Guide to Interpreting Genomic Heat Maps Summarizing Integration Site Distributions
|
|
|
- Eleanore Jones
- 6 years ago
- Views:
Transcription
1 Guide to Interpreting Genomic Heat Maps Summarizing Integration Site Distributions General Description We use heat maps to summarize the relationships of proviral distributions to genomic features. These maps were introduced in [1], which presents more background and examples of their uses. The goal of this Guide is to help new users interpret comparisons summarized in the genomic features heat maps. The heat maps summarize information on integration site data sets in columns and different genomic features in rows. Colored tiles indicate the intensity and direction of any departures from the distributions of random controls for each genomic feature in each integration site data set. This report covers heatmaps based on annotated genomic features. Other heat maps described elsewhere summarize integration intensity relative to epigenetic marks and bound proteins, measured by CHIP- seq. Heat maps can be interactive, so that clicking on the map allows statistical comparisons to specific genomic features, to specific integration site data sets or to matched random controls. These steps are explained below. For publication, one particular statistical comparison is chosen and presented as an image file. Abbreviations: MRC: Matched Random Control ROC: Receiver Operating Characteristic TSS: Transcription Start Site Generation of Tile Colors Tile color indicates whether a chosen genomic feature is favored or disfavored by an integrating element, typically a retroviral vector, under the conditions studied. We determine this bias by asking how frequently that feature coincides with an integration site as compared to random sites in the genome using a receiver operating characteristic (ROC) curve area. Generation of Matched Random Control sites: The calculation of ROC area is based on comparisons between true integration sites and computationally selected random sites. Because there is a bias in recovering integration sites when using a restriction enzyme based method, k different matched random control (MRC) sites are selected for each individual integration site. Usually k will be at least 3 MRCs per integration site. Each MRC site lies the same distance from a restriction enzyme recognition sequence as the corresponding integration site but is otherwise randomly distributed in the human genome (described in [1, 2]). For example, if the enzyme MseI is used to recover integration sites, and integration site A lies 120 bases from the nearest MseI site, all k MRCs selected for site A will also lie 120 bases from the nearest MseI site.
2 Calculation of ROC area for genomic feature J : The coincidence of genomic feature J with each integration site and matched random control site is measured (described for each feature below). Each integration site is then compared in a pair- wise fashion to its MRC sites, and a number is assigned indicating the relative rank of the integration site: 1 if the measurement of J is higher at the integration site than at the MRC site, 0 if the measurement of J is lower at the integration site than at the MRC site, 0.5 if the measurement of J is equal for the two sites. All rank values thus calculated for a dataset of integration sites (all k rank values for all integration sites) are averaged to obtain the overall ROC area for the feature measured. An ROC area between 0 and 0.5 indicates the genomic feature occurs less frequently at/near integration sites than at/near random sites in the genome and is therefore disfavored. An ROC area between 0.5 and 1 indicates the genomic feature is enriched at integration sites. An ROC area of exactly 0.5 indicates that integration sites in the dataset are neither enriched nor depleted with respect to the feature of interest. The ROC area is converted to a color tile according to the colorimetric scale at the bottom of the heat map. Positive associations (enrichment compared with random) are shown as increasing shades of red, negative associations (depletion compared with random) as increasing shades of blue, and no difference from random as white. Each tile represents a comparison to the randomly sampled controls for one genomic feature (row) in one experimental dataset (column). Note that we do not present the magnitude of effect in terms of the original units of measurement. We simply ask whether the average integration site has a higher rank for a given type of feature than its k matched random control sites. The color indicates the average quantile of each integration site relative to its random controls. This removes skewing effects contributed by non- normal distributions of the data and also reduces the effect of a few data points with extreme values for a feature. Statistics Statistical tests to determine whether the ROC areas calculated are significantly different from one another or from 0.5 (matched random controls). These are described for each genomic feature below and in [1] (Supplemental Material 3). All the tests rely on the variance- covariance matrix of the relative ranks of the integration sites to construct Wald- type test statistics. For comparisons between integration site sets for specific genomic features, the Wald statistics are calculated and referred to the Chi Square distribution to obtain p values as described in [3] (Supplemental Material 3). *=p<0.05, **=p<0.01, ***=p<0.001
3 The interactive heat maps allow comparison of integration site data sets to each other or genomic features to each other by clicking on appropriate column or row headings. If columns or rows are compared to a control, dashes overlay the control tiles. All tiles can be compared to matched random controls by clicking on the text to the upper right. Columns: Experimental Sets Each column of the heat map is a collection of colored tiles representing the preference of the vector for several genomic features under the condition tested (describe below). The vector/experimental condition is listed at the top of the column. Rows: Genomic Features Gene Boundaries Analysis of integration sites with respect to the transcription start and stop sites of genes as defined by indicated databases. 1. In Gene, Unigene: This tile indicates how frequently integration sites occur within genes in the Unigene database. The color of the tile can be interpreted as follows: Red indicates that the average integration site in the experimental set is more often found within a gene than matched random sites. Blue indicates that the average integration site is less often located within a gene than matched random sites. If the tile is white, the integration sites are no more or less likely to fall within a gene than matched random sites. ROC area calculation: For each integration site- MRC site comparison, the integration site is scored: 1 if the integration site lies in a gene and the MRC site is outside a gene, 0 if the MRC site is in a gene and the integration site is not, and 0.5 if both sites fall within a gene or outside a gene. ROC area is the average score across all comparisons (three per integration site) for all integration sites in the dataset. Statistical Test for difference from Random (ROC area = 0.5): A one degree of freedom Wald test based on the variance mentioned above. Significance: HIV- 1 has a known preference for integration within active transcription units [4], so a set of integration sites of HIV- 1 in wild type human cells will yield a red tile in this row. 2. In Gene, Refseq: Calculations and tile interpretations are identical to those for In Gene, Unigene with the exception that genes are defined by the Refseq database.
4 3. General Width: Indicates the relative width of the gene or intergenic space occupied by integration sites in the experimental set. The color of the tile can be interpreted as follows: Red indicates that the average space (gene width or intergenic width) occupied by integration sites in the set is larger than the average space occupied by matched random sites in the genome. Blue indicates that this average space is smaller for integration sites than for matched random sites. White indicates that this average space is the same for integration sites as it is for a collection of random sites in the genome. ROC area Calculation: For an integration site that falls within a gene (Refseq), the width of the gene is measured in base pairs. Comparisons are made only to those of the three MRC sites that are also in genes. For an integration site outside a gene, the width of the interval between the nearest genes on either side of the site is measured. Comparisons are made to the MRC sites outside genes. For each comparison the integration site is scored: 1 if the gene or intergenic interval within which an integration site lies is larger than that of the compared MRC site 0 if the interval is smaller for the integration site, and 0.5 if the intervals for the integration site and MRC site are matched in size. The ROC area is the average score of all integration sites in the dataset. Statistical Test for difference from Random Controls (ROC area = 0.5): A one Significance: The gene width values provide one measure of gene density, since gene dense regions are comprised of relatively short genes and intergenic regions. This value also correlates negatively with gene expression. Highly expressed genes tend to be shorter while less expressed genes are longer. Similarly, actively transcribed genes tend to cluster such that the intergenic width is shorter between expressed genes, longer between less expressed genes. HIV prefers to integrate in areas of the genome enriched for actively transcribed genes [4], therefore this tile will be blue for HIV- 1 in wild type cells. 4. Gene Width: Displays the relative width of the gene occupied by integration sites in the experimental set. Red indicates that when integration occurs in genes, there is a bias for larger genes. Blue, that integration is favored in shorter genes. The tile is white if integration shows no bias for either short or long genes. Calculations and interpretations are similar to those for General width ; however, for this tile, only those integration site- MRC site pairs in which both sites fall within genes are considered.
