Proteomics and Mass Spectrometry
|
|
|
- Ashlie Sullivan
- 5 years ago
- Views:
Transcription
1 Molecular Cell Biology Lecture. Oct. 18, 2018 Proteomics and Mass Spectrometry Ron Bose, MD PhD Biochemistry and Molecular Cell Biology Programs Lab: Couch Research Building (4515 McKinley), 3 rd floor Washington University School of Medicine
2 Introduction Definition of Proteomics: The large scale identification and characterization of proteins in a cell, tissue, or organism. Traditional Biochemistry Proteomics
3 Introduction Definition of Proteomics: The large scale identification and characterization of proteins in a cell, tissue, or organism. Well Established Methods for Proteomics 1. 2D-gels 2. Mass Spectrometry Methods still under development 1. Protein Arrays 2. Antibody Arrays 3. Proteome-wide coverage with Antibodies
4 Size 2 Dimensional Gel Electrophoresis First Dimension: pi by Isoelectric Focusing Second Dimension: MW by standard SDS-PAGE First Published in 1975 by Pat O Farrell Can separate at least 1,000 proteins Problems with run to run reproducibility limits the ability to easily compare multiple samples. Charge (pi) Solution to this problem: DIGE (Difference Imaging Gel Electrophoresis)
5 DIGE experiment Slide courtesy of Tracy Andacht
6 DIGE experiment Data from the labs of Tim Ley and Reid Townsend Bredemeyer et al., PNAS 101:11785, 2004
7 1. Protein solubility during Isoelectric Focusing. Membrane proteins often lost. 2. Size Limits difficulty with proteins >100 kd. 3. Identification of the proteins in each spot is tedious and slow. Use of robotics Limitations of DIGE 4. Individual spots typically contain several proteins. Intensity change is therefore the sum of the changes of each individual protein.
8 Principles of Mass Spectrometry The Importance of Mass: 1. The mass of a molecule is a fundamental physical property of a molecule. 2. Mass can be used to identify the molecule. Fragmentation provides Chemical Structure: If you fragment a molecule in a predictable manner and make measurements on the individual fragments, you can determine the chemical structure of the molecule.
9 1. Peptides and Proteins 2. Lipids Biological Applications of Mass Spectrometry 3. Oligosaccharides MALDI-TOF spectrum of a synthesized 25mer peptide. Measured mass= Da Calculated mass= Da
10 1. Peptides and Proteins 2. Lipids Biological Applications of Mass Spectrometry 3. Oligosaccharides Methodology to identify lipids by mass spectrometry. X. Han & R.W. Gross, Expert Review Proteomics 2:253, 2005
11 1. Peptides and Proteins 2. Lipids Biological Applications of Mass Spectrometry 3. Oligosaccharides: Analysis of Milk Tao et al., J. Dairy Sci 91:3768, 2008
12 Applications of Mass Spectrometry in the Physical Sciences Widely used in Analytical Chemistry and Organic Chemistry. Examples: Analyzing of drugs during chemical synthesis Identifying chemicals molecules or checking for contaminants. Environmental Measuring toxins such as PCB and Heavy Metals Geology Analyzing petroleum or petrochemicals
13 Applications of Mass Spectrometry in the Physical Sciences Space Exploration: Mars Curiosity Rover Sources: and Los Alamos National Laboratory
14 Applications of Mass Spectrometry in the Physical Sciences Space Exploration: Mars Curiosity Rover Sample Analysis at Mars (SAM) Instrument Suite 1. Mass Spectrometer 2. Gas Chromatograph 3. Laser Spectrometer Sources: and Los Alamos National Laboratory
15 Applications of Mass Spectrometry in the Physical Sciences Undersea Exploration: Deep Water Horizon Spill
16 Applications of Mass Spectrometry in the Physical Sciences Undersea Exploration: Deep Water Horizon Spill Scientific instruments used to measure the oil spill, including Mass Spectrometers for chemical analysis.
17 Applications of Mass Spectrometry in the Physical Sciences Anti Terrorism and Civil Defense: IonScan Mass Spectrometry Used at Airports and other facilities for the detection of Explosives and Narcotics. Manufacturer: Smiths Detection
18 Identifying a Protein by Mass Spectrometry on Its Tryptic Peptides Trypsin a protease that cleaves after basic residues (R or K). Protein of Interest: Slide courtesy of Andrew Link
19 Identifying a Protein by Mass Spectrometry on Its Tryptic Peptides Products from Trypsin digest. Average length of tryptic peptides = 10 aa residues Slide courtesy of Andrew Link
20 Identifying a Protein by Mass Spectrometry on Its Tryptic Peptides Select an Individual Peptide in the Mass Spectrometer Performed by adjusting the electrical fields in the mass spectrometer. Slide courtesy of Andrew Link
21 Identifying a Protein by Mass Spectrometry on Its Tryptic Peptides Impart energy to the peptide by colliding it with an inert gas (Argon or Helium). Slide courtesy of Andrew Link
22 Identifying a Protein by Mass Spectrometry on Its Tryptic Peptides Measure the masses of the fragment ions. Slide courtesy of Andrew Link
23 Identifying a Protein by Mass Spectrometry on Its Tryptic Peptides The mass difference between the peaks corresponds directly to the amino acid sequence. B-ions contain the N- terminus Slide courtesy of Andrew Link
24 Identifying a Protein by Mass Spectrometry on Its Tryptic Peptides Y-ions contain the C-terminus Slide courtesy of Andrew Link
25 Identifying a Protein by Mass Spectrometry on Its Tryptic Peptides The entire spectrum contains B-ions,Y-ions, and other fragment ions. Slide courtesy of Andrew Link
26 Identifying a Protein by Mass Spectrometry on Its Tryptic Peptides The puzzle: The B, Y, and other ions occur together and we cannot distinguish them just by simple inspection of the spectrum. Slide courtesy of Andrew Link
27 Identifying a Protein by Mass Spectrometry on Its Tryptic Peptides Actual spectra also have noise (either chemical noise or electrical noise). Slide courtesy of Andrew Link
28 Identifying a Protein by Mass Spectrometry on Its Tryptic Peptides The final spectrum: the interpretation requires experience and aid by software algorithms. Slide courtesy of Andrew Link
29 Software for Interpreting Peptide Mass Spectra Statistical Matching Work by statistically matching the measured spectra with the theoretical spectra of all possible tryptic peptides from an organism. 1. SeQuest 2. MASCOT 3. X! Tandem 4. OMSSA Requires a fully sequenced genome. De novo sequencing (determines a peptide sequence based on the spacings of the fragment ions). 1. PepNovo
30 Example of an Actual Spectrum Gross_9309HER4_8 #4181 RT: AV: 1 NL: 1.75E4 T: ITMS + c NSI d w Full ms @cid30.00 [ ] B Y 1 D py Imm Y D B 2 Y 3 G Y Q Y I m/z Y 6 V Peptide with phosphorylation on Y Y 7 L pylviqgddr Y 8
31 The Hardware for Peptide Mass Spectrometry Liquid Chromatography Pump Vacuum Pump Ionization Source Mass Analyzer Detector Different Types: Electrospray MALDI Time of Flight (TOF) Quadropole Ion Trap OrbiTrap Ion Cyclotron Resonance (ICR) Output: Spectra
32 The Hardware for Peptide Mass Spectrometry Liquid Chromatography Ionization Source Mass Analyzer and Detector Vacuum Pumps
33 Movie of MALDI TOF mass spectrometer. Movie of FT-ICR mass spectrometer.
34 Drilling Down to Low Abundance Proteins Limitations and Cautions of Proteomics: The Range of Protein Concentrations In Yeast Picotti et al., Cell Aug 21, 2009
35 Limitations and Cautions of Proteomics: The Range of Protein Concentrations In Human Plasma 3-4 log range of Mass Spectrometers Albumin 40 g/lc4 Complement 0.1 g/l Myoglobin < 100 mg/l TNFa < 1 ng/l Anderson & Anderson, MCP 1:845, 2002
36 Limitations and Cautions of Proteomics: The Range of Protein Concentrations In Human Plasma Depletion Remove abundant proteins that are not of interest to your experiment. Methods: Antibody based depletion, selective lysis technique, subcellular fractionation, etc. Enrichment Enrich for the proteins of interest. Methods Lysis techniques or subcellular fractionation, affinity-based enrichment (antibodies, resins, etc). Fractionation Reduce the complexity of your sample by separating the proteins into different fractions and running these fractions separately.
