Programmable RNA microstructures for coordinated delivery of sirnas
|
|
|
- Osborn Jonathan Hunt
- 6 years ago
- Views:
Transcription
1 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2016 Programmable RNA microstructures for coordinated delivery of sirnas Jaimie Marie Stewart, Mathias Viard, Hari K.K. Subramanian, Brandon K. Roark Kirill A. Afonin, and Elisa Franco Department of Bioengineering, University of California, Riverside, Riverside, CA 92521, USA. Basic Science Program, Leidos Biomedical Research, Frederick National Laboratory for Cancer Research, Frederick, MD 21702, USA. Department of Mechanical Engineering, University of California, Riverside, Riverside, CA 92521, USA. Nanoscale Science Program, Department of Chemistry University of North Carolina at Charlotte, Charlotte NC 28223, USA; The Center for Biomedical Engineering and Science, University of North Carolina at Charlotte, Charlotte NC 28223, USA.
2 Contents 1. Sequences 1.1 Genes 1.2 Genes for tiles with blunt ends 1.3 RNA sequences 2. Methods for preparation and analysis of RNA assemblies 2.1 Sample preparation 2.2 Atomic force Microscopy 2.3 Denaturing polyacrylamide gel electrophoresis 2.4 Non-denaturing polyacrylamide gel electrophoresis 2.5 RNA extraction 3. Tile and lattice purification 4. Structural analysis of tile and lattice assembly 5. Analysis of structural integrity with Lipofectamine Dynamic Light Scattering (DLS) experiments 7. Human blood serum stability assays 8. UV-melting experiments 9. Transfection experiments 10. Flow cytometry 11. Confocal microscopy of transfected cells 12. Gene silencing experiments 13. Viability assay 14. Effects of the presence of functional domains on assembly
3 1. Sequences RNA sequences for tile without sirna were identified by converting to RNA the (44+6) tile design from Ko et al. 1, as described by Stewart et al. 2. PAGE purified DNA sequences were ordered from IDT DNA (Coralville, IA, USA). Genes include 3 base sealing domains (5 end of non-template strand) to prevent breathing at the 5 end of the promoter site. All genes include the T7 promoter sequence. The +1 to +6 promoter region (transcription start site) was modified relative to the generally recommended sequence (GGGAGA) 3, to avoid altering or constraining the sequence content of RNA tiles. 1.1 Genes Strand Sa Sb Sc Sc-GFP Sc-PLK1 non template: 5 - TTC TAA TAC GAC TCA CTA TAG GTG CGA CTA TGC AAC CTG CCT GGC AAG ACC TAC GAT GGA CAC GGT AAC G -3, template: 5 - CGT TAC CGT GTC CAT CGT AGG TCT TGC CAG GCA GGT TGC ATA GTC GCA CCT ATA GTG AGT CGT ATT AGA A -3 non template: 5 - TTC TAA TAC GAC TCA CTA TAG TCT TGC CAG GCA CCA TCG TAG GTC TTG CCA GGC ACC ATC GTA G -3, template: 5 - CTA CGA TGG TGC CTG GCA AGA CCT ACG ATG GTG CCT GGC AAG ACT ATA GTG AGT CGT ATT AGA A -3 non template: 5 - TTC TAA TAC GAC TCA CTA TA GCA CCC GTT ACC GTG TGG TTG CAT AGT C -3, template: 5 - GAC TAT GCA ACC ACA CGG TAA CGG GTG CTA TAG TGA GTC GTA TTA GAA -3 non template: 5 - TTC TAA TAC GAC TCA CTA TAG CAC CCG TTA CCG TGT GGT TGC ATA GTC TTC GGT GGT GCA GAT GAA CTT CAG GGT CA -3, template: 5 - TGA CCC TGA AGT TCA TCT GCA CCA CCG AAG ACT ATG CAA CCA CAC GGT AAC GGG TGC TAT AGT GAG TCG TAT TAG AA 3 non template: 5'- TTC TAA TAC GAC TCA CTA TAG CAC CCG TTA CCG TGT GGT TGC ATA GTC TTC GTC ATT AAG CAG CTC GTT AAT GGT T -3, template: 5'- AAC CAT TAA CGA GCT GCT TAA TGA CGA AGA CTA TGC AAC CAC ACG GTA ACG GGT GCT ATA GTG AGT CGT ATT AGA A Genes for tiles with blunt ends Strand
4 Sa Sb Sc Sc-GFP Sc-PLK1 non template: 5 - TTC TAA TAC GAC TCA CTA TAG ACT ATG CAA CCT GCC TGG CAA GAC CTA CGA TGG ACA CGG TAA CG -3, template: 5 - CGT TAC CGT GTC CAT CGT AGG TCT TGC CAG GCA GGT TGC ATA GTC TAT AGT GAG TCG TAT TAG AA -3 non template: 5 - TTC TAA TAC GAC TCA CTA TAG TCT TGC CAG GCA CCA TCG TAG GTC TTG CCA GGC ACC ATC GTA G -3, template: 5 - CTA CGA TGG TGC CTG GCA AGA CCT ACG ATG GTG CCT GGC AAG ACT ATA GTG AGT CGT ATT AGA A -3 non template: 5 - TTC TAA TAC GAC TCA CTA TAC GTT ACC GTG TGG TTG CAT AGT C -3, template: 5 - GAC TAT GCA ACC ACA CGG TAA CGT ATA GTG AGT CGT ATT AGA A -3 non template: 5 - TTC TAA TAC GAC TCA CTA TAC GTT ACC GTG TGG TTG CAT AGT CTT CGG TGG TGC AGA TGA ACT TCA GGG TCA 3, template: 5 - TGA CCC TGA AGT TCA TCT GCA CCA CCG AAG ACT ATG CAA CCA CAC GGT AAC GTA TAG TGA GTC GTA TTA GAA 3 non template: 5'- TTC TAA TAC GAC TCA CTA TAC GTT ACC GTG TGG TTG CAT AGT CTT CGT CAT TAA GCA GCT CGT TAA TGG TT -3, template: 5'- AAC CAT TAA CGA GCT GCT TAA TGA CGA AGA CTA TGC AAC CAC ACG GTA ACG TAT AGT GAG TCG TAT TAG AA RNA sequences Strand Sa 5 - GGU GCG ACU AUG CAA CCU GC CUG GCA AGA CCU ACG AUG GAC ACG GUA ACG -3 Sb 5 - GUC UUG CCA GGC ACC AUC GUA GGU CUU GCC AGG CAC CAU CGU AG -3 Sc 5 - GCA CCC GUU ACC GUG UGG UUG CAU AGU C -3 Sc-GFP 5 GCA CCC GUU ACC GUG UGG UUG CAU AGU Cuu CGG UGG UGC AGA UGA ACU UCA GGG UCA 3 Sc-PLK1 5 GCA CCC GUU ACC GUG UGG UUG CAU AGU Cuu CGU CAU UAA GCA GCU CGU UAA UGG UU 3 GFP sense 5 pacc CUG AAG UUC AUC UGC ACC ACC G 3 PLK1 sense 5'- paac CAU UAA CGA GCU GCU UAA UGA CG -3' GFP sense 5 /5AlexF488N/ACC CUG AAG UUC AUC UGC ACC ACC G 3 labeled with Alexa488 PLK1 sense 5 /5AlexF546N/ CCA TTA ACG AGC TGC TTA ATG ACG 3 labeled with Alexa Methods 2.1 Sample preparation We prepared RNA nanostructures by slowly annealing extracted and purified RNA. RNA
