Gene Finding Genome Annotation
|
|
|
- Merilyn Charles
- 6 years ago
- Views:
Transcription
1 Gene Finding Genome Annotation
2 Gene finding is a cornerstone of genomic analysis Genome content and organization Differential expression analysis Epigenomics Population biology & evolution Medical genomics
3 Basic Approaches Computational Absolute rules: start and stop codons Statistical probabilities: which codon is a true start? Introns splice junctions codon usage Experimental Comparison with known genes/proteins (BLAST) Expressed sequence tags RNAseq data
4 Computational Gene Prediction Statistical properties of protein-coding genes differ from those of non-coding sequence Long ORFs On average stop codons should occur 3 times in every 64 codons (~1/21) Codon bias (human) codon Amino acid % ACA Thr 28 ACC Thr 36 ACG Thr 12 ACU Thr 24
5 Gene features tend to occur in specific sequence contexts a. Splice acceptor sites b. Splice donor sites c. Translation starts d. Splice acceptor sites for A. thaliana genes predicted using C. elegans parameters from Korf(2004)
6 Many of the ab initio gene finders use Hidden Markov Models (HMMs) HMMs Contain parameters defining probabilities that specific gene features occur in different sequence contexts They can be used to predict transcription start sites Intron splice junctions Poly-A addition sites promoters
7 Standard practice is to perform gene predictions with multiple programs We will run two programs in today s exercise: SNAP Korf (2004) Gene finding in novel genomes BMC Bioinformatics 5:59 AUGUSTUS Stanke et al (2004) AUGUSTUS: a web server for gene finding in eukaryotes. Nucl. Acids Research 32:W309
8 Gene validation Independent evidence that our candidate gene is, in fact, a gene Conserved protein motifs Blast matches Expressed sequence tags RNAseq reads
9 For today s exercise We will use the following evidences: Genes/proteins already identified in M.oryzae (many being well supported by blast, EST and other transcriptomic data) Splice junction information from the RNAseq mapping that we performed yesterday
10 Information overload!!! Results from: SNAP AUGUSTUS Magnaporthe genes Magnaporthe proteins RNAseq mapping data How are we going to make sense out of these highly redundant datasets?
11 Enter MAKER Synthesizes multiple forms of gene prediction data Predictions and evidences Outputs a single, consistent set of genes and gene models, including quality values Uses a standard gene annotation format GFF3 (related to the GTF format used yesterday) Results can be imported into a genome browser
12 GFF3 format seqid source type Start End Score Strand phase attributes ##gff-version 3 ##date Wed Jul 18 22:38: ##source gbrowse gbgff gff3 dumper ##sequence-region contig00001: contig00001 maker gene Name=snap_masked-contig00001-abinitgene-0.164;ID= contig00001 maker mrna Name=snap_masked-contig00001-abinitgene mRNA-1;Parent=215076;ID=215077;_QI=0%7C0%7C0%7C0%7C1%7C1%7C2%7C0%7C1128;_AED=1.00 contig00001 maker exon Parent=215077;ID= contig00001 maker exon Parent=215077;ID= contig00001 maker CDS Parent=215077;ID= contig00001 maker CDS Parent=215077;ID= contig00001 maker mrna Name=snap_masked-contig00001-abinitgene mRNA-1;ID=215077;_QI=0%7C0%7C0%7C0%7C1%7C1%7C2%7C0%7C1128;_AED=1.00 contig00001 maker exon Parent=215077;ID= contig00001 maker exon Parent=215077;ID= contig00001 maker CDS Parent=215077;ID= contig00001 maker CDS Parent=215077;ID= contig00001 maker gene Name=maker-contig00001-snap-gene ;ID= contig00001 maker mrna Name=maker-contig00001-snap-gene mrna-1;parent=215008;id=215009;_qi=0%7c0.5%7c0.33%7c1%7c0%7c0.33%7c3%7c0%7c285;_aed=0.06 contig00001 maker exon Parent=215009;ID= contig00001 maker exon Parent=215009;ID= contig00001 maker exon Parent=215009;ID= contig00001 maker CDS Parent=215009;ID= contig00001 maker CDS Parent=215009;ID= contig00001 maker CDS Parent=215009;ID= contig00001 maker mrna Name=maker-contig00001-snap-gene mrna-1;id=215009;_qi=0%7c0.5%7c0.33%7c1%7c0%7c0.33%7c3%7c0%7c285;_aed=0.06 contig00001 maker exon Parent=215009;ID= contig00001 maker exon Parent=215009;ID= contig00001 maker exon Parent=215009;ID= contig00001 maker CDS Parent=215009;ID= contig00001 maker CDS Parent=215009;ID= contig00001 maker CDS Parent=215009;ID=215015
13 Gene finding is an iterative process HMM SNAP AUGUSTUS GENE MODELS MAKER BLAST matches ESTs
14 Genome Browsers
15 Genome Browser Combines a genome database with interactive web pages Allows the user to retrieve and manipulate database record through a graphical user interface (GUI) Different types of information are displayed in an intuitive fashion in user-configurable tracks
16 GFF3 files are hard to interpret ##gff-version 3 ##date Wed Jul 18 22:38: ##source gbrowse gbgff gff3 dumper ##sequence-region contig00001: contig00001 maker gene Name=snap_masked-contig00001-abinitgene-0.164;ID= contig00001 maker mrna Name=snap_masked-contig00001-abinitgene mRNA-1;Parent=215076;ID=215077;_QI=0%7C0%7C0%7C0%7C1%7C1%7C2%7C0%7C1128;_AED=1.00 contig00001 maker exon Parent=215077;ID= contig00001 maker exon Parent=215077;ID= contig00001 maker CDS Parent=215077;ID= contig00001 maker CDS Parent=215077;ID= contig00001 maker mrna Name=snap_masked-contig00001-abinitgene mRNA-1;ID=215077;_QI=0%7C0%7C0%7C0%7C1%7C1%7C2%7C0%7C1128;_AED=1.00 contig00001 maker exon Parent=215077;ID= contig00001 maker exon Parent=215077;ID= contig00001 maker CDS Parent=215077;ID= contig00001 maker CDS Parent=215077;ID= contig00001 maker gene Name=maker-contig00001-snap-gene ;ID= contig00001 maker mrna Name=maker-contig00001-snap-gene mrna-1;parent=215008;id=215009;_qi=0%7c0.5%7c0.33%7c1%7c0%7c0.33%7c3%7c0%7c285;_aed=0.06 contig00001 maker exon Parent=215009;ID= contig00001 maker exon Parent=215009;ID= contig00001 maker exon Parent=215009;ID= contig00001 maker CDS Parent=215009;ID= contig00001 maker CDS Parent=215009;ID= contig00001 maker CDS Parent=215009;ID= contig00001 maker mrna Name=maker-contig00001-snap-gene mrna-1;id=215009;_qi=0%7c0.5%7c0.33%7c1%7c0%7c0.33%7c3%7c0%7c285;_aed=0.06 contig00001 maker exon Parent=215009;ID= contig00001 maker exon Parent=215009;ID= contig00001 maker exon Parent=215009;ID= contig00001 maker CDS Parent=215009;ID= contig00001 maker CDS Parent=215009;ID= contig00001 maker CDS Parent=215009;ID=215015
