Antibody-Antigen recognition. Structural Biology Weekend Seminar Annegret Kramer
|
|
|
- Ralf Bell
- 6 years ago
- Views:
Transcription
1 Antibody-Antigen recognition Structural Biology Weekend Seminar Annegret Kramer
2 Contents Function and structure of antibodies Features of antibody-antigen interfaces Examples of antibody-antigen interactions Cross reactivity Modeling of antibody-protein antigen interactions Anti-idiotypic antibodys Antibody-Antigen Recognition 2
3 Basics of antibodies 1 Antibodies = Immunoglobulins; 5 classes: IgA, IgD, IgE, IgG, IgM Homodimers 2 identical polypeptide chains (450 aa) heavy chains 2 identical polypeptide chains (250 aa) light chains Heavy & light chains divided into N-terminal variable & C-terminal constant parts Antibody-Antigen Recognition 3
4 Basics of antibodies 2 β-sheets together form a barrelshaped structure β-barrel structure of the immunoglobulin protein domain immunoglobulin fold Antibody-Antigen Recognition 4
5 Basics of antibodies 3 Each heavy chain: V H, C H 1, C H 2, C H 3 domains each domain: 2 anti-parallel β-sheets Each light chain: V L, C L domains Each V H and V L : 3 loops connecting the β- strands, highly variable in length & sequence comlementarity determining regions (CDRs) Antibody-Antigen Recognition 5
6 Fv Basics of antibodies 4 Fc Fab Fragments antigen binding (Fab): 2 identical fragments complete light chains + V H + C H 1 Fragment crystallizable (Fc): paired C H 2 + C H 3 Fragment variable (Fv): heterodimer V H + V L Antibody-Antigen Recognition 6
7 Basics of antibodies 5 Disulfide bridges link the hinge regions of the heavy chains & C H 1 + C L Antibody-Antigen Recognition 7
8 Central paradigm of antigenantibody recognition The 6 CDRs come together to form a binding surface 3 dimensional structure formed by the 6 CDRs recognizes & binds a complementary surface (epitope) on the antigen Sequences & structures of the CDRs determine the specifity of the surface Antibody-Antigen Recognition 8
9 Features of antibody-antigen interfaces 1 Richer in aromatic residues, esp. tyrosine, tryptophan than average protein surface Depleted in charged residues aspartate, glutamate, lysine, but enriched in arginine & aromatic residues Analysis of alanine mutations on binding energies thermodynamic hot spots due to multiple favorable interactions Antibody-Antigen Recognition 9
10 Features of antibody-antigen interfaces 2 Light and heavy chains make contacts with antigen (heavy chain more extensive) Specifity of binding determined by CDRs of V H and V L Contacting residues of the antigen: discontinuous in sequence, but form an epitope High complementarity of contacting surfaces Antibody-Antigen Recognition 10
11 Features of antibody-antigen interfaces 3 Surface areas of interaction: ca Å 2 Mediation of binding: v.d.waals, hydrogen bonds, salt bridges, occasional ion pairs Water molecules reinforce complementarity & interactions (supported by molecular dynamics simulations) Enthalpic forces (hydrogen bonds, v.d.waals) drive reaction Antibody-Antigen Recognition 11
12 Binding of a small antigen Complex between an Fab fragment of an antibody & its target phosphoryl-choline Residues from the antibody interact through H-bonding & electrostatic & v.d. Waals interactions Antibody-Antigen Recognition 12
13 Antibody-Protein Interactions Complex between an Fab fragment + lysozyme Binding surfaces are complementary in shape over a large area Glutamine 121 penetrates more deeply into the antibody combining site Antibody-Antigen Recognition 13
14 Fv--Mouse anti-hel monoclonal Antibody D1.3 complex with lysozyme 1.8 Å resolution 178 water molecules, 50 around interface with HEL Small conformational changes /side chain movements esp. in V H CDR3 Small rearrangement of V H and V L domains contacting residues closer to antigen by Å induced fit facilitating binding of antigen Antibody-Antigen Recognition 14
15 Mouse anti-hel (hen egg-white lysozyme) monoclonal Antibody D1.3 Stereo view of the VL CDR1 and CDR2 (thick bonds) & neighboring solvent molecules & lysozyme atoms from the Fv D1.3-HEL complex extent of hydration at the interface Antibody-Antigen Recognition 15
16 Mutant of Fv D1.3 (V L Trp92 Asp) complexed with HEL 1000-fold reduction in equilibrium binding constant wt: Trp92 extensive v.d.waals with HEL Gln121, Arg125 & Ile124 Mutant: rearrangement of H 2 O molecules at V L -HEL interface 2 H 2 O molecules fill hole created by Trp92 Asp Contact area lost: 150 Å 2-16 kj mol -1 enthalpy Antibody-Antigen Recognition 16
17 Mutant of Fv D1.3 (V L Trp92 Asp) complexed with HEL Antibody-Antigen Recognition 17
18 Mouse anti-hel Fab D salt links between V H glutamates (35 &50) and HEL arginines (45 & 68) Arg45 steric hindered by V L Trp94 from its preferred rotameric position 3 H 2 O molecules buried at interface H-bonds, stability & complementarity of complex Antibody-Antigen Recognition 18
19 Stereo view H-bonding scheme between Fab D44.1 and HEL Fab D44.1 (thick bonds) and HEL(lighter bonds); stacking of guanidino group of HEL Arg68 & indole ring of V H Trp54 Antibody-Antigen Recognition 19
20 Cross-reactivity Cross-reactions with closely related antigens frequently observed Antibodies binding better to antigens not used in challenging immune system heteroclitic binding Anti-HEL monoclonal antibody D11.15 which was raised against HEL, binds several avian lysozymes with higher affinity Antibody-Antigen Recognition 20
21 Fv of Anti-HEL D11.15 complexed with PHL (pheasant egg-white lysozyme) Comparison with D1.3-HEL structural model to explain high specifity (D1.3) & broader cross reactivity (D11.15) D1.3 and D11.15 obtained from the same individual mouse bind to partially overlapping epitopes Antibody-Antigen Recognition 21
22 Fv of Anti-HEL D11.15 complexed with PHL compared with D1.3-HEL Replacement at position 121, located in aromatic pocket between V H & V L domains prevents D1.3 from binding other lysozymes Tight fit in a central location of interface D1.3 cannot bind lysozymes not featuring Gln121 D11.15 binds an epitope shared by several lysozymes Here high specifity & cross reactivity depend on location of evolutionarily produced sequence changes in antigen Antibody-Antigen Recognition 22
23 Modeling of antibody-protein antigen interactions Statistical method for assessing the degree of complementarity (defined by packing density) Computer-aided modeling of antibody-antigen docking steric score (complementarity & docking) modified Lennard-Jones potential, consisting of a broad energy minimum simulating the likelihood of side-chain reorientation during binding Results marginal revealed by crystalline complexes Antibody-Antigen Recognition 23
