HHS Public Access Author manuscript Nat Methods. Author manuscript; available in PMC 2009 November 01.
|
|
|
- Noah Melton
- 6 years ago
- Views:
Transcription
1 Photoconversion in orange and red fluorescent proteins Gert-Jan Kremers 1, Kristin L. Hazelwood 2, Christopher S. Murphy 2, Michael W. Davidson 2, and David W. Piston 1 1 Department of Molecular Physiology and Biophysics, Vanderbilt University, 702 Light Hall Nashville, TN , USA 2 National High Magnetic Field Laboratory and Department of Biological Science, The Florida State University, Tallahassee, FL 32310, USA Abstract HHS Public Access Author manuscript Published in final edited form as: Nat Methods May ; 6(5): doi: /nmeth We report that photoconversion is fairly common among orange and red fluorescent proteins, as a screen of 12 variants yielded 8 that exhibit photoconversion. Specifically, three red fluorescent proteins can be switched into a green state, and two orange variants can be photoconverted to the far red. The orange highlighters are ideal for dual-probe highlighter applications, and they exhibit the most red-shifted excitation of all fluorescent protein described to date. Photoconvertable fluorescent proteins (pc-fps) exhibit a change in fluorescence excitation and emission spectra after excitation at a specific wavelength, and are thus useful as optical highlighters. Since the first description of a photoactivatable probe, PA-GFP1, which switches from a dark to a fluorescent state, many fluorescent proteins have been developed that permit photoconversion rather than just photoactivation. These fluorescent proteins switch between two colors (typically from green to red), both of which can be visualized. The palette of currently useful pc-fps includes PS-CFP2, Dendra3, 4, EosFP5, Kaede6, and KikGR7, all of which exhibit red-shifted fluorescence emission accompanied by a concomitant shift in the excitation maxima after irradiation with near-uv or deep blue light. The dimeric red fluorescent protein (RFP) Katushka and its monomeric variant mkate yield the greatest levels of fluorescence emission above 650 nm of all presently characterized RFPs8. Cells expressing Katushka or mkate were fluorescent upon two-photon excitation at 750 nm, although the fluorescence decreased rapidly, apparently due to photobleaching. However, subsequent imaging of the cells through a GFP emission filter showed a novel green fluorescence (Fig. 1a,b), indicating a photoconversion of the red fluorescent proteins to a green state. Further investigation revealed that red-to-green photoconversion also occured upon single-photon laser excitation at 405 and 561 nm (Table 1), and that the green Users may view, print, copy, and download text and data-mine the content in such documents, for the purposes of academic research, subject always to the full Conditions of use: Corresponding Author: Piston, D.W. (dave.piston@vanderbilt.edu). Author Contributions G.J.K. designed the experiments, performed part of the microscopy experiments, analyzed and organized all data collected by other authors, and prepared the manuscript; K.L.H., and C.S.M. constructed mammalian expression vectors and performed part of the microscopy experiments; M.W.D. constructed mammalian expression vectors and contributed to editing the manuscript; D.W.P. contributed to the conceptual design and manuscript preparation.
2 Kremers et al. Page 2 state had excitation and emission maxima of 495 and 518 nm, respectively. Katushka, mkate, mtagrfp9, and mtagrfp-t (a variant with increased photostability)10 are all derived from the same sea anemone protein, EqFP578. All four proteins share the same chromophore (Met-Tyr-Gly), but we observed no photoconversion of mtagrfp or mtagrfp-t at any excitation wavelength. HcRed1 is a far-red fluorescent protein and originally developed from a sea anemone chromoprotein11. We observed red-to-green photoconversion after illumination at 405, 543, or 750 nm two-photon excitation, but excitation at 561 nm produced the green fluorescent species with much better efficiency (Fig. 1c, Table 1 and Supplementary Fig. 1 online). This green species had an emission peak at 510 nm and was excited at 458 or 488 nm. We also tested several red mfruit fluorescent proteins derived from DsRed 10, 12. Neither the widely used RFP, mcherry, nor its brighter variant, mapple110, exhibited any photoconversion upon one-photon excitation ranging from 405 nm to 633 nm or two-photon excitation at 750 nm. Both mplum and mraspberry showed a 10 nm blue-shift in the emission maximum after prolonged illumination with 405 nm or 543 nm light (Supplementary Fig. 2 online). These spectral changes were too small for use in optical highlighter applications and therefore we did not further characterize photoconversion of these proteins. In contrast, we observed considerable photoconversion for the orange fluorescent proteins, morange1 and morange2. We performed a complete characterization for both variants, and the photoconversion characteristics were practically identical, so we report the details only for morange1. Photoconversion occured at 458 nm or 488 nm laser excitation but is far less efficient at 405 nm excitation. Since most pc-fps photoconvert upon near-uv excitation, which can be harmful to cells, this visible photoconversion may be advantageous for live cell applications. morange1 is not normally excited at 633 nm, but after photoconversion, we observed bright red fluorescence (Fig. 1d). Using a white-light continuum laser, we measured photoconverted excitation and emission spectra in live cells (Fig. 1e, f). The excitation and emission maxima of the far-red species were approximately 610 nm and 640 nm, respectively. To our knowledge, this species exhibits the most redshifted absorption peak (610 nm) of any fluorescent protein thus far characterized. Kusabira-Orange (mko)13 has a similar chromophore and spectral properties to morange, although the fluorescent proteins are unrelated. Previously, mko has been shown to photoconvert to a green fluorescent species under widefield illumination at 436 nm14, but we did not observe this blue-shift in fluorescence using laser excitation. However, excitation of mko using the 514 nm or 543 nm laser resulted in a minor emission red-shift (Supplementary Fig. 3 online). This fluorescence was dim, and it remains unclear whether this is photoconversion or the result of selective photobleaching of an mko subpopulation. The dynamics of each of these photoconversion reactions were studied using micro-droplets of purified protein. Photoconversion of all pc-fps, except Katushka and mkate, resulted in a mono-exponential increase in the photoconverted species. The photoconversion rate increased supralinearly with laser power (Fig. 2a), which indicated that these photoconversions were multi-photon phenomena. Photoconversion of the pc-fps described here required more light than photoconversion of Kaede and Dendra2. Photoconversion was
3 Kremers et al. Page 3 generally maximal at lower laser powers because of increased photobleaching at higher power (Supplementary Fig. 4 online). An exception was morange for which we observed the greatest photoconversion at the highest laser power. morange1 photoconversion was also not significantly affected by ph (Fig. 2b), which permits optical highlighting in a wide ph-range of cellular compartments. In comparison, Kaede photoconversion, like all other reported pc-fps, was highly ph dependent, and Kaede was nonfluorescent at ph Many of the photoconversion dynamics were complex (Supplementary Fig. 4 online). For example, Dendra2 showed an initial increase in green fluorescence upon photoconversion, and similar behavior has been observed for Kaede15. We also observed a novel blue fluorescent Kaede species during its photoconversion at high laser power. Finally, photoconversion of Katushka and mkate showed an initial lag in both the decrease of red fluorescence and increase of green fluorescence. Most pc-fps perform better under prolonged illumination at low intensities, but many applications require fast photoconversion (i.e. within seconds). For this purpose, we measured photoconversion efficiency of pc-fps in HeLa cells with the excitation set to reach a maximum increase within 14 seconds. The new pc-fps displayed a smaller increase in photoconverted fluorescence than Kaede and Dendra2 (Table 1). However, the contrast change for morange was comparable to Kaede. For applications using only 488 nm excitation, mkate outperformed Kaede in photoconverted fluorescence and contrast ratio. The morange fluorescent proteins have great potential as optical highlighters because of their monomeric nature, excellent photoconversion contrast, and relative ph-insensitivity of photoconversion. The red-shifted spectral properties also enabled their use in dual-probe optical highlighting together with the green photoswitchable fluorescent protein Dronpa16 to allow selective highlighting of 4 individual cell populations (Fig. 3a-d). We achieved this, by first switching off all Dronpa fluorescence, followed subsequently by photoconversion of morange and switching Dronpa on again in selected cells. To show cell viability, we followed photoconverted H2B-mOrange2 in cells for 140 minutes through mitosis (Fig 3e and Supplementary Movie online). While photoconverted morange was less photostable than Kaede and Dendra2 (Supplementary Note 1 online), the photostability was sufficient for time-lapse imaging. morange variants were also well-suited to laser scanning microscopy applications, because they are photoconverted at 488 nm and their contrast ratio was maximal at the highest excitation powers. This was in contrast to Kaede and Dendra2, for which the contrast ratio was maximal under low light conditions (Supplementary Fig. 4 online). In fact, we observed a smaller contrast ratio for Kaede after laser-based photoconversion (148-fold change; Table 1) compared to widefield photoconversion ( 2000-fold change)6. Existing green-to-red optical highlighters are prone to spectral bleedthrough of emission from the original species into the photoconverted species detector. Thus, sequential twochannel imaging with multiple excitation wavelengths is required to quantify the photoconverted signal. In contrast, red-to-green optical highlighters do not suffer from such spectral overlap. Although the contrast ratio was similar for mkate and Kaede when using single wavelength excitation (488 nm), the increase in photoconverted fluorescence was 5 times higher for mkate (Table 1).