17 MAKER genes & RNAseq reads in GBrowse
18 Genome Browsers for repeat definition Show is a track displaying the results of a genome blasted against itself
19 A plethora of genome browsers Annmap Apollo Genome Annotation Curation Tool Argo Genome Browser Avadis NGS BugView Celera Genome Browser Dalliance DiProGB DNAnexus Ensembl Gaggle Genome Browser GBrowse The Genomic HyperBrowser Genostar GenoBrowser GenPlay Integrated Genome Browser (IGB) Integrated Genome Viewer (IGV) Integrated Microbial Genomes (IMG) JBrowse (a JavaScript browser ) MGV - Microbial Genome Viewer MochiView Genome Browser NextBio Genome Browser Pathway Tools Genome Browser Savant Genome Browser SEED viewer UCSC Genome Bioinformatics Genome Browser Viral Genome Organizer (VGO) VISTA genome browser
20 Today s activity Learn how to use the Integrated Genome Browser Populate the browser with data: A Magnaporthe sequence contig MAKER annotations Mapped RNAseq reads RNAseqread heatmaps Explore the browser to get an idea of how it works and how the tracks can be manipulated/activated/deactivated
Outline. Introduction to ab initio and evidence-based gene finding. Prokaryotic gene predictions
 Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
Outline Introduction to ab initio and evidence-based gene finding Overview of computational gene predictions Different types of eukaryotic gene predictors Common types of gene prediction errors Wilson
Genome annotation. Erwin Datema (2011) Sandra Smit (2012, 2013)
 Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
Genome annotation Erwin Datema (2011) Sandra Smit (2012, 2013) Genome annotation AGACAAAGATCCGCTAAATTAAATCTGGACTTCACATATTGAAGTGATATCACACGTTTCTCTAAT AATCTCCTCACAATATTATGTTTGGGATGAACTTGTCGTGATTTGCCATTGTAGCAATCACTTGAA
Agenda. Web Databases for Drosophila. Gene annotation workflow. GEP Drosophila annotation projects 01/01/2018. Annotation adding labels to a sequence
 Agenda GEP annotation project overview Web Databases for Drosophila An introduction to web tools, databases and NCBI BLAST Web databases for Drosophila annotation UCSC Genome Browser NCBI / BLAST FlyBase
Agenda GEP annotation project overview Web Databases for Drosophila An introduction to web tools, databases and NCBI BLAST Web databases for Drosophila annotation UCSC Genome Browser NCBI / BLAST FlyBase
Genome Annotation. Stefan Prost 1. May 27th, States of America. Genome Annotation
 Genome Annotation Stefan Prost 1 1 Department of Integrative Biology, University of California, Berkeley, United States of America May 27th, 2015 Outline Genome Annotation 1 Repeat Annotation 2 Repeat
Genome Annotation Stefan Prost 1 1 Department of Integrative Biology, University of California, Berkeley, United States of America May 27th, 2015 Outline Genome Annotation 1 Repeat Annotation 2 Repeat
UCSC Genome Browser. Introduction to ab initio and evidence-based gene finding
 UCSC Genome Browser Introduction to ab initio and evidence-based gene finding Wilson Leung 06/2006 Outline Introduction to annotation ab initio gene finding Basics of the UCSC Browser Evidence-based gene
UCSC Genome Browser Introduction to ab initio and evidence-based gene finding Wilson Leung 06/2006 Outline Introduction to annotation ab initio gene finding Basics of the UCSC Browser Evidence-based gene
Outline. Annotation of Drosophila Primer. Gene structure nomenclature. Muller element nomenclature. GEP Drosophila annotation projects 01/04/2018
 Outline Overview of the GEP annotation projects Annotation of Drosophila Primer January 2018 GEP annotation workflow Practice applying the GEP annotation strategy Wilson Leung and Chris Shaffer AAACAACAATCATAAATAGAGGAAGTTTTCGGAATATACGATAAGTGAAATATCGTTCT
Outline Overview of the GEP annotation projects Annotation of Drosophila Primer January 2018 GEP annotation workflow Practice applying the GEP annotation strategy Wilson Leung and Chris Shaffer AAACAACAATCATAAATAGAGGAAGTTTTCGGAATATACGATAAGTGAAATATCGTTCT
Methods and Algorithms for Gene Prediction
 Methods and Algorithms for Gene Prediction Chaochun Wei 韦朝春 Sc.D. ccwei@sjtu.edu.cn http://cbb.sjtu.edu.cn/~ccwei Shanghai Jiao Tong University Shanghai Center for Bioinformation Technology 5/12/2011 K-J-C
Methods and Algorithms for Gene Prediction Chaochun Wei 韦朝春 Sc.D. ccwei@sjtu.edu.cn http://cbb.sjtu.edu.cn/~ccwei Shanghai Jiao Tong University Shanghai Center for Bioinformation Technology 5/12/2011 K-J-C
Outline. Gene Finding Questions. Recap: Prokaryotic gene finding Eukaryotic gene finding The human gene complement Regulation
 Tues, Nov 29: Gene Finding 1 Online FCE s: Thru Dec 12 Thurs, Dec 1: Gene Finding 2 Tues, Dec 6: PS5 due Project presentations 1 (see course web site for schedule) Thurs, Dec 8 Final papers due Project
Tues, Nov 29: Gene Finding 1 Online FCE s: Thru Dec 12 Thurs, Dec 1: Gene Finding 2 Tues, Dec 6: PS5 due Project presentations 1 (see course web site for schedule) Thurs, Dec 8 Final papers due Project
NCBI & Other Genome Databases. BME 110/BIOL 181 CompBio Tools
 NCBI & Other Genome Databases BME 110/BIOL 181 CompBio Tools Todd Lowe March 31, 2011 Admin Reading Dummies Ch 3 Assigned Review: "The impact of next-generation sequencing technology on genetics" by E.
NCBI & Other Genome Databases BME 110/BIOL 181 CompBio Tools Todd Lowe March 31, 2011 Admin Reading Dummies Ch 3 Assigned Review: "The impact of next-generation sequencing technology on genetics" by E.
