Host Cell Factor-1 Recruitment to E2F-Bound and Cell-Cycle-Control Genes Is Mediated by THAP11 and ZNF143
|
|
|
- Ethan Bates
- 6 years ago
- Views:
Transcription
1 Cell Reports, Volume 9 Supplemental Information Host Cell Factor-1 Recruitment to E2F-Bound and Cell-Cycle-Control Genes Is Mediated by THAP11 and ZNF143 J. Brandon Parker, Hanwei Yin, Aurimas Vinckevicius, and Debabrata Chakravarti
2 Supplemental Experimental Procedures Stable transfections of RBL1 promoter constructs Pools of T98G cells containing an integrated wildtype or E2F binding site mutant RBL1 promoter construct were created using the site-specific recombinase phic31 integrase and a modified luciferase reporter plasmid essentially as previously described (Hillman et al., 2012). To generate the luciferase reporter plasmid, the minimum 35bp attb sequence (GGTGCCAGGGCGTGCCCTTGGGCTCCCCGGGCGCG) was cloned into pgl4.21 (Promega) downstream of the puromycin selection cassette between restriction sites BstBI and SalI, creating pgl4.21-attb. Next, an RBL1 promoter fragment, corresponding to nucleotides -287 to +42 relative to the annotated transcription start site (NCBI RefSeq mrna NM_ ), was PCR-amplified from human genomic DNA and cloned into pgl4.21-attb between KpnI and EcoRV restriction enzyme sites. The -287/+42 RBL1 promoter containing the tandem E2F binding site mutations described by Burkhart et al. (2010) was prepared as synthetic dsdna (IDT) and cloned into pgl4.21-attb as described above. T98G cells in 35mm dishes were co-transfected with 1 g of a codon-optimized phic31 expression vector (Raymond et al., 2007) (Addgene plasmid #13795) and 100ng of either pgl4.21-attb RBL1-WT or pgl4.21-attb RBL1-E2F mutant plasmids using FugeneHD (Promega) transfection reagent. Approximately 24 hours post-transfection cells were split into 10cm dishes and selected for stable integration with 1 g/ml puromycin. After 10 days of selection the several hundred surviving colonies were trypsinized and RBL1-WT or RBL1-E2F mutant cells were maintained as separate pools under continuous puromycin selection.
3 Retroviral shrna transduction and sirna transfection Non-silencing control, THAP11, and HCFC1 psuper.retro.puro retroviral shrna expression vectors have been described previously (Parker et al., 2012). ZNF143, E2F1, E2F4, and DP1 shrnas were cloned into BglII/HindIII sites of psuper.retro.puro as described (Parker et al., 2012) using oligonucleotides containing the following shrna target sequences: ZNF143, GGACATGCTACAAGAGTAA; E2F1, GGGAGAAGTCACGCTATGA; E2F4, GGATTTACGACATTACCAA; DP1, GAGGAGACTTGAAAGAATA. Two shrnas targeting HPV18 E7 (E7-A, GGAGTTAATCATCAACATT; E7-B, GGAAGAAAACGATGAAATA) were cloned into psuper.retro.puro as described above. All retroviral shrna constructs are available at the plasmid repository Addgene ( /Debu_Chakravarti/). VSV-G pseudotyped retrovirus was produced and used to spin-infect cells as previously described (Parker et al., 2012). Two days post-transduction, cells were split into media containing 2 g/ml puromycin and selected for at least two days or as indicated. For knockdown using sirna, HeLa cells were transfected with 30nM nonsilencing control sirna (Thermo Scientific) or 15nM each of THAP11 and ZNF143 sirna using Lipofectamine RNAiMAX (Invitrogen) according to the manufacturer's instructions. Media was changed 24 hours post transfection, and transfected HeLa cells were harvested for RNA and protein isolation after 72 hour incubation. THAP11 and ZNF143 sirna oligonucleotides corresponding to the shrna targeting sequences were synthesized by Thermo Scientific and are as follows: ZNF143 sense, GGACAUGCUACAAGAGUAAUU; ZNF143 antisense,
4 UUACUCUUGUAGCAUGUCCUU; THAP11 sense, UGAUGGAAGUGAAGAUGAAUU; THAP11 antisense, UUCAUCUUCACUUCCAUCAUU. Immunoblots and Quantitative RT-PCR Whole cell extracts were prepared from cells expressing shrna four days posttransduction using modified RIPA buffer (20mM Tris-HCl ph 7.6, 150mM NaCl, 1mM EDTA, 1mM EGTA, 1% IGEPAL CA-630, 1% sodium deoxycholate, 0.25% SDS). Extracts were clarified by centrifugation at 20,000 g for 15 minutes at 4 C and protein concentrations determined by BCA assay (Pierce). Thirty micrograms of whole cell extract was separated by SDS-PAGE, transferred to nitrocellulose, and immunoblotted as described (Parker et al., 2012). Antibodies and working dilutions used for immunoblot are as follows: THAP11 (R&D Systems #MAB5727, 1:1000), HCFC1 (Bethyl Laboratories #A A, 1:5000), ZNF143 (Novus Biologicals #H M01, 1:2000), E2F1 (Millipore #05-379, 1:2000), DP1 (Santa Cruz Biotechnology #sc- 610x, 1:5000), GAPDH (Sigma #G-9545, 1:10,000), HPV18 E7 (Santa Cruz Biotechnology #sc-1590, 1:500). Immunoblots were developed by chemiluminescence using ECL Prime Western Blotting Detection Reagent (GE Healthcare). Total RNA was prepared from shrna expressing cells four days posttransduction or from sirna transfected cells 3 days post-transfection using Qiagen RNeasy Mini Kit. Total RNA (350ng) was reverse transcribed using qscript cdna Synthesis Kit (Quanta Biosciences) according to manufacturer s instructions. Quantitative PCR was performed on diluted cdna using an Applied Biosystems ABI PRISM 7900HT 384-well real time PCR machine in a final volume of 20 l using SYBR
5 green PCR master mix (Applied Biosystems). HPV18 E6/E7 transcripts were detected using the primer sequences described by Magaldi et al (2012). Immunoprecipitation SW620 cells in 15cm tissue culture dishes were rinsed three times with ice-cold PBS, scraped into PBS and collected by centrifugation at 500 g for five minutes at 4 C. Cells were washed once with Buffer A (10mM HEPES-KOH ph 7.6, 10mM KCl, 1.5mM MgCl 2 ) and then resuspended in 2.5 pellet cell volumes (PCV) of Buffer A plus 0.34M sucrose and Complete protease inhibitors (Roche). Cytoplasmic membranes were lysed by addition of 2.5 PCV of Buffer A, 0.34M sucrose, 0.2% Triton X-100 while gently mixing the cells by vortexing at half-maximum setting. Cells were incubated on ice for 10 minutes and nuclei were isolated by centrifugation at 2000 g for 5 minutes at 4 C. Nuclei were washed once with Buffer A without sucrose, and resuspended in 1 PCV of Buffer C (20mM HEPES-KOH ph 7.6, 420mM NaCl, 1.5mM MgCl 2, 0.2mM EDTA, 25% glycerol) supplemented with Complete protease inhibitors (Roche). Nuclei were extracted for one hour at 4 C with gentle inversion. Nuclear extracts were clarified by centrifugation (20,000 g for 15 minutes at 4 C), diluted with one volume of Buffer C without glycerol or NaCl, and adjusted to 0.2% Triton X-100. Nuclear extracts were reclarified by centrifugation to remove precipitates formed by dilution and immunoprecipitations were performed using the following antibodies: THAP11 (R&D Systems #MAB5727, 2 g), HCF-1 (Bethyl Laboratories #A A, 1 g). Immunoprecipitations were performed for two hours at 4 C with inversion. Protein G Dynabeads (20 l) were added and immunoprecipitation continued for an additional two hours. Beads were then washed four times with binding buffer (20mM HEPES-KOH ph
