Supporting Information
|
|
|
- Rose Gallagher
- 6 years ago
- Views:
Transcription
1 Supporting Information Zammarchi et al /pnas SI Materials and Methods STAT3 Minigene Construction. Human genomic DNA (Promega) was amplified by Platinum Pfx (Invitrogen) as per instructions for 35 cycles with an annealing temperature of 60 C and a 3-min extension time using STAT3-specific primers (I22F and I24R) with overhanging restriction sites for KpnI and BamHI (Promega), respectively. The amplified region spans the entire exon 22 (43 nt), intron 22 (280 nt), exon 23 (113 nt), intron 23 (988 nt), and 985 nt of exon 24, which include the full-length stop codon at position 56 of the exon. The entire exon 24, including the full UTR, is 2,477-nt long. The resulting PCR product was subcloned into pcdna3.1+ (Invitrogen). The ligation reaction was used to transform MAX Efficiency DH5a-competent cells, and colony PCR was used to screen for positive colonies that were then grown overnight at 37 C. Plasmid DNA was prepared (Qiagen) and the resulting vector was confirmed by restriction enzyme digestion and sequencing. Primer Sequences for STAT3 Minigene Construction. Introns 22F (I22F) 5 -CGACGGTACCTCCAGCTCTGCTTACTGAATGC- GA-3 Introns 24R (I24R) 5 -CGGTGGATCCGCTTAAAGCAC- CAAGGAGGCTGTT-3 Minigene Mutant Construction. Mutants were prepared by sitespecific mutagenesis using overlap extensions (1). Briefly, complementary PCR primers forward mutant (FM) and reverse mutant (RM) were designed to have either a deletion or a point mutation within the primer, flanked by at least 14 bp on each side. Two PCR reactions were performed using the FM with I24R reverse primer and RM with I22F forward primer. The resulting PCR products were gel-purified, and equal amounts were used as a template for a second PCR, with I22F and I24R primers. All PCR was performed with Platinum Pfx (Invitrogen), with an annealing temperature of 60 C and an extension time 1 min/kb as per instructions. The resulting products were then subcloned as previously described and verified by sequencing. Primer Sequences for STAT3 Minigene Deletion Mutants in WT Context. ΔA-F 5 -CTGCCCCCCATTTCCTACAGAAGCAATACCAT- TGACCTGCCGATG-3 ΔA-R 5 -CATCGGCAGGTCAATGGTATTGCTTCTGTAG- GAAATGGGGGGCAG-3 ΔB-F 5 -CCCCATTTCCTACAGAACGACCACCATTGAC- CTGCCGATGTCC-3 ΔB-R 5 -GGACATCGGCAGGTCAATGGTGGTCGTTCT- GTAGGAAATGGGG-3 ΔC-F 5 -CATTTCCTACAGAACGACCTGCAGCGACCTG- CCGATGTCCCC-3 ΔC-R 5 -GGGGACATCGGCAGGTCGCTGCAGGTCGTT- CTGTAGGAAATG-3 ΔD-F 5 -CTACAGAACGACCTGCAGCAATACCCCGATG- TCCCCCCGC-3 ΔD-R 5 -GCGGGGGGACATCGGGGTATTGCTGCAGG- TCGTTCTGTAG-3 ΔE-F 5 -ACGACCTGCAGCAATACCATTGACTCCCCCC- GCACTTTAGATTCATTGATG-3 ΔE-R 5 -CATCAATGAATCTAAAGTGCGGGGGGAGTC- AATGGTATTGCTGCAGGTCGT-3 ΔF-F 5 -GCAGCAATACCATTGACCTGCCGCGCACTTT- AGATTCATTGATGCAGTTTGGA-3 ΔF-R 5 -TCCAAACTGCATCAATGAATCTAAAGTGCG- CGGCAGGTCAATGGTATTGCTGC-3 ΔG-F 5 -AATACCATTGACCTGCCGATGTCCTTAGAT- TCATTGATGCAGTTTGGAAATAATGGTGAA-3 ΔG-R 5 -TTCACCATTATTTCCAAACTGCATCAATGAA- TCTAAGGACATCGGCAGGTCAATGGTATT-3 ΔA1-F 5 -CTGCCCCCCATTTCCTACAGAACCTGCAGC- AATACCATTGACCTG-3 ΔA1-R 5 -CAGGTCAATGGTATTGCTGCAGGTTCTGT- AGGAAATGGGGGGCAG-3 ΔA2-F 5 -TGCCCCCCATTTCCTACAGAACGTGCAGCA- ATACCATTGACCTGCC-3 ΔA2-R 5 -GGCAGGTCAATGGTATTGCTGCACGTTCTG- TAGGAAATGGGGGGCA-3 ΔA3-F 5 -GCCCCCCATTTCCTACAGAACGACCAGCAA- TACCATTGACCTGCCGATG-3 ΔA3-R 5 -CATCGGCAGGTCAATGGTATTGCTGGTCG- TTCTGTAGGAAATGGGGGGC-3 ΔB2-F 5 -CCCATTTCCTACAGAACGACCTGCAATACC- ATTGACCTGCCGATGTCCC-3 ΔB2-R 5 -GGGACATCGGCAGGTCAATGGTATTGCAG- GTCGTTCTGTAGGAAATGGG-3 ΔB3-F 5 -CATTTCCTACAGAACGACCTGCAGCACCAT- TGACCTGCCGATGTCC-3 ΔB3-R 5 -GGACATCGGCAGGTCAATGGTGCTGCAGG- TCGTTCTGTAGGAAATG-3 ΔH-F 5 -GCCGATGTCCCCCCGCACTTTAGAGTTTGGA- AATAATGGTGAAGGTGCTGAACCC-3 ΔH-R 5 -GGGTTCAGCACCTTCACCATTATTTCCAAAC- TCTAAAGTGCGGGGGGACATCGGC-3 ΔK-F 5 -CCCCCCGCACTTTAGATTCATTGATTAATGG- TGAAGGTGCTGAACCCTC-3 ΔK-R 5 -GAGGGTTCAGCACCTTCACCATTAATCAATG- AATCTAAAGTGCGGGG-3 ΔL-F 5 -CGCACTTTAGATTCATTGATGCAGTTTGGAG- GTGCTGAACCCTCAGCAG-3 ΔL-R 5 -CTGCTGAGGGTTCAGCACCTCCAAACTGCA- TCAATGAATCTAAAGTGCG-3 ΔM-F 5 -GCACTTTAGATTCATTGATGCAGTTTGGAA- ATAATGGACCCTCAGCAGGAGGGC-3 ΔM-R 5 -GCCCTCCTGCTGAGGGTCCATTATTTCCAA- ACTGCATCAATGAATCTAAAGTGC-3 ΔN-F 5 -GATGCAGTTTGGAAATAATGGTGAAGGTGC- AGGAGGGCAGTTTGGTGAGTATTTG-3 ΔN-R 5 -CAAATACTCACCAAACTGCCCTCCTGCACCT- TCACCATTATTTCCAAACTGCATC-3 ΔO-F 5 -GAAATAATGGTGAAGGTGCTGAACCCTCG- TTTGGTGAGTATTTGGTTGACAGACTTTGTC-3 ΔO-R 5 -GACAAAGTCTGTCAACCAAATACTCACCAA- ACGAGGGTTCAGCACCTTCACCATTATTTC-3 ΔH1-F 5 -CCGCACTTTAGAATTGATGCAGTTTGGAA-3 ΔH1-R 5 -TTCCAAACTGCATCAATTCTAAAGTGCGG-3 ΔH2-F 5 -CCGCACTTTAGATTCGATGCAGTTTGGA-3 ΔH2-R 5 -TCCAAACTGCATCGAATCTAAAGTGCGG-3 ΔH3-F 5 -CCGCACTTTAGATTCATTGCAGTTTGGAA-3 ΔH3-R 5 -TTCCAAACTGCAATGAATCTAAAGTGCGG-3 ΔK1-F 5 -CCGCACTTTAGATTCATTGATGTTTGGAAA- TAATGG-3 ΔK1-R 5 -CCATTATTTCCAAACATCAATGAATCTAAA- GTGCGG-3 ΔK2-F 5 -CCGCACTTTAGATTCATTGATGCATGGAAA- TA-3 1of10
2 ΔK2-R 5 -TATTTCCATGCATCAATGAATCTAAAGTGC- GG-3 ΔK3-F 5 -CGCACTTTAGATTCATTGATGCAGTTAAATA- ATGGTG-3 ΔK3-R 5 -CACCATTATTTAACTGCATCAATGAATCTA- AAGTGCG-3 Py^-F 5 -ATGTCCCCCCTCCCTTTAGATTCATTG-3 Py^-R 5 -CAATGAATCTAAAGGGAGGGGGGACAT-3 Transfection of Minigenes. Minigene transfections into HeLa cells were done with FuGENE 6 (Roche) as per instructions. Twentyfour hours after transfection, cells were collected for RNA extraction. RNA Extraction and RT-PCR. Total RNA was extracted using either the RNAqueous Kit (Ambion) (for cell lines) or TRIzol reagent (Invitrogen) (for tumor samples) according to the manufacturers instructions. Contaminating DNA was removed using TURBO DNA-free (Ambion) following product instruction. Total RNA was then reverse-transcribed with SuperScript III RT (Invitrogen) and the resulting cdna was used as template for PCR analysis. All PCR reactions were performed with Platinum Taq (Invitrogen) as per instructions with an annealing temperature of 60 C and an extension time of 30 s for a total of 30 cycles. Primers used to amplify STAT3 and GAPDH were as follows. STAT3 α/β splicing (minigene). Forward 5 -GCTAGCGTTTAAACTTAAGCTTGG-3 Reverse 5 -ACTCCGAGGTCAACTCCATGTCAA-3 STAT3 α/β splicing (endogenous). Forward 5 -TGCTACCAATATCCTGGTGTCTCC-3 Reverse 5 -ACTCCGAGGTCAACTCCATGTCAA-3 STAT3 exon 6 skipping. Forward 5 -TGGTGACGGAGAAGCAGCAGAT-3 Reverse 5 -TGCACGTACTCCATCGCTGACAAA-3 GAPDH primers. F5 -TGCACCACCAACTGCTTAGC-3 R5 -GGCATGGACTGTGGTCATGAG-3 Cell Culture. MDA-MB-435s, MDA-MB-468, MDA-MB-453, HeLa, A549,DU145,K562,4T1,andB16cellswerepurchasedfromthe American Type Culture Collection. Phoenix-Ampho packaging cells were a generous gift of Scott Lowe (Cold Spring Harbor Laboratory, Cold Spring Harbor, NY). Human breast cancer cell lines were grown in a mixture (1:1, vol/vol) of DMEM and Ham s F-12 medium supplemented with 1% nonessential amino acids (Invitrogen), 1% sodium pyruvate (Invitrogen), 1% penicillin/ streptomycin (Invitrogen), and 10% FBS (Sigma). HeLa and Phoenix-Ampho cells were grown in DMEM supplemented with 1% penicillin/streptomycin (Invitrogen) and 10% FBS (Sigma). A549, DU145, K562, 4T1, and B16 were grown in RPMI 1640 media supplemented with 1% penicillin/streptomycin (Invitrogen) and10%fbs(sigma).cellswereincubatedat37 Cina5%CO 2 atmosphere. Antibodies. Total-STAT3 (Cell Signaling; 9132), phospho-stat3 Y705 (Cell Signaling; 9131), total-stat1 (BD Biosciences; ), lens epithelium-derived growth factor (LEDGF) (BD Biosciences; ), peroxisomal biogenesis factor 1 (PEX1) (BD Biosciences; ), p300/cbp-associated factor (PCAF) (Cell Signaling; 3378), α-tubulin (Sigma; T5168), and actin (Sigma; A4700). Western Blot. Whole-cell lysates were prepared using M-PER protein extraction buffer (Thermo Scientific) with Complete Protease Inhibitor Mixture (Roche) as per instructions. Protein concentration was determined using the Bradford assay (Bio- Rad). Samples were subjected to an 8% SDS/PAGE, transferred to an Immobilon-FL PVDF membrane (Millipore), and blotted in 5% milk for 2 h at room temperature. Blots were incubated overnight at 4 C with the respective primary antibodies. Next, the blots were washed, incubated with the corresponding HRPconjugated secondary antibody, washed again, and detected using SuperSignal West Femto Substrate (Pierce). Morpholino Treatments. Morpholino oligos (Gene Tools) were mixed at the desired concentration with media plus 4% FBS and added to the cells. Endo-Porter (Gene Tools) was used as delivery reagent at a final concentration of 6 μm. Vivo-Morpholino oligos (2) were either intratumorally injected at a final concentration of 0.12 mg/30 μl or administered intravenously at 15 mg/kg. Treatments were administered twice a week for either 1 or 3 wk. Sequences of morpholinos were as follows. ST1, 5 -CTGCAGGTCGTTCTGTAGGAAAT-3 ST2, 5 -ATTGCTGCAGGTCGTTCTGTAGG-3 ST3, 5 -GGTATTGCTGCAGGTCGT-3 ST6, 5 -CATTTTCTGTTCTAGATCCTGTT-3 ST7, 5 -TTCACTTGCCTCCTTGACTCTTG-3 INV, 5 -GGATGTCTTGCTGGACGTCGTTA-3 Small Interfering RNA. Oligofectamine (Invitrogen) was used to transfect sirna into MDA-MB-435s cells as per