The eye-forming field in the diencephalon has been a classical
|
|
|
- Morgan Ferguson
- 6 years ago
- Views:
Transcription
1 Generation of Rx Pax6 neural retinal precursors from embryonic stem cells Hanako Ikeda*, Fumitaka Osakada*, Kiichi Watanabe*, Kenji Mizuseki*, Tomoko Haraguchi*, Hiroyuki Miyoshi, Daisuke Kamiya*, Yoshihito Honda, Noriaki Sasai*, Nagahisa Yoshimura, Masayo Takahashi, and Yoshiki Sasai* *Organogenesis and Neurogenesis Group, Center for Developmental Biology, RIKEN, Kobe , Japan; Department of Ophthalmology and Visual Sciences, Graduate School of Medicine, Department of Experimental Therapeutics, Translational Research Center, Kyoto University Hospital, and Department of Pharmacology, Graduate School of Pharmaceutical Sciences, Kyoto University, Kyoto , Japan; and BioResource Center, RIKEN, Tsukuba , Japan Edited by Fred H. Gage, The Salk Institute for Biological Studies, San Diego, CA, and approved June 22, 2005 (received for review January 3, 2005) We report directed differentiaion of retinal precursors in vitro from mouse ES cells. Six3 rostral brain progenitors are generated by culturing ES cells under serum-free suspension conditions (SFEB culture) in the presence of Wnt and Nodal antagonists (Dkk1 and LeftyA), and subsequently steered to differentiate into Rx cells (16%) by treatment with activin and serum. Consistent with the characteristics of early neural retinal precursors, the induced Rx cells coexpress Pax6 and the mitotic marker Ki67, but not Nestin. The ES cell-derived precursors efficiently generate cells with the photoreceptor phenotype (rhodopsin, recoverin ) when cocultured with embryonic retinal cells. Furthermore, organotypic culture studies demonstrate the selective integration and survival of ES cell-derived cells with the photoreceptor phenotype (marker expression and morphology) in the outer nuclear layer of the retina. Taken together, ES cells treated with SFEB Dkk1 LeftyA serum activin generate neural retinal precursors, which have the competence of photoreceptor differentiation. regenerative medicine differentiation photoreceptor induction Six3 The eye-forming field in the diencephalon has been a classical paradigm in experimental biology, and the mechanism of how the eye primordium is specified has attracted the interest of embryologists over many decades. In addition, most retinal degeneration diseases in humans are caused by impairment in the neural retina, particularly in photoreceptor cells (1, 2). Therefore, in vitro generation of neural retinal precursors, if successful, would greatly contribute to medical and pharmaceutical researches for retinal diseases. To date, however, only infrequent expression of photoreceptor markers in ES cell-derived neural tissues has been reported (3, 4). The development of the neural retina involves multiple regulatory steps during embryogenesis (see Fig. 1A). A number of transcription factors have been isolated and implicated in the control of the step-wise differentiation of vertebrate retinal tissues (5). In contrast, relatively little has been known about the molecular nature of extracellular factors that mediate tissue interactions to form the retinal tissues in the exact location. To facilitate the study on retinal development, we have attempted to establish an in vitro differentiation system for neural retinal precursors from mouse ES cells. In our previous reports, it has been shown that retinal pigment epithelial cells can be induced at a moderate efficiency from primate ES cells by coculturing with PA6 cells [stromal cell-derived inducing activity (SDIA) method] (6, 7). However, as described below in this study, the frequency of the generation of neural retinal cells in the SDIA system has proven to be low. In addition to the SDIA method, we have recently established another in vitro culture system that induces efficient neural differentiation from ES cells. In this method, floating aggregates of ES cells are cultured under serum-free conditions in the optimized medium without retinoic acid (RA) or exogenous growth factors (serum-free floating culture of embryoid body-like aggregates, SFEB) (8). A characteristic feature of the SFEB culture is that it efficiently induces differentiation of rostral-most CNS tissues. Particularly, the addition of the Wnt antagonist Dkk1 (9) and the Nodal antagonist LeftyA (10) to the SFEB culture during the first 5 days facilitates both neural differentiation (up to 90%) and rostral specification of neural tissues ( 35% are telencephalic tissues) (8). Here, we report efficient in vitro generation of neural retinal precursors from mouse ES cells by combining the SFEB culture and extracellular inductive signals. The ability of the ES cellderived progenitors to produce photoreceptors is also demonstrated. Materials and Methods Cell Culture. The methods of ES cell maintenance and the differentiation by the SDIA method have been described (11). For the SFEB method, dissociated ES cells (EB5 line) per milliliter were incubated in a bacterial-grade dish with ES differentiation medium (G-MEM, 5% KSR 0.1 mm nonessential amino acids 1 mm pyruvate 0.1 mm 2-mercaptoethanol) (8). Floating ES cell aggregates spontaneously formed within 1 day under this condition. After 5 days of suspension culture, the cell aggregates were replated en bloc at a density of aggregates per cm 2 on the poly(d-lysine) laminin fibronectin-coated culture slides. Each growth factor or FCS (JRH Biosciences) was added as indicated in the text and Fig. 1G. Recombinant proteins (Dkk1, LeftyA, activin-a, FGF, and Shh) were purchased fromr&dsystems.the ED 50 of the activin protein is ng ml (the ability to induce hemoglobin expression in K562 cells), and activin was used at sufficient doses (1 100 ng ml) in this study. BrdUrd uptake was examined by culturing cells with 5 g ml BrdUrd in medium for 12 h before fixation. For coculture experiments, SFEB Dkk1 LeftyA FCS activin (DLFA)- or SFEB RA-treated ES cells were dissociated with trypsin into single cells on day 5, labeled with 100 g ml of the rhodamine-labeling reagent [5-(6)-carboxytetramethylrhodamine, succinimidyl ester, which covalently binds to cell surface proteins, Molecular Probes] for 10 min, followed by washing three times with PBS. Anti-rhodamine antibody (see Supporting Text, which is published as supporting information on the PNAS web site) was used for enhancement of the detection when needed. For the genetic labeling using the Venus GFP lentivirus (see below), ES cells were infected with the virus 1 day before the start of the SFEB DLFA culture. These cells were dissociated on day 5 of differentiation and used for coculture experiments. This paper was submitted directly (Track II) to the PNAS office. Abbreviations: SDIA, stromal cell-derived inducing activity; SFEB, serum-free floating culture of embryoid body-like aggregates; RA, retinoic acid; En, embryonic day n; RPE, retinal pigment epithelium; ONL, outer nuclear layer; INL, inner nuclear layer; DLFA, Dkk1 LeftyA FCS activin. To whom correspondence should be addressed. sasaicdb@mub.biglobe.ne.jp by The National Academy of Sciences of the USA DEVELOPMENTAL BIOLOGY cgi doi pnas PNAS August 9, 2005 vol. 102 no
2 Immunohistochemistry, in Situ Hybridization, and Statistical Analysis. Immunohistochemistry was performed as described (11, 12). Detailed information on the use of antibodies and on the antibody production is available in Supporting Text. All images of immunostaining (including three-dimensional reconstruction) on ES cellderived cells were obtained with the LSM5 PASCAL (Zeiss) confocal microscope. Numbers of total cells were counted by staining nuclei with TOTO-3 or DAPI (Molecular Probes). The numbers of positive cells were counted in optical slice images prepared with confocal microscopy. Whole mount in situ hybridization was performed as described (13). A fragment of mouse Six3 cdna ( bp; database accession no. X90871) was amplified by RT-PCR and used as templates for making the RNA probe. For statistical analysis, colonies were examined in each experiment, and each experiment was performed at least three times. All statistical analyses were performed with INSTAT software (version 3.0, GraphPad, San Diego). The statistical significance of difference was determined by one-way ANOVA followed by Tukey s test for experiments in Fig. 1H. Data from Fig. 3B were evaluated by an unpaired t test. Probability values 5% were considered significant. Lentivirus Construction and Production. Replication-defective selfinactivating lentivirus vectors were used (14, 15). The DNA region encoding EGFP in the pcs-cag-egfp (a kind gift of K. Tanabe) was replaced with a fragment encoding Ires2-Venus (a kind gift of A. Miyawaki; psin-cag-ires-venus). A 0.9-kb mouse Crx cdna fragment containing the entire Crx coding region was amplified by PCR and subcloned into psin-cag-ires-venus (the psin-cag- Crx-Ires-Venus). These plasmids were cotransfected with the packaging plasmid (pcag-hivgp) and the VSV-G- and Revexpressing plasmid (pcmv-vsv-g-rsv-rev) into 293T cells. High-titer viral solution ( units ml) was prepared by ultracentrifugation at 80,000 g. Virus solution was applied to the SFEB DLFA, SFEB ( ), or SFEB RA-ES cells on day 7. The cells were analyzed after additional 10-day culture. Fig. 1. Efficient generation of Rx Pax6 retinal precursors from mouse ES cells by a modified SFEB method. (A) Multiple-step commitment in the development of retinal cells. Markers for respective differentiation steps are boxed. (B D) E10.5 mouse embryos immunostained with anti-rx (red) and anti-pax6 (green) antibodies. (B) Cross-section at the diencephalic region. oc, optic cup. A high-power image of the boxed ventral portion is shown in C. Cells in the ventral floor of the diencephalon express only Rx. (D) Cells in the neural retina express both Rx and Pax6. The pigment epithelium and lens cells are Pax6 and Rx. RPE, retinal pigment epithelium; NR, neural retina; L, lens vesicle. In addition to the neural retina, at this early stage, Rx Pax6 cells are found only transiently in the medial end of the optic stalk (ref. 22; Rx Pax6 expression there is shut off soon after). (E) In situ hybridization signals for Six3 in SFEB DL aggregates ( 60 70% cells in the positive aggregates were Six3 ). (F)NoSix3 expression in SFEB RA aggregates. (G) Schema of the modified SFEB culture. D, culture day. (H) Percentages of Rx Pax6 colonies (containing more than five double-positive cells) in SFEB-treated ES cells on day 8. The SFEB culture combined with Dkk1, LeftyA, FCS, and activin gave the highest efficiency for the differentiation of Rx Pax6 cells (**, P 0.01, Tukey s test). DL, SFEB combined with Dkk and Lefty; DLF, with Dkk, Lefty, and FCS; DLFA, with Dkk, Lefty, FCS, and activin. (I K) Immunocytochemical analyses of SFEB DLFAtreated ES cells on day 8 (I and J) and day 7 (K) with confocal microscopy. (I)A majority of colonies contained significant numbers of Rx cells (red). Nuclei were counterstained with TOTO-3 (blue). (J) Induced Rx cells frequently coexpressed Pax6 (green). (K) Nearly all Rx (red) cells were Six3 (green). (Scale bar, 100 m indand I and 20 m inj and K.) Retina Reaggregation Pellet and Explant Coculture. Retinas from 10 mouse embryos [embryonic day (E) 17.5] were excised and separated from other ocular tissues in Hanks balanced salt solution (HBSS), and then dissociated by incubation for 5 10 min at 37 C in 0.05% EDTA PBS with 2.5% trypsin. After being washed with retinal culture medium [66% E-MEM-Hepes (Sigma catalog no. M7278) 33% HBSS (Gibco catalog no ) 1% FCS supplemented with N mg/ml glucose 200 M L-glutamine 100 units/ml penicillin 100 g/ml streptomycin (Gibco)], embryonic retinal cells and rhodamine- or Venus-labeled ES cells (SFEB DLFA, SFEB RA) were mixed and centrifuged to produce reaggregated pellets as described (16). The pellets were cultured on 30-mm membranes (0.4- m diameter pore; Millicell Millipore catalog no. PICM03050) for 12 days at 37 C in a humidified atmosphere of 5% CO 2 with a medium change every other day. Retina explant cultures were prepared as described (17) with minor changes. Briefly, neural retinal tissues from E17.5 mouse embryos were excised and separated from other ocular tissues. About labeled ES cells were placed on the filter membrane before the retina was positioned on the membrane (with the vitreous side up). The retina was incubated in the retinal culture medium at 34 C in a humidified atmosphere of 5% CO 2 for 12 days. For immunohistochemistry, the retinal pellets or explants were fixed and sectioned (20 m thick). Results and Discussion Generation of Rx Pax6 Cells from ES Cells in Vitro. In early embryogenesis, the retinal primordia form within the rostral-most diencephalic region expressing Six3 (18) (Fig. 1A). The transcription factor Rx (an early bona fide marker for the eye field) plays an cgi doi pnas Ikeda et al.
