Why DNA Isn t Your Destiny
|
|
|
- Ambrose Floyd
- 5 years ago
- Views:
Transcription
1 Technical Journal Club April Why DNA Isn t Your Destiny Despoina Goniotaki Aguzzi Group
2 GENOME: a genome is an organism s complete set of DNA, including all of its genes. Each genome contains all of the information needed to build and maintain that organism T C G A modified by mashable.com modified by
3 From GENOME to EPIGENOME to CELLS to ORGANISMS trillion cells!!!! Cells listen for signals Types of signals 1. Direct Contact 2. Transmission (factor release) 3. Hormones 4. Combo modified by
4 From GENOME to EPIGENOME to CELLS to ORGANISMS Proteins Carry Signals to the DNA Gene Regulatory Proteins Have Two Functions Switch genes ON/OFF Recruit epi tag- ENZYMES Paradigm ORGANISM modified by
5 Force a rethink of the definition of a gene and of the minimum unit of heredity.
6 EPIGENOME: the epigenome comprises all of the chemical modifications (tags) added to the genome as a way to regulate the activity (expression) of all the genes. MODIFICATIONS II. Histone Modifications III. ncrna associated gene silencing I. DNA methylation modified by modified by
7 RESEARCH CONSORTIA BLOOD CELLS CELL LINES 111 CELLS & TISSUES
8 AIMS 1. How is genetic information interpreted by single cells? 2. Annotate cis-regulatory elements. 3. Create maps of epigenomic modifications. 4. Produce clinically usefull epigenetic information. 5. Dissect gene regulatory programs in development and disease.
9 METHODS
10 METHODS I. DNA methylation a) Sodium bisulfite modification b) Sequence-specific enzyme digestion c) Capture/quantification of methylated DNA
11 METHODS a) Sodium bisulfite modification
12 METHODS a) Sodium bisulfite modification Molecular Psychiatry (2007) 12, ; doi: /sj.mp
13 METHODS b) Sequence-specific enzyme digestion Molecular Psychiatry (2007) 12, ; doi: /sj.mp
14 METHODS c) Capture/quantification of methylated DNA (MeDIP)
15 METHODS II. Chromatin Modifications a) Chromatin Immunoprecipitation
16 METHODS a) Chromatin Immunoprecipitation (ChIP)
17 METHODS a) RNA-protein interactions (RIP)
18
19 MATERIAL
20 DATA SETS for each epigenome
21 Chromatin State Annotation across tissues Vertical Red Lines = PROMOTERS Dispersed Yellow Regions = ENHANCERS Promoters are primarily constitutive, while Enhancers are highly dynamic
22 Inactive state Active state Chromatin State and DNA methylation dynamics Chromatine States Human ES cells
23 Chromatin State and DNA methylation dynamics During Lineage Specification ES cell differentiation
24 Chromatin State and DNA methylation dynamics During Lineage Specification Skin Cells Keratinocytes Surface Ectoderm Melanocytes Neural Crest Fibroblasts Mesoderm Low overlap in DNA methylation & histone modification signatures.
25 Epigenome Relationships Lines with common developmental origins show similar epigenetic modification patterns.
26 CONCLUSION Common developmental origins can be a primary determinant of global DNA methylation patterns.
27 Cell Type differences in Chromatin states Most Constitutive Quiescent Regions Constitutive Active states H3K4me1 Tissue Specific Inactive states
28 CELL TYPE SPECIFICITY Chromatin states 1. Hematopoietic stem cells and Immune cells show a consistent and previously unrecognized depletion of active and bivalent promoters (TssA, TssBiv) and weakly transcribed states (TxWk). 2. ES cells and ips cells show enrichment of TssBiv, consistent with previous studies. They also show a depletion of ReprPCWk, possibly due to restriction of H3K27m3-establishing Polycomb proteins to promoter regions.
29 Epigenetic Dynamics - Regulators
30 Linking Regulators to tissues & cell types
31 Linking Epigenomic Enrichments to Disease traits
32
33
34 Analysis Pipeline
35 Analysis Pipeline Large number of sequences Variable size regions Sequences are greatly unbalanced for G-C content Motif Combinations
36 Analysis Pipeline -EPIGRAM Effect of sequence-set balancing (SSB) on sequence sets HOMER & EPIGRAM algorithms Full set of motifs
37 True positive rate Area under the curve Prediction Performance Receiver Operating Characteristic (ROC) curve False positive rate Motifs predictive of epigenetic modification Heat Map
38 Motif distribution Paradigm Motif distribution is correlated with H3K27ac variation
39 SUMMARY The first comprehensive catalog of DNA motifs Define the mechanisms by which DNA motifs orchestrate the epigenome In light of the genome editing technologies, these approaches can be used to guide locus-specific epigenome editing through alteration of the regulatory cis-elements.
40 OVERVIEW 1. How the epigenome affects gene expression? 2. How the epigenome changes during stem-cell differentiation (normal development)? 3. How the epigenome changes during disease?
41 CAVEATS 1. Studies are based on analysis of cell populations Cellular Variability within populations 2. Tissue Sample: Enhancer landscapes represent the composite of cell types that make up the tissue Use purified cell populations from in vivo sources 3. The DNA sequences found in cell specific enhancers provide clues for TF that regulate the enhancers these clues must be validated experimentally
42 Follow the Threads EPIGENOMIC NNOTATIONS EPI TAGS DEVELOPMENT REGULATORY MODELS EVOLUTION DISEASE BRAIN
43 Thank you!
NGS Approaches to Epigenomics
 I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
I519 Introduction to Bioinformatics, 2013 NGS Approaches to Epigenomics Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Background: chromatin structure & DNA methylation Epigenomic
What Can the Epigenome Teach Us About Cellular States and Diseases?
 What Can the Epigenome Teach Us About Cellular States and Diseases? (a computer scientist s view) Luca Pinello Outline Epigenetic: the code over the code What can we learn from epigenomic data? Resources
What Can the Epigenome Teach Us About Cellular States and Diseases? (a computer scientist s view) Luca Pinello Outline Epigenetic: the code over the code What can we learn from epigenomic data? Resources
Measuring Protein-DNA interactions
 Measuring Protein-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Transcription Factors are genetic switches 3 Regulation of Gene Expression by Transcription
Measuring Protein-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Transcription Factors are genetic switches 3 Regulation of Gene Expression by Transcription
DNA. bioinformatics. epigenetics methylation structural variation. custom. assembly. gene. tumor-normal. mendelian. BS-seq. prediction.