instructions. sirnas were purchased from the HTSCF (High-Throughput Screening Core facility) (Memorial Sloan-Kettering Cancer Center). As a control, we used an sirna oligo directed against GFP. sirnas against PEX1, PCAF, and CyclinC (CCNC) were used at a final concentration of 100 nm each and compared with MDA-MB-435s cells treated with 100 nm GFP sirna oligo. sirna against LEDGF was used at a final concentration of 50 nm and compared with MDA-MB-435s cells treated with 50 nm GFP sirna oligo. Each treatment was done in triplicate. Cells were treated for 72 h and then collected for quantitative (q)pcr/ Western blot and trypan blue analysis. Trypan Blue. Cells were treated with either the morpholino oligos or with the sirna oligos as described above. At the end of the treatment, cells were trypsinized, counted with a hemocytometer, and scored as dead/total. Data were analyzed by calculating the fold change in dead/total cells compared with the control treatment. Apoptosis and Cell-Cycle Analysis. Apoptosis was determined by annexin V/propidium iodide (AnxV/PI) staining. Cells ( ) were resuspended in AnxV/PI buffer (Annexin V Binding Buffer; Biolegend). Alexa Fluor 647-conjugated annexin V (Biolegend) was added to the cell suspension and incubated for 15 min at room temperature. Cells were further resuspended in AnxV/PI buffer, and PI (Sigma-Aldrich; 1 μg/ml) was added before flow analysis by Beckman Coulter cyan flow cytometer. For cell-cycle analysis, harvested cells ( ) were washed twice with PBS + 0.5% BSA and fixed in cold 70% (vol/vol) ethanol for 30 min. Cells were then washed twice and further treated with RNase (100 μg/ml; Sigma-Aldrich) and PI (50 μg/ ml). Samples were incubated at 37 C for 15 min and analyzed by flow cytometry. Data were analyzed using FlowJo software (Tree Star). ELISA. IL8 levels in MDA-MB-435s supernatants were measured with an IL8 ELISA Kit (Quantikine; R&D Systems) according to the manufacturer s protocol. Cell Proliferation Assay. Cells were plated in 96-well plates for 24 h and then treated with either 1 μm pan-jak inhibitor P6 (a tetracyclic pyridone; 2-tert-butyl-9-fluoro-3,6-dihydro-7H-benz[h]- imidaz[4,5-f]isoquinoline-7-one, pyridone 6) (3) or DMSO in association with 6 μm Vivo-Morpholino control oligonucleotide 2of10
3 inverted sequence (INV) or with just 6 μm Vivo-Morpholino oligonucleotides INV, ST2, and ST6 for 48 h. A CellTiter 96 AQ ueous One Solution Cell Proliferation Assay (Promega) was performed according to the manufacturer s instructions. Absorbance was measured at 490 nm using a microplate reader (Bio- Rad). TUNEL. TUNEL assay was carried out according to the manufacturer s instructions (Roche). Immunohistochemistry. Immunohistochemical staining was carried out on 5-μm sections from formalin-fixed, paraffin-embedded tumor specimens. Following routine deparaffinization and rehydration, the sections were subjected to staining with phospho- STAT3 (Y705: Cell Signaling, 9131; S727: Epitomics, ), total-stat3 (Cell Signaling; 9132), and caspase 3 (Cell Signaling; 9661) antibodies using the automated staining processor Discovery from Ventana Medical Systems. Quantification of Immunohistochemical Staining. TUNEL and cleaved caspase 3-stained cells were estimated in 10 randomly selected fields of tumor at 40 magnification, and the percentage of positive tumor cells was calculated as number of positive cells/ total number of cells 100. Affymetrix Oligonucleotide Microarray. MDA-MB-435s cells were treated with 16 μm ST2, ST6, or INV morpholino for 4 d and total RNA was extracted at the end of treatment. The quality of the RNA was ensured before labeling by analyzing ng of each sample using the RNA 6000 NanoAssay and a Bioanalyzer 2100 (Agilent). Samples with a 28S:18S ribosomal peak ratio of were considered suitable for labeling. For samples meeting this standard, 2 μg of total RNA was used for cdna synthesis using an oligo (dt)-t7 primer and the SuperScript Double-Stranded cdna Synthesis Kit (Invitrogen). Synthesis, linear amplification, and labeling of crna were accomplished by transcription in vitro using the MessageAmp arna Kit (Ambion) and biotinylated nucleotides (Enzo Diagnostics). Ten micrograms of labeled and fragmented crna was then hybridized to the mouse genome U133 plus 2.0 array (Affymetrix), which interrogates 39,000 transcripts, at 45 C for 16 h. Automated washing and staining were performed using the Affymetrix Fluidics Station 400 according to the manufacturer s protocols. Finally, chips were scanned with a high-numerical aperture and flying objective lens in the GS3000 scanner (Affymetrix). Raw expression data were analyzed using GCOS 1.4 (Affymetrix). Data were normalized to a target intensity of 500 to account for differences in global chip intensity. Real-Time Quantitative PCR. Primers for qpcr were designed for use with an annealing temperature of 60 C and a product size of less than 150 nucleotides through the oligo-design program at When possible, primers were designed to span exon exon junctions to prevent the amplification of contaminating genomic DNA. PCR reactions were performed per SYBR Green instructions (Invitrogen). Primers initially were tested for linearity with a template curve from 200 ng to 2 pg (10- fold dilutions) with retro-transcribed Universal Human RNA (Stratagene) and by checking the PCR product size on an agarose gel stained with ethidium bromide. A single gel band and a single peak on the melting curve were verified initially and used to confirm the PCR product thereafter. The ddc(t) method was used for qpcr determination (4). Hypoxanthine phosphoribosyl transferase (HPRT) and β-2-microglobulin (B2M) were used as housekeeping genes. Real-Time Primer Sequences. EGR1. F5 -AGCCCTACGAGCACCTGAC-3 R5 -AGCGGCCAGTATAGGTGATG-3 FAS. F5 -ATAAGCCCTGTCCTCCAGGTGAAA-3 R5 -ATTCTGGGTCCGGGTGCAGTTTAT-3 Hsp27. F5 -AAGGATGGCGTGGTGGAGATCA-3 R5 -TGATCTCGTTGGACTGCGTGGCTA-3 SOCS3. F5 -GAAGATCCCCCTGGTGTTGA-3 R5 -TTCCGACAGAGATGCTGAAGAGT-3 VEGF. F5 -TTTCTGCTGTCTTGGGTGCATTGG-3 R5 -AAGATGTCCACCAGGGTCTCGATT-3 IL8. F5 -AGAAACCACCGGAAGGAACCATCT-3 R5 -AATTTCTGTGTTGGCGCAGTGTGG-3 LEDGF. F5 -GTGTTTCAAAGGAAGATACCGACC-3 R5 -TTGCTGTTGTCACTACTCCTGCCT-3 CCNC. F5 -CAAGATCTGTTGAAGGAGCGCCAA-3 R5 -CTTGAAATATACCGTAGCAGTGGC-3 STAT1β. F5 -AGCCAATGCAACTTGATGGCCCTA-3 R5 -TCAACAAGTTCAGCTGTGATGGCG-3 PCAF. F5 -GGCAGACTTACAGCGAGTCTTTACCA-3 R5 -TTGCTTTAGGCACACTGCTTGGTG-3 PEX1. F5 -AGCTGCATGCTGTTTCCCTTGAAC-3 R5 -TATTCTCTTTGGCTCGGCGTGTCT-3 SURVIVIN. F5 -TTCGTCCGGTTGCGCTTTCC-3 R5 -TCGATGGCACGGCGCACTTT-3 B2M. F5 -GGCTATCCAGCGTACTCCAAA-3 R5 -CGGCAGGCATACTCATCTTTTT-3 HPRT. F5 -GACCAGTCAACAGGGGACAT-3 R5 -AACACTTCGTGGGGTCCTTTTC-3 Stable and Transient Overexpression. STAT1β, PCAF, IL8, and CCNC full-length cdnas were isolated by RT-PCR from MDA- MB-435s cells and subcloned into the retroviral vector plpc (5). The plpc vector is a retroviral expression vector with a puromycin resistance marker (a gift from S. Lowe, Cold Spring Harbor Laboratory, Cold Spring Harbor, NY). LEDGF cdna was a generous gift of Marja Jäättelä Danish Cancer Society, Copenhagen, Denmark, and it was also subcloned into the retroviral vector plpc. The plasmids were verified by sequence analysis to confirm the absence of mutations. To generate stable cell pools for STAT1β, PCAF, IL8, CCNC, and the empty vector (plpc), MDA-MB-435s cells were infected using the supernatant from transiently transfected Phoenix-Ampho packaging cells. Transfection of the Phoenix-Ampho packaging cells was done using FuGENE 6 (Roche) for a period of 48 h. After this time the media were filtered, supplemented with polybrene (4 μg/ml), and added to MDA-MB-435s cells. Cells were subjected to two rounds of infection every 12 h. After 24 h from the last infection, cells were trypsinized and selected with puromycin (2.5 μg/ml) for 72 h. After selection, cells were analyzed for protein expression by Western blot. To overexpress LEDGF, MDA-MB-435s cells were transiently transfected with either empty vector (plpc) or LEDGF using Lipofectamine 2000 (Invitrogen) following the manufacturer s instructions. Cells were analyzed for protein expression by Western blot. Xenograft Mouse Model. Athymic female mice 8 10 wk old were s.c. inoculated with MDA-MB-435s cells. When tumors reached an average of 100 mm 3 in size, mice were randomly assigned to treatment groups and injections of Vivo-Morpholino were administered as described in SI Morpholino Treatments. Tumor-bearing mice were assessed for weight loss and tumor volume twice weekly. Tumor volumes were determined by caliper measurement using the following equation: length (width) 2 π/6. Four days after the last treatment, mice were killed and tumors were fixed in formalin. All animal studies were conducted in accordance with guidelines approved by the Memorial Sloan- Kettering Cancer Center Institutional Animal Care and Use Committee and the Research Animal Resource Center. 3of10