37 Examples of Proteomic Experiments 1. Identification of Single Proteins 2. Identification of Proteins in the Nuclear Pore Complex 3. Identification of Proteins in the Secretory Pathway 4. Quantitative Measurement of Signal Transduction Pathways
38 Identification of Proteins in Single Bands Mary Olanich, a graduate student in Jason Weber lab, wanted to identify proteins binding to the untranslated regions (UTR) of the NPM mrna. She performed a pull-down assay with biotinylated NPM mrna. Protein bands obtained were visualized with a fluorescent protein stain. Single bands were cut from the gel and proteins ID ed by MS. Olanich et al., Oncogene 30(1):77-86, 2011.
39 ID of Nuclear Pore Complex Proteins Yeast Nuclear Pore Complexes are 50 MDa in size. Contain approximately 30 different proteins. Total number of proteins in the NPC is at least 456. Side View Top View Alber et al., Nature 450: , 2007 Yamada et al., Mol. Cell Proteomics 9: , 2010
40 Strategy to Identify NPC Proteins 1. Make a highly pure NPC prepation 2. Extensive fractionation and Mass Spec protein identification. 3. Validate results with: a. Immunofluorescence b. Epitope tagging c. Immuno-electron microscopy Rout et al., J Cell Bio 148: , 2000
41 Strategy to Identify NPC Proteins Hydroxyapatite Column Separation 200 kd 116 kd 97 kd 66 kd 45 kd 31 kd 21 kd 14 kd 6 kd Blue = Known NPC associating proteins Red = Proteins believed not to be NPC associated Rout et al., J Cell Bio 148: , 2000
42 Rout et al., J Cell Bio 148: , 2000 Strategy to Identify NPC Proteins Each band was cut out and digested with trypsin. Mass Spec analysis was done by looking at the MS spectra and the MS/MS spectra. MS spectrum of a mixture of 3 yeast proteins, all about 120 kd size, and trypsin auto-digestion peptides (marked by T). Each peak can be isolated in the Mass Spectrometer and then fragmented to give MS/MS spectra and peptide sequence information.
43 Identification of Secretory Pathway Proteins Started with a high quality preparation of Rough Microsomes (RM), Smooth Microsomes (SM), and Golgi apparatus (G). Fractionate the proteins on SDS-PAGE, cut thin slices of gel, digest with trypsin and run on Mass Spec. Gilchrist et al., Cell 127: , 2006
44 Identification of Secretory Pathway Proteins They identified over 1400 proteins and divided them into 23 functional categories. Semi-quantitative measurements of protein abundance were made by spectral counting (ie the number of observed spectra for a protein correlates with its abundance). Gilchrist et al., Cell 127: , 2006
45 Protein Quantitation with Mass Spectrometry In Western blots, each antigenantibody pair has a different affinity and response characteristics. Therefore, we can make comparison protein A in sample 1 vs.2 vs. 3, but not protein A vs. protein B in the same sample. Similarly, in Mass Spec, every peptide has its own ionization and detection characteristics. Protein A Protein B Protein C Sample 1 2 3
46 Protein Quantitation with Mass Spectrometry 1. Stable Isotope Labels based Quantitation Examples of Stable Isotopes: 13 C, 15 N, 2 H, 18 O Advantage of Stable Isotopes: They are easy separated and distinguished in the Mass Spec. Approach: An internal comparison within one Mass Spec run. Different samples can be labeled with different isotopes. Advantages: Precision of quantitation, less susceptible to artifacts in Mass Spec runs. Limitations: Cost of isotopes. Limited number of isotope combinations are feasible. 2. Label-free Quantitation No isotopes used.
47 Please, Consider the Following: Isotopes of Carbon Isotope Mass Abund ance in Nature 12 C 12 exactly Halflife Radioa ctivity release 98.9% Stable None 13 C % Stable None 14 C Trace 5,700 years b particle 11 C Nonnatural 20 min positron
48 Please, Consider the Following: Isotopes of Carbon Isotope Mass Abund ance in Nature 12 C 12 exactly Halflife Radioa ctivity release 98.9% Stable None 13 C % Stable None Commonly used in Mass Spectrometry for Quantitative Measurements 14 C Trace 5,700 years 11 C Nonnatural b particle 20 min positron DO NOT USE IN MASS SPEC.
49 Protein Quantitation with Mass Spectrometry Introduce Stable Isotope by Metabolic Labeling Control Treatment 1 Treatment 2 Mix Lysates Fractionate Proteins on SDS-PAGE Digest Bands with Trypsin Identify and Quantify Proteins by Mass Spec Bose et al., PNAS 103: , 2006
50 Protein Quantitation with Mass Spectrometry Protein VGQAQDILR Protein VAGQSSPSGIQSR FFEILSPVYR HDGAFLIR Protein Protein Key +0 Control 12 C-Arginine +6 Treatment 1 13 C 6 -Arginine +10 Treatment 2 13 C 15 6 N 4 -Arginine Bose et al., PNAS 103: , 2006
51 Protein Quantification with Mass Spectrometry Introduce Stable Isotope by Chemical Labeling Amine reactive tags itraq (Ross et al., MCP 3:1154, 2003) Cys reactive tags - ICAT Incorporating 18 O during Trypsin digestion
52 Studying EGFR Signal Transduction with Quantitative Proteomics Introduce Stable Isotope by Chemical Labeling Zhang et al., MCP 4: , 2005
53 Empty Vector Her2/neu Vehicle Gefitinib 1 mm Mapping Her2/neu Tyrosine Kinase Signaling using Quantitative Proteomics A. B. Her2 inhibitor (mm) kd 150 kd WB: Anti-pTyr 100 kd 75 kd Bose et al., PNAS 103:9773, 2006
54 Mapping Her2/neu Tyrosine Kinase Signaling using Quantitative Proteomics Empty vector cells Her2/neu cells +Her2 kinase inhibitor Her2/neu cells Mix Lysates Immunoaffinity Purify with Antiphosphotyrosine Antibodies Resolve on SDS-PAGE Digest Bands with Trypsin Identify and Quantify Proteins by LC-MS/MS Bose et al., PNAS 103:9773, 2006
55 Fold Change with Her2/neu Number of Proteins SILAC Quantitation of Protein Phosphorylation A. Fold Change with Her2/neu Mapping Her2/neu Tyrosine Kinase Signaling using Quantitative Proteomics Her2/neu B. Fold Change with Her2 kinase inhibitor 83 9 Fold Inhibition > No Change <0.5 Ratio 5 4 Fyb/ADAP Dok1 & STAT Axl & PLCg PI3kinase p85b subunit Number Protein of Proteins > <0.66 Fold Change with Her2/neu Series1 Series2 Series3 Bose et al., PNAS 103:9773, 2006
56 Mapping Her2/neu Tyrosine Kinase Signaling using Quantitative Proteomics Bose et al., PNAS 103:9773, 2006
57 Mapping Her2/neu Tyrosine Kinase Signaling using Quantitative Proteomics Bose et al., PNAS 103:9773, 2006
58 Mapping Her2/neu Tyrosine Kinase Signaling using Quantitative Proteomics Bayesian Network Analysis of Proteomic Results Bose et al., PNAS 103:9773, 2006
59 Studying Toll-Like Receptor Signaling in Macrophages using Quantitative Proteomics Results Identified 6900 phosphorylation sites on 1850 proteins. Changes with LPS: 24% of sites increased. 9% of sites decreased. Bone Marrow derived Macrophages + LPS (activator of TLR4) Measured the phosphorylation of 187 proteins annotated as transcriptional regulators. They linked proteomics measurements with changes in gene expression. Weintz et al., MSB 6:371, 2010
60 Studying Toll-Like Receptor Signaling in Macrophages using Quantitative Proteomics Weintz et al., MSB 6:371, 2010
61 Limitations and Cautions: Sizes of Proteomic Experiments A Medium sized Proteomic Experiment: Several hundred proteins time required: Months A Large Proteomic Experiment: A few thousand proteins time required: 1-3 YEARS. Proteomics cannot currently analyze as many genes as DNA microarray technology can! Proteomics is also highly technically demanding and often requires a lot of optimization and small scale testing before performing a large experiment.
62 Mass Spectrometry at Washington University Wash U receives NIH funding for the Biological and Biomedical Mass Spectrometry Research Resource. At least 8 labs at Wash U. perform biological mass spectrometry experiments. Available instruments on the Wash U medical campus, Wash U Danforth campus, and the Danforth Plant Science Center include: At least 30 mass spectrometers. 5 LTQ-OrbiTrap mass spectrometers (some of the latest and highest performance instruments).
