PPAR-γ - Peroxisome proliferator-ac4vated receptor gamma
|
|
|
- Mervyn Collins
- 6 years ago
- Views:
Transcription
1
2
3 PPAR-γ - Peroxisome proliferator-ac4vated receptor gamma
4 PPAR-γ - Peroxisome proliferator-ac4vated receptor gamma
5 C/EBPalpha - CCAAT/enhancer-binding protein alpha C/EBPα contains: - a func)onally related leucine zipper dimeriza)on domain (LZ) at its C-terminus and an adjacent highly conserved basic region (BR) that mediates sequence-specific DNA binding. - a transac)va)on domains (TADs) and a regulatory domain (RD) located in their N-terminal regions.
6 Cell differentiation
7 WNT pathway
8 Chroma4n structure and gene expression
9
10
11 Murine Adherent Fibroblast
12 in vitro osteogenesis model by exposing C3H10T1/2 cells to bone morphogene4c protein 2 (BMP2)
13 PPAR-γ mrna expression profile during C3H10T1/2 osteogenesis by real-4me PCR and western bloong
14 Adenovirus vector expressing sirna against PPAR-γ to down-regulate PPARc expression also used G3335, a specific antagonist of PPAR-γ, to block its ac4vity PPAR-γ contributed to C/EBPa expression during C3H10T1/2 osteogenesis at least in part
15 up-regula4on of PPAR-γ and C/EBPa expression These data suggest that PPAR-γ ac4vates C/EBPa expression in C3H10T1/2 cells
16 CLONING - TRANSFECTION - LUCIFERASE ASSAY Figura 21: Mappa circolare del veuore pgl3-basic. La regione del promotore -1785/-20 del 5 del gene SLC25A1 è stato inserito, mediante diges<one con le endonucleasi di restrizione BglII e HindIII, nella regione a monte del gene reporter per la luciferasi.
17 CLONING - TRANSFECTION - LUCIFERASE ASSAY
18 CLONING - TRANSFECTION - LUCIFERASE ASSAY
19 Binding of PPAR-γ to the 1286 bp/1065 bp promoter region to ac4vate C/EBPa expression a transient reporter assay was performed in C3H10T1/2 cells treated with BMP2, rosiglitazone and/or G3335 for 72 h
20 ChIP experiments
21 Binding of PPAR-γ to the 1286 bp/1065 bp promoter region to ac4vate C/EBPa expression ChIP experiments performed in C3H10T1/2 cells treated with BMP2, using PPARg an4body These data confirm binding of PPAR-γ to the 1286 bp/1065 bp region of the C/EBPa promoter.
22 Plasmid In vitro methyla4on: SAM treatment
23 DNA methyla4on and PPAR-γ -associated repression of HDAC1 in the 1286 bp/ 1065 bp region of the C/EBPa promoter In a previous report [2], DNA hypermethyla)on in the 1286 bp/1065 bp region of the C/ EBPa promoter was observed at the terminal stage (21 days) of osteogenesis of C3H10T1/2 cells The results showed that DNA methyla4on counters PPAR-γ binding to the 1286 bp/1065 bp region
24 DNA methyla4on and PPAR-γ -associated repression of HDAC1 in the 1286 bp/ 1065 bp region of the C/EBPa promoter In addi)on to DNA hypermethyla)on, hypoacetyla)on of histones 3 and 4 in the 1286 bp/1065 bp region of the C/EBPa promoter was also observed at the terminal stage of osteogenesis [2]. The acetyla4on level of histones 3 and 4 was reduced and HDAC1 binding was significantly enhanced in the 1286 bp/1065 bp region at the terminal stage of osteogenesis compared with the early stage
25 Ac4va4on of C/EBPa expression by PPAR-γ during adipogenesis of C3H10T1/2 cells C3H10T1/2 cells were induced to undergo adipogenesis by insulin, fetal bovine serum, methylisobutylxant hine and dexamethasone (IFMD protocol)
26 Luciferase reporter vector in which expression of luciferase was driven by the 1286 bp/1065 bp region of the C/EBPa promoter. A^er in vitro methyla)on, the vectors were transfected into C3H10T1/2 cells, and ChIP was performed in IFMD and control cultures
27 PPAR-γ and HDAC1 binding and the DNA methyla4on and histone acetyla4on status in the 1286 bp/1065 bp region of the C/EBPa promoter during in vitro osteogenesis of mouse BMSC (Bone marrow from BALB/c mice)
28 PPAR-γ and HDAC1 binding and the DNA methyla4on and histone acetyla4on status in the 1286 bp/1065 bp region of the C/EBPa promoter during in vitro osteogenesis of mouse BMSC (Bone marrow from BALB/c mice)
29 Balance between osteogenesis and adipogenesis of BMSC by modula4on of DNA methyla4on and histone acetyla4on The adipogenesis poten4al, which decreased at the terminal stage of osteogenesis, was recovered by 5ʹ-aza and TSA treatment
30 Model for C/EBPa expression regulated by PPAR-γ at the early and terminal stages of osteogenesis
31 The results of that study were interes4ng: DNA hypermethyla4on and hypoacetyla4on of histones 3 and 4 in the 1286 bp/1065 bp region of the C/EBPa promoter downregulated C/EBPa transcrip4on at the terminal stage of osteogenesis, and hypoacetyla4on of histones 3 and 4 depended on DNA hypermethyla4on through unknown molecular mechanisms. Ques4ons remain open regarding the balance between osteogenesis and adipogenesis regulated by C/EBPa that require elucida4on in more stringent, mechanis4c terms.
Decoding Chromatin States with Epigenome Data Advanced Topics in Computa8onal Genomics
 Decoding Chromatin States with Epigenome Data 02-715 Advanced Topics in Computa8onal Genomics HMMs for Decoding Chromatin States Epigene8c modifica8ons of the genome have been associated with Establishing
Decoding Chromatin States with Epigenome Data 02-715 Advanced Topics in Computa8onal Genomics HMMs for Decoding Chromatin States Epigene8c modifica8ons of the genome have been associated with Establishing
Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector.
 Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector. (a) Western blotting analysis and (b) qpcr analysis of eif6 expression in HEK293 T cells transfected with either
Fig. S1. eif6 expression in HEK293 transfected with shrna against eif6 or pcmv-eif6 vector. (a) Western blotting analysis and (b) qpcr analysis of eif6 expression in HEK293 T cells transfected with either
Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency.
 Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency. (a) Transfection of different concentration of GAS5-overexpressing
Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency. (a) Transfection of different concentration of GAS5-overexpressing
SUPPLEMENTARY INFORMATION
 Figure S1: Activation of the ATM pathway by I-PpoI. A. HEK293T cells were either untransfected, vector transfected, transfected with an I-PpoI expression vector, or subjected to 2Gy γ-irradiation. 24 hrs
Figure S1: Activation of the ATM pathway by I-PpoI. A. HEK293T cells were either untransfected, vector transfected, transfected with an I-PpoI expression vector, or subjected to 2Gy γ-irradiation. 24 hrs
The Transfection Collection TCF/LEF Transient Pack Wnt / -catenin Signaling Pathway Catalog #: 79273
 Data Sheet The Transfection Collection TCF/LEF Transient Pack Wnt / -catenin Signaling Pathway Catalog #: 79273 Background The Wnt / -catenin signaling pathway controls a large and diverse set of cell
Data Sheet The Transfection Collection TCF/LEF Transient Pack Wnt / -catenin Signaling Pathway Catalog #: 79273 Background The Wnt / -catenin signaling pathway controls a large and diverse set of cell
Table 1. Primers, annealing temperatures, and product sizes for PCR amplification.
 Table 1. Primers, annealing temperatures, and product sizes for PCR amplification. Gene Direction Primer sequence (5 3 ) Annealing Temperature Size (bp) BRCA1 Forward TTGCGGGAGGAAAATGGGTAGTTA 50 o C 292
Table 1. Primers, annealing temperatures, and product sizes for PCR amplification. Gene Direction Primer sequence (5 3 ) Annealing Temperature Size (bp) BRCA1 Forward TTGCGGGAGGAAAATGGGTAGTTA 50 o C 292
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
pdsipher and pdsipher -GFP shrna Vector User s Guide
 pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
pdsipher and pdsipher -GFP shrna Vector User s Guide NOTE: PLEASE READ THE ENTIRE PROTOCOL CAREFULLY BEFORE USE Page 1. Introduction... 1 2. Vector Overview... 1 3. Vector Maps 2 4. Materials Provided...
1. Primers for PCR to amplify hairpin stem-loop precursor mir-145 plus different flanking sequence from human genomic DNA.
 Supplemental data: 1. Primers for PCR to amplify hairpin stem-loop precursor mir-145 plus different flanking sequence from human genomic DNA. Strategy#1: 20nt at both sides: #1_BglII-Fd primer : 5 -gga
Supplemental data: 1. Primers for PCR to amplify hairpin stem-loop precursor mir-145 plus different flanking sequence from human genomic DNA. Strategy#1: 20nt at both sides: #1_BglII-Fd primer : 5 -gga
OmicsLink shrna Clones guaranteed knockdown even in difficult-to-transfect cells
 OmicsLink shrna Clones guaranteed knockdown even in difficult-to-transfect cells OmicsLink shrna clone collections consist of lentiviral, and other mammalian expression vector based small hairpin RNA (shrna)
OmicsLink shrna Clones guaranteed knockdown even in difficult-to-transfect cells OmicsLink shrna clone collections consist of lentiviral, and other mammalian expression vector based small hairpin RNA (shrna)
Estradiol-Estrogen Receptor α Mediates the Expression of the CXXC5 Gene through the Estrogen Response Element-Dependent Signaling Pathway
 Estradiol-Estrogen Receptor α Mediates the Expression of the CXXC5 Gene through the Estrogen Response Element-Dependent Signaling Pathway Pelin Yaşar, Gamze Ayaz and Mesut Muyan SUPPLEMENTARY INFORMATION
Estradiol-Estrogen Receptor α Mediates the Expression of the CXXC5 Gene through the Estrogen Response Element-Dependent Signaling Pathway Pelin Yaşar, Gamze Ayaz and Mesut Muyan SUPPLEMENTARY INFORMATION
Site-Directed Mutagenesis. Mutations in four Smad4 sites of mouse Gat1 promoter
 Supplement Supporting Materials and Methods Site-Directed Mutagenesis. Mutations in four Smad4 sites of mouse Gat1 promoter were independently generated using a two-step PCR method. The Smad4 binding site
Supplement Supporting Materials and Methods Site-Directed Mutagenesis. Mutations in four Smad4 sites of mouse Gat1 promoter were independently generated using a two-step PCR method. The Smad4 binding site
Table S1. Primers used in RT-PCR studies (all in 5 to 3 direction)
 Table S1. Primers used in RT-PCR studies (all in 5 to 3 direction) Epo Fw CTGTATCATGGACCACCTCGG Epo Rw TGAAGCACAGAAGCTCTTCGG Jak2 Fw ATCTGACCTTTCCATCTGGGG Jak2 Rw TGGTTGGGTGGATACCAGATC Stat5A Fw TTACTGAAGATCAAGCTGGGG
Table S1. Primers used in RT-PCR studies (all in 5 to 3 direction) Epo Fw CTGTATCATGGACCACCTCGG Epo Rw TGAAGCACAGAAGCTCTTCGG Jak2 Fw ATCTGACCTTTCCATCTGGGG Jak2 Rw TGGTTGGGTGGATACCAGATC Stat5A Fw TTACTGAAGATCAAGCTGGGG
Data Sheet. TCF/LEF Reporter Kit Wnt / -catenin signaling pathway Catalog #: 60500
 Data Sheet TCF/LEF Reporter Kit Wnt / -catenin signaling pathway Catalog #: 60500 Background The Wnt / -catenin signaling pathway controls a large and diverse set of cell fate decisions in embryonic development,
Data Sheet TCF/LEF Reporter Kit Wnt / -catenin signaling pathway Catalog #: 60500 Background The Wnt / -catenin signaling pathway controls a large and diverse set of cell fate decisions in embryonic development,
All organisms respond to cellular stress by shutting off the regular protein synthesis and
 1. INTRODUCTION All organisms respond to cellular stress by shutting off the regular protein synthesis and starting the synthesis of a new set of proteins called heat shock proteins or more generally,
1. INTRODUCTION All organisms respond to cellular stress by shutting off the regular protein synthesis and starting the synthesis of a new set of proteins called heat shock proteins or more generally,
Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan).
 1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
RayBio Human PPAR-gamma Transcription Factor Activity Assay Kit
 RayBio Human PPAR-gamma Transcription Factor Activity Assay Kit Catalog #: TFEH-PPARg User Manual Jan 5, 2018 3607 Parkway Lane, Suite 200 Norcross, GA 30092 Tel: 1-888-494-8555 (Toll Free) or 770-729-2992,
RayBio Human PPAR-gamma Transcription Factor Activity Assay Kit Catalog #: TFEH-PPARg User Manual Jan 5, 2018 3607 Parkway Lane, Suite 200 Norcross, GA 30092 Tel: 1-888-494-8555 (Toll Free) or 770-729-2992,
Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas.
 Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas. Target Gene Non-target / Control BTG2 BRCA1 NFE2L2 Target Sequence ON-TARGET plus Non-targeting sirna # 1 (Cat# D-001810-01-05) sirna1: GAACCGACAUGCUCCCGGA
Supplementary Table 1. Sequences for BTG2 and BRCA1 sirnas. Target Gene Non-target / Control BTG2 BRCA1 NFE2L2 Target Sequence ON-TARGET plus Non-targeting sirna # 1 (Cat# D-001810-01-05) sirna1: GAACCGACAUGCUCCCGGA
Genomic DNA fragments identified by ChIP-chip assay for MR [28] corresponding to two
![Genomic DNA fragments identified by ChIP-chip assay for MR [28] corresponding to two Genomic DNA fragments identified by ChIP-chip assay for MR [28] corresponding to two](/thumbs/89/98549522.jpg) Supplemental Materials and Methods Genomic DNA fragments identified by ChIP-chip assay for MR [28] corresponding to two upstream regions of the human Klf9 gene (-5139 to -5771 bp and -3875 to -4211 bp)
Supplemental Materials and Methods Genomic DNA fragments identified by ChIP-chip assay for MR [28] corresponding to two upstream regions of the human Klf9 gene (-5139 to -5771 bp and -3875 to -4211 bp)
Supplementary Information Supplementary Figure 1
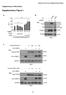 Bararia, Kwok et al, Supplementary Page 1 Supplementary Information Supplementary Figure 1 S1 Bararia, Kwok et al, Supplementary Page 2 Supplementary Figure 1 (cont.) S2 Bararia, Kwok et al, Supplementary
Bararia, Kwok et al, Supplementary Page 1 Supplementary Information Supplementary Figure 1 S1 Bararia, Kwok et al, Supplementary Page 2 Supplementary Figure 1 (cont.) S2 Bararia, Kwok et al, Supplementary
Investigating the regulation of mirna biogenesis and Argonaute2 by RNA binding proteins
 Investigating the regulation of mirna biogenesis and Argonaute2 by RNA binding proteins Patrick Peter Connerty Supervisor: Gyorgy Hutvagner Thesis Submitted for the Degree of Doctor of Philosophy (Science)
Investigating the regulation of mirna biogenesis and Argonaute2 by RNA binding proteins Patrick Peter Connerty Supervisor: Gyorgy Hutvagner Thesis Submitted for the Degree of Doctor of Philosophy (Science)
SUPPLEMENTAL MATERIAL
 SUPPLEMENTAL MATERIAL MATERIALS AND METHODS Generation of TSPOΔ/Δ murine embryonic fibroblasts Embryos were harvested from 13.5-day pregnant TSPOfl/fl mice. After dissection to eviscerate and remove the
SUPPLEMENTAL MATERIAL MATERIALS AND METHODS Generation of TSPOΔ/Δ murine embryonic fibroblasts Embryos were harvested from 13.5-day pregnant TSPOfl/fl mice. After dissection to eviscerate and remove the
UTR Reporter Vectors and Viruses
 UTR Reporter Vectors and Viruses 3 UTR, 5 UTR, Promoter Reporter (Version 1) Applied Biological Materials Inc. #1-3671 Viking Way Richmond, BC V6V 2J5 Canada Notice to Purchaser All abm products are for
UTR Reporter Vectors and Viruses 3 UTR, 5 UTR, Promoter Reporter (Version 1) Applied Biological Materials Inc. #1-3671 Viking Way Richmond, BC V6V 2J5 Canada Notice to Purchaser All abm products are for
Supplemental Table S1. RT-PCR primers used in this study
 Supplemental Table S1. RT-PCR primers used in this study -----------------------------------------------------------------------------------------------------------------------------------------------
Supplemental Table S1. RT-PCR primers used in this study -----------------------------------------------------------------------------------------------------------------------------------------------
Regulation of transcription by the MLL2 complex and MLL complex-associated AKAP95
 Supplementary Information Regulation of transcription by the complex and MLL complex-associated Hao Jiang, Xiangdong Lu, Miho Shimada, Yali Dou, Zhanyun Tang, and Robert G. Roeder Input HeLa NE IP lot:
Supplementary Information Regulation of transcription by the complex and MLL complex-associated Hao Jiang, Xiangdong Lu, Miho Shimada, Yali Dou, Zhanyun Tang, and Robert G. Roeder Input HeLa NE IP lot:
DNA, Genes and their Regula4on. Me#e Voldby Larsen PhD, Assistant Professor
 DNA, Genes and their Regula4on Me#e Voldby Larsen PhD, Assistant Professor Learning Objecer this talk, you should be able to Account for the structure of DNA and RNA including their similari
DNA, Genes and their Regula4on Me#e Voldby Larsen PhD, Assistant Professor Learning Objecer this talk, you should be able to Account for the structure of DNA and RNA including their similari
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
Position Description November 2018
 Position Description November 2018 MISSION The mission of St Vincent s Institute (SVI) is to carry out the highest quality laboratory based biomedical research, in order to make discoveries that will improve
Position Description November 2018 MISSION The mission of St Vincent s Institute (SVI) is to carry out the highest quality laboratory based biomedical research, in order to make discoveries that will improve
Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate
 Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
Supplementary Figure Legends Supplementary Fig. 1 related to Fig. 1 Clinical relevance of lncrna candidate BC041951 in gastric cancer. (A) The flow chart for selected candidate lncrnas in 660 up-regulated
OX40 MARKET LANDSCAPE
 OX40 Agonist OX40 MARKET LANDSCAPE OX40 (a.k.a. CD134) is a costimulatory molecule belonging to the TNF receptor family expressed primarily on activated effector T (T eff) cells and naive regulatory T
OX40 Agonist OX40 MARKET LANDSCAPE OX40 (a.k.a. CD134) is a costimulatory molecule belonging to the TNF receptor family expressed primarily on activated effector T (T eff) cells and naive regulatory T
HPV E6 oncoprotein targets histone methyltransferases for modulating specific. Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu,
 1 HPV E oncoprotein targets histone methyltransferases for modulating specific gene transcription 3 5 Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu, Cheng-Ming Chiang, Sheng-Chung
1 HPV E oncoprotein targets histone methyltransferases for modulating specific gene transcription 3 5 Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu, Cheng-Ming Chiang, Sheng-Chung
PS#2: We will begin at 10:30 am with Problem 1 in BH212. Assignments Problem 1- group 4 Problem 2- group 3 Problem 3- group 2 Problem 4- group 1
 PS#2: We will begin at 10:30 am with Problem 1 in BH212 Assignments Problem 1- group 4 Problem 2- group 3 Problem 3- group 2 Problem 4- group 1 Time limit: 25 min per group. The beginning: Designate one
PS#2: We will begin at 10:30 am with Problem 1 in BH212 Assignments Problem 1- group 4 Problem 2- group 3 Problem 3- group 2 Problem 4- group 1 Time limit: 25 min per group. The beginning: Designate one
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Old EXAM 1 BIO409/509 NAME. Please number your answers and write them on the attached, lined paper.