4 Statistical Analysis. The statistical significance of differences for both the morpholino and the sirna in vitro treatments was calculated using the stratified χ 2 test (6), and bars represent SEs. For both the ELISA and MTT assays a paired t test was used to establish statistically significant differences and bars represent SD. Differences were considered statistically significant at a P value of < Sambrook J, Russell DW (2001) Molecular Cloning: A Laboratory Manual (Cold Spring Harbor Lab, Cold Spring Harbor, NY), 3rd Ed, pp 13.26e Morcos PA, Li Y, Jiang S (2008) Vivo-Morpholinos: A non-peptide transporter delivers Morpholinos into a wide array of mouse tissues. Biotechniques 45:613e614, 616, 618 passim. 3. Pedranzini L, et al. (2006) Pyridone 6, a pan-janus-activated kinase inhibitor, induces growth inhibition of multiple myeloma cells. Cancer Res 66:9714e Dussault AA, Pouliot M (2006) Rapid and simple comparison of messenger RNA levels using real-time PCR. Biol Proced Online 8:1e de Stanchina E, et al. (1998) E1A signaling to p53 involves the p19(arf) tumor suppressor. Genes Dev 12:2434e Mantel N, Haenszel W (1959) Statistical aspects of the analysis of data from retrospective studies of disease. J Natl Cancer Inst 22:719e748. Fig. S1. Additional analysis of STAT3 exon 23 cis-elements. (A) RT-PCR analysis to assess the splicing pattern of endogenous STAT3 and transfected minigenes in HeLa cells using specific primers. Endo, endogenous; exo, exogenous. (B) Bioinformatic analysis of STAT3 exon 23 sequence using ESEfinder (1) ( cshl.edu/cgi-bin/tools/ese3/esefinder.cgi?process=home) and PESX (2) ( algorithms. The upper graph displays the predicted scores for functional binding of four common serine-arginine (SR) proteins (SRSF1, SRSF2, SRSF5, SRSF6), according to the ESEfinder algorithm, whereas in the PESX analysis, green sequences represent putative exonic splicing enhancers. Black bars at the bottom summarize the position of all deletion mutants (from this work) that had some effect on exon 23 splicing, thus identifying putative splicing regulatory elements in exon 23. Sequences boxed in yellow correspond to the region of exon 23 specifictotheαisoform, whereas sequences boxed in orange correspond to the region of exon 23 common to both α and β variants. Gray and black in the exon 23 sequence indicate codons in the α reading frame. (C) RT-PCR analysis of STAT3 α/β levels in HeLa cells transfected with the 12-nt (ΔH ΔO) exon 23 deletion mutants in STAT3 minigenes. wt, wild-type. (D) (Upper) Partial sequence of the wt and pyrimidine-enhanced (Py^) minigenes. Asterisks indicate the exact position of the mutations. (Lower) RT-PCR analysis of STAT3 α/β levels in both transfected minigenes in HeLa cells. (E) (Upper) Diagram of STAT3 minigene. Exon and intron sizes are indicated (nt). (Lower) Lines above and below the exon 23 sequence indicate the exact position of the 12-nt (ΔH ΔO) and 3-nt (ΔH1 ΔK3) deletions generated. (F and G) RT-PCR analysis of STAT3 α/β levels in HeLa cells transfected with the 12-nt (ΔH ΔO) and 3-nt (ΔH1 ΔK3) exon 23 deletion mutants, respectively, in the pyrimidine-enhanced minigenes. Py^, pyrimidine-enhanced. 4of10
5 1. Cartegni L, Wang J, Zhu Z, Zhang MQ, Krainer AR (2003) ESEfinder: A web resource to identify exonic splicing enhancers. Nucleic Acids Res 31:3568e Zhang XH, Chasin LA (2004) Computational definition of sequence motifs governing constitutive exon splicing. Genes Dev 18:1241e1250. Fig. S2. Specificity of morpholino treatment. RT-PCR analysis of STAT3α/β levels in human chronic myelogenous leukemia cell line K562, treated with increasing amount of ST3 and control INV compounds for 4 d at the indicated concentrations. NT, untreated. Fig. S3. Morpholino treatment of multiple cell lines. RT-PCR analysis of STAT3α/β levels in human lung cancer cell line A549, human prostate cancer cell line DU145, human chronic myelogenous leukemia cell line K562, mouse breast cancer cell line 4T1, and mouse melanoma cell line B16 treated with 16 μm ST2 for 4 d. Fig. S4. Sustainable splicing switch by ST2 treatment. RT-PCR analysis of STAT3 α/β levels in MDA-MB-435s cells treated with either INV or ST2 morpholino oligonucleotide for three consecutive treatments of 4 d each. Fig. S5. Side-by-side comparison of forced splicing-dependent nonsense-mediated decay and sirna effectiveness to knock down STAT3. Western blot (W.B.) of phospho-stat3 Y705 levels in lysates from treated MDA-MB-435s cells. Lanes 1 4 are a serial dilution of untreated cells (NT) from 100 to 12.5% of input. Lanes 5 7 are 100% of input from cells treated with 16 μm INV/ST2/ST6 oligos. Lane 8 is 100% of input from cells treated for 48 h with 100 nm sirna oligo against STAT3. Actin was used as loading control. Fig. S6. STAT3 Y705 phosphorylation status in different cell lines. Western blot analysis of phospho-stat3 Y705 and total levels in lysates from MDA-MB-453, MDA-MB-435s, and MDA-MB-468 cells. α-tubulin was used as loading control. 5of10
6 Fig. S7. ST2 but not ST6 treatment inhibits cell-cycle progression in MDA-MB-435s and MDA-MB-468 cells. The effect of ST2 or ST6 vivo-morpholino oligonucleotides on the cell cycle was evaluated by flow cytometric analysis. After 60 h of treatment, MDA-MB-435s cells treated with ST2 displayed an increase in the G2/M population (20.8% ± 0.28 SE) compared with the INV- (14% ± 1.4 SE) and ST6-treated cells (12.5% ± 3.5 SE). Similarly, in MDA-MB-468 cells, the G2/M population increased to 25% ± 1.4 SE following treatment with ST2 (INV: 9% ± 4.24 SE; ST6: 12% ± 0.7 SE). The observed increase of the cell-cycle population at the G2/M phase was accompanied by a decrease of cell population in the G1 phase of the cell cycle. All treatments were done at 4 μm, and the treatment was only for 60 h to ensure the cells were in exponential growth. (A) Representative profiles of the analysis of cell-cycle progression. MDA-MB-435s and MDA-MB-468 cells were treated with 4 μm INV, ST2, or ST6 for 60 h. The distribution and percentage of cells in the G1, S, and G2/M phase of the cell cycle are indicated. (B) Overlay of the cell-cycle profiles with ST2 treatment, associated with the increase in the G2/M population, showed in blue. ST6 is in red and control (INV) is in green. Fig. S8. ST2 but not ST6 treatment induces apoptosis in MDA-MB-435s and MDA-MB-468 cells. The effect of ST2 and ST6 treatment on apoptosis was evaluated with annexin V staining. After 60 h of treatment at 4 μm, an increase in the levels of apoptosis was observed in the MDA-MB-435s (9.33% apoptotic; ±1.76 SE) and MDA-MB-468 cells (20.6% apoptotic; ±0.5 SE), following ST2 treatment. (A) Representative scatter plot of flow analysis of MDA-MB-435s and MDA-MB-468 cells following treatment with INV, ST2, or ST6 at 4 μm for 60 h. Cells ( ) (including supernatant) were harvested and analyzed by AnxV/PI as detailed in SI Materials and Methods and data were analyzed with FlowJo software. The percentage of apoptotic cells is indicated. (B) Assessment of apoptotic population, as determined by AnxV/PI staining, in MDA-MB-435s and MDA-MB-468 cells treated with INV, ST2, or ST6 vivo-morpholinos (error bars indicate SE). 6of10
7 Fig. S9. Effectiveness of Vivo-Morpholino oligonucleotide treatment. (A) Side-by-side comparison of naked morpholinos and Vivo-Morpholino efficacy. RT- PCR analysis of STAT3α/β levels in MDA-MB-435s cells treated with either INV or ST2 morpholino plus Endo-Porter or with either INV or ST2 Vivo-Morpholino. All oligos were used at a final concentration of 16 μm for 4 d. EP, Endo-Porter. (B) Delivery of Vivo-Morpholino in a xenograft tumor model. RT-PCR analysis of STAT3α/β levels in intratumorally (IT) or intravenously (IV) treated xenograft tumors. MDA-MB-435s cells ( ) were s.c. inoculated in the flanks of female athymic mice. When tumors reached 100 mm 3 in volume size, mice (n = 3 per group) were randomly assigned to one of four treatment groups (Control-IV, ST2-IV, Control-IT, and ST2-IT). Animals were treated twice in 1 wk and, 4 d after the second treatment, tumors were collected and further analyzed. C, control. When similar experiments were attempted with MDA-MB-468 cells under comparable conditions (multiple times), the growth of the resulting tumors was unpredictable, with a high fraction of injections that did not take, whereas the ones that grew did so at very irregular rates, and in some cases the tumor grew but was highly necrotic (regardless of treatment). 7of10