24 Other attempts at computer simulation of antibody-antigen docking 1 Based on maximizing the surface area of interaction Docking arrangements showed only limited success Antibody-Antigen Recognition 24
25 Other attempts at computer simulation of antibody-antigen docking 2 Molecular dynamics analysis of D1.3-HEL association Many H 2 O molecules were attracted to antibody-antigen interface during the case of simulation Resulting models do not differ significantly from those obtained from X-ray crystal structures Antibody-Antigen Recognition 25
26 Anti-idiotypic antibodys (Ab2)...recognize an idiotope (=antigenic determinant that is unique to an antibody (Ab1)) Aim: Use of these mimics as sort of vaccines Antibody-Antigen Recognition 26
27 Complex between FabD1.3 & its idiotypic antibody FabE225 First crystallographic analysis of idiotypicanti-idiotope interactions 14 CDR residues from FabE225 interact with 10 CDR residues from FabD1.3 1 framework residue from FabE225, 3 framework residues from FabD1.3 Antibody-Antigen Recognition 27
28 References Braden BC, Poljak RJ: Structural features of the reactions between antibodies and protein antigens FASEB J.9, 9-16 (1995) Sundberg EJ, Mariuzza RA: Molecular recognition in antibody-antigen complexes Adv Prot Chem, Vol.61 (2003) Amzel LM, Gaffney BJ: Structural immunology: problems in molecular recognition FASEB J. 9, 7-8 (1995) Antibody-Antigen Recognition 28
29 Thank you for your attention Antibody-Antigen Recognition 29
Chapter 4. Antigen Recognition by B-cell and T-cell Receptors
 Chapter 4 Antigen Recognition by B-cell and T-cell Receptors Antigen recognition by BCR and TCR B cells 2 separate functions of immunoglobulin (Ig) bind pathogen & induce immune responses recruit cells
Chapter 4 Antigen Recognition by B-cell and T-cell Receptors Antigen recognition by BCR and TCR B cells 2 separate functions of immunoglobulin (Ig) bind pathogen & induce immune responses recruit cells
Molecular recognition
 Lecture 9 Molecular recognition Antoine van Oijen BCMP201 Spring 2008 Structural principles of binding 4 fundamental functions of proteins: 1) Binding 2) Catalysis 3) Switching 4) Structural All involve
Lecture 9 Molecular recognition Antoine van Oijen BCMP201 Spring 2008 Structural principles of binding 4 fundamental functions of proteins: 1) Binding 2) Catalysis 3) Switching 4) Structural All involve
Antibodies (Recommended reading: Abbas et al., 4th edition, Chapter 3; Chapter 4; Janeway et al., 5th edition, Chapter 3)
 HST 175 Antibodies (Recommended reading: Abbas et al., 4th edition, Chapter 3; Chapter 4; Janeway et al., 5th edition, Chapter 3) Antibodies protect us from a vast variety of pathogens. Indeed the antibody
HST 175 Antibodies (Recommended reading: Abbas et al., 4th edition, Chapter 3; Chapter 4; Janeway et al., 5th edition, Chapter 3) Antibodies protect us from a vast variety of pathogens. Indeed the antibody
It had been determined by several means that the proteins with antibody activity (immunoglobulins) had a molecular weight of approximately 150 kda.
 Immunology Dr. John J. Haddad Chapter 4 Antibody Structure and Function In 1890, Emil von Behring and Shibasaburo Kitasato showed that serum (the straw-colored liquid remaining after blood clots and the
Immunology Dr. John J. Haddad Chapter 4 Antibody Structure and Function In 1890, Emil von Behring and Shibasaburo Kitasato showed that serum (the straw-colored liquid remaining after blood clots and the
Antigen Antibody Binding
 Izumi Kumagai, Tohoku University, Sendai, Japan Kouhei Tsumoto, Tohoku University, Sendai, Japan Antibodies are a family of glycoproteins that bind specifically to foreign molecules (antigens). The binding
Izumi Kumagai, Tohoku University, Sendai, Japan Kouhei Tsumoto, Tohoku University, Sendai, Japan Antibodies are a family of glycoproteins that bind specifically to foreign molecules (antigens). The binding
Immunoglobulins. Light chain ~22-23 KDa whereas the heavy chain ~55-60 KDa
 Immunoglobulins Immunoglobulin (Ig) has a common name which is "Antibody (Ab)", but actually we should say Ig, why? Because the proteins, which are involved, are actually globular proteins "known as globulins"
Immunoglobulins Immunoglobulin (Ig) has a common name which is "Antibody (Ab)", but actually we should say Ig, why? Because the proteins, which are involved, are actually globular proteins "known as globulins"
Immunoglobulins. Even variable chain differ in variability of amino acid sequences:
 Revision: Immunoglobulin structure 2 light chain 25KDa and 2 heavy chain 50KDa (total=150kda) Heavy chain one quarter variable three quarter constant Even variable chain differ in variability of amino
Revision: Immunoglobulin structure 2 light chain 25KDa and 2 heavy chain 50KDa (total=150kda) Heavy chain one quarter variable three quarter constant Even variable chain differ in variability of amino
Attribution: University of Michigan Medical School, Department of Microbiology and Immunology
 Attribution: University of Michigan Medical School, Department of Microbiology and Immunology License: Unless otherwise noted, this material is made available under the terms of the Creative Commons Attribution
Attribution: University of Michigan Medical School, Department of Microbiology and Immunology License: Unless otherwise noted, this material is made available under the terms of the Creative Commons Attribution
Antibody Structure, and the Generation of B-cell Diversity. Chapter 4 5/1/17
 Antibody Structure, and the Generation of B-cell Diversity B cells recognize their antigen without needing an antigen presenting cell Chapter 4 Structure of Immunoglobulins Structure and function Immunoglobulin
Antibody Structure, and the Generation of B-cell Diversity B cells recognize their antigen without needing an antigen presenting cell Chapter 4 Structure of Immunoglobulins Structure and function Immunoglobulin
Immunoglobulins. Harper s biochemistry Chapter 49
 Immunoglobulins Harper s biochemistry Chapter 49 Immune system Detects and inactivates foreign molecules, viruses, bacteria and microorganisms Two components with 2 strategies B Lymphocytes (humoral immune
Immunoglobulins Harper s biochemistry Chapter 49 Immune system Detects and inactivates foreign molecules, viruses, bacteria and microorganisms Two components with 2 strategies B Lymphocytes (humoral immune
MOLECULAR RECOGNITION
 MOLECULAR RECOGNITION Bioanalytical Methods Classification 1. Biassay: molecular recognition, signal generation and detection in solution or on inert solid phase 2. Biosensor: molecular recognition system
MOLECULAR RECOGNITION Bioanalytical Methods Classification 1. Biassay: molecular recognition, signal generation and detection in solution or on inert solid phase 2. Biosensor: molecular recognition system