4 Kremers et al. Page 4 For the fluorescent proteins that exhibited red-to-green photoconversion, the spectral properties of the photoconverted green species resembled those of maturation intermediates. In fact, greening of DsRed fluorescence has been previously described17, but this effect was attributed to the presence of immature green DsRed chromophores within the DsRed tetramer. Our data suggest that this greening might be caused in part by photoconversion. If photoconversion yields an immature intermediate, then the photoconversion would be reversible as the immature intermediate matures again. Improving the maturation rate of these newly discovered pc-fps could thus yield novel reversible highlighters. It has also been shown that DsRed can photoconvert to a red-shifted fluorescent species upon high intensity irradiation at 532nm18. That photoconversion occurs through a sequential twophoton absorption process that involves decarboxylation of a glutamic acid residue adjacent to the chromophore. Since morange is derived from DsRed, it is tempting to speculate that a similar conversion mechanism is involved in morange photoconversion as well. If this speculation proves correct, structural determination of photoconverted morange might reveal new design principles for far red fluorescent proteins. Combined with previous reports, our data show that fluorescent protein photoconversion is fairly common, as eight out of twelve proteins investigated display this phenomenon. Our results demonstrate once more the complex photophysical characteristics of fluorescent proteins, and point toward the need for wariness in double labeling and fluorescence resonance energy transfer (FRET) experiments, where an unnoticed color-shift could lead to serious misinterpretations. Such a problem has already been reported for FRET experiments that use cyan and yellow fluorescent proteins and rely on acceptor photobleaching19. We also observed this blue-shifted photoconverted species for mvenus after laser-based bleaching (Supplementary Note 2 online). Further, the small spectral shifts observed for mplum and mraspberry would affect spectral unmixing experiments. Of course, detailed knowledge of the photophysics permits further uses for these fluorescent proteins. For instance, morange could also benefit superresolution microscopy approaches20 by leveraging the reduced autofluorescence in the red spectral region. Supplementary Material Acknowledgements Refer to Web version on PubMed Central for supplementary material. We thank D.M. Chudakov (Shemiakin-Ovchinnikov Institute of Bioorganic Chemistry, Moscow, Russia), A. Miyawaki (RIKEN, Wako, Saitama, Japan) and R.Y. Tsien (University of California at San Diego ) for providing plasmids encoding fluorescent protein variants. Measurements were performed in part at the Vanderbilt University Medical Center Cell Imaging Shared Resource (NIH grants CA68485, DK20593, DK58404) and at the QFM course at Mount Desert Island Biological Lab. Thanks to G. Daniels (Leica), J. Wailes (Zeiss), B. Burklow (Olympus), and S. Schwartz (Nikon) for assistance with their microscopes. Thanks to B. Livesay for help with the mvenus experiments. This work was supported by NIH Grant GM72048 (DWP).
5 Kremers et al. Page 5 Appendix Methods Cell culture Microscopy Power measurements HeLa and Cos7 cells were cultured in 35 mm glass bottom dishes (MatTek Corp.) or Delta T imaging chambers (Bioptechs) using Dulbecco s modified Eagle s medium (Invitrogen) supplemented with 10% fetal bovine serum, 100 U/ml penicillin, 100 μg/ml streptomycin and 2 mm L-glutamine. Cell were transfected using a transfection mix consisting either of 100 ng DNA and 1.5 μl Lipofectamine2000 (Invitrogen) in 100 μl Optimem (Invitrogen) media or 1 μg DNA with Effectene (Qiagen) in growth medium. Cells were imaged after 24 hours. Fluorescent proteins were expressed without a fusion partner or as histone H2B fusion proteins. The vector for Dronpa-mito was generated by substituting Dronpa for EGFP in pegfp-mito-7 (Clontech). Several laser scanning confocal microscopy platforms were used for data collection, including Zeiss LSM510 Meta and LSM710 (Zeiss), Olympus FV1000 (Olympus), Leica TCS SP5 X (Leica), and Nikon C1Si (Nikon). All microscopes were equipped with a Fluor design numerical aperture (NA) oil immersion objective. Imaging and photoconversion were done at 4 optical zoom using a 8 μs pixel dwell time and a 155 nm pixel size ( pixels). Red-to-green and green-to-red photoconversion was detected using the same filter settings. Green fluorescence was excited at 488 nm and detected through a nm bandpass filter. Red fluorescence was excited at 543 nm and detected through a 560 nm longpass emission filter. For single wavelength excitation applications green and red fluorescence were excited at 488 nm and detected simultaneous. For orangeto-red photoswitching, orange fluorescence was excited at 543 nm and detected through a nm bandpass emission filter. Far-red fluorescence was excited at 633 nm and detected through a 650 nm longpass emission filter. Dronpa fluorescence was detected using 488 nm excitation and a nm bandpass emission filter. Accurate power measurements of high NA oil immersion objectives are difficult to do directly. Therefore power measurements at the back aperture of the objective lens were done through a NA air objective using a Coherent Fieldmaster light meter (Coherent). During the measurement, the microscope was set to scan at its highest speed and maximum zoom. The measured average power was corrected for the off-time of the laser during scanning as well as the previously measured transmission efficiency of the objective lenses (90% for a NA air objective and 70% for a NA oil immersion objective). Photoconversion of protein purification Fluorescent proteins were expressed as his-tagged proteins in E. coli XL1Blue. Protein was purified using Talon metal affinity resin (Clontech) and buffer exchanged into phosphate buffered saline ph 7.4 using PD10 columns (GE Healthcare). Micro-droplets containing fluorescent protein were generated by mixing aqueous buffered fluorescent protein solutions
6 Kremers et al. Page 6 and octanol in a 1:10 volume ratio followed by sonication. Microscopy samples consisted of 4 μl emulsion between an object slide and a 22 mm square cover glass. Glassware was treated with methyltrimethoxysilane (Acros Organics) to prevent protein absorbance to the glass surface. The droplet size ( μm in diameter) was such that 3 to 10 droplets could be photoconverted simultaneously. Photoconversion was followed over time by sequential time-lapse imaging. Separate tracks were used to scan the original and photoconverted fluorescence, followed by a single scan with the photoconversion laser. Photoconversion rates were determined by fitting the increase in photoconverted fluorescence to a single exponential function. The rate constants were divided by the pixel dwell time (in seconds) to convert the values from scan -1 to s -1. ph dependence of photoconversion A series of titration buffers was prepared using 100 mm citric acid / Na citrate (ph 5.45), 100 mm KH 2 PO 4 / Na 2 HPO 4 (ph 6.72, 7.46, and 8.20), and 100 mm glycin / NaOH (ph 9.00). A 5-fold dilution of purified protein in each ph buffer was used to prepare microdroplet samples. Photoconversion was done as described for purified protein. The wavelength and laser power used for photoconversion were: morange1, 488 nm 632 μw; Kaede 405 nm 251 μw. Photoconversion in Mammalian cells Cells with medium fluorescence after transfection with equal amounts of plasmid DNA were selected for photoconversion. Initial and photoconverted fluorescence was imaged with the gain for each detector set at 850 volts. Laser power and duration of photoconversion were adjusted to reach a maximum increase in photoconverted fluorescence within 5 scan iterations. Changes in fluorescence were calculated after background subtraction, by dividing the fluorescence immediately after photoconversion by the fluorescence before photoactivation. In the mitosis experiments, cells expressing a fusion of morange2 to human histone H2B (morange2-h2b) were scanned to find cells entering prophase. One-half of the prophase nucleus was photoconverted and subsequently observed at lower laser power for a period ranging from 2 to 6 h. Photostability of photoconverted species in Mammalian cells Cells were transfected with equal amounts of plasmid DNA and cells with medium fluorescence were selected for photoconversion. Laser power and duration of photoconversion were selected to reach the maximum increase in photoconverted fluorescence within 5 scan iterations. The initial and photoconverted fluorescence was detected with detector gains set at 750 volts and 850 volts, respectively. The laser power for imaging the initial fluorescence as well as the photoconverted fluorescence immediately after photoconversion was adjusted to yield 2000 counts. The wavelength and power used for imaging the photoconverted species were: Dendra2, 543 nm 5.57 μw; Kaede, 543 nm 14 μw; Katushka and mkate, 488 nm 133 μw; morange 633 nm 312 μw.