8 Fig. S10. Histopathological analysis of Vivo-Morpholino-treated tumors. (A) Table summarizing the histopathological examination of PBS-, INV-, ST2-, and ST6-treated tumors such as their size (cm), estimated area of necrosis (%), and highest mitotic count per 20 high-power fields (hpf). (B) Representative results of the TUNEL staining of tumor sections from each of the four treated groups (40 magnification). (C) Representative results of the caspase 3 immunohistochemical staining of tumor sections from each of the four treated groups (40 magnification). Fig. S11. Validation of treatment efficacy of the samples used for qpcr/microarray analysis. (A) (Upper) RT-PCR analysis of STAT3 α/β levels in MDA-MB-435s cells treated with 16 μm INV, ST2, or ST6 oligos for 4 d. (Lower) RT-PCR analysis of STAT3 exon 6 skipping in MDA-MB-435s cells treated with 16 μm INV, ST2, or ST6 oligos for 4 d. (B) Western blot analysis of phospho-stat3 Y705 α/β levels in MDA-MB-435s cells treated with 16 μm INV, ST6, or ST2 for 4 d. α-tubulin was used as loading control. 8of10
9 Fig. S12. Microarray data from treated MDA-MB-435s cells. (A) Table representing significantly differentially expressed genes (fold change 2) filtered from the microarray dataset obtained by comparing ST2-treated versus INV-treated MDA-MB-435s cells. In red are up-regulated genes; in blue are down-regulated genes. (B) Table representing significantly differentially expressed genes (fold change 2) filtered from the microarray dataset obtained by comparing ST6- treated versus INV-treated MDA-MB-435s cells. In red are up-regulated genes; in blue are down-regulated genes. Fig. S13. Validation of STAT3β s transcriptional signature in the MDA-MB-468 cell line. cdnas from MDA-MB-468 cells treated with 16 μm ST2, ST6, or INV morpholinos for 4 d were analyzed by real-time PCR to validate STAT3β s transcriptional signature previously identified in MDA-MB-435s cells. Results are represented as a comparison between ST2 and INV treatments (blue) or between ST6 and INV treatments (red) and are expressed as ddc(t) values after normalization to HPRT expression levels. Data represent the average of at least three independent experiments. Bars represent SD. 9of10
10 Fig. S14. Comparison of (P6) pan-jak inhibitor with ST6 down-regulation of STAT3 in the MDA-MB-468 cell line. (A) MDA-MB-468 cells were treated for 48 h with 1 μm pan-jak inhibitor tetracyclic pyridone 6 (P6) or DMSO in combination with morpholino control treatment (INV), total STAT3 knockdown morpholino treatment (ST6), or STAT3 α/β splicing switch treatment (ST2) as indicated. All vivo-morpholinos were used at 6 μm. P6 was used in combination with INV to control for the baseline toxicity associated with morpholino treatment in MDA-MB-468 cells. However, the relative increase in cell death due to P6 versus DMSO is the same whether INV is present or not. Activation status of STAT3 and the effectiveness of treatment were monitored using an antibody against Y705 (Top), and total STAT3 levels were also determined by Western blot (Middle). Tubulin was used as loading control. (B) The effect of the various treatments on cell growth was measured using the MTT assay. Data represent the average of at least five independent experiments. Bars represent SD. Indicated P values were calculated by two-tailed Student s t test. The results indicate that lack of phospho-stat3 is not sufficient, per se, to inhibit growth even in cells described as highly sensitive to its depletion such as MDA-MB-468. Rather, another event must occur, such as the induction of STAT3β or the presumable inactivation of additional JAK substrates by pan-jak inhibitor P6. 10 of 10
Supporting Information
 Supporting Information Chakrabarty et al. 10.1073/pnas.1018001108 SI Materials and Methods Cell Lines. All cell lines were purchased from the American Type Culture Collection. Media and FBS were purchased
Supporting Information Chakrabarty et al. 10.1073/pnas.1018001108 SI Materials and Methods Cell Lines. All cell lines were purchased from the American Type Culture Collection. Media and FBS were purchased
Supplementary Information Alternative splicing of CD44 mrna by ESRP1 enhances lung colonization of metastatic cancer cell
 Supplementary Information Alternative splicing of CD44 mrna by ESRP1 enhances lung colonization of metastatic cancer cell Supplementary Figures S1-S3 Supplementary Methods Supplementary Figure S1. Identification
Supplementary Information Alternative splicing of CD44 mrna by ESRP1 enhances lung colonization of metastatic cancer cell Supplementary Figures S1-S3 Supplementary Methods Supplementary Figure S1. Identification
Supporting Information
 Supporting Information Casson et al. 10.1073/pnas.1421699112 Sl Materials and Methods Macrophage Infections. In experiments where macrophages were primed with LPS, cells were pretreated with 0.5 μg/ml
Supporting Information Casson et al. 10.1073/pnas.1421699112 Sl Materials and Methods Macrophage Infections. In experiments where macrophages were primed with LPS, cells were pretreated with 0.5 μg/ml
SUPPLEMENTARY INFORMATION
 (Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
(Supplementary Methods and Materials) GST pull-down assay GST-fusion proteins Fe65 365-533, and Fe65 538-700 were expressed in BL21 bacterial cells and purified with glutathione-agarose beads (Sigma).
Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan).
 1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and
 Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
Supplementary Figure Legend: Supplementary Fig. 1 Proteomic analysis of ATR-interacting proteins. ATR, ARID1A and ATRIP protein peptides identified from our mass spectrum analysis were shown. Supplementary
Supplemental Methods Cell lines and culture
 Supplemental Methods Cell lines and culture AGS, CL5, BT549, and SKBR were propagated in RPMI 64 medium (Mediatech Inc., Manassas, VA) supplemented with % fetal bovine serum (FBS, Atlanta Biologicals,
Supplemental Methods Cell lines and culture AGS, CL5, BT549, and SKBR were propagated in RPMI 64 medium (Mediatech Inc., Manassas, VA) supplemented with % fetal bovine serum (FBS, Atlanta Biologicals,
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
www.sciencemag.org/cgi/content/full/1137999/dc1 Supporting Online Material for Disrupting the Pairing Between let-7 and Enhances Oncogenic Transformation Christine Mayr, Michael T. Hemann, David P. Bartel*
SideStep Lysis and Stabilization Buffer
 SideStep Lysis and Stabilization Buffer INSTRUCTION MANUAL Catalog #400900 Revision B.0 For Research Use Only. Not for use in diagnostic procedures. 400900-12 LIMITED PRODUCT WARRANTY This warranty limits
SideStep Lysis and Stabilization Buffer INSTRUCTION MANUAL Catalog #400900 Revision B.0 For Research Use Only. Not for use in diagnostic procedures. 400900-12 LIMITED PRODUCT WARRANTY This warranty limits
Supplementary methods
 Supplementary methods Cell culture, infection, transfection, and RNA interference HEK293 cells and its derivatives were grown in DMEM supplemented with 10% FBS. Various constructs were introduced into
Supplementary methods Cell culture, infection, transfection, and RNA interference HEK293 cells and its derivatives were grown in DMEM supplemented with 10% FBS. Various constructs were introduced into
used at a final concentration of 5 ng/ml. Rabbit anti-bim and mouse anti-mkp2 antibodies were
 1 Supplemental Methods Reagents and chemicals: TGFβ was a generous gift from Genzyme Inc. (Cambridge, MA) and was used at a final concentration of 5 ng/ml. Rabbit anti-bim and mouse anti-mkp2 antibodies
1 Supplemental Methods Reagents and chemicals: TGFβ was a generous gift from Genzyme Inc. (Cambridge, MA) and was used at a final concentration of 5 ng/ml. Rabbit anti-bim and mouse anti-mkp2 antibodies
Supplementary Information: Materials and Methods. Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered
 Supplementary Information: Materials and Methods Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered saline (PBS) and lysed in TNN lysis buffer (50mM Tris at ph 8.0, 120mM NaCl
Supplementary Information: Materials and Methods Immunoblot and immunoprecipitation. Cells were washed in phosphate buffered saline (PBS) and lysed in TNN lysis buffer (50mM Tris at ph 8.0, 120mM NaCl
Reagents and cell culture Mcl-1 gene expression: real-time quantitative RT-PCR In vitro PP2A phosphatase assay Detection of Mcl-1 in vivo
 Reagents and cell culture Antibodies specific for caspase 3, PARP and GAPDH were purchased from Cell Signaling Technology Inc. (Beverly, MA). Caspase inhibitor z-vad-fmk and ROS scavenger N-acetyl-Lcysteine
Reagents and cell culture Antibodies specific for caspase 3, PARP and GAPDH were purchased from Cell Signaling Technology Inc. (Beverly, MA). Caspase inhibitor z-vad-fmk and ROS scavenger N-acetyl-Lcysteine
Mutagenesis and generation of expression vectors Gene expression profiling Lentiviral production and infection of TF1 cells
 Mutagenesis and generation of expression vectors Human c-kit wild-type cdna encoding the short c-kit isoform was excised from vector pbshkitwt (kind gift from Dr. L Gros, France) by Sal I-Acc65 I digestion.