AB. ANTIBODY CATALYSIS
 AB. ANTIBODY CATALYSIS By 1947 it had become clear that proteins are responsible for carrying out the majority of biological functions of interest. Two common modes of action by proteins were known. Receptors
AB. ANTIBODY CATALYSIS By 1947 it had become clear that proteins are responsible for carrying out the majority of biological functions of interest. Two common modes of action by proteins were known. Receptors
T-cell response. Taken from NIAID: s.aspx
 T-cell receptor T-cell response 1. Macrophage or dendritic cell digest antigen bacteria, virus 2. Fragments of Ag bind to major histo-compatiblity (MHC) proteins in macrophage. 3. MHC I-Ag fragment expressed
T-cell receptor T-cell response 1. Macrophage or dendritic cell digest antigen bacteria, virus 2. Fragments of Ag bind to major histo-compatiblity (MHC) proteins in macrophage. 3. MHC I-Ag fragment expressed
Immune system IgGs. Carla Cortinas, Eva Espigulé, Guillem Lopez-Grado, Margalida Roig, Valentina Salas. Group 2
 Immune system IgGs Carla Cortinas, Eva Espigulé, Guillem Lopez-Grado, Margalida Roig, Valentina Salas Group 2 Index 1. Introduction 1.1. 1.2. 1.3. 1.4. 2. Immunoglobulins IgG formation IgG subclasses Structural
Immune system IgGs Carla Cortinas, Eva Espigulé, Guillem Lopez-Grado, Margalida Roig, Valentina Salas Group 2 Index 1. Introduction 1.1. 1.2. 1.3. 1.4. 2. Immunoglobulins IgG formation IgG subclasses Structural
Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and
 Supplementary Tables Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and their side chain contacting residues in the second chain (B) a Interface Res. in Contacting
Supplementary Tables Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and their side chain contacting residues in the second chain (B) a Interface Res. in Contacting
Protein homology. Antigens & Antibodies I. Administrative issues:
 Administrative issues: Recommended text: Goldsby/Kuby Immunology, 6th edition (Note that Innate Immunity is not adequately covered in the 5th edition.) Text book reading assignments are to supplement the
Administrative issues: Recommended text: Goldsby/Kuby Immunology, 6th edition (Note that Innate Immunity is not adequately covered in the 5th edition.) Text book reading assignments are to supplement the
Immunoglobulins. Structure
 Immunoglobulins Structure Definitions Immunoglobulin is a generic term that refers to a diverse group of molecules found in the blood and tissue fluids They are soluble globulin molecules and they generally
Immunoglobulins Structure Definitions Immunoglobulin is a generic term that refers to a diverse group of molecules found in the blood and tissue fluids They are soluble globulin molecules and they generally
Chapter 2. Antibodies
 Chapter 2. Antibodies An iddy-biddy antibody Just nanometers long Saved the butt of a sumo man Hundreds of kilos strong Anonymous The main elements of the immune system are firstly antibodies, secondly
Chapter 2. Antibodies An iddy-biddy antibody Just nanometers long Saved the butt of a sumo man Hundreds of kilos strong Anonymous The main elements of the immune system are firstly antibodies, secondly
Basic concepts of molecular biology
 Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it Life The main actors in the chemistry of life are molecules called proteins nucleic acids Proteins: many different
Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it Life The main actors in the chemistry of life are molecules called proteins nucleic acids Proteins: many different
Structure formation and association of biomolecules. Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München
 Structure formation and association of biomolecules Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München Motivation Many biomolecules are chemically synthesized
Structure formation and association of biomolecules Prof. Dr. Martin Zacharias Lehrstuhl für Molekulardynamik (T38) Technische Universität München Motivation Many biomolecules are chemically synthesized
Prediction of Protein-Protein Binding Sites and Epitope Mapping. Copyright 2017 Chemical Computing Group ULC All Rights Reserved.
 Prediction of Protein-Protein Binding Sites and Epitope Mapping Epitope Mapping Antibodies interact with antigens at epitopes Epitope is collection residues on antigen Continuous (sequence) or non-continuous
Prediction of Protein-Protein Binding Sites and Epitope Mapping Epitope Mapping Antibodies interact with antigens at epitopes Epitope is collection residues on antigen Continuous (sequence) or non-continuous
Humoral Immune Response. Dr. Iman Hussein Shehata Professor of Medical Microbiology and Immunology
 Humoral Immune Response Dr. Iman Hussein Shehata Professor of Medical Microbiology and Immunology Intended Learning Outcomes By the end of this lesson the student is expected to: 1-Decribe the sequence
Humoral Immune Response Dr. Iman Hussein Shehata Professor of Medical Microbiology and Immunology Intended Learning Outcomes By the end of this lesson the student is expected to: 1-Decribe the sequence
Basic Antibody Structure. Multiple myeloma = cancerous plasma cells Monomer = 150,000. Chapter 4. Immunoglobulin Structure and Function
 Chapter 4. Immunoglobulin Structure and Function. Functional Regions. Types of chains. Constant & Variable regions 4. Glycoprotein * * * Heavy chain= 446 aa Light chain= 4aa Each heavy and light chain
Chapter 4. Immunoglobulin Structure and Function. Functional Regions. Types of chains. Constant & Variable regions 4. Glycoprotein * * * Heavy chain= 446 aa Light chain= 4aa Each heavy and light chain
Molecular design principles underlying β-strand swapping. in the adhesive dimerization of cadherins
 Supplementary information for: Molecular design principles underlying β-strand swapping in the adhesive dimerization of cadherins Jeremie Vendome 1,2,3,5, Shoshana Posy 1,2,3,5,6, Xiangshu Jin, 1,3 Fabiana
Supplementary information for: Molecular design principles underlying β-strand swapping in the adhesive dimerization of cadherins Jeremie Vendome 1,2,3,5, Shoshana Posy 1,2,3,5,6, Xiangshu Jin, 1,3 Fabiana
Nucleic acid and protein Flow of genetic information
 Nucleic acid and protein Flow of genetic information References: Glick, BR and JJ Pasternak, 2003, Molecular Biotechnology: Principles and Applications of Recombinant DNA, ASM Press, Washington DC, pages.
Nucleic acid and protein Flow of genetic information References: Glick, BR and JJ Pasternak, 2003, Molecular Biotechnology: Principles and Applications of Recombinant DNA, ASM Press, Washington DC, pages.