 Epigenomics T TM activation SNP target ncrna validation metagenomics genetics private RRBS-seq de novo trio RIP-seq exome mendelian comparative genomics DNA NGS ChIP-seq bioinformatics assembly tumor-normal
Epigenomics T TM activation SNP target ncrna validation metagenomics genetics private RRBS-seq de novo trio RIP-seq exome mendelian comparative genomics DNA NGS ChIP-seq bioinformatics assembly tumor-normal
2/10/17. Contents. Applications of HMMs in Epigenomics
 2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
2/10/17 I529: Machine Learning in Bioinformatics (Spring 2017) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Background:
ChIP-seq analysis 2/28/2018
 ChIP-seq analysis 2/28/2018 Acknowledgements Much of the content of this lecture is from: Furey (2012) ChIP-seq and beyond Park (2009) ChIP-seq advantages + challenges Landt et al. (2012) ChIP-seq guidelines
ChIP-seq analysis 2/28/2018 Acknowledgements Much of the content of this lecture is from: Furey (2012) ChIP-seq and beyond Park (2009) ChIP-seq advantages + challenges Landt et al. (2012) ChIP-seq guidelines
What is Epigenetics? Watch the video
 EPIGENETICS What is Epigenetics? The study of environmental factors on gene expression in DNA. The molecule is called methylation controls when genes are turned on. Methylation turns off genes. Acetylation
EPIGENETICS What is Epigenetics? The study of environmental factors on gene expression in DNA. The molecule is called methylation controls when genes are turned on. Methylation turns off genes. Acetylation
Genetics Journal Club YoSon Park Feb 11, Announcement: Marco Trizzino s seminar March 25 th, 2016!
 Genetics Journal Club YoSon Park Feb 11, 2016 Announcement: Marco Trizzino s seminar March 25 th, 2016! Craniofacial development Implications in brain development, bipedal postures, etc Most craniofacial
Genetics Journal Club YoSon Park Feb 11, 2016 Announcement: Marco Trizzino s seminar March 25 th, 2016! Craniofacial development Implications in brain development, bipedal postures, etc Most craniofacial
V23: the histone code
 V23: the histone code The DNA of eukaryotic organisms is packaged into chromatin, whose basic repeating unit is the nucleosome. A nucleosome is formed by wrapping 147 base pairs of DNA twice around an
V23: the histone code The DNA of eukaryotic organisms is packaged into chromatin, whose basic repeating unit is the nucleosome. A nucleosome is formed by wrapping 147 base pairs of DNA twice around an
2/19/13. Contents. Applications of HMMs in Epigenomics
 2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
2/19/13 I529: Machine Learning in Bioinformatics (Spring 2013) Contents Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Background:
Applications of HMMs in Epigenomics
 I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
I529: Machine Learning in Bioinformatics (Spring 2013) Applications of HMMs in Epigenomics Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Background:
Bioinformatics of Transcriptional Regulation
 Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Bioinformatics of Transcriptional Regulation Carl Herrmann IPMB & DKFZ c.herrmann@dkfz.de Wechselwirkung von Maßnahmen und Auswirkungen Einflussmöglichkeiten in einem Dialog From genes to active compounds
Integrating diverse datasets improves developmental enhancer prediction
 1 Integrating diverse datasets improves developmental enhancer prediction Genevieve D. Erwin 1, Rebecca M. Truty 1, Dennis Kostka 2, Katherine S. Pollard 1,3, John A. Capra 4 1 Gladstone Institute of Cardiovascular
1 Integrating diverse datasets improves developmental enhancer prediction Genevieve D. Erwin 1, Rebecca M. Truty 1, Dennis Kostka 2, Katherine S. Pollard 1,3, John A. Capra 4 1 Gladstone Institute of Cardiovascular
What we ll do today. Types of stem cells. Do engineered ips and ES cells have. What genes are special in stem cells?
 Do engineered ips and ES cells have similar molecular signatures? What we ll do today Research questions in stem cell biology Comparing expression and epigenetics in stem cells asuring gene expression
Do engineered ips and ES cells have similar molecular signatures? What we ll do today Research questions in stem cell biology Comparing expression and epigenetics in stem cells asuring gene expression
Do engineered ips and ES cells have similar molecular signatures?
 Do engineered ips and ES cells have similar molecular signatures? Comparing expression and epigenetics in stem cells George Bell, Ph.D. Bioinformatics and Research Computing 2012 Spring Lecture Series
Do engineered ips and ES cells have similar molecular signatures? Comparing expression and epigenetics in stem cells George Bell, Ph.D. Bioinformatics and Research Computing 2012 Spring Lecture Series
GATCGTGCACGATCTCGGCAATTCGGGATGCCGGCTCGTCACCGGTCGCT
 Problem. (pts) A. (5pts) Your colleague professor Eugene Mathew Lateed generated a genome-wide DNA methylation map for normal colon cells using MRE-seq and MeDIP-seq. In an intergenic region, he found
Problem. (pts) A. (5pts) Your colleague professor Eugene Mathew Lateed generated a genome-wide DNA methylation map for normal colon cells using MRE-seq and MeDIP-seq. In an intergenic region, he found
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Characterization of Hi-C/CHi-C dynamics and enhancer identification. (a) Scatterplot of Hi-C read counts supporting contacts between domain boundaries. Contacts enclosing domains
Supplementary Figure 1 Characterization of Hi-C/CHi-C dynamics and enhancer identification. (a) Scatterplot of Hi-C read counts supporting contacts between domain boundaries. Contacts enclosing domains
Mapping of variable DNA methylation across multiple cell types defines a dynamic regulatory landscape of the human genome
 G3: Genes Genomes Genetics Early Online, published on February 7, 26 as doi:.534/g3.5.25437 INVESTIGATIONS Mapping of variable DNA methylation across multiple cell types defines a dynamic regulatory landscape
G3: Genes Genomes Genetics Early Online, published on February 7, 26 as doi:.534/g3.5.25437 INVESTIGATIONS Mapping of variable DNA methylation across multiple cell types defines a dynamic regulatory landscape
DNA:CHROMATIN INTERACTIONS
 DNA:CHROMATIN INTERACTIONS Exploring transcription factor binding and the epigenomic landscape Chris Seward Introductions Cell and Developmental Biology PhD Candidate in Dr. Lisa Stubbs Laboratory Currently
DNA:CHROMATIN INTERACTIONS Exploring transcription factor binding and the epigenomic landscape Chris Seward Introductions Cell and Developmental Biology PhD Candidate in Dr. Lisa Stubbs Laboratory Currently
ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System
 ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System Liyan Pang, Ph.D. Application Scientist 1 Topics to be Covered Introduction What is ChIP-qPCR? Challenges Facing Biological
ChampionChIP Quick, High Throughput Chromatin Immunoprecipitation Assay System Liyan Pang, Ph.D. Application Scientist 1 Topics to be Covered Introduction What is ChIP-qPCR? Challenges Facing Biological
CELL BIOLOGY - CLUTCH CH. 7 - GENE EXPRESSION.