5 5. Distance to Start: Indicates whether integration is preferred near transcription start sites (TSS). A red tile means that integration sites in the experimental set are farther from TSSs than matched random sites in the genome and that gene start sites are disfavored for integration. Blue indicates that integration sites lie closer to gene starts than do random sites (gene start sites are favored). If the tile is white, TSSs are neither favored nor disfavored for integration. Note that in contrast to most features in the heat map, gene start sites are favored when this tile is blue, not red, indicating the shorter distance to start sites. ROC area Calculation: For sites that fall within genes, we measure the distance in base pairs to the TSS of that gene. For sites outside of genes, we measure the distance to the nearest transcription start site (according to RefSeq). An integration site is scored: 1 if the distance to the nearest TSS is larger for the integration site than for the compared MRC site 0 if the distance to the nearest TSS is smaller for the integration site 0.5 if the integration site and MRC site are equidistant from the nearest TSS. The ROC area is the average score of all integration sites in the dataset. Statistical Test for difference from Random Controls (ROC area = 0.5): A one Example/Significance: MLV and other gammaretroviruses show a strong preference for integration within gene promoters [5] and therefore, this tile would appear blue for an MLV infection of wild type cells. This makes such viruses relatively dangerous for use as gene therapy vectors, as integration in promoters may alter the control of important host genes [6-10]. By contrast, lentiviruses show little preference for gene starts and this tile appears white for HIV- 1 infection of wild type cells [2, 4]. This measurement is also influenced by the preference of the vector studied for integration within genes as well as width of gene or intergenic space occupied. 6. Distance to Boundary: Indicates whether integration is preferred near the boundaries (transcription start or end) of genes. As with Distance to Start blue indicates that gene boundaries are favored for integration. Red indicates that integration disfavors gene boundaries and white indicates that the virus has no preference for gene boundaries under the conditions studied. ROC area Calculation: Calculations are similar to those for Distance to Start except that for all sites, the distance in base pairs to the nearest gene boundary (start or end) is considered.
6 Statistical Test for difference from Random Controls (ROC area = 0.5): A one Significance: This measure is influenced by the preference of the vector studied for integration within genes, the width of gene or intergenic space occupied and the preference for transcription start sites (described above). 7. < 50 kb from Oncogene: Indicates how frequently integration sites occur within 50 kb of an oncogene (UCSC hg18 goldenpath database for genes, compared to the allonco cancer- related gene list at ). Red indicates that integration sites occur more frequently than random sites near oncogenes. Blue indicates that integration near oncogenes is disfavored. If the tile is white, the integration sites are no more or less likely to fall near an oncogene than matched random sites. ROC area calculation: An integration site is scored: 1 if the integration site falls < 50kb from an oncogene and the compared MRC site is 50 kb from an oncogene, 0 if the MRC site is <50 kb from an oncogene and integration site is not, and 0.5 if both sites lie within or outside 50 kb of an oncogene. ROC area is the average score for all integration sites in the dataset. Statistical Test for difference from Random Controls (ROC area = 0.5): A one Significance: Proximity of integration sites to oncogenes is important to consider when choosing vectors to use in gene therapy applications. DNase Sites Analysis of integration sites with respect to the location of DNase hypersensitive sites in the genome. We ask whether integration sites fall more or less frequently within the indicated distance of a DNase hypersensitive site (<1Mb, <100kb and <10kb) than would be expected for a random distribution (location of DNase sites from UCSC hg18 goldenpath database). Red indicates a preference for integration near a DNase hypersensitive site. Blue indicates that integration is disfavored near DNase hypersensitive sites. If the tile is white, the integration sites are no more or less likely to fall within the indicated distance of a DNase hypersensitive site than matched random sites. ROC area Calculation (eg <1Mb): An integration site is scored:
7 1 if the integration site falls within 1 Mb of a DNase hypersensitive site and the compared MRC site does not, 0 if the MRC site is <1 Mb from a DNase site and the integration site is not, 0.5 if both sites lie within or outside 1 Mb of a DNase hypersensitive site. ROC area is the average score for all integration sites in the dataset. Statistical Test for difference from Random Controls (ROC area = 0.5): A one degree of freedom Wald test based on the variance mentioned above Significance: DNase hypersensitive sites are exposed in more active chromatin and are protected in heterochromatin. Constitutively accessible DNase hypersensitive sites are surrogate markers for open active areas of chromatin. HIV- 1 prefers to integrate within actively transcribed regions, and therefore in regions rich in DNase sites. This preference appears most strongly at larger window sizes and serves as a marker for gene dense regions. HIV disfavors integration near DNAseI sites at very short distances, paralleling the disfavoring of CpG islands and gene start sites ([2, 11]. MLV, in contrast, strongly favors integration very near DNAseI hypersensitive sites [11]. CpG Islands Analysis of integration sites with respect to the location of CpG islands within the genome (UCSC hg18 goldenpath database). 1. CpG Density, 1Mb, 100kb, and 10kb: We ask whether integration sites fall within regions more or less dense in CpG islands than would be expected for a random site in the genome. To do so we consider the density of CpG islands within the indicated genomic intervals surrounding each integration site. A red tile indicates that integration occurs in regions enriched for CpG islands. Blue indicates that integration favors regions poor in CpG islands compared to random. White indicates that integration sites show no bias for or against regions enriched in CpG islands. ROC area Calculation (e.g. CpG Density, 1Mb): We count the number of CpG islands falling in a 1Mb window surrounding each integration and MRC site (ie within 500 kb of a site). An integration site is scored: 1 if more CpG islands lie in the 1Mb surrounding the integration site than in the same interval surrounding the compared MRC site, 0 if fewer CpG islands lie in the genomic interval around the integration site, 0.5 if same number of CpG islands fall in the interval surrounding each site. ROC area is the average score for all integration sites in the dataset. Statistical Test for difference from Random Controls (ROC area = 0.5): A one
8 2. <5kb and <1kb: Indicates whether integration sites fall more or less frequently within the indicated distance of a CpG island than would be expected for a random distribution (location of CpG islands from UCSC hg18 goldenpath database). Red indicates a preference for integration near a CpG island. Blue indicates that integration is disfavored near CpG islands. If the tile is white, the integration sites are no more or less likely to fall near CpG islands than matched random sites. ROC area Calculation (e.g. <5kb): An integration site is scored: 1 if the integration site falls <5kb from a CpG island and the compared MRC site does not, 0 if the MRC site is <5kb from a CpG island and the integration site is not, 0.5 if both sites lie within or outside 1 Mb of a CpG island. ROC area is the average score for all integration sites in the dataset. Statistical Test for difference from Random Controls (ROC area = 0.5): A one Significance: CpG islands are enriched in/near gene promoters, especially those of housekeeping genes, and are otherwise rare within the genome due to methylation and deamination of the cytosine. They are therefore markers of promoters when small genomic windows surrounding CpG islands are considered, and of gene dense regions when larger windows are considered. That is if a site is very close to a CpG island it is likely to be close to a gene promoter, and if a site is in a broad region enriched for CpG islands, it is likely to be in a more gene dense area of the genome. Gene Density Analysis of integration sites with respect to the local density of genes. We consider the indicated genomic interval surrounding each integration site (1Mb, 100kb, or 10kb) and ask whether integration sites fall within regions more or less dense in genes than would be expected for a random site in the genome. A red tile indicates that integration occurs in regions enriched for genes. Blue indicates that integration favors regions poor in genes compared to random. White indicates that integration sites show no bias for or against gene dense regions. ROC area Calculation (e.g. Density 1Mb): We count the number of genes (RefSeq) falling in a 1Mb window surrounding each integration and matched random control site (i.e. within 500 kb of a site). An integration site is scored: 1 if more genes lie in this interval for the integration site than for the compared MRC site, 0 if fewer genes lie in the interval around the integration site, and 0.5 if same number of genes fall in a 1Mb interval surrounding each site. ROC area is the average score for all integration sites in the dataset.