 Old EXAM 1 BIO409/509 NAME Please number your answers and write them on the attached, lined paper. 1) Describe euchromatin and heterochromatin. Which form of chromatin would the insulin gene be found in
Old EXAM 1 BIO409/509 NAME Please number your answers and write them on the attached, lined paper. 1) Describe euchromatin and heterochromatin. Which form of chromatin would the insulin gene be found in
Data Sheet. Application Monitor glucocorticoid signaling pathway activity. Screen activators or inhibitors of the glucocorticoid signaling pathway.
 Data Sheet GAL4 Reporter Kit (Glucocorticoid Receptor Pathway) Catalog #: w70533 Background The glucocorticoid signaling pathway plays an important role in development, fluid homeostasis, cognition, immune
Data Sheet GAL4 Reporter Kit (Glucocorticoid Receptor Pathway) Catalog #: w70533 Background The glucocorticoid signaling pathway plays an important role in development, fluid homeostasis, cognition, immune
Partial list of differentially expressed genes from cdna microarray, comparing MUC18-
 Supplemental Figure legends Table-1 Partial list of differentially expressed genes from cdna microarray, comparing MUC18- silenced and NT-transduced A375SM cells. Supplemental Figure 1 Effect of MUC-18
Supplemental Figure legends Table-1 Partial list of differentially expressed genes from cdna microarray, comparing MUC18- silenced and NT-transduced A375SM cells. Supplemental Figure 1 Effect of MUC-18
Transcriptional regulation of BRCA1 expression by a metabolic switch: Di, Fernandez, De Siervi, Longo, and Gardner. H3K4Me3
 ChIP H3K4Me3 enrichment.25.2.15.1.5 H3K4Me3 H3K4Me3 ctrl H3K4Me3 + E2 NS + E2 1. kb kb +82 kb Figure S1. Estrogen promotes entry of MCF-7 into the cell cycle but does not significantly change activation-associated
ChIP H3K4Me3 enrichment.25.2.15.1.5 H3K4Me3 H3K4Me3 ctrl H3K4Me3 + E2 NS + E2 1. kb kb +82 kb Figure S1. Estrogen promotes entry of MCF-7 into the cell cycle but does not significantly change activation-associated
Nucleosome Positioning and Organization Advanced Topics in Computa8onal Genomics
 Nucleosome Positioning and Organization 02-715 Advanced Topics in Computa8onal Genomics Nucleosome Core Nucleosome Core and Linker 147 bp DNA wrapping around nucleosome core Varying lengths of linkers
Nucleosome Positioning and Organization 02-715 Advanced Topics in Computa8onal Genomics Nucleosome Core Nucleosome Core and Linker 147 bp DNA wrapping around nucleosome core Varying lengths of linkers
Int. J. Mol. Sci. 2016, 17, 1259; doi: /ijms
 S1 of S5 Supplementary Materials: Fibroblast-Derived Extracellular Matrix Induces Chondrogenic Differentiation in Human Adipose-Derived Mesenchymal Stromal/Stem Cells in Vitro Kevin Dzobo, Taegyn Turnley,
S1 of S5 Supplementary Materials: Fibroblast-Derived Extracellular Matrix Induces Chondrogenic Differentiation in Human Adipose-Derived Mesenchymal Stromal/Stem Cells in Vitro Kevin Dzobo, Taegyn Turnley,
CRE/CREB Reporter Assay Kit camp/pka Cell Signaling Pathway Catalog #: 60611
 Data Sheet CRE/CREB Reporter Assay Kit camp/pka Cell Signaling Pathway Catalog #: 60611 Background The main role of the camp response element, or CRE, is mediating the effects of Protein Kinase A (PKA)
Data Sheet CRE/CREB Reporter Assay Kit camp/pka Cell Signaling Pathway Catalog #: 60611 Background The main role of the camp response element, or CRE, is mediating the effects of Protein Kinase A (PKA)
Myxoid Liposarcoma-Associated EWSR1-DDIT3 Selectively Represses Osteoblastic and Chondrocytic Transcription in Multipotent Mesenchymal Cells
 Myxoid Liposarcoma-Associated EWSR1-DDIT3 Selectively Represses Osteoblastic and Chondrocytic Transcription in Multipotent Mesenchymal Cells Kayo Suzuki 1, Yoshito Matsui 1 *, Mami Higashimoto 1, Yoshiharu
Myxoid Liposarcoma-Associated EWSR1-DDIT3 Selectively Represses Osteoblastic and Chondrocytic Transcription in Multipotent Mesenchymal Cells Kayo Suzuki 1, Yoshito Matsui 1 *, Mami Higashimoto 1, Yoshiharu
Gene Forward Primer Reverse Primer GAPDH ATCATCCCTGCCTCTACTGG GTCAGGTCCACCACTGACAC SSB1 AACTTCAGTGAGCCAAACCC GTTCTCAGAGGCTGGAGAGG
 Supplemental Data EXPERIMENTAL PROCEDURES Plasmids and Antibodies- Full length cdna of INT11 or INT12 were cloned into ps- Flag-SBP vector respectively. Anti-RNA pol II (RPB1) was purchased from Santa
Supplemental Data EXPERIMENTAL PROCEDURES Plasmids and Antibodies- Full length cdna of INT11 or INT12 were cloned into ps- Flag-SBP vector respectively. Anti-RNA pol II (RPB1) was purchased from Santa
OriGene GFC-Arrays for High-throughput Overexpression Screening of Human Gene Phenotypes
 OriGene GFC-Arrays for High-throughput Overexpression Screening of Human Gene Phenotypes High-throughput Gene Function Validation Tool Introduction sirna screening libraries enable scientists to identify
OriGene GFC-Arrays for High-throughput Overexpression Screening of Human Gene Phenotypes High-throughput Gene Function Validation Tool Introduction sirna screening libraries enable scientists to identify
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/362/ra12/dc1 Supplementary Materials for FOXP1 potentiates Wnt/β-catenin signaling in diffuse large B cell lymphoma Matthew P. Walker, Charles M. Stopford, Maria
www.sciencesignaling.org/cgi/content/full/8/362/ra12/dc1 Supplementary Materials for FOXP1 potentiates Wnt/β-catenin signaling in diffuse large B cell lymphoma Matthew P. Walker, Charles M. Stopford, Maria
Supplementary Figure 1. TSA (10 nmol/l), non-class-selective HDAC inhibitor, potentiates
 Supplementary Figure 1. TSA (10 nmol/l), non-class-selective HDAC inhibitor, potentiates vascular calcification (VC). (a) Von Kossa staining shows that TSA potentiated the Pi-induced VC. Scale bar, 100
Supplementary Figure 1. TSA (10 nmol/l), non-class-selective HDAC inhibitor, potentiates vascular calcification (VC). (a) Von Kossa staining shows that TSA potentiated the Pi-induced VC. Scale bar, 100
Making Deep Learning Understandable for Analyzing Sequen;al Data about Gene Regula;on. Dr. Yanjun Qi 2017/11/26
 Making Deep Learning Understandable for Analyzing Sequen;al Data about Gene Regula;on Dr. Yanjun Qi 2017/11/26 Roadmap ² Background of Machine Learning ² Background of Sequen?al Data about Gene Regula?on
Making Deep Learning Understandable for Analyzing Sequen;al Data about Gene Regula;on Dr. Yanjun Qi 2017/11/26 Roadmap ² Background of Machine Learning ² Background of Sequen?al Data about Gene Regula?on
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 Generation of the AARE-Gene system construct. (a) Position and sequence alignment of AAREs extracted from human Trb3, Chop or Atf3 promoters. AARE core sequences are boxed in grey.