Comparative Modeling Part 1. Jaroslaw Pillardy Computational Biology Service Unit Cornell Theory Center
 Comparative Modeling Part 1 Jaroslaw Pillardy Computational Biology Service Unit Cornell Theory Center Function is the most important feature of a protein Function is related to structure Structure is
Comparative Modeling Part 1 Jaroslaw Pillardy Computational Biology Service Unit Cornell Theory Center Function is the most important feature of a protein Function is related to structure Structure is
6-Foot Mini Toober Activity
 Big Idea The interaction between the substrate and enzyme is highly specific. Even a slight change in shape of either the substrate or the enzyme may alter the efficient and selective ability of the enzyme
Big Idea The interaction between the substrate and enzyme is highly specific. Even a slight change in shape of either the substrate or the enzyme may alter the efficient and selective ability of the enzyme
Specificity: Induced Fit
 Specificity: Induced Fit Conformational changes may occur upon ligand binding (Daniel Koshland in 1958) This adaptation is called the induced fit Induced fit allows for tighter binding of the ligand Induced
Specificity: Induced Fit Conformational changes may occur upon ligand binding (Daniel Koshland in 1958) This adaptation is called the induced fit Induced fit allows for tighter binding of the ligand Induced
Ch Biophysical Chemistry
 Ch 247.53. Biophysical Chemistry Nina Rosario L. Rojas 2012-2013 sem 1 Review of Protein Structure Why structure? Primary, secondary, tertiary structure Disulfide bonds scheme 2 STRUCTURE- REGULAR STRUCTURE
Ch 247.53. Biophysical Chemistry Nina Rosario L. Rojas 2012-2013 sem 1 Review of Protein Structure Why structure? Primary, secondary, tertiary structure Disulfide bonds scheme 2 STRUCTURE- REGULAR STRUCTURE
S uf6t<.. f\tj<t1&6t-'l
 Immunagens. An immune response is evoked by a foreign agent called antigen or immunogen. The distinction between these two terms is functional, an antigen is a compound that is capable of binding with
Immunagens. An immune response is evoked by a foreign agent called antigen or immunogen. The distinction between these two terms is functional, an antigen is a compound that is capable of binding with
 Problem: The GC base pairs are more stable than AT base pairs. Why? 5. Triple-stranded DNA was first observed in 1957. Scientists later discovered that the formation of triplestranded DNA involves a type
Problem: The GC base pairs are more stable than AT base pairs. Why? 5. Triple-stranded DNA was first observed in 1957. Scientists later discovered that the formation of triplestranded DNA involves a type
Protein-Protein Interactions I
 Biochemistry 412 Protein-Protein Interactions I March 23, 2007 Macromolecular Recognition by Proteins Protein folding is a process governed by intramolecular recognition. Protein-protein association is
Biochemistry 412 Protein-Protein Interactions I March 23, 2007 Macromolecular Recognition by Proteins Protein folding is a process governed by intramolecular recognition. Protein-protein association is
Fundamentals of Protein Structure
 Outline Fundamentals of Protein Structure Yu (Julie) Chen and Thomas Funkhouser Princeton University CS597A, Fall 2005 Protein structure Primary Secondary Tertiary Quaternary Forces and factors Levels
Outline Fundamentals of Protein Structure Yu (Julie) Chen and Thomas Funkhouser Princeton University CS597A, Fall 2005 Protein structure Primary Secondary Tertiary Quaternary Forces and factors Levels
GENETIC BASIS OF ANTIBODY STRUCTURE AND DIVERSITY. Steven J. Norris, Ph.D
 GENETIC BASIS OF ANTIBODY STRUCTURE AND DIVERSITY Steven J. Norris, Ph.D Topics I. General principles II. The heavy chain Ig locus and VDJ rearrangement III. Light chain rearrangement. IV. Mechanisms of
GENETIC BASIS OF ANTIBODY STRUCTURE AND DIVERSITY Steven J. Norris, Ph.D Topics I. General principles II. The heavy chain Ig locus and VDJ rearrangement III. Light chain rearrangement. IV. Mechanisms of
Chapter 4 ANTIBODY STRUCTURE AND FUNCTION
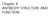 Chapter 4 ANTIBODY STRUCTURE AND FUNCTION Different way to depict an Ig molecule Y In both the heavy and light chain variable regions there is variability at every position and there are hypervariable
Chapter 4 ANTIBODY STRUCTURE AND FUNCTION Different way to depict an Ig molecule Y In both the heavy and light chain variable regions there is variability at every position and there are hypervariable
Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid
 Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
MCB 4211, Fall 2018, Practice Exam 1 Last, First name Student ID # Seat No. ***NOTE: Exam will have 40 multiple choice questions.
 MCB 4211, Fall 2018, Practice Exam 1 Last, First name Student ID # Seat No. ***NOTE: Exam 1 2018 will have 40 multiple choice questions. READ ALL THE CHOICES AND SELECT THE BEST 1. Which of the following
MCB 4211, Fall 2018, Practice Exam 1 Last, First name Student ID # Seat No. ***NOTE: Exam 1 2018 will have 40 multiple choice questions. READ ALL THE CHOICES AND SELECT THE BEST 1. Which of the following
a. Hypoxanthine was present in the media. MCB 4211, Fall 2018, Practice Exam 1 Last, First name Student ID # Seat No.
 MCB 4211, Fall 2018, Practice Exam 1 Last, First name Student ID # Seat No. ***NOTE: Exam 1 2018 will have 40 multiple choice questions. READ ALL THE CHOICES AND SELECT THE BEST 1. Which of the following
MCB 4211, Fall 2018, Practice Exam 1 Last, First name Student ID # Seat No. ***NOTE: Exam 1 2018 will have 40 multiple choice questions. READ ALL THE CHOICES AND SELECT THE BEST 1. Which of the following
Part 5 Antibody to protein recognition- structural principles in antibody design.
 Monday, January 14, 2013 Biophysics 204 Lecture Notes Robert Fletterick January 14, 2013 Part 5 Antibody to protein recognition- structural principles in antibody design. This section is about the structure,
Monday, January 14, 2013 Biophysics 204 Lecture Notes Robert Fletterick January 14, 2013 Part 5 Antibody to protein recognition- structural principles in antibody design. This section is about the structure,
Antibodies (Immunoglobulins)
 Antibodies (Immunoglobulins) The immune system plays a major role in the body s defense mechanisms against pathogens and other foreign bodies. It protects organisms from infection with a layered defense
Antibodies (Immunoglobulins) The immune system plays a major role in the body s defense mechanisms against pathogens and other foreign bodies. It protects organisms from infection with a layered defense
Basic concepts of molecular biology
 Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it What is life made of? 1665: Robert Hooke discovered that organisms are composed of individual compartments called cells
Basic concepts of molecular biology Gabriella Trucco Email: gabriella.trucco@unimi.it What is life made of? 1665: Robert Hooke discovered that organisms are composed of individual compartments called cells
ANTIBODIES. Agents of Immunity
 ANTIBODIES Agents of Immunity - Antibodies are: The Organization What are they? Protective agents of the immune system Neutralize foreign agents called antigens Essential part of the Adaptive Immune System
ANTIBODIES Agents of Immunity - Antibodies are: The Organization What are they? Protective agents of the immune system Neutralize foreign agents called antigens Essential part of the Adaptive Immune System
STRUCTURAL BIOLOGY. α/β structures Closed barrels Open twisted sheets Horseshoe folds
 STRUCTURAL BIOLOGY α/β structures Closed barrels Open twisted sheets Horseshoe folds The α/β domains Most frequent domain structures are α/β domains: A central parallel or mixed β sheet Surrounded by α
STRUCTURAL BIOLOGY α/β structures Closed barrels Open twisted sheets Horseshoe folds The α/β domains Most frequent domain structures are α/β domains: A central parallel or mixed β sheet Surrounded by α
The generation of lymphocyte antigen receptors (Chapter 5):
 The generation of lymphocyte antigen receptors (Chapter 5): 1. Ig Gene Rearrangement (somatic recombination). 2. Ig Somatic Hypermutation. 3. Ig Class Switching 4. T cell Receptor Gene Rearragement 1.
