism of action-1 receptors: mediators of hormone action, membrane associated vs. intracellular receptors: measurements of receptor - ligand interaction, mechanism surface-receptors: kinases, phosphatases,
Mechanism of action-1 receptors: mediators of hormone action, membrane associated vs. intracellular receptors: measurements of receptor - ligand interaction, mechanism surface-receptors: kinases, phosphatases,
Chem 465 Biochemistry II
 Chem 465 Biochemistry II Name: 2 points Multiple choice (4 points apiece): 1. Which of the following is not true of trna molecules? A) The 3'-terminal sequence is -CCA. B) Their anticodons are complementary
Chem 465 Biochemistry II Name: 2 points Multiple choice (4 points apiece): 1. Which of the following is not true of trna molecules? A) The 3'-terminal sequence is -CCA. B) Their anticodons are complementary
Chapter 9-II - Transcriptional Control of Gene Expression
 Chapter 9-II - Transcriptional Control of Gene Expression Transcriptional Control of Gene Expression 9.3 RNA Polymerase II Promoters and General Transcription Factors Three types of promoter sequences
Chapter 9-II - Transcriptional Control of Gene Expression Transcriptional Control of Gene Expression 9.3 RNA Polymerase II Promoters and General Transcription Factors Three types of promoter sequences
Gene Expression and Regulation - 1
 Gene Expression and Regulation - 1 We have been discussing the molecular structure of DNA and its function in DNA replication and in transcription. Earlier we discussed how genes interact in transmission
Gene Expression and Regulation - 1 We have been discussing the molecular structure of DNA and its function in DNA replication and in transcription. Earlier we discussed how genes interact in transmission
Complex Transcription Machinery
 Complex Transcription Machinery Subunits of the Basal Txn Apparatus Stepwise Assembly of the Pre-initiation Complex Interplay of Activators, Co-regulators and RNA Polymerase at the Promoter Divide and
Complex Transcription Machinery Subunits of the Basal Txn Apparatus Stepwise Assembly of the Pre-initiation Complex Interplay of Activators, Co-regulators and RNA Polymerase at the Promoter Divide and
Transcription in Prokaryotes. Jörg Bungert, PhD Phone:
 Transcription in Prokaryotes Jörg Bungert, PhD Phone: 352-273-8098 Email: jbungert@ufl.edu Objectives Understand the basic mechanism of transcription. Know the function of promoter elements and associating
Transcription in Prokaryotes Jörg Bungert, PhD Phone: 352-273-8098 Email: jbungert@ufl.edu Objectives Understand the basic mechanism of transcription. Know the function of promoter elements and associating
GENES AND CHROMOSOMES V. Lecture 7. Biology Department Concordia University. Dr. S. Azam BIOL 266/
 1 GENES AND CHROMOSOMES V Lecture 7 BIOL 266/4 2014-15 Dr. S. Azam Biology Department Concordia University 2 CELL NUCLEUS AND THE CONTROL OF GENE EXPRESSION An Overview of Gene Regulation in Eukaryotes
1 GENES AND CHROMOSOMES V Lecture 7 BIOL 266/4 2014-15 Dr. S. Azam Biology Department Concordia University 2 CELL NUCLEUS AND THE CONTROL OF GENE EXPRESSION An Overview of Gene Regulation in Eukaryotes
DNA & DNA : Protein Interactions BIBC 100
 DNA & DNA : Protein Interactions BIBC 100 Sequence = Information Alphabet = language L,I,F,E LIFE DNA = DNA code A, T, C, G CAC=Histidine CAG=Glutamine GGG=Glycine Protein = Protein code 20 a.a. LIVE EVIL
DNA & DNA : Protein Interactions BIBC 100 Sequence = Information Alphabet = language L,I,F,E LIFE DNA = DNA code A, T, C, G CAC=Histidine CAG=Glutamine GGG=Glycine Protein = Protein code 20 a.a. LIVE EVIL
Chapter 17 Lecture. Concepts of Genetics. Tenth Edition. Regulation of Gene Expression in Eukaryotes
 Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Transcription factors
 Transcription factors R. de Martin Department of Vascular Biology and Thrombosis Research Medical University of Vienna K63 A20, TTP, XIAP IκBα K48 IL-6, Il-8, MCP-1, ICAM1, SELE, ciap1/2, A20, IκBα Adapted