9 Statistical Test for difference from Random Controls (ROC area = 0.5): A one degree of freedom Wald test based on the variance mentioned above Significance/Example: HIV has a known preference for integration in regions of the chromosome that are enriched for genes. This may be because it prefers to integrate within active transcription units [4]. However it may also be independently guided to regions of chromatin enriched for genes. In either case, this preference is hypothesized to afford HIV better access to transcription factors. Expression Intensity Analysis of integration sites with respect to the local density of sets of genes on an Affymetrix Gene Chip. We consider density of total genes as well as density of highly expressed genes. For this analysis we use microarray data measuring relative expression of genes in the cell type used in the study. We note that manipulations to cells in certain experiments may change gene expression profiles, and this must be considered in interpretation of the data. 1. All Genes Density, 1Mb: Calculations and tile interpretations are identical to those for Density, 1Mb in the Gene Density section with the exception that the genes counted are loci identified by expression probe sets on the relevant Affymetrix GeneChip. 2. Top ½ Expression, 1Mb: Calculations and tile interpretations are identical to those for All Genes Density, 1Mb with the exception that only the top ½ most expressed loci are counted. 3. Top 1/16th Expression, 1Mb: Calculations and tile interpretations are identical to those for All Genes Density, 1Mb with the exception that only the top 1/16th most expressed loci are counted. Statistical Test for difference from Random Controls (ROC area = 0.5): A one degree of freedom Wald test based on the variance mentioned above Significance: This analysis allows us to consider whether the preference for total genes is the same as the preference for highly expressed genes. This would be the case if the tile colors are the same for all three measures. If highly expressed genes are preferred more than are genes in general, the Top 1/16th Expression and Top 1/2 expression tiles may be red shifted from the All Genes Density tile. GC Content Analysis of integration sites with respect to the local GC content. A red tile indicates that for the genomic interval considered (see below for discussion of window size) integration sites lie in regions that are GC- rich compared to random sites in the genome. Blue indicates
10 that integration is disfavored in GC rich regions (favored in AT rich regions). White indicates that the vector shows no bias for GC content at the genomic interval considered. ROC area Calculation: GC content is measured within the indicated interval surrounding each integration and matched random site. An integration site is ranked: 1 if the defined region surrounding an integration site is more GC rich than that surrounding the compared MRC site 0 if the integration site lies in a less GC rich region than the MRC site, and 0.5 if the integration and MRC sites are located in equally GC rich regions. ROC area is the average rank for all integration sites in the dataset. Statistical Test for difference from Random Controls (ROC area = 0.5): A one degree of freedom Wald test based on the variance mentioned above Significance: GC content is positively correlated with genes and therefore when considering broad windows surrounding integration sites, a high GC content may indicate a preference for genes/gene dense regions. However, due to local sequence constraints that affect the choice of integration sites of many viruses, small windows surrounding integration sites may show different effects. For example, HIV prefers to integrate in gene dense regions and, predictably, GC content is high if measured in large windows surrounding HIV- 1 integration sites in wild type cells. However if we consider only small windows surrounding integration sites, we find that HIV- 1 prefers relatively AT- rich loci within those large GC rich regions. This is likely to be secondary to HIV- 1 s preference for integration on nucleosomes which position in relatively AT rich DNA [12]. The AT- hooks of LEDGF- p75, a known tether for HIV integrase, may also contribute to this effect [13]. For this reason, we display ROC area tiles for GC content in several genomic intervals. References 1. Berry C, Hannenhalli S, Leipzig J, Bushman FD. (2006) Selection of target sites for mobile DNA integration in the human genome. PLoS Comput Biol 2(11): e Mitchell RS, Beitzel BF, Schroder AR, Shinn P, Chen H, et al. (2004) Retroviral DNA integration: ASLV, HIV, and MLV show distinct target site preferences. PLoS Biol 2(8): E Brady T, Lee YN, Ronen K, Malani N, Berry CC, et al. (2009) Integration target site selection by a resurrected human endogenous retrovirus. Genes Dev 23(5):
11 4. Schroder AR, Shinn P, Chen H, Berry C, Ecker JR, et al. (2002) HIV- 1 integration in the human genome favors active genes and local hotspots. Cell 110(4): Wu X, Li Y, Crise B, Burgess SM. (2003) Transcription start regions in the human genome are favored targets for MLV integration. Science 300(5626): Hacein- Bey- Abina S, Von Kalle C, Schmidt M, McCormack MP, Wulffraat N, et al. (2003) LMO2- associated clonal T cell proliferation in two patients after gene therapy for SCID- X1. Science 302(5644): Hacein- Bey- Abina S, von Kalle C, Schmidt M, Le Deist F, Wulffraat N, et al. (2003) A serious adverse event after successful gene therapy for X- linked severe combined immunodeficiency. N Engl J Med 348(3): Deichmann A, Hacein- Bey- Abina S, Schmidt M, Garrigue A, Brugman MH, et al. (2007) Vector integration is nonrandom and clustered and influences the fate of lymphopoiesis in SCID- X1 gene therapy. J Clin Invest 117(8): Hacein- Bey- Abina S, Garrigue A, Wang GP, Soulier J, Lim A, et al. (2008) Insertional oncogenesis in 4 patients after retrovirus- mediated gene therapy of SCID- X1. J Clin Invest 118(9): Stein S, Ott MG, Schultze- Strasser S, Jauch A, Burwinkel B, et al. (2010) Genomic instability and myelodysplasia with monosomy 7 consequent to EVI1 activation after gene therapy for chronic granulomatous disease. Nat Med 16(2): Lewinski MK, Yamashita M, Emerman M, Ciuffi A, Marshall H, et al. (2006) Retroviral DNA integration: Viral and cellular determinants of target- site selection. PLoS Pathog 2(6): e Wang GP, Ciuffi A, Leipzig J, Berry CC, Bushman FD. (2007) HIV integration site selection: Analysis by massively parallel pyrosequencing reveals association with epigenetic modifications. Genome Res 17(8): Ciuffi A, Llano M, Poeschla E, Hoffmann C, Leipzig J, et al. (2005) A role for LEDGF/p75 in targeting HIV DNA integration. Nat Med 11(12):
Selection of Target Sites for Mobile DNA Integration in the Human Genome
 Selection of Target Sites for Mobile DNA Integration in the Human Genome Charles Berry 1, Sridhar Hannenhalli 2, Jeremy Leipzig 3, Frederic D. Bushman 3* 1 Department of Family and Preventive Medicine,
Selection of Target Sites for Mobile DNA Integration in the Human Genome Charles Berry 1, Sridhar Hannenhalli 2, Jeremy Leipzig 3, Frederic D. Bushman 3* 1 Department of Family and Preventive Medicine,
Received 23 April 2008/Accepted 18 November 2008
 JOURNAL OF VIROLOGY, Feb. 2009, p. 1920 1929 Vol. 83, No. 4 0022-538X/09/$08.00 0 doi:10.1128/jvi.00856-08 Copyright 2009, American Society for Microbiology. All Rights Reserved. Genome Areas with High
JOURNAL OF VIROLOGY, Feb. 2009, p. 1920 1929 Vol. 83, No. 4 0022-538X/09/$08.00 0 doi:10.1128/jvi.00856-08 Copyright 2009, American Society for Microbiology. All Rights Reserved. Genome Areas with High
Journal of Computer Science & Systems Biology - Open Access
 J Comput Sci Syst Biol Volume 2(1): 117-123 (2009) - 117 Vectors and Integration in Gene Therapy: Statistical Considerations Alessandro Ambrosi 1, Clelia Di Serio 1* 1 University Centre of Statistics for
J Comput Sci Syst Biol Volume 2(1): 117-123 (2009) - 117 Vectors and Integration in Gene Therapy: Statistical Considerations Alessandro Ambrosi 1, Clelia Di Serio 1* 1 University Centre of Statistics for
Result Tables The Result Table, which indicates chromosomal positions and annotated gene names, promoter regions and CpG islands, is the best way for
 Result Tables The Result Table, which indicates chromosomal positions and annotated gene names, promoter regions and CpG islands, is the best way for you to discover methylation changes at specific genomic
Result Tables The Result Table, which indicates chromosomal positions and annotated gene names, promoter regions and CpG islands, is the best way for you to discover methylation changes at specific genomic
Epigenetics and DNase-Seq
 Epigenetics and DNase-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by Anthony
Epigenetics and DNase-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by Anthony
PRIMEGENSw3 User Manual
 PRIMEGENSw3 User Manual PRIMEGENSw3 is Web Server version of PRIMEGENS program to automate highthroughput primer and probe design. It provides three separate utilities to select targeted regions of interests
PRIMEGENSw3 User Manual PRIMEGENSw3 is Web Server version of PRIMEGENS program to automate highthroughput primer and probe design. It provides three separate utilities to select targeted regions of interests
NGS Approaches to Epigenomics
 I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
What Can the Epigenome Teach Us About Cellular States and Diseases?