7 Kremers et al. Page 7 Photoconversion of mvenus References mvenus protein purification and sample preparation were performed as described. Experiments were performed on a Zeiss LSM510 laser scanning confocal microscope equipped with a Chameleon mode-locked titanium sapphire laser oscillator (Chameleon, Coherent). Imaging was done using a Fluar 40x 1.3NA oil immersion objective at 4x optical zoom using a 6.4 μs pixel dwell time and a 110 nm pixel size ( pixels). Photoconversion in microdroplets was measured by time-lapse imaging. A single track was used to image the original and photoconverted fluorescence simultaneously upon twophoton excitation at 850 nm, followed by a single scan with the photoconversion laser at 514 nm. Yellow and photoconverted cyan fluorescence was detected through nm and nm bandpass emission filters, respectively. Photoconversion rates were determined by fitting the increase in photoconverted cyan fluorescence to a single exponential function. The rate constants were divided by the pixel dwell time (in s) to convert the values from scan -1 to s Patterson GH, Lippincott-Schwartz J. Science. 2002; 297: [PubMed: ] 2. Chudakov DM, et al. Nat Biotechnol. 2004; 22: [PubMed: ] 3. Gurskaya NG, et al. Nat Biotechnol. 2006; 24: [PubMed: ] 4. Chudakov DM, Lukyanov S, Lukyanov KA. Nat Protoc. 2007; 2: [PubMed: ] 5. Wiedenmann J, et al. Proc Natl Acad Sci U S A. 2004; 101: [PubMed: ] 6. Ando R, Hama H, Yamamoto-Hino M, Mizuno H, Miyawaki A. Proc Natl Acad Sci U S A. 2002; 99: [PubMed: ] 7. Tsutsui H, Karasawa S, Shimizu H, Nukina N, Miyawaki A. EMBO Rep. 2005; 6: [PubMed: ] 8. Shcherbo D, et al. Nat Methods. 2007; 4: [PubMed: ] 9. Merzlyak EM, et al. Nat Methods. 2007; 4: [PubMed: ] 10. Shaner NC, et al. Nat Methods. 2008; 5: [PubMed: ] 11. Gurskaya NG, et al. FEBS Lett. 2001; 507: [PubMed: ] 12. Shaner NC, et al. Nat Biotechnol. 2004; 22: [PubMed: ] 13. Karasawa S, Araki T, Nagai T, Mizuno H, Miyawaki A. Biochem J. 2004; 381: [PubMed: ] 14. Goedhart J, Vermeer JE, Adjobo-Hermans MJ, van Weeren L, Gadella TW Jr. PLoS ONE. 2007; 2:e1011. [PubMed: ] 15. Dittrich PS, Schafer SP, Schwille P. Biophys J. 2005; 89: [PubMed: ] 16. Habuchi S, et al. Proc Natl Acad Sci U S A. 2005; 102: [PubMed: ] 17. Marchant JS, Stutzmann GE, Leissring MA, LaFerla FM, Parker I. Nat Biotechnol. 2001; 19: [PubMed: ] 18. Habuchi S, et al. J Am Chem Soc. 2005; 127: [PubMed: ] 19. Valentin G, et al. Nat Methods. 2005; 2:801. [PubMed: ] 20. Betzig E, et al. Science. 2006; 313: [PubMed: ]
8 Kremers et al. Page 8 Figure 1. Photoconversion of mkate, HcRed1, and morange1 in HeLa cells. a) Cells expressing mkate were imaged before and after photoconversion induced with 750 nm two-photon excitation. Green and red fluorescence was collected simultaneously using 488 nm excitation. b) mkate emission spectra (ex 488 nm) showing gradual red to green photoconversion. c) HcRed1 emission spectra (ex 488 nm) showing gradual red to green photoconversion using 561 nm excitation. d) Cells expressing morange1 before and after photoconversion induced at 488 nm. e) Excitation spectrum of the morange1 far-red species
9 Kremers et al. Page 9 (black line) obtained by linear unmixing of the excitation spectra of a partially photoconverted cell (red line) and a non-converted cell (blue line). f) Emission spectra (ex 600 nm) of morange1 before and after photoconversion. Scale bars are 20 μm.
10 Kremers et al. Page 10 Figure 2. morange1 and Kaede photoconversion rates. a) Log-log plot of laser power dependence of photoconversion for Kaede (solid) and morange1 (dashed). Linear fits yield a slope for Kaede of 1.40 ± 0.07 (r = 0.997) and for morange1 of 1.59 ± 0.08 (r = 0.996). b) phdependence of Kaede (solid) and morange1 (dashed) photoconversion rates. Data points are averages of at least 5 measurements. Error bars indicate standard deviations.
11 Kremers et al. Page 11 Figure 3. Dual-probe optical highlighting with H2B-mOrange1 and Dronpa-mito. a) Before photoconversion cells have green labeled mitochondria and orange labeled nuclei (pseudocolored in red). b) Dronpa fluorescence is switched off with 74 μw 488 nm excitation; causing minimal photoconversion of morange1. c) morange1 is specifically photoconverted in the two upper nuclei with 742 μw 488 nm excitation (pseudo-colored blue). d) Finally, Dronpa fluorescence is switched on again in the two left-hand cells with 248 μw 405 nm illumination. Boxes indicate active regions of photoconversion. e) Time-lapse images from Supplementary Movie showing mitosis of a HeLa cell expressing H2B-mOrange2. Part of the nucleus was photoconverted during prophase (pseudo-colored in blue). The images in panel e are contrast-enhanced to help visualization. Scale bars are 20 μm.
12 Kremers et al. Page 12 Table 1 Photoconversion properties Color change Photoactivation wavelength and power Fold activation a Residual fluorescence b Change in contrast c e 27 (13) Katushka Red to green 750 nm 17.9 mw 6 ± 2 d 0.22 ± 0.07 (0.30 ± 0.02) 405 nm 1.4 mw 7 ± ± 0.03 (0.68 ± 0.03) 10 (10) 561 nm 0.32 mw 5 ± ± mkate Red to green 750 nm 17.9 mw 11 ± ± 0.08 (0.94 ± 0.09) 19 (12) 405 nm 1.4 mw 10 ± ± 0.03 (0.8 ± 0.6) 26 (13) 561 nm 0.32 mw 2.0 ± ± HcRed1 Red to green 561 nm 0.32 mw 26 ± ± morange1 Orange to far-red 488 nm 1.2 mw 16 ± ± morange2 Orange to far-red 488 nm 1.2 mw 16 ± ± Kaede Green to red 405 nm 0.38 mw 28 ± 10 (1.8 ± 0.1) 0.19 ± (10) Dendra2 Green to red 405 nm 0.25 mw 47 ± 26 (1.2 ± 0.4) 0.48 ± (2) a Fold increase in photoconverted fluorescence. b Fraction of original fluorescence remaining after photoconversion. c The change in contrast is determined by the increase in photoconverted fluorescence divided by the residual original fluorescence. d Values are mean ± s.d (n = 7). e Values between brackets are for single wavelength excitation (ex 488 nm).
Feb No. 22. Dmitriy Chudakov, Timo Zimmermann
 Feb. 2007 No. 22 Using photoactivatable fluorescent proteins Dendra2 and PS-CFP2 to track protein movement with Leica TCS SP5 Dmitriy Chudakov, Timo Zimmermann LAS AF: FRAP Wizard Live Data Mode Photoconversion
Feb. 2007 No. 22 Using photoactivatable fluorescent proteins Dendra2 and PS-CFP2 to track protein movement with Leica TCS SP5 Dmitriy Chudakov, Timo Zimmermann LAS AF: FRAP Wizard Live Data Mode Photoconversion
A guide to choosing fluorescent proteins
 A guide to choosing fluorescent proteins Nathan C Shaner 1,2, Paul A Steinbach 1,3 & Roger Y Tsien 1,3,4 The recent explosion in the diversity of available fluorescent proteins (FPs) 1 16 promises a wide
A guide to choosing fluorescent proteins Nathan C Shaner 1,2, Paul A Steinbach 1,3 & Roger Y Tsien 1,3,4 The recent explosion in the diversity of available fluorescent proteins (FPs) 1 16 promises a wide
Spectral Separation of Multifluorescence Labels with the LSM 510 META
 Microscopy from Carl Zeiss Spectral Separation of Multifluorescence Labels with the LSM 510 META Indians living in the South American rain forest can distinguish between almost 200 hues of green in their
Microscopy from Carl Zeiss Spectral Separation of Multifluorescence Labels with the LSM 510 META Indians living in the South American rain forest can distinguish between almost 200 hues of green in their
Rational design of true monomeric and bright photoconvertible
 Rational design of true monomeric and bright photoconvertible fluorescent proteins Mingshu Zhang, Hao Chang, Yongdeng Zhang, Junwei Yu, Lijie Wu, Wei Ji, Juanjuan Chen, Bei Liu, Jingze Lu, Yingfang Liu,
Rational design of true monomeric and bright photoconvertible fluorescent proteins Mingshu Zhang, Hao Chang, Yongdeng Zhang, Junwei Yu, Lijie Wu, Wei Ji, Juanjuan Chen, Bei Liu, Jingze Lu, Yingfang Liu,
D e c N o. 2 8
 D e c. 2 0 0 7 N o. 2 8 CONFOCAL APPLICATION LETTER resolution FRET Acceptor Photobleaching LAS AF Application Wizard FRET with Leica TCS SP5 LAS AF Version 1.7.0 Introduction Fluorescence Resonance Energy
D e c. 2 0 0 7 N o. 2 8 CONFOCAL APPLICATION LETTER resolution FRET Acceptor Photobleaching LAS AF Application Wizard FRET with Leica TCS SP5 LAS AF Version 1.7.0 Introduction Fluorescence Resonance Energy
Supplementary Material
 Supplementary Material Supplementary Table 1. Definitions of variants described in this work. Protein variant Substitutions and modifications relative to parent mtfp1 a = cfp484 b + delete all residues
Supplementary Material Supplementary Table 1. Definitions of variants described in this work. Protein variant Substitutions and modifications relative to parent mtfp1 a = cfp484 b + delete all residues
Photoactivatable mcherry for high-resolution two-color fluorescence microscopy
 Photoactivatable mcherry for high-resolution two-color fluorescence microscopy Fedor V Subach 1 3, George H Patterson,3, Suliana Manley, Jennifer M Gillette, Jennifer Lippincott-Schwartz & Vladislav V
Photoactivatable mcherry for high-resolution two-color fluorescence microscopy Fedor V Subach 1 3, George H Patterson,3, Suliana Manley, Jennifer M Gillette, Jennifer Lippincott-Schwartz & Vladislav V
Post-expansion antibody delivery, after epitope-preserving homogenization.