6 7.6, 150mM NaCl, 1.5mM MgCl 2, 0.2mM EDTA, 0.2% Triton X-100) and bound proteins eluted by boiling in 2x Laemmli buffer. Immunoprecipitated proteins were resolved by SDS-PAGE and immunoblotted with HCFC1 and THAP11 antibodies as described. Blots were developed by enhanced chemiluminescence using an appropriate anti-light chain specific HRP-conjugated secondary antibody (Jackson Immunoresearch). Flow Cytometry Cell cycle profile was determined by propidium iodide (PI) staining flow cytometry. Transduced cells (~ ) were harvested by trypsinization and collected by centrifugation at 500 g for 5 min at 4 C. Cell pellets were washed twice with icecold PBS, and resuspended in cell fixation buffer (200μL ice-cold PBS + 800μL ice-cold ethanol). After overnight fixation at -20 C, cells were recovered by centrifugation (500 g for 5 min at 4 C), washed twice using ice-cold PBS and incubated with 1mL PI staining buffer (50μg/mL propidium iodide, 0.1% Triton-X100, 2mg RNAse A) at 37 C for 30 minutes. Propidium iodide stained cells were analyzed by flow cytometry using a BD LSRFortessa Analyzer (BD Biosciences). Apoptosis was determined by Annexin V/PI staining flow cytometry. Adherent cells were harvested by trypsinization and collected by centrifugation (500 g for 5 min at 4 C), while floating cells were directly collected from medium by centrifugation (500 g for 5 min at 4 C). Adherent and floating cells were combined, washed using ice-cold PBS, and resuspended in Annexin binding buffer (10mM HEPES-KOH ph 7.6, 140mM NaCl, 5mM CaCl 2 ). Final concentration was adjusted to cells/ml. Next, 100μL of cell suspension was incubated with 5μL AlexaFluor 488 conjugated Annexin V (Invitrogen) and 0.5μL propidium iodide (1mg/mL) for 15 min at room temperature. After
7 the incubation period, each sample was gently diluted with 400μL of Annexin binding buffer on ice and analyzed by flow cytometry using a Beckman Coulter Epics XL Analyzer (BD Biosciences). In Silico Analyses To obtain the Thap11 DNA binding matrix, we performed de novo motif discovery analysis on previously-published mouse Thap11 ChIP-Seq data (Dejosez et al., 2010) using MEME software v4.8.1 (Bailey et al., 2009; Bailey and Elkan, 1994). The derived DNA binding matrix was used as input to MAST software v4.8.1 (Bailey et al., 2009; Bailey and Gribskov, 1998) to search for potential THAP11 binding sites within 400bp of ENCODE HA-E2F1 binding sites in HeLa-S3 cells (Consortium, 2011). The potential THAP11 binding sites were ranked by their similarity to the THAP11 DNA binding matrix and mapped to the nearest gene using R ChIPPeakAnno package (Zhu et al., 2010). ChIP-seq data was retrieved from the NCBI Sequence Read Archive (SRA, HCFC1 N [SRX092569], ZNF143 [SRX159127], THAP11 [SRX092574], E2F1 [SRX150563], ZNF143 (ENCODE) [SRX150515]. SRX092569, SRX159127, and SRX data were previously published by Michaud et al. (2013); SRX and SRX were deposited to SRA by the ENCODE consortium. FastQ data was extracted using fastq-dump (v2.3.4, SRA Toolkit, and concatenated into a single file where multiple sequencing files were provided per sample. Sequencing reads were mapped to the human hg19 genome using bowtie (v1.0.0, with -v 0 -m 1 -p 4 -t -S hg19 parameters followed by HOMER s (v4.3, maketagdirectory with -tbp 1 flag. Peaks
8 were called using HOMER s findpeaks.pl (-style factor, FDR of 0.001) and using respective ChIP-seq inputs for background filtering. Co-localized peaks within 500bp of each other (from peak centers) were found using intersectbed (bedtools, - wa flag). Distances between peaks were calculated from peak centers for co-localized peak pairs. Chromatin occupancy correlation analysis was performed only on colocalized peak pairs. Given such peak pairs, HCFC1 and corresponding transcription factor ChIP-seq tags in the 1000bp region surrounding the transcription factor were counted using HOMER s annotatepeaks.pl with size 1000 log parameters. Pearson product-moment correlation coefficient was calculated on inverse log 2 - transformed values using GraphPad. E2F1 peaks lacking THAP11 and ZNF143 peaks within 500bp were determined using intersectbed with v flag. These peaks were then intersected with HCFC1 peaks using intersectbed with f 0.5 wa flags to obtain E2F1/HCFC1 overlapping peaks devoid of nearby THAP11/ZNF143 binding. Supplemental References Bailey, T.L., and Elkan, C. (1994). Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc Int Conf Intell Syst Mol Biol 2, Magaldi, T.G., Almstead, L.L., Bellone, S., Prevatt, E.G., Santin, A.D., and DiMaio, D. (2012). Primary human cervical carcinoma cells require human papillomavirus E6 and E7 expression for ongoing proliferation. Virology 422, Zhu, L.J., Gazin, C., Lawson, N.D., Pages, H., Lin, S.M., Lapointe, D.S., and Green, M.R. (2010). ChIPpeakAnno: a Bioconductor package to annotate ChIP-seq and ChIPchip data. BMC Bioinformatics 11, 237.
9 Supplemental Data Figure S1 Related to Figure 1: HCF-1 occupancy at E2F target genes correlates with THAP11 and ZNF143 binding. Distribution of THAP11, HCF-1, ZNF143, E2F1, and DP1 on the promoter-proximal regions of E2F1 target genes in T98G cells as determined by ChIP-scanning assays.
10 A B Figure S2 Related to Figure 4: E2Fs are dispensable for HCF-1 promoter occupancy. (A) HCF-1 and E2F4 occupancy at the indicated promoters was determined by ChIP assays in HeLa cells expressing either control (shns) or E2F4 shrna. (B) Validation of HCF-1/E2F1-bound promoters as THAP11- and ZNF143-unbound was performed by ChIP. In panels A and B, values represent mean ± standard deviation of duplicate PCR reactions from a single experiment performed at least three times with similar results.
11 A B C D Figure S3 Related to Figure 5: THAP11 and ZNF143, but not E2F1, genome-wide chromatin occupancy correlates HCF-1 binding. (A) Co-localized ChIP-seq peaks were determined as peaks with centers located no more than 500 bases apart. (B) Absolute distances between HCF-1 and ZNF143 (ENCODE) peak centers are shown as boxand-whisker plots. Whiskers indicate 10% and 90% of the population. (C) Correlation of ZNF143 (ENCODE) tag counts with HCF-1 tag counts in overlapping regions. (D) Correlation of E2F1 tag counts with HCF-1 tag counts in overlapping regions devoid of ZNF143 or THAP11 peaks. (C and D) n = total number of peak pairs. r = Pearson product-moment correlation coefficient.
12 A C B Figure S4 Related to Figure 7: THAP11/ZNF143/HCF-1-dependent gene expression contributes to cell proliferation and cell cycle progression. (A) mrna expression changes of the same genes in Fig.7A were determined by quantitative RT-PCR in HeLa cells transfected with the control (sins) or THAP11 and ZNF143 sirna. Values represent the mean ± standard deviation of four independent experiments. Student s T-test p-values are indicated. (B) Representative immunoblot from cells in panel A. (C) Cell cycle analysis of T98G and SW620 cells collected four days post-transduction with the indicated shrnas.