The generation of lymphocyte antigen receptors (Chapter 5): 1. Ig Gene Rearrangement (somatic recombination). 2. Ig Somatic Hypermutation. 3. Ig Class Switching 4. T cell Receptor Gene Rearragement 1.
Protocol S1: Supporting Information
 Protocol S1: Supporting Information Basis for the specificity of the kinase domain of Abl for peptide substrates The crystal structures reported in this work were obtained using two different ATP analog-peptide
Protocol S1: Supporting Information Basis for the specificity of the kinase domain of Abl for peptide substrates The crystal structures reported in this work were obtained using two different ATP analog-peptide
Antibody Structure and Function
 Antibody Structure and Function Keri C. Smith, Ph.D. January 22, 2008 (or) Anatomy and Physiology of Antibodies Overview Physical properties of antibodies Structural and molecular features Differences
Antibody Structure and Function Keri C. Smith, Ph.D. January 22, 2008 (or) Anatomy and Physiology of Antibodies Overview Physical properties of antibodies Structural and molecular features Differences
IMMU 7630 Fall 2018 ANTIBODY STRUCTURE
 IMMU 7630 Fall 2018 ANTIBODY STRUCTURE ANTIBODY IS IMMUNOGLOBULIN. Almost 130 years ago it was observed that a new activity appeared in the blood plasma of animals or humans who had been immunized with
IMMU 7630 Fall 2018 ANTIBODY STRUCTURE ANTIBODY IS IMMUNOGLOBULIN. Almost 130 years ago it was observed that a new activity appeared in the blood plasma of animals or humans who had been immunized with
Packing of Secondary Structures
 7.88 Lecture Notes - 5 7.24/7.88J/5.48J The Protein Folding and Human Disease Packing of Secondary Structures Packing of Helices against sheets Packing of sheets against sheets Parallel Orthogonal Table:
7.88 Lecture Notes - 5 7.24/7.88J/5.48J The Protein Folding and Human Disease Packing of Secondary Structures Packing of Helices against sheets Packing of sheets against sheets Parallel Orthogonal Table:
Hapten - a small molecule that is antigenic but not (by itself) immunogenic.
 Chapter 4. Antigens Terminology: Antigen: Substances that can be recognized by the surface antibody (B cells) or by the TCR when associated with MHC molecules Immunogenicity VS Antigenicity: Immunogenicity
Chapter 4. Antigens Terminology: Antigen: Substances that can be recognized by the surface antibody (B cells) or by the TCR when associated with MHC molecules Immunogenicity VS Antigenicity: Immunogenicity
Solutions to 7.02 Quiz II 10/27/05
 Solutions to 7.02 Quiz II 10/27/05 Class Average = 83 Standard Deviation = 9 Range Grade % 87-100 A 43 74-86 B 39 55-73 C 17 > 54 D 1 Question 1 (56 points) While studying deep sea bacteria, you discover
Solutions to 7.02 Quiz II 10/27/05 Class Average = 83 Standard Deviation = 9 Range Grade % 87-100 A 43 74-86 B 39 55-73 C 17 > 54 D 1 Question 1 (56 points) While studying deep sea bacteria, you discover
RNA does not adopt the classic B-DNA helix conformation when it forms a self-complementary double helix
 Reason: RNA has ribose sugar ring, with a hydroxyl group (OH) If RNA in B-from conformation there would be unfavorable steric contact between the hydroxyl group, base, and phosphate backbone. RNA structure
Reason: RNA has ribose sugar ring, with a hydroxyl group (OH) If RNA in B-from conformation there would be unfavorable steric contact between the hydroxyl group, base, and phosphate backbone. RNA structure
IMMUNOLOGY Receptors of T cells are TCR T Cell Receptors which are present on the cell surface of T lymphocytes.
 IMMUNOLOGY - 4 - What is an ANTIGEN? It is a molecule that can be recognized by a receptor and combine with it specifically and the receptor here is the one either produced by B cells or T cells: Receptors
IMMUNOLOGY - 4 - What is an ANTIGEN? It is a molecule that can be recognized by a receptor and combine with it specifically and the receptor here is the one either produced by B cells or T cells: Receptors
Immunoglobulins: Structure and Function
 Immunoglobulins: Structure and Function Immunoglobulins:Structure and Function Definition: Glycoprotein molecules that are produced by plasma cells in response to an immunogen and which function as antibodies
Immunoglobulins: Structure and Function Immunoglobulins:Structure and Function Definition: Glycoprotein molecules that are produced by plasma cells in response to an immunogen and which function as antibodies
Protein-Protein Interactions I
 Biochemistry 412 Protein-Protein Interactions I March 11, 2008 Macromolecular Recognition by Proteins Protein folding is a process governed by intramolecular recognition. Protein-protein association is
Biochemistry 412 Protein-Protein Interactions I March 11, 2008 Macromolecular Recognition by Proteins Protein folding is a process governed by intramolecular recognition. Protein-protein association is
Antibody Structure. Antibodies
 Antibodies Secreted by B lymphocytes Great diversity and specificity: >10 9 different antibodies; can distinguish between very similar molecules Tag particles for clearance/destruction Protect against
Antibodies Secreted by B lymphocytes Great diversity and specificity: >10 9 different antibodies; can distinguish between very similar molecules Tag particles for clearance/destruction Protect against
Antibody Structure supports Function
 Antibodies Secreted by B lymphocytes Great diversity and specificity: >10 9 different antibodies; can distinguish between very similar molecules Tag particles for clearance/destruction Protect against
Antibodies Secreted by B lymphocytes Great diversity and specificity: >10 9 different antibodies; can distinguish between very similar molecules Tag particles for clearance/destruction Protect against
Proteins Higher Order Structures
 Proteins Higher Order Structures Dr. Mohammad Alsenaidy Department of Pharmaceutics College of Pharmacy King Saud University Office: AA 101 msenaidy@ksu.edu.sa Previously on PHT 426!! Protein Structures
Proteins Higher Order Structures Dr. Mohammad Alsenaidy Department of Pharmaceutics College of Pharmacy King Saud University Office: AA 101 msenaidy@ksu.edu.sa Previously on PHT 426!! Protein Structures
Introduction to Antibody Structure/Function. Med Chem 528
 Introduction to Antibody Structure/Function Med Chem 528 Origins of antibodies Product of the adaptive immune system B cells (antibody based immunity) T cells (cell based immunity) Pre-exposure protects