3 essential role for the specification of the retinal primordium in the Six3 rostral CNS (19 21). During early embryogenesis (E10.5), Rx expression coincides with Pax6 expression (22) in the neural retinal progenitors (Fig. 1 B and D), whereas Rx cells in the floor of the ventral diencephalon are Pax6 (Fig. 1C). The retinal pigment epithelium (RPE) is Rx and Pax6 (Fig. 1D). Thus, the neural retinal lineage during early development is characterized by the Rx Pax6 coexpression (Pax6 expression gradually decreases as the neural retina matures). To understand the suitability of the SFEB-based approach (8) for the induction of retinal tissues from ES cells, we first examined the expression of the rostral-most CNS marker Six3. On culture day 5, strong expression of Six3 was found in SFEB-treated ES cell aggregates cultured in the absence and presence of Dkk1 (100 ng ml) plus LeftyA (500 ng ml) (in 82% and 87% of aggregates, respectively; Fig. 1E and data not shown), but not in those cultured with the caudalizing factor RA (0.2 M, during days 3 5; Fig. 1F; ref. 23). Because retinal progenitors arise from the Six3 rostral CNS tissue, we next attempted to induce Rx Pax6 cells from SFEB-induced neural precursors by modifying the culture conditions. Because extracellular patterning signals that determine the induction of retinal primordia in the embryo have not yet been elucidated, we experimentally searched for soluble factors that induced Rx Pax6 expression by testing a number of candidate factors for such activities (Fig. 1 G and H and data not shown). Among them, the most obvious enhancement was seen with treatment of 5% FCS during days 3 5 (Fig. 1H, lane 4). Furthermore, activin-a treatment (100 ng ml) during days 4 6 increased the induction when combined with Dkk1, LeftyA, and FCS treatments (Fig. 1H, lane 5; little effects were seen with activin treatment alone at ng ml, data not shown). We also tested the effects of Shh, Wnt, BMP4, Nodal (without LeftyA), IGF, FGF-1, FGF-2, or FGF antagonists during days 3 6, but observed only marginal effects, if any, on Rx induction in this culture system (data not shown). Hereafter, ES cells treated with SFEB and Dkk1 LeftyA FCS activin are referred to as SFEB DLFA cells. As a whole, Rx cells were found in % of SFEB DLFA cells, whereas 1% Rx cells were generated in SFEB DL-treated ES cells without FCS and activin. At the colony level, % colonies of SFEB DLFA cells were Rx (Fig. 1I) and % colonies contained Rx Pax6 cells (Fig. 1H). In the Rx Pax6 colonies, % were Rx Pax6 cells on average (Fig. 1J), whereas Rx Pax6 cells occupied %. In all, % cells of the culture were strongly positive for both Rx and Pax6. Such frequent induction of Rx Pax6 cells is not commonly seen in other ES cell differentiation systems. For instance, ES cell-derived neural precursors induced by coculture with PA6 cells (SDIA method) (11) generated few Rx Pax6 colonies ( 3%) even upon treatment with serum or activin (data not shown). These findings show that the SFEB DLFA treatment preferentially induces differentiation of Rx Pax6 retinal progenitor-like cells from ES cells. SFEB DLFA-Induced Rx Cells Exhibit a Marker Expression Profile Indistinguishable from That of Embryonic Neural Retinal Progenitors. Consistent with the idea of retinal progenitor differentiation, most of ES cell-derived Rx cells coexpressed Six3 on day 7 (Fig. 1K), which is strongly expressed in the developing neural retina of the E11 mice (18). Next, we further analyzed the nature of Rx cells present in SFEB DLFA cells with multiple eye markers. Otx2 (22) is coexpressed with Rx in the embryonic neural retina (Fig. 2A and data not shown), whereas the RPE is Rx Otx2. Consistent with the in vivo coexpression, nearly all Rx cells in SFEB DLFA cells were Otx2 ( %; Fig. 2B). In the early neural retina (E10.5; Fig. 2C), most Rx cells are proliferating progenitors, which are positive for the mitotic marker Ki67. Similarly, Rx cells in SFEB DLFA cells were Ki67 ( %; Fig. 2D), TuJ1 ( %; postmitotic neuronal marker; Fig. 2E), and positive Fig. 2. Rx cells induced by SFEB DLFA exhibit typical marker characteristics of retinal progenitors. Immunostaining of E10.5 mouse optic cups (A, C, G, I, and K) and SFEB DLFA-treated ES cells (day 8) (B, D F, H, J, and L; confocal images) with anti-rx antibodies (red) [raised in mice (A, B, I, and J) and rabbits (C H, K, and L)] and anti-otx2 (A and B), Ki67 (C and D), TuJ (E), BrdUrd (F), Nestin (G and H), Sox1 (I and J), or Mitf (K and L) antibodies (green). (A) The neural retinal cells (especially in distal portion) coexpress Rx and Otx2. (B) Most ES cell-derived Rx cells coexpressed Otx2. (C and D) Ki67 expression in Rx neural retinal cells (C) and Rx ES cells (D). (E) ES cell-derived Rx cells were negative for TuJ1. (F) Most ES cellderived Rx cells were BrdUrd after 12 h of BrdUrd exposure. (G and H) Lack of Nestin staining in Rx cells of the retina (G) and the ES cell-derived Rx cells (H). (I)Rx retinal cells are Sox1, whereas the cells in the lens vesicle and the neural epithelium of the adjacent diencephalon (arrow) are Sox1.(J)Rx cells induced by SFEB DLFA were negative for Sox1. (K) Mitf cells are Rx except for some cells in the peripheral marginal zone of the embryonic retina on E10.5 (doublepositive cells are more abundant in earlier days). (L) A large portion of the Mitf cells induce by SFEB DLFA were negative for Rx. (Scale bar in K, 100 m for A, C, G, I, and K;inL,20 m, for B, D-F, H, J, and L.) for BrdUrd uptake ( % after 12 h exposure; Fig. 2F), indicating that Rx cells produced by SFEB DLFA are mitotically active. In the embryo, unlike most of the neural progenitors in the CNS, Rx neural retinal precursors do not express Nestin (24) or DEVELOPMENTAL BIOLOGY Ikeda et al. PNAS August 9, 2005 vol. 102 no
4 Sox1 (25) (Fig. 2 G and I). Similarly, most Rx cells in SFEB DLFA cells were Nestin (Fig. 2H) and Sox1 (Fig. 2 J), although they were mitotically active. Collectively, a high proportion of Rx cells produced by SFEB DLFA exhibit characteristics consistent with those of progenitors in the developing neural retina, at least with regard to the marker expression profile tested here. We next asked whether differentiation of another component of the retina, the RPE, was observed in the SFEB DLFA culture, and examined the expression of the early RPE marker Mitf (22) (Fig. 2K). In the SFEB DLFA culture, % of ES cell aggregates were positive for Mitf, whereas % of ES cell aggregates treated with SFEB alone were Mitf. Among Mitf SFEB DLFAtreated cell aggregates, % cells expressed Mitf (i.e., 1% of total cells in culture). Consistent with the in vivo expression profile of RPE markers (22), most Mitf cells in the SFEB DLFA culture were Pax6 (Fig. 6, which is published as supporting information on the PNAS web site). In the SFEB DLFA culture, the majority of Mitf cells were Rx on day 8, although Rx cells were frequently found in the close vicinity of Mitf cell clusters (Fig. 2L). None of Mitf cells on day 12 coexpressed Rx (data not shown). Induction of Rhodopsin Expression by Crx Overexpression in SFEB DLFA-Treated ES Cells. Next, we asked whether SFEB DLFAtreated ES cells could generate the photoreceptor cell, a key component of the neural retina, after a long period culture [adherent culture on poly(d-lysine) laminin fibronectin-coated culture slides in the differentiation medium]. On day 12, % of the colonies contained small clusters of cells positive for the early photoreceptor precursor marker Crx (26, 27) ( 0.5% of total cells in culture; Fig. 7 A and B, which is published as supporting information on the PNAS web site). These Crx cells coexpressed Otx2, consistent with the colocalization in the developing retina (28) (Fig. 7C). On day 17, % of the colonies contained small cell clusters positive for the late photoreceptor marker rhodopsin ( 0.5% of total cells; Fig. 7D). We tested several soluble factors that reportedly have positive effects on in vitro differentiation of late embryonic or neonatal retinal progenitors into photoreceptors (refs. 29 and 30; Shh, FGF-1, FGF-2, Taurine, and RA). No significant enhancement of Crx rhodopsin induction in SFEB DLFA cells was observed with these factors (data not shown; each added from day 9 in serum-free differentiation medium or in 1% FCS containing retinal culture medium). We then asked whether the inefficient Crx rhodopsin induction reflected a small number of progenitors with the competence of photoreceptor differentiation or the lack of proper inductive cues in culture. To date, the exact molecular nature of in vivo signals that induce Crx photoreceptor precursors from Rx retinal progenitors remains elusive. Therefore, we first tested whether SFEB DLFA cells contained sensitized precursors that had the competence to express rhodopsin in response to Crx overexpression. Previous studies have indicated that Crx overexpression induces photoreceptor differentiation in immature retinal tissue (27) and iris-derived precursors (31). Lentivirus-mediated Crx gene transfer (Fig. 3A) remarkably increased the appearance of rhodopsin cells in SFEB DLFA-treated ES cell colonies ( % of Crx virus-infected cells vs % of control Venus-GFP (32) virus-infected cells, P 0.001; Fig. 3 B D). In contrast, forced Crx expression generated only a few rhodopsin cells in SFEB-treated ( % of infected cells; Fig. 3B) and SFEB RA-treated ( %; Fig. 7 E and F) ES cells. No detectable levels of rhodopsin proteins were induced in COS-7 or 293T cells by Crx overexpression. (10 days after infection of the Crx virus; data not shown). These findings demonstrate that the SFEB DLFA-induced progenitors contain sensitized precursors with the ability to express rhodopsin in the presence of Crx, as do immature retinal cells. Fig. 3. Overexpression of Crx efficiently induces rhodopsin cells in SFEB DLFA cells. (A) Lentivirus vectors. CAG, chicken -actin promoter with CMV-IE enhancer; IRES, internal ribosomal unit; RRE, rev responsive element; cppt, central polypurine tract; WPRE, woodchuck hepatitis virus posttranscriptional regulatory element. (B D) ES cells treated with SFEB DLFA (C and D) or SFEB ( ) were infected with the control (Venus only, C) or Crx Venusexpressing (D) lentivirus. (B) Proportions of the rhodopsin cells per Venus cells. The data represent the averages of three independent experiments (***, P 0.001, t test). (C) SFEB DLFA-treated ES cells infected with Venusexpressing virus (green) rarely expressed rhodopsin (red). (D) Rhodopsin induction in SFEB DLFA-treated ES cells infected with the Crx Venus-expressing virus. (Scale bar in D, 20 m, for C and D.) Photoreceptor Differentiation in SFEB DLFA-Treated ES Cells by Coculture with Embryonic Retinal Cells. To obtain further evidence for the generation of neural retinal progenitors from ES cells, we tested whether photoreceptor differentiation could be efficiently induced by mimicking the in vivo environment in culture. We first performed the reaggregation coculture assay (16). In this assay, when cocultured with more mature retinal tissues in vitro, early embryonic retinal progenitors efficiently differentiate into photoreceptors in a fashion similar to in vivo development (16). SFEB DLFAtreated ES cells ( cells) were dissociated on day 5, labeled with the rhodamine dye, mixed with dissociated E17.5 mouse retinal cells (1:50), and spun down to make coaggregates of cells (Fig. 4A). The reaggregated cell pellets were subsequently cultured for 12 days on the culture insert (porous filter membrane) in E-MEM HBSS 1% FCS N2 medium. Under these conditions, surviving cells labeled with rhodamine were found in the cell pellets. In these cell aggregates (where endogenous photoreceptors spontaneously became sorted together and made clusters), 38 6% of the surviving SFEB DLFA cells were found in the areas rich in embryo-derived photoreceptor cells. Interestingly, confocal microscopic analysis showed that 36 3% of ES cell-derived cells in these areas expressed the photoreceptor markers rhodopsin and recoverin (33) (Fig. 4 B F),whereas these markers were rarely observed cgi doi pnas Ikeda et al.