Mutagenesis and generation of expression vectors Human c-kit wild-type cdna encoding the short c-kit isoform was excised from vector pbshkitwt (kind gift from Dr. L Gros, France) by Sal I-Acc65 I digestion.
Fig. S1 TGF RI inhibitor SB effectively blocks phosphorylation of Smad2 induced by TGF. FET cells were treated with TGF in the presence of
 Fig. S1 TGF RI inhibitor SB525334 effectively blocks phosphorylation of Smad2 induced by TGF. FET cells were treated with TGF in the presence of different concentrations of SB525334. Cells were lysed and
Fig. S1 TGF RI inhibitor SB525334 effectively blocks phosphorylation of Smad2 induced by TGF. FET cells were treated with TGF in the presence of different concentrations of SB525334. Cells were lysed and
supplementary information
 DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
To determine MRK-003 IC50 values, cell lines were plated in triplicate in 96-well plates at 3 x
 Supplementary methods: Cellular growth assay and cell cycle analysis To determine MRK-003 IC50 values, cell lines were plated in triplicate in 96-well plates at 3 x 10 3 cells/well (except for TALL1 and
Supplementary methods: Cellular growth assay and cell cycle analysis To determine MRK-003 IC50 values, cell lines were plated in triplicate in 96-well plates at 3 x 10 3 cells/well (except for TALL1 and
SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes
 WHITE PAPER SuperScript IV Reverse Transcriptase SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes Abstract Reverse transcriptases (RTs) from avian myeloblastosis virus
WHITE PAPER SuperScript IV Reverse Transcriptase SuperScript IV Reverse Transcriptase as a better alternative to AMV-based enzymes Abstract Reverse transcriptases (RTs) from avian myeloblastosis virus
RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the
 Supplementary Methods RT-PCR and real-time PCR analysis RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the manufacturer s protocol and quantified by measuring the
Supplementary Methods RT-PCR and real-time PCR analysis RNA was isolated using NucleoSpin RNA II (Macherey-Nagel, Bethlehem, PA) according to the manufacturer s protocol and quantified by measuring the
MeCP2. MeCP2/α-tubulin. GFP mir1-1 mir132
 Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
Rapid Method for the Purification of Total RNA from Formalin- Fixed Paraffin-Embedded (FFPE) Tissue Samples
 Application Note 17 RNA Sample Preparation Rapid Method for the Purification of Total RNA from Formalin- Fixed Paraffin-Embedded (FFPE) Tissue Samples M. Melmogy 1, V. Misic 1, B. Lam, PhD 1, C. Dobbin,
Application Note 17 RNA Sample Preparation Rapid Method for the Purification of Total RNA from Formalin- Fixed Paraffin-Embedded (FFPE) Tissue Samples M. Melmogy 1, V. Misic 1, B. Lam, PhD 1, C. Dobbin,
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
SANTA CRUZ BIOTECHNOLOGY, INC.
 TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
TECHNICAL SERVICE GUIDE: Western Blotting 2. What size bands were expected and what size bands were detected? 3. Was the blot blank or was a dark background or non-specific bands seen? 4. Did this same
Efficient Method for Isolation of High Quality Concentrated Cellular RNA with Extremely Low Levels of Genomic DNA Contamination Application
 Efficient Method for Isolation of High Quality Concentrated Cellular RNA with Extremely Low Levels of Genomic DNA Contamination Application Gene Expression Authors Ilgar Abbaszade, Claudia Robbins, John
Efficient Method for Isolation of High Quality Concentrated Cellular RNA with Extremely Low Levels of Genomic DNA Contamination Application Gene Expression Authors Ilgar Abbaszade, Claudia Robbins, John
Gene Expression Technology
 Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
Gene Expression Technology Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Gene expression Gene expression is the process by which information from a gene
B. ADM: C. D. Apoptosis: 1.68% 2.99% 1.31% Figure.S1,Li et al. number. invaded cells. HuH7 BxPC-3 DLD-1.
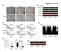 A. - Figure.S1,Li et al. B. : - + - + - + E-cadherin CK19 α-sma vimentin β -actin C. D. Apoptosis: 1.68% 2.99% 1.31% - : - + - + - + Apoptosis: 48.33% 45.32% 44.59% E. invaded cells number 400 300 200
A. - Figure.S1,Li et al. B. : - + - + - + E-cadherin CK19 α-sma vimentin β -actin C. D. Apoptosis: 1.68% 2.99% 1.31% - : - + - + - + Apoptosis: 48.33% 45.32% 44.59% E. invaded cells number 400 300 200
HeLa cells stably transfected with an empty pcdna3 vector (HeLa Neo) or with a plasmid
 SUPPLEMENTAL MATERIALS AND METHODS Cell culture, transfection and treatments. HeLa cells stably transfected with an empty pcdna3 vector (HeLa Neo) or with a plasmid encoding vmia (HeLa vmia) 1 were cultured
SUPPLEMENTAL MATERIALS AND METHODS Cell culture, transfection and treatments. HeLa cells stably transfected with an empty pcdna3 vector (HeLa Neo) or with a plasmid encoding vmia (HeLa vmia) 1 were cultured
Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit
 Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Application Note 13 RNA Sample Preparation Non-Organic-Based Isolation of Mammalian microrna using Norgen s microrna Purification Kit B. Lam, PhD 1, P. Roberts, MSc 1 Y. Haj-Ahmad, M.Sc., Ph.D 1,2 1 Norgen
Isolation of total nucleic acids from FFPE tissues using FormaPure DNA
 APPLICATION NOTE Isolation of total nucleic acids from FFPE tissues using FormaPure DNA Jung Hoon Doh, Ph.D. Senior Application Scientist Beckman Coulter Life Sciences, Indianapolis, IN USA Summary Extensive
APPLICATION NOTE Isolation of total nucleic acids from FFPE tissues using FormaPure DNA Jung Hoon Doh, Ph.D. Senior Application Scientist Beckman Coulter Life Sciences, Indianapolis, IN USA Summary Extensive
pdsipher and pdsipher -GFP shrna Vector User s Guide
 pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by. 5 AGACACAAACACCAUUGUCACACUCCACAGC; Rand-2 OMe,
 Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Materials and methods Oligonucleotides and DNA constructs RNA oligonucleotides and 2 -O-methylated oligonucleotides were synthesized by Dharmacon Inc. (Lafayette, CO). The sequences were: 122-2 OMe, 5
Supplementary Information (Ha, et. al) Supplementary Figures Supplementary Fig. S1
 Supplementary Information (Ha, et. al) Supplementary Figures Supplementary Fig. S1 a His-ORMDL3 ~ 17 His-ORMDL3 GST-ORMDL3 - + - + IPTG GST-ORMDL3 ~ b Integrated Density (ORMDL3/ -actin) 0.4 0.3 0.2 0.1
Supplementary Information (Ha, et. al) Supplementary Figures Supplementary Fig. S1 a His-ORMDL3 ~ 17 His-ORMDL3 GST-ORMDL3 - + - + IPTG GST-ORMDL3 ~ b Integrated Density (ORMDL3/ -actin) 0.4 0.3 0.2 0.1
Supporting Information
 Supporting Information Su et al. 10.1073/pnas.1211604110 SI Materials and Methods Cell Culture and Plasmids. Tera-1 and Tera-2 cells (ATCC: HTB- 105/106) were maintained in McCoy s 5A medium with 15% FBS
Supporting Information Su et al. 10.1073/pnas.1211604110 SI Materials and Methods Cell Culture and Plasmids. Tera-1 and Tera-2 cells (ATCC: HTB- 105/106) were maintained in McCoy s 5A medium with 15% FBS
Supplementary Figure 1: Overexpression of EBV-encoded proteins Western blot analysis of the expression levels of EBV-encoded latency III proteins in
 Supplementary Figure 1: Overexpression of EBV-encoded proteins Western blot analysis of the expression levels of EBV-encoded latency III proteins in BL2 cells. The Ponceau S staining of the membranes or
Supplementary Figure 1: Overexpression of EBV-encoded proteins Western blot analysis of the expression levels of EBV-encoded latency III proteins in BL2 cells. The Ponceau S staining of the membranes or
Supplementary Fig.1. Over-expression of RNase L in stable polyclonal cell line
 Supplemental Data mrna Protein A kda 75 40 NEO/vector NEO/RNase L RNASE L β-actin RNASE L β-actin B % of control (neo vector), normalized 240 ** 200 160 120 80 40 0 Neo Neo/RNase L RNase L Protein Supplementary
Supplemental Data mrna Protein A kda 75 40 NEO/vector NEO/RNase L RNASE L β-actin RNASE L β-actin B % of control (neo vector), normalized 240 ** 200 160 120 80 40 0 Neo Neo/RNase L RNase L Protein Supplementary
CITE-seq & Cell Hashing Protocol
 CITE-seq & Cell Hashing Protocol For experiments involving Cell Hashing, use cost per cell calculator to plan experiments, determine number of hashes, number of cells to load, expected doublet rates (detected
CITE-seq & Cell Hashing Protocol For experiments involving Cell Hashing, use cost per cell calculator to plan experiments, determine number of hashes, number of cells to load, expected doublet rates (detected
Cell Hashing Protocol
 Cell Hashing Protocol For experiments involving Cell Hashing, use cost per cell calculator to plan experiments, determine number of hashes, number of cells to load, expected doublet rates (detected and
Cell Hashing Protocol For experiments involving Cell Hashing, use cost per cell calculator to plan experiments, determine number of hashes, number of cells to load, expected doublet rates (detected and
Te c htips. Simple Approaches for Optimization of RT-PCR TECH TIP 206
 Te c htips TECH TIP 206 Fueling Innovation USB Corporation 26111 Miles Road Cleveland, Ohio 44128 800.321.9322 www.usbweb.com Simple Approaches for Optimization of RT-PCR Reverse transcription-polymerase
Te c htips TECH TIP 206 Fueling Innovation USB Corporation 26111 Miles Road Cleveland, Ohio 44128 800.321.9322 www.usbweb.com Simple Approaches for Optimization of RT-PCR Reverse transcription-polymerase
Application Note 18 RNA/DNA/Protein Sample Preparation METHODS AND MATERIALS INTRODUCTION
 Application Note 18 /DNA/Protein Sample Preparation Sequential Purification of, DNA and Protein from a Single Sample using 's /DNA/Protein Purification Kit and Comparison to a Market B. Lam, PhD 1, C.