Supplementary Figure 1 Generation of the AARE-Gene system construct. (a) Position and sequence alignment of AAREs extracted from human Trb3, Chop or Atf3 promoters. AARE core sequences are boxed in grey.
Supplementary Information. HBx-upregulated lncrna UCA1 promotes cell growth and tumorigenesis
 Supplementary Information HBx-upregulated lncrna UCA1 promotes cell growth and tumorigenesis by recruiting EZH2 and repressing p27kip1/cdk2 signaling Jiao-Jiao Hu 1, Wei Song 1, Shao-Dan Zhang 1, Xiao-Hui
Supplementary Information HBx-upregulated lncrna UCA1 promotes cell growth and tumorigenesis by recruiting EZH2 and repressing p27kip1/cdk2 signaling Jiao-Jiao Hu 1, Wei Song 1, Shao-Dan Zhang 1, Xiao-Hui
mir-24-mediated down-regulation of H2AX suppresses DNA repair
 Supplemental Online Material mir-24-mediated down-regulation of H2AX suppresses DNA repair in terminally differentiated blood cells Ashish Lal 1,4, Yunfeng Pan 2,4, Francisco Navarro 1,4, Derek M. Dykxhoorn
Supplemental Online Material mir-24-mediated down-regulation of H2AX suppresses DNA repair in terminally differentiated blood cells Ashish Lal 1,4, Yunfeng Pan 2,4, Francisco Navarro 1,4, Derek M. Dykxhoorn
SureSilencing sirna Array Technology Overview
 SureSilencing sirna Array Technology Overview Pathway-Focused sirna-based RNA Interference Topics to be Covered Who is SuperArray? Brief Introduction to RNA Interference Challenges Facing RNA Interference
SureSilencing sirna Array Technology Overview Pathway-Focused sirna-based RNA Interference Topics to be Covered Who is SuperArray? Brief Introduction to RNA Interference Challenges Facing RNA Interference
a KYSE270-CON KYSE270-Id1
 a KYSE27-CON KYSE27- shcon shcon sh b Human Mouse CD31 Relative MVD 3.5 3 2.5 2 1.5 1.5 *** *** c KYSE15 KYSE27 sirna (nm) 5 1 Id2 Id2 sirna 5 1 sirna (nm) 5 1 Id2 sirna 5 1 Id2 [h] (pg per ml) d 3 2 1
a KYSE27-CON KYSE27- shcon shcon sh b Human Mouse CD31 Relative MVD 3.5 3 2.5 2 1.5 1.5 *** *** c KYSE15 KYSE27 sirna (nm) 5 1 Id2 Id2 sirna 5 1 sirna (nm) 5 1 Id2 sirna 5 1 Id2 [h] (pg per ml) d 3 2 1
Regulatory Framework for Gene Therapies Incorpora:ng Human Genome Edi:ng
 Regulatory Framework for Gene Therapies Incorpora:ng Human Genome Edi:ng Anna Kwilas, Ph.D. CMC Reviewer, Gene Therapy Branch Division of Cellular and Gene Therapies Office of Tissues and Advanced Therapies
Regulatory Framework for Gene Therapies Incorpora:ng Human Genome Edi:ng Anna Kwilas, Ph.D. CMC Reviewer, Gene Therapy Branch Division of Cellular and Gene Therapies Office of Tissues and Advanced Therapies
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/9/eaat5401/dc1 Supplementary Materials for GLK-IKKβ signaling induces dimerization and translocation of the AhR-RORγt complex in IL-17A induction and autoimmune
advances.sciencemag.org/cgi/content/full/4/9/eaat5401/dc1 Supplementary Materials for GLK-IKKβ signaling induces dimerization and translocation of the AhR-RORγt complex in IL-17A induction and autoimmune
Molecular Biology (BIOL 4320) Exam #1 March 12, 2002
 Molecular Biology (BIOL 4320) Exam #1 March 12, 2002 Name KEY SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number.
Molecular Biology (BIOL 4320) Exam #1 March 12, 2002 Name KEY SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number.
supplementary information
 DOI: 10.1038/ncb2116 Figure S1 CDK phosphorylation of EZH2 in cells. (a) Comparison of candidate CDK phosphorylation sites on EZH2 with known CDK substrates by multiple sequence alignments. (b) CDK1 and
DOI: 10.1038/ncb2116 Figure S1 CDK phosphorylation of EZH2 in cells. (a) Comparison of candidate CDK phosphorylation sites on EZH2 with known CDK substrates by multiple sequence alignments. (b) CDK1 and
FEBRUARY 12, 2010 VOLUME 285 NUMBER 7 JOURNAL OF BIOLOGICAL CHEMISTRY 4847
 THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 285, NO. 7, pp. 4847 4858, February 12, 2010 2010 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. The Tumor Suppressor
THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 285, NO. 7, pp. 4847 4858, February 12, 2010 2010 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. The Tumor Suppressor
Data Sheet. SBE Reporter Kit (TGFβ/SMAD signaling pathway) Catalog #: 60654
 Data Sheet SBE Reporter Kit (TGFβ/SMAD signaling pathway) Catalog #: 60654 Background The transforming growth factor beta (TGFβ) signaling pathway is involved in a diverse range of cell processes such
Data Sheet SBE Reporter Kit (TGFβ/SMAD signaling pathway) Catalog #: 60654 Background The transforming growth factor beta (TGFβ) signaling pathway is involved in a diverse range of cell processes such
supplementary information
 DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
DOI: 1.138/ncb1839 a b Control 1 2 3 Control 1 2 3 Fbw7 Smad3 1 2 3 4 1 2 3 4 c d IGF-1 IGF-1Rβ IGF-1Rβ-P Control / 1 2 3 4 Real-time RT-PCR Relative quantity (IGF-1/ mrna) 2 1 IGF-1 1 2 3 4 Control /
supplementary information
 Figure S1 ZEB1 full length mrna. (a) Analysis of the ZEB1 mrna using the UCSC genome browser (http://genome.ucsc.edu) revealed truncation of the annotated Refseq sequence (NM_030751). The probable terminus
Figure S1 ZEB1 full length mrna. (a) Analysis of the ZEB1 mrna using the UCSC genome browser (http://genome.ucsc.edu) revealed truncation of the annotated Refseq sequence (NM_030751). The probable terminus
TITLE: Restoration of Wild-Type Activity to Mutant p53 in Prostate Cancer: A Novel Therapeutic Approach
 AD Award Number: W81XWH-05-1-0109 TITLE: Restoration of Wild-Type Activity to Mutant p53 in Prostate Cancer: A Novel Therapeutic Approach PRINCIPAL INVESTIGATOR: James Manfredi, Ph.D. CONTRACTING ORGANIZATION:
AD Award Number: W81XWH-05-1-0109 TITLE: Restoration of Wild-Type Activity to Mutant p53 in Prostate Cancer: A Novel Therapeutic Approach PRINCIPAL INVESTIGATOR: James Manfredi, Ph.D. CONTRACTING ORGANIZATION:
CELL BIOLOGY - CLUTCH CH. 7 - GENE EXPRESSION.