5 Fig. 4. Efficient differentiation of rhodopsin cells by coculture with embryonic retinal tissues. Coculture of the SFEB DLFA-treated ES cells with embryonic retinal cells under the reaggregation pellet coculture. (B H) A number of rhodopsin (green) SFEB DLFA cells (rhodamine labeled; red) were found (B; magnified view, C and D) in the rhodopsin cell-rich cluster of the embryonic retinal cells. Large arrowheads indicate ES cell derivatives that were not incorporated in the rhodopsin cluster of the recipient retinal cells (B). (E and F) Recoverin expression (green; overlay in F) in rhodamine SFEB DLFA cells (red) after reaggregation coculture. (G and H) SFEB DLFA-treated ES cells (red) were cocultured with embryonic retinal cells (expressing GFP). Rhodopsin (blue)-expressing ES cells (red) in G did not express GFP (green) in H.(I L) ES cell-derived cells genetically labeled with Venus were detected with anti-gfp antibody (green). (I and J) Some Venus (green) rhodopsin (red) ES-derived cells had processes (small arrows; overlay in J). White lines mark the direction of the reslicing done along the z axis (I and J ). (K and L) ES-derived cells (green) also expressed recoverin (red; overlay in L). Small arrowheads in C, H, and I indicate the margin of the ES cell. Note that rhodopsin (membrane protein) was localized in the periphery of the cell. All pictures were obtained with confocal microscopy. (Scale bar, 20 m inb; and 10 m inl, for C L.) in rhodamine-labeled cells staying away from the photoreceptorrich regions (arrowheads in Fig. 4B). These observations are consistent with a previous coculture study with neonatal retinal tissues (30) showing that E11 mouse retinal progenitors differentiate into photoreceptors only when they are located closely to neonate-derived rod cell clusters. In contrast to SFEB DLFA cells, the reaggregate coculture experiments using SFEB RA-treated ES cells (containing no Rx Pax6 cells) did not result in significant generation of rhodopsin cells ( % in the embryo-derived photoreceptor-rich areas). Fig. 5. Integration of rhodopsin cells into the outer nuclear layer. (A) Confocal images of organotypic coculture using SFEB DLFA cells (red) and embryonic retinal explants. (B) Integration of ES cell (red, arrows)-derived rhodopsin (green) cells into the ONL. Retinal explants were counterstained with TOTO-3 (blue). Large arrowheads indicate ES cells that were not integrated into the retinal layers. (C and D) High-magnification pictures of the ES-derived cell indicated with the white arrow in B (C, and overlay D). Small arrowheads indicate rhodopsin signals in the periphery of the cell body, and the bracket indicates strong localization of rhodopsin in the outer segmentlike structure. (E) The outer segment-like morphology (indicated with the bracket) and axon-like structure (indicated with a small arrow) in the ES cell derivative in the ONL. (F) Expression of glutamine synthetase (a marker for Müller glia, green) in the ES cell-derived cell of the INL. (G) Proportions of SFEB DLFA cells integrated into each layer of the retinal explant. Few cells were found in the OPL and IPL. G, ganglion cell layer; IPL, inner plexiform layer; INL, inner nuclear layer; OPL, outer plexiform layer; ONL, outer nuclear layer. (Scale bar, 20 m inband F; and 10 m inc E.) These observations show that cells with the photoreceptor-like phenotype are efficiently generated from SFEB DLFA cells in the presence of retina-derived environmental cues (36% of SFEB DLFA cells in the vicinity of photoreceptor clusters, 14% in the whole reaggregates), but not in the absence of them ( 0.5%). Rhodopsin cells did not appear to be generated by cell fusion, because none of these cells showed double (or large) nuclei (n 50). Consistently, when the embryonic retinal tissues were obtained from mice that constitutively expressed GFP (a kind gift of S. Yamada; ref. 34), embryo-derived GFP signals were not colocalized with the rhodamine tracer (green and red in Fig. 4 G and H). In addition, to rule out the possibility of artifacts caused by the rhodamine dye, we confirmed the cell lineage and expression of photoreceptor markers by using ES cells prelabeled with Venus- GFP. Confocal microscopic analysis showed that the ES cellderived Venus signals were detected in rhodopsin cells (Fig. 4 I and J; I and J show reconstructed slice images along the z axis) and in recoverin cells (Fig. 4 K and L). Integration of ES Cell-Derived Photoreceptors into the Proper Retinal Layer by Organotypic Culture. Finally, we performed organotypic culture experiments to test whether ES cell-derived rhodopsin DEVELOPMENTAL BIOLOGY Ikeda et al. PNAS August 9, 2005 vol. 102 no
6 cells had the ability to integrate into the proper location of the retina. Explants of the embryonic neural retina (E17.5) were cultured on a porous filter insert with the vitreous side up as previously (17). A drop of single cell suspension of SFEB DLFAtreated ES cells (day 5; rhodamine-labeled) was placed between the explant and the filter (Fig. 5A). After 12 days of organotypic coculture, a number of ES cell-derived rhodamine cells ( cells per explant) had integrated in various layers of the retinal explant. A total of 10% of the ES cell-derived cells that had integrated into the retinal tissue were located in the outer nuclear layer (ONL), where endogenous photoreceptors lay. Interestingly, a significant proportion of the rhodamine cells present in the outer nuclear layer expressed rhodopsin (37%; Fig. 5B), whereas few rhodopsin cells were found in the other layers. Similar observations were obtained when ES cells were labeled genetically with Venus-GFP, indicating that these findings were not artifacts caused by the dye labeling (Fig. 8, which is published as supporting information on the PNAS web site). Interestingly, the rhodamine rhodopsin cells possessed a characteristic protrusion on the outer side, which is reminiscent of the photoreceptor outer segment (brackets in Fig. 5 B, D, and E). These findings suggest that ES cell-derived rhodopsin photoreceptor-like cells are preferentially incorporated into the exact layer expected for photoreceptor cells and survive there. Another interpretation could be that immature retinal progenitors are preferentially induced to differentiate into photoreceptors after incorporation into the ONL. Taken together with the studies of differentiation markers (Figs. 1 and 2), these observations indicate that SFEB DLFA-treatment promote ES cell differentiation into neural retinal precursors, which have the competence of generating cells with the photoreceptor phenotype. In addition to the ONL, a large proportion ( 75%) of surviving SFEB DLFA cells were found in the inner nuclear layer (INL; Fig. 5G). ES cell derivatives incorporated in the INL expressed the Müller glia marker glutamine synthetase (GS) (26% of cells in the INL; Fig. 5F; ref. 35) and the rod bipolar cell marker PKC (10% of cells in the INL; refs. 35 and 36). In contrast, a relatively small number of surviving SFEB DLFA cells in the ganglion cell layer expressed the ganglion cell marker Islet-1 (37) (9% in this layer and 1% in the explant, data not shown). The relatively low frequency may reflect the fact that ganglion cell differentiation starts much earlier in the embryo than on E17.5 (38). Prospects. In this study, we have demonstrated that Rx Pax6 neural retinal precursors are generated at a significant frequency from ES cells in the SFEB culture combined with Dkk1, LeftyA, FCS, and activin treatment. By applying extracellular inductive signals (or environment), ES cells were steered to differentiate stepwise into Six3 rostral CNS progenitors (Fig. 1E), Rx Pax6 neural retinal progenitors (Fig. 1J), and photoreceptor precursors (Figs. 4 and 5). This report demonstrates systematic induction of neural retinal precursors from ES cells. The signaling mechanism of FCS and activin and their in vivo relevance to the induction of the retinal primordia should be interesting in future investigation. It is important to understand whether these factors act mainly in the induction of Rx Pax6 progenitors or in their growth and maintenance (or both). Further analysis on the effects of additional signals should be crucial for obtaining more selective induction of Rx Pax6 progenitors. Successful generation of ES cell-derived neural retinal progenitors raises the future possibility for its applications in both basic and medical researches on the retina. To this end, it is challenging but important to further improve the efficiency of photoreceptor induction in chemically defined culture by identifying the molecular nature of retina-derived promoting factors (39) for photoreceptor differentiation. In addition, for future functional analysis and transplantation studies, the establishment of a selective purification system of photoreceptor precursors should be crucial. We are grateful to members of the Sasai laboratory for discussion; to S. Nakagawa and H. Kawasaki for comments on this work; to A. Miyawaki (RIKEN), H. Enomoto (RIKEN), and K. Tanabe (RIKEN) for expression plasmids and advice; to F. Vaccarino (Yale University, New Haven, CT) for Otx2 antibody; to S. Yamada (Kyoto University) and T. Suzuki (Kyoto University) for GFP mice; to R. Ladher for advice on rhodamine labeling; to T. Akagi for advice on organotypic culture; and to A. Nishiyama, T. Katayama, M. Kawada, Y. Nakazawa, M. Matsumura, and K. Fukushima for excellent technical assistance. This work was supported by grants-in-aid from the Ministry of Education, Culture, Sports, Science, and Technology (Japan) (to Y.S. and M.T.), the Kobe Cluster Project (to Y.S.), and the Leading Project (to Y.S. and M.T.). 1. Adler, R., Curcio, C., Hicks, D., Price, D. & Wong, F. (1999) Mol. Vis. 5, Delyfer, M.-N., Léveillard, T., Mohand-Säid, S., Hicks, D., Picaud, S. & Sahel, J.