TIGR THE INSTITUTE FOR GENOMIC RESEARCH
 Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
Introduction to Genome Annotation: Overview of What You Will Learn This Week C. Robin Buell May 21, 2007 Types of Annotation Structural Annotation: Defining genes, boundaries, sequence motifs e.g. ORF,
Agenda. Annotation of Drosophila. Muller element nomenclature. Annotation: Adding labels to a sequence. GEP Drosophila annotation projects 01/03/2018
 Agenda Annotation of Drosophila January 2018 Overview of the GEP annotation project GEP annotation strategy Types of evidence Analysis tools Web databases Annotation of a single isoform (walkthrough) Wilson
Agenda Annotation of Drosophila January 2018 Overview of the GEP annotation project GEP annotation strategy Types of evidence Analysis tools Web databases Annotation of a single isoform (walkthrough) Wilson
Computational gene finding. Devika Subramanian Comp 470
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) The biological context Lec 1 Lec 2 Lec 3 Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) The biological context Lec 1 Lec 2 Lec 3 Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Bioinformatics for Proteomics. Ann Loraine
 Bioinformatics for Proteomics Ann Loraine aloraine@uab.edu What is bioinformatics? The science of collecting, processing, organizing, storing, analyzing, and mining biological information, especially data
Bioinformatics for Proteomics Ann Loraine aloraine@uab.edu What is bioinformatics? The science of collecting, processing, organizing, storing, analyzing, and mining biological information, especially data
Gene Identification in silico
 Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Gene Identification in silico Nita Parekh, IIIT Hyderabad Presented at National Seminar on Bioinformatics and Functional Genomics, at Bioinformatics centre, Pondicherry University, Feb 15 17, 2006. Introduction
Genscan. The Genscan HMM model Training Genscan Validating Genscan. (c) Devika Subramanian,
 Genscan The Genscan HMM model Training Genscan Validating Genscan (c) Devika Subramanian, 2009 96 Gene structure assumed by Genscan donor site acceptor site (c) Devika Subramanian, 2009 97 A simple model
Genscan The Genscan HMM model Training Genscan Validating Genscan (c) Devika Subramanian, 2009 96 Gene structure assumed by Genscan donor site acceptor site (c) Devika Subramanian, 2009 97 A simple model
Annotating 7G24-63 Justin Richner May 4, Figure 1: Map of my sequence
 Annotating 7G24-63 Justin Richner May 4, 2005 Zfh2 exons Thd1 exons Pur-alpha exons 0 40 kb 8 = 1 kb = LINE, Penelope = DNA/Transib, Transib1 = DINE = Novel Repeat = LTR/PAO, Diver2 I = LTR/Gypsy, Invader
Annotating 7G24-63 Justin Richner May 4, 2005 Zfh2 exons Thd1 exons Pur-alpha exons 0 40 kb 8 = 1 kb = LINE, Penelope = DNA/Transib, Transib1 = DINE = Novel Repeat = LTR/PAO, Diver2 I = LTR/Gypsy, Invader
From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow
 From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow Technical Overview Import VCF Introduction Next-generation sequencing (NGS) studies have created unanticipated challenges with
From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow Technical Overview Import VCF Introduction Next-generation sequencing (NGS) studies have created unanticipated challenges with
Genome and DNA Sequence Databases. BME 110: CompBio Tools Todd Lowe April 5, 2007
 Genome and DNA Sequence Databases BME 110: CompBio Tools Todd Lowe April 5, 2007 Admin Reading: Chapters 2 & 3 Notes available in PDF format on-line (see class calendar page): http://www.soe.ucsc.edu/classes/bme110/spring07/bme110-calendar.html
Genome and DNA Sequence Databases BME 110: CompBio Tools Todd Lowe April 5, 2007 Admin Reading: Chapters 2 & 3 Notes available in PDF format on-line (see class calendar page): http://www.soe.ucsc.edu/classes/bme110/spring07/bme110-calendar.html
BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers
 BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html UCSC
BCHM 6280 Tutorial: Gene specific information using NCBI, Ensembl and genome viewers Web resources: NCBI database: http://www.ncbi.nlm.nih.gov/ Ensembl database: http://useast.ensembl.org/index.html UCSC
MAKER: An easy-to-use annotation pipeline designed for emerging model organism genomes
 Resource MAKER: An easy-to-use annotation pipeline designed for emerging model organism genomes Brandi L. Cantarel, 1 Ian Korf, 2 Sofia M.C. Robb, 3 Genis Parra, 2 Eric Ross, 4 Barry Moore, 1 Carson Holt,
Resource MAKER: An easy-to-use annotation pipeline designed for emerging model organism genomes Brandi L. Cantarel, 1 Ian Korf, 2 Sofia M.C. Robb, 3 Genis Parra, 2 Eric Ross, 4 Barry Moore, 1 Carson Holt,
Interpreting RNA-seq data (Browser Exercise II)
 Interpreting RNA-seq data (Browser Exercise II) In previous exercises, you spent some time learning about gene pages and examining genes in the context of the GBrowse genome browser. It is important to
Interpreting RNA-seq data (Browser Exercise II) In previous exercises, you spent some time learning about gene pages and examining genes in the context of the GBrowse genome browser. It is important to
RNA-Seq with the Tuxedo Suite
 RNA-Seq with the Tuxedo Suite Monica Britton, Ph.D. Sr. Bioinformatics Analyst September 2015 Workshop The Basic Tuxedo Suite References Trapnell C, et al. 2009 TopHat: discovering splice junctions with
RNA-Seq with the Tuxedo Suite Monica Britton, Ph.D. Sr. Bioinformatics Analyst September 2015 Workshop The Basic Tuxedo Suite References Trapnell C, et al. 2009 TopHat: discovering splice junctions with
From assembled genome to annotated genome
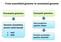 From assembled genome to annotated genome Procaryotic genomes Eucaryotic genomes Genome annotation servers (web based) 1. RAST 2. NCBI Gene prediction pipeline: Maker Function annotation pipeline: Blast2GO
From assembled genome to annotated genome Procaryotic genomes Eucaryotic genomes Genome annotation servers (web based) 1. RAST 2. NCBI Gene prediction pipeline: Maker Function annotation pipeline: Blast2GO
Make the protein through the genetic dogma process.
 Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
Make the protein through the genetic dogma process. Coding Strand 5 AGCAATCATGGATTGGGTACATTTGTAACTGT 3 Template Strand mrna Protein Complete the table. DNA strand DNA s strand G mrna A C U G T A T Amino
COMPUTER RESOURCES II:
 COMPUTER RESOURCES II: Using the computer to analyze data, using the internet, and accessing online databases Bio 210, Fall 2006 Linda S. Huang, Ph.D. University of Massachusetts Boston In the first computer
COMPUTER RESOURCES II: Using the computer to analyze data, using the internet, and accessing online databases Bio 210, Fall 2006 Linda S. Huang, Ph.D. University of Massachusetts Boston In the first computer
Machine Learning Methods for RNA-seq-based Transcriptome Reconstruction
 Machine Learning Methods for RNA-seq-based Transcriptome Reconstruction Gunnar Rätsch Friedrich Miescher Laboratory Max Planck Society, Tübingen, Germany NGS Bioinformatics Meeting, Paris (March 24, 2010)
Machine Learning Methods for RNA-seq-based Transcriptome Reconstruction Gunnar Rätsch Friedrich Miescher Laboratory Max Planck Society, Tübingen, Germany NGS Bioinformatics Meeting, Paris (March 24, 2010)
Steve Thompson 9/25/03. Introduction to BioInformatics BSC4933/5936. Florida State University Department of Biology
 Introduction to BioInformatics BSC4933/5936 Florida State University Department of Biology www.bio.fsu.edu October 2, 2003 Genomics Steven M. Thompson Florida State University School of Computational Science
Introduction to BioInformatics BSC4933/5936 Florida State University Department of Biology www.bio.fsu.edu October 2, 2003 Genomics Steven M. Thompson Florida State University School of Computational Science
Genome Annotation. What Does Annotation Describe??? Genome duplications Genes Mobile genetic elements Small repeats Genetic diversity
 Genome Annotation Genome Sequencing Costliest aspect of sequencing the genome o But Devoid of content Genome must be annotated o Annotation definition Analyzing the raw sequence of a genome and describing
Genome Annotation Genome Sequencing Costliest aspect of sequencing the genome o But Devoid of content Genome must be annotated o Annotation definition Analyzing the raw sequence of a genome and describing
Genomic region (ENCODE) Gene definitions
 DNA From genes to proteins Bioinformatics Methods RNA PROMOTER ELEMENTS TRANSCRIPTION Iosif Vaisman mrna SPLICE SITES SPLICING Email: ivaisman@gmu.edu START CODON STOP CODON TRANSLATION PROTEIN From genes
DNA From genes to proteins Bioinformatics Methods RNA PROMOTER ELEMENTS TRANSCRIPTION Iosif Vaisman mrna SPLICE SITES SPLICING Email: ivaisman@gmu.edu START CODON STOP CODON TRANSLATION PROTEIN From genes
Leonardo Mariño-Ramírez, PhD NCBI / NLM / NIH. BIOL 7210 A Computational Genomics 2/18/2015
 Leonardo Mariño-Ramírez, PhD NCBI / NLM / NIH BIOL 7210 A Computational Genomics 2/18/2015 The $1,000 genome is here! http://www.illumina.com/systems/hiseq-x-sequencing-system.ilmn Bioinformatics bottleneck
Leonardo Mariño-Ramírez, PhD NCBI / NLM / NIH BIOL 7210 A Computational Genomics 2/18/2015 The $1,000 genome is here! http://www.illumina.com/systems/hiseq-x-sequencing-system.ilmn Bioinformatics bottleneck
About Strand NGS. Strand Genomics, Inc All rights reserved.
 About Strand NGS Strand NGS-formerly known as Avadis NGS, is an integrated platform that provides analysis, management and visualization tools for next-generation sequencing data. It supports extensive
About Strand NGS Strand NGS-formerly known as Avadis NGS, is an integrated platform that provides analysis, management and visualization tools for next-generation sequencing data. It supports extensive
AGILENT S BIOINFORMATICS ANALYSIS SOFTWARE
 ACCELERATING PROGRESS IS IN OUR GENES AGILENT S BIOINFORMATICS ANALYSIS SOFTWARE GENESPRING GENE EXPRESSION (GX) MASS PROFILER PROFESSIONAL (MPP) PATHWAY ARCHITECT (PA) See Deeper. Reach Further. BIOINFORMATICS
ACCELERATING PROGRESS IS IN OUR GENES AGILENT S BIOINFORMATICS ANALYSIS SOFTWARE GENESPRING GENE EXPRESSION (GX) MASS PROFILER PROFESSIONAL (MPP) PATHWAY ARCHITECT (PA) See Deeper. Reach Further. BIOINFORMATICS
GREG GIBSON SPENCER V. MUSE
 A Primer of Genome Science ience THIRD EDITION TAGCACCTAGAATCATGGAGAGATAATTCGGTGAGAATTAAATGGAGAGTTGCATAGAGAACTGCGAACTG GREG GIBSON SPENCER V. MUSE North Carolina State University Sinauer Associates, Inc.
A Primer of Genome Science ience THIRD EDITION TAGCACCTAGAATCATGGAGAGATAATTCGGTGAGAATTAAATGGAGAGTTGCATAGAGAACTGCGAACTG GREG GIBSON SPENCER V. MUSE North Carolina State University Sinauer Associates, Inc.
Year III Pharm.D Dr. V. Chitra
 Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Year III Pharm.D Dr. V. Chitra 1 Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Only one strand of DNA serves
Eukaryotic Gene Prediction. Wei Zhu May 2007
 Eukaryotic Gene Prediction Wei Zhu May 2007 In nature, nothing is perfect... - Alice Walker Gene Structure What is Gene Prediction? Gene prediction is the problem of parsing a sequence into nonoverlapping
Eukaryotic Gene Prediction Wei Zhu May 2007 In nature, nothing is perfect... - Alice Walker Gene Structure What is Gene Prediction? Gene prediction is the problem of parsing a sequence into nonoverlapping
132 Grundlagen der Bioinformatik, SoSe 14, D. Huson, June 22, This exposition is based on the following source, which is recommended reading:
 132 Grundlagen der Bioinformatik, SoSe 14, D. Huson, June 22, 214 1 Gene Prediction Using HMMs This exposition is based on the following source, which is recommended reading: 1. Chris Burge and Samuel
132 Grundlagen der Bioinformatik, SoSe 14, D. Huson, June 22, 214 1 Gene Prediction Using HMMs This exposition is based on the following source, which is recommended reading: 1. Chris Burge and Samuel
Genomics and Transcriptomics of Spirodela polyrhiza
 Genomics and Transcriptomics of Spirodela polyrhiza Doug Bryant Bioinformatics Core Facility & Todd Mockler Group, Donald Danforth Plant Science Center Desired Outcomes High-quality genomic reference sequence
Genomics and Transcriptomics of Spirodela polyrhiza Doug Bryant Bioinformatics Core Facility & Todd Mockler Group, Donald Danforth Plant Science Center Desired Outcomes High-quality genomic reference sequence
Grundlagen der Bioinformatik, SoSe 11, D. Huson, July 4, This exposition is based on the following source, which is recommended reading:
 Grundlagen der Bioinformatik, SoSe 11, D. Huson, July 4, 211 155 12 Gene Prediction Using HMMs This exposition is based on the following source, which is recommended reading: 1. Chris Burge and Samuel
Grundlagen der Bioinformatik, SoSe 11, D. Huson, July 4, 211 155 12 Gene Prediction Using HMMs This exposition is based on the following source, which is recommended reading: 1. Chris Burge and Samuel
RNA-Seq Software, Tools, and Workflows
 RNA-Seq Software, Tools, and Workflows Monica Britton, Ph.D. Sr. Bioinformatics Analyst September 1, 2016 Some mrna-seq Applications Differential gene expression analysis Transcriptional profiling Assumption:
RNA-Seq Software, Tools, and Workflows Monica Britton, Ph.D. Sr. Bioinformatics Analyst September 1, 2016 Some mrna-seq Applications Differential gene expression analysis Transcriptional profiling Assumption:
MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE?
 MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE? Lesson Plan: Title Introduction to the Genome Browser: what is a gene? JOYCE STAMM Objectives Demonstrate basic skills in using the UCSC Genome
MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE? Lesson Plan: Title Introduction to the Genome Browser: what is a gene? JOYCE STAMM Objectives Demonstrate basic skills in using the UCSC Genome
Ensembl Tools. EBI is an Outstation of the European Molecular Biology Laboratory.
 Ensembl Tools EBI is an Outstation of the European Molecular Biology Laboratory. Questions? We ve muted all the mics Ask questions in the Chat box in the webinar interface I will check the Chat box periodically
Ensembl Tools EBI is an Outstation of the European Molecular Biology Laboratory. Questions? We ve muted all the mics Ask questions in the Chat box in the webinar interface I will check the Chat box periodically
Using Expressing Sequence Tags to Improve Gene Structure Annotation
 Washington University in St. Louis Washington University Open Scholarship All Computer Science and Engineering Research Computer Science and Engineering Report Number: WUCS-2006-25 2006-05-01 Using Expressing
Washington University in St. Louis Washington University Open Scholarship All Computer Science and Engineering Research Computer Science and Engineering Report Number: WUCS-2006-25 2006-05-01 Using Expressing
Annotating Fosmid 14p24 of D. Virilis chromosome 4
 Lo 1 Annotating Fosmid 14p24 of D. Virilis chromosome 4 Lo, Louis April 20, 2006 Annotation Report Introduction In the first half of Research Explorations in Genomics I finished a 38kb fragment of chromosome
Lo 1 Annotating Fosmid 14p24 of D. Virilis chromosome 4 Lo, Louis April 20, 2006 Annotation Report Introduction In the first half of Research Explorations in Genomics I finished a 38kb fragment of chromosome
Gene prediction in novel fungal genomes using an ab initio algorithm with unsupervised training
 Methods Gene prediction in novel fungal genomes using an ab initio algorithm with unsupervised training Vardges Ter-Hovhannisyan, 1,4 Alexandre Lomsadze, 2,4 Yury O. Chernoff, 1 and Mark Borodovsky 2,3,5
Methods Gene prediction in novel fungal genomes using an ab initio algorithm with unsupervised training Vardges Ter-Hovhannisyan, 1,4 Alexandre Lomsadze, 2,4 Yury O. Chernoff, 1 and Mark Borodovsky 2,3,5
DNA is normally found in pairs, held together by hydrogen bonds between the bases
 Bioinformatics Biology Review The genetic code is stored in DNA Deoxyribonucleic acid. DNA molecules are chains of four nucleotide bases Guanine, Thymine, Cytosine, Adenine DNA is normally found in pairs,
Bioinformatics Biology Review The genetic code is stored in DNA Deoxyribonucleic acid. DNA molecules are chains of four nucleotide bases Guanine, Thymine, Cytosine, Adenine DNA is normally found in pairs,
Finding Genes, Building Search Strategies and Visiting a Gene Page
 Finding Genes, Building Search Strategies and Visiting a Gene Page 1. Finding a gene using text search. For this exercise use http://www.plasmodb.org a. Find all possible kinases in Plasmodium. Hint: use
Finding Genes, Building Search Strategies and Visiting a Gene Page 1. Finding a gene using text search. For this exercise use http://www.plasmodb.org a. Find all possible kinases in Plasmodium. Hint: use
Gene-centered resources at NCBI
 COURSE OF BIOINFORMATICS a.a. 2014-2015 Gene-centered resources at NCBI We searched Accession Number: M60495 AT NCBI Nucleotide Gene has been implemented at NCBI to organize information about genes, serving
COURSE OF BIOINFORMATICS a.a. 2014-2015 Gene-centered resources at NCBI We searched Accession Number: M60495 AT NCBI Nucleotide Gene has been implemented at NCBI to organize information about genes, serving
Finding Genes, Building Search Strategies and Visiting a Gene Page
 Finding Genes, Building Search Strategies and Visiting a Gene Page 1. Finding a gene using text search. For this exercise use http://www.plasmodb.org a. Find all possible kinases in Plasmodium. Hint: use
Finding Genes, Building Search Strategies and Visiting a Gene Page 1. Finding a gene using text search. For this exercise use http://www.plasmodb.org a. Find all possible kinases in Plasmodium. Hint: use
NCBI web resources I: databases and Entrez
 NCBI web resources I: databases and Entrez Yanbin Yin Most materials are downloaded from ftp://ftp.ncbi.nih.gov/pub/education/ 1 Homework assignment 1 Two parts: Extract the gene IDs reported in table
NCBI web resources I: databases and Entrez Yanbin Yin Most materials are downloaded from ftp://ftp.ncbi.nih.gov/pub/education/ 1 Homework assignment 1 Two parts: Extract the gene IDs reported in table
Eukaryotic Gene Structure
 Eukaryotic Gene Structure Terminology Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Gene Basic physical and
Eukaryotic Gene Structure Terminology Genome entire genetic material of an individual Transcriptome set of transcribed sequences Proteome set of proteins encoded by the genome 2 Gene Basic physical and
What we ll do today. Types of stem cells. Do engineered ips and ES cells have. What genes are special in stem cells?
 Do engineered ips and ES cells have similar molecular signatures? What we ll do today Research questions in stem cell biology Comparing expression and epigenetics in stem cells asuring gene expression
Do engineered ips and ES cells have similar molecular signatures? What we ll do today Research questions in stem cell biology Comparing expression and epigenetics in stem cells asuring gene expression
Do engineered ips and ES cells have similar molecular signatures?