Introduction to Antibody Structure/Function Med Chem 528 Origins of antibodies Product of the adaptive immune system B cells (antibody based immunity) T cells (cell based immunity) Pre-exposure protects
Topic (7): Antibodies and Antigens
 Topic (7): Antibodies and Antigens INTRODUCTION Antibodies (Abs) are one of the three classes of molecules able to differentiate between antigens [Ags] (the other two are T-cell receptor [TCR] and major
Topic (7): Antibodies and Antigens INTRODUCTION Antibodies (Abs) are one of the three classes of molecules able to differentiate between antigens [Ags] (the other two are T-cell receptor [TCR] and major
Bioinformatics. ONE Introduction to Biology. Sami Khuri Department of Computer Science San José State University Biology/CS 123A Fall 2012
 Bioinformatics ONE Introduction to Biology Sami Khuri Department of Computer Science San José State University Biology/CS 123A Fall 2012 Biology Review DNA RNA Proteins Central Dogma Transcription Translation
Bioinformatics ONE Introduction to Biology Sami Khuri Department of Computer Science San José State University Biology/CS 123A Fall 2012 Biology Review DNA RNA Proteins Central Dogma Transcription Translation
Kinetics of Association of Anti-lysozyme Monoclonal Antibody D44.1 and Hen-Egg Lysozyme
 2954 Biophysical Journal Volume 79 December 2000 2954 2965 Kinetics of Association of Anti-lysozyme Monoclonal Antibody D44.1 and Hen-Egg Lysozyme Gioia Altobelli* and Shankar Subramaniam* *National Center
2954 Biophysical Journal Volume 79 December 2000 2954 2965 Kinetics of Association of Anti-lysozyme Monoclonal Antibody D44.1 and Hen-Egg Lysozyme Gioia Altobelli* and Shankar Subramaniam* *National Center
11 questions for a total of 120 points
 Your Name: BYS 201, Final Exam, May 3, 2010 11 questions for a total of 120 points 1. 25 points Take a close look at these tables of amino acids. Some of them are hydrophilic, some hydrophobic, some positive
Your Name: BYS 201, Final Exam, May 3, 2010 11 questions for a total of 120 points 1. 25 points Take a close look at these tables of amino acids. Some of them are hydrophilic, some hydrophobic, some positive
Protein Folding Problem I400: Introduction to Bioinformatics
 Protein Folding Problem I400: Introduction to Bioinformatics November 29, 2004 Protein biomolecule, macromolecule more than 50% of the dry weight of cells is proteins polymer of amino acids connected into
Protein Folding Problem I400: Introduction to Bioinformatics November 29, 2004 Protein biomolecule, macromolecule more than 50% of the dry weight of cells is proteins polymer of amino acids connected into
Hmwk # 8 : DNA-Binding Proteins : Part II
 The purpose of this exercise is : Hmwk # 8 : DNA-Binding Proteins : Part II 1). to examine the case of a tandem head-to-tail homodimer binding to DNA 2). to view a Zn finger motif 3). to consider the case
The purpose of this exercise is : Hmwk # 8 : DNA-Binding Proteins : Part II 1). to examine the case of a tandem head-to-tail homodimer binding to DNA 2). to view a Zn finger motif 3). to consider the case
Andrea s SI Session PCB 3233
 Practice Test Test 2 1. A pathogen invades a tissue. Which cell of the immune system is more likely to respond first? a. Neutrophil b. T Cell c. B Cell d. Macrophage 2. The receptor for C3b is? a. CR1
Practice Test Test 2 1. A pathogen invades a tissue. Which cell of the immune system is more likely to respond first? a. Neutrophil b. T Cell c. B Cell d. Macrophage 2. The receptor for C3b is? a. CR1
4/10/2011. Rosetta software package. Rosetta.. Conformational sampling and scoring of models in Rosetta.
 Rosetta.. Ph.D. Thomas M. Frimurer Novo Nordisk Foundation Center for Potein Reseach Center for Basic Metabilic Research Breif introduction to Rosetta Rosetta docking example Rosetta software package Breif
Rosetta.. Ph.D. Thomas M. Frimurer Novo Nordisk Foundation Center for Potein Reseach Center for Basic Metabilic Research Breif introduction to Rosetta Rosetta docking example Rosetta software package Breif
Immunoglobulins. (1 of 2)
 Immunoglobulins (1 of 2) Immunoglobulins (Igs) = antibodies Each B cell synthesizes Igs of single specificity for a specific epitope B cell receptors (BCRs) are the Igs on B cell surface Humoral immunity
Immunoglobulins (1 of 2) Immunoglobulins (Igs) = antibodies Each B cell synthesizes Igs of single specificity for a specific epitope B cell receptors (BCRs) are the Igs on B cell surface Humoral immunity
Biomolecular chemistry. 7. Antibodies: structure and function
 154 Biomolecular chemistry 7. Antibodies: structure and function Suggested reading: Sections 5.1 to 5.3 of Mikkelsen and Cortón, Bioanalytical Chemistry Primary Source Material Biochemistry Chapter 33:
154 Biomolecular chemistry 7. Antibodies: structure and function Suggested reading: Sections 5.1 to 5.3 of Mikkelsen and Cortón, Bioanalytical Chemistry Primary Source Material Biochemistry Chapter 33:
Structural bioinformatics
 Structural bioinformatics Why structures? The representation of the molecules in 3D is more informative New properties of the molecules are revealed, which can not be detected by sequences Eran Eyal Plant
Structural bioinformatics Why structures? The representation of the molecules in 3D is more informative New properties of the molecules are revealed, which can not be detected by sequences Eran Eyal Plant
Algorithms in Bioinformatics ONE Transcription Translation
 Algorithms in Bioinformatics ONE Transcription Translation Sami Khuri Department of Computer Science San José State University sami.khuri@sjsu.edu Biology Review DNA RNA Proteins Central Dogma Transcription
Algorithms in Bioinformatics ONE Transcription Translation Sami Khuri Department of Computer Science San José State University sami.khuri@sjsu.edu Biology Review DNA RNA Proteins Central Dogma Transcription
2018 Protein Modeling Exam Key
 2018 Protein Modeling Exam Key Multiple Choice: 1. Which of the following amino acids has a negative charge at ph 7? a. Gln b. Glu c. Ser d. Cys 2. Which of the following is an example of secondary structure?
2018 Protein Modeling Exam Key Multiple Choice: 1. Which of the following amino acids has a negative charge at ph 7? a. Gln b. Glu c. Ser d. Cys 2. Which of the following is an example of secondary structure?