Application Note 18 /DNA/Protein Sample Preparation Sequential Purification of, DNA and Protein from a Single Sample using 's /DNA/Protein Purification Kit and Comparison to a Market B. Lam, PhD 1, C.
Cell death analysis using the high content bioimager BD PathwayTM 855 instrument (BD
 Supplemental information Materials and Methods: Cell lines, reagents and antibodies: Wild type (A3) and caspase-8 -/- (I9.2) Jurkat cells were cultured in RPMI 164 medium (Life Technologies) supplemented
Supplemental information Materials and Methods: Cell lines, reagents and antibodies: Wild type (A3) and caspase-8 -/- (I9.2) Jurkat cells were cultured in RPMI 164 medium (Life Technologies) supplemented
Supplemental Material Igreja and Izaurralde 1. CUP promotes deadenylation and inhibits decapping of mrna targets. Catia Igreja and Elisa Izaurralde
 Supplemental Material Igreja and Izaurralde 1 CUP promotes deadenylation and inhibits decapping of mrna targets Catia Igreja and Elisa Izaurralde Supplemental Materials and methods Functional assays and
Supplemental Material Igreja and Izaurralde 1 CUP promotes deadenylation and inhibits decapping of mrna targets Catia Igreja and Elisa Izaurralde Supplemental Materials and methods Functional assays and
Roche Molecular Biochemicals Technical Note No. LC 12/2000
 Roche Molecular Biochemicals Technical Note No. LC 12/2000 LightCycler Absolute Quantification with External Standards and an Internal Control 1. General Introduction Purpose of this Note Overview of Method
Roche Molecular Biochemicals Technical Note No. LC 12/2000 LightCycler Absolute Quantification with External Standards and an Internal Control 1. General Introduction Purpose of this Note Overview of Method
Transcriptional Regulation of Pro-apoptotic Protein Kinase C-delta: Implications for Oxidative Stress-induced Neuronal Cell Death
 SUPPLEMENTAL DATA Transcriptional Regulation of Pro-apoptotic Protein Kinase C-delta: Implications for Oxidative Stress-induced Neuronal Cell Death Huajun Jin 1, Arthi Kanthasamy 1, Vellareddy Anantharam
SUPPLEMENTAL DATA Transcriptional Regulation of Pro-apoptotic Protein Kinase C-delta: Implications for Oxidative Stress-induced Neuronal Cell Death Huajun Jin 1, Arthi Kanthasamy 1, Vellareddy Anantharam
monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal antibody was examined in
 Supplementary information Supplementary figures Supplementary Figure 1 Determination of the s pecificity of in-house anti-rhbdd1 mouse monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal
Supplementary information Supplementary figures Supplementary Figure 1 Determination of the s pecificity of in-house anti-rhbdd1 mouse monoclonal antibody. (a) The specificity of the anti-rhbdd1 monoclonal
Rapid Method for the Purification of Total RNA from Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Samples
 Application Note 17 RNA Sample Preparation Rapid Method for the Purification of Total RNA from Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Samples M. Melmogy 1, V. Misic 1, B. Lam, PhD 1, C. Dobbin,
Application Note 17 RNA Sample Preparation Rapid Method for the Purification of Total RNA from Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Samples M. Melmogy 1, V. Misic 1, B. Lam, PhD 1, C. Dobbin,
Supplementary Information
 Supplementary Information Supplementary Figures Figure S1. Study of mgtl translation in vitro. (A) Detection of 5 LR RNA using wild-type and anti-sd (91-95) substituted templates in a transcription-translation
Supplementary Information Supplementary Figures Figure S1. Study of mgtl translation in vitro. (A) Detection of 5 LR RNA using wild-type and anti-sd (91-95) substituted templates in a transcription-translation
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1 shrna mediated knockdown of ZRSR2 in K562 and 293T cells. (a) ZRSR2 transcript levels in stably transduced K562 cells were determined using qrt-pcr. GAPDH was
Supplementary Figure 1 Supplementary Fig. 1 shrna mediated knockdown of ZRSR2 in K562 and 293T cells. (a) ZRSR2 transcript levels in stably transduced K562 cells were determined using qrt-pcr. GAPDH was
Supporting Information
 Supporting Information Development of a 2,4-Dinitrotoluene-Responsive Synthetic Riboswitch in E. coli cells Molly E. Davidson, Svetlana V. Harbaugh, Yaroslav G. Chushak, Morley O. Stone, Nancy Kelley-
Supporting Information Development of a 2,4-Dinitrotoluene-Responsive Synthetic Riboswitch in E. coli cells Molly E. Davidson, Svetlana V. Harbaugh, Yaroslav G. Chushak, Morley O. Stone, Nancy Kelley-
ASPP1 Fw GGTTGGGAATCCACGTGTTG ASPP1 Rv GCCATATCTTGGAGCTCTGAGAG
 Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
QUANTITATIVE RT-PCR PROTOCOL (SYBR Green I) (Last Revised: April, 2007)
 QUANTITATIVE RT-PCR PROTOCOL (SYBR Green I) (Last Revised: April, 007) Please contact Center for Plant Genomics (CPG) facility manager Hailing Jin (hljin@iastate.edu) regarding questions or corrections.
QUANTITATIVE RT-PCR PROTOCOL (SYBR Green I) (Last Revised: April, 007) Please contact Center for Plant Genomics (CPG) facility manager Hailing Jin (hljin@iastate.edu) regarding questions or corrections.
IKK is a therapeutic target in KRAS-induced lung cancer with disrupted p53 activity
 IKK is a therapeutic target in KRAS-induced lung cancer with disrupted p5 activity H6 5 5 H58 A59 H6 H58 A59 anti-ikkα anti-ikkβ anti-panras anti-gapdh anti-ikkα anti-ikkβ anti-panras anti-gapdh anti-ikkα
IKK is a therapeutic target in KRAS-induced lung cancer with disrupted p5 activity H6 5 5 H58 A59 H6 H58 A59 anti-ikkα anti-ikkβ anti-panras anti-gapdh anti-ikkα anti-ikkβ anti-panras anti-gapdh anti-ikkα
Mammosphere formation assay. Mammosphere culture was performed as previously described (13,
 Supplemental Text Materials and Methods Mammosphere formation assay. Mammosphere culture was performed as previously described (13, 17). For co-culture with fibroblasts and treatment with CM or CCL2, fibroblasts
Supplemental Text Materials and Methods Mammosphere formation assay. Mammosphere culture was performed as previously described (13, 17). For co-culture with fibroblasts and treatment with CM or CCL2, fibroblasts
NUCLEIC ACID PURIFICATION KITS FAST SIMPLE QUALITY CONVENIENT REPRODUCIBLE
 NUCLEIC ACID PURIFICATION KITS FAST SIMPLE QUALITY CONVENIENT REPRODUCIBLE COMPANY PROFILE Since its founding in 1998,, Inc. has been at the forefront of nucleic acid purification by offering products
NUCLEIC ACID PURIFICATION KITS FAST SIMPLE QUALITY CONVENIENT REPRODUCIBLE COMPANY PROFILE Since its founding in 1998,, Inc. has been at the forefront of nucleic acid purification by offering products
SUPPLEMENTAL FIGURES AND TABLES
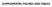 SUPPLEMENTAL FIGURES AND TABLES A B Flag-ALDH1A1 IP: α-ac HEK293T WT 91R 128R 252Q 367R 41/ 419R 435R 495R 412R C Flag-ALDH1A1 NAM IP: HEK293T + + - + D NAM - + + E Relative ALDH1A1 activity 1..8.6.4.2
SUPPLEMENTAL FIGURES AND TABLES A B Flag-ALDH1A1 IP: α-ac HEK293T WT 91R 128R 252Q 367R 41/ 419R 435R 495R 412R C Flag-ALDH1A1 NAM IP: HEK293T + + - + D NAM - + + E Relative ALDH1A1 activity 1..8.6.4.2
Emanuela Tumini, Sonia Barroso, Carmen Pérez Calero and Andrés Aguilera
 SUPPLEMENTARY INFORMATION Roles of human POLD1 and POLD3 in genome stability Emanuela Tumini, Sonia Barroso, Carmen Pérez Calero and Andrés Aguilera SUPPLEMENTARY METHODS Cell proliferation After sirna
SUPPLEMENTARY INFORMATION Roles of human POLD1 and POLD3 in genome stability Emanuela Tumini, Sonia Barroso, Carmen Pérez Calero and Andrés Aguilera SUPPLEMENTARY METHODS Cell proliferation After sirna
Score winning cdna yields with SuperScript III RT
 Score winning cdna yields with RT SuperScript III offers: Higher cdna yields Higher thermal stability Longer half-life than any other RT you could use Announcing SuperScript III Reverse Transcriptase How
Score winning cdna yields with RT SuperScript III offers: Higher cdna yields Higher thermal stability Longer half-life than any other RT you could use Announcing SuperScript III Reverse Transcriptase How
Supplementary Methods
 Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in
 Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
Supplementary Materials and Methods Barrier function assays At E17.5, the embryos were rinsed in phosphate-buffered saline (PBS) and immersed in acidic X-gal mix (100 mm phosphate buffer at ph4.3, 3 mm
SUPPLEMENTAL MATERIALS SIRTUIN 1 PROMOTES HYPEROXIA-INDUCED LUNG EPITHELIAL DEATH INDEPENDENT OF NRF2 ACTIVATION
 SUPPLEMENTAL MATERIALS SIRTUIN PROMOTES HYPEROXIA-INDUCED LUNG EPITHELIAL DEATH INDEPENDENT OF NRF ACTIVATION Haranatha R. Potteti*, Subbiah Rajasekaran*, Senthilkumar B. Rajamohan*, Chandramohan R. Tamatam,
SUPPLEMENTAL MATERIALS SIRTUIN PROMOTES HYPEROXIA-INDUCED LUNG EPITHELIAL DEATH INDEPENDENT OF NRF ACTIVATION Haranatha R. Potteti*, Subbiah Rajasekaran*, Senthilkumar B. Rajamohan*, Chandramohan R. Tamatam,