Transcription factors R. de Martin Department of Vascular Biology and Thrombosis Research Medical University of Vienna K63 A20, TTP, XIAP IκBα K48 IL-6, Il-8, MCP-1, ICAM1, SELE, ciap1/2, A20, IκBα Adapted
Resources. This lecture Campbell and Farrell's Biochemistry, Chapter 11
 Transcription Resources This lecture Campbell and Farrell's Biochemistry, Chapter 11 2 Definition of a gene The entire nucleic acid sequence that is necessary for the synthesis of a functional polypeptide
Transcription Resources This lecture Campbell and Farrell's Biochemistry, Chapter 11 2 Definition of a gene The entire nucleic acid sequence that is necessary for the synthesis of a functional polypeptide
Eukaryotic & Prokaryotic Transcription. RNA polymerases
 Eukaryotic & Prokaryotic Transcription RNA polymerases RNA Polymerases A. E. coli RNA polymerase 1. core enzyme = ββ'(α)2 has catalytic activity but cannot recognize start site of transcription ~500,000
Eukaryotic & Prokaryotic Transcription RNA polymerases RNA Polymerases A. E. coli RNA polymerase 1. core enzyme = ββ'(α)2 has catalytic activity but cannot recognize start site of transcription ~500,000
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs
 Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
Chapter 8 DNA Recognition in Prokaryotes by Helix-Turn-Helix Motifs 1. Helix-turn-helix proteins 2. Zinc finger proteins 3. Leucine zipper proteins 4. Beta-scaffold factors 5. Others λ-repressor AND CRO
Differential Gene Expression
 Biology 4361 - Developmental Biology Differential Gene Expression June 18, 2009 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Biology 4361 - Developmental Biology Differential Gene Expression June 18, 2009 Differential Gene Expression Overview Chromatin structure Gene anatomy RNA processing and protein production Initiating transcription:
Mechanism of action-1
 Mechanism of action-1 receptors: mediators of hormone action, membrane associated vs. intracellular receptors: measurements of receptor - ligand interactions, regulation mechanisms surface - receptors:
Mechanism of action-1 receptors: mediators of hormone action, membrane associated vs. intracellular receptors: measurements of receptor - ligand interactions, regulation mechanisms surface - receptors:
30 Gene expression: Transcription
 30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
30 Gene expression: Transcription Gene structure. o Exons coding region of DNA. o Introns non-coding region of DNA. o Introns are interspersed between exons of a single gene. o Promoter region helps enzymes
Hmwk # 8 : DNA-Binding Proteins : Part II
 The purpose of this exercise is : Hmwk # 8 : DNA-Binding Proteins : Part II 1). to examine the case of a tandem head-to-tail homodimer binding to DNA 2). to view a Zn finger motif 3). to consider the case
The purpose of this exercise is : Hmwk # 8 : DNA-Binding Proteins : Part II 1). to examine the case of a tandem head-to-tail homodimer binding to DNA 2). to view a Zn finger motif 3). to consider the case
M1 - Biochemistry. Nucleic Acid Structure II/Transcription I
 M1 - Biochemistry Nucleic Acid Structure II/Transcription I PH Ratz, PhD (Resources: Lehninger et al., 5th ed., Chapters 8, 24 & 26) 1 Nucleic Acid Structure II/Transcription I Learning Objectives: 1.
M1 - Biochemistry Nucleic Acid Structure II/Transcription I PH Ratz, PhD (Resources: Lehninger et al., 5th ed., Chapters 8, 24 & 26) 1 Nucleic Acid Structure II/Transcription I Learning Objectives: 1.
Old EXAM 1 BIO409/509 NAME. Please number your answers and write them on the attached, lined paper.
 Old EXAM 1 BIO409/509 NAME Please number your answers and write them on the attached, lined paper. 1) Describe euchromatin and heterochromatin. Which form of chromatin would the insulin gene be found in
Old EXAM 1 BIO409/509 NAME Please number your answers and write them on the attached, lined paper. 1) Describe euchromatin and heterochromatin. Which form of chromatin would the insulin gene be found in
Nucleic Acid Structure. Nucleic Acid Sequence Abbreviations. Sequence Abbreviations, con t.