-A. (2004) Biol. Cell 96, Zhao, X., Liu, J. & Ahmad, I. (2002) Biochem. Biophys. Res. Commun. 297, Hirano, M., Yamamoto, A., Yoshimura, N., Tokunaga, T., Motohashi, T., Ishizaki, K., Yoshida, H., Okazaki, K., Yamazaki, H., Hayashi, S., et al. (2003) Dev. Dyn. 228, Chow, R. L. & Lang, R. A. (2001) Annu. Rev. Cell Dev. Biol. 17, Kawasaki, H., Suemori, H., Mizuseki, K., Watanabe, K., Urano, F., Ichinose, H., Haruta, M., Takahashi, M., Yoshikawa, K., Nishikawa, S.-I., et al. (2002) Proc. Natl. Acad. Sci. USA 99, Haruta, M., Sasai, Y., Kawasaki, H., Amemiya, K., Ooto, S., Kitada, M., Suemori, H., Nakatsuji, N., Ide, C., Honda, Y., et al. (2004) Invest. Ophthalmol. Vis. Sci. 45, Watanabe, K., Kamiya, D., Nishiyama, A., Katayama, T., Nozaki, S., Kawasaki, H., Watanabe, Y., Mizuseki, K. & Sasai, Y. (2005) Nat. Neurosci. 8, Glinka, A., Wu, W., Delius, H., Monaghan, A. P., Blumenstock, C. & Niehrs, C. (1998) Nature 391, Sakuma, R., Ohnishi, Y., Meno, C., Fujii, H., Juan, H., Takeuchi, J., Ogura, T., Li, E., Miyazono, K. & Hamada, H. (2002) Genes Cells 7, Kawasaki, H., Mizuseki, K., Nishikawa, S., Kaneko, S., Kuwana, Y., Nakanishi, S., Nishikawa, S.-I. & Sasai, Y. (2000) Neuron 28, Mizuseki, K., Sakamoto, T., Watanabe, K., Muguruma, K., Ikeya, M., Nishiyama, A., Arakawa, A., Suemori, H., Nakatsuji, N., Kawasaki, H., et al. (2003) Proc. Natl. Acad. Sci. USA 100, Chitnis, A., Henrique, D., Lewis, J., Ish-Horowicz, D. & Kintner, C. (1995) Nature 375, Miyoshi, H., Takahashi, M., Gage, F. H. & Verma, I. M. (1997) Proc. Natl. Acad. Sci. USA 94, Miyoshi, H., Blömer, U., Takahashi, M., Gage, F. H. & Verma, I. M. (1998) J. Virol. 72, Watanabe, T. & Raff, M. C. (1990) Neuron 4, Hatakeyama, J., Tomita, K., Inoue, T. & Kageyama, R. (2001) Development (Cambridge, U.K.) 128, Oliver, G., Mailhos, A., Wehr, R., Copeland, N. G., Jenkins, N. A. & Gruss, P. (1995) Development (Cambridge, U.K.) 121, Lagutin, O. V., Zhu, C. C., Kobayashi, D., Topczewski, J., Shimamura, K., Puelles, L., Russell, H. R. C., McKinnon, P. J., Solnica-Krezel, L. & Oliver, G. (2003) Genes Dev. 17, Mathers, P. H., Grinberg, A., Mahon, K. A. & Jamrich, M. (1997) Nature 387, Furukawa, T., Kozak, C. A. & Cepko, C. L. (1997) Proc. Natl. Acad. Sci. USA 94, Bäumer, N., Marquardt, T., Stoykova, A., Spieler, D., Treichel, D., Ashery-Padan, R. & Gruss, P. (2003) Development (Cambridge, U.K.) 130, Durston, A. J., van der Wees, J., Pijnappel, W. W. M. & Godsave, S. F. (1998) Curr. Top. Dev. Biol. 40, Yang, J., Bian, W., Gao, X., Chen, L. & Jing, N. (2000) Mech. Dev. 94, Collignon, J., Sockanathan, S., Hacker, A., Cohen-Tannoudji, M., Norris, D., Rastan, S., Stevanovic, M., Goodfellow, P. & Lovell-Badge, R. (1996) Development (Cambridge, U.K.) 122, Chen, S., Wang, Q.-L., Nie, Z., Sun, H., Lennon, G., Copeland, N. G., Gilbert, D. J., Jenkins, N. A. & Zack, D. J. (1997) Neuron 19, Furukawa, T., Morrow, E. M. & Cepko, C. L. (1997) Cell 91, Nishida, A., Furukawa, A., Koike, C., Tano, Y., Aizawa, S., Matsuo, I. & Furukawa, T. (2003) Nat. Neurosci. 6, Levine, E. M., Fuhrmann, S. & Reh, T. A. (2000) Cell. Mol. Life Sci. 57, Reh, T. A. (1992) J. Neurobiol. 23, Haruta, M., Kosaka, M., Kanegae, Y., Saito, I., Inoue, T., Kageyama, R., Nishida, A., Honda, Y. & Takahashi, M. (2001) Nat. Neurosci. 4, Nagai, T., Ibata, K., Park, E. S., Kubota, M., Mikoshiba, K. & Miyawaki, A. (2002) Nat. Biotechnol. 20, McGinnis, J. F., Stepanik, P. L., Chen, W., Elias, R., Cao, W. & Lerious, V. (1999) J. Neurosci. Res. 55, Ikawa, M., Yamada, S., Nakanishi, T. & Okabe, M. (1999) Curr. Top. Dev. Biol. 44, Haverkamp, S. & Wässle, H. (2000) J. Comp. Neurol. 424, Greferath, U., Grünert, U. & Wässle, H. (1990) J. Comp. Neurol. 301, Cayouette, M., Barres, B. A. & Raff, M. (2003) Neuron 40, Marquardt, T. & Gruss, P. (2002) Trends Neurosci. 25, Watanabe, T. & Raff, M. C. (1992) Development (Cambridge, U.K.) 114, cgi doi pnas Ikeda et al.
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/323/5911/256/dc1 Supporting Online Material for HDAC4 Regulates Neuronal Survival in Normal and Diseased Retinas Bo Chen and Constance L. Cepko E-mail: bochen@genetics.med.harvard.edu,
www.sciencemag.org/cgi/content/full/323/5911/256/dc1 Supporting Online Material for HDAC4 Regulates Neuronal Survival in Normal and Diseased Retinas Bo Chen and Constance L. Cepko E-mail: bochen@genetics.med.harvard.edu,
Notch1 (forward: 5 -TGCCTGTGCACACCATTCTGC-3, reverse: Notch2 (forward: 5 -ATGCACCATGACATCGTTCG-3, reverse:
 Supplementary Information Supplementary Methods. RT-PCR. For mouse ES cells, the primers used were: Notch1 (forward: 5 -TGCCTGTGCACACCATTCTGC-3, reverse: 5 -CAATCAGAGATGTTGGAATGC-3 ) 1, Notch2 (forward:
Supplementary Information Supplementary Methods. RT-PCR. For mouse ES cells, the primers used were: Notch1 (forward: 5 -TGCCTGTGCACACCATTCTGC-3, reverse: 5 -CAATCAGAGATGTTGGAATGC-3 ) 1, Notch2 (forward:
Self-organization of neural patterns and structures in 3D culture of stem cells
 Invited Paper Self-organization of neural patterns and structures in 3D culture of stem cells Yoshiki Sasai Organogenesis and Neurogenesis Group, RIKEN Center for Developmental Biology, Kobe 650-0047,
Invited Paper Self-organization of neural patterns and structures in 3D culture of stem cells Yoshiki Sasai Organogenesis and Neurogenesis Group, RIKEN Center for Developmental Biology, Kobe 650-0047,
SUPPLEMENTARY INFORMATION
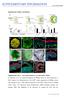 doi:10.1038/nature09941 Supplementary Figures and legends Supplementary Fig. 1 Eye-cup formation in vivo and in ESC culture. a, Schematic of in vivo retinal development. b, SFEBq culture for retinal induction.
doi:10.1038/nature09941 Supplementary Figures and legends Supplementary Fig. 1 Eye-cup formation in vivo and in ESC culture. a, Schematic of in vivo retinal development. b, SFEBq culture for retinal induction.
The retina is a component of the central nervous system
 Potential for neural regeneration after neurotoxic injury in the adult mammalian retina Sotaro Ooto*, Tadamichi Akagi*, Ryoichiro Kageyama, Joe Akita*, Michiko Mandai*, Yoshihito Honda*, and Masayo Takahashi*
Potential for neural regeneration after neurotoxic injury in the adult mammalian retina Sotaro Ooto*, Tadamichi Akagi*, Ryoichiro Kageyama, Joe Akita*, Michiko Mandai*, Yoshihito Honda*, and Masayo Takahashi*
Supplementary Fig. S1. Schematic representation of mouse lines Pax6 fl/fl and mrx-cre used in this study. (A) To generate Pax6 fl/ fl
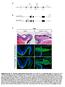 Supplementary Fig. S1. Schematic representation of mouse lines Pax6 fl/fl and mrx-cre used in this study. (A) To generate Pax6 fl/ fl, loxp sites flanking exons 3-6 (red arrowheads) were introduced into
Supplementary Fig. S1. Schematic representation of mouse lines Pax6 fl/fl and mrx-cre used in this study. (A) To generate Pax6 fl/ fl, loxp sites flanking exons 3-6 (red arrowheads) were introduced into
Culture and differentiation of embryonic stem cells. Hong-Lin Su Department of Life Sciences, National Chung-Hsing University
 Culture and differentiation of embryonic stem cells Hong-Lin Su Department of Life Sciences, National Chung-Hsing University Topics-ES cell maintenance Establishment Culture condition, including the serum,
Culture and differentiation of embryonic stem cells Hong-Lin Su Department of Life Sciences, National Chung-Hsing University Topics-ES cell maintenance Establishment Culture condition, including the serum,
Over the past several years, much progress has been made in
 Neural conversion of ES cells by an inductive activity on human amniotic membrane matrix Morio Ueno*, Michiru Matsumura*, Kiichi Watanabe*, Takahiro Nakamura, Fumitaka Osakada*, Masayo Takahashi, Hiroshi
Neural conversion of ES cells by an inductive activity on human amniotic membrane matrix Morio Ueno*, Michiru Matsumura*, Kiichi Watanabe*, Takahiro Nakamura, Fumitaka Osakada*, Masayo Takahashi, Hiroshi
Supplementary Figure 1. Retinogenesis during human embryonic development and in 3-D retinal cups (RCs) derived from hipsc. a, Main steps leading to
 Supplementary Figure 1. Retinogenesis during human embryonic development and in 3-D retinal cups (RCs) derived from hipsc. a, Main steps leading to the formation of the human retina in vivo. b, Diagram
Supplementary Figure 1. Retinogenesis during human embryonic development and in 3-D retinal cups (RCs) derived from hipsc. a, Main steps leading to the formation of the human retina in vivo. b, Diagram
Protocol Using a Dox-Inducible Polycistronic m4f2a Lentivirus to Reprogram MEFs into ips Cells
 STEMGENT Page 1 OVERVIEW The following protocol describes the transduction and reprogramming of one well of Oct4-GFP mouse embryonic fibroblasts (MEF) using the Dox Inducible Reprogramming Polycistronic
STEMGENT Page 1 OVERVIEW The following protocol describes the transduction and reprogramming of one well of Oct4-GFP mouse embryonic fibroblasts (MEF) using the Dox Inducible Reprogramming Polycistronic
Supplementary Figure 1. Soft fibrin gels promote growth and organized mesodermal differentiation. Representative images of single OGTR1 ESCs cultured
 Supplementary Figure 1. Soft fibrin gels promote growth and organized mesodermal differentiation. Representative images of single OGTR1 ESCs cultured in 90-Pa 3D fibrin gels for 5 days in the presence
Supplementary Figure 1. Soft fibrin gels promote growth and organized mesodermal differentiation. Representative images of single OGTR1 ESCs cultured in 90-Pa 3D fibrin gels for 5 days in the presence
Supplementary Data. Supplementary Methods Three-step protocol for spontaneous differentiation of mouse induced pluripotent stem (embryonic stem) cells
 Supplementary Data Supplementary Methods Three-step protocol for spontaneous differentiation of mouse induced pluripotent stem (embryonic stem) cells Mouse induced pluripotent stem cells (ipscs) were cultured
Supplementary Data Supplementary Methods Three-step protocol for spontaneous differentiation of mouse induced pluripotent stem (embryonic stem) cells Mouse induced pluripotent stem cells (ipscs) were cultured