 Do engineered ips and ES cells have similar molecular signatures? Comparing expression and epigenetics in stem cells George Bell, Ph.D. Bioinformatics and Research Computing 2012 Spring Lecture Series
Do engineered ips and ES cells have similar molecular signatures? Comparing expression and epigenetics in stem cells George Bell, Ph.D. Bioinformatics and Research Computing 2012 Spring Lecture Series
RNA-Sequencing analysis
 RNA-Sequencing analysis Markus Kreuz 25. 04. 2012 Institut für Medizinische Informatik, Statistik und Epidemiologie Content: Biological background Overview transcriptomics RNA-Seq RNA-Seq technology Challenges
RNA-Sequencing analysis Markus Kreuz 25. 04. 2012 Institut für Medizinische Informatik, Statistik und Epidemiologie Content: Biological background Overview transcriptomics RNA-Seq RNA-Seq technology Challenges
Gene Prediction: Preliminary Results
 Gene Prediction: Preliminary Results Outline Preliminary Pipeline Programs Program Comparison Tests Metrics Gene Prediction Tools: Usage + Results GeneMarkS Glimmer 3.0 Prodigal BLAST ncrna Prediction
Gene Prediction: Preliminary Results Outline Preliminary Pipeline Programs Program Comparison Tests Metrics Gene Prediction Tools: Usage + Results GeneMarkS Glimmer 3.0 Prodigal BLAST ncrna Prediction
Guided tour to Ensembl
 Guided tour to Ensembl Introduction Introduction to the Ensembl project Walk-through of the browser Variations and Functional Genomics Comparative Genomics BioMart Ensembl Genome browser http://www.ensembl.org
Guided tour to Ensembl Introduction Introduction to the Ensembl project Walk-through of the browser Variations and Functional Genomics Comparative Genomics BioMart Ensembl Genome browser http://www.ensembl.org
user s guide Question 3
 Question 3 During a positional cloning project aimed at finding a human disease gene, linkage data have been obtained suggesting that the gene of interest lies between two sequence-tagged site markers.
Question 3 During a positional cloning project aimed at finding a human disease gene, linkage data have been obtained suggesting that the gene of interest lies between two sequence-tagged site markers.
Lecture 11: Gene Prediction
 Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
 Genome Annotation Genome annotation What is the function of each part of the genome? Where are the genes? What is the mrna sequence (transcription, splicing) What is the protein sequence? What does
Genome Annotation Genome annotation What is the function of each part of the genome? Where are the genes? What is the mrna sequence (transcription, splicing) What is the protein sequence? What does
Introduction to the UCSC genome browser
 Introduction to the UCSC genome browser Dominik Beck NHMRC Peter Doherty and CINSW ECR Fellow, Senior Lecturer Lowy Cancer Research Centre, UNSW and Centre for Health Technology, UTS SYDNEY NSW AUSTRALIA
Introduction to the UCSC genome browser Dominik Beck NHMRC Peter Doherty and CINSW ECR Fellow, Senior Lecturer Lowy Cancer Research Centre, UNSW and Centre for Health Technology, UTS SYDNEY NSW AUSTRALIA
Training materials.
 Training materials Ensembl training materials are protected by a CC BY license http://creativecommons.org/licenses/by/4.0/ If you wish to re-use these materials, please credit Ensembl for their creation
Training materials Ensembl training materials are protected by a CC BY license http://creativecommons.org/licenses/by/4.0/ If you wish to re-use these materials, please credit Ensembl for their creation
Introduction to Bioinformatics
 Introduction to Bioinformatics Changhui (Charles) Yan Old Main 401 F http://www.cs.usu.edu www.cs.usu.edu/~cyan 1 How Old Is The Discipline? "The term bioinformatics is a relatively recent invention, not
Introduction to Bioinformatics Changhui (Charles) Yan Old Main 401 F http://www.cs.usu.edu www.cs.usu.edu/~cyan 1 How Old Is The Discipline? "The term bioinformatics is a relatively recent invention, not
Sequence Based Function Annotation. Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University
 Sequence Based Function Annotation Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Usage scenarios for sequence based function annotation Function prediction of newly cloned
Sequence Based Function Annotation Qi Sun Bioinformatics Facility Biotechnology Resource Center Cornell University Usage scenarios for sequence based function annotation Function prediction of newly cloned
Integration of data management and analysis for genome research
 Integration of data management and analysis for genome research Volker Brendel Deparment of Zoology & Genetics and Department of Statistics Iowa State University 2112 Molecular Biology Building Ames, Iowa
Integration of data management and analysis for genome research Volker Brendel Deparment of Zoology & Genetics and Department of Statistics Iowa State University 2112 Molecular Biology Building Ames, Iowa
Chapter 12 Packet DNA 1. What did Griffith conclude from his experiment? 2. Describe the process of transformation.
 Chapter 12 Packet DNA and RNA Name Period California State Standards covered by this chapter: Cell Biology 1. The fundamental life processes of plants and animals depend on a variety of chemical reactions
Chapter 12 Packet DNA and RNA Name Period California State Standards covered by this chapter: Cell Biology 1. The fundamental life processes of plants and animals depend on a variety of chemical reactions
Using geneid to Identify Genes
 Using geneid to Identify Genes UNIT 4.3 The gene-prediction program geneid is based on a simple hierarchical design: (1) search splicing signals, start codons, and stop codons, (2) build and score candidate
Using geneid to Identify Genes UNIT 4.3 The gene-prediction program geneid is based on a simple hierarchical design: (1) search splicing signals, start codons, and stop codons, (2) build and score candidate
Multiple choice questions (numbers in brackets indicate the number of correct answers)
 1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
1 Multiple choice questions (numbers in brackets indicate the number of correct answers) February 1, 2013 1. Ribose is found in Nucleic acids Proteins Lipids RNA DNA (2) 2. Most RNA in cells is transfer
RNA Genomics II. BME 110: CompBio Tools Todd Lowe & Andrew Uzilov May 17, 2011
 RNA Genomics II BME 110: CompBio Tools Todd Lowe & Andrew Uzilov May 17, 2011 1 TIME Why RNA? An evolutionary perspective The RNA World hypotheses: life arose as self-replicating non-coding RNA (ncrna)
RNA Genomics II BME 110: CompBio Tools Todd Lowe & Andrew Uzilov May 17, 2011 1 TIME Why RNA? An evolutionary perspective The RNA World hypotheses: life arose as self-replicating non-coding RNA (ncrna)
DNA makes RNA makes Proteins. The Central Dogma
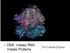 DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
user s guide Question 3
 Question 3 During a positional cloning project aimed at finding a human disease gene, linkage data have been obtained suggesting that the gene of interest lies between two sequence-tagged site markers.
Question 3 During a positional cloning project aimed at finding a human disease gene, linkage data have been obtained suggesting that the gene of interest lies between two sequence-tagged site markers.
Computational analysis of non-coding RNA. Andrew Uzilov BME110 Tue, Nov 16, 2010
 Computational analysis of non-coding RNA Andrew Uzilov auzilov@ucsc.edu BME110 Tue, Nov 16, 2010 1 Corrected/updated talk slides are here: http://tinyurl.com/uzilovrna redirects to: http://users.soe.ucsc.edu/~auzilov/bme110/fall2010/
Computational analysis of non-coding RNA Andrew Uzilov auzilov@ucsc.edu BME110 Tue, Nov 16, 2010 1 Corrected/updated talk slides are here: http://tinyurl.com/uzilovrna redirects to: http://users.soe.ucsc.edu/~auzilov/bme110/fall2010/
Themes: RNA and RNA Processing. Messenger RNA (mrna) What is a gene? RNA is very versatile! RNA-RNA interactions are very important!
 Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Hands-On Four Investigating Inherited Diseases
 Hands-On Four Investigating Inherited Diseases The purpose of these exercises is to introduce bioinformatics databases and tools. We investigate an important human gene and see how mutations give rise
Hands-On Four Investigating Inherited Diseases The purpose of these exercises is to introduce bioinformatics databases and tools. We investigate an important human gene and see how mutations give rise
SSA Signal Search Analysis II
 SSA Signal Search Analysis II SSA other applications - translation In contrast to translation initiation in bacteria, translation initiation in eukaryotes is not guided by a Shine-Dalgarno like motif.
SSA Signal Search Analysis II SSA other applications - translation In contrast to translation initiation in bacteria, translation initiation in eukaryotes is not guided by a Shine-Dalgarno like motif.
MATH 5610, Computational Biology
 MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
MATH 5610, Computational Biology Lecture 2 Intro to Molecular Biology (cont) Stephen Billups University of Colorado at Denver MATH 5610, Computational Biology p.1/24 Announcements Error on syllabus Class
Big Idea 3C Basic Review
 Big Idea 3C Basic Review 1. A gene is a. A sequence of DNA that codes for a protein. b. A sequence of amino acids that codes for a protein. c. A sequence of codons that code for nucleic acids. d. The end
Big Idea 3C Basic Review 1. A gene is a. A sequence of DNA that codes for a protein. b. A sequence of amino acids that codes for a protein. c. A sequence of codons that code for nucleic acids. d. The end
Bioinformatics in next generation sequencing projects
 Bioinformatics in next generation sequencing projects Rickard Sandberg Assistant Professor Department of Cell and Molecular Biology Karolinska Institutet May 2013 Standard sequence library generation Illumina
Bioinformatics in next generation sequencing projects Rickard Sandberg Assistant Professor Department of Cell and Molecular Biology Karolinska Institutet May 2013 Standard sequence library generation Illumina
Browsing Genes and Genomes with Ensembl
 Browsing Genes and Genomes with Ensembl Victoria Newman Ensembl Outreach Officer EMBL-EBI Objectives What is Ensembl? What type of data can you get in Ensembl? How to navigate the Ensembl browser website.
Browsing Genes and Genomes with Ensembl Victoria Newman Ensembl Outreach Officer EMBL-EBI Objectives What is Ensembl? What type of data can you get in Ensembl? How to navigate the Ensembl browser website.
Gene Prediction Chengwei Luo, Amanda McCook, Nadeem Bulsara, Phillip Lee, Neha Gupta, and Divya Anjan Kumar
 Gene Prediction Chengwei Luo, Amanda McCook, Nadeem Bulsara, Phillip Lee, Neha Gupta, and Divya Anjan Kumar Gene Prediction Introduction Protein-coding gene prediction RNA gene prediction Modification
Gene Prediction Chengwei Luo, Amanda McCook, Nadeem Bulsara, Phillip Lee, Neha Gupta, and Divya Anjan Kumar Gene Prediction Introduction Protein-coding gene prediction RNA gene prediction Modification
Gene Signal Estimates from Exon Arrays
 Gene Signal Estimates from Exon Arrays I. Introduction: With exon arrays like the GeneChip Human Exon 1.0 ST Array, researchers can examine the transcriptional profile of an entire gene (Figure 1). Being
Gene Signal Estimates from Exon Arrays I. Introduction: With exon arrays like the GeneChip Human Exon 1.0 ST Array, researchers can examine the transcriptional profile of an entire gene (Figure 1). Being
RNA Genomics. BME 110: CompBio Tools Todd Lowe May 14, 2010
 RNA Genomics BME 110: CompBio Tools Todd Lowe May 14, 2010 Admin WebCT quiz on Tuesday cover reading, using Jalview & Pfam Homework #3 assigned today due next Friday (8 days) In Genomes, Two Types of Genes
RNA Genomics BME 110: CompBio Tools Todd Lowe May 14, 2010 Admin WebCT quiz on Tuesday cover reading, using Jalview & Pfam Homework #3 assigned today due next Friday (8 days) In Genomes, Two Types of Genes
Wednesday, November 22, 17. Exons and Introns
 Exons and Introns Introns and Exons Exons: coded regions of DNA that get transcribed and translated into proteins make up 5% of the genome Introns and Exons Introns: non-coded regions of DNA Must be removed
Exons and Introns Introns and Exons Exons: coded regions of DNA that get transcribed and translated into proteins make up 5% of the genome Introns and Exons Introns: non-coded regions of DNA Must be removed
Gene Structure & Gene Finding Part II
 Gene Structure & Gene Finding Part II David Wishart david.wishart@ualberta.ca 30,000 metabolite Gene Finding in Eukaryotes Eukaryotes Complex gene structure Large genomes (0.1 to 10 billion bp) Exons and
Gene Structure & Gene Finding Part II David Wishart david.wishart@ualberta.ca 30,000 metabolite Gene Finding in Eukaryotes Eukaryotes Complex gene structure Large genomes (0.1 to 10 billion bp) Exons and
BIOINFORMATICS Introduction
 BIOINFORMATICS Introduction Mark Gerstein, Yale University bioinfo.mbb.yale.edu/mbb452a 1 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu What is Bioinformatics? (Molecular) Bio -informatics One idea
BIOINFORMATICS Introduction Mark Gerstein, Yale University bioinfo.mbb.yale.edu/mbb452a 1 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu What is Bioinformatics? (Molecular) Bio -informatics One idea
Motif Discovery from Large Number of Sequences: a Case Study with Disease Resistance Genes in Arabidopsis thaliana
 Motif Discovery from Large Number of Sequences: a Case Study with Disease Resistance Genes in Arabidopsis thaliana Irfan Gunduz, Sihui Zhao, Mehmet Dalkilic and Sun Kim Indiana University, School of Informatics
Motif Discovery from Large Number of Sequences: a Case Study with Disease Resistance Genes in Arabidopsis thaliana Irfan Gunduz, Sihui Zhao, Mehmet Dalkilic and Sun Kim Indiana University, School of Informatics
Product Applications for the Sequence Analysis Collection
 Product Applications for the Sequence Analysis Collection Pipeline Pilot Contents Introduction... 1 Pipeline Pilot and Bioinformatics... 2 Sequence Searching with Profile HMM...2 Integrating Data in a
Product Applications for the Sequence Analysis Collection Pipeline Pilot Contents Introduction... 1 Pipeline Pilot and Bioinformatics... 2 Sequence Searching with Profile HMM...2 Integrating Data in a
Introduction to RNA sequencing