Roche Molecular Biochemicals Technical Note No. LC 10/2000
 Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Roche Molecular Biochemicals Technical Note No. LC 10/2000 LightCycler Overview of LightCycler Quantification Methods 1. General Introduction Introduction Content Definitions This Technical Note will introduce
Guide-it sgrna In Vitro Transcription and Screening Systems User Manual
 Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 632638, 632639, 632635, 632636, 632637 (040618) 1290 Terra Bella Avenue, Mountain View, CA 94043, USA U.S. Technical Support:
Guide-it sgrna In Vitro Transcription and Screening Systems User Manual Cat. Nos. 632638, 632639, 632635, 632636, 632637 (040618) 1290 Terra Bella Avenue, Mountain View, CA 94043, USA U.S. Technical Support:
SUPPORTING INFORMATION. A cleavage-responsive stem-loop hairpin for assaying guide RNA activity
 SUPPORTING INFORMATION A cleavage-responsive stem-loop hairpin for assaying guide RNA activity Tara R. deboer 1, Noreen Wauford 1, Jing-Yi Chung, Miguel Salvador Torres Perez, and Niren Murthy* University
SUPPORTING INFORMATION A cleavage-responsive stem-loop hairpin for assaying guide RNA activity Tara R. deboer 1, Noreen Wauford 1, Jing-Yi Chung, Miguel Salvador Torres Perez, and Niren Murthy* University
Blimp-1/PRDM1 rabbit monoclonal antibody (C14A4) was purchased from Cell Signaling
 1 2 3 4 5 6 7 8 9 10 11 Supplementary Methods Antibodies Blimp-1/PRDM1 rabbit monoclonal antibody (C14A4) was purchased from Cell Signaling (Danvers, MA) and used at 1:1000 to detect the total PRDM1 protein
1 2 3 4 5 6 7 8 9 10 11 Supplementary Methods Antibodies Blimp-1/PRDM1 rabbit monoclonal antibody (C14A4) was purchased from Cell Signaling (Danvers, MA) and used at 1:1000 to detect the total PRDM1 protein
Single cell imaging of Bruton's Tyrosine Kinase using an irreversible inhibitor
 SUPPLEMENTARY INFORMATION Single cell imaging of Bruton's Tyrosine Kinase using an irreversible inhibitor Anna Turetsky 1,a, Eunha Kim 1,a, Rainer H. Kohler 1, Miles A. Miller 1, Ralph Weissleder 1,2,
SUPPLEMENTARY INFORMATION Single cell imaging of Bruton's Tyrosine Kinase using an irreversible inhibitor Anna Turetsky 1,a, Eunha Kim 1,a, Rainer H. Kohler 1, Miles A. Miller 1, Ralph Weissleder 1,2,
Schematic representation of the endogenous PALB2 locus and gene-disruption constructs
 Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Supplementary Figure 1. Expressions of stem cell markers decreased in TRCs on 2D plastic. TRCs were cultured on plastic for 1, 3, 5, or 7 days,
 Supplementary Figure 1. Expressions of stem cell markers decreased in TRCs on 2D plastic. TRCs were cultured on plastic for 1, 3, 5, or 7 days, respectively, and their mrnas were quantified by real time
Supplementary Figure 1. Expressions of stem cell markers decreased in TRCs on 2D plastic. TRCs were cultured on plastic for 1, 3, 5, or 7 days, respectively, and their mrnas were quantified by real time
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Table 1. Primers, annealing temperatures, and product sizes for PCR amplification.
 Table 1. Primers, annealing temperatures, and product sizes for PCR amplification. Gene Direction Primer sequence (5 3 ) Annealing Temperature Size (bp) BRCA1 Forward TTGCGGGAGGAAAATGGGTAGTTA 50 o C 292
Table 1. Primers, annealing temperatures, and product sizes for PCR amplification. Gene Direction Primer sequence (5 3 ) Annealing Temperature Size (bp) BRCA1 Forward TTGCGGGAGGAAAATGGGTAGTTA 50 o C 292
PrimePCR Assay Validation Report
 Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID protein kinase N1 PKN1 Human The protein encoded by this gene belongs to the protein
Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID protein kinase N1 PKN1 Human The protein encoded by this gene belongs to the protein
SUPPLEMENTARY MATERIAL AND METHODS
 SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
SUPPLEMENTARY MATERIAL AND METHODS Amplification of HEV ORF1, ORF2 and ORF3 genome regions Total RNA was extracted from 200 µl EDTA plasma using Cobas AmpliPrep total nucleic acid isolation kit (Roche,
P HENIX. PHENIX PCR Enzyme Guide Tools For Life Science Discovery RESEARCH PRODUCTS
 PHENIX PCR Enzyme Guide PHENIX offers a broad line of premium quality PCR Enzymes. This PCR Enzyme Guide will help simplify your polymerase selection process. Each DNA Polymerase has different characteristics
PHENIX PCR Enzyme Guide PHENIX offers a broad line of premium quality PCR Enzymes. This PCR Enzyme Guide will help simplify your polymerase selection process. Each DNA Polymerase has different characteristics
Protocol CRISPR Genome Editing In Cell Lines
 Protocol CRISPR Genome Editing In Cell Lines Protocol 2: HDR donor plasmid applications (gene knockout, gene mutagenesis, gene tagging, Safe Harbor ORF knock-in) Notes: 1. sgrna validation: GeneCopoeia
Protocol CRISPR Genome Editing In Cell Lines Protocol 2: HDR donor plasmid applications (gene knockout, gene mutagenesis, gene tagging, Safe Harbor ORF knock-in) Notes: 1. sgrna validation: GeneCopoeia
Percent survival. Supplementary fig. S3 A.
 Supplementary fig. S3 A. B. 100 Percent survival 80 60 40 20 Ml 0 0 100 C. Fig. S3 Comparison of leukaemia incidence rate in the triple targeted chimaeric mice and germline-transmission translocator mice
Supplementary fig. S3 A. B. 100 Percent survival 80 60 40 20 Ml 0 0 100 C. Fig. S3 Comparison of leukaemia incidence rate in the triple targeted chimaeric mice and germline-transmission translocator mice
supplementary information
 DOI: 10.1038/ncb1862 Figure S1 Identification of SCAI as a Dia1 associating factor. (a) Identification of SCAI (Riken cdna 930041I02) as a Dia1-FH3 interacting protein. Mouse brain lysate was incubated
DOI: 10.1038/ncb1862 Figure S1 Identification of SCAI as a Dia1 associating factor. (a) Identification of SCAI (Riken cdna 930041I02) as a Dia1-FH3 interacting protein. Mouse brain lysate was incubated
LysoTracker Red DND-99 (Invitrogen) was used as a marker of lysosome or acidic
 information MATERIAL AND METHODS Lysosome staining LysoTracker Red DND-99 (Invitrogen) was used as a marker of lysosome or acidic compartments, according to the manufacturer s protocol. Plasmid independent
information MATERIAL AND METHODS Lysosome staining LysoTracker Red DND-99 (Invitrogen) was used as a marker of lysosome or acidic compartments, according to the manufacturer s protocol. Plasmid independent
Supplementary Fig. 1. (A) Working model. The pluripotency transcription factor OCT4
 SUPPLEMENTARY FIGURE LEGENDS Supplementary Fig. 1. (A) Working model. The pluripotency transcription factor OCT4 directly up-regulates the expression of NIPP1 and CCNF that together inhibit protein phosphatase
SUPPLEMENTARY FIGURE LEGENDS Supplementary Fig. 1. (A) Working model. The pluripotency transcription factor OCT4 directly up-regulates the expression of NIPP1 and CCNF that together inhibit protein phosphatase
PrimePCR Assay Validation Report
 Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID protein phosphatase, Mg2+/Mn2+ dependent, 1A PPM1A Human The protein encoded by
Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID protein phosphatase, Mg2+/Mn2+ dependent, 1A PPM1A Human The protein encoded by
The Wnt-5a-derived hexapeptide Foxy-5 inhibits breast cancer. metastasis in vivo by targeting cell motility
 SUPPLEMENTARY MATERIAL: The Wnt-5a-derived hexapeptide Foxy-5 inhibits breast cancer metastasis in vivo by targeting cell motility Annette Säfholm, Johanna Tuomela, Jeanette Rosenkvist, Janna Dejmek, Pirkko
SUPPLEMENTARY MATERIAL: The Wnt-5a-derived hexapeptide Foxy-5 inhibits breast cancer metastasis in vivo by targeting cell motility Annette Säfholm, Johanna Tuomela, Jeanette Rosenkvist, Janna Dejmek, Pirkko
Comparative Analysis of Argonaute-Dependent Small RNA Pathways in Drosophila
 Molecular Cell, Volume 32 Supplemental Data Comparative Analysis of Argonaute-Dependent Small RNA Pathways in Drosophila Rui Zhou, Ikuko Hotta, Ahmet M. Denli, Pengyu Hong, Norbert Perrimon, and Gregory
Molecular Cell, Volume 32 Supplemental Data Comparative Analysis of Argonaute-Dependent Small RNA Pathways in Drosophila Rui Zhou, Ikuko Hotta, Ahmet M. Denli, Pengyu Hong, Norbert Perrimon, and Gregory
Figure S Relative MUC4 transcript level* CD18/HPAF CD18/HPAF-Scr CD18/HPAF-siMUC4
 Figure S1 Relative MUC4 transcript level* 1.4 1.2 1 0.8 0.6 0.4 0.2 0 CD18/HPAF CD18/HPAF-Scr CD18/HPAF-siMUC4 Figure S2 * * CD18/HPAF-Scr CD18/HPAF-siMUC4 CD18/HPAF-Scr CD18/HPAF-siMUC4 Figure S3 CD18/HPAF-Scr
Figure S1 Relative MUC4 transcript level* 1.4 1.2 1 0.8 0.6 0.4 0.2 0 CD18/HPAF CD18/HPAF-Scr CD18/HPAF-siMUC4 Figure S2 * * CD18/HPAF-Scr CD18/HPAF-siMUC4 CD18/HPAF-Scr CD18/HPAF-siMUC4 Figure S3 CD18/HPAF-Scr
supplementary information