 What Can the Epigenome Teach Us About Cellular States and Diseases? (a computer scientist s view) Luca Pinello Outline Epigenetic: the code over the code What can we learn from epigenomic data? Resources
What Can the Epigenome Teach Us About Cellular States and Diseases? (a computer scientist s view) Luca Pinello Outline Epigenetic: the code over the code What can we learn from epigenomic data? Resources
MODULE TSS1: TRANSCRIPTION START SITES INTRODUCTION (BASIC)
 MODULE TSS1: TRANSCRIPTION START SITES INTRODUCTION (BASIC) Lesson Plan: Title JAMIE SIDERS, MEG LAAKSO & WILSON LEUNG Identifying transcription start sites for Peaked promoters using chromatin landscape,
MODULE TSS1: TRANSCRIPTION START SITES INTRODUCTION (BASIC) Lesson Plan: Title JAMIE SIDERS, MEG LAAKSO & WILSON LEUNG Identifying transcription start sites for Peaked promoters using chromatin landscape,
Go to Bottom Left click WashU Epigenome Browser. Click
 Now you are going to look at the Human Epigenome Browswer. It has a more sophisticated but weirder interface than the UCSC Genome Browser. All the data that you will view as tracks is in reality just files
Now you are going to look at the Human Epigenome Browswer. It has a more sophisticated but weirder interface than the UCSC Genome Browser. All the data that you will view as tracks is in reality just files
Galaxy Platform For NGS Data Analyses
 Galaxy Platform For NGS Data Analyses Weihong Yan wyan@chem.ucla.edu Collaboratory Web Site http://qcb.ucla.edu/collaboratory http://collaboratory.lifesci.ucla.edu Workshop Outline ü Day 1 UCLA galaxy
Galaxy Platform For NGS Data Analyses Weihong Yan wyan@chem.ucla.edu Collaboratory Web Site http://qcb.ucla.edu/collaboratory http://collaboratory.lifesci.ucla.edu Workshop Outline ü Day 1 UCLA galaxy
Reviewers' Comments: Reviewer #1 (Remarks to the Author)
 Reviewers' Comments: Reviewer #1 (Remarks to the Author) In this study, Rosenbluh et al reported direct comparison of two screening approaches: one is genome editing-based method using CRISPR-Cas9 (cutting,
Reviewers' Comments: Reviewer #1 (Remarks to the Author) In this study, Rosenbluh et al reported direct comparison of two screening approaches: one is genome editing-based method using CRISPR-Cas9 (cutting,
Exam 1 from a Past Semester
 Exam from a Past Semester. Provide a brief answer to each of the following questions. a) What do perfect match and mismatch mean in the context of Affymetrix GeneChip technology? Be as specific as possible
Exam from a Past Semester. Provide a brief answer to each of the following questions. a) What do perfect match and mismatch mean in the context of Affymetrix GeneChip technology? Be as specific as possible
Complete Sample to Analysis Solutions for DNA Methylation Discovery using Next Generation Sequencing
 Complete Sample to Analysis Solutions for DNA Methylation Discovery using Next Generation Sequencing SureSelect Human/Mouse Methyl-Seq Kyeong Jeong PhD February 5, 2013 CAG EMEAI DGG/GSD/GFO Agilent Restricted
Complete Sample to Analysis Solutions for DNA Methylation Discovery using Next Generation Sequencing SureSelect Human/Mouse Methyl-Seq Kyeong Jeong PhD February 5, 2013 CAG EMEAI DGG/GSD/GFO Agilent Restricted
Compositional Analysis of Homogeneous Regions in Human Genomic DNA
 Compositional Analysis of Homogeneous Regions in Human Genomic DNA Eric C. Rouchka 1 and David J. States 2 WUCS-2002-2 March 19, 2002 1 Department of Computer Science Washington University Campus Box 1045
Compositional Analysis of Homogeneous Regions in Human Genomic DNA Eric C. Rouchka 1 and David J. States 2 WUCS-2002-2 March 19, 2002 1 Department of Computer Science Washington University Campus Box 1045
Gene expression analysis. Biosciences 741: Genomics Fall, 2013 Week 5. Gene expression analysis
 Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
2/10/17. Contents. Applications of HMMs in Epigenomics
 2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
Coherence analysis discriminates between retroviral integration patterns in CD34 + cells transduced under differing clinical trial conditions
 Citation: Molecular Therapy Methods & Clinical Development (2015) 2, 15015; doi:10.1038/mtm.2015.15 All rights reserved 2329-0501/15 www.nature.com/mtm Article Coherence analysis discriminates between
Citation: Molecular Therapy Methods & Clinical Development (2015) 2, 15015; doi:10.1038/mtm.2015.15 All rights reserved 2329-0501/15 www.nature.com/mtm Article Coherence analysis discriminates between
a previous report, Lee and colleagues showed that Oct4 and Sox2 bind to DXPas34 and Xite
 SUPPLEMENTARY INFORMATION doi:1.138/nature9496 a 1 Relative RNA levels D12 female ES cells 24h Control sirna 24h sirna Oct4 5 b,4 Oct4 Xist (wt) Xist (mut) Tsix 3' (wt) Tsix 5' (wt) Beads Oct4,2 c,2 Xist
SUPPLEMENTARY INFORMATION doi:1.138/nature9496 a 1 Relative RNA levels D12 female ES cells 24h Control sirna 24h sirna Oct4 5 b,4 Oct4 Xist (wt) Xist (mut) Tsix 3' (wt) Tsix 5' (wt) Beads Oct4,2 c,2 Xist
The ENCODE Encyclopedia. & Variant Annotation Using RegulomeDB and HaploReg
 The ENCODE Encyclopedia & Variant Annotation Using RegulomeDB and HaploReg Jill E. Moore Weng Lab University of Massachusetts Medical School October 10, 2015 Where s the Encyclopedia? ENCODE: Encyclopedia
The ENCODE Encyclopedia & Variant Annotation Using RegulomeDB and HaploReg Jill E. Moore Weng Lab University of Massachusetts Medical School October 10, 2015 Where s the Encyclopedia? ENCODE: Encyclopedia
Experimental validation of candidates of tissuespecific and CpG-island-mediated alternative polyadenylation in mouse
 Karin Fleischhanderl; Martina Fondi Experimental validation of candidates of tissuespecific and CpG-island-mediated alternative polyadenylation in mouse 108 - Biotechnologie Abstract --- Keywords: Alternative
Karin Fleischhanderl; Martina Fondi Experimental validation of candidates of tissuespecific and CpG-island-mediated alternative polyadenylation in mouse 108 - Biotechnologie Abstract --- Keywords: Alternative
Simultaneous profiling of transcriptome and DNA methylome from a single cell
 Additional file 1: Supplementary materials Simultaneous profiling of transcriptome and DNA methylome from a single cell Youjin Hu 1, 2, Kevin Huang 1, 3, Qin An 1, Guizhen Du 1, Ganlu Hu 2, Jinfeng Xue
Additional file 1: Supplementary materials Simultaneous profiling of transcriptome and DNA methylome from a single cell Youjin Hu 1, 2, Kevin Huang 1, 3, Qin An 1, Guizhen Du 1, Ganlu Hu 2, Jinfeng Xue
Gene Signal Estimates from Exon Arrays
 Gene Signal Estimates from Exon Arrays I. Introduction: With exon arrays like the GeneChip Human Exon 1.0 ST Array, researchers can examine the transcriptional profile of an entire gene (Figure 1). Being
Gene Signal Estimates from Exon Arrays I. Introduction: With exon arrays like the GeneChip Human Exon 1.0 ST Array, researchers can examine the transcriptional profile of an entire gene (Figure 1). Being
Supplementary Table 1: Oligo designs. A list of ATAC-seq oligos used for PCR.