 !! www.clutchprep.com CONCEPT: CONTROL OF GENE EXPRESSION BASICS Gene expression is the process through which cells selectively to express some genes and not others Every cell in an organism is a clone
!! www.clutchprep.com CONCEPT: CONTROL OF GENE EXPRESSION BASICS Gene expression is the process through which cells selectively to express some genes and not others Every cell in an organism is a clone
Next-generation sequencing technologies
 Next-generation sequencing technologies Illumina: Summary https://www.youtube.com/watch?v=fcd6b5hraz8 Illumina platforms: Benchtop sequencers https://www.illumina.com/systems/sequencing-platforms.html
Next-generation sequencing technologies Illumina: Summary https://www.youtube.com/watch?v=fcd6b5hraz8 Illumina platforms: Benchtop sequencers https://www.illumina.com/systems/sequencing-platforms.html
Chapter 17 Lecture. Concepts of Genetics. Tenth Edition. Regulation of Gene Expression in Eukaryotes
 Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Chapter 17 Lecture Concepts of Genetics Tenth Edition Regulation of Gene Expression in Eukaryotes Chapter Contents 17.1 Eukaryotic Gene Regulation Can Occur at Any of the Steps Leading from DNA to Protein
Supplementary Figure 2
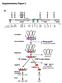 Supplementary Figure 2 a SBS-C1 SBS-C2 SBS-C3 SBS-C4 SBS-C5 SBS-C6 SBS-C7 SBS-C8 SBS-C9 LCR CNS-1 CNS-2 Il5 Rad50 Il13 Il4 Kif3a Sept8 0 50 100 150 200kb Sau3AI 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17
Supplementary Figure 2 a SBS-C1 SBS-C2 SBS-C3 SBS-C4 SBS-C5 SBS-C6 SBS-C7 SBS-C8 SBS-C9 LCR CNS-1 CNS-2 Il5 Rad50 Il13 Il4 Kif3a Sept8 0 50 100 150 200kb Sau3AI 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17
Role of polycomb proteins in gene transcription, stem cell and human diseases Dr. Luciano Di Croce
 Role of Polycomb Proteins in Gene Transcription, Stem Cells and Human Diseases 1 Luciano Di Croce, PhD Gene Regulation, Stem Cell and Cancer Department ICREA and Center for Genomic Regulation Barcelona
Role of Polycomb Proteins in Gene Transcription, Stem Cells and Human Diseases 1 Luciano Di Croce, PhD Gene Regulation, Stem Cell and Cancer Department ICREA and Center for Genomic Regulation Barcelona
The Role of Enhancers in Genetic and Epigenetic Control of Gene Expression
 The Role of Enhancers in Genetic and Epigenetic Control of Gene Expression J. Wesley Pike Department of Biochemistry University of Wisconsin-Madison, Madison, Wisconsin Encode Users Mee?ng-2016 Stanford
The Role of Enhancers in Genetic and Epigenetic Control of Gene Expression J. Wesley Pike Department of Biochemistry University of Wisconsin-Madison, Madison, Wisconsin Encode Users Mee?ng-2016 Stanford
Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup
 Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup 1. Introduction The data produced by IHEC is illustrated in Figure 1. Figure 1. The space of epigenomic
Data and Metadata Models Recommendations Version 1.2 Developed by the IHEC Metadata Standards Workgroup 1. Introduction The data produced by IHEC is illustrated in Figure 1. Figure 1. The space of epigenomic
Gene Regulation in Eukaryotes. Dr. Syahril Abdullah Medical Genetics Laboratory
 Gene Regulation in Eukaryotes Dr. Syahril Abdullah Medical Genetics Laboratory syahril@medic.upm.edu.my Lecture Outline 1. The Genome 2. Overview of Gene Control 3. Cellular Differentiation in Higher Eukaryotes
Gene Regulation in Eukaryotes Dr. Syahril Abdullah Medical Genetics Laboratory syahril@medic.upm.edu.my Lecture Outline 1. The Genome 2. Overview of Gene Control 3. Cellular Differentiation in Higher Eukaryotes
Value Correct Answer Feedback. Student Response. A. Dicer enzyme. complex. C. the Dicer-RISC complex D. none of the above
 1 RNA mediated interference is a post-transcriptional gene silencing mechanism Which component of the RNAi pathway have been implicated in cleavage of the target mrna? A Dicer enzyme B the RISC-siRNA complex
1 RNA mediated interference is a post-transcriptional gene silencing mechanism Which component of the RNAi pathway have been implicated in cleavage of the target mrna? A Dicer enzyme B the RISC-siRNA complex
MODULE TSS1: TRANSCRIPTION START SITES INTRODUCTION (BASIC)
 MODULE TSS1: TRANSCRIPTION START SITES INTRODUCTION (BASIC) Lesson Plan: Title JAMIE SIDERS, MEG LAAKSO & WILSON LEUNG Identifying transcription start sites for Peaked promoters using chromatin landscape,
MODULE TSS1: TRANSCRIPTION START SITES INTRODUCTION (BASIC) Lesson Plan: Title JAMIE SIDERS, MEG LAAKSO & WILSON LEUNG Identifying transcription start sites for Peaked promoters using chromatin landscape,
Integrating Diverse Datasets Improves Developmental Enhancer Prediction
 Integrating Diverse Datasets Improves Developmental Enhancer Prediction Genevieve D. Erwin 1,2, Nir Oksenberg 2,3, Rebecca M. Truty 1, Dennis Kostka 4, Karl K. Murphy 2,3, Nadav Ahituv 2,3, Katherine S.
Integrating Diverse Datasets Improves Developmental Enhancer Prediction Genevieve D. Erwin 1,2, Nir Oksenberg 2,3, Rebecca M. Truty 1, Dennis Kostka 4, Karl K. Murphy 2,3, Nadav Ahituv 2,3, Katherine S.