Neural Stem Cell Characterization Kit
 Neural Stem Cell Characterization Kit Catalog No. SCR019 FOR RESEARCH USE ONLY Not for use in diagnostic procedures USA & Canada Phone: +1(800) 437-7500 Fax: +1 (951) 676-9209 Europe +44 (0) 23 8026 2233
Neural Stem Cell Characterization Kit Catalog No. SCR019 FOR RESEARCH USE ONLY Not for use in diagnostic procedures USA & Canada Phone: +1(800) 437-7500 Fax: +1 (951) 676-9209 Europe +44 (0) 23 8026 2233
Nestin expression during mouse eye and lens development
 Mechanisms of Development 94 (2000) 287±291 www.elsevier.com/locate/modo Gene expression pattern Nestin expression during mouse eye and lens development Jing Yang 1,2, Wei Bian 1, Xiang Gao, Leping Chen,
Mechanisms of Development 94 (2000) 287±291 www.elsevier.com/locate/modo Gene expression pattern Nestin expression during mouse eye and lens development Jing Yang 1,2, Wei Bian 1, Xiang Gao, Leping Chen,
Sumitomo Bakelite Co., Ltd. S-BIO Business Division
 Sumitomo Bakelite Co., Ltd. S-BIO Business Division - 1 - ES, ips Spheroid Formation with PrimeSurface MSC - 2 - The Influence of Spheroid Size on Stem Cell Differentiation The size of spheroids are known
Sumitomo Bakelite Co., Ltd. S-BIO Business Division - 1 - ES, ips Spheroid Formation with PrimeSurface MSC - 2 - The Influence of Spheroid Size on Stem Cell Differentiation The size of spheroids are known
Department of Clinical Application, Cebter for ips cell Research and Application, 53 Kawahara-cho Shogoin Sakyo-ku, Kyoto,
 Department of Clinical Application, Cebter for ips cell Research and Application, Kyoto University 53 Kawahara-cho Shogoin Sakyo-ku, Kyoto, 606-8507 JAPAN Phone: 81-75-366-7066, Fax: 81-75-366-7071 Daisuke
Department of Clinical Application, Cebter for ips cell Research and Application, Kyoto University 53 Kawahara-cho Shogoin Sakyo-ku, Kyoto, 606-8507 JAPAN Phone: 81-75-366-7066, Fax: 81-75-366-7071 Daisuke
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature22047 Supplementary discussion Overall developmental timeline and brain regionalization in human whole-brain organoids We first defined the timeline of generation of broadly-defined cell
doi:10.1038/nature22047 Supplementary discussion Overall developmental timeline and brain regionalization in human whole-brain organoids We first defined the timeline of generation of broadly-defined cell
GENESDEV/2007/ Supplementary Figure 1 Elkabetz et al.,
 GENESDEV/2007/089581 Supplementary Figure 1 Elkabetz et al., GENESDEV/2007/089581 Supplementary Figure 2 Elkabetz et al., GENESDEV/2007/089581 Supplementary Figure 3 Elkabetz et al., GENESDEV/2007/089581
GENESDEV/2007/089581 Supplementary Figure 1 Elkabetz et al., GENESDEV/2007/089581 Supplementary Figure 2 Elkabetz et al., GENESDEV/2007/089581 Supplementary Figure 3 Elkabetz et al., GENESDEV/2007/089581
SUPPLEMENTARY INFORMATION
 a before amputation regeneration regenerated limb DERMIS SKELETON MUSCLE SCHWANN CELLS EPIDERMIS DERMIS SKELETON MUSCLE SCHWANN CELLS EPIDERMIS developmental origin: lateral plate mesoderm presomitic mesoderm
a before amputation regeneration regenerated limb DERMIS SKELETON MUSCLE SCHWANN CELLS EPIDERMIS DERMIS SKELETON MUSCLE SCHWANN CELLS EPIDERMIS developmental origin: lateral plate mesoderm presomitic mesoderm
Protocol Reprogramming MEFs using the Dox Inducible Reprogramming Lentivirus Set: Mouse OKSM
 STEMGENT Page 1 OVERVIEW The following protocol describes the reprogramming of one well of mouse embryonic fibroblasts (MEFs) into induced pluripotent stem (ips) cells in a 6-well format. Transduction
STEMGENT Page 1 OVERVIEW The following protocol describes the reprogramming of one well of mouse embryonic fibroblasts (MEFs) into induced pluripotent stem (ips) cells in a 6-well format. Transduction
Supplementary Figure 1 (related to Figure 2) Activation of microglia at early stages after light lesion. In the unlesioned retina, resident microglia
 Supplementary Figure 1 (related to Figure 2) Activation of microglia at early stages after light lesion. In the unlesioned retina, resident microglia labeled with the specific marker 4C4 (magenta) are
Supplementary Figure 1 (related to Figure 2) Activation of microglia at early stages after light lesion. In the unlesioned retina, resident microglia labeled with the specific marker 4C4 (magenta) are
SUPPLEMENTARY NOTE 3. Supplememtary Note 3, Wehr et al., Monitoring Regulated Protein-Protein Interactions Using Split-TEV 1
 SUPPLEMENTARY NOTE 3 Fluorescent Proteolysis-only TEV-Reporters We generated TEV reporter that allow visualizing TEV activity at the membrane and in the cytosol of living cells not relying on the addition
SUPPLEMENTARY NOTE 3 Fluorescent Proteolysis-only TEV-Reporters We generated TEV reporter that allow visualizing TEV activity at the membrane and in the cytosol of living cells not relying on the addition
Figure S1 is related to Figure 1B, showing more details of outer segment of
 Supplemental Information Supplementary Figure legends and Figures Figure S1. Electron microscopic images in Sema4A +/+ and Sema4A / retinas Figure S1 is related to Figure 1B, showing more details of outer
Supplemental Information Supplementary Figure legends and Figures Figure S1. Electron microscopic images in Sema4A +/+ and Sema4A / retinas Figure S1 is related to Figure 1B, showing more details of outer
Transfection of Mouse ES Cells and Mouse ips cells using the Stemfect 2.0 -mesc Transfection Reagent
 APPLICATION NOTE Page 1 Transfection of Mouse ES Cells and Mouse ips cells using the Stemfect 2.0 -mesc Transfection Reagent Authors: Amelia L. Cianci 1, Xun Cheng 1 and Kerry P. Mahon 1,2 1 Stemgent Inc.,
APPLICATION NOTE Page 1 Transfection of Mouse ES Cells and Mouse ips cells using the Stemfect 2.0 -mesc Transfection Reagent Authors: Amelia L. Cianci 1, Xun Cheng 1 and Kerry P. Mahon 1,2 1 Stemgent Inc.,
Chapter 2. Generation of a Three-Dimensional Retinal Tissue from Self-Organizing Human ESC Culture
 Chapter 2 Generation of a Three-Dimensional Retinal Tissue from Self-Organizing Human ESC Culture Atsushi Kuwahara, Tokushige Nakano, and Mototsugu Eiraku Abstract A three-dimensional (3D) tissue generated
Chapter 2 Generation of a Three-Dimensional Retinal Tissue from Self-Organizing Human ESC Culture Atsushi Kuwahara, Tokushige Nakano, and Mototsugu Eiraku Abstract A three-dimensional (3D) tissue generated
Supplementary Protocol. sirna transfection methodology and performance
 Supplementary Protocol sirna transfection methodology and performance sirna oligonucleotides, DNA construct and cell line. Chemically synthesized 21 nt RNA duplexes were obtained from Ambion Europe, Ltd.
Supplementary Protocol sirna transfection methodology and performance sirna oligonucleotides, DNA construct and cell line. Chemically synthesized 21 nt RNA duplexes were obtained from Ambion Europe, Ltd.
Guided differentiation of ES cell into mesenchymal stem cell
 Guided differentiation of ES cell into mesenchymal stem cell Takumi Era Division of Molecular Neurobiology Institute of Molecular Embryology and Genetics, Kumamoto University Differentiation pathways of
Guided differentiation of ES cell into mesenchymal stem cell Takumi Era Division of Molecular Neurobiology Institute of Molecular Embryology and Genetics, Kumamoto University Differentiation pathways of
GM130 Is Required for Compartmental Organization of Dendritic Golgi Outposts
 Current Biology, Volume 24 Supplemental Information GM130 Is Required for Compartmental Organization of Dendritic Golgi Outposts Wei Zhou, Jin Chang, Xin Wang, Masha G. Savelieff, Yinyin Zhao, Shanshan
Current Biology, Volume 24 Supplemental Information GM130 Is Required for Compartmental Organization of Dendritic Golgi Outposts Wei Zhou, Jin Chang, Xin Wang, Masha G. Savelieff, Yinyin Zhao, Shanshan
Pre-made Lentiviral Particles for Nuclear Permeant CRE Recombinase Expression
 Pre-made Lentiviral Particles for Nuclear Permeant CRE Recombinase Expression LVP336 LVP336-PBS LVP339 LVP339-PBS LVP297 LVP297-PBS LVP013 LVP013-PBS LVP338 LVP338-PBS LVP027 LVP027-PBS LVP337 LVP337-PBS
Pre-made Lentiviral Particles for Nuclear Permeant CRE Recombinase Expression LVP336 LVP336-PBS LVP339 LVP339-PBS LVP297 LVP297-PBS LVP013 LVP013-PBS LVP338 LVP338-PBS LVP027 LVP027-PBS LVP337 LVP337-PBS
To isolate single GNS 144 cell clones, cells were plated at a density of 1cell/well
 Supplemental Information: Supplemental Methods: Cell culture To isolate single GNS 144 cell clones, cells were plated at a density of 1cell/well in 96 well Primaria plates in GNS media and incubated at
Supplemental Information: Supplemental Methods: Cell culture To isolate single GNS 144 cell clones, cells were plated at a density of 1cell/well in 96 well Primaria plates in GNS media and incubated at
Quantitative real-time RT-PCR analysis of the expression levels of E-cadherin
 Supplementary Information 1 Quantitative real-time RT-PCR analysis of the expression levels of E-cadherin and ribosomal protein L19 (RPL19) mrna in cleft and bud epithelial cells of embryonic salivary
Supplementary Information 1 Quantitative real-time RT-PCR analysis of the expression levels of E-cadherin and ribosomal protein L19 (RPL19) mrna in cleft and bud epithelial cells of embryonic salivary
In vitro differentiation of retinal cells from human pluripotent stem cells by small-molecule induction
 Research Article 3169 In vitro differentiation of retinal cells from human pluripotent stem cells by small-molecule induction Fumitaka Osakada 1,2,3, *, Zi-Bing Jin 1, Yasuhiko Hirami 1, Hanako Ikeda 1,2,
Research Article 3169 In vitro differentiation of retinal cells from human pluripotent stem cells by small-molecule induction Fumitaka Osakada 1,2,3, *, Zi-Bing Jin 1, Yasuhiko Hirami 1, Hanako Ikeda 1,2,
Flow cytometry Stained cells were analyzed and sorted by SORP FACS Aria (BD Biosciences).