Supplementary Information: Materials and Methods. Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered
 Supplementary Information: Materials and Methods Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered saline (PBS) and lysed in TNN lysis buffer (50mM Tris at ph 8.0, 120mM NaCl
Supplementary Information: Materials and Methods Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered saline (PBS) and lysed in TNN lysis buffer (50mM Tris at ph 8.0, 120mM NaCl
Supporting Information
 Supporting Information Su et al. 10.1073/pnas.1211604110 SI Materials and Methods Cell Culture and Plasmids. Tera-1 and Tera-2 cells (ATCC: HTB- 105/106) were maintained in McCoy s 5A medium with 15% FBS
Supporting Information Su et al. 10.1073/pnas.1211604110 SI Materials and Methods Cell Culture and Plasmids. Tera-1 and Tera-2 cells (ATCC: HTB- 105/106) were maintained in McCoy s 5A medium with 15% FBS
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Transcriptional regulation of BRCA1 expression by a metabolic switch: Di, Fernandez, De Siervi, Longo, and Gardner. H3K4Me3
 ChIP H3K4Me3 enrichment.25.2.15.1.5 H3K4Me3 H3K4Me3 ctrl H3K4Me3 + E2 NS + E2 1. kb kb +82 kb Figure S1. Estrogen promotes entry of MCF-7 into the cell cycle but does not significantly change activation-associated
ChIP H3K4Me3 enrichment.25.2.15.1.5 H3K4Me3 H3K4Me3 ctrl H3K4Me3 + E2 NS + E2 1. kb kb +82 kb Figure S1. Estrogen promotes entry of MCF-7 into the cell cycle but does not significantly change activation-associated
RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the
 Supplementary Methods RT-PCR and real-time PCR analysis RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the manufacturer s protocol and quantified by measuring the
Supplementary Methods RT-PCR and real-time PCR analysis RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the manufacturer s protocol and quantified by measuring the
supplementary information
 DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
Quantitative and non-quantitative RT-PCR. cdna was generated from 500ng RNA (iscript;
 Supplemental Methods Quantitative and non-quantitative RT-PCR. cdna was generated from 500ng RNA (iscript; Bio-Rad, Hercules, CA, USA) and standard RT-PCR experiments were carried out using the 2X GoTaq
Supplemental Methods Quantitative and non-quantitative RT-PCR. cdna was generated from 500ng RNA (iscript; Bio-Rad, Hercules, CA, USA) and standard RT-PCR experiments were carried out using the 2X GoTaq
Fig. S1 TGF RI inhibitor SB effectively blocks phosphorylation of Smad2 induced by TGF. FET cells were treated with TGF in the presence of
 Fig. S1 TGF RI inhibitor SB525334 effectively blocks phosphorylation of Smad2 induced by TGF. FET cells were treated with TGF in the presence of different concentrations of SB525334. Cells were lysed and
Fig. S1 TGF RI inhibitor SB525334 effectively blocks phosphorylation of Smad2 induced by TGF. FET cells were treated with TGF in the presence of different concentrations of SB525334. Cells were lysed and
Supplementary Material
 Supplementary Material Supplementary Methods Cell synchronization. For synchronized cell growth, thymidine was added to 30% confluent U2OS cells to a final concentration of 2.5mM. Cells were incubated
Supplementary Material Supplementary Methods Cell synchronization. For synchronized cell growth, thymidine was added to 30% confluent U2OS cells to a final concentration of 2.5mM. Cells were incubated
1. Cross-linking and cell harvesting
 ChIP is a powerful tool that allows the specific matching of proteins or histone modifications to regions of the genome. Chromatin is isolated and antibodies to the antigen of interest are used to determine
ChIP is a powerful tool that allows the specific matching of proteins or histone modifications to regions of the genome. Chromatin is isolated and antibodies to the antigen of interest are used to determine
Arf6 Activation Assay Kit
 A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Arf6 Activation Assay Kit Catalog Number: 82401 20 assays NewEast Biosciences 1 Table of Content Product
A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Arf6 Activation Assay Kit Catalog Number: 82401 20 assays NewEast Biosciences 1 Table of Content Product
Cdc42 Activation Assay Kit
 A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Cdc42 Activation Assay Kit Catalog Number: 80701 20 assays 1 Table of Content Product Description 3 Assay
A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Cdc42 Activation Assay Kit Catalog Number: 80701 20 assays 1 Table of Content Product Description 3 Assay
Supplementary Materials and Methods
 Supplementary Materials and Methods sirna sequences used in this study The sequences of Stealth Select RNAi for ALK and FLOT-1 were as follows: ALK sense no.1 (ALK): 5 -AAUACUGACAGCCACAGGCAAUGUC-3 ; ALK
Supplementary Materials and Methods sirna sequences used in this study The sequences of Stealth Select RNAi for ALK and FLOT-1 were as follows: ALK sense no.1 (ALK): 5 -AAUACUGACAGCCACAGGCAAUGUC-3 ; ALK
SANTA CRUZ BIOTECHNOLOGY, INC.
 TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
Supplemental Data. Noncoding Transcription by RNA Polymerase Pol IVb/Pol V Mediates Transcriptional Silencing of Overlapping and Adjacent Genes
 Cell, Volume 135 Supplemental Data Noncoding Transcription by RNA Polymerase Pol IVb/Pol V Mediates Transcriptional Silencing of Overlapping and Adjacent Genes Andrzej T. Wierzbicki, Jeremy R. Haag, and
Cell, Volume 135 Supplemental Data Noncoding Transcription by RNA Polymerase Pol IVb/Pol V Mediates Transcriptional Silencing of Overlapping and Adjacent Genes Andrzej T. Wierzbicki, Jeremy R. Haag, and
Apoptosis assay: Apoptotic cells were identified by Annexin V-Alexa Fluor 488 and Propidium
 Apoptosis assay: Apoptotic cells were identified by Annexin V-Alexa Fluor 488 and Propidium Iodide (Invitrogen, Carlsbad, CA) staining. Briefly, 2x10 5 cells were washed once in cold PBS and resuspended
Apoptosis assay: Apoptotic cells were identified by Annexin V-Alexa Fluor 488 and Propidium Iodide (Invitrogen, Carlsbad, CA) staining. Briefly, 2x10 5 cells were washed once in cold PBS and resuspended
RheB Activation Assay Kit
 A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based RheB Activation Assay Kit Catalog Number: 81201 20 assays NewEast Biosciences 1 FAX: 610-945-2008 Table
A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based RheB Activation Assay Kit Catalog Number: 81201 20 assays NewEast Biosciences 1 FAX: 610-945-2008 Table
Rab5 Activation Assay Kit
 A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Rab5 Activation Assay Kit Catalog Number: 83701 20 assays 24 Whitewoods Lane 1 Table of Content Product
A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Rab5 Activation Assay Kit Catalog Number: 83701 20 assays 24 Whitewoods Lane 1 Table of Content Product