ANNOUNCEMENTS. HW2 is due Thursday 2/4 by 12:00 pm. Office hours: Monday 12:50 1:20 (ECCH 134)
 ANNOUNCEMENTS HW2 is due Thursday 2/4 by 12:00 pm. Office hours: Monday 12:50 1:20 (ECCH 134) Lectures 6 8 Outline Stem Cell Division - symmetric - asymmetric Stem cell lineage potential - pluripotent
ANNOUNCEMENTS HW2 is due Thursday 2/4 by 12:00 pm. Office hours: Monday 12:50 1:20 (ECCH 134) Lectures 6 8 Outline Stem Cell Division - symmetric - asymmetric Stem cell lineage potential - pluripotent
Epigenetics and DNase-Seq
 Epigenetics and DNase-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by Anthony
Epigenetics and DNase-Seq BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by Anthony
Go to Bottom Left click WashU Epigenome Browser. Click
 Now you are going to look at the Human Epigenome Browswer. It has a more sophisticated but weirder interface than the UCSC Genome Browser. All the data that you will view as tracks is in reality just files
Now you are going to look at the Human Epigenome Browswer. It has a more sophisticated but weirder interface than the UCSC Genome Browser. All the data that you will view as tracks is in reality just files
V8: Modelling the switching of cell fate
 V8: Modelling the switching of cell fate Aim : develop models for the architecture of coupled epigenetic and genetic networks which describe large changes in cellular identity (e.g., induction of pluripotency
V8: Modelling the switching of cell fate Aim : develop models for the architecture of coupled epigenetic and genetic networks which describe large changes in cellular identity (e.g., induction of pluripotency
The B cell identity factor Pax5 regulates distinct transcriptional programs in early and late B lymphopoiesis
 Manuscript EMBO-2011-80422 The B cell identity factor Pax5 regulates distinct transcriptional programs in early and late B lymphopoiesis Roger Revilla-i-Domingo, Ivan Bilic, Bojan Vilagos, Hiromi Tagoh,
Manuscript EMBO-2011-80422 The B cell identity factor Pax5 regulates distinct transcriptional programs in early and late B lymphopoiesis Roger Revilla-i-Domingo, Ivan Bilic, Bojan Vilagos, Hiromi Tagoh,
Bio5488 Practice Midterm (2018) 1. Next-gen sequencing
 1. Next-gen sequencing 1. You have found a new strain of yeast that makes fantastic wine. You d like to sequence this strain to ascertain the differences from S. cerevisiae. To accurately call a base pair,
1. Next-gen sequencing 1. You have found a new strain of yeast that makes fantastic wine. You d like to sequence this strain to ascertain the differences from S. cerevisiae. To accurately call a base pair,
Exploring Genome-wide Organization of Chromatin Structure by ChIP Alon Goren, Ph.D.
 Exploring Genome-wide Organization of Chromatin Structure by ChIP Alon Goren, Ph.D. Broad Institute of MIT & Harvard, MGH Pathology Harvard Medical School Webinar Overview Introduction to chromatin, ChIP
Exploring Genome-wide Organization of Chromatin Structure by ChIP Alon Goren, Ph.D. Broad Institute of MIT & Harvard, MGH Pathology Harvard Medical School Webinar Overview Introduction to chromatin, ChIP
The ChIP-Seq project. Giovanna Ambrosini, Philipp Bucher. April 19, 2010 Lausanne. EPFL-SV Bucher Group
 The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
The ChIP-Seq project Giovanna Ambrosini, Philipp Bucher EPFL-SV Bucher Group April 19, 2010 Lausanne Overview Focus on technical aspects Description of applications (C programs) Where to find binaries,
Complete Sample to Analysis Solutions for DNA Methylation Discovery using Next Generation Sequencing
 Complete Sample to Analysis Solutions for DNA Methylation Discovery using Next Generation Sequencing SureSelect Human/Mouse Methyl-Seq Kyeong Jeong PhD February 5, 2013 CAG EMEAI DGG/GSD/GFO Agilent Restricted
Complete Sample to Analysis Solutions for DNA Methylation Discovery using Next Generation Sequencing SureSelect Human/Mouse Methyl-Seq Kyeong Jeong PhD February 5, 2013 CAG EMEAI DGG/GSD/GFO Agilent Restricted
Epigenetic CHaracterization and Observation (ECHO) Proposers Day
 Epigenetic CHaracterization and Observation (ECHO) Proposers Day Eric Van Gieson, Ph.D. Program Manager Biological Technologies Office (BTO) February 23, 2018 ECHO Program Overview Determining Exposure
Epigenetic CHaracterization and Observation (ECHO) Proposers Day Eric Van Gieson, Ph.D. Program Manager Biological Technologies Office (BTO) February 23, 2018 ECHO Program Overview Determining Exposure
Stem Cel s Key Words:
 Stem Cells Key Words: Embryonic stem cells, Adult stem cells, ips cells, self-renewal, differentiation, pluripotent, multipotent, Inner cell mass, Nuclear transfer (Therapeutic cloning), Feeder cells,
Stem Cells Key Words: Embryonic stem cells, Adult stem cells, ips cells, self-renewal, differentiation, pluripotent, multipotent, Inner cell mass, Nuclear transfer (Therapeutic cloning), Feeder cells,
Control of Eukaryotic Gene Expression (Learning Objectives)
 Control of Eukaryotic Gene Expression (Learning Objectives) 1. Compare and contrast chromatin and chromosome: composition, proteins involved and level of packing. Explain the structure and function of
Control of Eukaryotic Gene Expression (Learning Objectives) 1. Compare and contrast chromatin and chromosome: composition, proteins involved and level of packing. Explain the structure and function of
Chapter 18. Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression 2007-2008 Control of Prokaryotic (Bacterial) Genes 2007- Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
Chapter 18 Regulation of Gene Expression 2007-2008 Control of Prokaryotic (Bacterial) Genes 2007- Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have
E i p g i e g n e e n t e i t c i s