 BC 4054 Spring 2001 Chapter 11 & 12 Review Lecture otes Slide 1 ucleic Acid Structure Linear polymer of nucleotides Phosphodiester linkage between 3 and 5 positions See Figure 11.17 Slide 2 ucleic Acid
BC 4054 Spring 2001 Chapter 11 & 12 Review Lecture otes Slide 1 ucleic Acid Structure Linear polymer of nucleotides Phosphodiester linkage between 3 and 5 positions See Figure 11.17 Slide 2 ucleic Acid
32 Gene regulation in Eukaryotes Lecture Outline 11/28/05. Gene Regulation in Prokaryotes and Eukarykotes
 3 Gene regulation in Eukaryotes Lecture Outline /8/05 Gene regulation in eukaryotes Chromatin remodeling More kinds of control elements Promoters, Enhancers, and Silencers Combinatorial control Cell-specific
3 Gene regulation in Eukaryotes Lecture Outline /8/05 Gene regulation in eukaryotes Chromatin remodeling More kinds of control elements Promoters, Enhancers, and Silencers Combinatorial control Cell-specific
Chromatographic Separation of the three forms of RNA Polymerase II.
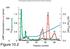 Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Chromatographic Separation of the three forms of RNA Polymerase II. α-amanitin α-amanitin bound to Pol II Function of the three enzymes. Yeast Pol II. RNA Polymerase Subunit Structures 10-7 Subunit structure.
Transcription Regulation And Gene Expression in Eukaryotes FS 2016 Graduate Course G2 P Matthias and RG Clerc Pharmazentrum Hörsaal 2 16h15-18h00
 Transcription Regulation And Gene Expression in Eukaryotes FS 2016 Graduate Course G2 P Matthias and RG Clerc Pharmazentrum Hörsaal 2 16h15-18h00 MODULAR STRUCTURE OF UPSTREAM TRANSCRIPTION FACTORS Experimental
Transcription Regulation And Gene Expression in Eukaryotes FS 2016 Graduate Course G2 P Matthias and RG Clerc Pharmazentrum Hörsaal 2 16h15-18h00 MODULAR STRUCTURE OF UPSTREAM TRANSCRIPTION FACTORS Experimental
Section C: The Control of Gene Expression
 Section C: The Control of Gene Expression 1. Each cell of a multicellular eukaryote expresses only a small fraction of its genes 2. The control of gene expression can occur at any step in the pathway from
Section C: The Control of Gene Expression 1. Each cell of a multicellular eukaryote expresses only a small fraction of its genes 2. The control of gene expression can occur at any step in the pathway from
Introduction to Bioinformatics Online Course: IBT
 Introduction to Bioinformatics Online Course: IBT Multiple Sequence Alignment Building Multiple Sequence Alignment Lec5: Interpreting your MSA Using Logos Using Logos - Logos are a terrific way to generate
Introduction to Bioinformatics Online Course: IBT Multiple Sequence Alignment Building Multiple Sequence Alignment Lec5: Interpreting your MSA Using Logos Using Logos - Logos are a terrific way to generate
From mechanism to medicne
 From mechanism to medicne a look at proteins and drug design Chem 342 δ δ δ+ M 2009 δ+ δ+ δ M Drug Design - an Iterative Approach @ DSU Structural Analysis of Receptor Structural Analysis of Ligand-Receptor
From mechanism to medicne a look at proteins and drug design Chem 342 δ δ δ+ M 2009 δ+ δ+ δ M Drug Design - an Iterative Approach @ DSU Structural Analysis of Receptor Structural Analysis of Ligand-Receptor
Control of Gene Expression
 Control of Gene Expression 1 How Gene Regulation Works 2 Control of Gene Expression Controlling gene expression is often accomplished by controlling transcription initiation Regulatory proteins bind to
Control of Gene Expression 1 How Gene Regulation Works 2 Control of Gene Expression Controlling gene expression is often accomplished by controlling transcription initiation Regulatory proteins bind to