 Introduction to RNA sequencing Bioinformatics perspective Olga Dethlefsen NBIS, National Bioinformatics Infrastructure Sweden November 2017 Olga (NBIS) RNA-seq November 2017 1 / 49 Outline Why sequence
Introduction to RNA sequencing Bioinformatics perspective Olga Dethlefsen NBIS, National Bioinformatics Infrastructure Sweden November 2017 Olga (NBIS) RNA-seq November 2017 1 / 49 Outline Why sequence
Computational Biology and Bioinformatics
 Computational Biology and Bioinformatics Computational biology Development of algorithms to solve problems in biology Bioinformatics Application of computational biology to the analysis and management
Computational Biology and Bioinformatics Computational biology Development of algorithms to solve problems in biology Bioinformatics Application of computational biology to the analysis and management
Glossary of Commonly used Annotation Terms
 Glossary of Commonly used Annotation Terms Akela a general use server for the annotation group as well as other groups throughout TIGR. Annotation Notebook a link from the gene list page that is associated
Glossary of Commonly used Annotation Terms Akela a general use server for the annotation group as well as other groups throughout TIGR. Annotation Notebook a link from the gene list page that is associated
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Types of Databases - By Scope
 Biological Databases Bioinformatics Workshop 2009 Chi-Cheng Lin, Ph.D. Department of Computer Science Winona State University clin@winona.edu Biological Databases Data Domains - By Scope - By Level of
Biological Databases Bioinformatics Workshop 2009 Chi-Cheng Lin, Ph.D. Department of Computer Science Winona State University clin@winona.edu Biological Databases Data Domains - By Scope - By Level of
Assessing De-Novo Transcriptome Assemblies
 Assessing De-Novo Transcriptome Assemblies Shawn T. O Neil Center for Genome Research and Biocomputing Oregon State University Scott J. Emrich University of Notre Dame 100K Contigs, Perfect 1M Contigs,
Assessing De-Novo Transcriptome Assemblies Shawn T. O Neil Center for Genome Research and Biocomputing Oregon State University Scott J. Emrich University of Notre Dame 100K Contigs, Perfect 1M Contigs,
Improved annotation with de novo transcriptome assembly in four social amoeba species
 Singh et al. BMC Genomics (2017) 18:120 DOI 10.1186/s12864-017-3505-0 RESEARCH ARTICLE Improved annotation with de novo transcriptome assembly in four social amoeba species Open Access Reema Singh 1,2,
Singh et al. BMC Genomics (2017) 18:120 DOI 10.1186/s12864-017-3505-0 RESEARCH ARTICLE Improved annotation with de novo transcriptome assembly in four social amoeba species Open Access Reema Singh 1,2,
Introduction to Bioinformatics CPSC 265. What is bioinformatics? Textbooks
 Introduction to Bioinformatics CPSC 265 Thanks to Jonathan Pevsner, Ph.D. Textbooks Johnathan Pevsner, who I stole most of these slides from (thanks!) has written a textbook, Bioinformatics and Functional
Introduction to Bioinformatics CPSC 265 Thanks to Jonathan Pevsner, Ph.D. Textbooks Johnathan Pevsner, who I stole most of these slides from (thanks!) has written a textbook, Bioinformatics and Functional
Sanger vs Next-Gen Sequencing
 Tools and Algorithms in Bioinformatics GCBA815/MCGB815/BMI815, Fall 2017 Week-8: Next-Gen Sequencing RNA-seq Data Analysis Babu Guda, Ph.D. Professor, Genetics, Cell Biology & Anatomy Director, Bioinformatics
Tools and Algorithms in Bioinformatics GCBA815/MCGB815/BMI815, Fall 2017 Week-8: Next-Gen Sequencing RNA-seq Data Analysis Babu Guda, Ph.D. Professor, Genetics, Cell Biology & Anatomy Director, Bioinformatics
Gene Prediction: Statistical Approaches
 Gene Prediction: Statistical Approaches Outline Codons Discovery of Split Genes Exons and Introns Splicing Open Reading Frames Codon Usage Splicing Signals TestCode Gene Prediction: Computational Challenge
Gene Prediction: Statistical Approaches Outline Codons Discovery of Split Genes Exons and Introns Splicing Open Reading Frames Codon Usage Splicing Signals TestCode Gene Prediction: Computational Challenge
Protein Synthesis & Gene Expression
 DNA provides the instructions for how to build proteins Each gene dictates how to build a single protein in prokaryotes The sequence of nucleotides (AGCT) in DNA dictates the order of amino acids that
DNA provides the instructions for how to build proteins Each gene dictates how to build a single protein in prokaryotes The sequence of nucleotides (AGCT) in DNA dictates the order of amino acids that
Genes and gene finding
 Genes and gene finding Ben Langmead Department of Computer Science You are free to use these slides. If you do, please sign the guestbook (www.langmead-lab.org/teaching-materials), or email me (ben.langmead@gmail.com)
Genes and gene finding Ben Langmead Department of Computer Science You are free to use these slides. If you do, please sign the guestbook (www.langmead-lab.org/teaching-materials), or email me (ben.langmead@gmail.com)
measuring gene expression December 5, 2017
 measuring gene expression December 5, 2017 transcription a usually short-lived RNA copy of the DNA is created through transcription RNA is exported to the cytoplasm to encode proteins some types of RNA
measuring gene expression December 5, 2017 transcription a usually short-lived RNA copy of the DNA is created through transcription RNA is exported to the cytoplasm to encode proteins some types of RNA
Reference genomes and common file formats
 Reference genomes and common file formats Overview Reference genomes and GRC Fasta and FastQ (unaligned sequences) SAM/BAM (aligned sequences) Summarized genomic features BED (genomic intervals) GFF/GTF
Reference genomes and common file formats Overview Reference genomes and GRC Fasta and FastQ (unaligned sequences) SAM/BAM (aligned sequences) Summarized genomic features BED (genomic intervals) GFF/GTF
MAKING WHOLE GENOME ALIGNMENTS USABLE FOR BIOLOGISTS. EXAMPLES AND SAMPLE ANALYSES.
 MAKING WHOLE GENOME ALIGNMENTS USABLE FOR BIOLOGISTS. EXAMPLES AND SAMPLE ANALYSES. Table of Contents Examples 1 Sample Analyses 5 Examples: Introduction to Examples While these examples can be followed
MAKING WHOLE GENOME ALIGNMENTS USABLE FOR BIOLOGISTS. EXAMPLES AND SAMPLE ANALYSES. Table of Contents Examples 1 Sample Analyses 5 Examples: Introduction to Examples While these examples can be followed