 Epigenetics Introduction to Epigenetics Classical Genetics: The Central Dogma The Human Genome Project (HGP) Key findings: approx. 24,000 genes in human being all human races are 99.99% alike Challenges:
Epigenetics Introduction to Epigenetics Classical Genetics: The Central Dogma The Human Genome Project (HGP) Key findings: approx. 24,000 genes in human being all human races are 99.99% alike Challenges:
Lecture 14. ENCODE. Michael Schatz. March 15, 2017 JHU : Applied Comparative Genomics
 Lecture 14. ENCODE Michael Schatz March 15, 2017 JHU 601.749: Applied Comparative Genomics Project Proposal! Due March 15 HW6: Due March 29 *-seq in 4 short vignettes RNA-seq Methyl-seq ChIP-seq Hi-C Chromatin
Lecture 14. ENCODE Michael Schatz March 15, 2017 JHU 601.749: Applied Comparative Genomics Project Proposal! Due March 15 HW6: Due March 29 *-seq in 4 short vignettes RNA-seq Methyl-seq ChIP-seq Hi-C Chromatin
Whole Transcriptome Analysis of Illumina RNA- Seq Data. Ryan Peters Field Application Specialist
 Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Whole Transcriptome Analysis of Illumina RNA- Seq Data Ryan Peters Field Application Specialist Partek GS in your NGS Pipeline Your Start-to-Finish Solution for Analysis of Next Generation Sequencing Data
Il trascrittoma dei mammiferi
 29 Novembre 2005 Il trascrittoma dei mammiferi dott. Manuela Gariboldi Gruppo di ricerca IFOM: Genetica molecolare dei tumori (responsabile dott. Paolo Radice) Copyright 2005 IFOM Fondazione Istituto FIRC
29 Novembre 2005 Il trascrittoma dei mammiferi dott. Manuela Gariboldi Gruppo di ricerca IFOM: Genetica molecolare dei tumori (responsabile dott. Paolo Radice) Copyright 2005 IFOM Fondazione Istituto FIRC
Ebola therapy protects severely ill monkeys
 Propojení výuky oborů Molekulární a buněčné biologie a Ochrany a tvorby životního prostředí OPVK (CZ.1.07/2.2.00/28.0032) Ebola therapy protects severely ill monkeys Nature 514, 41 43 2 October 2014 Published
Propojení výuky oborů Molekulární a buněčné biologie a Ochrany a tvorby životního prostředí OPVK (CZ.1.07/2.2.00/28.0032) Ebola therapy protects severely ill monkeys Nature 514, 41 43 2 October 2014 Published
THE ANALYSIS OF CHROMATIN CONDENSATION STATE AND TRANSCRIPTIONAL ACTIVITY USING DNA MICROARRAYS 1. INTRODUCTION
 JOURNAL OF MEDICAL INFORMATICS & TECHNOLOGIES Vol.6/2003, ISSN 1642-6037 Piotr WIDŁAK *, Krzysztof FUJAREWICZ ** DNA microarrays, chromatin, transcription THE ANALYSIS OF CHROMATIN CONDENSATION STATE AND
JOURNAL OF MEDICAL INFORMATICS & TECHNOLOGIES Vol.6/2003, ISSN 1642-6037 Piotr WIDŁAK *, Krzysztof FUJAREWICZ ** DNA microarrays, chromatin, transcription THE ANALYSIS OF CHROMATIN CONDENSATION STATE AND
The Plant Cell, March 2013, American Society of Plant Biologists. All rights reserved.
 Epigenetics (TTPB4) Teaching Guide Overview Epigenetic modifications of DNA and chromatin affect the activity of genes and transposons. Epigenetic controls affect processes as diverse as time-of-flowering,
Epigenetics (TTPB4) Teaching Guide Overview Epigenetic modifications of DNA and chromatin affect the activity of genes and transposons. Epigenetic controls affect processes as diverse as time-of-flowering,
7. For What Molecule Do Genes Contain The
 7. For What Molecule Do Genes Contain The Instructions For Building For what molecule do genes contain the instructions for building? Are there many or few acetyl molecules attached to the genes associated
7. For What Molecule Do Genes Contain The Instructions For Building For what molecule do genes contain the instructions for building? Are there many or few acetyl molecules attached to the genes associated
Multi-omics in biology: integration of omics techniques
 31/07/17 Летняя школа по биоинформатике 2017 Multi-omics in biology: integration of omics techniques Konstantin Okonechnikov Division of Pediatric Neurooncology German Cancer Research Center (DKFZ) 2 Short
31/07/17 Летняя школа по биоинформатике 2017 Multi-omics in biology: integration of omics techniques Konstantin Okonechnikov Division of Pediatric Neurooncology German Cancer Research Center (DKFZ) 2 Short
Epigenetics, Environment and Human Health
 Epigenetics, Environment and Human Health A. Karim Ahmed National Council for Science and the Environment Washington, DC May, 2015 Epigenetics A New Biological Paradigm A Question about Cells: All cells
Epigenetics, Environment and Human Health A. Karim Ahmed National Council for Science and the Environment Washington, DC May, 2015 Epigenetics A New Biological Paradigm A Question about Cells: All cells
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Introduction to genome biology Lisa Stubbs We ve found most genes; but what about the rest of the genome? Genome size* 12 Mb 95 Mb 170 Mb 1500 Mb 2700 Mb 3200 Mb #coding genes ~7000 ~20000 ~14000 ~26000
Title: Genome-Wide Predictions of Transcription Factor Binding Events using Multi- Dimensional Genomic and Epigenomic Features Background
 Title: Genome-Wide Predictions of Transcription Factor Binding Events using Multi- Dimensional Genomic and Epigenomic Features Team members: David Moskowitz and Emily Tsang Background Transcription factors
Title: Genome-Wide Predictions of Transcription Factor Binding Events using Multi- Dimensional Genomic and Epigenomic Features Team members: David Moskowitz and Emily Tsang Background Transcription factors
Basic Concepts and History of Genetic Engineering. Mitesh Shrestha
 Basic Concepts and History of Genetic Engineering Mitesh Shrestha Genetic Engineering AKA gene manipulation, gene cloning, recombinant DNA technology, genetic modification, and the new genetics. A technique