Myers Lab ChIP-seq Protocol v Modified January 10, 2014
 Myers Lab ChIP-seq Protocol V011014 1 Contact information: Dr. Florencia Pauli Behn HudsonAlpha Institute for Biotechnology 601 Genome Way Huntsville, AL 35806 Telephone: 256-327-5229 Email: fpauli@hudsonalpha.org
Myers Lab ChIP-seq Protocol V011014 1 Contact information: Dr. Florencia Pauli Behn HudsonAlpha Institute for Biotechnology 601 Genome Way Huntsville, AL 35806 Telephone: 256-327-5229 Email: fpauli@hudsonalpha.org
Cell extracts and western blotting RNA isolation and real-time PCR Chromatin immunoprecipitation (ChIP)
 Cell extracts and western blotting Cells were washed with ice-cold phosphate-buffered saline (PBS) and lysed with lysis buffer. 1 Total cell extracts were separated by SDS-PAGE and transferred to nitrocellulose
Cell extracts and western blotting Cells were washed with ice-cold phosphate-buffered saline (PBS) and lysed with lysis buffer. 1 Total cell extracts were separated by SDS-PAGE and transferred to nitrocellulose
SUPPLEMENTARY INFORMATION
 The Supplementary Information (SI) Methods Cell culture and transfections H1299, U2OS, 293, HeLa cells were maintained in DMEM medium supplemented with 10% fetal bovine serum. H1299 and 293 cells were
The Supplementary Information (SI) Methods Cell culture and transfections H1299, U2OS, 293, HeLa cells were maintained in DMEM medium supplemented with 10% fetal bovine serum. H1299 and 293 cells were
Gα 13 Activation Assay Kit
 A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Gα 13 Activation Assay Kit Catalog Number: 80401 20 assays NewEast Biosciences 1 Table of Content Product
A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Gα 13 Activation Assay Kit Catalog Number: 80401 20 assays NewEast Biosciences 1 Table of Content Product
RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by. 5 AGACACAAACACCAUUGUCACACUCCACAGC; Rand-2 OMe,
 Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Gα i Activation Assay Kit
 A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Gα i Activation Assay Kit Catalog Number 80301 20 assays NewEast Biosciences, Inc 1 Table of Content Product
A helping hand for your research Product Manual Configuration-specific Monoclonal Antibody Based Gα i Activation Assay Kit Catalog Number 80301 20 assays NewEast Biosciences, Inc 1 Table of Content Product
Ren Lab ENCODE in situ HiC Protocol for Tissue
 Ren Lab ENCODE in situ HiC Protocol for Tissue Pulverization, Crosslinking of Tissue Note: Ensure the samples are kept frozen on dry ice throughout pulverization. 1. Pour liquid nitrogen into a mortar
Ren Lab ENCODE in situ HiC Protocol for Tissue Pulverization, Crosslinking of Tissue Note: Ensure the samples are kept frozen on dry ice throughout pulverization. 1. Pour liquid nitrogen into a mortar
SUPPLEMENTAL MATERIALS SIRTUIN 1 PROMOTES HYPEROXIA-INDUCED LUNG EPITHELIAL DEATH INDEPENDENT OF NRF2 ACTIVATION
 SUPPLEMENTAL MATERIALS SIRTUIN PROMOTES HYPEROXIA-INDUCED LUNG EPITHELIAL DEATH INDEPENDENT OF NRF ACTIVATION Haranatha R. Potteti*, Subbiah Rajasekaran*, Senthilkumar B. Rajamohan*, Chandramohan R. Tamatam,
SUPPLEMENTAL MATERIALS SIRTUIN PROMOTES HYPEROXIA-INDUCED LUNG EPITHELIAL DEATH INDEPENDENT OF NRF ACTIVATION Haranatha R. Potteti*, Subbiah Rajasekaran*, Senthilkumar B. Rajamohan*, Chandramohan R. Tamatam,
Supplementary Information
 Supplementary Information Table of contents: Pages: Supplementary Methods 2-3 Supplementary Figure 1 4 Supplementary Figure 2 5 Supplementary Figure 3 6 Supplementary Figure 4 7 Supplementary Figure 5
Supplementary Information Table of contents: Pages: Supplementary Methods 2-3 Supplementary Figure 1 4 Supplementary Figure 2 5 Supplementary Figure 3 6 Supplementary Figure 4 7 Supplementary Figure 5
ASPP1 Fw GGTTGGGAATCCACGTGTTG ASPP1 Rv GCCATATCTTGGAGCTCTGAGAG
 Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental material
 Supplemental material THE JOURNAL OF CELL BIOLOGY Taylor et al., http://www.jcb.org/cgi/content/full/jcb.201403021/dc1 Figure S1. Representative images of Cav 1a -YFP mutants with and without LMB treatment.
Supplemental material THE JOURNAL OF CELL BIOLOGY Taylor et al., http://www.jcb.org/cgi/content/full/jcb.201403021/dc1 Figure S1. Representative images of Cav 1a -YFP mutants with and without LMB treatment.
Chromatin Immunoprecipitation (ChIP)
 de Lange Lab Chromatin Immunoprecipitation (ChIP) Required Solutions IP Wash A 0.1% SDS 1% Triton X-100 2 mm EDTA ph 8.0 20 mm Tris-HCl ph 8.0 150 mm NaCl 1 mm PMSF 1 µg/ml Leupeptin 1 µg/ml Aprotinin
de Lange Lab Chromatin Immunoprecipitation (ChIP) Required Solutions IP Wash A 0.1% SDS 1% Triton X-100 2 mm EDTA ph 8.0 20 mm Tris-HCl ph 8.0 150 mm NaCl 1 mm PMSF 1 µg/ml Leupeptin 1 µg/ml Aprotinin
RNP-IP (Modified Method)-Getting Majority RNA from RNA Binding Protein in the Cytoplasm Fengzhi Liu *
 RNP-IP (Modified Method)-Getting Majority RNA from RNA Binding Protein in the Cytoplasm Fengzhi Liu * School of Biomedical Sciences, Thomas Jefferson University, Philadelphia, USA *For correspondence:
RNP-IP (Modified Method)-Getting Majority RNA from RNA Binding Protein in the Cytoplasm Fengzhi Liu * School of Biomedical Sciences, Thomas Jefferson University, Philadelphia, USA *For correspondence:
ChIP protocol Chromatin fragmentation using the Covaris S2 sonicator by Ethan Ford (version 12/1/11) X- link Cells
 ChIP protocol Chromatin fragmentation using the Covaris S2 sonicator by Ethan Ford (version 12/1/11) X- link Cells 1. Grow six 15 cm plates of HeLa cells to 90% confluency. 2. Remove media and add wash
ChIP protocol Chromatin fragmentation using the Covaris S2 sonicator by Ethan Ford (version 12/1/11) X- link Cells 1. Grow six 15 cm plates of HeLa cells to 90% confluency. 2. Remove media and add wash
ChIPmentation CeMM v1.14 (September 2016)
 ChIPmentation CeMM v1.14 (September 2016) This protocol works well for efficient histone modification and transcription factor antibodies. For less efficient antibodies sonication different sonication-,
ChIPmentation CeMM v1.14 (September 2016) This protocol works well for efficient histone modification and transcription factor antibodies. For less efficient antibodies sonication different sonication-,
HiChIP Protocol Mumbach et al. (CHANG), p. 1 Chang Lab, Stanford University. HiChIP Protocol
 HiChIP Protocol Mumbach et al. (CHANG), p. 1 HiChIP Protocol Citation: Mumbach et. al., HiChIP: Efficient and sensitive analysis of protein-directed genome architecture. Nature Methods (2016). Cell Crosslinking
HiChIP Protocol Mumbach et al. (CHANG), p. 1 HiChIP Protocol Citation: Mumbach et. al., HiChIP: Efficient and sensitive analysis of protein-directed genome architecture. Nature Methods (2016). Cell Crosslinking
IgG TrueBlot Protocol for Mouse, Rabbit or Goatderived Antibodies - For Research Use Only