 Ad1_noMX: Ad2.1_TAAGGCGA Ad2.2_CGTACTAG Ad2.3_AGGCAGAA Ad2.4_TCCTGAGC Ad2.5_GGACTCCT Ad2.6_TAGGCATG Ad2.7_CTCTCTAC Ad2.8_CAGAGAGG Ad2.9_GCTACGCT Ad2.10_CGAGGCTG Ad2.11_AAGAGGCA Ad2.12_GTAGAGGA Ad2.13_GTCGTGAT
Ad1_noMX: Ad2.1_TAAGGCGA Ad2.2_CGTACTAG Ad2.3_AGGCAGAA Ad2.4_TCCTGAGC Ad2.5_GGACTCCT Ad2.6_TAGGCATG Ad2.7_CTCTCTAC Ad2.8_CAGAGAGG Ad2.9_GCTACGCT Ad2.10_CGAGGCTG Ad2.11_AAGAGGCA Ad2.12_GTAGAGGA Ad2.13_GTCGTGAT
Bioinformatics of Transcriptional Regulation
 Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Microarray Ordering Guide
 Catalog Microarray Ordering Guide Gene Expression Microarrays G4112F 014850 4 5 Whole Human Genome Microarray Kit, 4x44K 41,000+ unique genes and transcripts represented, all with public domain annotations.
Catalog Microarray Ordering Guide Gene Expression Microarrays G4112F 014850 4 5 Whole Human Genome Microarray Kit, 4x44K 41,000+ unique genes and transcripts represented, all with public domain annotations.
Lecture 9. Eukaryotic gene regulation: DNA METHYLATION
 Lecture 9 Eukaryotic gene regulation: DNA METHYLATION Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation sequences Transcriptional
Lecture 9 Eukaryotic gene regulation: DNA METHYLATION Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation sequences Transcriptional
Lab 1: A review of linear models
 Lab 1: A review of linear models The purpose of this lab is to help you review basic statistical methods in linear models and understanding the implementation of these methods in R. In general, we need
Lab 1: A review of linear models The purpose of this lab is to help you review basic statistical methods in linear models and understanding the implementation of these methods in R. In general, we need
Figure S1. nuclear extracts. HeLa cell nuclear extract. Input IgG IP:ORC2 ORC2 ORC2. MCM4 origin. ORC2 occupancy
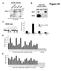 A nuclear extracts B HeLa cell nuclear extract Figure S1 ORC2 (in kda) 21 132 7 ORC2 Input IgG IP:ORC2 32 ORC C D PRKDC ORC2 occupancy Directed against ORC2 C-terminus (sc-272) MCM origin 2 2 1-1 -1kb
A nuclear extracts B HeLa cell nuclear extract Figure S1 ORC2 (in kda) 21 132 7 ORC2 Input IgG IP:ORC2 32 ORC C D PRKDC ORC2 occupancy Directed against ORC2 C-terminus (sc-272) MCM origin 2 2 1-1 -1kb
Applications of HMMs in Epigenomics
 I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
ChIP-seq/Functional Genomics/Epigenomics. CBSU/3CPG/CVG Next-Gen Sequencing Workshop. Josh Waterfall. March 31, 2010
 ChIP-seq/Functional Genomics/Epigenomics CBSU/3CPG/CVG Next-Gen Sequencing Workshop Josh Waterfall March 31, 2010 Outline Introduction to ChIP-seq Control data sets Peak/enriched region identification
ChIP-seq/Functional Genomics/Epigenomics CBSU/3CPG/CVG Next-Gen Sequencing Workshop Josh Waterfall March 31, 2010 Outline Introduction to ChIP-seq Control data sets Peak/enriched region identification
Nature Genetics: doi: /ng Supplementary Figure 1. H3K27ac HiChIP enriches enhancer promoter-associated chromatin contacts.
 Supplementary Figure 1 H3K27ac HiChIP enriches enhancer promoter-associated chromatin contacts. (a) Schematic of chromatin contacts captured in H3K27ac HiChIP. (b) Loop call overlap for cohesin HiChIP
Supplementary Figure 1 H3K27ac HiChIP enriches enhancer promoter-associated chromatin contacts. (a) Schematic of chromatin contacts captured in H3K27ac HiChIP. (b) Loop call overlap for cohesin HiChIP
Introduction to the UCSC genome browser
 Introduction to the UCSC genome browser Dominik Beck NHMRC Peter Doherty and CINSW ECR Fellow, Senior Lecturer Lowy Cancer Research Centre, UNSW and Centre for Health Technology, UTS SYDNEY NSW AUSTRALIA
Introduction to the UCSC genome browser Dominik Beck NHMRC Peter Doherty and CINSW ECR Fellow, Senior Lecturer Lowy Cancer Research Centre, UNSW and Centre for Health Technology, UTS SYDNEY NSW AUSTRALIA
Title: Genome-Wide Predictions of Transcription Factor Binding Events using Multi- Dimensional Genomic and Epigenomic Features Background
 Title: Genome-Wide Predictions of Transcription Factor Binding Events using Multi- Dimensional Genomic and Epigenomic Features Team members: David Moskowitz and Emily Tsang Background Transcription factors
Title: Genome-Wide Predictions of Transcription Factor Binding Events using Multi- Dimensional Genomic and Epigenomic Features Team members: David Moskowitz and Emily Tsang Background Transcription factors
Unit 8: Genomics Guided Reading Questions (150 pts total)
 Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 18 The Genetics of Viruses and Bacteria Unit 8: Genomics Guided
Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 18 The Genetics of Viruses and Bacteria Unit 8: Genomics Guided
Nature Methods: doi: /nmeth.4396
 Supplementary Figure 1 Comparison of technical replicate consistency between and across the standard ATAC-seq method, DNase-seq, and Omni-ATAC. (a) Heatmap-based representation of ATAC-seq quality control
Supplementary Figure 1 Comparison of technical replicate consistency between and across the standard ATAC-seq method, DNase-seq, and Omni-ATAC. (a) Heatmap-based representation of ATAC-seq quality control
Supplementary Figure 1 Strategy for parallel detection of DHSs and adjacent nucleosomes
 Supplementary Figure 1 Strategy for parallel detection of DHSs and adjacent nucleosomes DNase I cleavage DNase I DNase I digestion Sucrose gradient enrichment Small Large F1 F2...... F9 F1 F1 F2 F3 F4
Supplementary Figure 1 Strategy for parallel detection of DHSs and adjacent nucleosomes DNase I cleavage DNase I DNase I digestion Sucrose gradient enrichment Small Large F1 F2...... F9 F1 F1 F2 F3 F4
Suppl. Table S1. Characteristics of DHS regions analyzed by bisulfite sequencing. No. CpGs analyzed in the amplicon. Genomic location specificity
 Suppl. Table S1. Characteristics of DHS regions analyzed by bisulfite sequencing. DHS/GRE Genomic location Tissue specificity DHS type CpG density (per 100 bp) No. CpGs analyzed in the amplicon CpG within
Suppl. Table S1. Characteristics of DHS regions analyzed by bisulfite sequencing. DHS/GRE Genomic location Tissue specificity DHS type CpG density (per 100 bp) No. CpGs analyzed in the amplicon CpG within
Modern Epigenomics. Histone Code
 Modern Epigenomics Histone Code Ting Wang Department of Genetics Center for Genome Sciences and Systems Biology Washington University Dragon Star 2012 Changchun, China July 2, 2012 DNA methylation + Histone
Modern Epigenomics Histone Code Ting Wang Department of Genetics Center for Genome Sciences and Systems Biology Washington University Dragon Star 2012 Changchun, China July 2, 2012 DNA methylation + Histone
INTRODUCTION. Research Article Retroviral vector integrations are cell-specific
 Integration profile of retroviral vector in gene therapy treated patients is cell-specific according to gene expression and chromatin conformation of target cell Luca Biasco 1,2, Alessandro Ambrosi 3,
Integration profile of retroviral vector in gene therapy treated patients is cell-specific according to gene expression and chromatin conformation of target cell Luca Biasco 1,2, Alessandro Ambrosi 3,
Nature Biotechnology: doi: /nbt Supplementary Figure 1. sndrop-seq overview.
 Supplementary Figure 1 sndrop-seq overview. A. sndrop-seq method showing modifications needed to process nuclei, including bovine serum albumin (BSA) coating and droplet heating to ensure complete nuclear
Supplementary Figure 1 sndrop-seq overview. A. sndrop-seq method showing modifications needed to process nuclei, including bovine serum albumin (BSA) coating and droplet heating to ensure complete nuclear
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Targeted RNA sequencing reveals the deep complexity of the human transcriptome.