63 Summary (Part 1) 1. There is wide spread use of mass spectrometry in both the biological and physical sciences. 2. Proteins are usually digested into peptides. Peptide sequence is determined by fragmentation in the Mass Spectrometer. 3. Protein abundance and enrichment or fractionation methods are critical to consider in the planning of proteomic experiments. 4. Proteomics can identify proteins and map their posttranslational modifications. Components of protein complexes and intracellular pathways can be analyzed by proteomics.
64 Summary (Part 2) 5. Quantitative proteomics can be performed by incorporating stable isotopes into proteins or by using label-free quantitation methods. 6. Proteomics cannot analyze as many genes as DNA microarray technology. Further, proteomics is highly technically demanding and often requires a lot of optimization. 7. Many labs at Wash U. use mass spec and proteomics. Wash U. has a lot of the necessary equipment and expertise to conduct mass spectrometry experiments.
Proteomics and some of its Mass Spectrometric Applications
 Proteomics and some of its Mass Spectrometric Applications What? Large scale screening of proteins, their expression, modifications and interactions by using high-throughput approaches 2 1 Why? The number
Proteomics and some of its Mass Spectrometric Applications What? Large scale screening of proteins, their expression, modifications and interactions by using high-throughput approaches 2 1 Why? The number
Computing with large data sets
 Computing with large data sets Richard Bonneau, spring 2009 Lecture 14 (week 8): genomics 1 Central dogma Gene expression DNA RNA Protein v22.0480: computing with data, Richard Bonneau Lecture 14 places
Computing with large data sets Richard Bonneau, spring 2009 Lecture 14 (week 8): genomics 1 Central dogma Gene expression DNA RNA Protein v22.0480: computing with data, Richard Bonneau Lecture 14 places
MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1. Lecture 25: Mass Spectrometry Applications
![MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1. Lecture 25: Mass Spectrometry Applications MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1. Lecture 25: Mass Spectrometry Applications](/thumbs/74/71368870.jpg) MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1 Lecture 25: Mass Spectrometry Applications Measuring Protein Abundance o ICAT o DIGE Identifying Post-Translational Modifications Protein-protein
MBios 478: Mass Spectrometry Applications [Dr. Wyrick] Slide #1 Lecture 25: Mass Spectrometry Applications Measuring Protein Abundance o ICAT o DIGE Identifying Post-Translational Modifications Protein-protein
Exam MOL3007 Functional Genomics
 Faculty of Medicine Department of Cancer Research and Molecular Medicine Exam MOL3007 Functional Genomics Thursday December 20 th 9.00-13.00 ECTS credits: 7.5 Number of pages (included front-page): 5 Supporting
Faculty of Medicine Department of Cancer Research and Molecular Medicine Exam MOL3007 Functional Genomics Thursday December 20 th 9.00-13.00 ECTS credits: 7.5 Number of pages (included front-page): 5 Supporting
Proteomics. Proteomics is the study of all proteins within organism. Challenges
 Proteomics Proteomics is the study of all proteins within organism. Challenges 1. The proteome is larger than the genome due to alternative splicing and protein modification. As we have said before we
Proteomics Proteomics is the study of all proteins within organism. Challenges 1. The proteome is larger than the genome due to alternative splicing and protein modification. As we have said before we
Proteomics: A Challenge for Technology and Information Science. What is proteomics?
 Proteomics: A Challenge for Technology and Information Science CBCB Seminar, November 21, 2005 Tim Griffin Dept. Biochemistry, Molecular Biology and Biophysics tgriffin@umn.edu What is proteomics? Proteomics
Proteomics: A Challenge for Technology and Information Science CBCB Seminar, November 21, 2005 Tim Griffin Dept. Biochemistry, Molecular Biology and Biophysics tgriffin@umn.edu What is proteomics? Proteomics
SGN-6106 Computational Systems Biology I
 SGN-6106 Computational Systems Biology I A View of Modern Measurement Systems in Cell Biology Kaisa-Leena Taattola 21.2.2007 1 The cell a complex system (Source: Lehninger Principles of Biochemistry 4th
SGN-6106 Computational Systems Biology I A View of Modern Measurement Systems in Cell Biology Kaisa-Leena Taattola 21.2.2007 1 The cell a complex system (Source: Lehninger Principles of Biochemistry 4th
Strategies in proteomics
 Strategies in proteomics Systems biology - understand cellpathways, network, and complex interacting (includes Genomics, Proteomics, Metabolomics..) Biological processes - characterize protein complexes,
Strategies in proteomics Systems biology - understand cellpathways, network, and complex interacting (includes Genomics, Proteomics, Metabolomics..) Biological processes - characterize protein complexes,
Strategies for Quantitative Proteomics. Atelier "Protéomique Quantitative" La Grande Motte, France - June 26, 2007
 Strategies for Quantitative Proteomics Atelier "Protéomique Quantitative", France - June 26, 2007 Bruno Domon, Ph.D. Institut of Molecular Systems Biology ETH Zurich Zürich, Switzerland OUTLINE Introduction
Strategies for Quantitative Proteomics Atelier "Protéomique Quantitative", France - June 26, 2007 Bruno Domon, Ph.D. Institut of Molecular Systems Biology ETH Zurich Zürich, Switzerland OUTLINE Introduction
Proteomics and Cancer
 Proteomics and Cancer Japan Society for the Promotion of Science (JSPS) Science Dialogue Program at Niitsu Senior High School Niitsu, Niigata September 4th 2006 Vladimir Valera, M.D, PhD JSPS Postdoctoral
Proteomics and Cancer Japan Society for the Promotion of Science (JSPS) Science Dialogue Program at Niitsu Senior High School Niitsu, Niigata September 4th 2006 Vladimir Valera, M.D, PhD JSPS Postdoctoral
Lecture 5: 8/31. CHAPTER 5 Techniques in Protein Biochemistry
 Lecture 5: 8/31 CHAPTER 5 Techniques in Protein Biochemistry Chapter 5 Outline The proteome is the entire set of proteins expressed and modified by a cell under a particular set of biochemical conditions.
Lecture 5: 8/31 CHAPTER 5 Techniques in Protein Biochemistry Chapter 5 Outline The proteome is the entire set of proteins expressed and modified by a cell under a particular set of biochemical conditions.
Quantitative mass spec based proteomics
 Quantitative mass spec based proteomics Tuula Nyman Institute of Biotechnology tuula.nyman@helsinki.fi THE PROTEOME The complete protein complement expressed by a genome or by a cell or a tissue type (M.
Quantitative mass spec based proteomics Tuula Nyman Institute of Biotechnology tuula.nyman@helsinki.fi THE PROTEOME The complete protein complement expressed by a genome or by a cell or a tissue type (M.
Really high sensitivity mass spectrometry and Discovery and analysis of protein complexes
 Really high sensitivity mass spectrometry and Discovery and analysis of protein complexes The PRIME lab and AMS Importance of protein complexes in biology Methods for isolation of protein complexes In
Really high sensitivity mass spectrometry and Discovery and analysis of protein complexes The PRIME lab and AMS Importance of protein complexes in biology Methods for isolation of protein complexes In
ENCODE DCC Antibody Validation Document
 ENCODE DCC Antibody Validation Document Date of Submission 09/12/12 Name: Trupti Kawli Email: trupti@stanford.edu Lab Snyder Antibody Name: SREBP1 (sc-8984) Target: SREBP1 Company/ Source: Santa Cruz Biotechnology
ENCODE DCC Antibody Validation Document Date of Submission 09/12/12 Name: Trupti Kawli Email: trupti@stanford.edu Lab Snyder Antibody Name: SREBP1 (sc-8984) Target: SREBP1 Company/ Source: Santa Cruz Biotechnology
Mass Spectrometry and Proteomics - Lecture 6 - Matthias Trost Newcastle University
 Mass Spectrometry and Proteomics - Lecture 6 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk Previously Quantitation techniques Label-free TMT SILAC Peptide identification and FDR 194 Lecture
Mass Spectrometry and Proteomics - Lecture 6 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk Previously Quantitation techniques Label-free TMT SILAC Peptide identification and FDR 194 Lecture
Biology: the basis for smart proteomic approaches to protein analysis
 QuickTime and a TIFF (Uncompressed) decompressor are needed to see this picture. Biology: the basis for smart proteomic approaches to protein analysis Helen Kim, PhD Department of Pharmacology & Toxicology
QuickTime and a TIFF (Uncompressed) decompressor are needed to see this picture. Biology: the basis for smart proteomic approaches to protein analysis Helen Kim, PhD Department of Pharmacology & Toxicology
Proteomics And Cancer Biomarker Discovery. Dr. Zahid Khan Institute of chemical Sciences (ICS) University of Peshawar. Overview. Cancer.
