RNA secondary structure prediction and analysis
|
|
|
- Susanna Nicholson
- 6 years ago
- Views:
Transcription
1 RNA secondary structure prediction and analysis 1
2 Resources Lecture Notes from previous years: Takis Benos Covariance algorithm: Eddy and Durbin, Nucleic Acids Research, v22: 11, 2079 Useful lecture slides from Larry Ruzzo, U of Washington and Phillip Compeau, UCSD 2
3 Outline RNA Folding Dynamic Programming for RNA secondary structure prediction Nussinov et al and Zucker et al algorithms Covariance Model Eddy and Durbin 3
4 Various types of RNA messenger RNA (mrna) transfer RNA (trna) Ribosomal RNA (rrna) small interfering RNA (sirna) micro RNA (mirna) small nuclear RNA (snrna) small nucleolar RNA (snorna) guide RNA (grna) efference RNA(eRNA) 4
5 Non coding RNA (ncrna) RNA that isn t translated into protein Includes: trna, rrna, snrna, snorna, mirna, grna, erna, prna, tmrna What about mrna? 5 methylated cap 5 -UTR (un-translated regions) CDS (coding sequence) 3 -UTR Poly-A tail mrna contrains untranslated regions (5 UTR, 3 UTR), but UTRs are not considered ncrna 5
6 RNA Basics RNA bases: A, C, G, U Watson-Crick Pair A-U (~ 2 kcal/mol) G-C (~ 3 kcal/mol) Wobble pair G-U (~ 1 kcal/mol) Non-Canonical pairs (modified suitably) Bases can only pair with one other base Two hydrogen bonds 6
7 RNA Basics RNA bases: A, C, G, U Canonical Base Pairs A-U G-C G-U Bases can only pair with one other base Three hydrogen bonds => more stable 7
8 RNA Basics RNA bases: A, C, G, U Canonical Base Pairs A-U G-C G-U Bases can only pair with one other base wobble pairing 8
9 9
10 10
11 11
12 RNA Secondary Structure/Motifs stem Pseudoknot Single stranded Interior Loop Bulge loop Multiloop or junction Hairpin loop 12
13 13
14 14
15 15
16 RNA importance Ribozymes (RNA enzymes) Retroviruses Effects on transcription, translation, splicing.. Functional RNAs: rrna, trna, snrna, snorna, microrna, RNAi, riboswitches, regulatory elements in 3 and 5 UTR 16
17 RNA motifs Function depends on sequence and secondary structure Functions include Protein binding Basepairing to another RNA Modifying a nucleic acid bond Types: Single-strand regions Helices (or stems) Bulges Hairpin loops Internal loops Junctions 17
18 RNA Motifs Regulatory Effects Regulations of translations Processing of RNA Catalytic modification of other RNAs Transport and position in the cell Stability of RNA-transcript Expression of encoded proteins 18
19 Why predict structures? Knowing the shape of a biomolecule is invaluable in drug design and understanding disease mechanisms Current physical methods (X-Ray, NMR) are too expensive and timeconsuming Predict shape from sequence of bases Four basic structures: helices, loops, bulges and junctions 19
20 RNA secondary structure What makes RNA fold? Problem: given an RNA sequence, find the set of base pairs that is correct or optimal Maximize number of base pairs (Nussinov et al) Minimize energy (Zucker et al) Search problem: very high number of possible structures Algorithm: dynamic programming Cannot handle pseudoknots 20
21 21
22 22
23 23
24 24
25 25
26 26
27 Base Pair Maximization: Dynamic Programming S(i, j) is the folding of the RNA subsequence of the strand from index i to index j which results in the highest number of base pairs. Recurrence: Images Sean Eddy 27
28 Base Pair Maximization: Dynamic Programming S(i, j) is the folding of the RNA subsequence of the strand from index i to index j which results in the highest number of base pairs. Recurrence: S i, j max Images Sean Eddy 28
29 Base Pair Maximization: Dynamic Programming S(i, j) is the folding of the RNA subsequence of the strand from index i to index j which results in the highest number of base pairs. Recurrence: Base pair at i and j S i, j max S i 1, j 1 1 (if i, j base pair) Images Sean Eddy 29
30 Base Pair Maximization: Dynamic Programming S(i, j) is the folding of the RNA subsequence of the strand from index i to index j which results in the highest number of base pairs. Recurrence: Base pair at i and j Unmatched at i S i, j max S i 1, j 1 1 (if i, j base pair) S i 1, j Images Sean Eddy 30
31 Base Pair Maximization: Dynamic Programming S(i, j) is the folding of the RNA subsequence of the strand from index i to index j which results in the highest number of base pairs. Recurrence: Base pair at i and j Unmatched at i Unmatched at j S i, j max S i 1, j 1 1 (if i, j base pair) S i 1, j S i, j 1 Images Sean Eddy 31
32 Base Pair Maximization: Dynamic Programming S(i, j) is the folding of the RNA subsequence of the strand from index i to index j which results in the highest number of base pairs. Recurrence: Base pair at i and j Unmatched at i Unmatched at j S i, j max Images Sean Eddy S i 1, j 1 1 (if i, j base pair) S i 1, j S i, j 1 max 1 k j S i, k S k 1, j Bifurcation 32
33 Base Pair Maximization: Dynamic Programming Alignment Method: Align RNA strand to itself Score increases for feasible base pairs Each score independent of overall structure Bifurcation adds extra dimension Images Sean Eddy Images Sean Eddy 33