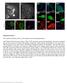 Supplementary Figure 1 Post-expansion antibody delivery, after epitope-preserving homogenization. (a, b) Wide-field fluorescence images of Thy1-YFP-expressing mouse brain hemisphere slice before expansion
Supplementary Figure 1 Post-expansion antibody delivery, after epitope-preserving homogenization. (a, b) Wide-field fluorescence images of Thy1-YFP-expressing mouse brain hemisphere slice before expansion
Fluorescent protein. Origin of fluorescent proteins. Structural and biophysical analysis of FPs. Application in molecular biology and biotechnology
 Origin of fluorescent proteins types of FPs mechanism of fluorescence Fluorescent protein Structural and biophysical analysis of FPs Application in molecular biology and biotechnology Variety of organisms
Origin of fluorescent proteins types of FPs mechanism of fluorescence Fluorescent protein Structural and biophysical analysis of FPs Application in molecular biology and biotechnology Variety of organisms
Fluorescence Light Microscopy for Cell Biology
 Fluorescence Light Microscopy for Cell Biology Why use light microscopy? Traditional questions that light microscopy has addressed: Structure within a cell Locations of specific molecules within a cell
Fluorescence Light Microscopy for Cell Biology Why use light microscopy? Traditional questions that light microscopy has addressed: Structure within a cell Locations of specific molecules within a cell
Supplementary Figure 1. Design of linker truncation library between Nluc and either Venus or mneongreen
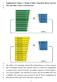 1 2 3 Supplementary Figure 1. Design of linker truncation library between Nluc and either Venus or mneongreen 4 5 6 7 8 9 10 The cdna of C-terminally deleted FPs (mneongreen or Venus) mutants and N-terminally
1 2 3 Supplementary Figure 1. Design of linker truncation library between Nluc and either Venus or mneongreen 4 5 6 7 8 9 10 The cdna of C-terminally deleted FPs (mneongreen or Venus) mutants and N-terminally
Improving the photostability of bright monomeric orange and red fluorescent proteins
 Improving the photostability of bright monomeric orange and red fluorescent proteins Nathan C Shaner 1,5, Michael Z Lin 1,2, Michael R McKeown 1,2, Paul A Steinbach 1,2, Kristin L Hazelwood 4, Michael
Improving the photostability of bright monomeric orange and red fluorescent proteins Nathan C Shaner 1,5, Michael Z Lin 1,2, Michael R McKeown 1,2, Paul A Steinbach 1,2, Kristin L Hazelwood 4, Michael
Bright far-red fluorescent protein for whole-body imaging
 Bright far-red fluorescent protein for whole-body imaging Dmitry Shcherbo, Ekaterina M Merzlyak, Tatiana V Chepurnykh, Arkady F Fradkov, Galina V Ermakova, Elena A Solovieva, Konstantin A Lukyanov, Ekaterina
Bright far-red fluorescent protein for whole-body imaging Dmitry Shcherbo, Ekaterina M Merzlyak, Tatiana V Chepurnykh, Arkady F Fradkov, Galina V Ermakova, Elena A Solovieva, Konstantin A Lukyanov, Ekaterina
Localization Microscopy
 Localization Microscopy Theory, Sample Prep & Practical Considerations Patrina Pellett & Ann McEvoy Applications Scientist GE Healthcare, Cell Technologies May 27 th, 2015 Localization Microscopy Talk
Localization Microscopy Theory, Sample Prep & Practical Considerations Patrina Pellett & Ann McEvoy Applications Scientist GE Healthcare, Cell Technologies May 27 th, 2015 Localization Microscopy Talk
Improving the photostability of bright monomeric orange and red fluorescent proteins
 ARTICLES Improving the photostability of bright monomeric orange and red fluorescent proteins 2008 Nature Publishing Group h ttp://www.nature.com/natur e method s Nathan C Shaner 1,5, Michael Z Lin 1,2,
ARTICLES Improving the photostability of bright monomeric orange and red fluorescent proteins 2008 Nature Publishing Group h ttp://www.nature.com/natur e method s Nathan C Shaner 1,5, Michael Z Lin 1,2,
 Introduction to Fluorescent Proteins The discovery of green fluorescent protein in the early 1960s ultimately heralded a new era in cell biology by enabling investigators to apply molecular cloning methods,
Introduction to Fluorescent Proteins The discovery of green fluorescent protein in the early 1960s ultimately heralded a new era in cell biology by enabling investigators to apply molecular cloning methods,
Fluorescence microscopy
 Fluorescence microscopy 1 Fluorescence microscopies basic fluorescence, fluorophores Deconvolution Confocal Two-photon/multi-photon 4Pi Light sheet Total internal reflection STED FRAP/FLIP/FCS FRET PALM/STORM/iPALM
Fluorescence microscopy 1 Fluorescence microscopies basic fluorescence, fluorophores Deconvolution Confocal Two-photon/multi-photon 4Pi Light sheet Total internal reflection STED FRAP/FLIP/FCS FRET PALM/STORM/iPALM
Confocal Microscopes. Evolution of Imaging
 Confocal Microscopes and Evolution of Imaging Judi Reilly Hans Richter Massachusetts Institute of Technology Environment, Health & Safety Office Radiation Protection What is Confocal? Pinhole diaphragm
Confocal Microscopes and Evolution of Imaging Judi Reilly Hans Richter Massachusetts Institute of Technology Environment, Health & Safety Office Radiation Protection What is Confocal? Pinhole diaphragm
Special Techniques 1. Mark Scott FILM Facility
 Special Techniques 1 Mark Scott FILM Facility SPECIAL TECHNIQUES Multi-photon microscopy Second Harmonic Generation FRAP FRET FLIM In-vivo imaging TWO-PHOTON MICROSCOPY Alternative to confocal and deconvolution
Special Techniques 1 Mark Scott FILM Facility SPECIAL TECHNIQUES Multi-photon microscopy Second Harmonic Generation FRAP FRET FLIM In-vivo imaging TWO-PHOTON MICROSCOPY Alternative to confocal and deconvolution
Optimization of fluorescent proteins for novel quantitative multiparameter microscopy approaches Kremers, G.J.
 UvA-DARE (Digital Academic Repository) Optimization of fluorescent proteins for novel quantitative multiparameter microscopy approaches Kremers, G.J. Link to publication Citation for published version
UvA-DARE (Digital Academic Repository) Optimization of fluorescent proteins for novel quantitative multiparameter microscopy approaches Kremers, G.J. Link to publication Citation for published version
F* techniques: FRAP, FLIP, FRET, FLIM,
 F* techniques: FRAP, FLIP, FRET, FLIM, FCS Antonia Göhler March 2015 Fluorescence explained in the Bohr model Absorption of light (blue) causes an electron to move to a higher energy orbit. After a particular
F* techniques: FRAP, FLIP, FRET, FLIM, FCS Antonia Göhler March 2015 Fluorescence explained in the Bohr model Absorption of light (blue) causes an electron to move to a higher energy orbit. After a particular
Product Manual. RFP ELISA Kit. Catalog Number. FOR RESEARCH USE ONLY Not for use in diagnostic procedures
 Product Manual RFP ELISA Kit Catalog Number AKR-122 96 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Red fluorescent protein (DsRed) is a spontaneously fluorescent protein
Product Manual RFP ELISA Kit Catalog Number AKR-122 96 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Red fluorescent protein (DsRed) is a spontaneously fluorescent protein
Low Background D-A-D Type Fluorescent Probe for Imaging of Biothiols
 Electronic Supplementary Material (ESI) for Journal of Materials Chemistry B. This journal is The Royal Society of Chemistry 2018 Electronic supplementary information Low Background D-A-D Type Fluorescent
Electronic Supplementary Material (ESI) for Journal of Materials Chemistry B. This journal is The Royal Society of Chemistry 2018 Electronic supplementary information Low Background D-A-D Type Fluorescent
Chapter 4. the biological community to assay for protein-protein interactions. FRET describes the
 31 Chapter 4 Determination of nachr stoichiometry using Normalized Försters Resonance Energy Transfer (NFRET) Försters resonance energy transfer (FRET) has become a technique widely used in the biological
31 Chapter 4 Determination of nachr stoichiometry using Normalized Försters Resonance Energy Transfer (NFRET) Försters resonance energy transfer (FRET) has become a technique widely used in the biological
Supporting Information
 Electronic Supplementary Material (ESI) for Materials Chemistry Frontiers. This journal is the Partner Organisations 2017 Supporting Information Supramolecular Conjugated Polymer Materials for Organelle
Electronic Supplementary Material (ESI) for Materials Chemistry Frontiers. This journal is the Partner Organisations 2017 Supporting Information Supramolecular Conjugated Polymer Materials for Organelle
Aryl-thioether substituted nitrobenzothiadiazole probe for selective detection of cysteine and homocysteine
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2015 Electronic Supplementary Information (ESI) for Chemical Communications This journal is The Royal
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2015 Electronic Supplementary Information (ESI) for Chemical Communications This journal is The Royal
λ N -GFP: an RNA reporter system for live-cell imaging
 λ N -GFP: an RNA reporter system for live-cell imaging Nathalie Daigle & Jan Ellenberg Supplementary Figures and Text: Supplementary Figure 1 localization in the cytoplasm. 4 λ N22-3 megfp-m9 serves as
λ N -GFP: an RNA reporter system for live-cell imaging Nathalie Daigle & Jan Ellenberg Supplementary Figures and Text: Supplementary Figure 1 localization in the cytoplasm. 4 λ N22-3 megfp-m9 serves as
spectrum of applications beyond the simple tracking of tagged biomolecules in living cells is now becoming fully appreciated.