Plant Molecular and Cellular Biology Lecture 9: Nuclear Genome Organization: Chromosome Structure, Chromatin, DNA Packaging, Mitosis Gary Peter
 Plant Molecular and Cellular Biology Lecture 9: Nuclear Genome Organization: Chromosome Structure, Chromatin, DNA Packaging, Mitosis Gary Peter 9/16/2008 1 Learning Objectives 1. List and explain how DNA
Plant Molecular and Cellular Biology Lecture 9: Nuclear Genome Organization: Chromosome Structure, Chromatin, DNA Packaging, Mitosis Gary Peter 9/16/2008 1 Learning Objectives 1. List and explain how DNA
Computational Biology I LSM5191 (2003/4)
 Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
Computational Biology I LSM5191 (2003/4) Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression I Flow of information: DNA to polypeptide DNA Start Exon1 Intron Exon2 Termination
Molecular Cell Biology - Problem Drill 01: Introduction to Molecular Cell Biology
 Molecular Cell Biology - Problem Drill 01: Introduction to Molecular Cell Biology Question No. 1 of 10 1. Which statement describes how an organism is organized from most simple to most complex? Question
Molecular Cell Biology - Problem Drill 01: Introduction to Molecular Cell Biology Question No. 1 of 10 1. Which statement describes how an organism is organized from most simple to most complex? Question
Chapter 24: Promoters and Enhancers
 Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
Chapter 24: Promoters and Enhancers A typical gene transcribed by RNA polymerase II has a promoter that usually extends upstream from the site where transcription is initiated the (#1) of transcription
Chromatin Structure and its Effects on Transcription
 Chromatin Structure and its Effects on Transcription Epigenetics 2014 by Nigel Atkinson The University of Texas at Austin From Weaver 4th edition and Armstrong 1st edition What is the point? DNA is not
Chromatin Structure and its Effects on Transcription Epigenetics 2014 by Nigel Atkinson The University of Texas at Austin From Weaver 4th edition and Armstrong 1st edition What is the point? DNA is not
The Little Things About the Little Things Inside of Us The Eukaryotic Genome and Its Expression
 The Little Things About the Little Things Inside of Us The Eukaryotic Genome and Its Expression What Are the Characteristics of the Eukaryotic Genome? Key differences between eukaryotic and prokaryotic
The Little Things About the Little Things Inside of Us The Eukaryotic Genome and Its Expression What Are the Characteristics of the Eukaryotic Genome? Key differences between eukaryotic and prokaryotic
Expression of the genome. Books: 1. Molecular biology of the gene: Watson et al 2. Genetics: Peter J. Russell
 Expression of the genome Books: 1. Molecular biology of the gene: Watson et al 2. Genetics: Peter J. Russell 1 Transcription 1. Francis Crick (1956) named the flow of information from DNA RNA protein the
Expression of the genome Books: 1. Molecular biology of the gene: Watson et al 2. Genetics: Peter J. Russell 1 Transcription 1. Francis Crick (1956) named the flow of information from DNA RNA protein the
DNA Transcription. Dr Aliwaini
 DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
Mechanisms of Transcription. School of Life Science Shandong University
 Mechanisms of Transcription School of Life Science Shandong University Ch 12: Mechanisms of Transcription 1. RNA polymerase and the transcription cycle 2. The transcription cycle in bacteria 3. Transcription
Mechanisms of Transcription School of Life Science Shandong University Ch 12: Mechanisms of Transcription 1. RNA polymerase and the transcription cycle 2. The transcription cycle in bacteria 3. Transcription