 Proteomics And Cancer Biomarker Discovery Dr. Zahid Khan Institute of chemical Sciences (ICS) University of Peshawar Overview Proteomics Cancer Aims Tools Data Base search Challenges Summary 1 Overview
Proteomics And Cancer Biomarker Discovery Dr. Zahid Khan Institute of chemical Sciences (ICS) University of Peshawar Overview Proteomics Cancer Aims Tools Data Base search Challenges Summary 1 Overview
Advances in analytical biochemistry and systems biology: Proteomics
 Advances in analytical biochemistry and systems biology: Proteomics Brett Boghigian Department of Chemical & Biological Engineering Tufts University July 29, 2005 Proteomics The basics History Current
Advances in analytical biochemistry and systems biology: Proteomics Brett Boghigian Department of Chemical & Biological Engineering Tufts University July 29, 2005 Proteomics The basics History Current
NPTEL VIDEO COURSE PROTEOMICS PROF. SANJEEVA SRIVASTAVA
 LECTURE-24 Quantitative proteomics: Stable isotope labeling by amino acids in cell culture (SILAC) TRANSCRIPT Welcome to the proteomics course. In today s lecture we will talk about Quantitative proteomics:
LECTURE-24 Quantitative proteomics: Stable isotope labeling by amino acids in cell culture (SILAC) TRANSCRIPT Welcome to the proteomics course. In today s lecture we will talk about Quantitative proteomics:
LECTURE-3. Protein Chemistry to proteomics HANDOUT. Proteins are the most dynamic and versatile macromolecules in a living cell, which
 LECTURE-3 Protein Chemistry to proteomics HANDOUT PREAMBLE Proteins are the most dynamic and versatile macromolecules in a living cell, which regulates essential activities of the cell. The classical protein
LECTURE-3 Protein Chemistry to proteomics HANDOUT PREAMBLE Proteins are the most dynamic and versatile macromolecules in a living cell, which regulates essential activities of the cell. The classical protein
What are proteomics? And what can they tell us about seed maturation and germination?
 Danseed Symposium Kobæk Strand, Skelskør 14 March, 2017 What are proteomics? And what can they tell us about seed maturation and germination? Ian Max Møller Department of Molecular Biology and Genetics
Danseed Symposium Kobæk Strand, Skelskør 14 March, 2017 What are proteomics? And what can they tell us about seed maturation and germination? Ian Max Møller Department of Molecular Biology and Genetics
Basic protein and peptide science for proteomics. Henrik Johansson
 Basic protein and peptide science for proteomics Henrik Johansson Proteins are the main actors in the cell Membranes Transport and storage Chemical factories DNA Building proteins Structure Proteins mediate
Basic protein and peptide science for proteomics Henrik Johansson Proteins are the main actors in the cell Membranes Transport and storage Chemical factories DNA Building proteins Structure Proteins mediate
Stable Isotope Labeling with Amino Acids in Cell Culture. Thermo Scientific Pierce SILAC Protein Quantitation Kits
 Stable Isotope Labeling with Amino Acids in Cell Culture Thermo Scientific Pierce SILAC Protein Quantitation Kits Quantitative Analysis of Differential Protein Expression Stable isotope labeling using
Stable Isotope Labeling with Amino Acids in Cell Culture Thermo Scientific Pierce SILAC Protein Quantitation Kits Quantitative Analysis of Differential Protein Expression Stable isotope labeling using
Kinetics Review. Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets)
 Quiz 1 Kinetics Review Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets) I will post the problems with solutions on Toolkit for those that can t make
Quiz 1 Kinetics Review Tonight at 7 PM Phys 204 We will do two problems on the board (additional ones than in the problem sets) I will post the problems with solutions on Toolkit for those that can t make
2D separations and analysis of proteins in biological samples
 January 18, 2005 2D separations and analysis of proteins in biological samples Helen Kim 934-3880 helenkim@uab.edu McCallum Building, room 460 Jan 18, 2005 HelenKim/UAB/PharmTox 1 Learning objectives 2-D
January 18, 2005 2D separations and analysis of proteins in biological samples Helen Kim 934-3880 helenkim@uab.edu McCallum Building, room 460 Jan 18, 2005 HelenKim/UAB/PharmTox 1 Learning objectives 2-D
Appendix. Table of contents
 Appendix Table of contents 1. Appendix figures 2. Legends of Appendix figures 3. References Appendix Figure S1 -Tub STIL (41) DVFFYQADDEHYIPR (55) (64) AVLLDLEPR (72) 1 451aa (1071) YLNENQLSQLSVTR (1084)
Appendix Table of contents 1. Appendix figures 2. Legends of Appendix figures 3. References Appendix Figure S1 -Tub STIL (41) DVFFYQADDEHYIPR (55) (64) AVLLDLEPR (72) 1 451aa (1071) YLNENQLSQLSVTR (1084)
Genomics, Transcriptomics and Proteomics
 Genomics, Transcriptomics and Proteomics GENES ARNm PROTEINS Genomics Transcriptomics Proteomics (10 6 protein forms?) Maturation PTMs Partners Localisation EDyP Lab : «Exploring the Dynamics of Proteomes»
Genomics, Transcriptomics and Proteomics GENES ARNm PROTEINS Genomics Transcriptomics Proteomics (10 6 protein forms?) Maturation PTMs Partners Localisation EDyP Lab : «Exploring the Dynamics of Proteomes»
Overview. Tools for Protein Sample Preparation, 2-D Electrophoresis, and Imaging and Analysis
 Expression Proteomics // Tools for Protein Separation and Analysis www.expressionproteomics.com 1 2 3 4 Overview Tools for Protein Sample Preparation, 2-D Electrophoresis, and Imaging and Analysis overview
Expression Proteomics // Tools for Protein Separation and Analysis www.expressionproteomics.com 1 2 3 4 Overview Tools for Protein Sample Preparation, 2-D Electrophoresis, and Imaging and Analysis overview
ProMass HR Applications!