5 strands were transcribed; gel extracted and purified using the RNA extraction protocol. Strands were added in a 2:1:2:2 ratio (Sa:Sb:Sc-antisense:Sense) (Figure S1) to a solution of 1X tris-acetate-edta (TAE) and 12.5 mm MgCl 2. The solution was then annealed from 70 C to 22 C over 24 hours. 2.2 Atomic force microscopy AFM images were obtained in tapping mode using 1X TAE and 12.5 mm MgCl 2 buffer with a Digital Instruments Multimode AFM, equipped with a Nanoscope III controller. Sharp Nitride Lever (SNL) tips from Bruker with a nominal spring constant of 0.24 N/m were used for imaging, with a drive frequency of 9-10 khz. 2.3 Denaturing polyacrylamide gel electrophoresis Pre-mix was prepared by adding 42 g of urea to 25 ml of nanopure water (for a final volume of 100 ml), the mixture was then heated until the urea completely dissolved. This mixture was allowed to cool to room temperature, then a 40% (v/v) 19:1 acrylamide/bis-acrylamide solution was added in the appropriate volume for the desired percentage. The pre-mix was added in appropriate ratios with TBE and nanopure water, ammonium persulfate (APS), and tetramethylethylenediamine (TEMED) to start polymerization. Gels were cast in cm, 1 mm thick disposable mini gel cassettes (Thermo Scientific, #NC2010) and allowed to polymerize for at least 2 hours before electrophoresis. Gels were run at room temperature at 100 V in 1X TBE unless otherwise noted. After electrophoresis the gels were stained in SYBR Gold Nucleic Acid Gel Stain or ethidium bromide and then imaged using the Bio-Rad ChemiDoc MP system. 2.4 Non-denaturing polyacrylamide gel electrophoresis 40% solution of 19:1 acrylamide/bis-acrylamide, TAE, MgCl mm (final concentration), APS, and TEMED were added together at appropriate concentrations for the desired polyacrylamide percentage, then cast in cm, 1 mm thick disposable mini gel cassettes (Thermo Scientific, #NC2010) and allowed to polymerize for at least 2 hours before electrophoresis. Gels were run at 4 C at 150 V in 1X TBE buffer. After electrophoresis gels were stained in SYBR Gold Nucleic Acid Gel Stain or ethidium bromide and then imaged using the Bio-Rad ChemiDoc MP system. 2.5 RNA extraction Using the AmpliScribe-T7-Flash Transcription Kit the following components were mixed at room temperature: RNase-free water, µg gene, AmpliScribe T7-Flash 1X Reaction Buffer (Epicentre, #ASF3507), 9 mm NTPs, 40 U/µL RiboGuard RNase Inhibitor (Epicentre, #RG90925), and 1 unit of AmpliScribe T7-Flash Enzyme Solution (Epicentre, #ASF3507). This mix was incubated at 37 C for 4 hours. After incubation, 20 µl of loading dye was added to the 20 µl transcription solution, and 8 µl of the transcription/dye mix was added to each of the middle 5 lanes of the gel. The gel was run at 100 V at room temperature in 1X TBE. After electrophoresis the gel was stained in 80 ml 1X TBE and 1 µl SYBR Gold Nucleic Acid Gel Stain (Thermo Scientific, # S-11494) for minutes. The gel was then placed on a TLC silica gel 60 W F254S aluminum sheet (EMD Millipore, #
6 ) covered in plastic wrap. The gel was illuminated with UV light and the desired RNA band was cut out and chopped into equal amounts and placed into 0.5 ml DNA Lobind tubes (Eppendorf, # ), then 200 µl of 0.3 M sodium acetate at ph 5.3 was added to the Lobind tubes. The samples were then incubated at 42 C for ~20 hours. After incubation, the sodium acetate was removed and placed into 1.7 ml RNase/DNase free tubes. The old Lobind tubes were rinsed with 100 µl of 0.3 M sodium acetate ph 5.3, which was added to the new samples. Then 1 ml of freezer cold 100% ETOH and 1 µl of glycogen were added into each tube and the sample was incubated at -20 C for 20 hours. Next, the samples were spun at 13,500 rpm at 4 C for 15 minutes. The white precipitate pellet (RNA) at the bottom of the tube was located and the supernatant was carefully pipetted out of the tubes avoiding removal of the pellet. Then, 500 µl of 70% freezer cold ETOH was added to the tubes and spun at 13,500 rpm at 4 C for 5 minutes. The supernatant was carefully decanted again with a pipette. This washing procedure was repeated a third time. After the last wash, as much supernatant was removed as possible, then the samples were opened and placed in the vacuum concentrator and allowed to spin at room temperature for 15 minutes. The samples were then re-suspended in µl of Ambion nuclease free water (Ambion, #AM9932). 3. Tile and lattice purification Structures were annealed in 1X TAE / 12.5 mm MgCl 2 by adding strands in proper stoichiometry from 70 C to room temperature over 24 hours. Structures were then electrophoresed through 15% non-denaturing PAGE ran at 150 V, at 4 C in 1X TBE. For each sample the correct bands, which incorporated the sense strand into the tile or lattice was extracted. 3-4 bands were placed into one tube and covered in 90 µl of 1X TAE / 12.5 mm MgCl 2 and allowed to incubate at 4 C for hours. 4. Structural analysis of tile and lattice assembly To verify the sense strand attachment to the tile, the sense strand for both GFP and PLK1 were labeled with Alexa 488 and Alexa 546, respectively at the 5 ends. We annealed individual tiles without sticky ends and lattices and tubular structures with fluorescently labeled sense strands. The sense strand targeted against GFP was labeled with Alexa 488 and the sense strand targeted against PLK1 was labeled with Alexa 546. We performed non-denaturing PAGE studies on these structures and tracked the labeled sense strand to determine its location in the tile and lattice (Figure S2). Interestingly, for the individual tiles we observed three prominent bands, which incorporated all of the tile strands (Figure S2A, S2B). We hypothesize that the upper band has two sense strands attached, the next lower band has only one sense attached, and the third lowest band has no sense strands attached. For individual tile assembly with no sticky ends, there appears to be a small amount of tiles that polymerize at the top of the well, this could be due to the anti-sense strands interacting. For the lanes including annealed tiles, we observe a fairly prominent band at the bottom of the wells, which we expect is composed by tiles polymerized into lattices.