 Introduction to Fluorescent Proteins From http://www.microscopyu.com/print/articles/livecellim aging/fpintro-print.html The discovery of green fluorescent protein in the early 1960s ultimately heralded
Introduction to Fluorescent Proteins From http://www.microscopyu.com/print/articles/livecellim aging/fpintro-print.html The discovery of green fluorescent protein in the early 1960s ultimately heralded
Cell Surface-Anchored Fluorescent Aptamer Sensor Enables. Imaging of Chemical Transmitter Dynamics
 Supporting Information for Cell Surface-Anchored Fluorescent Aptamer Sensor Enables Imaging of Chemical Transmitter Dynamics Takeshi Tokunaga, Shigeyuki Namiki, Katsuhiro Yamada, Takahiro Imaishi, Hiroshi
Supporting Information for Cell Surface-Anchored Fluorescent Aptamer Sensor Enables Imaging of Chemical Transmitter Dynamics Takeshi Tokunaga, Shigeyuki Namiki, Katsuhiro Yamada, Takahiro Imaishi, Hiroshi
Super-resolution Microscopy
 Semr oc kwhi t epaperser i es : 1. Introduction Super-resolution Microscopy Fluorescence microscopy has revolutionized the study of biological samples. Ever since the invention of fluorescence microscopy
Semr oc kwhi t epaperser i es : 1. Introduction Super-resolution Microscopy Fluorescence microscopy has revolutionized the study of biological samples. Ever since the invention of fluorescence microscopy
Design for Manufacturability (DFM) in the Life Sciences
 T E C H N I C A L N O T E Design for Manufacturability (DFM) in the Life Sciences Fluorescence Spectroscopy Product Platform Realized with TracePro TM Suite of Opto-Mechanical Design Software Tools Authors:
T E C H N I C A L N O T E Design for Manufacturability (DFM) in the Life Sciences Fluorescence Spectroscopy Product Platform Realized with TracePro TM Suite of Opto-Mechanical Design Software Tools Authors:
A fusion tag enabling optical marking and tracking of proteins
 Journal of Microscopy, Vol. 222, Pt 1 April 2006, pp. 15 21 Received 20 March 2005; accepted 21 October 2005 A fusion tag enabling optical marking and tracking of proteins Blackwell Publishing Ltd and
Journal of Microscopy, Vol. 222, Pt 1 April 2006, pp. 15 21 Received 20 March 2005; accepted 21 October 2005 A fusion tag enabling optical marking and tracking of proteins Blackwell Publishing Ltd and
Imaging of BacMam Transfected U-2 OS Cells
 A p p l i c a t i o n N o t e Imaging of BacMam Transfected U-2 OS Cells Optimization of Transfection Conditions Using the Cytation 3 Multi- Mode Reader and Gen5 Data Analysis Software Paul Held Ph. D.
A p p l i c a t i o n N o t e Imaging of BacMam Transfected U-2 OS Cells Optimization of Transfection Conditions Using the Cytation 3 Multi- Mode Reader and Gen5 Data Analysis Software Paul Held Ph. D.
Janos Szabad Department of Biology University of Szeged 6720 Szeged, Somogyi str
 Janos Szabad Department of Biology University of Szeged 6720 Szeged, Somogyi str. 4. E-mail: szabad.janos@med.u-szeged.hu - Through the use of antibodies - against the protein - against a fusion partner
Janos Szabad Department of Biology University of Szeged 6720 Szeged, Somogyi str. 4. E-mail: szabad.janos@med.u-szeged.hu - Through the use of antibodies - against the protein - against a fusion partner
A quantitative protocol for intensity-based live cell FRET imaging.
 A quantitative protocol for intensity-based live cell FRET imaging. Kaminski CF, Rees EJ, Schierle GS. Methods Mol Biol. 2014; 1076:445-454. Department of Chemical Engineering and Biotechnology, Pembroke
A quantitative protocol for intensity-based live cell FRET imaging. Kaminski CF, Rees EJ, Schierle GS. Methods Mol Biol. 2014; 1076:445-454. Department of Chemical Engineering and Biotechnology, Pembroke
Photon Upconversion Sensitized Nanoprobes for
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2014 Supporting Information Photon Upconversion Sensitized Nanoprobes for Sensing and Imaging of ph
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2014 Supporting Information Photon Upconversion Sensitized Nanoprobes for Sensing and Imaging of ph
Confocal Microscopy Analyzes Cells
 Choosing Filters for Fluorescence A Laurin Publication Photonic Solutions for Biotechnology and Medicine November 2002 Confocal Microscopy Analyzes Cells Reprinted from the November 2002 issue of Biophotonics
Choosing Filters for Fluorescence A Laurin Publication Photonic Solutions for Biotechnology and Medicine November 2002 Confocal Microscopy Analyzes Cells Reprinted from the November 2002 issue of Biophotonics
BIO 315 Lab Exam I. Section #: Name:
 Section #: Name: Also provide this information on the computer grid sheet given to you. (Section # in special code box) BIO 315 Lab Exam I 1. In labeling the parts of a standard compound light microscope
Section #: Name: Also provide this information on the computer grid sheet given to you. (Section # in special code box) BIO 315 Lab Exam I 1. In labeling the parts of a standard compound light microscope
Supplementary Information
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2015 Supplementary Information Title: Mitochondria-targeted fluorescent thermometer monitors intracellular
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2015 Supplementary Information Title: Mitochondria-targeted fluorescent thermometer monitors intracellular
Quantum Dot-Fluorescent Protein FRET Probe for Monitoring Intracellular ph. Supporting Information
 Quantum Dot-Fluorescent Protein FRET Probe for Monitoring Intracellular ph Allison M. Dennis, Won Jong Rhee, Steven N. Dublin, David Sotto, and Gang Bao Supporting Information Protein Engineering, Expression,
Quantum Dot-Fluorescent Protein FRET Probe for Monitoring Intracellular ph Allison M. Dennis, Won Jong Rhee, Steven N. Dublin, David Sotto, and Gang Bao Supporting Information Protein Engineering, Expression,
Electronic Supplementary Information
 Electronic Supplementary Information FRET-based probing to gain direct information on sirna sustainability in live cells: Asymmetric degradation of sirna strands Seonmi Shin, a Hyun-Mi Kwon, b Kyung-Sik
Electronic Supplementary Information FRET-based probing to gain direct information on sirna sustainability in live cells: Asymmetric degradation of sirna strands Seonmi Shin, a Hyun-Mi Kwon, b Kyung-Sik
Advanced fluorescence microscopy techniques
 Practice-oriented, student-friendly modernization of the biomedical education for strengthening the international competitiveness of the rural Hungarian universities TÁMOP-4.1.1.C-13/1/KONV-2014-0001 Advanced
Practice-oriented, student-friendly modernization of the biomedical education for strengthening the international competitiveness of the rural Hungarian universities TÁMOP-4.1.1.C-13/1/KONV-2014-0001 Advanced
Improving membrane voltage measurements using FRET with new fluorescent proteins
 correction notice Nat. Methods 5, 683 685 (2008) Improving membrane voltage measurements using FRET with new fluorescent proteins Hidekazu Tsutsui, Satoshi Karasawa, Yasushi Okamura & Atsushi Miyawaki
correction notice Nat. Methods 5, 683 685 (2008) Improving membrane voltage measurements using FRET with new fluorescent proteins Hidekazu Tsutsui, Satoshi Karasawa, Yasushi Okamura & Atsushi Miyawaki
Supplementary Information
 Supplementary Information Supplemental Figure 1. VVD-III purifies in a reduced state. (a) The cell pellet of VVD-III (VVD 36 C108A:M135I:M165I) is green compared to VVD-I (wild type VVD 36) due to the
Supplementary Information Supplemental Figure 1. VVD-III purifies in a reduced state. (a) The cell pellet of VVD-III (VVD 36 C108A:M135I:M165I) is green compared to VVD-I (wild type VVD 36) due to the
SUPPLEMENTARY INFORMATION
 a 14 12 Densitometry (AU) 1 8 6 4 2 t b 16 NMHC-IIA GAPDH NMHC-IIB Densitometry (AU) 14 12 1 8 6 4 2 1 nm 1 nm 1 nm 1 nm sirna 1 nm 1 nm Figure S1 S4 Quantification of protein levels. (a) The microtubule
a 14 12 Densitometry (AU) 1 8 6 4 2 t b 16 NMHC-IIA GAPDH NMHC-IIB Densitometry (AU) 14 12 1 8 6 4 2 1 nm 1 nm 1 nm 1 nm sirna 1 nm 1 nm Figure S1 S4 Quantification of protein levels. (a) The microtubule
Supporting Information
 Supporting Information Azido Push Pull Fluorogens Photoactivate to Produce Bright Fluorescent Labels Samuel J. Lord, Hsiao-lu D. Lee, Reichel Samuel, Ryan Weber, Na Liu, Nicholas R. Conley, Michael A.