 Mice C57BL/6-Ly5.1 or -Ly5.2 congenic mice were used for LSK transduction and competitive repopulation assays. Animal care was in accordance with the guidelines of Keio University for animal and recombinant
Mice C57BL/6-Ly5.1 or -Ly5.2 congenic mice were used for LSK transduction and competitive repopulation assays. Animal care was in accordance with the guidelines of Keio University for animal and recombinant
Supplemental Figure 1 (Figure S1), related to Figure 1 Figure S1 provides evidence to demonstrate Nfatc1Cre is a mouse line that directed gene
 Developmental Cell, Volume 25 Supplemental Information Brg1 Governs a Positive Feedback Circuit in the Hair Follicle for Tissue Regeneration and Repair Yiqin Xiong, Wei Li, Ching Shang, Richard M. Chen,
Developmental Cell, Volume 25 Supplemental Information Brg1 Governs a Positive Feedback Circuit in the Hair Follicle for Tissue Regeneration and Repair Yiqin Xiong, Wei Li, Ching Shang, Richard M. Chen,
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Generation of NSCs from hpscs in SDC medium.
 Supplementary Figure 1 Generation of NSCs from hpscs in SDC medium. (a) Q-PCR of pluripotent markers (OCT4, NANOG), neural markers (SOX1, PAX6, N-Cadherin), markers for the other germ layers (T, EOMES,
Supplementary Figure 1 Generation of NSCs from hpscs in SDC medium. (a) Q-PCR of pluripotent markers (OCT4, NANOG), neural markers (SOX1, PAX6, N-Cadherin), markers for the other germ layers (T, EOMES,
All quality control test results are reported on a lot specific Certificate of Analysis which is available at or upon request.
 PRIME-XV Neural Basal Medium PRIME-XV Neural Basal Medium is a chemically-defined basal medium optimized for the culture and maintenance of neuronal cells when supplemented with PRIME-XV IS21 Supplement
PRIME-XV Neural Basal Medium PRIME-XV Neural Basal Medium is a chemically-defined basal medium optimized for the culture and maintenance of neuronal cells when supplemented with PRIME-XV IS21 Supplement
Protocol Reprogramming Human Fibriblasts using the Dox Inducible Reprogramming Polycistronic Lentivirus Set: Human 4F2A LoxP
 STEMGENT Page 1 OVERVIEW The following protocol describes the reprogramming of one well of BJ Human Fibroblasts (BJ cells) into induced pluripotent stem (ips) cells in a 6-well format. Transduction efficiency
STEMGENT Page 1 OVERVIEW The following protocol describes the reprogramming of one well of BJ Human Fibroblasts (BJ cells) into induced pluripotent stem (ips) cells in a 6-well format. Transduction efficiency
Input. Pulldown GST. GST-Ahi1
 A kd 250 150 100 75 50 37 25 -GST-Ahi1 +GST-Ahi1 kd 148 98 64 50 37 22 GST GST-Ahi1 C kd 250 148 98 64 50 Control Input Hap1A Hap1 Pulldown GST GST-Ahi1 Hap1A Hap1 Hap1A Hap1 Figure S1. (A) Ahi1 western
A kd 250 150 100 75 50 37 25 -GST-Ahi1 +GST-Ahi1 kd 148 98 64 50 37 22 GST GST-Ahi1 C kd 250 148 98 64 50 Control Input Hap1A Hap1 Pulldown GST GST-Ahi1 Hap1A Hap1 Hap1A Hap1 Figure S1. (A) Ahi1 western
Xeno-Free Systems for hesc & hipsc. Facilitating the shift from Stem Cell Research to Clinical Applications
 Xeno-Free Systems for hesc & hipsc Facilitating the shift from Stem Cell Research to Clinical Applications NutriStem Defined, xeno-free (XF), serum-free media (SFM) specially formulated for growth and
Xeno-Free Systems for hesc & hipsc Facilitating the shift from Stem Cell Research to Clinical Applications NutriStem Defined, xeno-free (XF), serum-free media (SFM) specially formulated for growth and
In general, 8- to 10-week-old adult females were ovariectomized and rested for 10 days
 Animal injections and tissue processing In general, 8- to 10-week-old adult females were ovariectomized and rested for 10 days before they received any injections. Mice were injected with sesame seed oil
Animal injections and tissue processing In general, 8- to 10-week-old adult females were ovariectomized and rested for 10 days before they received any injections. Mice were injected with sesame seed oil
REGENERATIVE MEDICINE
 REGENERATIVE MEDICINE Defining the Integration Capacity of Embryonic Stem Cell-Derived Photoreceptor Precursors EMMA L. WEST, a ANAI GONZALEZ-CORDERO, a CLAIRE HIPPERT, a FUMITAKA OSAKADA, b JUAN PEDRO
REGENERATIVE MEDICINE Defining the Integration Capacity of Embryonic Stem Cell-Derived Photoreceptor Precursors EMMA L. WEST, a ANAI GONZALEZ-CORDERO, a CLAIRE HIPPERT, a FUMITAKA OSAKADA, b JUAN PEDRO
Sarker et al. Supplementary Material. Subcellular Fractionation
 Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
TISSUE ENGINEERING AND REGENERATIVE MEDICINE
 TISSUE ENGINEERING AND REGENERATIVE MEDICINE Integration-Free Induced Pluripotent Stem Cells Derived from Retinitis Pigmentosa Patient for Disease Modeling ZI-BING JIN, a,b SATOSHI OKAMOTO, b PING XIANG,
TISSUE ENGINEERING AND REGENERATIVE MEDICINE Integration-Free Induced Pluripotent Stem Cells Derived from Retinitis Pigmentosa Patient for Disease Modeling ZI-BING JIN, a,b SATOSHI OKAMOTO, b PING XIANG,
To examine the silencing effects of Celsr3 shrna, we co transfected 293T cells with expression
 Supplemental figures Supplemental Figure. 1. Silencing expression of Celsr3 by shrna. To examine the silencing effects of Celsr3 shrna, we co transfected 293T cells with expression plasmids for the shrna
Supplemental figures Supplemental Figure. 1. Silencing expression of Celsr3 by shrna. To examine the silencing effects of Celsr3 shrna, we co transfected 293T cells with expression plasmids for the shrna
Supplementary Figure 1. Generation of B2M -/- ESCs. Nature Biotechnology: doi: /nbt.3860
 Supplementary Figure 1 Generation of B2M -/- ESCs. (a) Maps of the B2M alleles in cells with the indicated B2M genotypes. Probes and restriction enzymes used in Southern blots are indicated (H, Hind III;
Supplementary Figure 1 Generation of B2M -/- ESCs. (a) Maps of the B2M alleles in cells with the indicated B2M genotypes. Probes and restriction enzymes used in Southern blots are indicated (H, Hind III;
Antibiotic Resistance: Ampicillin Bacterial Backbone: pbluescriptksii(+), Agilent Technologies
 G0656 pfiv3.2rsvmcs Coordinates Plasmid Features 1-682 CMV/5 LTR hybrid 958-1468 Partial Gag 1494-1653 Central Polypurine Tract 1695-2075 RSV Promoter 2104-2222 Multiple Cloning Sites 2244-2839 WPRE 2868-3009
G0656 pfiv3.2rsvmcs Coordinates Plasmid Features 1-682 CMV/5 LTR hybrid 958-1468 Partial Gag 1494-1653 Central Polypurine Tract 1695-2075 RSV Promoter 2104-2222 Multiple Cloning Sites 2244-2839 WPRE 2868-3009
Supporting Information
 Supporting Information Patel et al. 10.1073/pnas. SI Materials and Methods Cells and Reagents. Hepatocyte-derived H35 cells were grown in high glucose DMEM (Gibco) media, supplemented with 10% FBS, 2%
Supporting Information Patel et al. 10.1073/pnas. SI Materials and Methods Cells and Reagents. Hepatocyte-derived H35 cells were grown in high glucose DMEM (Gibco) media, supplemented with 10% FBS, 2%
Specification of the Retinal Fate of Mouse Embryonic Stem Cells by Ectopic Expression of Rx/rax, a Homeobox Gene
 MOLECULAR AND CELLULAR BIOLOGY, May 2004, p. 4513 4521 Vol. 24, No. 10 0270-7306/04/$08.00 0 DOI: 10.1128/MCB.24.10.4513 4521.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved.
MOLECULAR AND CELLULAR BIOLOGY, May 2004, p. 4513 4521 Vol. 24, No. 10 0270-7306/04/$08.00 0 DOI: 10.1128/MCB.24.10.4513 4521.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved.
Supplemental Data. Self-Organized Formation of Polarized. Cortical Tissues from ESCs and Its Active. Manipulation by Extrinsic Signals
 Cell Stem Cell, Volume 3 Supplemental Data Self-Organized Formation of Polarized Cortical Tissues from ESCs and Its Active Manipulation by Extrinsic Signals Mototsugu Eiraku, Kiichi Watanabe, Mami Matsuo-Takasaki,
Cell Stem Cell, Volume 3 Supplemental Data Self-Organized Formation of Polarized Cortical Tissues from ESCs and Its Active Manipulation by Extrinsic Signals Mototsugu Eiraku, Kiichi Watanabe, Mami Matsuo-Takasaki,
The Effects of Scaffold Rigidity on Retinal Pigment Epithelial Cells. Corina White Symposium on Biomaterials Science 24 October 2016
 The Effects of Scaffold Rigidity on Retinal Pigment Epithelial Cells Corina White Symposium on Biomaterials Science 24 October 2016 Background Physiology The retina is the light-responsive tissue layer
The Effects of Scaffold Rigidity on Retinal Pigment Epithelial Cells Corina White Symposium on Biomaterials Science 24 October 2016 Background Physiology The retina is the light-responsive tissue layer
A Survey of Genetic Methods
 IBS 8102 Cell, Molecular, and Developmental Biology A Survey of Genetic Methods January 24, 2008 DNA RNA Hybridization ** * radioactive probe reverse transcriptase polymerase chain reaction RT PCR DNA
IBS 8102 Cell, Molecular, and Developmental Biology A Survey of Genetic Methods January 24, 2008 DNA RNA Hybridization ** * radioactive probe reverse transcriptase polymerase chain reaction RT PCR DNA
Nr2e1 regulates retinal lamination and the development of Müller glia, S-cones, and glycineric amacrine cells during retinogenesis
 Corso-Díaz and Simpson Molecular Brain (2015) 8:37 DOI 10.1186/s13041-015-0126-x RESEARCH Nr2e1 regulates retinal lamination and the development of Müller glia, S-cones, and glycineric amacrine cells during
Corso-Díaz and Simpson Molecular Brain (2015) 8:37 DOI 10.1186/s13041-015-0126-x RESEARCH Nr2e1 regulates retinal lamination and the development of Müller glia, S-cones, and glycineric amacrine cells during
Imaging of BacMam Transfected U-2 OS Cells
 A p p l i c a t i o n N o t e Imaging of BacMam Transfected U-2 OS Cells Optimization of Transfection Conditions Using the Cytation 3 Multi- Mode Reader and Gen5 Data Analysis Software Paul Held Ph. D.
A p p l i c a t i o n N o t e Imaging of BacMam Transfected U-2 OS Cells Optimization of Transfection Conditions Using the Cytation 3 Multi- Mode Reader and Gen5 Data Analysis Software Paul Held Ph. D.