 ProMass HR Applications! ProMass HR Features Ø ProMass HR includes features for high resolution data processing. Ø ProMass HR includes the standard ProMass deconvolution algorithm as well as the full Positive
ProMass HR Applications! ProMass HR Features Ø ProMass HR includes features for high resolution data processing. Ø ProMass HR includes the standard ProMass deconvolution algorithm as well as the full Positive
Proteomics software at MSI. Pratik Jagtap Minnesota Supercomputing institute
 Proteomics software at MSI. Pratik Jagtap Minnesota Supercomputing institute http://www.mass.msi.umn.edu/ Proteomics software at MSI. proteomics : emerging technology proteomics workflow search algorithms
Proteomics software at MSI. Pratik Jagtap Minnesota Supercomputing institute http://www.mass.msi.umn.edu/ Proteomics software at MSI. proteomics : emerging technology proteomics workflow search algorithms
Proteomics. Manickam Sugumaran. Department of Biology University of Massachusetts Boston, MA 02125
 Proteomics Manickam Sugumaran Department of Biology University of Massachusetts Boston, MA 02125 Genomic studies produced more than 75,000 potential gene sequence targets. (The number may be even higher
Proteomics Manickam Sugumaran Department of Biology University of Massachusetts Boston, MA 02125 Genomic studies produced more than 75,000 potential gene sequence targets. (The number may be even higher
基于质谱的蛋白质药物定性定量分析技术及应用
 基于质谱的蛋白质药物定性定量分析技术及应用 蛋白质组学质谱数据分析 单抗全蛋白测序及鉴定 Bioinformatics Solutions Inc. Waterloo, ON, Canada 上海 中国 2017 1 3 PEAKS Studio GUI Introduction Overview of PEAKS Studio GUI Project View Tasks, running info,
基于质谱的蛋白质药物定性定量分析技术及应用 蛋白质组学质谱数据分析 单抗全蛋白测序及鉴定 Bioinformatics Solutions Inc. Waterloo, ON, Canada 上海 中国 2017 1 3 PEAKS Studio GUI Introduction Overview of PEAKS Studio GUI Project View Tasks, running info,
Extracting Pure Proteins from Cells
 Extracting Pure Proteins from Cells 0 Purification techniques focus mainly on size & charge 0 The first step is homogenization (grinding, Potter Elvejhem homogenizer, sonication, freezing and thawing,
Extracting Pure Proteins from Cells 0 Purification techniques focus mainly on size & charge 0 The first step is homogenization (grinding, Potter Elvejhem homogenizer, sonication, freezing and thawing,
ichemistry SOLUTIONS Integrated chemistries to boost mass spectrometry workflows
 ichemistry SOLUTIONS Integrated chemistries to boost mass spectrometry workflows ichemistry SOLUTIONS ichemistry solutions ichemistry solutions are the world s only reagents and consumables that are customdesigned
ichemistry SOLUTIONS Integrated chemistries to boost mass spectrometry workflows ichemistry SOLUTIONS ichemistry solutions ichemistry solutions are the world s only reagents and consumables that are customdesigned
Finding Genes with Genomics Technologies
 PLNT2530 Plant Biotechnology (2018) Unit 7 Finding Genes with Genomics Technologies Unless otherwise cited or referenced, all content of this presenataion is licensed under the Creative Commons License
PLNT2530 Plant Biotechnology (2018) Unit 7 Finding Genes with Genomics Technologies Unless otherwise cited or referenced, all content of this presenataion is licensed under the Creative Commons License
Využití cílené proteomiky pro kontrolu falšování potravin: identifikace peptidových markerů v mase pomocí LC- Q Exactive MS/MS
 Využití cílené proteomiky pro kontrolu falšování potravin: identifikace peptidových markerů v mase pomocí LC- Q Exactive MS/MS Michal Godula Ph.D. Thermo Fisher Scientific The world leader in serving science
Využití cílené proteomiky pro kontrolu falšování potravin: identifikace peptidových markerů v mase pomocí LC- Q Exactive MS/MS Michal Godula Ph.D. Thermo Fisher Scientific The world leader in serving science
Lecture 8: Affinity Chromatography-III
 Lecture 8: Affinity Chromatography-III Key words: Chromatography; Affinity chromatography; Protein Purification During this lecture, we shall be studying few more examples of affinity chromatography. The
Lecture 8: Affinity Chromatography-III Key words: Chromatography; Affinity chromatography; Protein Purification During this lecture, we shall be studying few more examples of affinity chromatography. The
Proteomics Background and clinical utility
 Proteomics Background and clinical utility H.H. Helgason MD Antoni van Leeuwenhoek Hospital The Netherlands Cancer Institute Amsterdam Introduction Background Definitions Protein biomarkers Technical aspects
Proteomics Background and clinical utility H.H. Helgason MD Antoni van Leeuwenhoek Hospital The Netherlands Cancer Institute Amsterdam Introduction Background Definitions Protein biomarkers Technical aspects
Protein analysis. Dr. Mamoun Ahram Summer semester, Resources This lecture Campbell and Farrell s Biochemistry, Chapters 5
 Protein analysis Dr. Mamoun Ahram Summer semester, 2015-2016 Resources This lecture Campbell and Farrell s Biochemistry, Chapters 5 Bases of protein separation Proteins can be purified on the basis Solubility
Protein analysis Dr. Mamoun Ahram Summer semester, 2015-2016 Resources This lecture Campbell and Farrell s Biochemistry, Chapters 5 Bases of protein separation Proteins can be purified on the basis Solubility
ENCODE DCC Antibody Validation Document
 ENCODE DCC Antibody Validation Document Date of Submission 06/15/2012 Name: Trupti Kawli Email: trupti@stanford.edu Lab Snyder Antibody Name: CDP (sc6327) Target: CDP Company/ Source: Santa Cruz Catalog
ENCODE DCC Antibody Validation Document Date of Submission 06/15/2012 Name: Trupti Kawli Email: trupti@stanford.edu Lab Snyder Antibody Name: CDP (sc6327) Target: CDP Company/ Source: Santa Cruz Catalog
Laboratory Water Quality Affects Protein Separation by 2D Gel Electrophoresis
 Laboratory Water Quality Affects Protein Separation by 2D Gel Electrophoresis 2D gel electrophoresis remains a dominant technique in proteomics. Obtaining high quality gels requires careful and tedious
Laboratory Water Quality Affects Protein Separation by 2D Gel Electrophoresis 2D gel electrophoresis remains a dominant technique in proteomics. Obtaining high quality gels requires careful and tedious
Post-translational modification
 Protein expression Western blotting, is a widely used and accepted technique to detect levels of protein expression in a cell or tissue extract. This technique measures protein levels in a biological sample
Protein expression Western blotting, is a widely used and accepted technique to detect levels of protein expression in a cell or tissue extract. This technique measures protein levels in a biological sample
Mass Spectrometry-based Methods of Proteome Analysis
 1 Mass Spectrometry-based Methods of Proteome Analysis Boris L. Zybailov and Michael P. Washburn Stowers Institute for Medical Research, Kansas City, MO, USA 1 Principles and Instrumentation 3 1.1 Proteome
1 Mass Spectrometry-based Methods of Proteome Analysis Boris L. Zybailov and Michael P. Washburn Stowers Institute for Medical Research, Kansas City, MO, USA 1 Principles and Instrumentation 3 1.1 Proteome
FACTORS THAT AFFECT PROTEIN IDENTIFICATION BY MASS SPECTROMETRY HAOFEI TIFFANY WANG. (Under the Direction of Ron Orlando) ABSTRACT
 FACTORS THAT AFFECT PROTEIN IDENTIFICATION BY MASS SPECTROMETRY by HAOFEI TIFFANY WANG (Under the Direction of Ron Orlando) ABSTRACT Mass spectrometry combined with database search utilities is a valuable
FACTORS THAT AFFECT PROTEIN IDENTIFICATION BY MASS SPECTROMETRY by HAOFEI TIFFANY WANG (Under the Direction of Ron Orlando) ABSTRACT Mass spectrometry combined with database search utilities is a valuable
Exam MOL3007 Functional Genomics
 Faculty of Medicine Department of Cancer Research and Molecular Medicine Exam MOL3007 Functional Genomics Tuesday May 29 th 9.00-13.00 ECTS credits: 7.5 Number of pages (included front-page): 5 Supporting
Faculty of Medicine Department of Cancer Research and Molecular Medicine Exam MOL3007 Functional Genomics Tuesday May 29 th 9.00-13.00 ECTS credits: 7.5 Number of pages (included front-page): 5 Supporting
Bioinformatics (Lec 19) Picture Copyright: the National Museum of Health
 3/29/05 1 Picture Copyright: AccessExcellence @ the National Museum of Health PCR 3/29/05 2 Schematic outline of a typical PCR cycle Target DNA Primers dntps DNA polymerase 3/29/05 3 Gel Electrophoresis
3/29/05 1 Picture Copyright: AccessExcellence @ the National Museum of Health PCR 3/29/05 2 Schematic outline of a typical PCR cycle Target DNA Primers dntps DNA polymerase 3/29/05 3 Gel Electrophoresis
Selected Techniques Part I
 1 Selected Techniques Part I Gel Electrophoresis Can be both qualitative and quantitative Qualitative About what size is the fragment? How many fragments are present? Is there in insert or not? Quantitative