Supporting Information Azido Push Pull Fluorogens Photoactivate to Produce Bright Fluorescent Labels Samuel J. Lord, Hsiao-lu D. Lee, Reichel Samuel, Ryan Weber, Na Liu, Nicholas R. Conley, Michael A.
Active targeting of the nucleus using non-peptidic boronate tags
 Supporting Information Active targeting of the nucleus using non-peptidic boronate tags Rui Tang 1, Ming Wang 2, Moumita Ray 1, Ying Jiang 1, Ziwen Jiang 1, Qiaobing Xu 2 *, Vincent M. Rotello 1 * 1 Department
Supporting Information Active targeting of the nucleus using non-peptidic boronate tags Rui Tang 1, Ming Wang 2, Moumita Ray 1, Ying Jiang 1, Ziwen Jiang 1, Qiaobing Xu 2 *, Vincent M. Rotello 1 * 1 Department
Fluorescence Nanoscopy
 Fluorescence Nanoscopy Keith A. Lidke University of New Mexico panda3.phys.unm.edu/~klidke/index.html Optical Microscopy http://en.wikipedia.org/wiki/k%c3%b6hler_illumination 30 µm Fluorescent Probes Michalet
Fluorescence Nanoscopy Keith A. Lidke University of New Mexico panda3.phys.unm.edu/~klidke/index.html Optical Microscopy http://en.wikipedia.org/wiki/k%c3%b6hler_illumination 30 µm Fluorescent Probes Michalet
STORM/PALM. Super Resolution Microscopy 10/31/2011. Looking into microscopic world of life
 Super Resolution Microscopy STORM/PALM Bo Huang Department of Pharmaceutical Chemistry, UCSF CSHL Quantitative Microscopy, 1/31/211 Looking into microscopic world of life 1 µm 1 µm 1 nm 1 nm 1 nm 1 Å Naked
Super Resolution Microscopy STORM/PALM Bo Huang Department of Pharmaceutical Chemistry, UCSF CSHL Quantitative Microscopy, 1/31/211 Looking into microscopic world of life 1 µm 1 µm 1 nm 1 nm 1 nm 1 Å Naked
Photoactivatable mcherry for high-resolution two-color fluorescence microscopy
 nature methods Photoactivatable mcherry for high-resolution two-color fluorescence microscopy Fedor V Subach, George H Patterson, Suliana Manley, Jennifer M Gillette, Jennifer Lippincott-Schwartz & Vladislav
nature methods Photoactivatable mcherry for high-resolution two-color fluorescence microscopy Fedor V Subach, George H Patterson, Suliana Manley, Jennifer M Gillette, Jennifer Lippincott-Schwartz & Vladislav
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10016 Supplementary discussion on binding site density for protein complexes on the surface: The density of biotin sites on the chip is ~10 3 biotin-peg per µm 2. The biotin sites are
doi:10.1038/nature10016 Supplementary discussion on binding site density for protein complexes on the surface: The density of biotin sites on the chip is ~10 3 biotin-peg per µm 2. The biotin sites are
Accessorize Your Imaging. Ian Clements Invitrogen Corp
 Accessorize Your Imaging Ian Clements Invitrogen Corp Imaging Tools and Accessories Antifade Reagents Prolong Gold SlowFade Gold Image-iT FX Signal Enhancer FocalCheck Microscope Test Slides Fluorescent
Accessorize Your Imaging Ian Clements Invitrogen Corp Imaging Tools and Accessories Antifade Reagents Prolong Gold SlowFade Gold Image-iT FX Signal Enhancer FocalCheck Microscope Test Slides Fluorescent
Advanced fluorescence microscopy techniques
 Practice-oriented, student-friendly modernization of the biomedical education for strengthening the international competitiveness of the rural Hungarian universities TÁMOP-4.1.1.C-13/1/KONV-2014-0001 Advanced
Practice-oriented, student-friendly modernization of the biomedical education for strengthening the international competitiveness of the rural Hungarian universities TÁMOP-4.1.1.C-13/1/KONV-2014-0001 Advanced
Workshop advanced light microscopy
 Workshop advanced light microscopy Multi-mode confocal laser scanning microscope Jan Willem Borst Laboratory of Biochemistry Biomolecular Networks www.bic.wur.nl MicroSpectroscopy Centre Wageningen Microspectroscopy
Workshop advanced light microscopy Multi-mode confocal laser scanning microscope Jan Willem Borst Laboratory of Biochemistry Biomolecular Networks www.bic.wur.nl MicroSpectroscopy Centre Wageningen Microspectroscopy
A bright and photostable photoconvertible fluorescent protein
 correction notice Nat. Methods 6, 131 133 (2009) A bright and photostable photoconvertible fluorescent protein Sean A McKinney, Christopher S Murphy, Kristin L Hazelwood, Michael W Davidson & Loren L Looger
correction notice Nat. Methods 6, 131 133 (2009) A bright and photostable photoconvertible fluorescent protein Sean A McKinney, Christopher S Murphy, Kristin L Hazelwood, Michael W Davidson & Loren L Looger
Aoki et al.,
 JCB: SUPPLEMENTAL MATERIAL Aoki et al., http://www.jcb.org/cgi/content/full/jcb.200609017/dc1 Supplemental materials and methods Calculation of the raw number of translocated proteins First, the average
JCB: SUPPLEMENTAL MATERIAL Aoki et al., http://www.jcb.org/cgi/content/full/jcb.200609017/dc1 Supplemental materials and methods Calculation of the raw number of translocated proteins First, the average
ab CytoPainter ER Staining Kit Red Fluorescence
 ab139482 CytoPainter ER Staining Kit Red Fluorescence Instructions for Use Designed to detect Human endoplasmic reticulum by microscopy. This product is for research use only and is not intended for diagnostic
ab139482 CytoPainter ER Staining Kit Red Fluorescence Instructions for Use Designed to detect Human endoplasmic reticulum by microscopy. This product is for research use only and is not intended for diagnostic
Microscopy from Carl Zeiss
 Microscopy from Carl Zeiss LSM 710 In Tune with Your Application Enjoy new freedom in selecting fluorescent dyes with In Tune, the new laser system for the LSM 710. Whatever the wavelength, you can match
Microscopy from Carl Zeiss LSM 710 In Tune with Your Application Enjoy new freedom in selecting fluorescent dyes with In Tune, the new laser system for the LSM 710. Whatever the wavelength, you can match
BIO 315 Lab Exam I. Section #: Name:
 Section #: Name: Also provide this information on the computer grid sheet given to you. (Section # in special code box) BIO 315 Lab Exam I 1. In labeling the parts of a standard compound light microscope
Section #: Name: Also provide this information on the computer grid sheet given to you. (Section # in special code box) BIO 315 Lab Exam I 1. In labeling the parts of a standard compound light microscope
.acceptor photobleaching; fluorescence lifetime imaging. Imaging probes for microscopy
 Imaging probes for microscopy Fluorescent proteins for FRET microscopy: Monitoring protein interactions in living cells Richard N. Day 1) and Michael W. Davidson 2) The discovery and engineering of novel
Imaging probes for microscopy Fluorescent proteins for FRET microscopy: Monitoring protein interactions in living cells Richard N. Day 1) and Michael W. Davidson 2) The discovery and engineering of novel
Spectral imaging and its use in the measurement of Förster resonance energy transfer in living cells
 Laboratory Techniques in Biochemistry and Molecular Biology, Volume 33 FRET and FLIM Techniques T. W. J. Gadella (Editor) CHAPTER 8 Spectral imaging and its use in the measurement of Förster resonance
Laboratory Techniques in Biochemistry and Molecular Biology, Volume 33 FRET and FLIM Techniques T. W. J. Gadella (Editor) CHAPTER 8 Spectral imaging and its use in the measurement of Förster resonance
Concept review: Fluorescence
 16 Concept review: Fluorescence Some definitions: Chromophore. The structural feature of a molecule responsible for the absorption of UV or visible light. Fluorophore. A chromophore that remits an absorbed
16 Concept review: Fluorescence Some definitions: Chromophore. The structural feature of a molecule responsible for the absorption of UV or visible light. Fluorophore. A chromophore that remits an absorbed
Obtaining More Accurate Signals: Spatiotemporal Imaging of Cancer Sites Enabled by a Photoactivatable Aptamer-Based Strategy
 Supporting Information Obtaining More Accurate Signals: Spatiotemporal Imaging of Cancer Sites Enabled by a Photoactivatable Aptamer-Based Strategy Heng Xiao,,, Yuqi Chen,, Erfeng Yuan,, Wei Li, Zhuoran
Supporting Information Obtaining More Accurate Signals: Spatiotemporal Imaging of Cancer Sites Enabled by a Photoactivatable Aptamer-Based Strategy Heng Xiao,,, Yuqi Chen,, Erfeng Yuan,, Wei Li, Zhuoran
Labeling cells with photoconvertible fluorescent proteins in zebrafish. Instructors: Debbie Yelon and Xin-Xin Zeng
 Labeling cells with photoconvertible fluorescent proteins in zebrafish Instructors: Debbie Yelon (dyelon@ucsd.edu) and Xin-Xin Zeng (xxzeng@ucsd.edu) Introduction: Photoconvertible (aka photoswitchable,