B. Incorrect! Centromeric DNA is largely heterochromatin, which is inactive DNA.
 MCAT Biology - Problem Drill 06: Molecular Biology of Eukaryotes Question No. 1 of 10 1. Which type of DNA would have the highest level of expression? Question #01 (A) Heterochromatin. (B) Centromeric
MCAT Biology - Problem Drill 06: Molecular Biology of Eukaryotes Question No. 1 of 10 1. Which type of DNA would have the highest level of expression? Question #01 (A) Heterochromatin. (B) Centromeric
Spring 2006 Biochemistry 302 Exam 2
 Name Spring 2006 Biochemistry 302 Exam 2 Directions: This exam has 45 questions/problems totaling 110 points. Check to make sure you have seven pages. Some questions have multiple parts so read each one
Name Spring 2006 Biochemistry 302 Exam 2 Directions: This exam has 45 questions/problems totaling 110 points. Check to make sure you have seven pages. Some questions have multiple parts so read each one
Gene Regulation in Eukaryotes
 Gene Regulation in Eukaryotes The latest estimates are that a human cell, a eukaryotic cell, contains 20,000 25,000 genes. Some of these are expressed in all cells all the time. These so-called housekeeping
Gene Regulation in Eukaryotes The latest estimates are that a human cell, a eukaryotic cell, contains 20,000 25,000 genes. Some of these are expressed in all cells all the time. These so-called housekeeping
Biochemistry 674, Fall, 1995: Nucleic Acids Exam II: November 16, 1995 Your Name Here: PCR A C
 CHM 674 Exam II, 1995 1 iochemistry 674, Fall, 1995: Nucleic cids Prof. Jason Kahn Exam II: November 16, 1995 Your Name Here: This exam has five questions worth 20 points each. nswer all five. You do not
CHM 674 Exam II, 1995 1 iochemistry 674, Fall, 1995: Nucleic cids Prof. Jason Kahn Exam II: November 16, 1995 Your Name Here: This exam has five questions worth 20 points each. nswer all five. You do not
17.5 Eukaryotic Transcription Initiation Is Regulated by Transcription Factors That Bind to Cis-Acting Sites
 17.5 Eukaryotic Transcription Initiation Is Regulated by Transcription Factors That Bind to Cis-Acting Sites 1 Section 17.5 Transcription regulatory proteins, transcription factors, target cis-acting sites
17.5 Eukaryotic Transcription Initiation Is Regulated by Transcription Factors That Bind to Cis-Acting Sites 1 Section 17.5 Transcription regulatory proteins, transcription factors, target cis-acting sites
BETA STRAND Prof. Alejandro Hochkoeppler Department of Pharmaceutical Sciences and Biotechnology University of Bologna
 Prof. Alejandro Hochkoeppler Department of Pharmaceutical Sciences and Biotechnology University of Bologna E-mail: a.hochkoeppler@unibo.it C-ter NH and CO groups: right, left, right (plane of the slide)
Prof. Alejandro Hochkoeppler Department of Pharmaceutical Sciences and Biotechnology University of Bologna E-mail: a.hochkoeppler@unibo.it C-ter NH and CO groups: right, left, right (plane of the slide)
Figure S1 Glucocorticoid receptor-green fluorescent protein (GR-GFP) reporter construct.
 Figure S1 Glucocorticoid receptor-green fluorescent protein (GR-GFP) reporter construct. (A) Schematic display of the construct is presented detailing binding sites and regions of interest within the GR
Figure S1 Glucocorticoid receptor-green fluorescent protein (GR-GFP) reporter construct. (A) Schematic display of the construct is presented detailing binding sites and regions of interest within the GR
AP Biology Gene Expression/Biotechnology REVIEW
 AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
AP Biology Gene Expression/Biotechnology REVIEW Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Gene expression can be a. regulated before transcription.
Eukaryotic Gene Expression John O. Thomas
 Eukaryotic Gene Expression John O. Thomas I) RNA polymerases A) There are four RNA polymerases in human cells. 1) RNA polymerase I, located in the nucleolar region of the nucleus, is responsible for the
Eukaryotic Gene Expression John O. Thomas I) RNA polymerases A) There are four RNA polymerases in human cells. 1) RNA polymerase I, located in the nucleolar region of the nucleus, is responsible for the
DNA Transcription. Visualizing Transcription. The Transcription Process
 DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
DNA Transcription By: Suzanne Clancy, Ph.D. 2008 Nature Education Citation: Clancy, S. (2008) DNA transcription. Nature Education 1(1) If DNA is a book, then how is it read? Learn more about the DNA transcription
Bi 8 Lecture 7. Ellen Rothenberg 26 January Reading: Ch. 3, pp ; panel 3-1
 Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Bi 8 Lecture 7 PROTEIN STRUCTURE, Functional analysis, and evolution Ellen Rothenberg 26 January 2016 Reading: Ch. 3, pp. 109-134; panel 3-1 (end with free amine) aromatic, hydrophobic small, hydrophilic
Regulatory Dynamics in Engineered Gene Networks
 Regulatory Dynamics in Engineered Gene Networks The Physico-chemical Foundation of Transcriptional Regulation with Applications to Systems Biology Mads Kærn Boston University Center for BioDynamics Center
Regulatory Dynamics in Engineered Gene Networks The Physico-chemical Foundation of Transcriptional Regulation with Applications to Systems Biology Mads Kærn Boston University Center for BioDynamics Center
Chapter 8 Lecture Outline. Transcription, Translation, and Bioinformatics
 Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
DNA. Is a molecule that encodes the genetic instructions used in the development and functioning of all known living organisms and many viruses.