34 Base Pair Maximization: Dynamic Programming Alignment Method: Align RNA strand to itself Score increases for feasible base pairs Each score independent of overall structure Bifurcation adds extra dimension Initialize first two diagonals to 0 Images Sean Eddy Images Sean Eddy 34
35 Base Pair Maximization: Dynamic Programming Alignment Method: Align RNA strand to itself Score increases for feasible base pairs Each score independent of overall structure Bifurcation adds extra dimension Fill in squares sweeping diagonally Images Sean Eddy Images Sean Eddy 35
36 Base Pair Maximization: Dynamic Programming Alignment Method: Align RNA strand to itself Score increases for feasible base pairs Each score independent of overall structure Bifurcation adds extra dimension Fill in squares sweeping diagonally Images Sean Eddy Images Sean Eddy 36
37 Base Pair Maximization: Dynamic Programming Alignment Method: Align RNA strand to itself Score increases for feasible base pairs Each score independent of overall structure Bifurcation adds extra dimension Bases cannot pair Images Sean Eddy Images Sean Eddy 37
38 Base Pair Maximization: Dynamic Programming Alignment Method: Align RNA strand to itself Score increases for feasible base pairs Each score independent of overall structure Bifurcation adds extra dimension Bases can pair, similar to matched alignment Images Sean Eddy Images Sean Eddy 38
39 Base Pair Maximization: Dynamic Programming Alignment Method: Align RNA strand to itself Score increases for feasible base pairs Each score independent of overall structure Bifurcation adds extra dimension Dynamic Programming possible paths Images Sean Eddy Images Sean Eddy 39
40 Base Pair Maximization: Dynamic Programming Alignment Method: Align RNA strand to itself Score increases for feasible base pairs Each score independent of overall structure Bifurcation adds extra dimension S(i, j 1) Dynamic Programming possible paths Images Sean Eddy Images Sean Eddy 40
41 Base Pair Maximization: Dynamic Programming Alignment Method: Align RNA strand to itself Score increases for feasible base pairs Each score independent of overall structure Bifurcation adds extra dimension S(i + 1, j) Dynamic Programming possible paths Images Sean Eddy Images Sean Eddy 41
42 Base Pair Maximization: Dynamic Programming Alignment Method: Align RNA strand to itself Score increases for feasible base pairs Each score independent of overall structure Bifurcation adds extra dimension Dynamic Programming possible paths Images Sean Eddy S(i + 1, j 1) +1 42
43 Base Pair Maximization: Dynamic Programming Alignment Method: Align RNA strand to itself Score increases for feasible base pairs Each score independent of overall structure Bifurcation adds extra dimension Bifurcation add values for all k Images Sean Eddy 43
44 Base Pair Maximization: Dynamic Programming Alignment Method: Align RNA strand to itself Score increases for feasible base pairs Each score independent of overall structure Bifurcation adds extra dimension Bifurcation add values for all k Images Sean Eddy 44
45 Base Pair Maximization: Dynamic Programming Alignment Method: Align RNA strand to itself Score increases for feasible base pairs Each score independent of overall structure Bifurcation adds extra dimension Bifurcation add values for all k Images Sean Eddy 45
46 Base Pair Maximization: Dynamic Programming Alignment Method: Align RNA strand to itself Score increases for feasible base pairs Each score independent of overall structure Bifurcation adds extra dimension Bifurcation add values for all k Images Sean Eddy 46
47 Base Pair Maximization: Drawbacks Base pair maximization will not necessarily lead to the most stable structure. It may create structure with many interior loops or hairpins which are energetically unfavorable. Results comparable to aligning sequences with scattered matches not biologically reasonable. 47
48 Energy Minimization Thermodynamic Stability Estimated using experimental techniques. Theory : Most Stable = Most likely No pseudoknots due to algorithm limitations. Attempts to maximize the score, taking thermodynamics into account. MFOLD and ViennaRNA 48
49 49
50 50
51 51
52 52
53 Energy Minimization Results Linear RNA strand folded back on itself to create secondary structure Circularized representation uses this requirement Arcs represent base pairing Images David Mount 53
54 Energy Minimization Results All loops must have exactly three bases in them. Equivalent to having at least three base pairs between arc endpoints. Images David Mount 54
55 Energy Minimization Results All loops must have exactly three bases in them. Exception: Location where beginning and end of RNA come together in circularized representation. Images David Mount 55