Labeling cells with photoconvertible fluorescent proteins in zebrafish Instructors: Debbie Yelon (dyelon@ucsd.edu) and Xin-Xin Zeng (xxzeng@ucsd.edu) Introduction: Photoconvertible (aka photoswitchable,
SUPPLEMENTARY FIGURES
 SYNERGISTIC STRATEGY FOR MULTICOLOR TWO-PHOTON MICROSCOPY: APPLICATION TO THE ANALYSIS OF GERMINAL CENTER REACTIONS IN VIVO ASYLKHAN RAKHYMZHAN, RUTH LEBEN, HANNA ZIMMERMANN, ROBERT GÜNTHER, PEGGY MEX,
SYNERGISTIC STRATEGY FOR MULTICOLOR TWO-PHOTON MICROSCOPY: APPLICATION TO THE ANALYSIS OF GERMINAL CENTER REACTIONS IN VIVO ASYLKHAN RAKHYMZHAN, RUTH LEBEN, HANNA ZIMMERMANN, ROBERT GÜNTHER, PEGGY MEX,
Photoactivatable fluorescein derivatives with. azidomethyl caging groups for tracing. oligonucleotides in living human cells
 Supporting Information Photoactivatable fluorescein derivatives with azidomethyl caging groups for tracing oligonucleotides in living human cells Kazuhiro Furukawa, a,b Hiroshi Abe, * a Satoshi Tsuneda,
Supporting Information Photoactivatable fluorescein derivatives with azidomethyl caging groups for tracing oligonucleotides in living human cells Kazuhiro Furukawa, a,b Hiroshi Abe, * a Satoshi Tsuneda,
Fast, three-dimensional super-resolution imaging of live cells
 Nature Methods Fast, three-dimensional super-resolution imaging of live cells Sara A Jones, Sang-Hee Shim, Jiang He & Xiaowei Zhuang Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure 3
Nature Methods Fast, three-dimensional super-resolution imaging of live cells Sara A Jones, Sang-Hee Shim, Jiang He & Xiaowei Zhuang Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure 3
Fluorescent labeling of the nuclear envelope by localizing GFP on the inner nuclear membrane
 Supporting Information Fluorescent labeling of the nuclear envelope by localizing GFP on the inner nuclear membrane Toshiyuki Taniyama 1, Natsumi Tsuda 1, and Shinji Sueda 1,2,* 1 Department of Bioscience
Supporting Information Fluorescent labeling of the nuclear envelope by localizing GFP on the inner nuclear membrane Toshiyuki Taniyama 1, Natsumi Tsuda 1, and Shinji Sueda 1,2,* 1 Department of Bioscience
Photoswitchable cyan fluorescent protein for protein tracking
 24 Nature Publishing Group http://www.nature.com/naturebiotechnology Photoswitchable cyan fluorescent protein for protein tracking Dmitriy M Chudakov 1,4, Vladislav V Verkhusha 2,4, Dmitry B Staroverov
24 Nature Publishing Group http://www.nature.com/naturebiotechnology Photoswitchable cyan fluorescent protein for protein tracking Dmitriy M Chudakov 1,4, Vladislav V Verkhusha 2,4, Dmitry B Staroverov
More on fluorescence
 More on fluorescence Last class Fluorescence Absorption emission Jablonski diagrams This class More on fluorescence Common fluorophores Jablonski diagrams to spectra Properties of fluorophores Excitation
More on fluorescence Last class Fluorescence Absorption emission Jablonski diagrams This class More on fluorescence Common fluorophores Jablonski diagrams to spectra Properties of fluorophores Excitation
Challenges to measuring intracellular Ca 2+ Calmodulin: nature s Ca 2+ sensor
 Calcium Signals in Biological Systems Lecture 3 (2/9/0) Measuring intracellular Ca 2+ signals II: Genetically encoded Ca 2+ sensors Henry M. Colecraft, Ph.D. Challenges to measuring intracellular Ca 2+
Calcium Signals in Biological Systems Lecture 3 (2/9/0) Measuring intracellular Ca 2+ signals II: Genetically encoded Ca 2+ sensors Henry M. Colecraft, Ph.D. Challenges to measuring intracellular Ca 2+
Photoactivatable. Fluorescent Proteins. Effective tool to monitor cellular events in real time
 Section B Photoactivatable Fluorescent Proteins Effective tool to monitor cellular events in real time t r a c k i n g c e l l m o v e m e n t m o n i t o r i n g c o m m u n i c a t i o n p a t h w a
Section B Photoactivatable Fluorescent Proteins Effective tool to monitor cellular events in real time t r a c k i n g c e l l m o v e m e n t m o n i t o r i n g c o m m u n i c a t i o n p a t h w a
FLUORESCENCE. Matyas Molnar and Dirk Pacholsky
 FLUORESCENCE Matyas Molnar and Dirk Pacholsky 1 Information This lecture contains images and information from the following internet homepages http://micro.magnet.fsu.edu/primer/index.html http://www.microscopyu.com/
FLUORESCENCE Matyas Molnar and Dirk Pacholsky 1 Information This lecture contains images and information from the following internet homepages http://micro.magnet.fsu.edu/primer/index.html http://www.microscopyu.com/
Fluorescence Microscopy
 Fluorescence Microscopy Dr. Arne Seitz Swiss Institute of Technology (EPFL) Faculty of Life Sciences Head of BIOIMAGING AND OPTICS BIOP arne.seitz@epfl.ch Fluorescence Microscopy Why do we need fluorescence
Fluorescence Microscopy Dr. Arne Seitz Swiss Institute of Technology (EPFL) Faculty of Life Sciences Head of BIOIMAGING AND OPTICS BIOP arne.seitz@epfl.ch Fluorescence Microscopy Why do we need fluorescence
Illumatool ΤΜ Tunable Light System: A Non-Destructive Light Source For Molecular And Cellular Biology Applications. John Fox, Lightools Research.
 Illumatool ΤΜ Tunable Light System: A Non-Destructive Light Source For Molecular And Cellular Biology Applications. John Fox, Lightools Research. Fluorescent dyes and proteins are basic analytical tools
Illumatool ΤΜ Tunable Light System: A Non-Destructive Light Source For Molecular And Cellular Biology Applications. John Fox, Lightools Research. Fluorescent dyes and proteins are basic analytical tools
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Nanogap Engineerable Raman-Active Nanodumbbells for Single-Molecule Detection Dong-Kwon Lim 1,, Ki-Seok Jeon 2,, Hyung Min Kim 2, Jwa-Min Nam 1, *, and Yung Doug Suh 2, * 1 Department
SUPPLEMENTARY INFORMATION Nanogap Engineerable Raman-Active Nanodumbbells for Single-Molecule Detection Dong-Kwon Lim 1,, Ki-Seok Jeon 2,, Hyung Min Kim 2, Jwa-Min Nam 1, *, and Yung Doug Suh 2, * 1 Department
The Green Fluorescent Protein. w.chem.uwec.edu/chem412_s99/ppt/green.ppt
 The Green Fluorescent Protein w.chem.uwec.edu/chem412_s99/ppt/green.ppt www.chem.uwec.edu/chem412_s99/ppt/green.ppt Protein (gene) is from a jellyfish: Aequorea victoria www.chem.uwec.edu/chem412_s99/ppt/green.ppt
The Green Fluorescent Protein w.chem.uwec.edu/chem412_s99/ppt/green.ppt www.chem.uwec.edu/chem412_s99/ppt/green.ppt Protein (gene) is from a jellyfish: Aequorea victoria www.chem.uwec.edu/chem412_s99/ppt/green.ppt
Lab 1: Ensemble Fluorescence Basics
 Lab 1: Ensemble Fluorescence Basics This laboratory module is divided into two sections. The first one is on organic fluorophores, and the second one is on ensemble measurement of FRET (Fluorescence Resonance
Lab 1: Ensemble Fluorescence Basics This laboratory module is divided into two sections. The first one is on organic fluorophores, and the second one is on ensemble measurement of FRET (Fluorescence Resonance
Gene$cally encoded fluorescent proteins
 Gene$cally encoded fluorescent proteins Overview: The fluorescent proteins (FPs) for live cell imaging: Origins and Limitations: 2 Richard N. Day. Ph.D. Indiana University! School of Medicine! Department
Gene$cally encoded fluorescent proteins Overview: The fluorescent proteins (FPs) for live cell imaging: Origins and Limitations: 2 Richard N. Day. Ph.D. Indiana University! School of Medicine! Department
A Cell-Surface-Anchored Ratiometric I-Motif Sensor for. Extracellular ph Detection
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 216 Electronic Supplementary Information (ESI) A Cell-Surface-Anchored Ratiometric I-Motif Sensor for
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 216 Electronic Supplementary Information (ESI) A Cell-Surface-Anchored Ratiometric I-Motif Sensor for
Supplementary Information
 Supplementary Information Fluorescent protein choice The dual fluorescence channel ratiometric promoter characterization plasmid we designed for this study required a pair of fluorescent proteins with
Supplementary Information Fluorescent protein choice The dual fluorescence channel ratiometric promoter characterization plasmid we designed for this study required a pair of fluorescent proteins with
Lab 5: Optical trapping and single molecule fluorescence
 Lab 5: Optical trapping and single molecule fluorescence PI: Matt Lang Lab Instructor: Jorge Ferrer Summary Optical tweezers are an excellent experimental tool to study the biophysics of single molecule
Lab 5: Optical trapping and single molecule fluorescence PI: Matt Lang Lab Instructor: Jorge Ferrer Summary Optical tweezers are an excellent experimental tool to study the biophysics of single molecule