 IgG TrueBlot Protocol for Mouse, Rabbit or Goatderived Antibodies - For Research Use Only Introduction The IgG TrueBlot for mouse, rabbit, or goat-derived antibodies represents unique series of respective
IgG TrueBlot Protocol for Mouse, Rabbit or Goatderived Antibodies - For Research Use Only Introduction The IgG TrueBlot for mouse, rabbit, or goat-derived antibodies represents unique series of respective
MeCP2. MeCP2/α-tubulin. GFP mir1-1 mir132
 Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
supplementary information
 DOI: 10.1038/ncb1862 Figure S1 Identification of SCAI as a Dia1 associating factor. (a) Identification of SCAI (Riken cdna 930041I02) as a Dia1-FH3 interacting protein. Mouse brain lysate was incubated
DOI: 10.1038/ncb1862 Figure S1 Identification of SCAI as a Dia1 associating factor. (a) Identification of SCAI (Riken cdna 930041I02) as a Dia1-FH3 interacting protein. Mouse brain lysate was incubated
Anti-HB-EGF (Human) mab
 Page 1 For Research Use Only. Not for use in diagnostic procedures. CODE No. D308-3 Anti-HB-EGF (Human) mab CLONALITY CLONE ISOTYPE QUANTITY SOURCE IMMUNOGEN FORMURATION STORAGE Monoclonal 3H4 Mouse IgG1
Page 1 For Research Use Only. Not for use in diagnostic procedures. CODE No. D308-3 Anti-HB-EGF (Human) mab CLONALITY CLONE ISOTYPE QUANTITY SOURCE IMMUNOGEN FORMURATION STORAGE Monoclonal 3H4 Mouse IgG1
Supplementary Information: Materials and Methods. GST and GST-p53 were purified according to standard protocol after
 Supplementary Information: Materials and Methods Recombinant protein expression and in vitro kinase assay. GST and GST-p53 were purified according to standard protocol after induction with.5mm IPTG for
Supplementary Information: Materials and Methods Recombinant protein expression and in vitro kinase assay. GST and GST-p53 were purified according to standard protocol after induction with.5mm IPTG for
Supplementary methods
 Supplementary methods Cell culture, infection, transfection, and RNA interference HEK293 cells and its derivatives were grown in DMEM supplemented with 10% FBS. Various constructs were introduced into
Supplementary methods Cell culture, infection, transfection, and RNA interference HEK293 cells and its derivatives were grown in DMEM supplemented with 10% FBS. Various constructs were introduced into
SUPPLEMENTARY INFORMATION. LIN-28 co-transcriptionally binds primary let-7 to regulate mirna maturation in C. elegans
 SUPPLEMENTARY INFORMATION LIN-28 co-transcriptionally binds primary let-7 to regulate mirna maturation in C. elegans Priscilla M. Van Wynsberghe 1, Zoya S. Kai 1, Katlin B. Massirer 2-4, Victoria H. Burton
SUPPLEMENTARY INFORMATION LIN-28 co-transcriptionally binds primary let-7 to regulate mirna maturation in C. elegans Priscilla M. Van Wynsberghe 1, Zoya S. Kai 1, Katlin B. Massirer 2-4, Victoria H. Burton
Nascent Chromatin Capture. The NCC protocol is designed for SILAC-based massspectrometry
 Extended experimental procedure Nascent Chromatin Capture. The NCC protocol is designed for SILAC-based massspectrometry analysis of nascent versus mature chromatin composition (1). It requires a large
Extended experimental procedure Nascent Chromatin Capture. The NCC protocol is designed for SILAC-based massspectrometry analysis of nascent versus mature chromatin composition (1). It requires a large
SUPPLEMENTAL MATERIAL. Supplemental Methods. Cumate solution was from System Biosciences. Human complement C1q and complement
 SUPPLEMENTAL MATERIAL Supplemental Methods Reagents Cumate solution was from System Biosciences. Human complement Cq and complement C-esterase inhibitor (C-INH) were from Calbiochem. C-INH (Berinert) for
SUPPLEMENTAL MATERIAL Supplemental Methods Reagents Cumate solution was from System Biosciences. Human complement Cq and complement C-esterase inhibitor (C-INH) were from Calbiochem. C-INH (Berinert) for
SUPPLEMENTARY INFORMATION
 (Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
(Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
Supplementary Figures S1-S5. a b c
 Supplementary Figures S1-S5 a b c Supplementary Figure S1. Generation of IL-17RD-deficient mice. (a) Schematic shows the murine il-17rd gene and the genetrap targeting vector, containing a promoter-less
Supplementary Figures S1-S5 a b c Supplementary Figure S1. Generation of IL-17RD-deficient mice. (a) Schematic shows the murine il-17rd gene and the genetrap targeting vector, containing a promoter-less
SOD1 as a Molecular Switch for Initiating the Homeostatic ER Stress Response under Zinc Deficiency
 Molecular Cell, Volume 52 Supplemental Information SOD1 as a Molecular Switch for Initiating the Homeostatic ER Stress Response under Zinc Deficiency Kengo Homma, Takao Fujisawa, Naomi Tsuburaya, Namiko
Molecular Cell, Volume 52 Supplemental Information SOD1 as a Molecular Switch for Initiating the Homeostatic ER Stress Response under Zinc Deficiency Kengo Homma, Takao Fujisawa, Naomi Tsuburaya, Namiko
Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan).
 1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
ChIP-seq Protocol for RNA-Binding Proteins
 ChIP-seq: Cells were grown according to the approved ENCODE cell culture protocols. Cells were fixed in 1% formaldehyde and resuspended in lysis buffer. Chromatin was sheared to 100-500 bp using a Branson
ChIP-seq: Cells were grown according to the approved ENCODE cell culture protocols. Cells were fixed in 1% formaldehyde and resuspended in lysis buffer. Chromatin was sheared to 100-500 bp using a Branson
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.aab3291/dc1 Supplementary Materials for DNA tumor virus oncogenes antagonize the cgas-sting DNA sensing pathway Laura Lau, Elizabeth E. Gray, Rebecca L. Brunette,
www.sciencemag.org/cgi/content/full/science.aab3291/dc1 Supplementary Materials for DNA tumor virus oncogenes antagonize the cgas-sting DNA sensing pathway Laura Lau, Elizabeth E. Gray, Rebecca L. Brunette,
Supplementary information
 Supplementary information Table of Content: Supplementary Results... 2 Supplementary Figure S1: Experimental validation of AP-MS results by coimmunprecipitation Western blot analysis.... 3 Supplementary
Supplementary information Table of Content: Supplementary Results... 2 Supplementary Figure S1: Experimental validation of AP-MS results by coimmunprecipitation Western blot analysis.... 3 Supplementary
Anti-CLOCK (Mouse) mab
 Page 1 For Research Use Only. Not for use in diagnostic procedures. Anti-CLOCK (Mouse) mab CODE No. D349-3 CLONALITY CLONE ISOTYPE QUANTITY SOURCE IMMUNOGEN FORMURATION STORAGE Monoclonal CLSP4 Mouse IgG1
Page 1 For Research Use Only. Not for use in diagnostic procedures. Anti-CLOCK (Mouse) mab CODE No. D349-3 CLONALITY CLONE ISOTYPE QUANTITY SOURCE IMMUNOGEN FORMURATION STORAGE Monoclonal CLSP4 Mouse IgG1