7 5. Analysis of structural integrity with Lipofectamine 2000 To ensure that our structures remain intact during transfection, we performed experiments to determine the integrity of our RNA structures in the presence of Lipofectamine 2000 (L2K, Thermo Scientific # ) (Figure S3). Four samples were prepared: Sa+Sb (1:1), Sa+Sb (2:1), and Sa+Sb+Sc (2:1:2) were slowly annealed from 70 C to 22 C over 24 hr in 1X TAE and 12.5 mm MgCl µl of sample was added to 1 µl of L2K and allowed to incubate at room temperature for 30 minutes, then non-denaturing PAGE was performed with 10% gel at 150 V at 4 C in 1X TBE (Figure S2B). Next 1 µl of Triton X100 (Sigma Aldrich #X100) was added to the sample to remove L2K, and allowed to incubate at room temperature for 30 minutes. Then, nondenaturing PAGE was performed with 10% gel at 150 V at 4 C in 1X TBE (Figure S3C). Because L2K is positively charged, DNA associated with is unable to travel through the electrophoresis gel properly. However, when the detergent is introduced and L2K is removed, structures can migrate through the gel. Based on the consistent height of the band in lane 4 of the gels in Fig. 2A and 2C, we conclude that minimum to no damage occurs to our structures when associated to L2K. 6. Dynamic Light Scattering (DLS) experiments For DLS experiments, 100 µl of sample solutions (prepared in 0.2 µm filtered buffer) of assembled 1 µm tiles and lattices were used. The samples were measured at 25 C with a Nano-series Zetasizer ZS (Malvern Instruments Ltd) equipped with a red (633 nm) laser. Readings with polydispersity index (PdI) below 0.2 were used. Results are shown in Figure S4. 7. Human blood serum stability assays Freshly drawn human whole blood was spun down and serum was collected, aliquoted, frozen -80 C and later used for blood serum degradation experiments. Alexa-488 labeled tiles and lattices (1 µm final) were incubated with 10% (v/v) human blood serum at 37 C for various time points. Degradation time courses were quenched on dry ice and analyzed on native-page, loaded in reverse order. The disappearance of fluorescent bands corresponding to the labeled tiles and lattices were quantified and analyzed. Results are shown in Figure S5. 8. UV-melting experiments. Experiments were carried out at 260 nm on an Agilent 8453 spectrophotometer coupled with the Agilent Peltier Temperature Controller. The temperature was gradually increased and the absorbance was recorded every 1 C with 10 seconds equilibration time. The assembled RNA tiles and lattices (70 µl of 250 nm) were analyzed using Starna Cells sub-micro quartz fluorometer cell cuvette. The data was processed by a Boltzmann sigmoidal curve fit using Origin Pro 2016 Graphing and Analysis software. Results are shown in Figure S6. 9. Transfection experiments To evaluate the delivery of functionalized RNA tiles and lattices, the human breast cancer cell line MDA-MB-231 (with or without GFP) was grown in D-MEM medium
8 (Gibco BRL) supplemented with 10% FBS and penicillin-streptomycin (pen-strep) in a 5% CO 2 incubator. All transfections were performed using Lipofectamine 2000 (L2K) purchased from Invitrogen. 100X transfection solutions were pre-incubated at room temperature with L2K. Prior to each transfection, the cell media was replaced with OPTI-MEM with added RNA/L2K complexes at a final concentration of 1X. The cells were incubated for 4 hours followed by the media change (D-MEM, 10%FCS, 1% penstrep). Results are shown in Figure S Flow cytometry For flow cytometry experiments, MDA-MB-231 cells (with or without GFP) grown in 24- well plates were transfected. The cells were lifted with cell dissociation buffer 24h later to assess their level of Alexa 488 (Alexa 546) fluorescence or 72h later to assess their GFP signal for uptake and silencing experiments respectively. The measurements were performed through fluorescence activated cell sorting on a FACSCalibur instrument (BD Biosciences, San Jose). At least 20,000 events were collected and analyzed with the CellQuest software to retrieve the geometric mean fluorescence intensity (gmfi) and the standard error of the mean. 11. Confocal microscopy of transfected cells All experiments were performed using a LSM 710 confocal microscope (Carl Zeiss, Oberkochen) and a Plan-Apochromat 63x/1.40 Oil lens. MDA-MB-231 cells plated in glass bottom dishes (Ibidi, Madison) were transfected with fluorescently labeled nanoconstructs. Upon 4 hour incubation at 37 C, the cells were washed three times with PBS and fixed with 4% paraformaldehyde for 20 minutes at room temperature. For Alexa 488 imaging, the 488 nm line of an argon laser was used as excitation and the emission was collected between 493 and 557 nm. For Alexa 546 imaging, a DPSS 561 laser was used for excitation and emission was collected between 566 and 680 nm. For silencing experiments, MDA-MB-231 egfp cells were used and visualized 72 hours post transfection upon fixation. For GFP imaging, the 488 nm line of an argon laser was used as excitation and the emission was collected between 499 and 644 nm. 12. Gene silencing experiments To assess the function of sirnas, we performed specific gene silencing experiments. GFP expressing breast cancer cells (MDA-MB-231/GFP) were transfected as mentioned in the Transfection experiments section with constructs carrying anti-gfp sirnas and free DS RNAs. The GFP silencing was observed for 10 days with a faster fluorescence recovery for cells treated with functionalized lattices (Figure S8). 13. Viability assay The viability of cells upon transfection was assessed through the CellTiter Blue assay (Promega, Madison) following manufacturer s protocol. Briefly, upon addition of the CellTiter blue reagents to the cells at a 1 to 6 ratio in DMEM, the fluorescence of the resorufin forming compound was measured (560 ex/590 em) after 1-2 hour incubation at 37 C.
9 14. Effects of the presence of functional domains on assembly In regards to structure formation we found that depending on the linked anti-sense and attached sense strand that the structures that form may either be lattices or tubular structures (Figure S9). We observed that when both Dicer substrate (DS) RNAs attached are targeted against GFP, tubular structures form. When both DS RNAs attached are targeted against PLK1, only lattices are observed. When DS RNAs attached are targeted against GFP and PLK1, lattices form. Tubular structures exhibit a left-handed chirality; however, the cause of this chirality is unknown. References 1. Ko, S. H.; Su, M.; Zhang, C.; Ribbe, A. E.; Jiang, W.; Mao, C. Nature chemistry 2010, 2, (12), Stewart, J. M.; Subramanian, H. K.; Franco, E. Submitted. Manuscript can be downloaded here: Milligan, J. F.; Groebe, D. R.; Witherell, G. W.; Uhlenbeck, O. C. Nucleic acids research 1987, 15, (21),
10 Figures Figure S1: Scheme and 3D model of the RNA DX tile. A. Tile without DSRNA domains. B. Tile with DSRNA domains.
11 Figure S2. 10% non-denaturing PAGE analysis of tiles. A. Unpurified samples: (1) Tile with GFP (labeled) and PLK1 (unlabeled) senses. (2) Tile with GFP (unlabeled) and PLK1 (labeled) senses. (3) Tile with GFP (labeled) and PLK1 (labeled) senses (4) Lattice with GFP (labeled) and PLK1 (unlabeled) senses (5) Lattice with GFP (unlabeled) and PLK1 (labeled) senses (6) Lattice with GFP (labeled) and PLK1 (labeled) senses. B. SYBR gold stain of A. C. Purified samples: (1) Tile with GFP (labeled) and PLK1 (unlabeled) senses (upper band) (2) Tile with GFP (labeled) and PLK1 (unlabeled) senses (lower band) (3) Tile with GFP (unlabeled) and PLK1 (labeled) senses (upper band) (4) Tile with GFP (unlabeled) and PLK1 (labeled) senses (lower band) (5) Tile with GFP (labeled) and PLK1 (labeled) senses (upper band) (6) Tile with GFP (labeled) and PLK1 (labeled) senses (lower band) (7) Lattices with GFP (labeled) and PLK1 (unlabeled) senses (8) Lattices with GFP (unlabeled) and PLK1 (labeled) senses (9) Lattices with GFP (labeled) and PLK1 (labeled) senses.
12 Figure S3: 10% non-denaturing PAGE. A. Assembled structures. B. Assembled structures associated with Lipofectamine C. Assembled structures associated with Lipofectamine 2000 and subsequently washed, with Trition X100.
13 Figure S4. Dynamic light scattering (DLS) experiment. The formation of RNA tiles and corresponding lattices functionalized with DS RNAs against GFP and PLK1 were confirmed by ethidium bromine total staining native-page (37.5:1) and further analyzed by DLS. Size distribution by intensity and raw correlation data are shown for each sample. Note that the size distribution and the polydispersity index (PdI) increases dramatically for functionalized lattices.
14 Figure S5 Relative stabilities of functionalized tiles and lattices in human blood serum. A. Fluorescently labeled tiles and lattices treated with 10% (v/v) serum for different amounts of time were visualized on native-page (37.5:1). B. Relative stabilities were estimated based on the quantification of the fluorescent bands. Error bars denote ± SD, N=3.
15 Figure S6. UV melting of DS RNA functionalized tiles and lattices. Melting profiles measured by UV-melt for tiles and lattices functionalized with DS RNAs.