Conformational changes in IgE contribute to its. uniquely slow dissociation rate from receptor FcεRI
 Conformational changes in IgE contribute to its uniquely slow dissociation rate from receptor FcεRI M.D. Holdom, A.M. Davies, J.E. Nettleship, S.C. Bagby, B. Dhaliwal, E. Girardi, J. Hunt, H.J. Gould,
Conformational changes in IgE contribute to its uniquely slow dissociation rate from receptor FcεRI M.D. Holdom, A.M. Davies, J.E. Nettleship, S.C. Bagby, B. Dhaliwal, E. Girardi, J. Hunt, H.J. Gould,
Introduction to Proteins
 Introduction to Proteins Lecture 4 Module I: Molecular Structure & Metabolism Molecular Cell Biology Core Course (GSND5200) Matthew Neiditch - Room E450U ICPH matthew.neiditch@umdnj.edu What is a protein?
Introduction to Proteins Lecture 4 Module I: Molecular Structure & Metabolism Molecular Cell Biology Core Course (GSND5200) Matthew Neiditch - Room E450U ICPH matthew.neiditch@umdnj.edu What is a protein?
In mice: V H ~250 D ~15 J ~5. L 1 -V H1 -L 2 -V H2 -L 3 -V H3 ---L n -V Hn ----D 1 -D 2 -D 3 ---D n ----J 1 -J J n V L ~250 J ~4
 Lecture 14 ymes Cont. ymes preferentially stabilize transitions states versus ground states ite directed mutagenesis (Tyr ta synthetase) Transition state analogue inhibitors Design of protein active sites
Lecture 14 ymes Cont. ymes preferentially stabilize transitions states versus ground states ite directed mutagenesis (Tyr ta synthetase) Transition state analogue inhibitors Design of protein active sites
LECTURE: 22 IMMUNOGLOBULIN DIVERSITIES LEARNING OBJECTIVES: The student should be able to:
 LECTURE: 22 Title IMMUNOGLOBULIN DIVERSITIES LEARNING OBJECTIVES: The student should be able to: Identify the chromosome that contains the gene segments that encode the surface immunoglobulin heavy chain
LECTURE: 22 Title IMMUNOGLOBULIN DIVERSITIES LEARNING OBJECTIVES: The student should be able to: Identify the chromosome that contains the gene segments that encode the surface immunoglobulin heavy chain
Chapter 5. Genetic Models. Organization and Expression of Immunoglobulin Genes 3. The two-gene model: Models to Explain Antibody Diversity
 Chapter 5 Organization and Expression of Immunoglobulin Genes 3 4 5 6 Genetic Models How to account for: ) Vast diversity of antibody specificities ) Presence of Variable regions at the amino end of Heavy
Chapter 5 Organization and Expression of Immunoglobulin Genes 3 4 5 6 Genetic Models How to account for: ) Vast diversity of antibody specificities ) Presence of Variable regions at the amino end of Heavy
Antibodies and Antigens in the Blood Bank 9/7/2015 NAHLA BAKHAMIS 1
 Antibodies and Antigens in the Blood Bank NAHLA BAKHAMIS 9/7/2015 NAHLA BAKHAMIS 1 Outline Antibodies structure, classes and functions Most important Abs in the blood bank effective roles of Abs Zeta potential
Antibodies and Antigens in the Blood Bank NAHLA BAKHAMIS 9/7/2015 NAHLA BAKHAMIS 1 Outline Antibodies structure, classes and functions Most important Abs in the blood bank effective roles of Abs Zeta potential
Protein Structure Prediction by Constraint Logic Programming
 MPRI C2-19 Protein Structure Prediction by Constraint Logic Programming François Fages, Constraint Programming Group, INRIA Rocquencourt mailto:francois.fages@inria.fr http://contraintes.inria.fr/ Molecules
MPRI C2-19 Protein Structure Prediction by Constraint Logic Programming François Fages, Constraint Programming Group, INRIA Rocquencourt mailto:francois.fages@inria.fr http://contraintes.inria.fr/ Molecules
Antibodies and Antigens In the blood bank
 Antibodies and Antigens In the blood bank 1 Nice game!! http://nobelprize.org/ 2 Karl Landsteiner discovered blood groups in 1901. Awarded Nobel Prize for Physiology or Medicine in 1930 3 Why we study
Antibodies and Antigens In the blood bank 1 Nice game!! http://nobelprize.org/ 2 Karl Landsteiner discovered blood groups in 1901. Awarded Nobel Prize for Physiology or Medicine in 1930 3 Why we study
Τάσος Οικονόµου ιαλεξη 8. Kινηση, λειτουργια, ελεγχος.
 Τάσος Οικονόµου ιαλεξη 8 Kινηση, λειτουργια, ελεγχος http://ecoserver.imbb.forth.gr/bio321.htm εν ξεχνω. Cell The peptide bond Polypeptides are stabilized by: 1. Covalent bonds= amide bond 2. Noncovalent,
Τάσος Οικονόµου ιαλεξη 8 Kινηση, λειτουργια, ελεγχος http://ecoserver.imbb.forth.gr/bio321.htm εν ξεχνω. Cell The peptide bond Polypeptides are stabilized by: 1. Covalent bonds= amide bond 2. Noncovalent,
SUPPLEMENTARY INFORMATION
 ARTICLE NUMBER: 16054 DOI: 10.1038/NMICROBIOL.2016.54 Microbially cleaved immunoglobulins are sensed by the innate immune receptor LILRA2 Kouyuki Hirayasu, Fumiji Saito, Tadahiro Suenaga, Kyoko Shida,
ARTICLE NUMBER: 16054 DOI: 10.1038/NMICROBIOL.2016.54 Microbially cleaved immunoglobulins are sensed by the innate immune receptor LILRA2 Kouyuki Hirayasu, Fumiji Saito, Tadahiro Suenaga, Kyoko Shida,
Antibodies. Immunoglobulin (Ig) is a synonym for antibody. Most antibodies are found in the gamma globulin fraction of serum.