 !! www.clutchprep.com CONCEPT: CONTROL OF GENE EXPRESSION BASICS Gene expression is the process through which cells selectively to express some genes and not others Every cell in an organism is a clone
!! www.clutchprep.com CONCEPT: CONTROL OF GENE EXPRESSION BASICS Gene expression is the process through which cells selectively to express some genes and not others Every cell in an organism is a clone
Supplemental Methods Cell lines and culture
 Supplemental Methods Cell lines and culture AGS, CL5, BT549, and SKBR were propagated in RPMI 64 medium (Mediatech Inc., Manassas, VA) supplemented with % fetal bovine serum (FBS, Atlanta Biologicals,
Supplemental Methods Cell lines and culture AGS, CL5, BT549, and SKBR were propagated in RPMI 64 medium (Mediatech Inc., Manassas, VA) supplemented with % fetal bovine serum (FBS, Atlanta Biologicals,
Molecular Cloning. Joseph Sambrook. David W. Russell A LABORATORY MANUAL COLD SPRING HARBOR LABORATORY PRESS VOLUME.
 VOLUME Molecular Cloning A LABORATORY MANUAL THIRD EDITION www.molecularcloning.com Joseph Sambrook PETER MACCALLUM CANCER INSTITUTE AND THE UNIVERSITY OF MELBOURNE, AUSTRALIA David W. Russell UNIVERSITY
VOLUME Molecular Cloning A LABORATORY MANUAL THIRD EDITION www.molecularcloning.com Joseph Sambrook PETER MACCALLUM CANCER INSTITUTE AND THE UNIVERSITY OF MELBOURNE, AUSTRALIA David W. Russell UNIVERSITY
Supplementary Information
 Supplementary Information Sam68 modulates the promoter specificity of NF-κB and mediates expression of CD25 in activated T cells Kai Fu 1, 6, Xin Sun 1, 6, Wenxin Zheng 1, 6, Eric M. Wier 1, Andrea Hodgson
Supplementary Information Sam68 modulates the promoter specificity of NF-κB and mediates expression of CD25 in activated T cells Kai Fu 1, 6, Xin Sun 1, 6, Wenxin Zheng 1, 6, Eric M. Wier 1, Andrea Hodgson
Legend for Supplemental Figures and Tables
 Legend for Supplemental Figures and Tables Supplemental Fig. 1. Negative regulation of the CYP27B1 promoter in a ligand-dependent manner (A) OK-P cells were transfected with pcdna-trα, pcdna-trβ1 or pcdna3
Legend for Supplemental Figures and Tables Supplemental Fig. 1. Negative regulation of the CYP27B1 promoter in a ligand-dependent manner (A) OK-P cells were transfected with pcdna-trα, pcdna-trβ1 or pcdna3
sherwood - UltramiR shrna Collections
 sherwood - UltramiR shrna Collections Incorporating advances in shrna design and processing for superior potency and specificity sherwood - UltramiR shrna Collections Enabling Discovery Across the Genome
sherwood - UltramiR shrna Collections Incorporating advances in shrna design and processing for superior potency and specificity sherwood - UltramiR shrna Collections Enabling Discovery Across the Genome
DNA Technology and Genomics. Ch. 20
 DNA Technology and Genomics Ch. 20 Gene8c engineering Gene$c engineering: the direct manipula8on of genes for prac8cal purposes When genes from two different sources are combined in vitro into the same
DNA Technology and Genomics Ch. 20 Gene8c engineering Gene$c engineering: the direct manipula8on of genes for prac8cal purposes When genes from two different sources are combined in vitro into the same
Supplementary Figure Legends. Figure-S1 Molecular basis of ASS1 deficiency in myxofibrosarcoma: (A)
 Supplementary Figure Legends Figure-S1 Molecular basis of ASS1 deficiency in myxofibrosarcoma: (A) High-resolution oligonucleotide-based array comparative genomic hybridization shows normal DNA copy number
Supplementary Figure Legends Figure-S1 Molecular basis of ASS1 deficiency in myxofibrosarcoma: (A) High-resolution oligonucleotide-based array comparative genomic hybridization shows normal DNA copy number
Data Sheet. CRE/CREB Reporter Assay Kit (camp/pka Cell Signaling Pathway) Catalog #: 60611
 Data Sheet CRE/CREB Reporter Assay Kit (camp/pka Cell Signaling Pathway) Catalog #: 60611 Background The main role of the camp response element, or CRE, is mediating the effects of Protein Kinase A (PKA)
Data Sheet CRE/CREB Reporter Assay Kit (camp/pka Cell Signaling Pathway) Catalog #: 60611 Background The main role of the camp response element, or CRE, is mediating the effects of Protein Kinase A (PKA)
Cis ac$ng transcrip$onal elements with nega$ve regulatory func$on
 Cis ac$ng transcrip$onal elements with nega$ve regulatory func$on Laura Elnitski, PhD Na$onal Human Genome Research Ins$tute Na$onal Ins$tutes of Health 2 Non coding regulatory signals affec$ng transcrip$on
Cis ac$ng transcrip$onal elements with nega$ve regulatory func$on Laura Elnitski, PhD Na$onal Human Genome Research Ins$tute Na$onal Ins$tutes of Health 2 Non coding regulatory signals affec$ng transcrip$on
Chapter 14 Regulation of Transcription
 Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
7.012 Problem Set 5. Question 1