16 Figure S7. Co-localization experiments for RNA tiles and lattices. Human breast cancer cells (MDA-MB-231) were transfected with gel purified tiles and lattices labeled with two different dyes (Alexa 488 and Alexa 546). Image numbers correspond to: differential interference contrast (DIC) images (1), Alexa 488 emission (2), and Alexa 546 emission (3). Images (1+2+3) are superposition of three different images.
17 Figure S8. Specific gene silencing triggered by functionalized RNA tiles and lattices. Silencing was observed for 10 days. Error bars denote ± S.E.M.
18 A B C D E F 0 nm 8 nm Figure S9. Overview of AFM images of assembled structures. A. RNA DX tile with 2 DS RNAs targeted against green fluorescent protein (GFP). B. RNA DX tile with 2 DS RNAs targeted against PLK1. C. RNA DX tile with 2 DS RNAs targeted against GFP and PLK1. D. RNA DX tile with 1 DS RNA targeted against GFP and anti-sense PLK1. E. RNA DX tile with 1 DS RNA targeted against PLK1 and anti-sense GFP. F. RNA DX tile with GFP anti-sense and PLK1 anti-sense. Scale bar: 200 nm.
PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells
 Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Supplementary Information for: PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells Ju Hye Jang 1, Hyun Kim 2, Mi Jung Jang 2, Ju Hyun Cho 1,2,* 1 Research Institute
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH).
 Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Bisulfite Treatment of DNA Dilute DNA sample to 2µg DNA in 50µl ddh 2 O. Add 5µl of 3N NaOH to DNA sample (final concentration 0.3N NaOH). Incubate in a 37ºC water bath for 30 minutes. To 55µl samples
Y-chromosomal haplogroup typing Using SBE reaction
 Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Schematic of multiplex PCR followed by SBE reaction Multiplex PCR Exo SAP purification SBE reaction 5 A 3 ddatp ddgtp 3 T 5 A G 3 T 5 3 5 G C 5 3 3 C 5 ddttp ddctp 5 T 3 T C 3 A 5 3 A 5 5 C 3 3 G 5 3 G
Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC
 Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supplementary Appendixes Supplement 1: Sequences of Capture Probes. Capture probes were /5AmMC6/CTG TAG GTG CGG GTG GAC GTA GTC ACG TAG CTC CGG CTG GA-3 for vimentin, /5AmMC6/TCC CTC GCG CGT GGC TTC CGC
Supporting Information
 Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supporting Information Table S1. Oligonucleotide sequences used in this work Oligo DNA A B C D CpG-A CpG-B CpG-C CpG-D Sequence 5 ACA TTC CTA AGT CTG AAA CAT TAC AGC TTG CTA CAC GAG AAG AGC CGC CAT AGT
Supporting Information
 Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
Supporting Information Transfection of DNA Cages into Mammalian Cells Email: a.turberfield@physics.ox.ac.uk Table of Contents Supporting Figure 1 DNA tetrahedra used in transfection experiments 2 Supporting
Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis
 1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
1 2 3 4 5 6 7 8 9 10 11 12 Figure S1. Characterization of the irx9l-1 mutant. (A) Diagram of the Arabidopsis IRX9L gene drawn based on information from TAIR (the Arabidopsis Information Research). Exons
Supplementary. Table 1: Oligonucleotides and Plasmids. complementary to positions from 77 of the SRα '- GCT CTA GAG AAC TTG AAG TAC AGA CTG C
 Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
Supplementary Table 1: Oligonucleotides and Plasmids 913954 5'- GCT CTA GAG AAC TTG AAG TAC AGA CTG C 913955 5'- CCC AAG CTT ACA GTG TGG CCA TTC TGC TG 223396 5'- CGA CGC GTA CAG TGT GGC CAT TCT GCT G
DNA sentences. How are proteins coded for by DNA? Materials. Teacher instructions. Student instructions. Reflection
 DNA sentences How are proteins coded for by DNA? Deoxyribonucleic acid (DNA) is the molecule of life. DNA is one of the most recognizable nucleic acids, a double-stranded helix. The process by which DNA
DNA sentences How are proteins coded for by DNA? Deoxyribonucleic acid (DNA) is the molecule of life. DNA is one of the most recognizable nucleic acids, a double-stranded helix. The process by which DNA
Supplemental Data Supplemental Figure 1.
 Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Supplemental Data Supplemental Figure 1. Silique arrangement in the wild-type, jhs, and complemented lines. Wild-type (WT) (A), the jhs1 mutant (B,C), and the jhs1 mutant complemented with JHS1 (Com) (D)
Cat. # Product Size DS130 DynaExpress TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1
 Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Product Name: Kit Component TA PCR Cloning Kit (ptakn-2) Cat. # Product Size DS130 TA PCR Cloning Kit (ptakn-2) 20 reactions Box 1 (-20 ) ptakn-2 Vector, linearized 20 µl (50 ng/µl) 1 2 Ligation Buffer
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR
 Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Lecture 10, 20/2/2002: The process of solution development - The CODEHOP strategy for automatic design of consensus-degenerate primers for PCR 1 The problem We wish to clone a yet unknown gene from a known
Supplementary Figures
 Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Supplementary Figures Supplementary Fig. 1 Characterization of GSCs. a. Immunostaining of primary GSC spheres from GSC lines. Nestin (neural progenitor marker, red), TLX (green). Merged images of nestin,
Hes6. PPARα. PPARγ HNF4 CD36
 SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
SUPPLEMENTARY INFORMATION Supplementary Table Positions and Sequences of ChIP primers -63 AGGTCACTGCCA -79 AGGTCTGCTGTG Hes6-0067 GGGCAaAGTTCA ACOT -395 GGGGCAgAGTTCA PPARα -309 GGCTCAaAGTTCAaGTTCA CPTa
Phosphate and R2D2 Restrict the Substrate Specificity of Dicer-2, an ATP- Driven Ribonuclease
 Supplemental Information Molecular Cell, Volume 42 hosphate and R2D2 Restrict the Substrate Specificity of Dicer-2, an AT- Driven Ribonuclease Elif Sarinay Cenik, Ryuya Fukunaga, Gang Lu, Robert Dutcher,
Supplemental Information Molecular Cell, Volume 42 hosphate and R2D2 Restrict the Substrate Specificity of Dicer-2, an AT- Driven Ribonuclease Elif Sarinay Cenik, Ryuya Fukunaga, Gang Lu, Robert Dutcher,
Supporting information for Biochemistry, 1995, 34(34), , DOI: /bi00034a013
 Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Supporting information for Biochemistry, 1995, 34(34), 10807 10815, DOI: 10.1021/bi00034a013 LESNIK 10807-1081 Terms & Conditions Electronic Supporting Information files are available without a subscription
Codon Bias with PRISM. 2IM24/25, Fall 2007
 Codon Bias with PRISM 2IM24/25, Fall 2007 from RNA to protein mrna vs. trna aminoacid trna anticodon mrna codon codon-anticodon matching Watson-Crick base pairing A U and C G binding first two nucleotide
Codon Bias with PRISM 2IM24/25, Fall 2007 from RNA to protein mrna vs. trna aminoacid trna anticodon mrna codon codon-anticodon matching Watson-Crick base pairing A U and C G binding first two nucleotide
Supplementary Information. Construction of Lasso Peptide Fusion Proteins
 Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Supplementary Information Construction of Lasso Peptide Fusion Proteins Chuhan Zong 1, Mikhail O. Maksimov 2, A. James Link 2,3 * Departments of 1 Chemistry, 2 Chemical and Biological Engineering, and
Supplemental Information. Human Senataxin Resolves RNA/DNA Hybrids. Formed at Transcriptional Pause Sites. to Promote Xrn2-Dependent Termination
 Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
Supplemental Information Molecular Cell, Volume 42 Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination Konstantina Skourti-Stathaki, Nicholas
PROTEIN SYNTHESIS Study Guide
 PART A. Read the following: PROTEIN SYNTHESIS Study Guide Protein synthesis is the process used by the body to make proteins. The first step of protein synthesis is called Transcription. It occurs in the
PART A. Read the following: PROTEIN SYNTHESIS Study Guide Protein synthesis is the process used by the body to make proteins. The first step of protein synthesis is called Transcription. It occurs in the
RPA-AB RPA-C Supplemental Figure S1: SDS-PAGE stained with Coomassie Blue after protein purification.
 RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
RPA-AB RPA-C (a) (b) (c) (d) (e) (f) Supplemental Figure S: SDS-PAGE stained with Coomassie Blue after protein purification. (a) RPA; (b) RPA-AB; (c) RPA-CDE; (d) RPA-CDE core; (e) RPA-DE; and (f) RPA-C
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of
 Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Supplementary Figure 1. Localization of MST1 in RPE cells. Proliferating or ciliated HA- MST1 expressing RPE cells (see Fig. 5b for establishment of the cell line) were immunostained for HA, acetylated
Supporting Information. Copyright Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, 2006
 Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2006 Supporting Information for Expanding the Genetic
PCR analysis was performed to show the presence and the integrity of the var1csa and var-
 Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
Supplementary information: Methods: Table S1: Primer Name Nucleotide sequence (5-3 ) DBL3-F tcc ccg cgg agt gaa aca tca tgt gac tg DBL3-R gac tag ttt ctt tca ata aat cac tcg c DBL5-F cgc cct agg tgc ttc
ΔPDD1 x ΔPDD1. ΔPDD1 x wild type. 70 kd Pdd1. Pdd3
 Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Supplemental Fig. S1 ΔPDD1 x wild type ΔPDD1 x ΔPDD1 70 kd Pdd1 50 kd 37 kd Pdd3 Supplemental Fig. S1. ΔPDD1 strains express no detectable Pdd1 protein. Western blot analysis of whole-protein extracts
Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR
 Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Supplemental Dataset Supplemental Table 1. Mutant ADAMTS3 alleles detected in HEK293T clone 4C2. DNA sequence Amino acid sequence WT CCTGTCACTTTGGTTGATAGC MVLLSLWLIAAALVEVR Allele 1 CCTGTC------------------GATAGC
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Multiplexed Detection of Lung Cancer Cells at the Single-Molecule
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Multiplexed Detection of Lung Cancer Cells at the Single-Molecule
Supplementary Figure 1A A404 Cells +/- Retinoic Acid
 Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
Supplementary Figure 1A A44 Cells +/- Retinoic Acid 1 1 H3 Lys4 di-methylation SM-actin VEC cfos (-) RA (+) RA 14 1 1 8 6 4 H3 Lys79 di-methylation SM-actin VEC cfos (-) RA (+) RA Supplementary Figure
MacBlunt PCR Cloning Kit Manual
 MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
MacBlunt PCR Cloning Kit Manual Shipping and Storage MacBlunt PCR Cloning Kits are shipped on dry ice. Each kit contains a box with cloning reagents and an attached bag with Eco-Blue Competent Cells (optional).
Disease and selection in the human genome 3
 Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Disease and selection in the human genome 3 Ka/Ks revisited Please sit in row K or forward RBFD: human populations, adaptation and immunity Neandertal Museum, Mettman Germany Sequence genome Measure expression
Amplified Analysis of DNA by the Autonomous Assembly of Polymers Consisting of DNAzyme Wires
 Supporting information for the paper: Amplified Analysis of DNA by the Autonomous Assembly of Polymers Consisting of DNAzyme Wires Fuan Wang, Johann Elbaz, Ron Orbach, Nimrod Magen and Itamar Willner*
Supporting information for the paper: Amplified Analysis of DNA by the Autonomous Assembly of Polymers Consisting of DNAzyme Wires Fuan Wang, Johann Elbaz, Ron Orbach, Nimrod Magen and Itamar Willner*
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Supporting Online Information
 Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Supporting Online Information Isolation of Human Genomic DNA Sequences with Expanded Nucleobase Selectivity Preeti Rathi, Sara Maurer, Grzegorz Kubik and Daniel Summerer* Department of Chemistry and Chemical
Wet Lab Tutorial: Genelet Circuits
 Wet Lab Tutorial: Genelet Circuits DNA 17 This tutorial will introduce the in vitro transcriptional circuits toolkit. The tutorial will focus on the design, synthesis, and experimental testing of a synthetic
Wet Lab Tutorial: Genelet Circuits DNA 17 This tutorial will introduce the in vitro transcriptional circuits toolkit. The tutorial will focus on the design, synthesis, and experimental testing of a synthetic
DNA Computing Circuits Using Libraries of. DNAzyme Subunits
 SUPPLEMENTARY INFORMATION Supplementary Information for the paper: DNA Computing Circuits Using Libraries of DNAzyme Subunits Johann Elbaz a, Oleg lioubashevski a, Fuan Wang a, Françoise Remacle b, Raphael
SUPPLEMENTARY INFORMATION Supplementary Information for the paper: DNA Computing Circuits Using Libraries of DNAzyme Subunits Johann Elbaz a, Oleg lioubashevski a, Fuan Wang a, Françoise Remacle b, Raphael
Materials Protein synthesis kit. This kit consists of 24 amino acids, 24 transfer RNAs, four messenger RNAs and one ribosome (see below).
 Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Protein Synthesis Instructions The purpose of today s lab is to: Understand how a cell manufactures proteins from amino acids, using information stored in the genetic code. Assemble models of four very
Arabidopsis actin depolymerizing factor AtADF4 mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB
 Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Arabidopsis actin depolymerizing factor mediates defense signal transduction triggered by the Pseudomonas syringae effector AvrPphB Files in this Data Supplement: Supplemental Table S1 Supplemental Table
Molecular Level of Genetics
 Molecular Level of Genetics Most of the molecules found in humans and other living organisms fall into one of four categories: 1. carbohydrates (sugars and starches) 2. lipids (fats, oils, and waxes) 3.
Molecular Level of Genetics Most of the molecules found in humans and other living organisms fall into one of four categories: 1. carbohydrates (sugars and starches) 2. lipids (fats, oils, and waxes) 3.
Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were
 1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
1 Supplemental methods 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 18 19 21 22 23 Quantitative reverse-transcription PCR. Transcript levels of flgs, flgr, flia and flha were monitored by quantitative reverse-transcription
Legends for supplementary figures 1-3
 High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
High throughput resistance profiling of Plasmodium falciparum infections based on custom dual indexing and Illumina next generation sequencing-technology Sidsel Nag 1,2 *, Marlene D. Dalgaard 3, Poul-Erik
Gene synthesis by circular assembly amplification
 Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Gene synthesis by circular assembly amplification Duhee Bang & George M Church Supplementary figures and text: Supplementary Figure 1. Dpo4 gene (1.05kb) construction by various methods. Supplementary
Surface modification of a DNA tetrahedron
 Surface modification of a DNA tetrahedron Chuan Zhang, Min Su, Yu He, Yujun Leng, Alexander E. Ribbe, Guansong Wang ±, Wen Jiang & Chengde Mao * Department of Chemistry, Markey Center for Structural Biology
Surface modification of a DNA tetrahedron Chuan Zhang, Min Su, Yu He, Yujun Leng, Alexander E. Ribbe, Guansong Wang ±, Wen Jiang & Chengde Mao * Department of Chemistry, Markey Center for Structural Biology
ORFs and genes. Please sit in row K or forward
 ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
ORFs and genes Please sit in row K or forward https://www.flickr.com/photos/teseum/3231682806/in/photostream/ Question: why do some strains of Vibrio cause cholera and others don t? Methods Mechanisms
SUPPORTING INFORMATION
 SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
SUPPORTING INFORMATION Investigation of the Biosynthesis of the Lasso Peptide Chaxapeptin Using an E. coli-based Production System Helena Martin-Gómez, Uwe Linne, Fernando Albericio, Judit Tulla-Puche,*
A green method of staining DNA in polyacrylamide gel electrophoresis based on fluorescent copper nanoclusters synthesized in situ
 Electronic Supplementary Material A green method of staining DNA in polyacrylamide gel electrophoresis based on fluorescent copper nanoclusters synthesized in situ Xiaoli Zhu 1, Hai Shi 1, Yalan Shen 1,
Electronic Supplementary Material A green method of staining DNA in polyacrylamide gel electrophoresis based on fluorescent copper nanoclusters synthesized in situ Xiaoli Zhu 1, Hai Shi 1, Yalan Shen 1,
Supplementary Information
 Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
Supplementary Information A general solution for opening double-stranded DNA for isothermal amplification Gangyi Chen, Juan Dong, Yi Yuan, Na Li, Xin Huang, Xin Cui* and Zhuo Tang* Supplementary Materials
Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR
 Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
Supplementary Methods Antibodies Anti-Pim-1 (Cat#3247), anti-met (Cat#3127), anti-ron (Cat#2654), Anti-EGFR (Cat#2646), anti-igf1r (Cat#3018), anti-insr (Cat#3020), anti-akt (pan, Cat#4691), anti-phospho-akt
NAME:... MODEL ANSWER... STUDENT NUMBER:... Maximum marks: 50. Internal Examiner: Hugh Murrell, Computer Science, UKZN
 COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
COMP710, Bioinformatics with Julia, Test One, Thursday the 20 th of April, 2017, 09h30-11h30 1 NAME:...... MODEL ANSWER... STUDENT NUMBER:...... Maximum marks: 50 Internal Examiner: Hugh Murrell, Computer
SUPPLEMENTARY INFORMATION
 1. RNA/DNA sequences used in this study 2. Height and stiffness measurements on hybridized molecules 3. Stiffness maps at varying concentrations of target DNA 4. Stiffness measurements on RNA/DNA hybrids.
1. RNA/DNA sequences used in this study 2. Height and stiffness measurements on hybridized molecules 3. Stiffness maps at varying concentrations of target DNA 4. Stiffness measurements on RNA/DNA hybrids.
SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer
 TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
TEACHER S GUIDE SAY IT WITH DNA: Protein Synthesis Activity by Larry Flammer SYNOPSIS This activity uses the metaphor of decoding a secret message for the Protein Synthesis process. Students teach themselves
strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular
![strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular strain devoid of the aox1 gene [1]. Thus, the identification of AOX1 in the intracellular](/thumbs/72/66361212.jpg) Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Additional file 2 Identification of AOX1 in P. pastoris GS115 with a Mut s phenotype Results and Discussion The HBsAg producing strain was originally identified as a Mut s (methanol utilization slow) strain
Supplemental material
 Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Supplemental material Diversity of O-antigen repeat-unit structures can account for the substantial sequence variation of Wzx translocases Yaoqin Hong and Peter R. Reeves School of Molecular Bioscience,
Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURB
 Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURBO DNA-free Kit (Ambion). One µg of total RNA was reverse
Supplementary Methods Quantitative RT-PCR. For mrna, total RNA was prepared using TRIzol reagent (Invitrogen) and genomic DNA was eliminated with TURBO DNA-free Kit (Ambion). One µg of total RNA was reverse
Multi-Amplified Sensing of MicroRNA by a Small DNA Fragment-Driven Enzymatic Cascade Reaction
 Supporting Information for Multi-Amplified Sensing of MicroRNA by a Small DNA Fragment-Driven Enzymatic Cascade Reaction Eunjung Kim, Philip D. Howes, Spencer W. Crowder, and Molly M. Stevens* Department
Supporting Information for Multi-Amplified Sensing of MicroRNA by a Small DNA Fragment-Driven Enzymatic Cascade Reaction Eunjung Kim, Philip D. Howes, Spencer W. Crowder, and Molly M. Stevens* Department
PCR-based Markers and Cut Flower Longevity in Carnation
 PCRbased Markers and Cut Flower Longevity in Carnation Laura De Benedetti, Luca Braglia, Simona Bruna, Gianluca Burchi *, Antonio Mercuri and Tito Schiva Istituto Sperimentale per la Floricoltura, Corso
PCRbased Markers and Cut Flower Longevity in Carnation Laura De Benedetti, Luca Braglia, Simona Bruna, Gianluca Burchi *, Antonio Mercuri and Tito Schiva Istituto Sperimentale per la Floricoltura, Corso
Supporting Information. for. Angew. Chem. Int. Ed. Z Wiley-VCH 2004
 Supporting Information for Angew. Chem. Int. Ed. Z53445 Wiley-VC 2004 69451 Weinheim, Germany Small Interfering RNA Expression in Cells Based on an Efficiently Constructed Dumbbell-shaped DNA Masumi Taki,
Supporting Information for Angew. Chem. Int. Ed. Z53445 Wiley-VC 2004 69451 Weinheim, Germany Small Interfering RNA Expression in Cells Based on an Efficiently Constructed Dumbbell-shaped DNA Masumi Taki,
Chapter 3: Information Storage and Transfer in Life
 Chapter 3: Information Storage and Transfer in Life The trapped scientist examples are great for conceptual purposes, but they do not accurately model how information in life changes because they do not
Chapter 3: Information Storage and Transfer in Life The trapped scientist examples are great for conceptual purposes, but they do not accurately model how information in life changes because they do not
Dierks Supplementary Fig. S1
 Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
Dierks Supplementary Fig. S1 ITK SYK PH TH K42R wt K42R (kinase deficient) R29C E42K Y323F R29C E42K Y323F (reduced phospholipid binding) (enhanced phospholipid binding) (reduced Cbl binding) E42K Y323F
hcd1tg/hj1tg/ ApoE-/- hcd1tg/hj1tg/ ApoE+/+
 ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
ApoE+/+ ApoE-/- ApoE-/- H&E (1x) Supplementary Figure 1. No obvious pathology is observed in the colon of diseased ApoE-/me. Colon samples were fixed in 1% formalin and laid out in Swiss rolls for paraffin
Table S1. Bacterial strains (Related to Results and Experimental Procedures)
 Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
Table S1. Bacterial strains (Related to Results and Experimental Procedures) Strain number Relevant genotype Source or reference 1045 AB1157 Graham Walker (Donnelly and Walker, 1989) 2458 3084 (MG1655)
DNA Tetrahedron-Based Molecular Beacon for Tumor-Related
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2015 Electronic Supporting Information (ESI) DNA Tetrahedron-Based Molecular Beacon for Tumor-Related