 Is a molecule that encodes the genetic instructions used in the development and functioning of all known living organisms and many viruses. Genetic information is encoded as a sequence of nucleotides (guanine,
Is a molecule that encodes the genetic instructions used in the development and functioning of all known living organisms and many viruses. Genetic information is encoded as a sequence of nucleotides (guanine,
16 Regulation of Gene Expression
 16 Regulation of Gene Expression The prokaryotic bacterium Escherichia coli can use a wide range of different nutrients to sustain growth. It accomplishes this partially by turning on and off expression
16 Regulation of Gene Expression The prokaryotic bacterium Escherichia coli can use a wide range of different nutrients to sustain growth. It accomplishes this partially by turning on and off expression
3.1 Storing Information. Polypeptides. Protein Structure. Helical Secondary Structure. Types of Protein Structure. Molecular Biology Fourth Edition
 Lecture PowerPoint to accompany Molecular Biology Fourth Edition Robert F. Weaver Chapter 3 An Introduction to Gene Function 3.1 Storing Information Producing a protein from DNA information involves both
Lecture PowerPoint to accompany Molecular Biology Fourth Edition Robert F. Weaver Chapter 3 An Introduction to Gene Function 3.1 Storing Information Producing a protein from DNA information involves both
GENETICS - CLUTCH CH.10 TRANSCRIPTION.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF TRANSCRIPTION Transcription is the process of using DNA as a template to RNA RNA polymerase is the enzyme that transcribes DNA - There are many different types
!! www.clutchprep.com CONCEPT: OVERVIEW OF TRANSCRIPTION Transcription is the process of using DNA as a template to RNA RNA polymerase is the enzyme that transcribes DNA - There are many different types
All Rights Reserved. U.S. Patents 6,471,520B1; 5,498,190; 5,916, North Market Street, Suite CC130A, Milwaukee, WI 53202
 Secondary Structure In the previous protein folding activity, you created a hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously folds
Secondary Structure In the previous protein folding activity, you created a hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously folds
BIOCHEMISTRY REVIEW. Overview of Biomolecules. Chapter 10 Nucleic Acids
 BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 10 Nucleic Acids 2 3 DNA vs RNA DNA RNA deoxyribose ribose A, C, G, T A, C, G, U 10 3 10 8 nucleotides 10 2 10 4 nucleotides nucleus cytoplasm double-stranded
BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 10 Nucleic Acids 2 3 DNA vs RNA DNA RNA deoxyribose ribose A, C, G, T A, C, G, U 10 3 10 8 nucleotides 10 2 10 4 nucleotides nucleus cytoplasm double-stranded
However, only a fraction of these genes are transcribed in an individual cell at any given time.
 All cells in an organism contain the same set of genes. However, only a fraction of these genes are transcribed in an individual cell at any given time. It is the pattern of gene expression that determines
All cells in an organism contain the same set of genes. However, only a fraction of these genes are transcribed in an individual cell at any given time. It is the pattern of gene expression that determines
Contact information Blaine Bartholomew Office: Neckers Bldg., Rm. 211 Phone:
 Contact information Blaine Bartholomew Office: Neckers Bldg., Rm. 211 Phone: 453-6437 Email: bbartholomew@siumed.edu Class notes http://www.siumed.edu/~bbartholomew/hbtxn.html Other references Principles
Contact information Blaine Bartholomew Office: Neckers Bldg., Rm. 211 Phone: 453-6437 Email: bbartholomew@siumed.edu Class notes http://www.siumed.edu/~bbartholomew/hbtxn.html Other references Principles
Gene Regulation Biology
 Gene Regulation Biology Potential and Limitations of Cell Re-programming in Cancer Research Eric Blanc KCL April 13, 2010 Eric Blanc (KCL) Gene Regulation Biology April 13, 2010 1 / 21 Outline 1 The Central
Gene Regulation Biology Potential and Limitations of Cell Re-programming in Cancer Research Eric Blanc KCL April 13, 2010 Eric Blanc (KCL) Gene Regulation Biology April 13, 2010 1 / 21 Outline 1 The Central
Independent Study Guide The Blueprint of Life, from DNA to Protein (Chapter 7)
 Independent Study Guide The Blueprint of Life, from DNA to Protein (Chapter 7) I. General Principles (Chapter 7 introduction) a. Morse code distinct series of dots and dashes encode the 26 letters of the
Independent Study Guide The Blueprint of Life, from DNA to Protein (Chapter 7) I. General Principles (Chapter 7 introduction) a. Morse code distinct series of dots and dashes encode the 26 letters of the
Genetics Biology 331 Exam 3B Spring 2015
 Genetics Biology 331 Exam 3B Spring 2015 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) DNA methylation may be a significant mode of genetic regulation
Genetics Biology 331 Exam 3B Spring 2015 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) DNA methylation may be a significant mode of genetic regulation
Molecular Cell Biology - Problem Drill 09: Gene Expression in Prokaryotes
 Molecular Cell Biology - Problem Drill 09: Gene Expression in Prokaryotes Question No. 1 of 10 1. Which of the following statements about gene expression in prokaryotes is correct? Question #1 (A) In prokaryotes,
Molecular Cell Biology - Problem Drill 09: Gene Expression in Prokaryotes Question No. 1 of 10 1. Which of the following statements about gene expression in prokaryotes is correct? Question #1 (A) In prokaryotes,
RNA: Structure & Synthesis. Amr S. Moustafa, M.D.; Ph.D.