Immunofluorescence Confocal Microscopy of 3D Cultures Grown on Alvetex
 Immunofluorescence Confocal Microscopy of 3D Cultures Grown on Alvetex 1.0. Introduction Immunofluorescence uses the recognition of cellular targets by fluorescent dyes or antigen-specific antibodies coupled
Immunofluorescence Confocal Microscopy of 3D Cultures Grown on Alvetex 1.0. Introduction Immunofluorescence uses the recognition of cellular targets by fluorescent dyes or antigen-specific antibodies coupled
Gene Therapy for MYO7A USH1B. Shannon Boye, Ph.D. Assistant Professor Department of Ophthalmology University of Florida
 Gene Therapy for MYO7A USH1B Shannon Boye, Ph.D. Assistant Professor Department of Ophthalmology University of Florida Hearing Vision Vestibular Function (balance) Type 1 Type 2 Type 3 Profound deafness
Gene Therapy for MYO7A USH1B Shannon Boye, Ph.D. Assistant Professor Department of Ophthalmology University of Florida Hearing Vision Vestibular Function (balance) Type 1 Type 2 Type 3 Profound deafness
(a) Scheme depicting the strategy used to generate the ko and conditional alleles. (b) RT-PCR for
 Supplementary Figure 1 Generation of Diaph3 ko mice. (a) Scheme depicting the strategy used to generate the ko and conditional alleles. (b) RT-PCR for different regions of Diaph3 mrna from WT, heterozygote
Supplementary Figure 1 Generation of Diaph3 ko mice. (a) Scheme depicting the strategy used to generate the ko and conditional alleles. (b) RT-PCR for different regions of Diaph3 mrna from WT, heterozygote
Lab Module 7: Cell Adhesion
 Lab Module 7: Cell Adhesion Tissues are made of cells and materials secreted by cells that occupy the spaces between the individual cells. This material outside of cells is called the Extracellular Matrix
Lab Module 7: Cell Adhesion Tissues are made of cells and materials secreted by cells that occupy the spaces between the individual cells. This material outside of cells is called the Extracellular Matrix
Mouse/Rat Neural Stem Cell Functional Identification Kit
 Mouse/Rat Neural Stem Cell Functional Identification Kit Catalog Number SC013 Reagents for the identification of mouse/rat Neural Stem Cells (NSCs) by in vitro functional differentiation. This package
Mouse/Rat Neural Stem Cell Functional Identification Kit Catalog Number SC013 Reagents for the identification of mouse/rat Neural Stem Cells (NSCs) by in vitro functional differentiation. This package
TISSUE-SPECIFIC STEM CELLS
 TISSUE-SPECIFIC STEM CELLS Adult Ciliary Epithelial Cells, Previously Identified as Retinal Stem Cells with Potential for Retinal Repair, Fail to Differentiate into New Rod Photoreceptors SARA GUALDONI,
TISSUE-SPECIFIC STEM CELLS Adult Ciliary Epithelial Cells, Previously Identified as Retinal Stem Cells with Potential for Retinal Repair, Fail to Differentiate into New Rod Photoreceptors SARA GUALDONI,
bfgf Supports Human ES Cell Self- Renewal
 APPLICATION NOTE Page 1 bfgf Supports Human ES Cell Self- Renewal Authors: Dongmei Wu, Wen Xiong, Yan Gao, Kristine Guerrero, Yi Chen, Liming Yang, Yang Liu, and Shuyuan Yao 1 Stemgent, Inc., 10575 Roselle
APPLICATION NOTE Page 1 bfgf Supports Human ES Cell Self- Renewal Authors: Dongmei Wu, Wen Xiong, Yan Gao, Kristine Guerrero, Yi Chen, Liming Yang, Yang Liu, and Shuyuan Yao 1 Stemgent, Inc., 10575 Roselle
Supplement Figure S1:
 Supplement Figure S1: A, Sequence of Xcadherin-11 Morpholino 1 (Xcad-11MO) and 2 (Xcad-11 MO2) and control morpholino in comparison to the Xcadherin-11 sequence. The Xcad-11MO binding sequence spans the
Supplement Figure S1: A, Sequence of Xcadherin-11 Morpholino 1 (Xcad-11MO) and 2 (Xcad-11 MO2) and control morpholino in comparison to the Xcadherin-11 sequence. The Xcad-11MO binding sequence spans the
Figure legends for supplement
 Figure legends for supplement Supplemental Figure 1 Characterization of purified and recombinant proteins Relevant fractions related the final stage of the purification protocol(bingham et al., 1998; Toba
Figure legends for supplement Supplemental Figure 1 Characterization of purified and recombinant proteins Relevant fractions related the final stage of the purification protocol(bingham et al., 1998; Toba
Human ESC-derived Neural Progenitor Expansion Starter Panel. Reagents for the expansion of human neural progenitors.
 Human ESC-derived Neural Progenitor Expansion Starter Panel Catalog Number SC016 Reagents for the expansion of human neural progenitors. This package insert must be read in its entirety before using this
Human ESC-derived Neural Progenitor Expansion Starter Panel Catalog Number SC016 Reagents for the expansion of human neural progenitors. This package insert must be read in its entirety before using this
Plasmid DNA transfection of SW480 human colorectal cancer cells with the Biontex K2 Transfection System
 Plasmid DNA transfection of human colorectal cancer cells with the Biontex K2 Transfection System Stephanie Hehlgans and Franz Rödel, Department of Radiotherapy and Oncology, Goethe- University Frankfurt,
Plasmid DNA transfection of human colorectal cancer cells with the Biontex K2 Transfection System Stephanie Hehlgans and Franz Rödel, Department of Radiotherapy and Oncology, Goethe- University Frankfurt,
A population of Nestin expressing progenitors in the cerebellum exhibits increased tumorigenicity
 A population of Nestin expressing progenitors in the cerebellum exhibits increased tumorigenicity Peng Li 1,2, Fang Du 1, Larra W. Yuelling 1, Tiffany Lin 3, Renata E. Muradimova 1, Rossella Tricarico
A population of Nestin expressing progenitors in the cerebellum exhibits increased tumorigenicity Peng Li 1,2, Fang Du 1, Larra W. Yuelling 1, Tiffany Lin 3, Renata E. Muradimova 1, Rossella Tricarico
Certificate of Analysis
 Certificate of Analysis Catalog No. Amount Lot Number 631986 10 μg Specified on product label. Product Information plvx-ef1α-mcherry-n1 is a lentiviral expression vector that can be used to generate high-titer
Certificate of Analysis Catalog No. Amount Lot Number 631986 10 μg Specified on product label. Product Information plvx-ef1α-mcherry-n1 is a lentiviral expression vector that can be used to generate high-titer
At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in
 Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
Stem Cel s Key Words:
 Stem Cells Key Words: Embryonic stem cells, Adult stem cells, ips cells, self-renewal, differentiation, pluripotent, multipotent, Inner cell mass, Nuclear transfer (Therapeutic cloning), Feeder cells,
Stem Cells Key Words: Embryonic stem cells, Adult stem cells, ips cells, self-renewal, differentiation, pluripotent, multipotent, Inner cell mass, Nuclear transfer (Therapeutic cloning), Feeder cells,
Adenoviral Expression Systems. Lentivirus is not the only choice for gene delivery. Adeno-X
 Adenoviral Expression Systems Lentivirus is not the only choice for gene delivery 3 Adeno-X Why choose adenoviral gene delivery? Table I: Adenoviral vs. Lentiviral Gene Delivery Lentivirus Adenovirus Infects
Adenoviral Expression Systems Lentivirus is not the only choice for gene delivery 3 Adeno-X Why choose adenoviral gene delivery? Table I: Adenoviral vs. Lentiviral Gene Delivery Lentivirus Adenovirus Infects
G0752 pfiv3.2mcswtiresegfp
 G0752 pfiv3.2mcswtiresegfp Plasmid Features Coordinates Feature 1-682 CMV/5 LTR Hybrid 958-1468 Partial Gag 1494-1653 Central Polypurine Tract 1653-1738 Multiple Cloning Sites 1739-2325 wtires 2326-3045
G0752 pfiv3.2mcswtiresegfp Plasmid Features Coordinates Feature 1-682 CMV/5 LTR Hybrid 958-1468 Partial Gag 1494-1653 Central Polypurine Tract 1653-1738 Multiple Cloning Sites 1739-2325 wtires 2326-3045
ipsc Reprogramming from Human Peripheral Blood Using Sendai Virus Mediated Gene Transfer
 ipsc Reprogramming from Human Peripheral Blood Using Sendai Virus Mediated Gene Transfer Wenli Yang 1, Jason A. Mills 2, Spencer Sullivan 3, Ying Liu 1, Deborah L. French 2 and Paul Gadue 2, 1 Institute
ipsc Reprogramming from Human Peripheral Blood Using Sendai Virus Mediated Gene Transfer Wenli Yang 1, Jason A. Mills 2, Spencer Sullivan 3, Ying Liu 1, Deborah L. French 2 and Paul Gadue 2, 1 Institute
Nature Biotechnology: doi: /nbt.4166
 Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
OriGene GFC-Arrays for High-throughput Overexpression Screening of Human Gene Phenotypes
 OriGene GFC-Arrays for High-throughput Overexpression Screening of Human Gene Phenotypes High-throughput Gene Function Validation Tool Introduction sirna screening libraries enable scientists to identify
OriGene GFC-Arrays for High-throughput Overexpression Screening of Human Gene Phenotypes High-throughput Gene Function Validation Tool Introduction sirna screening libraries enable scientists to identify
Cytotoxicity of Botulinum Neurotoxins Reveals a Direct Role of
 Supplementary Information Cytotoxicity of Botulinum Neurotoxins Reveals a Direct Role of Syntaxin 1 and SNAP-25 in Neuron Survival Lisheng Peng, Huisheng Liu, Hongyu Ruan, William H. Tepp, William H. Stoothoff,
Supplementary Information Cytotoxicity of Botulinum Neurotoxins Reveals a Direct Role of Syntaxin 1 and SNAP-25 in Neuron Survival Lisheng Peng, Huisheng Liu, Hongyu Ruan, William H. Tepp, William H. Stoothoff,
Mag4C-Lv Kit - Results
 Mag4C-Lv Kit - Results Mag4C-Lv Kit Magnetic capture for viral concentration & storage buffer for superior viral preservation OZ Biosciences is delighted to announce the launching of a new product Mag4C-Lv
Mag4C-Lv Kit - Results Mag4C-Lv Kit Magnetic capture for viral concentration & storage buffer for superior viral preservation OZ Biosciences is delighted to announce the launching of a new product Mag4C-Lv
supplementary information
 DOI: 10.1038/ncb1919 Mori et al. Supplementary Figure 1a-b a Characterization of the mdrg cells Anti-NF150 DIC (89%: N=154) 100µm Anti-S100 DIC 100µm b (9.1%: N=144) Characterization of the cortical neurons
DOI: 10.1038/ncb1919 Mori et al. Supplementary Figure 1a-b a Characterization of the mdrg cells Anti-NF150 DIC (89%: N=154) 100µm Anti-S100 DIC 100µm b (9.1%: N=144) Characterization of the cortical neurons
ViraBind Lentivirus Concentration and Purification Kit
 Product Manual ViraBind Lentivirus Concentration and Purification Kit Catalog Number VPK-091 VPK-091-5 5 preps 25 preps FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Lentivirus
Product Manual ViraBind Lentivirus Concentration and Purification Kit Catalog Number VPK-091 VPK-091-5 5 preps 25 preps FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Lentivirus
G0411 pfiv3.2mcsegfp. Plasmid Features
 G0411 pfiv3.2mcsegfp Plasmid Features Coordinates Feature CMV/5 LTR hybrid 1:682 Major Splice Donor 926:957 Partial gag 958:1468 Central Polypurine Tract 1494:1653 Multiple cloning sites 1653:1714 egfp
G0411 pfiv3.2mcsegfp Plasmid Features Coordinates Feature CMV/5 LTR hybrid 1:682 Major Splice Donor 926:957 Partial gag 958:1468 Central Polypurine Tract 1494:1653 Multiple cloning sites 1653:1714 egfp
Supporting Information
 Supporting Information Nowakowski et al. 10.1073/pnas.1219385110 SI Materials and Methods Primary Antibodies. Primary antibodies used in this study were raised against BrdU (rat, 1:50; Abcam), cleaved
Supporting Information Nowakowski et al. 10.1073/pnas.1219385110 SI Materials and Methods Primary Antibodies. Primary antibodies used in this study were raised against BrdU (rat, 1:50; Abcam), cleaved
Supporting Information
 Supporting Information Ho et al. 10.1073/pnas.0808899106 SI Materials and Methods In immunostaining, antibodies from Developmental Studies Hybridoma ank were -FasII (mouse, 1:200), - (mouse, 1:100), and
Supporting Information Ho et al. 10.1073/pnas.0808899106 SI Materials and Methods In immunostaining, antibodies from Developmental Studies Hybridoma ank were -FasII (mouse, 1:200), - (mouse, 1:100), and
Plasmid DNA transfection of LN-229 human glioblastoma cells with the Biontex K2 Transfection System
 Plasmid DNA transfection of human glioblastoma cells with the Biontex K2 Transfection System Stephanie Hehlgans and Franz Rödel, Department of Radiotherapy and Oncology, Goethe- University Frankfurt, Theodor-Stern-Kai
Plasmid DNA transfection of human glioblastoma cells with the Biontex K2 Transfection System Stephanie Hehlgans and Franz Rödel, Department of Radiotherapy and Oncology, Goethe- University Frankfurt, Theodor-Stern-Kai
Mag4C-Ad Kit - Results
 Mag4C-Ad Kit - Results Mag4C-Ad Kit Magnetic capture for viral concentration & storage buffer for superior viral preservation OZ Biosciences is delighted to announce the launching of a new product Mag4C-Ad
Mag4C-Ad Kit - Results Mag4C-Ad Kit Magnetic capture for viral concentration & storage buffer for superior viral preservation OZ Biosciences is delighted to announce the launching of a new product Mag4C-Ad
Components Neural induction basal medium (Cat. # ) 100 ml x 5 Neural induction medium supplement 50x (Cat. # ) 2 ml x 5
 HPSC Neural Induction Medium (PSCNIM) Catalog #5931 Introduction Human embryonic stem cells (hesc) and induced pluripotent stem cells (hipsc) possess the capability of differentiating into all derivatives
HPSC Neural Induction Medium (PSCNIM) Catalog #5931 Introduction Human embryonic stem cells (hesc) and induced pluripotent stem cells (hipsc) possess the capability of differentiating into all derivatives
Table of Contents. 2.1 NeuroCult NCFC Assay Kit (Rat) Components Additional Required Reagents Required Equipment...