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2015 Electronic Supporting Information (ESI) DNA Tetrahedron-Based Molecular Beacon for Tumor-Related
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
www.sciencesignaling.org/cgi/content/full/10/494/eaan6284/dc1 Supplementary Materials for Activation of master virulence regulator PhoP in acidic ph requires the Salmonella-specific protein UgtL Jeongjoon
Universal Split Spinach Aptamer (USSA) for Nucleic Acid Analysis and DNA Computation
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2017 Supporting materials Universal Split Spinach Aptamer (USSA) for Nucleic Acid Analysis and DNA Computation
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2017 Supporting materials Universal Split Spinach Aptamer (USSA) for Nucleic Acid Analysis and DNA Computation
Overexpression Normal expression Overexpression Normal expression. 26 (21.1%) N (%) P-value a N (%)
 SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
SUPPLEMENTARY TABLES Table S1. Alteration of ZNF322A protein expression levels in relation to clinicopathological parameters in 123 Asian and 74 Caucasian lung cancer patients. Asian patients Caucasian
Supporting Information. Trifluoroacetophenone-Linked Nucleotides and DNA for Studying of DNA-protein Interactions by 19 F NMR Spectroscopy
 Supporting Information Trifluoroacetophenone-Linked Nucleotides and DNA for Studying of DNA-protein Interactions by 19 F NMR Spectroscopy Agata Olszewska, Radek Pohl and Michal Hocek # * Institute of Organic
Supporting Information Trifluoroacetophenone-Linked Nucleotides and DNA for Studying of DNA-protein Interactions by 19 F NMR Spectroscopy Agata Olszewska, Radek Pohl and Michal Hocek # * Institute of Organic
Lecture 19A. DNA computing
 Lecture 19A. DNA computing What exactly is DNA (deoxyribonucleic acid)? DNA is the material that contains codes for the many physical characteristics of every living creature. Your cells use different
Lecture 19A. DNA computing What exactly is DNA (deoxyribonucleic acid)? DNA is the material that contains codes for the many physical characteristics of every living creature. Your cells use different
evaluated with UAS CLB eliciting UAS CIT -N Libraries increase in the
 Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Supplemental Data. Bennett et al. (2010). Plant Cell /tpc
 BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
BRN1 ---------MSSSNGGVPPGFRFHPTDEELLHYYLKKKISYEKFEMEVIKEVDLNKIEPWDLQDRCKIGSTPQNEWYFFSHKDRKYPTGS 81 BRN2 --------MGSSSNGGVPPGFRFHPTDEELLHYYLKKKISYQKFEMEVIREVDLNKLEPWDLQERCKIGSTPQNEWYFFSHKDRKYPTGS 82 SMB
Luo et al. Supplemental Figures and Materials and Methods
 Luo et al. Supplemental Figures and Materials and Methods The supplemental figures demonstrate that nuclear NFAT is situated at PODs, overexpressed PML does not increase NFAT nuclear localization, and
Luo et al. Supplemental Figures and Materials and Methods The supplemental figures demonstrate that nuclear NFAT is situated at PODs, overexpressed PML does not increase NFAT nuclear localization, and
Nongenetic Reprogramming of the Ligand Specificity. of Growth Factor Receptors by Bispecific DNA Aptamers
 Supporting Information For Nongenetic Reprogramming of the Ligand Specificity of Growth Factor Receptors by Bispecific DNA Aptamers Ryosuke Ueki,* Saki Atsuta, Ayaka Ueki and Shinsuke Sando* Department
Supporting Information For Nongenetic Reprogramming of the Ligand Specificity of Growth Factor Receptors by Bispecific DNA Aptamers Ryosuke Ueki,* Saki Atsuta, Ayaka Ueki and Shinsuke Sando* Department
A netlike rolling circle nucleic acid amplification technique
 Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2014 Supplementary Information A netlike rolling circle nucleic acid amplification technique Xiaoli Zhu,
Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2014 Supplementary Information A netlike rolling circle nucleic acid amplification technique Xiaoli Zhu,
A high efficient electrochemiluminescence resonance energy. transfer system in one nanostructure: its application for
 Supporting Information for A high efficient electrochemiluminescence resonance energy transfer system in one nanostructure: its application for ultrasensitive detection of microrna in cancer cells Zhaoyang
Supporting Information for A high efficient electrochemiluminescence resonance energy transfer system in one nanostructure: its application for ultrasensitive detection of microrna in cancer cells Zhaoyang
Engineering D66N mutant using quick change site directed mutagenesis. Harkewal Singh 09/01/2010
 Engineering D66N mutant using quick change site directed mutagenesis Harkewal Singh 09/01/2010 1 1- What is quick change site directed mutagenesis? 2- An overview of the kit contents. 3- A brief information
Engineering D66N mutant using quick change site directed mutagenesis Harkewal Singh 09/01/2010 1 1- What is quick change site directed mutagenesis? 2- An overview of the kit contents. 3- A brief information
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system
 Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Converting rabbit hybridoma into recombinant antibodies with effective transient production in an optimized human expression system Dr. Tim Welsink Molecular Biology Transient Gene Expression OUTLINE Short
Supplementary Figure 1. The level of pri-mir-8 gradually decreases while those of BR-C and E74 increase during 3rd instar larval development.
 qrt-pcr RT-PCR Relative pri-mir-8 level 1.2 1.0 0.8 0.6 0.4 0.2 0.0 Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) BR-C E74 mtl rrna Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) Supplementary Figure 1.
qrt-pcr RT-PCR Relative pri-mir-8 level 1.2 1.0 0.8 0.6 0.4 0.2 0.0 Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) BR-C E74 mtl rrna Early 3rd (72h) Mid 3rd (96h) Late 3rd (119h) Supplementary Figure 1.
Supplementary Figure 1. Ratiometric fluorescence visualization of DNA cleavage by
 Supplementary Figure 1. Ratiometric fluorescence visualization of DNA cleavage by Cas9:gRNA. (a) A labeled by Cy3 and Cy5 with an inter-probe distance of > 30 bp was tethered to a PEG-coated surface via
Supplementary Figure 1. Ratiometric fluorescence visualization of DNA cleavage by Cas9:gRNA. (a) A labeled by Cy3 and Cy5 with an inter-probe distance of > 30 bp was tethered to a PEG-coated surface via
11th Meeting of the Science Working Group. Lima, Peru, October 2012 SWG-11-JM-11
 11th Meeting of the Science Working Group Lima, Peru, 15-19 October 2012 Russian population genetics studies of jack mackerel in the South Pacific P.K.Afanasiev M.A.Rabchun A.I.Glubokov Introduction. In
11th Meeting of the Science Working Group Lima, Peru, 15-19 October 2012 Russian population genetics studies of jack mackerel in the South Pacific P.K.Afanasiev M.A.Rabchun A.I.Glubokov Introduction. In
Supporting Information
 Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
Supporting Information Barderas et al. 10.1073/pnas.0801221105 SI Text: Docking of gastrin to Constructed scfv Models Interactive predocking of the 4-WL-5 motif into the central pocket observed in the
Multiplexing Genome-scale Engineering
 Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
Multiplexing Genome-scale Engineering Harris Wang, Ph.D. Department of Systems Biology Department of Pathology & Cell Biology http://wanglab.c2b2.columbia.edu Rise of Genomics An Expanding Toolbox Esvelt
G+C content. 1 Introduction. 2 Chromosomes Topology & Counts. 3 Genome size. 4 Replichores and gene orientation. 5 Chirochores.
 1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
1 Introduction 2 Chromosomes Topology & Counts 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 7 Codon usage 121 marc.bailly-bechet@univ-lyon1.fr Bacterial genome structures Introduction
Human Gene,cs 06: Gene Expression. Diversity of cell types. How do cells become different? 9/19/11. neuron
 Human Gene,cs 06: Gene Expression 20110920 Diversity of cell types neuron How do cells become different? A. Each type of cell has different DNA in its nucleus B. Each cell has different genes C. Each type
Human Gene,cs 06: Gene Expression 20110920 Diversity of cell types neuron How do cells become different? A. Each type of cell has different DNA in its nucleus B. Each cell has different genes C. Each type
SUPPORTING INFORMATION FILE
 Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules Nan Liu, Mengxi Wang, Wen Deng, Christine S. Schmidt, Weihua Qin, Heinrich Leonhardt and Fabio Spada Department of
Supplemental Table 1. Primers used for PCR.
 Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
Supplemental Table 1. Primers used for PCR. Gene Type Primer Sequence Genotyping and semi-quantitative RT-PCR F 5 -TTG CCC GAT CAC CAT CTG TA-3 rwa1-1 R 5 -TGT AGC GAT CAA GGC CTG ATC TAA-3 LB 5 -TAG CAT
SUPPLEMENTARY INFORMATION
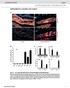 doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
doi: 10.1038/nature07182 SUPPLEMENTAL FIGURES AND TABLES Fig. S1. myf5-expressing cells give rise to brown fat depots and skeletal muscle (a) Perirenal BAT from control (cre negative) and myf5-cre:r26r3-yfp
A 2D graphical representation of the sequences of DNA based on triplets and its application
 Zou et al. EURASIP Journal on Bioinformatics and Systems Biology 2014, 2014:1 RESEARCH Open Access A 2D graphical representation of the sequences of DNA based on triplets and its application Sai Zou, Lei
Zou et al. EURASIP Journal on Bioinformatics and Systems Biology 2014, 2014:1 RESEARCH Open Access A 2D graphical representation of the sequences of DNA based on triplets and its application Sai Zou, Lei
Circular bivalent aptamers enable in vivo stability and recognition
 Supporting Information Circular bivalent aptamers enable in vivo stability and recognition Hailan Kuai,, Zilong Zhao,, Liuting Mo, Hui Liu, Xiaoxiao Hu, Ting Fu, Xiaobing Zhang, Weihong Tan*,, Molecular
Supporting Information Circular bivalent aptamers enable in vivo stability and recognition Hailan Kuai,, Zilong Zhao,, Liuting Mo, Hui Liu, Xiaoxiao Hu, Ting Fu, Xiaobing Zhang, Weihong Tan*,, Molecular
Lecture 11: Gene Prediction
 Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
Lecture 11: Gene Prediction Study Chapter 6.11-6.14 1 Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and end positions of genes in a genome Where are
II 0.95 DM2 (RPP1) DM3 (At3g61540) b
 Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
Table S2. F 2 Segregation Ratios at 16 C, Related to Figure 2 Cross n c Phenotype Model e 2 Locus A Locus B Normal F 1 -like Enhanced d Uk-1/Uk-3 149 64 36 49 DM2 (RPP1) DM1 (SSI4) a Bla-1/Hh-0 F 3 111
Sequencing of DNA lesions facilitated by site-specific excision via base. excision repair DNA glycosylases yielding ligatable gaps
 Supporting information Sequencing of DNA lesions facilitated by site-specific excision via base excision repair DNA glycosylases yielding ligatable gaps Jan Riedl, Aaron M. Fleming, and Cynthia J. Burrows*
Supporting information Sequencing of DNA lesions facilitated by site-specific excision via base excision repair DNA glycosylases yielding ligatable gaps Jan Riedl, Aaron M. Fleming, and Cynthia J. Burrows*
Supplemental Data. Lin28 Mediates the Terminal Uridylation. of let-7 Precursor MicroRNA. Molecular Cell, Volume 32
 Molecular Cell, Volume 32 Supplemental Data Lin28 Mediates the Terminal Uridylation of let-7 Precursor MicroRNA Inha Heo, Chirlmin Joo, Jun Cho, Minju Ha, Jinju Han, and V. Narry Kim Figure S1. Endogenous
Molecular Cell, Volume 32 Supplemental Data Lin28 Mediates the Terminal Uridylation of let-7 Precursor MicroRNA Inha Heo, Chirlmin Joo, Jun Cho, Minju Ha, Jinju Han, and V. Narry Kim Figure S1. Endogenous
Event-specific Method for the Quantification of Soybean SYHT0H2 by Real-time PCR. Validated Method
 EUROPEAN COMMISSION JOINT RESEARCH CENTRE Institute for Health and Consumer Protection Molecular Biology and Genomics Unit Event-specific Method for the Quantification of Soybean SYHT0H2 by Real-time PCR
EUROPEAN COMMISSION JOINT RESEARCH CENTRE Institute for Health and Consumer Protection Molecular Biology and Genomics Unit Event-specific Method for the Quantification of Soybean SYHT0H2 by Real-time PCR
Evolution of protein coding sequences
 Evolution of protein coding sequences Kinds of nucleo-de subs-tu-ons Given 2 nucleo-de sequences, how their similari-es and differences arose from a common ancestor? We assume A the common ancestor: Single
Evolution of protein coding sequences Kinds of nucleo-de subs-tu-ons Given 2 nucleo-de sequences, how their similari-es and differences arose from a common ancestor? We assume A the common ancestor: Single
I. Things to keep in mind before you start this protocol
 Inverse PCR and Sequencing Protocol on 5 Fly Preps For recovery of sequences flanking piggybac elements This protocol is an adaptation of "Inverse PCR and Cycle Sequencing Protocols" by E. Jay Rehm Berkeley
Inverse PCR and Sequencing Protocol on 5 Fly Preps For recovery of sequences flanking piggybac elements This protocol is an adaptation of "Inverse PCR and Cycle Sequencing Protocols" by E. Jay Rehm Berkeley
An enzyme-free DNA walker that moves on the surface of. functionalized magnetic microparticles and its biosensing analysis
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry
Supplementary Information
 Supplementary Information Microbead-based biomimetic synthetic neighbors enhance survival and function of rat pancreatic β-cells Wei Li, a Samuel Lee, b Minglin Ma, a, f Soo Min Kim, b Patrick Guye, c
Supplementary Information Microbead-based biomimetic synthetic neighbors enhance survival and function of rat pancreatic β-cells Wei Li, a Samuel Lee, b Minglin Ma, a, f Soo Min Kim, b Patrick Guye, c
Electronic Supplementary Information
 Electronic Supplementary Information Ultrasensitive and Selective DNA Detection Based on Nicking Endonuclease Assisted Signal Amplification and Its Application in Cancer Cells Detection Sai Bi, Jilei Zhang,
Electronic Supplementary Information Ultrasensitive and Selective DNA Detection Based on Nicking Endonuclease Assisted Signal Amplification and Its Application in Cancer Cells Detection Sai Bi, Jilei Zhang,
Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers
 DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
DNA Research 9, 163 171 (2002) Search for and Analysis of Single Nucleotide Polymorphisms (SNPs) in Rice (Oryza sativa, Oryza rufipogon) and Establishment of SNP Markers Shinobu Nasu, Junko Suzuki, Rieko
Homework. A bit about the nature of the atoms of interest. Project. The role of electronega<vity
 Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
Homework Why cited articles are especially useful. citeulike science citation index When cutting and pasting less is more. Project Your protein: I will mail these out this weekend If you haven t gotten
Fishy Amino Acid Codon. UUU Phe UCU Ser UAU Tyr UGU Cys. UUC Phe UCC Ser UAC Tyr UGC Cys. UUA Leu UCA Ser UAA Stop UGA Stop
 Fishy Code Slips Fish 1 GGTTATAGAGGTACTACC Fish 2 GGCTTCAGAGGTACTACC Fish 3 CATAGCAGAGGTACTACC Fish 4 GGTTATTCTGTCTTATTG Fish 5 GGCTTCTCTGTCTTATTG Fish 6 CATAGCGCTGCAACTACC Fishy Amino Acid Codon UUU Phe
Fishy Code Slips Fish 1 GGTTATAGAGGTACTACC Fish 2 GGCTTCAGAGGTACTACC Fish 3 CATAGCAGAGGTACTACC Fish 4 GGTTATTCTGTCTTATTG Fish 5 GGCTTCTCTGTCTTATTG Fish 6 CATAGCGCTGCAACTACC Fishy Amino Acid Codon UUU Phe