Basic Concepts and History of Genetic Engineering Mitesh Shrestha Genetic Engineering AKA gene manipulation, gene cloning, recombinant DNA technology, genetic modification, and the new genetics. A technique
Chapter 11: Regulation of Gene Expression
 Chapter Review 1. It has long been known that there is probably a genetic link for alcoholism. Researchers studying rats have begun to elucidate this link. Briefly describe the genetic mechanism found
Chapter Review 1. It has long been known that there is probably a genetic link for alcoholism. Researchers studying rats have begun to elucidate this link. Briefly describe the genetic mechanism found
Applications of ChIP. November 05, David Grotsky, PhD Scientific Support Specialist - Epigenetics
 Applications of ChIP November 05, 2014 David Grotsky, PhD Scientific Support Specialist - Epigenetics Overview Introduction to chromatin and ChIP The histone code hypothesis What we learn from ChIP-on-chip
Applications of ChIP November 05, 2014 David Grotsky, PhD Scientific Support Specialist - Epigenetics Overview Introduction to chromatin and ChIP The histone code hypothesis What we learn from ChIP-on-chip
WORKSHOP. Transcriptional circuitry and the regulatory conformation of the genome. Ofir Hakim Faculty of Life Sciences
 WORKSHOP Transcriptional circuitry and the regulatory conformation of the genome Ofir Hakim Faculty of Life Sciences Chromosome conformation capture (3C) Most GR Binding Sites Are Distant From Regulated
WORKSHOP Transcriptional circuitry and the regulatory conformation of the genome Ofir Hakim Faculty of Life Sciences Chromosome conformation capture (3C) Most GR Binding Sites Are Distant From Regulated
Lecture 9. Eukaryotic gene regulation: DNA METHYLATION
 Lecture 9 Eukaryotic gene regulation: DNA METHYLATION Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation sequences Transcriptional
Lecture 9 Eukaryotic gene regulation: DNA METHYLATION Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation sequences Transcriptional
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017
 Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Functional Genomics Overview RORY STARK PRINCIPAL BIOINFORMATICS ANALYST CRUK CAMBRIDGE INSTITUTE 18 SEPTEMBER 2017 Agenda What is Functional Genomics? RNA Transcription/Gene Expression Measuring Gene
Gene Expression Microarrays. For microarrays, purity of the RNA was further assessed by
 Supplemental Methods Gene Expression Microarrays. For microarrays, purity of the RNA was further assessed by an Agilent 2100 Bioanalyzer. 500 ng of RNA was reverse transcribed into crna and biotin-utp
Supplemental Methods Gene Expression Microarrays. For microarrays, purity of the RNA was further assessed by an Agilent 2100 Bioanalyzer. 500 ng of RNA was reverse transcribed into crna and biotin-utp
Green Center Computational Core ChIP- Seq Pipeline, Just a Click Away
 Green Center Computational Core ChIP- Seq Pipeline, Just a Click Away Venkat Malladi Computational Biologist Computational Core Cecil H. and Ida Green Center for Reproductive Biology Science Introduc
Green Center Computational Core ChIP- Seq Pipeline, Just a Click Away Venkat Malladi Computational Biologist Computational Core Cecil H. and Ida Green Center for Reproductive Biology Science Introduc
Decoding Chromatin States with Epigenome Data Advanced Topics in Computa8onal Genomics
 Decoding Chromatin States with Epigenome Data 02-715 Advanced Topics in Computa8onal Genomics HMMs for Decoding Chromatin States Epigene8c modifica8ons of the genome have been associated with Establishing
Decoding Chromatin States with Epigenome Data 02-715 Advanced Topics in Computa8onal Genomics HMMs for Decoding Chromatin States Epigene8c modifica8ons of the genome have been associated with Establishing
Gene Regulation Biology
 Gene Regulation Biology Potential and Limitations of Cell Re-programming in Cancer Research Eric Blanc KCL April 13, 2010 Eric Blanc (KCL) Gene Regulation Biology April 13, 2010 1 / 21 Outline 1 The Central
Gene Regulation Biology Potential and Limitations of Cell Re-programming in Cancer Research Eric Blanc KCL April 13, 2010 Eric Blanc (KCL) Gene Regulation Biology April 13, 2010 1 / 21 Outline 1 The Central
Next Workshop on Epigenetic Profiling ChIP & MCIp
 Next Workshop on Epigenetic Profiling ChIP & MCIp 18-21st October 2010 Organized by the Division of Epigenomics and Cancer Risk Factors DKFZ Heidelberg, and Diagenode s training team Have you heard about
Next Workshop on Epigenetic Profiling ChIP & MCIp 18-21st October 2010 Organized by the Division of Epigenomics and Cancer Risk Factors DKFZ Heidelberg, and Diagenode s training team Have you heard about
HebbPlot: an intelligent tool for learning and visualizing chromatin mark signatures
 Girgis et al. BMC Bioinformatics (2018) 19:310 https://doi.org/10.1186/s12859-018-2312-1 SOFTWARE Open Access HebbPlot: an intelligent tool for learning and visualizing chromatin mark signatures Hani Z.
Girgis et al. BMC Bioinformatics (2018) 19:310 https://doi.org/10.1186/s12859-018-2312-1 SOFTWARE Open Access HebbPlot: an intelligent tool for learning and visualizing chromatin mark signatures Hani Z.
Edexcel (B) Biology A-level
 Edexcel (B) Biology A-level Topic 7: Modern Genetics Notes Using Gene Sequencing Genome = all of an organism s DNA, including mitochondrial/chloroplast DNA. Polymerase chain reaction (PCR) is used to amplify
Edexcel (B) Biology A-level Topic 7: Modern Genetics Notes Using Gene Sequencing Genome = all of an organism s DNA, including mitochondrial/chloroplast DNA. Polymerase chain reaction (PCR) is used to amplify
Einführung in die Genetik
 Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
Reads to Discovery. Visualize Annotate Discover. Small DNA-Seq ChIP-Seq Methyl-Seq. MeDIP-Seq. RNA-Seq. RNA-Seq.