 Targeted RNA sequencing reveals the deep complexity of the human transcriptome. Tim R. Mercer 1, Daniel J. Gerhardt 2, Marcel E. Dinger 1, Joanna Crawford 1, Cole Trapnell 3, Jeffrey A. Jeddeloh 2,4, John
Targeted RNA sequencing reveals the deep complexity of the human transcriptome. Tim R. Mercer 1, Daniel J. Gerhardt 2, Marcel E. Dinger 1, Joanna Crawford 1, Cole Trapnell 3, Jeffrey A. Jeddeloh 2,4, John
2/19/13. Contents. Applications of HMMs in Epigenomics
 2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
Introduction to Microarray Analysis
 Introduction to Microarray Analysis Methods Course: Gene Expression Data Analysis -Day One Rainer Spang Microarrays Highly parallel measurement devices for gene expression levels 1. How does the microarray
Introduction to Microarray Analysis Methods Course: Gene Expression Data Analysis -Day One Rainer Spang Microarrays Highly parallel measurement devices for gene expression levels 1. How does the microarray
GATCGTGCACGATCTCGGCAATTCGGGATGCCGGCTCGTCACCGGTCGCT
 Problem. (pts) A. (5pts) Your colleague professor Eugene Mathew Lateed generated a genome-wide DNA methylation map for normal colon cells using MRE-seq and MeDIP-seq. In an intergenic region, he found
Problem. (pts) A. (5pts) Your colleague professor Eugene Mathew Lateed generated a genome-wide DNA methylation map for normal colon cells using MRE-seq and MeDIP-seq. In an intergenic region, he found
From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow
 From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow Technical Overview Import VCF Introduction Next-generation sequencing (NGS) studies have created unanticipated challenges with
From Variants to Pathways: Agilent GeneSpring GX s Variant Analysis Workflow Technical Overview Import VCF Introduction Next-generation sequencing (NGS) studies have created unanticipated challenges with
nature methods A paired-end sequencing strategy to map the complex landscape of transcription initiation
 nature methods A paired-end sequencing strategy to map the complex landscape of transcription initiation Ting Ni, David L Corcoran, Elizabeth A Rach, Shen Song, Eric P Spana, Yuan Gao, Uwe Ohler & Jun
nature methods A paired-end sequencing strategy to map the complex landscape of transcription initiation Ting Ni, David L Corcoran, Elizabeth A Rach, Shen Song, Eric P Spana, Yuan Gao, Uwe Ohler & Jun
Supplement to: The Genomic Sequence of the Chinese Hamster Ovary (CHO)-K1 cell line
 Supplement to: The Genomic Sequence of the Chinese Hamster Ovary (CHO)-K1 cell line Table of Contents SUPPLEMENTARY TEXT:... 2 FILTERING OF RAW READS PRIOR TO ASSEMBLY:... 2 COMPARATIVE ANALYSIS... 2 IMMUNOGENIC
Supplement to: The Genomic Sequence of the Chinese Hamster Ovary (CHO)-K1 cell line Table of Contents SUPPLEMENTARY TEXT:... 2 FILTERING OF RAW READS PRIOR TO ASSEMBLY:... 2 COMPARATIVE ANALYSIS... 2 IMMUNOGENIC
NimbleGen Arrays and LightCycler 480 System: A Complete Workflow for DNA Methylation Biomarker Discovery and Validation.
 Cancer Research Application Note No. 9 NimbleGen Arrays and LightCycler 480 System: A Complete Workflow for DNA Methylation Biomarker Discovery and Validation Tomasz Kazimierz Wojdacz, PhD Institute of
Cancer Research Application Note No. 9 NimbleGen Arrays and LightCycler 480 System: A Complete Workflow for DNA Methylation Biomarker Discovery and Validation Tomasz Kazimierz Wojdacz, PhD Institute of
Supporting Information
 Supporting Information Ho et al. 1.173/pnas.81288816 SI Methods Sequences of shrna hairpins: Brg shrna #1: ccggcggctcaagaaggaagttgaactcgagttcaacttccttcttgacgnttttg (TRCN71383; Open Biosystems). Brg shrna
Supporting Information Ho et al. 1.173/pnas.81288816 SI Methods Sequences of shrna hairpins: Brg shrna #1: ccggcggctcaagaaggaagttgaactcgagttcaacttccttcttgacgnttttg (TRCN71383; Open Biosystems). Brg shrna
Novel principles of gamma-retroviral insertional transcription activation in murine leukemia virus-induced end-stage tumors
 Sokol et al. Retrovirology 2014, 11:36 RESEARCH Open Access Novel principles of gamma-retroviral insertional transcription activation in murine leukemia virus-induced end-stage tumors Martin Sokol 1, Matthias
Sokol et al. Retrovirology 2014, 11:36 RESEARCH Open Access Novel principles of gamma-retroviral insertional transcription activation in murine leukemia virus-induced end-stage tumors Martin Sokol 1, Matthias
The ChIP-Seq project. Giovanna Ambrosini, Philipp Bucher. April 19, 2010 Lausanne. EPFL-SV Bucher Group
 The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
Introduction to human genomics and genome informatics
 Introduction to human genomics and genome informatics Session 1 Prince of Wales Clinical School Dr Jason Wong ARC Future Fellow Head, Bioinformatics & Integrative Genomics Adult Cancer Program, Lowy Cancer
Introduction to human genomics and genome informatics Session 1 Prince of Wales Clinical School Dr Jason Wong ARC Future Fellow Head, Bioinformatics & Integrative Genomics Adult Cancer Program, Lowy Cancer
Single-cell sequencing
 Single-cell sequencing Harri Lähdesmäki Department of Computer Science Aalto University December 5, 2017 Contents Background & Motivation Single cell sequencing technologies Single cell sequencing data
Single-cell sequencing Harri Lähdesmäki Department of Computer Science Aalto University December 5, 2017 Contents Background & Motivation Single cell sequencing technologies Single cell sequencing data
Mate-pair library data improves genome assembly
 De Novo Sequencing on the Ion Torrent PGM APPLICATION NOTE Mate-pair library data improves genome assembly Highly accurate PGM data allows for de Novo Sequencing and Assembly For a draft assembly, generate
De Novo Sequencing on the Ion Torrent PGM APPLICATION NOTE Mate-pair library data improves genome assembly Highly accurate PGM data allows for de Novo Sequencing and Assembly For a draft assembly, generate
797Trex megadomain(s) Per403 megadomain(s) chr1: chr1: chr12: chr12:
 Table S1. Megadomains that overlap in all BRD4-NUT NMC cell lines and tissue. chromosome union region start union region end 797 megadomain(s) chr1 17197632 17247632 chr1:1721332-1724232 chr12 851264 8598264
Table S1. Megadomains that overlap in all BRD4-NUT NMC cell lines and tissue. chromosome union region start union region end 797 megadomain(s) chr1 17197632 17247632 chr1:1721332-1724232 chr12 851264 8598264
Value Correct Answer Feedback. Student Response. A. Dicer enzyme. complex. C. the Dicer-RISC complex D. none of the above
 1 RNA mediated interference is a post-transcriptional gene silencing mechanism Which component of the RNAi pathway have been implicated in cleavage of the target mrna? A Dicer enzyme B the RISC-siRNA complex
1 RNA mediated interference is a post-transcriptional gene silencing mechanism Which component of the RNAi pathway have been implicated in cleavage of the target mrna? A Dicer enzyme B the RISC-siRNA complex
Analysis of a Tiling Regulation Study in Partek Genomics Suite 6.6
 Analysis of a Tiling Regulation Study in Partek Genomics Suite 6.6 The example data set used in this tutorial consists of 6 technical replicates from the same human cell line, 3 are SP1 treated, and 3
Analysis of a Tiling Regulation Study in Partek Genomics Suite 6.6 The example data set used in this tutorial consists of 6 technical replicates from the same human cell line, 3 are SP1 treated, and 3
Microarray Data Analysis in GeneSpring GX 11. Month ##, 200X
 Microarray Data Analysis in GeneSpring GX 11 Month ##, 200X Agenda Genome Browser GO GSEA Pathway Analysis Network building Find significant pathways Extract relations via NLP Data Visualization Options
Microarray Data Analysis in GeneSpring GX 11 Month ##, 200X Agenda Genome Browser GO GSEA Pathway Analysis Network building Find significant pathways Extract relations via NLP Data Visualization Options
Computational gene finding
 Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
Computational gene finding Devika Subramanian Comp 470 Outline (3 lectures) Lec 1 Lec 2 Lec 3 The biological context Markov models and Hidden Markov models Ab-initio methods for gene finding Comparative
ChIP-seq and RNA-seq. Farhat Habib
 ChIP-seq and RNA-seq Farhat Habib fhabib@iiserpune.ac.in Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions
ChIP-seq and RNA-seq Farhat Habib fhabib@iiserpune.ac.in Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions
Supplementary Figures
 Supplementary Figures A B Supplementary Figure 1. Examples of discrepancies in predicted and validated breakpoint coordinates. A) Most frequently, predicted breakpoints were shifted relative to those derived
Supplementary Figures A B Supplementary Figure 1. Examples of discrepancies in predicted and validated breakpoint coordinates. A) Most frequently, predicted breakpoints were shifted relative to those derived
Chromatin. Structure and modification of chromatin. Chromatin domains
 Chromatin Structure and modification of chromatin Chromatin domains 2 DNA consensus 5 3 3 DNA DNA 4 RNA 5 ss RNA forms secondary structures with ds hairpins ds forms 6 of nucleic acids Form coiling bp/turn
Chromatin Structure and modification of chromatin Chromatin domains 2 DNA consensus 5 3 3 DNA DNA 4 RNA 5 ss RNA forms secondary structures with ds hairpins ds forms 6 of nucleic acids Form coiling bp/turn
A Brief History. Bootstrapping. Bagging. Boosting (Schapire 1989) Adaboost (Schapire 1995)
 A Brief History Bootstrapping Bagging Boosting (Schapire 1989) Adaboost (Schapire 1995) What s So Good About Adaboost Improves classification accuracy Can be used with many different classifiers Commonly
A Brief History Bootstrapping Bagging Boosting (Schapire 1989) Adaboost (Schapire 1995) What s So Good About Adaboost Improves classification accuracy Can be used with many different classifiers Commonly
Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate
 Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Conserved elements with potential to form polymorphic G-quadruplex structures in the first intron of human genes
 Published online 10 January 2008 Nucleic Acids Research, 2008, Vol. 36, No. 4 1321 1333 doi:10.1093/nar/gkm1138 Conserved elements with potential to form polymorphic G-quadruplex structures in the first
Published online 10 January 2008 Nucleic Acids Research, 2008, Vol. 36, No. 4 1321 1333 doi:10.1093/nar/gkm1138 Conserved elements with potential to form polymorphic G-quadruplex structures in the first
Gene Therapy for Fanconi Anaemia Hope or hype? Dr Phil Ancliff Great Ormond Street Hospital / UCL Institute of Child Health Twycross Zoo October 2017
 Gene Therapy for Fanconi Anaemia Hope or hype? Dr Phil Ancliff Great Ormond Street Hospital / UCL Institute of Child Health Twycross Zoo October 2017 Gene therapy in Seattle Grace Miranda Bailey had the
Gene Therapy for Fanconi Anaemia Hope or hype? Dr Phil Ancliff Great Ormond Street Hospital / UCL Institute of Child Health Twycross Zoo October 2017 Gene therapy in Seattle Grace Miranda Bailey had the
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Chapter 2 The Structure of Genes and Genomes. Electron micrograph of a metaphase chromosome
 Chapter 2 The Structure of Genes and Genomes Electron micrograph of a metaphase chromosome Genetic information is stored in double stranded DNA DNA structure, building blocks: (A) Bases DNA structure,
Chapter 2 The Structure of Genes and Genomes Electron micrograph of a metaphase chromosome Genetic information is stored in double stranded DNA DNA structure, building blocks: (A) Bases DNA structure,
Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C
 CORRECTION NOTICE Nat. Genet. 47, 598 606 (2015) Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C Borbala Mifsud, Filipe Tavares-Cadete, Alice N Young, Robert Sugar,
CORRECTION NOTICE Nat. Genet. 47, 598 606 (2015) Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C Borbala Mifsud, Filipe Tavares-Cadete, Alice N Young, Robert Sugar,
Statistical Methods for Network Analysis of Biological Data
 The Protein Interaction Workshop, 8 12 June 2015, IMS Statistical Methods for Network Analysis of Biological Data Minghua Deng, dengmh@pku.edu.cn School of Mathematical Sciences Center for Quantitative
The Protein Interaction Workshop, 8 12 June 2015, IMS Statistical Methods for Network Analysis of Biological Data Minghua Deng, dengmh@pku.edu.cn School of Mathematical Sciences Center for Quantitative
Lecture 8: Predicting and analyzing metagenomic composition from 16S survey data
 Lecture 8: Predicting and analyzing metagenomic composition from 16S survey data What can we tell about the taxonomic and functional stability of microbiota? Why? Nature. 2012; 486(7402): 207 214. doi:10.1038/nature11234
Lecture 8: Predicting and analyzing metagenomic composition from 16S survey data What can we tell about the taxonomic and functional stability of microbiota? Why? Nature. 2012; 486(7402): 207 214. doi:10.1038/nature11234
Zhang et al., RepID facilitates replication Initiation. Supplemental Information:
 Supplemental Information: a b 1 Supplementary Figure 1 (a) DNA sequence of all the oligonucleotides used in this study. Only one strand is shown. The unshaded nucleotide sequences show changes from the
Supplemental Information: a b 1 Supplementary Figure 1 (a) DNA sequence of all the oligonucleotides used in this study. Only one strand is shown. The unshaded nucleotide sequences show changes from the
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017
 Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Comparative eqtl analyses within and between seven tissue types suggest mechanisms underlying cell type specificity of eqtls
 Comparative eqtl analyses within and between seven tissue types suggest mechanisms underlying cell type specificity of eqtls, Duke University Christopher D Brown, University of Pennsylvania November 9th,
Comparative eqtl analyses within and between seven tissue types suggest mechanisms underlying cell type specificity of eqtls, Duke University Christopher D Brown, University of Pennsylvania November 9th,
V21: Analysis of DNA methylation data
 V21: Analysis of DNA methylation data Epigenetics refers to alternate phenotypic states that are not based on differences in genotype, and are potentially reversible, but are generally stably maintained
V21: Analysis of DNA methylation data Epigenetics refers to alternate phenotypic states that are not based on differences in genotype, and are potentially reversible, but are generally stably maintained
Supplemental Figure 1.
 Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Supplemental Data. Charron et al. Dynamic landscapes of four histone modifications during de-etiolation in Arabidopsis. Plant Cell (2009). 10.1105/tpc.109.066845 Supplemental Figure 1. Immunodetection
Complete draft sequence 2001
 Genomes: What we know and what we don t know Complete draft sequence 2001 November11, 2009 Dr. Stefan Maas, BioS Lehigh U. What we know Raw genome data The range of genome sizes in the animal & plant kingdoms
Genomes: What we know and what we don t know Complete draft sequence 2001 November11, 2009 Dr. Stefan Maas, BioS Lehigh U. What we know Raw genome data The range of genome sizes in the animal & plant kingdoms
Gene Expression Microarrays. For microarrays, purity of the RNA was further assessed by
 Supplemental Methods Gene Expression Microarrays. For microarrays, purity of the RNA was further assessed by an Agilent 2100 Bioanalyzer. 500 ng of RNA was reverse transcribed into crna and biotin-utp
Supplemental Methods Gene Expression Microarrays. For microarrays, purity of the RNA was further assessed by an Agilent 2100 Bioanalyzer. 500 ng of RNA was reverse transcribed into crna and biotin-utp
Package COCOA. November 6, 2018
 Version 1.1.0 Date 2018-10-29 Title Coordinate Covariation Analysis Package COCOA November 6, 2018 COCOA is a method for understanding variation among samples and can be used with data that includes genomic
Version 1.1.0 Date 2018-10-29 Title Coordinate Covariation Analysis Package COCOA November 6, 2018 COCOA is a method for understanding variation among samples and can be used with data that includes genomic
Deep Sequencing technologies
 Deep Sequencing technologies Gabriela Salinas 30 October 2017 Transcriptome and Genome Analysis Laboratory http://www.uni-bc.gwdg.de/index.php?id=709 Microarray and Deep-Sequencing Core Facility University
Deep Sequencing technologies Gabriela Salinas 30 October 2017 Transcriptome and Genome Analysis Laboratory http://www.uni-bc.gwdg.de/index.php?id=709 Microarray and Deep-Sequencing Core Facility University
MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE?
 MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE? Lesson Plan: Title Introduction to the Genome Browser: what is a gene? JOYCE STAMM Objectives Demonstrate basic skills in using the UCSC Genome
MODULE 1: INTRODUCTION TO THE GENOME BROWSER: WHAT IS A GENE? Lesson Plan: Title Introduction to the Genome Browser: what is a gene? JOYCE STAMM Objectives Demonstrate basic skills in using the UCSC Genome
Final exam: Introduction to Bioinformatics and Genomics DUE: Friday June 29 th at 4:00 pm
 Final exam: Introduction to Bioinformatics and Genomics DUE: Friday June 29 th at 4:00 pm Exam description: The purpose of this exam is for you to demonstrate your ability to use the different biomolecular
Final exam: Introduction to Bioinformatics and Genomics DUE: Friday June 29 th at 4:00 pm Exam description: The purpose of this exam is for you to demonstrate your ability to use the different biomolecular
Permutation Clustering of the DNA Sequence Facilitates Understanding of the Nonlinearly Organized Genome
 RESEARCH PROPOSAL Permutation Clustering of the DNA Sequence Facilitates Understanding of the Nonlinearly Organized Genome Qiao JIN School of Medicine, Tsinghua University Advisor: Prof. Xuegong ZHANG
RESEARCH PROPOSAL Permutation Clustering of the DNA Sequence Facilitates Understanding of the Nonlinearly Organized Genome Qiao JIN School of Medicine, Tsinghua University Advisor: Prof. Xuegong ZHANG
University of Groningen
 University of Groningen Accounting for immunoprecipitation efficiencies in the statistical analysis of ChIP-seq data Bao, Yanchun; Vinciotti, Veronica; Wit, Ernst; 't Hoen, Peter A C Published in: Bmc
University of Groningen Accounting for immunoprecipitation efficiencies in the statistical analysis of ChIP-seq data Bao, Yanchun; Vinciotti, Veronica; Wit, Ernst; 't Hoen, Peter A C Published in: Bmc
Runs of Homozygosity Analysis Tutorial
 Runs of Homozygosity Analysis Tutorial Release 8.7.0 Golden Helix, Inc. March 22, 2017 Contents 1. Overview of the Project 2 2. Identify Runs of Homozygosity 6 Illustrative Example...............................................
Runs of Homozygosity Analysis Tutorial Release 8.7.0 Golden Helix, Inc. March 22, 2017 Contents 1. Overview of the Project 2 2. Identify Runs of Homozygosity 6 Illustrative Example...............................................
ChroMoS Guide (version 1.2)
 ChroMoS Guide (version 1.2) Background Genome-wide association studies (GWAS) reveal increasing number of disease-associated SNPs. Since majority of these SNPs are located in intergenic and intronic regions
ChroMoS Guide (version 1.2) Background Genome-wide association studies (GWAS) reveal increasing number of disease-associated SNPs. Since majority of these SNPs are located in intergenic and intronic regions
Analysis of ChIP-seq data with R / Bioconductor
 Analysis of ChIP-seq data with R / Bioconductor Martin Morgan Bioconductor / Fred Hutchinson Cancer Research Center Seattle, WA, USA 8-10 June 2009 ChIP-seq Chromatin immunopreciptation to enrich sample
Analysis of ChIP-seq data with R / Bioconductor Martin Morgan Bioconductor / Fred Hutchinson Cancer Research Center Seattle, WA, USA 8-10 June 2009 ChIP-seq Chromatin immunopreciptation to enrich sample
Understanding transcriptional regulation by integrative analysis of transcription factor binding data
 Understanding transcriptional regulation by integrative analysis of transcription factor binding data Cheng et al. 2012 Shu Yang Feb. 21, 2013 1 / 26 Introduction 2 / 26 DNA-binding Proteins sequence-specific
Understanding transcriptional regulation by integrative analysis of transcription factor binding data Cheng et al. 2012 Shu Yang Feb. 21, 2013 1 / 26 Introduction 2 / 26 DNA-binding Proteins sequence-specific
Poly(dA:dT)-rich DNAs are highly flexible in the context of DNA looping: Supporting Information
 Poly(dA:dT)-rich DNAs are highly flexible in the context of DNA looping: Supporting Information Stephanie Johnson, Yi-Ju Chen, and Rob Phillips List of Figures S1 No-promoter looping sequences used in
Poly(dA:dT)-rich DNAs are highly flexible in the context of DNA looping: Supporting Information Stephanie Johnson, Yi-Ju Chen, and Rob Phillips List of Figures S1 No-promoter looping sequences used in
Genomics and Gene Recognition Genes and Blue Genes
 Genomics and Gene Recognition Genes and Blue Genes November 3, 2004 Eukaryotic Gene Structure eukaryotic genomes are considerably more complex than those of prokaryotes eukaryotic cells have organelles
Genomics and Gene Recognition Genes and Blue Genes November 3, 2004 Eukaryotic Gene Structure eukaryotic genomes are considerably more complex than those of prokaryotes eukaryotic cells have organelles
Satellite Education Workshop (SW4): Epigenomics: Design, Implementation and Analysis for RNA-seq and Methyl-seq Experiments
 Satellite Education Workshop (SW4): Epigenomics: Design, Implementation and Analysis for RNA-seq and Methyl-seq Experiments Saturday March 17, 2012 Orlando, Florida Workshop Description: This full day
Satellite Education Workshop (SW4): Epigenomics: Design, Implementation and Analysis for RNA-seq and Methyl-seq Experiments Saturday March 17, 2012 Orlando, Florida Workshop Description: This full day
The Human Genome Project has always been something of a misnomer, implying the existence of a single human genome
 The Human Genome Project has always been something of a misnomer, implying the existence of a single human genome Of course, every person on the planet with the exception of identical twins has a unique
The Human Genome Project has always been something of a misnomer, implying the existence of a single human genome Of course, every person on the planet with the exception of identical twins has a unique
Gene Expression. Chapters 11 & 12: Gene Conrtrol and DNA Technology. Cloning. Honors Biology Fig
 Chapters & : Conrtrol and Technology Honors Biology 0 Cloning Produced by asexual reproduction and so it is genetically identical to the parent st large cloned mammal: Dolly the sheep Animals that are
Chapters & : Conrtrol and Technology Honors Biology 0 Cloning Produced by asexual reproduction and so it is genetically identical to the parent st large cloned mammal: Dolly the sheep Animals that are
SUPPLEMENTARY INFORMATION
 Supplementary Discussion Interestingly, a recent study demonstrated that knockdown of Tet1 alone would lead to dysregulation of differentiation genes, but only minor defects in ES maintenance and no change
Supplementary Discussion Interestingly, a recent study demonstrated that knockdown of Tet1 alone would lead to dysregulation of differentiation genes, but only minor defects in ES maintenance and no change
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Current SCID Gene Transfer Experience
 Current Perspectives on Gene Transfer for X-SCID Recombinant DNA Advisory Committee Safety Symposium March 15, 2005 Discussion Questions Current SCID Gene Transfer Experience 1. What data is available
Current Perspectives on Gene Transfer for X-SCID Recombinant DNA Advisory Committee Safety Symposium March 15, 2005 Discussion Questions Current SCID Gene Transfer Experience 1. What data is available
Technical Note. GeneChip Mouse Expression Set 430. GeneChip Mouse Expression Set 430 Design
 GeneChip Mouse Expression Set 43 AFFYMETRIX PRODUCT FAMILY > RNA ARRAYS AND REAGENTS > Technical Note Array Design and Performance of the GeneChip Mouse Expression Set 43 The mouse is a popular model system
GeneChip Mouse Expression Set 43 AFFYMETRIX PRODUCT FAMILY > RNA ARRAYS AND REAGENTS > Technical Note Array Design and Performance of the GeneChip Mouse Expression Set 43 The mouse is a popular model system
user s guide Question 1
 Question 1 How does one find a gene of interest and determine that gene s structure? Once the gene has been located on the map, how does one easily examine other genes in that same region? doi:10.1038/ng966
Question 1 How does one find a gene of interest and determine that gene s structure? Once the gene has been located on the map, how does one easily examine other genes in that same region? doi:10.1038/ng966