B. ADM: C. D. Apoptosis: 1.68% 2.99% 1.31% Figure.S1,Li et al. number. invaded cells. HuH7 BxPC-3 DLD-1.
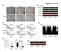 A. - Figure.S1,Li et al. B. : - + - + - + E-cadherin CK19 α-sma vimentin β -actin C. D. Apoptosis: 1.68% 2.99% 1.31% - : - + - + - + Apoptosis: 48.33% 45.32% 44.59% E. invaded cells number 400 300 200
A. - Figure.S1,Li et al. B. : - + - + - + E-cadherin CK19 α-sma vimentin β -actin C. D. Apoptosis: 1.68% 2.99% 1.31% - : - + - + - + Apoptosis: 48.33% 45.32% 44.59% E. invaded cells number 400 300 200
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Chromatin immunoprecipitation (ChIP) ES and FACS-sorted GFP+/Flk1+ cells were fixed in 1% formaldehyde and sonicated until fragments of an average
 Chromatin immunoprecipitation (ChIP) ES and FACS-sorted cells were fixed in % formaldehyde and sonicated until fragments of an average size of 5 bp were obtained. Soluble, sheared chromatin was diluted
Chromatin immunoprecipitation (ChIP) ES and FACS-sorted cells were fixed in % formaldehyde and sonicated until fragments of an average size of 5 bp were obtained. Soluble, sheared chromatin was diluted
Supplemental Data. LMO4 Controls the Balance between Excitatory. and Inhibitory Spinal V2 Interneurons
 Neuron, Volume 61 Supplemental Data LMO4 Controls the Balance between Excitatory and Inhibitory Spinal V2 Interneurons Kaumudi Joshi, Seunghee Lee, Bora Lee, Jae W. Lee, and Soo-Kyung Lee Supplemental
Neuron, Volume 61 Supplemental Data LMO4 Controls the Balance between Excitatory and Inhibitory Spinal V2 Interneurons Kaumudi Joshi, Seunghee Lee, Bora Lee, Jae W. Lee, and Soo-Kyung Lee Supplemental
ATAC-seq Protocol Kaestner Lab
 ATAC-seq Protocol Kaestner Lab Reagents 1X PBS Nuclease-free H 2O NP-40 10% (Sigma/Roche, catalog # 11332473001), store at 4 o C Tween-20 10% (Sigma/Roche, catalog # 11332465001), store at 4 o C Digitonin
ATAC-seq Protocol Kaestner Lab Reagents 1X PBS Nuclease-free H 2O NP-40 10% (Sigma/Roche, catalog # 11332473001), store at 4 o C Tween-20 10% (Sigma/Roche, catalog # 11332465001), store at 4 o C Digitonin
Product Datasheet. Histone H4 [Dimethyl Lys20] Antibody NB SS. Unit Size: mg
![Product Datasheet. Histone H4 [Dimethyl Lys20] Antibody NB SS. Unit Size: mg Product Datasheet. Histone H4 [Dimethyl Lys20] Antibody NB SS. Unit Size: mg](/thumbs/88/115416970.jpg) Product Datasheet Histone H4 [Dimethyl Lys20] Antibody NB21-2089SS Unit Size: 0.025 mg Store at 4C short term. Aliquot and store at -20C long term. Avoid freeze-thaw cycles. Protocols, Publications, Related
Product Datasheet Histone H4 [Dimethyl Lys20] Antibody NB21-2089SS Unit Size: 0.025 mg Store at 4C short term. Aliquot and store at -20C long term. Avoid freeze-thaw cycles. Protocols, Publications, Related
Supporting Information
 Supporting Information He et al. 10.1073/pnas.1116302108 SI Methods Cell Culture. Mouse J774A.1 and RAW 264.7 macrophages were obtained from ATCC and were cultured in MEM supplemented with 10% FS (Sigma)
Supporting Information He et al. 10.1073/pnas.1116302108 SI Methods Cell Culture. Mouse J774A.1 and RAW 264.7 macrophages were obtained from ATCC and were cultured in MEM supplemented with 10% FS (Sigma)
HPV E6 oncoprotein targets histone methyltransferases for modulating specific. Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu,
 1 HPV E oncoprotein targets histone methyltransferases for modulating specific gene transcription 3 5 Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu, Cheng-Ming Chiang, Sheng-Chung
1 HPV E oncoprotein targets histone methyltransferases for modulating specific gene transcription 3 5 Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu, Cheng-Ming Chiang, Sheng-Chung
Online Supplementary Information
 Online Supplementary Information NLRP4 negatively regulates type I interferon signaling by targeting TBK1 for degradation via E3 ubiquitin ligase DTX4 Jun Cui 1,4,6,7, Yinyin Li 1,5,6,7, Liang Zhu 1, Dan
Online Supplementary Information NLRP4 negatively regulates type I interferon signaling by targeting TBK1 for degradation via E3 ubiquitin ligase DTX4 Jun Cui 1,4,6,7, Yinyin Li 1,5,6,7, Liang Zhu 1, Dan
The preparation of native chromatin from cultured human cells.
 Native chromatin immunoprecipitation protocol The preparation of native chromatin from cultured human cells. All solutions need to be ice cold. Sucrose containing solutions must be made up fresh on the
Native chromatin immunoprecipitation protocol The preparation of native chromatin from cultured human cells. All solutions need to be ice cold. Sucrose containing solutions must be made up fresh on the
For immunoprecipitation, cells were serum-starved for 2 hours and treated with
 Supplementary Materials and Methods Immunoprecipitation and Immunoblot Analyses For immunoprecipitation, cells were serum-starved for 2 hours and treated with 12.5ng/ml TGF-β1 for 1 hour and cell lysates
Supplementary Materials and Methods Immunoprecipitation and Immunoblot Analyses For immunoprecipitation, cells were serum-starved for 2 hours and treated with 12.5ng/ml TGF-β1 for 1 hour and cell lysates
Western Blot Tool Box
 Western Blot Tool Box BOX12/BOX12-03 V1.1 Store at 2-8 C For research use only Introduction The Western Blot Tool Box is designed to conveniently provide reagents/buffers needed for Western blotting, from
Western Blot Tool Box BOX12/BOX12-03 V1.1 Store at 2-8 C For research use only Introduction The Western Blot Tool Box is designed to conveniently provide reagents/buffers needed for Western blotting, from
Protein and transcriptome quantitation using BD AbSeq Antibody-Oligonucleotide
 Protein and transcriptome quantitation using BD AbSeq Antibody-Oligonucleotide technology and the 10X Genomics Chromium Single Cell Gene Expression Solution Jocelyn G. Olvera, Brigid S. Boland, John T.
Protein and transcriptome quantitation using BD AbSeq Antibody-Oligonucleotide technology and the 10X Genomics Chromium Single Cell Gene Expression Solution Jocelyn G. Olvera, Brigid S. Boland, John T.