 Reads to Discovery RNA-Seq Small DNA-Seq ChIP-Seq Methyl-Seq RNA-Seq MeDIP-Seq www.strand-ngs.com Analyze Visualize Annotate Discover Data Import Alignment Vendor Platforms: Illumina Ion Torrent Roche
Reads to Discovery RNA-Seq Small DNA-Seq ChIP-Seq Methyl-Seq RNA-Seq MeDIP-Seq www.strand-ngs.com Analyze Visualize Annotate Discover Data Import Alignment Vendor Platforms: Illumina Ion Torrent Roche
Distinct epigenetic features of differentiation regulated replication origins
 DOI 10.1186/s13072-016-0067-3 Epigenetics & Chromatin RESEARCH Open Access Distinct epigenetic features of differentiation regulated replication origins Owen K. Smith 1, RyanGuk Kim 2, Haiqing Fu 1, Melvenia
DOI 10.1186/s13072-016-0067-3 Epigenetics & Chromatin RESEARCH Open Access Distinct epigenetic features of differentiation regulated replication origins Owen K. Smith 1, RyanGuk Kim 2, Haiqing Fu 1, Melvenia
Section C: The Control of Gene Expression
 Section C: The Control of Gene Expression 1. Each cell of a multicellular eukaryote expresses only a small fraction of its genes 2. The control of gene expression can occur at any step in the pathway from
Section C: The Control of Gene Expression 1. Each cell of a multicellular eukaryote expresses only a small fraction of its genes 2. The control of gene expression can occur at any step in the pathway from
PhysicsAndMathsTutor.com. Question Number. Answer Additional guidance Mark. 1(a) 1. reference to stem cells being {totipotent / pluripotent} ;
 1(a) 1. reference to stem cells being {totipotent / pluripotent} ; 2. can specialise or differentiate / can give rise to {differentiated / specialised} cells ; 3. idea that these can replace damaged cells
1(a) 1. reference to stem cells being {totipotent / pluripotent} ; 2. can specialise or differentiate / can give rise to {differentiated / specialised} cells ; 3. idea that these can replace damaged cells
Experimental validation of candidates of tissuespecific and CpG-island-mediated alternative polyadenylation in mouse
 Karin Fleischhanderl; Martina Fondi Experimental validation of candidates of tissuespecific and CpG-island-mediated alternative polyadenylation in mouse 108 - Biotechnologie Abstract --- Keywords: Alternative
Karin Fleischhanderl; Martina Fondi Experimental validation of candidates of tissuespecific and CpG-island-mediated alternative polyadenylation in mouse 108 - Biotechnologie Abstract --- Keywords: Alternative
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali
 Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
Le proteine regolative variano nei vari tipi cellulari e in funzione degli stimoli ambientali Tipo cellulare 1 Tipo cellulare 2 Tipo cellulare 3 DNA-protein Crosslink Lisi Frammentazione Immunopurificazione
Einführung in die Genetik
 Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Twitter: @PlantDevTUM, #genetiktum FB: Plant Development TUM Prof. Dr. Claus Schwechheimer
Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Twitter: @PlantDevTUM, #genetiktum FB: Plant Development TUM Prof. Dr. Claus Schwechheimer
Frumkin, 2e Part 1: Methods and Paradigms. Chapter 6: Genetics and Environmental Health
 Frumkin, 2e Part 1: Methods and Paradigms Chapter 6: Genetics and Environmental Health Genetics Genetics, the study of individual genes, has expanded to include genomics, which is the study of all the
Frumkin, 2e Part 1: Methods and Paradigms Chapter 6: Genetics and Environmental Health Genetics Genetics, the study of individual genes, has expanded to include genomics, which is the study of all the
Bi8 Lecture 19. Review and Practice Questions March 8th 2016
 Bi8 Lecture 19 Review and Practice Questions March 8th 2016 Common Misconceptions Where Words Matter Common Misconceptions DNA vs RNA vs Protein Su(H) vs Su(H) Replication Transcription Transcribed vs
Bi8 Lecture 19 Review and Practice Questions March 8th 2016 Common Misconceptions Where Words Matter Common Misconceptions DNA vs RNA vs Protein Su(H) vs Su(H) Replication Transcription Transcribed vs
Hitting the mark: specificity analysis of histone antibodies
 APPLICATION NOTE Histone modification antibodies Hitting the mark: specificity analysis of histone antibodies Introduction The nucleosome, composed of the histones H2A, H2B, H3, and H4, is the fundamental
APPLICATION NOTE Histone modification antibodies Hitting the mark: specificity analysis of histone antibodies Introduction The nucleosome, composed of the histones H2A, H2B, H3, and H4, is the fundamental
Infiltrating immune cells do not differ between T2E and non-t2e samples and represent a small fraction of total cellularity.
 Supplementary Figure 1 Infiltrating immune cells do not differ between T2E and non-t2e samples and represent a small fraction of total cellularity. (a) ESTIMATE immune score from mrna expression array
Supplementary Figure 1 Infiltrating immune cells do not differ between T2E and non-t2e samples and represent a small fraction of total cellularity. (a) ESTIMATE immune score from mrna expression array
Supplementary Information. Supplementary Figures
 Fib skin 01 Fib skin 02 Fib skin 03 Ker skin 01 Ker skin 02 Ker skin 03 Mel skin 01 Mel skin 02 Mel skin 03 Fib skin 01 Fib skin 02 Fib skin 03 Ker skin 01 Ker skin 02 Ker skin 03 Mel skin 01 Mel skin
Fib skin 01 Fib skin 02 Fib skin 03 Ker skin 01 Ker skin 02 Ker skin 03 Mel skin 01 Mel skin 02 Mel skin 03 Fib skin 01 Fib skin 02 Fib skin 03 Ker skin 01 Ker skin 02 Ker skin 03 Mel skin 01 Mel skin
Genome 541 Gene regulation and epigenomics Lecture 3 Integrative analysis of genomics assays
 Genome 541 Gene regulation and epigenomics Lecture 3 Integrative analysis of genomics assays Please consider both the forward and reverse strands (i.e. reverse compliment sequence). You do not need to
Genome 541 Gene regulation and epigenomics Lecture 3 Integrative analysis of genomics assays Please consider both the forward and reverse strands (i.e. reverse compliment sequence). You do not need to
a previous report, Lee and colleagues showed that Oct4 and Sox2 bind to DXPas34 and Xite
 SUPPLEMENTARY INFORMATION doi:1.138/nature9496 a 1 Relative RNA levels D12 female ES cells 24h Control sirna 24h sirna Oct4 5 b,4 Oct4 Xist (wt) Xist (mut) Tsix 3' (wt) Tsix 5' (wt) Beads Oct4,2 c,2 Xist
SUPPLEMENTARY INFORMATION doi:1.138/nature9496 a 1 Relative RNA levels D12 female ES cells 24h Control sirna 24h sirna Oct4 5 b,4 Oct4 Xist (wt) Xist (mut) Tsix 3' (wt) Tsix 5' (wt) Beads Oct4,2 c,2 Xist
Non-conserved intronic motifs in human and mouse are associated with a conserved set of functions
 Non-conserved intronic motifs in human and mouse are associated with a conserved set of functions Aristotelis Tsirigos Bioinformatics & Pattern Discovery Group IBM Research Outline. Discovery of DNA motifs
Non-conserved intronic motifs in human and mouse are associated with a conserved set of functions Aristotelis Tsirigos Bioinformatics & Pattern Discovery Group IBM Research Outline. Discovery of DNA motifs
Nature Genetics: doi: /ng Supplementary Figure 1. ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets.
 Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
ChIP-seq/Functional Genomics/Epigenomics. CBSU/3CPG/CVG Next-Gen Sequencing Workshop. Josh Waterfall. March 31, 2010
 ChIP-seq/Functional Genomics/Epigenomics CBSU/3CPG/CVG Next-Gen Sequencing Workshop Josh Waterfall March 31, 2010 Outline Introduction to ChIP-seq Control data sets Peak/enriched region identification
ChIP-seq/Functional Genomics/Epigenomics CBSU/3CPG/CVG Next-Gen Sequencing Workshop Josh Waterfall March 31, 2010 Outline Introduction to ChIP-seq Control data sets Peak/enriched region identification
ChIP-Seq Tools. J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015
 ChIP-Seq Tools J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA or
ChIP-Seq Tools J Fass UCD Genome Center Bioinformatics Core Wednesday September 16, 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA or
Escape from epigenetic silencing of lactase gene expression triggered by a single nucleotide change
 Escape from epigenetic silencing of lactase gene expression triggered by a single nucleotide change Dallas M. Swallow and Jesper T. Troelsen The importance of subtle gene regulation and epigenetics in
Escape from epigenetic silencing of lactase gene expression triggered by a single nucleotide change Dallas M. Swallow and Jesper T. Troelsen The importance of subtle gene regulation and epigenetics in
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday 15 June 2015 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind DNA
The ENCODE Encyclopedia. & Variant Annotation Using RegulomeDB and HaploReg
 The ENCODE Encyclopedia & Variant Annotation Using RegulomeDB and HaploReg Jill E. Moore Weng Lab University of Massachusetts Medical School October 10, 2015 Where s the Encyclopedia? ENCODE: Encyclopedia
The ENCODE Encyclopedia & Variant Annotation Using RegulomeDB and HaploReg Jill E. Moore Weng Lab University of Massachusetts Medical School October 10, 2015 Where s the Encyclopedia? ENCODE: Encyclopedia
MCDB 1041 Class 21 Splicing and Gene Expression
 MCDB 1041 Class 21 Splicing and Gene Expression Learning Goals Describe the role of introns and exons Interpret the possible outcomes of alternative splicing Relate the generation of protein from DNA to
MCDB 1041 Class 21 Splicing and Gene Expression Learning Goals Describe the role of introns and exons Interpret the possible outcomes of alternative splicing Relate the generation of protein from DNA to
NimbleGen Arrays and LightCycler 480 System: A Complete Workflow for DNA Methylation Biomarker Discovery and Validation.
 Cancer Research Application Note No. 9 NimbleGen Arrays and LightCycler 480 System: A Complete Workflow for DNA Methylation Biomarker Discovery and Validation Tomasz Kazimierz Wojdacz, PhD Institute of
Cancer Research Application Note No. 9 NimbleGen Arrays and LightCycler 480 System: A Complete Workflow for DNA Methylation Biomarker Discovery and Validation Tomasz Kazimierz Wojdacz, PhD Institute of
APPLICATION NOTE. Abstract. Introduction
 From minuscule amounts to magnificent results: reliable ChIP-seq data from 1, cells with the True MicroChIP and the MicroPlex Library Preparation kits Abstract Diagenode has developed groundbreaking solutions
From minuscule amounts to magnificent results: reliable ChIP-seq data from 1, cells with the True MicroChIP and the MicroPlex Library Preparation kits Abstract Diagenode has developed groundbreaking solutions
ChIP-Seq Data Analysis. J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014
 ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
ChIP-Seq Data Analysis J Fass UCD Genome Center Bioinformatics Core Wednesday December 17, 2014 What s the Question? Where do Transcription Factors (TFs) bind genomic DNA 1? (Where do other things bind
Computational Challenges of Medical Genomics
 Talk at the VSC User Workshop Neusiedl am See, 27 February 2012 [cbock@cemm.oeaw.ac.at] http://medical-epigenomics.org (lab) http://www.cemm.oeaw.ac.at (institute) Introducing myself to Vienna s scientific
Talk at the VSC User Workshop Neusiedl am See, 27 February 2012 [cbock@cemm.oeaw.ac.at] http://medical-epigenomics.org (lab) http://www.cemm.oeaw.ac.at (institute) Introducing myself to Vienna s scientific
The Nucleus, DNA and Chromatin Structure
 The Nucleus, DNA and Chromatin Structure Size of the human genom: The diploid human genome contains approximately 6 billion base pairs of DNA packaged into 23 chromosome pairs. Each diploid cell therefore
The Nucleus, DNA and Chromatin Structure Size of the human genom: The diploid human genome contains approximately 6 billion base pairs of DNA packaged into 23 chromosome pairs. Each diploid cell therefore
Advanced Cell Biology. Lecture 17
 Advanced Cell Biology. Lecture 17 Alexey Shipunov Minot State University February 25, 2013 Shipunov (MSU) Advanced Cell Biology. Lecture 17 February 25, 2013 1 / 29 Outline Questions and answers DNA DNA
Advanced Cell Biology. Lecture 17 Alexey Shipunov Minot State University February 25, 2013 Shipunov (MSU) Advanced Cell Biology. Lecture 17 February 25, 2013 1 / 29 Outline Questions and answers DNA DNA
Introduction to genome biology
 Introduction to genome biology Lisa Stubbs Deep transcritpomes for traditional model species from ENCODE (and modencode) Deep RNA-seq and chromatin analysis on 147 human cell types, as well as tissues,
Introduction to genome biology Lisa Stubbs Deep transcritpomes for traditional model species from ENCODE (and modencode) Deep RNA-seq and chromatin analysis on 147 human cell types, as well as tissues,
Reprogramming of the Genome by Toxic Injury Ken Ramos, MD, PhD, ATS
 Reprogramming of the Genome by Toxic Injury Ken Ramos, MD, PhD, ATS Professor of dicine, University of Arizona Health Sciences Center & Associate Vice President for Precision Health Sciences Office of
Reprogramming of the Genome by Toxic Injury Ken Ramos, MD, PhD, ATS Professor of dicine, University of Arizona Health Sciences Center & Associate Vice President for Precision Health Sciences Office of