 RNA: Structure & Synthesis By Amr S. Moustafa, M.D.; Ph.D. Objectives The differences between DNA and RNA The structure and functions of RNAs RNA synthesis (Transcription) Post-transcriptional events (modifications)
RNA: Structure & Synthesis By Amr S. Moustafa, M.D.; Ph.D. Objectives The differences between DNA and RNA The structure and functions of RNAs RNA synthesis (Transcription) Post-transcriptional events (modifications)
Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site.
 Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site. Still having trouble understanding the material? Check
Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site. Still having trouble understanding the material? Check
Chapter 18. Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression 2007-2008 Control of Prokaryotic (Bacterial) Genes 2007- Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
Chapter 18 Regulation of Gene Expression 2007-2008 Control of Prokaryotic (Bacterial) Genes 2007- Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
DNA Prokaryote Transcription Steps (updated February 2013)
 URS AACTGT ATATTA - 35-10 transcription Pribnow Box discriminator +1 AGGAGGT TTA TCCTCCA ATT Gene C TGA TAG ACT ATC rho or GC hairpin loop transcription termination DNA Prokaryote Transcription Steps (updated
URS AACTGT ATATTA - 35-10 transcription Pribnow Box discriminator +1 AGGAGGT TTA TCCTCCA ATT Gene C TGA TAG ACT ATC rho or GC hairpin loop transcription termination DNA Prokaryote Transcription Steps (updated
Prokaryotic Transcription
 Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
Prokaryotic Transcription Transcription Basics DNA is the genetic material Nucleic acid Capable of self-replication and synthesis of RNA RNA is the middle man Nucleic acid Structure and base sequence are
PART FOUR - ANSWERS PART FOUR: GENE REGULATION ANSWERS. Answers to questions from Chapter 15 on Positive and negative control of the lac operon
 PART FOUR: GENE REGULATION ANSWERS Answers to questions from Chapter 15 on Positive and negative control of the lac operon 15.1 laci, lacz, lacy, laca 15.2 The lac operon is negatively regulated by a repressor,
PART FOUR: GENE REGULATION ANSWERS Answers to questions from Chapter 15 on Positive and negative control of the lac operon 15.1 laci, lacz, lacy, laca 15.2 The lac operon is negatively regulated by a repressor,
DOWNLOAD OR READ : EXAMPLES OF REGULATORY PROTEINS THAT CONTROL THE CELL CYCLE PDF EBOOK EPUB MOBI
 DOWNLOAD OR READ : EXAMPLES OF REGULATORY PROTEINS THAT CONTROL THE CELL CYCLE PDF EBOOK EPUB MOBI Page 1 Page 2 examples of regulatory proteins that control the cell cycle examples of regulatory proteins
DOWNLOAD OR READ : EXAMPLES OF REGULATORY PROTEINS THAT CONTROL THE CELL CYCLE PDF EBOOK EPUB MOBI Page 1 Page 2 examples of regulatory proteins that control the cell cycle examples of regulatory proteins
TRANSCRIPTION AND PROCESSING OF RNA
 TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
TRANSCRIPTION AND PROCESSING OF RNA 1. The steps of gene expression. 2. General characterization of transcription: steps, components of transcription apparatus. 3. Transcription of eukaryotic structural
Chapter 11: Regulation of Gene Expression
 Chapter Review 1. It has long been known that there is probably a genetic link for alcoholism. Researchers studying rats have begun to elucidate this link. Briefly describe the genetic mechanism found
Chapter Review 1. It has long been known that there is probably a genetic link for alcoholism. Researchers studying rats have begun to elucidate this link. Briefly describe the genetic mechanism found