Analysing protein protein interactions using a GST-fusion protein to pull down the interacting target from the cell lysate Hong Wang and Xin Zeng
 Analysing protein protein interactions using a GST-fusion protein to pull down the interacting target from the cell lysate Hong Wang and Xin Zeng Department of Molecular Genetics, Biochemistry and Microbiology,
Analysing protein protein interactions using a GST-fusion protein to pull down the interacting target from the cell lysate Hong Wang and Xin Zeng Department of Molecular Genetics, Biochemistry and Microbiology,
OPPF-UK Standard Protocols: Mammalian Expression
 OPPF-UK Standard Protocols: Mammalian Expression Joanne Nettleship joanne@strubi.ox.ac.uk Table of Contents 1. Materials... 3 2. Cell Maintenance... 4 3. 24-Well Transient Expression Screen... 5 4. DNA
OPPF-UK Standard Protocols: Mammalian Expression Joanne Nettleship joanne@strubi.ox.ac.uk Table of Contents 1. Materials... 3 2. Cell Maintenance... 4 3. 24-Well Transient Expression Screen... 5 4. DNA
ChIP cross-link Gold. ChIP grade reagent I Separately available. Cat. No. C Version 1 I 06.15
 ChIP cross-link Gold ChIP grade reagent I Separately available Cat. No. C01019027 Version 1 I 06.15 Contacts diagenode headquarters Diagenode s.a. BELGIUM EUROPE LIEGE SCIENCE PARK Rue Bois Saint-Jean,
ChIP cross-link Gold ChIP grade reagent I Separately available Cat. No. C01019027 Version 1 I 06.15 Contacts diagenode headquarters Diagenode s.a. BELGIUM EUROPE LIEGE SCIENCE PARK Rue Bois Saint-Jean,
Comparative Analysis of Argonaute-Dependent Small RNA Pathways in Drosophila
 Molecular Cell, Volume 32 Supplemental Data Comparative Analysis of Argonaute-Dependent Small RNA Pathways in Drosophila Rui Zhou, Ikuko Hotta, Ahmet M. Denli, Pengyu Hong, Norbert Perrimon, and Gregory
Molecular Cell, Volume 32 Supplemental Data Comparative Analysis of Argonaute-Dependent Small RNA Pathways in Drosophila Rui Zhou, Ikuko Hotta, Ahmet M. Denli, Pengyu Hong, Norbert Perrimon, and Gregory
Chromatin Immunoprecipitation (ChIP) Assay (PROT11)
 Chromatin Immunoprecipitation (ChIP) Assay (PROT11) Antigone Kouskouti & Irene Kyrmizi Institute of Molecular Biology and Biotechnology FORTH 1527 Vassilika Vouton 711 10, Heraklion, Crete, Greece Email
Chromatin Immunoprecipitation (ChIP) Assay (PROT11) Antigone Kouskouti & Irene Kyrmizi Institute of Molecular Biology and Biotechnology FORTH 1527 Vassilika Vouton 711 10, Heraklion, Crete, Greece Email
Intracellular receptors specify complex patterns of gene expression that are cell and gene
 SUPPLEMENTAL RESULTS AND DISCUSSION Some HPr-1AR ARE-containing Genes Are Unresponsive to Androgen Intracellular receptors specify complex patterns of gene expression that are cell and gene specific. For
SUPPLEMENTAL RESULTS AND DISCUSSION Some HPr-1AR ARE-containing Genes Are Unresponsive to Androgen Intracellular receptors specify complex patterns of gene expression that are cell and gene specific. For
used at a final concentration of 5 ng/ml. Rabbit anti-bim and mouse anti-mkp2 antibodies were
 1 Supplemental Methods Reagents and chemicals: TGFβ was a generous gift from Genzyme Inc. (Cambridge, MA) and was used at a final concentration of 5 ng/ml. Rabbit anti-bim and mouse anti-mkp2 antibodies
1 Supplemental Methods Reagents and chemicals: TGFβ was a generous gift from Genzyme Inc. (Cambridge, MA) and was used at a final concentration of 5 ng/ml. Rabbit anti-bim and mouse anti-mkp2 antibodies
Chromatin Immunoprecipitation (ChIP) Assay*
 Chromatin Immunoprecipitation (ChIP) Assay* Chromatin Immunoprecipitation (ChIP) assays are used to evaluate the association of proteins with specific DNA regions. The technique involves crosslinking of
Chromatin Immunoprecipitation (ChIP) Assay* Chromatin Immunoprecipitation (ChIP) assays are used to evaluate the association of proteins with specific DNA regions. The technique involves crosslinking of
CELLSCRIPT RNA for Translation in Cells
 TM RNA for Translation in Cells Cat. No. C-ACM04037 INTRODUCTION The AmpliCap-Max T7 High Yield Message Maker Kit produces capped RNA by in vitro transcription using T7 RNA polymerase and the standard
TM RNA for Translation in Cells Cat. No. C-ACM04037 INTRODUCTION The AmpliCap-Max T7 High Yield Message Maker Kit produces capped RNA by in vitro transcription using T7 RNA polymerase and the standard
Supplementary Figure 1. GST pull-down analysis of the interaction of GST-cIAP1 (A, B), GSTcIAP1
 Legends Supplementary Figure 1. GST pull-down analysis of the interaction of GST- (A, B), GST mutants (B) or GST- (C) with indicated proteins. A, B, Cell lysate from untransfected HeLa cells were loaded
Legends Supplementary Figure 1. GST pull-down analysis of the interaction of GST- (A, B), GST mutants (B) or GST- (C) with indicated proteins. A, B, Cell lysate from untransfected HeLa cells were loaded
Supplemental Methods Cell lines and culture
 Supplemental Methods Cell lines and culture AGS, CL5, BT549, and SKBR were propagated in RPMI 64 medium (Mediatech Inc., Manassas, VA) supplemented with % fetal bovine serum (FBS, Atlanta Biologicals,
Supplemental Methods Cell lines and culture AGS, CL5, BT549, and SKBR were propagated in RPMI 64 medium (Mediatech Inc., Manassas, VA) supplemented with % fetal bovine serum (FBS, Atlanta Biologicals,
Document S1. Supplemental Experimental Procedures and Three Figures (see next page)
 Supplemental Data Document S1. Supplemental Experimental Procedures and Three Figures (see next page) Table S1. List of Candidate Genes Identified from the Screen. Candidate genes, corresponding dsrnas
Supplemental Data Document S1. Supplemental Experimental Procedures and Three Figures (see next page) Table S1. List of Candidate Genes Identified from the Screen. Candidate genes, corresponding dsrnas
Propidium Iodide. Catalog Number: Structure: Molecular Formula: C 27H 34I 2N 4. Molecular Weight: CAS #
 Catalog Number: 195458 Propidium Iodide Structure: Molecular Formula: C 27H 34I 2N 4 Molecular Weight: 668.45 CAS # 25535-16-4 Physical Description: Dark red crystals Description: Reagent used for the
Catalog Number: 195458 Propidium Iodide Structure: Molecular Formula: C 27H 34I 2N 4 Molecular Weight: 668.45 CAS # 25535-16-4 Physical Description: Dark red crystals Description: Reagent used for the
ab Ran Activation Assay Kit
 ab173247 Ran Activation Assay Kit Instructions for Use For the simple and fast measurement of Ran activation. This product is for research use only and is not intended for diagnostic use. Version 1 Last
ab173247 Ran Activation Assay Kit Instructions for Use For the simple and fast measurement of Ran activation. This product is for research use only and is not intended for diagnostic use. Version 1 Last
Figure S1. Fold activation. Development Supplementary information. δ51+s/p5 Nes+S/P5 Nes+BS/BP5
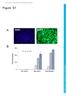 Figure S A DAPI EGFP B 50 0 5 20 Fold activation 00 50 0 δ5+s/p5 Nes+S/P5 Nes+BS/BP5 C Anti-ZIC2 Anti-H2 Anti-OTX2 Anti-H2 + ZIC2 + OTX2 50 40 30 20 0 + ZIC2 40 30 20 0 +OTX2 0 0 0.2 0.4 0.6 0.8.2.4.6.8
Figure S A DAPI EGFP B 50 0 5 20 Fold activation 00 50 0 δ5+s/p5 Nes+S/P5 Nes+BS/BP5 C Anti-ZIC2 Anti-H2 Anti-OTX2 Anti-H2 + ZIC2 + OTX2 50 40 30 20 0 + ZIC2 40 30 20 0 +OTX2 0 0 0.2 0.4 0.6 0.8.2.4.6.8
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling
 Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Protocol(Research use only)
 Immunohistochemistry (without pretreatment) p2 Immunohistochemistry (Microwave pretreatment) p3 Immunohistochemistry (Autoclave pretreatment) p4 Immunohistochemistry (Trypsin pretreatment) p5 Immunohistochemistry
Immunohistochemistry (without pretreatment) p2 Immunohistochemistry (Microwave pretreatment) p3 Immunohistochemistry (Autoclave pretreatment) p4 Immunohistochemistry (Trypsin pretreatment) p5 Immunohistochemistry
Description of supplementary material file