 Antibodies Introduction Antibodies are a class of serum proteins which are induced following contact with antigen. They bind specifically with antigen which induced their formation. Immunoglobulin (Ig)
Antibodies Introduction Antibodies are a class of serum proteins which are induced following contact with antigen. They bind specifically with antigen which induced their formation. Immunoglobulin (Ig)
A therapeutic battle: Antibodies vs. Aptamers
 NS109 - Research paper Nanoscience master program A therapeutic battle: Antibodies vs. Aptamers Jan Hidding* 1 Abstract The goal of creating drugs that specifically target their pathogen has been around
NS109 - Research paper Nanoscience master program A therapeutic battle: Antibodies vs. Aptamers Jan Hidding* 1 Abstract The goal of creating drugs that specifically target their pathogen has been around
produces an RNA copy of the coding region of a gene
 1. Transcription Gene Expression The expression of a gene into a protein occurs by: 1) Transcription of a gene into RNA produces an RNA copy of the coding region of a gene the RNA transcript may be the
1. Transcription Gene Expression The expression of a gene into a protein occurs by: 1) Transcription of a gene into RNA produces an RNA copy of the coding region of a gene the RNA transcript may be the
OpenStax-CNX module: m Antibodies * OpenStax. Abstract
 OpenStax-CNX module: m44823 1 Antibodies * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you will be able to:
OpenStax-CNX module: m44823 1 Antibodies * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you will be able to:
3D Structure of Biologics in a Convenient Immunoassay Format
 3D Structure of Biologics in a Convenient Immunoassay Format Xing Wang, Ph.D. Array Bridge Inc. 5/25/2017 1 Topics Covered Today Why New Technologies? Technology Development. Case Studies for Novel and
3D Structure of Biologics in a Convenient Immunoassay Format Xing Wang, Ph.D. Array Bridge Inc. 5/25/2017 1 Topics Covered Today Why New Technologies? Technology Development. Case Studies for Novel and
Virulence factors: name them and explain what they do, how do you calculate how virulent something is
 General Microbiology Final Exam Study Guide Spring 2017 Ginny L. Please bring various colored writing utensils! Fair warning, I am not a TA or teacher and I have not seen your final exam, this is to cover
General Microbiology Final Exam Study Guide Spring 2017 Ginny L. Please bring various colored writing utensils! Fair warning, I am not a TA or teacher and I have not seen your final exam, this is to cover
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
CHAPTER 3 ANTIBODY STRUCTURE I
 CHAPTER 3 ANTIBODY STRUCTURE I See APPENDIX: (3) OUCHTERLONY ANALYSIS; (6), EQUILIBRIUM DIALYSIS; (7) CROSS-REACTIVITY Electrophoretic separation of serum proteins identifies the GAMMA-GLOBULIN fraction
CHAPTER 3 ANTIBODY STRUCTURE I See APPENDIX: (3) OUCHTERLONY ANALYSIS; (6), EQUILIBRIUM DIALYSIS; (7) CROSS-REACTIVITY Electrophoretic separation of serum proteins identifies the GAMMA-GLOBULIN fraction
Proteins: Wide range of func2ons. Polypep2des. Amino Acid Monomers
 Proteins: Wide range of func2ons Proteins coded in DNA account for more than 50% of the dry mass of most cells Protein func9ons structural support storage transport cellular communica9ons movement defense
Proteins: Wide range of func2ons Proteins coded in DNA account for more than 50% of the dry mass of most cells Protein func9ons structural support storage transport cellular communica9ons movement defense
Nature inspired design of motif specific antibody scaffolds *
 Nature inspired design of motif specific antibody scaffolds * Nature Biotechnology, October 2013, Vol. 31, #10 * Scientific information in the following slides is from this article, if not stated otherwise
Nature inspired design of motif specific antibody scaffolds * Nature Biotechnology, October 2013, Vol. 31, #10 * Scientific information in the following slides is from this article, if not stated otherwise
AdvanceBio HIC: a Hydrophobic HPLC Column for Monoclonal Antibody (mab) Variant Analysis
 Application Note Biologics Development AdvanceBio HIC: a Hydrophobic HPLC Column for Monoclonal Antibody (mab) Variant Analysis Using the Agilent 16 Infinity II Bio-Inert LC Authors Andrew Coffey and Sandeep
Application Note Biologics Development AdvanceBio HIC: a Hydrophobic HPLC Column for Monoclonal Antibody (mab) Variant Analysis Using the Agilent 16 Infinity II Bio-Inert LC Authors Andrew Coffey and Sandeep
Antibody - Antigen Reactions: ABO and D typing Antibody screening and identification
 Antibody - Antigen Reactions: ABO and D typing Antibody screening and identification Basics of antigen/ antibody reactions Why is the ABO group so special? D antigen it s complicated! Antibody screen Antibody
Antibody - Antigen Reactions: ABO and D typing Antibody screening and identification Basics of antigen/ antibody reactions Why is the ABO group so special? D antigen it s complicated! Antibody screen Antibody
Adaptive Immunity: Specific Defenses of the Host
 PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College C H A P T E R 17 Adaptive Immunity: Specific Defenses of the Host The Adaptive Immune System Adaptive immunity:
PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College C H A P T E R 17 Adaptive Immunity: Specific Defenses of the Host The Adaptive Immune System Adaptive immunity:
Generation of Recombinant Antibodies and Means for Increasing Their Affinity
 ISSN 0006-2979, Biochemistry (Moscow), 2010, Vol. 75, No. 13, pp. 1584-1605. Pleiades Publishing, Ltd., 2010. Original Russian Text E. P. Altshuler, D. V. Serebryanaya, A. G. Katrukha, 2010, published
ISSN 0006-2979, Biochemistry (Moscow), 2010, Vol. 75, No. 13, pp. 1584-1605. Pleiades Publishing, Ltd., 2010. Original Russian Text E. P. Altshuler, D. V. Serebryanaya, A. G. Katrukha, 2010, published
Building an Antibody Homology Model
 Application Guide Antibody Modeling: A quick reference guide to building Antibody homology models, and graphical identification and refinement of CDR loops in Discovery Studio. Francisco G. Hernandez-Guzman,
Application Guide Antibody Modeling: A quick reference guide to building Antibody homology models, and graphical identification and refinement of CDR loops in Discovery Studio. Francisco G. Hernandez-Guzman,
Protein Structure Analysis
 BINF 731 Protein Structure Analysis http://binf.gmu.edu/vaisman/binf731/ Secondary Structure: Computational Problems Secondary structure characterization Secondary structure assignment Secondary structure
BINF 731 Protein Structure Analysis http://binf.gmu.edu/vaisman/binf731/ Secondary Structure: Computational Problems Secondary structure characterization Secondary structure assignment Secondary structure
Custom Antibody Services. Antibodies Designed Just for You. HuCAL Recombinant Monoclonal Antibody Generation Service
 Custom Antibody Services Antibodies Designed Just for You HuCAL Recombinant Monoclonal Antibody Generation Service Antibodies Designed Just for You What if there was a technology that would allow you to
Custom Antibody Services Antibodies Designed Just for You HuCAL Recombinant Monoclonal Antibody Generation Service Antibodies Designed Just for You What if there was a technology that would allow you to
1 Name. 1. (3 pts) What is apoptosis and how does it differ from necrosis? Which is more likely to trigger inflammation?
 1 Name MCB 150 Midterm Eam #1 (100 points total) Please write your full name on each page of the eam!! The eam consists of 17 questions (6 pages). Each has a different point count as indicated. Please
1 Name MCB 150 Midterm Eam #1 (100 points total) Please write your full name on each page of the eam!! The eam consists of 17 questions (6 pages). Each has a different point count as indicated. Please