 DOI: 10.1038/ncb2116 Figure S1 CDK phosphorylation of EZH2 in cells. (a) Comparison of candidate CDK phosphorylation sites on EZH2 with known CDK substrates by multiple sequence alignments. (b) CDK1 and
DOI: 10.1038/ncb2116 Figure S1 CDK phosphorylation of EZH2 in cells. (a) Comparison of candidate CDK phosphorylation sites on EZH2 with known CDK substrates by multiple sequence alignments. (b) CDK1 and
Supplementary Information
 Supplementary Information Supplementary Figures Supplementary Figure 1. MLK1-4 phosphorylate MEK in the presence of RAF inhibitors. (a) H157 cells were transiently transfected with Flag- or HA-tagged MLK1-4
Supplementary Information Supplementary Figures Supplementary Figure 1. MLK1-4 phosphorylate MEK in the presence of RAF inhibitors. (a) H157 cells were transiently transfected with Flag- or HA-tagged MLK1-4
Impact of Nutraceuticals on TERT gene encoded protein
 Impact of Nutraceuticals on TERT gene encoded protein Xu Liu Department of Biological Sciences Fordham University, Bronx, New York, 10458 Abstract Telomerase is a Ribonucleo-protein polymerase that plays
Impact of Nutraceuticals on TERT gene encoded protein Xu Liu Department of Biological Sciences Fordham University, Bronx, New York, 10458 Abstract Telomerase is a Ribonucleo-protein polymerase that plays
Partial list of differentially expressed genes from cdna microarray, comparing MUC18-
 Supplemental Figure legends Table-1 Partial list of differentially expressed genes from cdna microarray, comparing MUC18- silenced and NT-transduced A375SM cells. Supplemental Figure 1 Effect of MUC-18
Supplemental Figure legends Table-1 Partial list of differentially expressed genes from cdna microarray, comparing MUC18- silenced and NT-transduced A375SM cells. Supplemental Figure 1 Effect of MUC-18
b alternative classical none
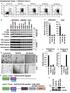 Supplementary Figure. 1: Related to Figure.1 a d e b alternative classical none NIK P-IkBa Total IkBa Tubulin P52 (Lighter) P52 (Darker) RelB (Lighter) RelB (Darker) HDAC1 Control-Sh RelB-Sh NF-kB2-Sh
Supplementary Figure. 1: Related to Figure.1 a d e b alternative classical none NIK P-IkBa Total IkBa Tubulin P52 (Lighter) P52 (Darker) RelB (Lighter) RelB (Darker) HDAC1 Control-Sh RelB-Sh NF-kB2-Sh
Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna
 Supplemental Materials and Methods Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna (Mission shrna, Sigma) against mouse MMP14 were transfected into HEK293 cells using FuGene6
Supplemental Materials and Methods Short hairpin RNA (shrna) against MMP14. Lentiviral plasmids containing shrna (Mission shrna, Sigma) against mouse MMP14 were transfected into HEK293 cells using FuGene6
Flow cytometric determination of apoptosis by annexin V/propidium iodide double staining.
 Supplementary materials and methods Flow cytometric determination of apoptosis by annexin V/propidium iodide double staining. Cells were analyzed for phosphatidylserine exposure by an annexin-v FITC/propidium
Supplementary materials and methods Flow cytometric determination of apoptosis by annexin V/propidium iodide double staining. Cells were analyzed for phosphatidylserine exposure by an annexin-v FITC/propidium
Recombinant DNA Technology
 History of recombinant DNA technology Recombinant DNA Technology (DNA cloning) Majid Mojarrad Recombinant DNA technology is one of the recent advances in biotechnology, which was developed by two scientists
History of recombinant DNA technology Recombinant DNA Technology (DNA cloning) Majid Mojarrad Recombinant DNA technology is one of the recent advances in biotechnology, which was developed by two scientists
Supplementary Information
 Journal : Nature Biotechnology Supplementary Information Targeted genome engineering in human cells with RNA-guided endonucleases Seung Woo Cho, Sojung Kim, Jong Min Kim, and Jin-Soo Kim* National Creative
Journal : Nature Biotechnology Supplementary Information Targeted genome engineering in human cells with RNA-guided endonucleases Seung Woo Cho, Sojung Kim, Jong Min Kim, and Jin-Soo Kim* National Creative
SUPPLEMENTARY INFORMATION
 doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
Supplementary Figure 1 - Characterization of rbag3 binding on macrophages cell surface.
 Supplementary Figure 1 - Characterization of rbag3 binding on macrophages cell surface. (a) Human PDAC cell lines were treated as indicated in Figure 1 panel F. Cells were analyzed for FITC-rBAG3 binding
Supplementary Figure 1 - Characterization of rbag3 binding on macrophages cell surface. (a) Human PDAC cell lines were treated as indicated in Figure 1 panel F. Cells were analyzed for FITC-rBAG3 binding
ProtoScript. First Strand cdna Synthesis Kit DNA AMPLIFICATION & PCR. Instruction Manual. NEB #E6300S/L 30/150 reactions Version 2.
 DNA AMPLIFICATION & PCR ProtoScript First Strand cdna Synthesis Kit Instruction Manual NEB #E6300S/L 30/150 reactions Version 2.2 11/16 be INSPIRED drive DISCOVERY stay GENUINE This product is intended
DNA AMPLIFICATION & PCR ProtoScript First Strand cdna Synthesis Kit Instruction Manual NEB #E6300S/L 30/150 reactions Version 2.2 11/16 be INSPIRED drive DISCOVERY stay GENUINE This product is intended
The Agilent Total RNA Isolation Kit
 Better RNA purity. Better data. Without DNase treatment. The Agilent Total RNA Isolation Kit Now you can isolate highly purified, intact RNA without DNase treatment Introducing the Agilent Total RNA Isolation
Better RNA purity. Better data. Without DNase treatment. The Agilent Total RNA Isolation Kit Now you can isolate highly purified, intact RNA without DNase treatment Introducing the Agilent Total RNA Isolation
QUICK-Clone TM User Manual. cdna
 QUICK-Clone TM User Manual cdna PT1150-1 (PR752268) Published 25 May 2007 Table of Contents I. Introduction 3 II. Applications Discussion 4 A. Primer Design 4 B. Setting up the PCR Reaction 4 C. Example
QUICK-Clone TM User Manual cdna PT1150-1 (PR752268) Published 25 May 2007 Table of Contents I. Introduction 3 II. Applications Discussion 4 A. Primer Design 4 B. Setting up the PCR Reaction 4 C. Example
OneTaq One-Step RT-PCR Kit
 POLYMERASES & AMPLIFICATION OneTaq One-Step RT-PCR Kit Instruction Manual NEB #E5315S 30 reactions Version 2.0 6/18 be INSPIRED drive DISCOVERY stay GENUINE This product is intended for research purposes
POLYMERASES & AMPLIFICATION OneTaq One-Step RT-PCR Kit Instruction Manual NEB #E5315S 30 reactions Version 2.0 6/18 be INSPIRED drive DISCOVERY stay GENUINE This product is intended for research purposes
Hossain_Supplemental Figure 1
 Hossain_Supplemental Figure 1 GFP-PACT GFP-PACT Motif I GFP-PACT Motif II A. MG132 (1µM) GFP Tubulin GFP-PACT Pericentrin GFP-PACT GFP-PACT Pericentrin Fig. S1. Expression and localization of Orc1 PACT
Hossain_Supplemental Figure 1 GFP-PACT GFP-PACT Motif I GFP-PACT Motif II A. MG132 (1µM) GFP Tubulin GFP-PACT Pericentrin GFP-PACT GFP-PACT Pericentrin Fig. S1. Expression and localization of Orc1 PACT
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/317/5837/477/dc1 Supporting Online Material for AAV Vector Integration Sites in Mouse Hepatocellular Carcinoma Anthony Donsante, Daniel G. Miller, Yi Li, Carole Vogler,
www.sciencemag.org/cgi/content/full/317/5837/477/dc1 Supporting Online Material for AAV Vector Integration Sites in Mouse Hepatocellular Carcinoma Anthony Donsante, Daniel G. Miller, Yi Li, Carole Vogler,
Document S1. Supplemental Experimental Procedures and Three Figures (see next page)
 Supplemental Data Document S1. Supplemental Experimental Procedures and Three Figures (see next page) Table S1. List of Candidate Genes Identified from the Screen. Candidate genes, corresponding dsrnas
Supplemental Data Document S1. Supplemental Experimental Procedures and Three Figures (see next page) Table S1. List of Candidate Genes Identified from the Screen. Candidate genes, corresponding dsrnas
Purification and Sequencing of DNA Guides from Prokaryotic Argonaute Daan C. Swarts *, Edze R. Westra, Stan J. J. Brouns and John van der Oost
 Purification and Sequencing of DNA Guides from Prokaryotic Argonaute Daan C. Swarts *, Edze R. Westra, Stan J. J. Brouns and John van der Oost Department of Agrotechnology and Food Sciences, Wageningen
Purification and Sequencing of DNA Guides from Prokaryotic Argonaute Daan C. Swarts *, Edze R. Westra, Stan J. J. Brouns and John van der Oost Department of Agrotechnology and Food Sciences, Wageningen
Purification of Concentrated, High-Quality RNA from Cells and Tissues
 Purification of Concentrated, High-Quality RNA from Cells and Tissues ReliaPrep RNA Cell and Tissue Miniprep Systems Don Smith, Promega Corporation Products Featured: ReliaPrep RNA Cell Miniprep System
Purification of Concentrated, High-Quality RNA from Cells and Tissues ReliaPrep RNA Cell and Tissue Miniprep Systems Don Smith, Promega Corporation Products Featured: ReliaPrep RNA Cell Miniprep System