 Description of supplementary material file In the supplementary results we show that the VHL-fibronectin interaction is indirect, mediated by fibronectin binding to COL4A2. This provides additional information
Description of supplementary material file In the supplementary results we show that the VHL-fibronectin interaction is indirect, mediated by fibronectin binding to COL4A2. This provides additional information
Rapid amplification of cdna ends (RACE)
 Rapid amplification of cdna ends (RACE) Rapid amplification of cdna ends (RACE) is a technique used in molecular biology to obtain the full length sequence of an RNA transcript found within a cell. RACE
Rapid amplification of cdna ends (RACE) Rapid amplification of cdna ends (RACE) is a technique used in molecular biology to obtain the full length sequence of an RNA transcript found within a cell. RACE
Electrophoretic Mobility Shift Assay (EMSA). Nuclear extracts were. oligonucleotide spanning the NF-kB site (5 -GATCC-
 SUPPLEMENTARY MATERIALS AND METHODS Electrophoretic Mobility Shift Assay (EMSA). Nuclear extracts were prepared as previously described. (1) A [ 32 P] datp-labeled doublestranded oligonucleotide spanning
SUPPLEMENTARY MATERIALS AND METHODS Electrophoretic Mobility Shift Assay (EMSA). Nuclear extracts were prepared as previously described. (1) A [ 32 P] datp-labeled doublestranded oligonucleotide spanning
QuickExtract RNA Extraction Kit
 Cat. Nos. QER09015 and QER090150 Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook (facebook.com/epicentrebio), and Twitter (@EpicentreBio). www.epicentre.com Lit. # 299 12/2012 1
Cat. Nos. QER09015 and QER090150 Connect with Epicentre on our blog (epicentral.blogspot.com), Facebook (facebook.com/epicentrebio), and Twitter (@EpicentreBio). www.epicentre.com Lit. # 299 12/2012 1
EPIGENTEK. EpiQuik Chromatin Immunoprecipitation Kit. Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Chromatin Immunoprecipitation Kit Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Chromatin Immunoprecipitation Kit is suitable for combining the specificity of
EpiQuik Chromatin Immunoprecipitation Kit Base Catalog # P-2002 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Chromatin Immunoprecipitation Kit is suitable for combining the specificity of
Sarker et al. Supplementary Material. Subcellular Fractionation
 Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Supplemental Information. PARP1 Represses PAP and Inhibits Polyadenylation during Heat Shock
 Molecular Cell, Volume 49 Supplemental Information PARP1 Represses PAP and Inhibits Polyadenylation during Heat Shock Dafne Campigli Di Giammartino, Yongsheng Shi, and James L. Manley Supplemental Information
Molecular Cell, Volume 49 Supplemental Information PARP1 Represses PAP and Inhibits Polyadenylation during Heat Shock Dafne Campigli Di Giammartino, Yongsheng Shi, and James L. Manley Supplemental Information
EPIGENTEK. EpiQuik Tissue Chromatin Immunoprecipitation Kit. Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Tissue Chromatin Immunoprecipitation Kit Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Tissue Chromatin Immunoprecipitation Kit is suitable for combining the specificity
EpiQuik Tissue Chromatin Immunoprecipitation Kit Base Catalog # P-2003 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE The EpiQuik Tissue Chromatin Immunoprecipitation Kit is suitable for combining the specificity
Supplementary Figure 1. HiChIP provides confident 1D factor binding information.
 Supplementary Figure 1 HiChIP provides confident 1D factor binding information. a, Reads supporting contacts called using the Mango pipeline 19 for GM12878 Smc1a HiChIP and GM12878 CTCF Advanced ChIA-PET
Supplementary Figure 1 HiChIP provides confident 1D factor binding information. a, Reads supporting contacts called using the Mango pipeline 19 for GM12878 Smc1a HiChIP and GM12878 CTCF Advanced ChIA-PET
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1154040/dc1 Supporting Online Material for Selective Blockade of MicroRNA Processing by Lin-28 Srinivas R. Viswanathan, George Q. Daley,* Richard I. Gregory* *To whom
www.sciencemag.org/cgi/content/full/1154040/dc1 Supporting Online Material for Selective Blockade of MicroRNA Processing by Lin-28 Srinivas R. Viswanathan, George Q. Daley,* Richard I. Gregory* *To whom
Cell Hashing Protocol
 Cell Hashing Protocol For experiments involving Cell Hashing, use cost per cell calculator to plan experiments, determine number of hashes, number of cells to load, expected doublet rates (detected and
Cell Hashing Protocol For experiments involving Cell Hashing, use cost per cell calculator to plan experiments, determine number of hashes, number of cells to load, expected doublet rates (detected and
Blimp-1/PRDM1 rabbit monoclonal antibody (C14A4) was purchased from Cell Signaling
 1 2 3 4 5 6 7 8 9 10 11 Supplementary Methods Antibodies Blimp-1/PRDM1 rabbit monoclonal antibody (C14A4) was purchased from Cell Signaling (Danvers, MA) and used at 1:1000 to detect the total PRDM1 protein
1 2 3 4 5 6 7 8 9 10 11 Supplementary Methods Antibodies Blimp-1/PRDM1 rabbit monoclonal antibody (C14A4) was purchased from Cell Signaling (Danvers, MA) and used at 1:1000 to detect the total PRDM1 protein
SUPPLEMENTARY EXPEMENTAL PROCEDURES
 SUPPLEMENTARY EXPEMENTAL PROCEDURES Plasmids- Total RNAs were extracted from HeLaS3 cells and reverse-transcribed using Superscript III Reverse Transcriptase (Invitrogen) to obtain DNA template for the
SUPPLEMENTARY EXPEMENTAL PROCEDURES Plasmids- Total RNAs were extracted from HeLaS3 cells and reverse-transcribed using Superscript III Reverse Transcriptase (Invitrogen) to obtain DNA template for the
High Pure RNA Isolation Kit for isolation of total RNA from 50 samples Cat. No
 for isolation of total RNA from 50 samples Cat. No. 1 88 665 Principle A single reagent lyses the sample lysis and inactivates RNase. In the presence of a chaotropic salt (guanidine HCl), the released
for isolation of total RNA from 50 samples Cat. No. 1 88 665 Principle A single reagent lyses the sample lysis and inactivates RNase. In the presence of a chaotropic salt (guanidine HCl), the released
Cells were plated and allowed to adhere to culture plates for 24hrs and then treated as specified.
 Supplementary Methods Flow cytometry Cells were plated and allowed to adhere to culture plates for 24hrs and then treated as specified. Subsequently, cells were harvested in 1x PBS containing 0.1% BSA.
Supplementary Methods Flow cytometry Cells were plated and allowed to adhere to culture plates for 24hrs and then treated as specified. Subsequently, cells were harvested in 1x PBS containing 0.1% BSA.
RhoC Activation Assay Kit
 Product Manual RhoC Activation Assay Kit Catalog Number STA-403-C 20 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Small GTP-binding proteins (or GTPases) are a family
Product Manual RhoC Activation Assay Kit Catalog Number STA-403-C 20 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Small GTP-binding proteins (or GTPases) are a family
Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas.
 Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas. Target Gene Non-target / Control BTG2 BRCA1 NFE2L2 Target Sequence ON-TARGET plus Non-targeting sirna # 1 (Cat# D-001810-01-05) sirna1: GAACCGACAUGCUCCCGGA
Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas. Target Gene Non-target / Control BTG2 BRCA1 NFE2L2 Target Sequence ON-TARGET plus Non-targeting sirna # 1 (Cat# D-001810-01-05) sirna1: GAACCGACAUGCUCCCGGA
XactEdit Cas9 Nuclease with NLS User Manual
 XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
XactEdit Cas9 Nuclease with NLS User Manual An RNA-guided recombinant endonuclease for efficient targeted DNA cleavage Catalog Numbers CE1000-50K, CE1000-50, CE1000-250, CE1001-250, CE1001-1000 Table of