 i Table of Contents 1.0 Overview of the NeuroCult NCFC Assay 2.0 Materials 2.1 NeuroCult NCFC Assay Kit (Rat) Components... 4 2.2 Additional Required Reagents... 4 2.3 Required Equipment... 4 3.0 Preparation
i Table of Contents 1.0 Overview of the NeuroCult NCFC Assay 2.0 Materials 2.1 NeuroCult NCFC Assay Kit (Rat) Components... 4 2.2 Additional Required Reagents... 4 2.3 Required Equipment... 4 3.0 Preparation
ViraBind Lentivirus Concentration and Purification Kit
 Product Manual ViraBind Lentivirus Concentration and Purification Kit Catalog Number VPK-090 2 preps FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Lentivirus vectors based on
Product Manual ViraBind Lentivirus Concentration and Purification Kit Catalog Number VPK-090 2 preps FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Lentivirus vectors based on
Chapter 20 Recombinant DNA Technology. Copyright 2009 Pearson Education, Inc.
 Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
Selected Techniques Part I
 1 Selected Techniques Part I Gel Electrophoresis Can be both qualitative and quantitative Qualitative About what size is the fragment? How many fragments are present? Is there in insert or not? Quantitative
1 Selected Techniques Part I Gel Electrophoresis Can be both qualitative and quantitative Qualitative About what size is the fragment? How many fragments are present? Is there in insert or not? Quantitative
Therapeutic Cell Replacement. Steven McLoon Department of Neuroscience University of Minnesota
 Therapeutic Cell Replacement Steven McLoon Department of Neuroscience University of Minnesota 1 Neuronal Death Neurons are lost due to four main causes: Trauma Toxin Hypoxia (typically loss of air or blood
Therapeutic Cell Replacement Steven McLoon Department of Neuroscience University of Minnesota 1 Neuronal Death Neurons are lost due to four main causes: Trauma Toxin Hypoxia (typically loss of air or blood
Supplementary Figure S1. Np95 interacts with de novo methyltransferases Dnmt3a and 3b. (A) Co-immunoprecipitation of endogenous Dnmt3a2 (left and
 Supplementary Figure S1. Np95 interacts with de novo methyltransferases Dnmt3a and 3b. (A) Co-immunoprecipitation of endogenous Dnmt3a2 (left and right), Dnmt3b isoforms (left) and Dnmt1 (right) with GFP-Np95
Supplementary Figure S1. Np95 interacts with de novo methyltransferases Dnmt3a and 3b. (A) Co-immunoprecipitation of endogenous Dnmt3a2 (left and right), Dnmt3b isoforms (left) and Dnmt1 (right) with GFP-Np95
Sabrina Jedlicka. Interactions of Neurons and Materials
 Sabrina Jedlicka Interactions of Neurons and Materials Developmental Cell Biology Some Perspective Stem cells: Undifferentiated cells that are capable of proliferation, self-maintenance and differentiation
Sabrina Jedlicka Interactions of Neurons and Materials Developmental Cell Biology Some Perspective Stem cells: Undifferentiated cells that are capable of proliferation, self-maintenance and differentiation
Supplemental Figures Supplemental Figure 1
 Supplemental Figures Supplemental Figure 1 The de novo and salvage purine synthesis pathways are shown converging on GMP synthesis (A). Enzymes and intermediates not involved in MPA action and/or guanosine
Supplemental Figures Supplemental Figure 1 The de novo and salvage purine synthesis pathways are shown converging on GMP synthesis (A). Enzymes and intermediates not involved in MPA action and/or guanosine
Nature Medicine doi: /nm.2548
 Supplementary Table 1: Genotypes of offspring and embryos from matings of Pmm2 WT/F118L mice with Pmm2 WT/R137H mice total events Pmm2 WT/WT Pmm2 WT/R137H Pmm2 WT/F118L Pmm2 R137H/F118L offspring 117 (100%)
Supplementary Table 1: Genotypes of offspring and embryos from matings of Pmm2 WT/F118L mice with Pmm2 WT/R137H mice total events Pmm2 WT/WT Pmm2 WT/R137H Pmm2 WT/F118L Pmm2 R137H/F118L offspring 117 (100%)
StemXVivo. Mouse Oligodendrocyte Differentiation Kit. Catalog Number SC004
 StemXVivo Mouse Oligodendrocyte Differentiation Kit Catalog Number SC004 For the differentiation of mouse pluripotent stem cells to oligodendrocytes. This package insert must be read in its entirety before
StemXVivo Mouse Oligodendrocyte Differentiation Kit Catalog Number SC004 For the differentiation of mouse pluripotent stem cells to oligodendrocytes. This package insert must be read in its entirety before
Table S1. Components for knockout serum replacer medium (KSR medium) (500 ml) Component Supplier Catalogue number
 Table S1. Components for knockout serum replacer medium (KSR medium) (500 ml) Component Supplier Catalogue number Stock Concentration Final concentration Volume used Advanced DMEM/F12 Invitrogen 12634010-78%
Table S1. Components for knockout serum replacer medium (KSR medium) (500 ml) Component Supplier Catalogue number Stock Concentration Final concentration Volume used Advanced DMEM/F12 Invitrogen 12634010-78%
Panx2 expression modulates neuronal differentiation SUPPLEMENTAL DATA. Figure Legends
 Panx2 expression modulates neuronal differentiation SUPPLEMENTAL DATA Figure Legends Suppl. Fig. S1. Antigenic determinants of the Panx2 antibodies employed in this study. (A) Schematic of mouse Panx2
Panx2 expression modulates neuronal differentiation SUPPLEMENTAL DATA Figure Legends Suppl. Fig. S1. Antigenic determinants of the Panx2 antibodies employed in this study. (A) Schematic of mouse Panx2
Sélection Internationale Ens Ulm 2012, Cell Biology
 Sélection Internationale Ens Ulm 2012, Cell Biology Please read the whole subject before starting. Questions 1-8 and 12-14 are general biology questions. In questions 9, 10, 11 and15, we ask you to interpret
Sélection Internationale Ens Ulm 2012, Cell Biology Please read the whole subject before starting. Questions 1-8 and 12-14 are general biology questions. In questions 9, 10, 11 and15, we ask you to interpret
T H E J O U R N A L O F C E L L B I O L O G Y
 Supplemental material Thompson et al., http://www.jcb.org/cgi/content/full/jcb.200909067/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Modification-specific antibodies do not detect unmodified
Supplemental material Thompson et al., http://www.jcb.org/cgi/content/full/jcb.200909067/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Modification-specific antibodies do not detect unmodified
Subsets of retinal progenitors display temporally regulated and distinct biases in the fates of their progeny
 Development 124, 1119-1131 (1997) Printed in Great Britain The Company of Biologists Limited 1997 DEV1122 1119 Subsets of retinal progenitors display temporally regulated and distinct biases in the fates
Development 124, 1119-1131 (1997) Printed in Great Britain The Company of Biologists Limited 1997 DEV1122 1119 Subsets of retinal progenitors display temporally regulated and distinct biases in the fates
B. ADM: C. D. Apoptosis: 1.68% 2.99% 1.31% Figure.S1,Li et al. number. invaded cells. HuH7 BxPC-3 DLD-1.
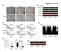 A. - Figure.S1,Li et al. B. : - + - + - + E-cadherin CK19 α-sma vimentin β -actin C. D. Apoptosis: 1.68% 2.99% 1.31% - : - + - + - + Apoptosis: 48.33% 45.32% 44.59% E. invaded cells number 400 300 200
A. - Figure.S1,Li et al. B. : - + - + - + E-cadherin CK19 α-sma vimentin β -actin C. D. Apoptosis: 1.68% 2.99% 1.31% - : - + - + - + Apoptosis: 48.33% 45.32% 44.59% E. invaded cells number 400 300 200
Rat Cortical Stem Cells
 Rat Cortical Stem Cells Catalog Number NSC001 For cortical stem cell expansion and differentiation into neurons, astrocytes, and oligodendrocytes. This package insert must be read in its entirety before
Rat Cortical Stem Cells Catalog Number NSC001 For cortical stem cell expansion and differentiation into neurons, astrocytes, and oligodendrocytes. This package insert must be read in its entirety before