Multiplexed 3D FRET imaging in deep tissue of live embryos Ming Zhao, Xiaoyang Wan, Yu Li, Weibin Zhou and Leilei Peng
 Scientific Reports Multiplexed 3D FRET imaging in deep tissue of live embryos Ming Zhao, Xiaoyang Wan, Yu Li, Weibin Zhou and Leilei Peng 1 Supplementary figures and notes Supplementary Figure S1 Volumetric
Scientific Reports Multiplexed 3D FRET imaging in deep tissue of live embryos Ming Zhao, Xiaoyang Wan, Yu Li, Weibin Zhou and Leilei Peng 1 Supplementary figures and notes Supplementary Figure S1 Volumetric
Graphene oxide-enhanced cytoskeleton imaging and mitosis tracking
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2017 Supplementary information for Graphene oxide-enhanced cytoskeleton imaging and mitosis tracking
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2017 Supplementary information for Graphene oxide-enhanced cytoskeleton imaging and mitosis tracking
Supplementary Materials: A Toolbox of Genetically Encoded FRET-Based Biosensors for Rapid L-Lysine Analysis
 S1 of S9 Supplementary Materials: A Toolbox of Genetically Encoded FRET-Based Biosensors for Rapid L-Lysine Analysis Victoria Steffen, Julia Otten, Susann Engelmann, Andreas Radek, Michael Limberg, Bernd
S1 of S9 Supplementary Materials: A Toolbox of Genetically Encoded FRET-Based Biosensors for Rapid L-Lysine Analysis Victoria Steffen, Julia Otten, Susann Engelmann, Andreas Radek, Michael Limberg, Bernd
PALM/STORM, BALM, STED
 PALM/STORM, BALM, STED Last class 2-photon Intro to PALM/STORM Cyanine dyes/dronpa This class Finish localization super-res BALM STED Localization microscopy Intensity Bins = pixels xx 2 = ss2 + aa 2 /12
PALM/STORM, BALM, STED Last class 2-photon Intro to PALM/STORM Cyanine dyes/dronpa This class Finish localization super-res BALM STED Localization microscopy Intensity Bins = pixels xx 2 = ss2 + aa 2 /12
Characterizing Phenotypes of Bacteria by Staining Method
 Experiment 3 Laboratory to Biology III Diversity of Microorganisms / Wintersemester / page 1 Experiment 3 Characterizing Phenotypes of Bacteria by Staining Method Advisor NN Reading Chapters in BBOM 9
Experiment 3 Laboratory to Biology III Diversity of Microorganisms / Wintersemester / page 1 Experiment 3 Characterizing Phenotypes of Bacteria by Staining Method Advisor NN Reading Chapters in BBOM 9
CENTRAL FACILITY BIOOPTICS MFPL
 CENTRAL FACILITY BIOOPTICS MFPL LIVE IMAGING CONSIDERATIONS Why Live Imaging? You can (somehow) believe what you see no fixation artefacts Follow the order of events in real-time Monitor dynamics: kinetics,
CENTRAL FACILITY BIOOPTICS MFPL LIVE IMAGING CONSIDERATIONS Why Live Imaging? You can (somehow) believe what you see no fixation artefacts Follow the order of events in real-time Monitor dynamics: kinetics,
Characterizing Phenotypes of Bacteria by Staining Method
 Experiment 3 Laboratory to Biology III Diversity of Microorganisms / Wintersemester / page 1 Experiment Characterizing Phenotypes of Bacteria by Staining Method Advisor Reading NN Chapters 3.1, 3.7, 3.8,
Experiment 3 Laboratory to Biology III Diversity of Microorganisms / Wintersemester / page 1 Experiment Characterizing Phenotypes of Bacteria by Staining Method Advisor Reading NN Chapters 3.1, 3.7, 3.8,
Creation of circularly permutated yellow fluorescent proteins using fluorescence screening and a tandem fusion template
 Biotechnology Letters (2006) 28: 471 475 Ó Springer 2006 DOI 10.1007/s10529-006-0007-6 Creation of circularly permutated yellow fluorescent proteins using fluorescence screening and a tandem fusion template
Biotechnology Letters (2006) 28: 471 475 Ó Springer 2006 DOI 10.1007/s10529-006-0007-6 Creation of circularly permutated yellow fluorescent proteins using fluorescence screening and a tandem fusion template
Green Fluorescent Protein-like Proteins in Reef Anthozoa Animals
 CELL STRUCTURE AND FUNCTION 27: 343 347 (2002) 2002 by Japan Society for Cell Biology REVIEW Green Fluorescent Protein-like Proteins in Reef Anthozoa Animals Atsushi Miyawaki Laboratory for Cell Function
CELL STRUCTURE AND FUNCTION 27: 343 347 (2002) 2002 by Japan Society for Cell Biology REVIEW Green Fluorescent Protein-like Proteins in Reef Anthozoa Animals Atsushi Miyawaki Laboratory for Cell Function
How to use the SP5 confocal microscope
 How to use the SP5 confocal microscope Mailfert Sébastien (mailfert@ciml.univ-mrs.fr) Tel : 9126 Imaging Immunity (ImagImm) photonic microscopy facility Centre d Immunologie de Marseille-Luminy 2016 /
How to use the SP5 confocal microscope Mailfert Sébastien (mailfert@ciml.univ-mrs.fr) Tel : 9126 Imaging Immunity (ImagImm) photonic microscopy facility Centre d Immunologie de Marseille-Luminy 2016 /
Partha Roy
 Fluorescence microscopy http://micro.magnet.fsu.edu/primer/index.html Partha Roy 1 Lecture Outline Definition of fluorescence Common fluorescent reagents Construction ti of a fluorescence microscope Optical
Fluorescence microscopy http://micro.magnet.fsu.edu/primer/index.html Partha Roy 1 Lecture Outline Definition of fluorescence Common fluorescent reagents Construction ti of a fluorescence microscope Optical
A Brief History of Light Microscopy And How It Transformed Biomedical Research
 A Brief History of Light Microscopy And How It Transformed Biomedical Research Suewei Lin Office: Interdisciplinary Research Building 8A08 Email: sueweilin@gate.sinica.edu.tw TEL: 2789-9315 Microscope
A Brief History of Light Microscopy And How It Transformed Biomedical Research Suewei Lin Office: Interdisciplinary Research Building 8A08 Email: sueweilin@gate.sinica.edu.tw TEL: 2789-9315 Microscope
Welcome! openmicberkeley.wordpress.com. Open Berkeley
 Welcome! openmicberkeley.wordpress.com Agenda Jen Lee: Introduction to FRET Marla Feller: Using FRET sensors to look at time resolved measurements Becky Lamason: Using FRET to determine if a bacterial
Welcome! openmicberkeley.wordpress.com Agenda Jen Lee: Introduction to FRET Marla Feller: Using FRET sensors to look at time resolved measurements Becky Lamason: Using FRET to determine if a bacterial
Sub-micron scale patterning of fluorescent. silver nanoclusters using low-power laser
 Sub-micron scale patterning of fluorescent silver nanoclusters using low-power laser Puskal Kunwar 1,*, Jukka Hassinen 2, Godofredo Bautista 1, Robin H. A. Ras 2, and Juha Toivonen 1 1 Tampere University
Sub-micron scale patterning of fluorescent silver nanoclusters using low-power laser Puskal Kunwar 1,*, Jukka Hassinen 2, Godofredo Bautista 1, Robin H. A. Ras 2, and Juha Toivonen 1 1 Tampere University
Dino-Lite knowledge & education. Fluorescence Microscopes
 Dino-Lite knowledge & education Fluorescence Microscopes Dino-Lite Fluorescence models Smallest fluorescence microscope in the world Revolution to biomedical and educational applications Flexible Easy
Dino-Lite knowledge & education Fluorescence Microscopes Dino-Lite Fluorescence models Smallest fluorescence microscope in the world Revolution to biomedical and educational applications Flexible Easy
ab CytoPainter ER Staining Kit Red Fluorescence
 ab139482 CytoPainter ER Staining Kit Red Fluorescence Instructions for Use Designed to detect Human endoplasmic reticulum by microscopy. This product is for research use only and is not intended for diagnostic
ab139482 CytoPainter ER Staining Kit Red Fluorescence Instructions for Use Designed to detect Human endoplasmic reticulum by microscopy. This product is for research use only and is not intended for diagnostic
Single-molecule specific mislocalization of red fluorescent proteins in live Escherichia coli
 University of Wollongong Research Online Faculty of Science, Medicine and Health - Papers Faculty of Science, Medicine and Health 2016 Single-molecule specific mislocalization of red fluorescent proteins
University of Wollongong Research Online Faculty of Science, Medicine and Health - Papers Faculty of Science, Medicine and Health 2016 Single-molecule specific mislocalization of red fluorescent proteins
Protein localization in electron micrographs using fluorescence nanoscopy
 Nature Methods Protein localization in electron micrographs using fluorescence nanoscopy Shigeki Watanabe, Annedore Punge, Gunther Hollopeter, Katrin I Willig, Robert John Hobson, M Wayne Davis, Stefan
Nature Methods Protein localization in electron micrographs using fluorescence nanoscopy Shigeki Watanabe, Annedore Punge, Gunther Hollopeter, Katrin I Willig, Robert John Hobson, M Wayne Davis, Stefan