1 Selected Techniques Part I Gel Electrophoresis Can be both qualitative and quantitative Qualitative About what size is the fragment? How many fragments are present? Is there in insert or not? Quantitative
CCRD Proteomics facility RIH-Brown University
 CCRD Proteomics facility RIH-Brown University Experimental design TO Publication Nagib Ahsan, PhD Where is the CCRD Proteomics Facility COBRE CCRD Proteomics Core Facility 1 Hoppin St, Providence, 02903
CCRD Proteomics facility RIH-Brown University Experimental design TO Publication Nagib Ahsan, PhD Where is the CCRD Proteomics Facility COBRE CCRD Proteomics Core Facility 1 Hoppin St, Providence, 02903
Instructions SILAC Protein Quantitation Kits
 Instructions SILAC Protein Quantitation Kits CIL Catalog No. Description DMEM LYS C SILAC Protein Quantitation Kit DMEM (Dulbecco s Modified Eagle s Medium) Kit contains: SILAC DMEM Media, 2 x 500 ml Dialyzed
Instructions SILAC Protein Quantitation Kits CIL Catalog No. Description DMEM LYS C SILAC Protein Quantitation Kit DMEM (Dulbecco s Modified Eagle s Medium) Kit contains: SILAC DMEM Media, 2 x 500 ml Dialyzed
Introduction to Proteomics
 Introduction to Proteomics Tasso Miliotis, PhD AstraZeneca R&D Gothenburg Translational Sciences tasso.miliotis@astrazeneca.com 1 Site Management Mölndal May 2008 Outline Drug Discovery AZ R&D Sample Preparation
Introduction to Proteomics Tasso Miliotis, PhD AstraZeneca R&D Gothenburg Translational Sciences tasso.miliotis@astrazeneca.com 1 Site Management Mölndal May 2008 Outline Drug Discovery AZ R&D Sample Preparation
Proteomics 6/4/2009 WESTERN BLOT ANALYSIS
 SDS-PAGE (PolyAcrylamide Gel Electrophoresis) Proteomics WESTERN BLOT ANALYSIS Presented by: Nuvee Prapasarakul Veterinary Microbiology Chulalongkorn University Proteomics has been said to be the next
SDS-PAGE (PolyAcrylamide Gel Electrophoresis) Proteomics WESTERN BLOT ANALYSIS Presented by: Nuvee Prapasarakul Veterinary Microbiology Chulalongkorn University Proteomics has been said to be the next
ProteinPilot Report for ProteinPilot Software
 ProteinPilot Report for ProteinPilot Software Detailed Analysis of Protein Identification / Quantitation Results Automatically Sean L Seymour, Christie Hunter SCIEX, USA Powerful mass spectrometers like
ProteinPilot Report for ProteinPilot Software Detailed Analysis of Protein Identification / Quantitation Results Automatically Sean L Seymour, Christie Hunter SCIEX, USA Powerful mass spectrometers like
Improving Productivity with Applied Biosystems GPS Explorer
 Product Bulletin TOF MS Improving Productivity with Applied Biosystems GPS Explorer Software Purpose GPS Explorer Software is the application layer software for the Applied Biosystems 4700 Proteomics Discovery
Product Bulletin TOF MS Improving Productivity with Applied Biosystems GPS Explorer Software Purpose GPS Explorer Software is the application layer software for the Applied Biosystems 4700 Proteomics Discovery
Innovations for Protein Research. Protein Research. Powerful workflows built on solid science
 Innovations for Protein Research Protein Research Powerful workflows built on solid science Your partner in discovering innovative solutions for quantitative protein and biomarker research Applied Biosystems/MDS
Innovations for Protein Research Protein Research Powerful workflows built on solid science Your partner in discovering innovative solutions for quantitative protein and biomarker research Applied Biosystems/MDS
PROTEIN AND PROTEOME ANALYTICS IDENTIFICATION, CHARACTERIZATION AND EXPRESSION ANALYSIS OF PROTEINS
 FRAUNHOFER INSTITUTE FOR INTERFACIAL ENGINEERING AND BIOTECHNOLOGY PROTEIN AND PROTEOME ANALYTICS IDENTIFICATION, CHARACTERIZATION AND EXPRESSION ANALYSIS OF PROTEINS 1 1 2 2 PROTEINS PROTEOMICS Proteins,
FRAUNHOFER INSTITUTE FOR INTERFACIAL ENGINEERING AND BIOTECHNOLOGY PROTEIN AND PROTEOME ANALYTICS IDENTIFICATION, CHARACTERIZATION AND EXPRESSION ANALYSIS OF PROTEINS 1 1 2 2 PROTEINS PROTEOMICS Proteins,
27041, Week 02. Review of Week 01
 27041, Week 02 Review of Week 01 The human genome sequencing project (HGP) 2 CBS, Department of Systems Biology Systems Biology and emergent properties 3 CBS, Department of Systems Biology Different model
27041, Week 02 Review of Week 01 The human genome sequencing project (HGP) 2 CBS, Department of Systems Biology Systems Biology and emergent properties 3 CBS, Department of Systems Biology Different model
Motivation From Protein to Gene
 MOLECULAR BIOLOGY 2003-4 Topic B Recombinant DNA -principles and tools Construct a library - what for, how Major techniques +principles Bioinformatics - in brief Chapter 7 (MCB) 1 Motivation From Protein
MOLECULAR BIOLOGY 2003-4 Topic B Recombinant DNA -principles and tools Construct a library - what for, how Major techniques +principles Bioinformatics - in brief Chapter 7 (MCB) 1 Motivation From Protein
Spectrum Mill MS Proteomics Workbench. Comprehensive tools for MS proteomics
 Spectrum Mill MS Proteomics Workbench Comprehensive tools for MS proteomics Meeting the challenge of proteomics data analysis Mass spectrometry is a core technology for proteomics research, but large-scale
Spectrum Mill MS Proteomics Workbench Comprehensive tools for MS proteomics Meeting the challenge of proteomics data analysis Mass spectrometry is a core technology for proteomics research, but large-scale
Key questions of proteomics. Bioinformatics 2. Proteomics. Foundation of proteomics. What proteins are there? Protein digestion
 s s Key questions of proteomics What proteins are there? Bioinformatics 2 Lecture X roteomics How much is there of each of the proteins? - Absolute quantitation - Stoichiometry What (modification/splice)
s s Key questions of proteomics What proteins are there? Bioinformatics 2 Lecture X roteomics How much is there of each of the proteins? - Absolute quantitation - Stoichiometry What (modification/splice)
New Approaches to Quantitative Proteomics Analysis
 New Approaches to Quantitative Proteomics Analysis Chris Hodgkins, Market Development Manager, SCIEX ANZ 2 nd November, 2017 Who is SCIEX? Founded by Dr. Barry French & others: University of Toronto Introduced
New Approaches to Quantitative Proteomics Analysis Chris Hodgkins, Market Development Manager, SCIEX ANZ 2 nd November, 2017 Who is SCIEX? Founded by Dr. Barry French & others: University of Toronto Introduced
Medicinal Chemistry of Modern Antibiotics
 Chemistry 259 Medicinal Chemistry of Modern Antibiotics Spring 2012 Lecture 5: Modern Target Discovery & MOA Thomas Hermann Department of Chemistry & Biochemistry University of California, San Diego Drug
Chemistry 259 Medicinal Chemistry of Modern Antibiotics Spring 2012 Lecture 5: Modern Target Discovery & MOA Thomas Hermann Department of Chemistry & Biochemistry University of California, San Diego Drug
Medicinal Chemistry of Modern Antibiotics
 Chemistry 259 Medicinal Chemistry of Modern Antibiotics Spring 2008 Lecture 5: Modern Target Discovery & MOA Thomas Hermann Department of Chemistry & Biochemistry University of California, San Diego Drug
Chemistry 259 Medicinal Chemistry of Modern Antibiotics Spring 2008 Lecture 5: Modern Target Discovery & MOA Thomas Hermann Department of Chemistry & Biochemistry University of California, San Diego Drug
Mass Spectrometry. Kyle Chau and Andrew Gioe
 Mass Spectrometry Kyle Chau and Andrew Gioe Computation of Molecular Mass - Mass Spectrum is a plot of intensity as a function of masscharge ratio, m/z. - Mass-charge ratio can be determined through accelerating
Mass Spectrometry Kyle Chau and Andrew Gioe Computation of Molecular Mass - Mass Spectrum is a plot of intensity as a function of masscharge ratio, m/z. - Mass-charge ratio can be determined through accelerating
CBI Toolbox Tour 2015
 CBI Toolbox Tour 2015 Thermophoresis (NanoTemper) NT.115 & NT.LabelFree Images: NanoTemper Circular Dichroism Jasco J-1500 Spectrometer Six Position Turreted Peltier Temperature Control System Automated
CBI Toolbox Tour 2015 Thermophoresis (NanoTemper) NT.115 & NT.LabelFree Images: NanoTemper Circular Dichroism Jasco J-1500 Spectrometer Six Position Turreted Peltier Temperature Control System Automated
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
What is a proteome and how is the link between an organisms genome and a proteome.
 Answer to exam questions Bi2014 autumn 2013. Question 1a. What is a proteome and how is the link between an organisms genome and a proteome. The answer should discuss the following topics: What is a genome,
Answer to exam questions Bi2014 autumn 2013. Question 1a. What is a proteome and how is the link between an organisms genome and a proteome. The answer should discuss the following topics: What is a genome,
Supplementary Figure 1. Processing and quality control for recombinant proteins.