 Name Section 7.012 Problem Set 5 Question 1 While studying the problem of infertility, you attempt to isolate a hypothetical rabbit gene that accounts for the prolific reproduction of rabbits. After much
Name Section 7.012 Problem Set 5 Question 1 While studying the problem of infertility, you attempt to isolate a hypothetical rabbit gene that accounts for the prolific reproduction of rabbits. After much
Gene regulation V Biochemistry 302. March 6, 2006
 Gene regulation V Biochemistry 302 March 6, 2006 Common structural motifs associated with transcriptional regulatory proteins Helix-turn-helix Prokaryotic repressors and activators Eukaryotic homeodomain
Gene regulation V Biochemistry 302 March 6, 2006 Common structural motifs associated with transcriptional regulatory proteins Helix-turn-helix Prokaryotic repressors and activators Eukaryotic homeodomain
ES, MEF, F9, C2C12, Hela, HEK293, Jurkat, LNCaP, RAG, and MCF7 cells were
 Supplemental Materials and Methods Cell culture and stable cell lines ES, MEF, F9, C2C12, Hela, HEK293, Jurkat, LNCaP, RAG, and MCF7 cells were maintained in DMEM medium containing 10% fetal bovine serum
Supplemental Materials and Methods Cell culture and stable cell lines ES, MEF, F9, C2C12, Hela, HEK293, Jurkat, LNCaP, RAG, and MCF7 cells were maintained in DMEM medium containing 10% fetal bovine serum
Transcriptional regulation of IFN-l genes in Hepatitis C virus-infected hepatocytes via IRF-3 IRF-7 NF- B complex
 POSTER PRESENTATION Transcriptional regulation of IFN-l genes in Hepatitis C virus-infected hepatocytes via IRF-3 IRF-7 NF- B complex Hai-Chon Lee *, Je-In Youn, Kyungwha Lee, Hwanyul Yong, Seung-Yong
POSTER PRESENTATION Transcriptional regulation of IFN-l genes in Hepatitis C virus-infected hepatocytes via IRF-3 IRF-7 NF- B complex Hai-Chon Lee *, Je-In Youn, Kyungwha Lee, Hwanyul Yong, Seung-Yong
Supplementary Figures and Figure legends
 Supplementary Figures and Figure legends 3 Supplementary Figure 1. Conditional targeting construct for the murine Satb1 locus with a modified FLEX switch. Schematic of the wild type Satb1 locus; the conditional
Supplementary Figures and Figure legends 3 Supplementary Figure 1. Conditional targeting construct for the murine Satb1 locus with a modified FLEX switch. Schematic of the wild type Satb1 locus; the conditional
A Survey of Genetic Methods
 IBS 8102 Cell, Molecular, and Developmental Biology A Survey of Genetic Methods January 24, 2008 DNA RNA Hybridization ** * radioactive probe reverse transcriptase polymerase chain reaction RT PCR DNA
IBS 8102 Cell, Molecular, and Developmental Biology A Survey of Genetic Methods January 24, 2008 DNA RNA Hybridization ** * radioactive probe reverse transcriptase polymerase chain reaction RT PCR DNA
Recombination Lecture, Dr. Aguilera 2/17/2014
 Lymphocytes and Antigen Receptors Thymus T-Cells Lymph nodes Spleen } T+B-cells Paper Presentation: Bone Marrow Stem cells and B-cells Nat. Rev. Immunol. STEM CELL CLP Committed Lymphocyte Precursor T-cells
Lymphocytes and Antigen Receptors Thymus T-Cells Lymph nodes Spleen } T+B-cells Paper Presentation: Bone Marrow Stem cells and B-cells Nat. Rev. Immunol. STEM CELL CLP Committed Lymphocyte Precursor T-cells
Introducing new DNA into the genome requires cloning the donor sequence, delivery of the cloned DNA into the cell, and integration into the genome.
 Key Terms Chapter 32: Genetic Engineering Cloning describes propagation of a DNA sequence by incorporating it into a hybrid construct that can be replicated in a host cell. A cloning vector is a plasmid
Key Terms Chapter 32: Genetic Engineering Cloning describes propagation of a DNA sequence by incorporating it into a hybrid construct that can be replicated in a host cell. A cloning vector is a plasmid
AP Biology Day 34. Monday, November 14, 2016
 AP Biology Day 34 Monday, November 14, 2016 Essen%al knowledge standards 3.A.1: DNA, and in some cases RNA, is the primary source of heritable informa%on 3.A.1.e: Gene%c engineering techniques can manipulate
AP Biology Day 34 Monday, November 14, 2016 Essen%al knowledge standards 3.A.1: DNA, and in some cases RNA, is the primary source of heritable informa%on 3.A.1.e: Gene%c engineering techniques can manipulate
RNA-Guided Gene Activation by CRISPR-Cas9-Based Transcription Factors
 Supplementary Information RNA-Guided Gene Activation by CRISPR-Cas9-Based Transcription Factors Pablo Perez-Pinera 1, Daniel D. Kocak 1, Christopher M. Vockley 2,3, Andrew F. Adler 1, Ami M. Kabadi 1,
Supplementary Information RNA-Guided Gene Activation by CRISPR-Cas9-Based Transcription Factors Pablo Perez-Pinera 1, Daniel D. Kocak 1, Christopher M. Vockley 2,3, Andrew F. Adler 1, Ami M. Kabadi 1,
Chapter 20 Recombinant DNA Technology. Copyright 2009 Pearson Education, Inc.
 Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
Chapter 20 Recombinant DNA Technology Copyright 2009 Pearson Education, Inc. 20.1 Recombinant DNA Technology Began with Two Key Tools: Restriction Enzymes and DNA Cloning Vectors Recombinant DNA refers
Experimental Tools and Resources Available in Arabidopsis. Manish Raizada, University of Guelph, Canada
 Experimental Tools and Resources Available in Arabidopsis Manish Raizada, University of Guelph, Canada Community website: The Arabidopsis Information Resource (TAIR) at http://www.arabidopsis.org Can order
Experimental Tools and Resources Available in Arabidopsis Manish Raizada, University of Guelph, Canada Community website: The Arabidopsis Information Resource (TAIR) at http://www.arabidopsis.org Can order
INVESTIGATION OF THE BINDING SPECIFICITY OF IGF-IR USING MONOCLONAL ANTIBODIES
 INVESTIGATION OF THE BINDING SPECIFICITY OF IGF-IR USING MONOCLONAL ANTIBODIES By Mehrnaz Keyhanfar, Pharm.D. A thesis submitted to the University of Adelaide, South Australia in fulfilment of the requirements
INVESTIGATION OF THE BINDING SPECIFICITY OF IGF-IR USING MONOCLONAL ANTIBODIES By Mehrnaz Keyhanfar, Pharm.D. A thesis submitted to the University of Adelaide, South Australia in fulfilment of the requirements
Functional characterisation
 Me Me Ac Ac Functional characterisation How can we know if measured changes in DNA methylation and function (phenotype) and linked, and in what way? DNMT Dianne Ford Professor of Molecular Nutritional
Me Me Ac Ac Functional characterisation How can we know if measured changes in DNA methylation and function (phenotype) and linked, and in what way? DNMT Dianne Ford Professor of Molecular Nutritional
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the
 Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Supplementary Figure S1
 Supplementary Figure S1 Luciferase ctivity (%) 5 4 3 2 1 HeLa E2 OHT Figure S1. SENP2 represses estradiol-dependent transcriptional activity in HeLa cells. HeLa cells were transiently transfected with
Supplementary Figure S1 Luciferase ctivity (%) 5 4 3 2 1 HeLa E2 OHT Figure S1. SENP2 represses estradiol-dependent transcriptional activity in HeLa cells. HeLa cells were transiently transfected with
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX
 Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Transfection of CRISPR/Cas9 Nuclease NLS ribonucleoprotein (RNP) into adherent mammalian cells using Lipofectamine RNAiMAX INTRODUCTION The CRISPR/Cas genome editing system consists of a single guide RNA
Comparative Analysis of Argonaute-Dependent Small RNA Pathways in Drosophila
 Molecular Cell, Volume 32 Supplemental Data Comparative Analysis of Argonaute-Dependent Small RNA Pathways in Drosophila Rui Zhou, Ikuko Hotta, Ahmet M. Denli, Pengyu Hong, Norbert Perrimon, and Gregory
Molecular Cell, Volume 32 Supplemental Data Comparative Analysis of Argonaute-Dependent Small RNA Pathways in Drosophila Rui Zhou, Ikuko Hotta, Ahmet M. Denli, Pengyu Hong, Norbert Perrimon, and Gregory
Khaled_Fig. S MITF TYROSINASE R²= MITF PDE4D R²= Variance from mean mrna expression
 Khaled_Fig. S Variance from mean mrna expression.8 2 MITF.6 TYROSINASE.4 R²=.73.2.8.6.4.2.8.6.4.2.8.6.4.2 MALME3M SKMEL28 UACC257 MITF PDE4D R²=.63 4.5 5 MITF 3.5 4 PDE4B 2.5 3 R²=.2.5 2.5.8.6.4.2.8.6.4.2
Khaled_Fig. S Variance from mean mrna expression.8 2 MITF.6 TYROSINASE.4 R²=.73.2.8.6.4.2.8.6.4.2.8.6.4.2 MALME3M SKMEL28 UACC257 MITF PDE4D R²=.63 4.5 5 MITF 3.5 4 PDE4B 2.5 3 R²=.2.5 2.5.8.6.4.2.8.6.4.2
C) Based on your knowledge of transcription, what factors do you expect to ultimately identify? (1 point)
 Question 1 Unlike bacterial RNA polymerase, eukaryotic RNA polymerases need additional assistance in finding a promoter and transcription start site. You wish to reconstitute accurate transcription by
Question 1 Unlike bacterial RNA polymerase, eukaryotic RNA polymerases need additional assistance in finding a promoter and transcription start site. You wish to reconstitute accurate transcription by
Other Polymerase Chain Reac3on (PCR) Methods
 Other Polymerase Chain Reac3on (PCR) Methods What is PCR? The Reaction Components 1) Target DNA - contains the sequence to be amplified. 2) Pair of Primers - oligonucleotides that define the sequence to
Other Polymerase Chain Reac3on (PCR) Methods What is PCR? The Reaction Components 1) Target DNA - contains the sequence to be amplified. 2) Pair of Primers - oligonucleotides that define the sequence to
ASPP1 Fw GGTTGGGAATCCACGTGTTG ASPP1 Rv GCCATATCTTGGAGCTCTGAGAG
 Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
TRANSGENIC ANIMALS. -transient transfection of cells -stable transfection of cells. - Two methods to produce transgenic animals:
 TRANSGENIC ANIMALS -transient transfection of cells -stable transfection of cells - Two methods to produce transgenic animals: 1- DNA microinjection - random insertion 2- embryonic stem cell-mediated gene
TRANSGENIC ANIMALS -transient transfection of cells -stable transfection of cells - Two methods to produce transgenic animals: 1- DNA microinjection - random insertion 2- embryonic stem cell-mediated gene
ERYTHROPOIESIS MEDIATED BY GATA-1-FOG-1-Mi2 COMPLEX BINDING AT ACCEPTED THE -566 GATA SITE.
 MCB Accepts, published online ahead of print on 17 March 2008 Mol. Cell. Biol. doi:10.1128/mcb.01858-07 Copyright 2008, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
MCB Accepts, published online ahead of print on 17 March 2008 Mol. Cell. Biol. doi:10.1128/mcb.01858-07 Copyright 2008, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
Document S1. Supplemental Experimental Procedures and Three Figures (see next page)
 Supplemental Data Document S1. Supplemental Experimental Procedures and Three Figures (see next page) Table S1. List of Candidate Genes Identified from the Screen. Candidate genes, corresponding dsrnas
Supplemental Data Document S1. Supplemental Experimental Procedures and Three Figures (see next page) Table S1. List of Candidate Genes Identified from the Screen. Candidate genes, corresponding dsrnas
Biosc10 schedule reminders
 Biosc10 schedule reminders Review of molecular biology basics DNA Is each person s DNA the same, or unique? What does DNA look like? What are the three parts of each DNA nucleotide Which DNA bases pair,
Biosc10 schedule reminders Review of molecular biology basics DNA Is each person s DNA the same, or unique? What does DNA look like? What are the three parts of each DNA nucleotide Which DNA bases pair,
In any cell type, a large part of the genes are kept inactive (not expressed) by gene silencing
 Part 1.week 4 8. Chromatin 9. Epigenetics and imprintingi 10. Post-transcriptional control and proteomics 8.1 Nucleosomes 8.2 Covalent modification of histones 8.2 The histone code 8.3 Nucleosome transitions
Part 1.week 4 8. Chromatin 9. Epigenetics and imprintingi 10. Post-transcriptional control and proteomics 8.1 Nucleosomes 8.2 Covalent modification of histones 8.2 The histone code 8.3 Nucleosome transitions
TRIM31 is recruited to mitochondria after infection with SeV.
 Supplementary Figure 1 TRIM31 is recruited to mitochondria after infection with SeV. (a) Confocal microscopy of TRIM31-GFP transfected into HEK293T cells for 24 h followed with SeV infection for 6 h. MitoTracker
Supplementary Figure 1 TRIM31 is recruited to mitochondria after infection with SeV. (a) Confocal microscopy of TRIM31-GFP transfected into HEK293T cells for 24 h followed with SeV infection for 6 h. MitoTracker
peg10, an imprinted gene, plays a crucial role in adipocyte differentiation
 FEBS Letters 581 (2007) 4272 4278 peg10, an imprinted gene, plays a crucial role in adipocyte differentiation Tomoaki Hishida, Kumiko Naito, Shigehiro Osada, Makoto Nishizuka, Masayoshi Imagawa * Department
FEBS Letters 581 (2007) 4272 4278 peg10, an imprinted gene, plays a crucial role in adipocyte differentiation Tomoaki Hishida, Kumiko Naito, Shigehiro Osada, Makoto Nishizuka, Masayoshi Imagawa * Department