BIOLOGY. Chapter 16 GenesExpression
 BIOLOGY Chapter 16 GenesExpression CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 18 Gene Expression 2014 Pearson Education, Inc. Figure 16.1 Differential Gene Expression results
BIOLOGY Chapter 16 GenesExpression CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 18 Gene Expression 2014 Pearson Education, Inc. Figure 16.1 Differential Gene Expression results
Announcement Structure Analysis
 Announcement Structure Analysis BSC 4439/BSC 5436: Biomedical Informatics: Structure Analysis Spring 2019, CB117 Monday and Wednesday 12:00 1:15pm Office hour: Monday and Wednesday 1:15 2pm Topics include
Announcement Structure Analysis BSC 4439/BSC 5436: Biomedical Informatics: Structure Analysis Spring 2019, CB117 Monday and Wednesday 12:00 1:15pm Office hour: Monday and Wednesday 1:15 2pm Topics include
Chapter 3. DNA, RNA, and Protein Synthesis
 Chapter 3. DNA, RNA, and Protein Synthesis 4. Transcription Gene Expression Regulatory region (promoter) 5 flanking region Upstream region Coding region 3 flanking region Downstream region Transcription
Chapter 3. DNA, RNA, and Protein Synthesis 4. Transcription Gene Expression Regulatory region (promoter) 5 flanking region Upstream region Coding region 3 flanking region Downstream region Transcription
 The replication of DNA Kornberg 1957 Meselson and Stahl 1958 Cairns 1963 Okazaki 1968 DNA Replication The driving force for DNA synthesis. The addition of a nucleotide to a growing polynucleotide
The replication of DNA Kornberg 1957 Meselson and Stahl 1958 Cairns 1963 Okazaki 1968 DNA Replication The driving force for DNA synthesis. The addition of a nucleotide to a growing polynucleotide
Transcription Gene regulation
 Transcription Gene regulation The machine that transcribes a gene is composed of perhaps 50 proteins, including RNA polymerase, the enzyme that converts DNA code into RNA code. A crew of transcription
Transcription Gene regulation The machine that transcribes a gene is composed of perhaps 50 proteins, including RNA polymerase, the enzyme that converts DNA code into RNA code. A crew of transcription
Gene Regulation in Eukaryotes. Dr. Syahril Abdullah Medical Genetics Laboratory
 Gene Regulation in Eukaryotes Dr. Syahril Abdullah Medical Genetics Laboratory syahril@medic.upm.edu.my Lecture Outline 1. The Genome 2. Overview of Gene Control 3. Cellular Differentiation in Higher Eukaryotes
Gene Regulation in Eukaryotes Dr. Syahril Abdullah Medical Genetics Laboratory syahril@medic.upm.edu.my Lecture Outline 1. The Genome 2. Overview of Gene Control 3. Cellular Differentiation in Higher Eukaryotes
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION
 M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
M I C R O B I O L O G Y WITH DISEASES BY TAXONOMY, THIRD EDITION Chapter 7 Microbial Genetics Lecture prepared by Mindy Miller-Kittrell, University of Tennessee, Knoxville The Structure and Replication
DNA Evolution of knowledge about gene. Contains information about RNAs and proteins. Polynucleotide chains; Double stranded molecule;
 Evolution of knowledge about gene G. Mendel Hereditary factors W.Johannsen, 1909 G.W.Beadle, E.L.Tatum, 1945 Ingram, 1957 Actual concepts The gene hereditary unit located in chromosomes Hypotheses One
Evolution of knowledge about gene G. Mendel Hereditary factors W.Johannsen, 1909 G.W.Beadle, E.L.Tatum, 1945 Ingram, 1957 Actual concepts The gene hereditary unit located in chromosomes Hypotheses One