 Supplementary Figure 1 Processing and quality control for recombinant proteins. (a) Schematic representation of processing of a recombinant protein library. Recombinant proteins were generated individually.
Supplementary Figure 1 Processing and quality control for recombinant proteins. (a) Schematic representation of processing of a recombinant protein library. Recombinant proteins were generated individually.
11/22/13. Proteomics, functional genomics, and systems biology. Biosciences 741: Genomics Fall, 2013 Week 11
 Proteomics, functional genomics, and systems biology Biosciences 741: Genomics Fall, 2013 Week 11 1 Figure 6.1 The future of genomics Functional Genomics The field of functional genomics represents the
Proteomics, functional genomics, and systems biology Biosciences 741: Genomics Fall, 2013 Week 11 1 Figure 6.1 The future of genomics Functional Genomics The field of functional genomics represents the
Non-covalent or Native Mass Spectrometry. We can ionize intact protein complexes using ESI!!
 Non-covalent or Native Mass Spectrometry We can ionize intact protein complexes using ESI!! What Can We Learn? Stoichiometry of complex Relative affinity and topology Rates of subunit exchange Assembly
Non-covalent or Native Mass Spectrometry We can ionize intact protein complexes using ESI!! What Can We Learn? Stoichiometry of complex Relative affinity and topology Rates of subunit exchange Assembly
Supplementary Figure 1. Immunoprecipitation of synthetic SUMOm-remnant peptides using UMO monoclonal antibody. (a) LC-MS analyses of tryptic
 Supplementary Figure 1. Immunoprecipitation of synthetic SUMOm-remnant peptides using UMO 1-7-7 monoclonal antibody. (a) LC-MS analyses of tryptic digest from HEK293 cells spiked with 6 SUMOmremnant peptides
Supplementary Figure 1. Immunoprecipitation of synthetic SUMOm-remnant peptides using UMO 1-7-7 monoclonal antibody. (a) LC-MS analyses of tryptic digest from HEK293 cells spiked with 6 SUMOmremnant peptides
Chapter 5: Proteins: Primary Structure
 Instant download and all chapters Test Bank Fundamentals of Biochemistry Life at the Molecular Level 4th Edition Donald Voet https://testbanklab.com/download/test-bank-fundamentals-biochemistry-life-molecular-level-
Instant download and all chapters Test Bank Fundamentals of Biochemistry Life at the Molecular Level 4th Edition Donald Voet https://testbanklab.com/download/test-bank-fundamentals-biochemistry-life-molecular-level-
BST 226 Statistical Methods for Bioinformatics David M. Rocke. February 10, 2014 BST 226 Statistical Methods for Bioinformatics 1
 BST 226 Statistical Methods for Bioinformatics David M. Rocke February 10, 2014 BST 226 Statistical Methods for Bioinformatics 1 Proteomics by LS/MS/MS Proteins are digested with trypsin or other proteases
BST 226 Statistical Methods for Bioinformatics David M. Rocke February 10, 2014 BST 226 Statistical Methods for Bioinformatics 1 Proteomics by LS/MS/MS Proteins are digested with trypsin or other proteases
rapiflex Innovation with Integrity Designed for Molecules that Matter. MALDI TOF/TOF
 rapiflex Designed for Molecules that Matter. Innovation with Integrity MALDI TOF/TOF rapiflex TM The first MALDI-TOF/TOF that adapts to your needs. The rapiflex is the most advanced MALDI TOF/TOF system
rapiflex Designed for Molecules that Matter. Innovation with Integrity MALDI TOF/TOF rapiflex TM The first MALDI-TOF/TOF that adapts to your needs. The rapiflex is the most advanced MALDI TOF/TOF system
Topic 2: Proteins. 2-1 specific proteins can be purified from cell extracts. Molecular Biology and Public Health ( 分子生物学与公共卫生 )
 HAPTER20: Techniques of Molecular Biology Molecular Biology and Public Health ( 分子生物学与公共卫生 ) -------Protein manipulation ( 蛋白操作 ) Topic 2: Proteins 1. Protein purification ( 蛋白质纯化 ) 2. Affinity chromatography
HAPTER20: Techniques of Molecular Biology Molecular Biology and Public Health ( 分子生物学与公共卫生 ) -------Protein manipulation ( 蛋白操作 ) Topic 2: Proteins 1. Protein purification ( 蛋白质纯化 ) 2. Affinity chromatography
Systems Biology and Systems Medicine
 CHAPTER 5 Systems Biology and Systems Medicine INTRODUCTION A new approach to biology, termed systems biology, has emerged over the past 15 years or so an approach that looks - - have transformed systems
CHAPTER 5 Systems Biology and Systems Medicine INTRODUCTION A new approach to biology, termed systems biology, has emerged over the past 15 years or so an approach that looks - - have transformed systems
A Highly Accurate Mass Profiling Approach to Protein Biomarker Discovery Using HPLC-Chip/ MS-Enabled ESI-TOF MS
 Application Note PROTEOMICS METABOLOMICS GENOMICS INFORMATICS GLYILEVALCYSGLUGLNALASERLEUASPARG CYSVALLYSPROLYSPHETYRTHRLEUHISLYS A Highly Accurate Mass Profiling Approach to Protein Biomarker Discovery
Application Note PROTEOMICS METABOLOMICS GENOMICS INFORMATICS GLYILEVALCYSGLUGLNALASERLEUASPARG CYSVALLYSPROLYSPHETYRTHRLEUHISLYS A Highly Accurate Mass Profiling Approach to Protein Biomarker Discovery
Peptide enrichment and fractionation
 Protein sample preparation Successful analysis of low-abundance proteins and/or the identification of posttranslationally modified peptides often require several steps: enrichment, fractionation, and/or
Protein sample preparation Successful analysis of low-abundance proteins and/or the identification of posttranslationally modified peptides often require several steps: enrichment, fractionation, and/or
NPTEL
 NPTEL Syllabus Proteomics: Principles and Techniques - Video course COURSE OUTLINE An introduction to proteomics: Basics of protein structure and function, An overview of systems biology, Evolution from
NPTEL Syllabus Proteomics: Principles and Techniques - Video course COURSE OUTLINE An introduction to proteomics: Basics of protein structure and function, An overview of systems biology, Evolution from
The Open2Dprot Proteomics Project for n-dimensional Protein Expression Data Analysis. The Open2Dprot Project. Introduction
 The Open2Dprot Proteomics Project for n-dimensional Protein Expression Data Analysis http://open2dprot.sourceforge.net/ Revised 2-05-2006 * (cf. 2D-LC) Introduction There is a need for integrated proteomics
The Open2Dprot Proteomics Project for n-dimensional Protein Expression Data Analysis http://open2dprot.sourceforge.net/ Revised 2-05-2006 * (cf. 2D-LC) Introduction There is a need for integrated proteomics
A New Strategy for Quantitative Proteomics Using Isotope-Coded Protein Labels
 ICPL TM -Isotope Coded Label A New Strategy for Quantitative Proteomics Using Isotope-Coded Labels Alexander Schmidt Department for Max-Planck-Institute of Biochemistry Spotfire User Conference 2004, Cologne
ICPL TM -Isotope Coded Label A New Strategy for Quantitative Proteomics Using Isotope-Coded Labels Alexander Schmidt Department for Max-Planck-Institute of Biochemistry Spotfire User Conference 2004, Cologne
Isotopic Resolution of Chromatographically Separated IdeS Subunits Using the X500B QTOF System
 Isotopic Resolution of Chromatographically Separated IdeS Subunits Using the X500B QTOF System Fan Zhang 2, Sean McCarthy 1 1 SCIEX, MA, USA, 2 SCIEX, CA USA Analysis of protein subunits using high resolution
Isotopic Resolution of Chromatographically Separated IdeS Subunits Using the X500B QTOF System Fan Zhang 2, Sean McCarthy 1 1 SCIEX, MA, USA, 2 SCIEX, CA USA Analysis of protein subunits using high resolution
Faster, easier, flexible proteomics solutions
 Agilent HPLC-Chip LC/MS Faster, easier, flexible proteomics solutions Our measure is your success. products applications soft ware services Phospho- Anaysis Intact Glycan & Glycoprotein Agilent s HPLC-Chip
Agilent HPLC-Chip LC/MS Faster, easier, flexible proteomics solutions Our measure is your success. products applications soft ware services Phospho- Anaysis Intact Glycan & Glycoprotein Agilent s HPLC-Chip
Identification of human serum proteins detectable after Albumin removal with Vivapure Anti-HSA Kit
 Andreas Kocourek, Pieter Eyckerman, Robert Zeidler, Pascal Bolon and Birgit Thome-Kromer Introduction The analysis of the complete proteome is a major interest of many researchers. Of particular importance
Andreas Kocourek, Pieter Eyckerman, Robert Zeidler, Pascal Bolon and Birgit Thome-Kromer Introduction The analysis of the complete proteome is a major interest of many researchers. Of particular importance
Comparability Analysis of Protein Therapeutics by Bottom-Up LC-MS with Stable Isotope-Tagged Reference Standards
 Comparability Analysis of Protein Therapeutics by Bottom-Up LC-MS with Stable Isotope-Tagged Reference Standards 16 September 2011 Abbott Bioresearch Center, Worcester, MA USA Manuilov, A. V., C. H. Radziejewski
Comparability Analysis of Protein Therapeutics by Bottom-Up LC-MS with Stable Isotope-Tagged Reference Standards 16 September 2011 Abbott Bioresearch Center, Worcester, MA USA Manuilov, A. V., C. H. Radziejewski
Purification: Step 1. Lecture 11 Protein and Peptide Chemistry. Cells: Break them open! Crude Extract
 Purification: Step 1 Lecture 11 Protein and Peptide Chemistry Cells: Break them open! Crude Extract Total contents of cell Margaret A. Daugherty Fall 2003 Big Problem: Crude extract is not the natural
Purification: Step 1 Lecture 11 Protein and Peptide Chemistry Cells: Break them open! Crude Extract Total contents of cell Margaret A. Daugherty Fall 2003 Big Problem: Crude extract is not the natural
Purification: Step 1. Protein and Peptide Chemistry. Lecture 11. Big Problem: Crude extract is not the natural environment. Cells: Break them open!
 Lecture 11 Protein and Peptide Chemistry Margaret A. Daugherty Fall 2003 Purification: Step 1 Cells: Break them open! Crude Extract Total contents of cell Big Problem: Crude extract is not the natural
Lecture 11 Protein and Peptide Chemistry Margaret A. Daugherty Fall 2003 Purification: Step 1 Cells: Break them open! Crude Extract Total contents of cell Big Problem: Crude extract is not the natural
Expression Proteomics: principles and examples of application
 Expression Proteomics: principles and examples of application Miguel Teixeira IBB Institute for Biotechnology and Bioengineering, Centro de Eng. Biológica e Química, IST Functional Genomics and Bioinformatics
Expression Proteomics: principles and examples of application Miguel Teixeira IBB Institute for Biotechnology and Bioengineering, Centro de Eng. Biológica e Química, IST Functional Genomics and Bioinformatics
From Proteomics to Systems Biology. Integration of omics - information
 From Proteomics to Systems Biology Integration of omics - information Outline and learning objectives Omics science provides global analysis tools to study entire systems How to obtain omics - data What
From Proteomics to Systems Biology Integration of omics - information Outline and learning objectives Omics science provides global analysis tools to study entire systems How to obtain omics - data What
Capabilities & Services
 Capabilities & Services Accelerating Research & Development Table of Contents Introduction to DHMRI 3 Services and Capabilites: Genomics 4 Proteomics & Protein Characterization 5 Metabolomics 6 In Vitro
Capabilities & Services Accelerating Research & Development Table of Contents Introduction to DHMRI 3 Services and Capabilites: Genomics 4 Proteomics & Protein Characterization 5 Metabolomics 6 In Vitro
Protein purification prior to proteomics analysis
 Protein purification prior to proteomics analysis Stephen Barnes, PhD Department of Pharmacology & Toxicology and Mass Spectrometry Shared Facility, UAB The dynamic range of protein abundances Proteins
Protein purification prior to proteomics analysis Stephen Barnes, PhD Department of Pharmacology & Toxicology and Mass Spectrometry Shared Facility, UAB The dynamic range of protein abundances Proteins
Proteomics. Areas of Application for Proteomics. Most Commonly Used Proteomics Techniques: Limitations: Examples
 Proteomics Areas of Application for Proteomics Most Commonly Used Proteomics Techniques: Antibody arrays Protein activity arrays 2-D gels ICAT technology SELDI Limitations: protein sources surfaces and
Proteomics Areas of Application for Proteomics Most Commonly Used Proteomics Techniques: Antibody arrays Protein activity arrays 2-D gels ICAT technology SELDI Limitations: protein sources surfaces and
Spectral Counting Approaches and PEAKS
 Spectral Counting Approaches and PEAKS INBRE Proteomics Workshop, April 5, 2017 Boris Zybailov Department of Biochemistry and Molecular Biology University of Arkansas for Medical Sciences 1. Introduction
Spectral Counting Approaches and PEAKS INBRE Proteomics Workshop, April 5, 2017 Boris Zybailov Department of Biochemistry and Molecular Biology University of Arkansas for Medical Sciences 1. Introduction
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Outline and learning objectives. From Proteomics to Systems Biology. Integration of omics - information
 From to Systems Biology Outline and learning objectives Omics science provides global analysis tools to study entire systems How to obtain omics - What can we learn Limitations Integration of omics - In-class
From to Systems Biology Outline and learning objectives Omics science provides global analysis tools to study entire systems How to obtain omics - What can we learn Limitations Integration of omics - In-class
PROTEOMICS. Timothy Palzkill Baylor College of Medicine. KLUWER ACADEMIC PUBLISHERS New York / Boston / Dordrecht / London / Moscow
 PROTEOMICS PROTEOMICS by Timothy Palzkill Baylor College of Medicine KLUWER ACADEMIC PUBLISHERS New York / Boston / Dordrecht / London / Moscow ebook ISBN: 0-306-46895-6 Print ISBN: 0-792-37565-3 2002
PROTEOMICS PROTEOMICS by Timothy Palzkill Baylor College of Medicine KLUWER ACADEMIC PUBLISHERS New York / Boston / Dordrecht / London / Moscow ebook ISBN: 0-306-46895-6 Print ISBN: 0-792-37565-3 2002
Blotting Techniques (Southern blot, Northern blot, Western blot, and Eastern blot)
 Blotting Techniques (Southern blot, Northern blot, Western blot, and Eastern blot) Masheal Aljumaah SEP 2018 Learning Objectives: What is blotting? Blotting Techniques Types. Applications for each technique.
Blotting Techniques (Southern blot, Northern blot, Western blot, and Eastern blot) Masheal Aljumaah SEP 2018 Learning Objectives: What is blotting? Blotting Techniques Types. Applications for each technique.
Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells.
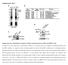 Supplementary Fig. 1 Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells. (a) FRTL-5 cells were treated with 1 mm dibutyryl camp for 24 h, and the lysates
Supplementary Fig. 1 Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells. (a) FRTL-5 cells were treated with 1 mm dibutyryl camp for 24 h, and the lysates
Supplementary Figure S1 Supplementary Figure S2 Supplementary Figure S3. Supplementary Figure S4
 Supplementary Figure S1 Supplementary Figure S2 Supplementary Figure S3 Supplementary Figure S4 Supplementary Figure S5 Supplementary Figure S6 Supplementary Figure S7 Supplementary Figure S8 Supplementary
Supplementary Figure S1 Supplementary Figure S2 Supplementary Figure S3 Supplementary Figure S4 Supplementary Figure S5 Supplementary Figure S6 Supplementary Figure S7 Supplementary Figure S8 Supplementary
Quantification of Isotope Encoded Proteins in 2D Gels
 Quantification of Isotope Encoded Proteins in 2D Gels Using Surface Enhanced Resonance Raman Giselle M. Knudsen 1, Brandon M. Davis 2, Shirshendu K. Deb 1, Yvette Loethen 2, Ravindra Gudihal 1, Pradeep
Quantification of Isotope Encoded Proteins in 2D Gels Using Surface Enhanced Resonance Raman Giselle M. Knudsen 1, Brandon M. Davis 2, Shirshendu K. Deb 1, Yvette Loethen 2, Ravindra Gudihal 1, Pradeep
Chapter 9 Proteomics. From genomics to proteomics
 Chapter 9 Proteomics From genomics to proteomics From structural genomics, functional genomics to proteomics http://www.nature.com/scitable/topicpage/the-proteome-discovering-the-structure-and-function-613
Chapter 9 Proteomics From genomics to proteomics From structural genomics, functional genomics to proteomics http://www.nature.com/scitable/topicpage/the-proteome-discovering-the-structure-and-function-613