56 Trouble with Pseudoknots Pseudoknots cause a breakdown in the dynamic programming algorithm. In order to form a pseudoknot, checks must be made to ensure base is not already paired this breaks down the recurrence relations. Images David Mount 56
57 Trouble with Pseudoknots Pseudoknots cause a breakdown in the dynamic programming algorithm. In order to form a pseudoknot, checks must be made to ensure base is not already paired this breaks down the recurrence relations. Images David Mount 57
58 Trouble with Pseudoknots Pseudoknots cause a breakdown in the dynamic programming algorithm. In order to form a pseudoknot, checks must be made to ensure base is not already paired this breaks down the recurrence relations. Images David Mount 58
59 Energy Minimization: Drawbacks Computes only one optimal structure. Optimal solution may not represent the biologically correct solution. 59
60 Alternative Algorithms - Covariaton Expect areas of base pairing in trna to be covarying between various species. Find this by sequence alignment. Base pairing creates same stable trna structure in organisms. 60
61 Sequence Alignment to Determine Structure Bases pair in order to form backbones and determine the secondary structure. Aligning bases based on their ability to pair with each other gives an algorithmic approach to determining the optimal structure. 61
62 62
63 63
64 Alternative Algorithms - Covariaton Expect areas of base pairing in trna to be covarying between various species. Base pairing creates same stable trna structure in organisms. Mutation in one base yields pairing impossible and breaks down structure. 64
65 Alternative Algorithms - Covariaton Expect areas of base pairing in trna to be covarying between various species. Base pairing creates same stable trna structure in organisms. Mutation in one base yields pairing impossible and breaks down structure. Covariation ensures ability to base pair is maintained and RNA structure is conserved. 65
66 66
67 67
68 Binary Tree Representation of RNA Secondary Structure Representation of RNA structure using Binary tree Nodes represent Base pair if two bases are shown Loop if base and gap (dash) are shown Pseudoknots still not represented Tree does not permit varying sequences Mismatches Insertions & Deletions Images Eddy et al. 68
69 Binary Tree Representation of RNA Secondary Structure Representation of RNA structure using Binary tree Nodes represent Base pair if two bases are shown Loop if base and gap (dash) are shown Pseudoknots still not represented Tree does not permit varying sequences Mismatches Insertions & Deletions Images Eddy et al. 69
70 Binary Tree Representation of RNA Secondary Structure Representation of RNA structure using Binary tree Nodes represent Base pair if two bases are shown Loop if base and gap (dash) are shown Pseudoknots still not represented Tree does not permit varying sequences Mismatches Insertions & Deletions Images Eddy et al. 70
71 Binary Tree Representation of RNA Secondary Structure Representation of RNA structure using Binary tree Nodes represent Base pair if two bases are shown Loop if base and gap (dash) are shown Pseudoknots still not represented Tree does not permit varying sequences Mismatches Insertions & Deletions Images Eddy et al. 71
72 Covariance Model Covariance Model: HMM which permits flexible alignment to an RNA structure emission and transition probabilities Model trees based on finite number of states Match states sequence conforms to the model: MATP: State in which bases are paired in the model and sequence. MATL & MATR: State in which either right or left bulges in the sequence and the model. Deletion State in which there is deletion in the sequence when compared to the model. Insertion State in which there is an insertion relative to model. 72
73 Covariance Model Covariance Model: HMM which permits flexible alignment to an RNA structure emission and transition probabilities Transitions have probabilities. Varying probability: Enter insertion, remain in current state, etc. Bifurcation: No probability, describes path. 73
74 Covariance Model (CM) Training Algorithm S(i, j) = Score at indices i and j in RNA when aligned to the Covariance Model. S i, j max S i 1, j 1 M i, j S i 1, j S i, j 1 max i k j S i, k S k 1, j Frequencies obtained by aligning model to training data consists of sample sequences. Reflect values which optimize alignment of sequences to model. 74
75 Covariance Model (CM) Training Algorithm S(i, j) = Score at indices i and j in RNA when aligned to the Covariance Model. S i 1, j 1 M i, j S i, j max S i 1, j S i, j 1 max i k j S i, k S k 1, j Frequency of seeing the symbols (A, C, G, U) together in locations i and j depending on symbol. Frequencies obtained by aligning model to training data consists of sample sequences. Reflect values which optimize alignment of sequences to model. 75
76 Covariance Model (CM) Training Algorithm S(i, j) = Score at indices i and j in RNA when aligned to the Covariance Model. S i 1, j 1 M i, j S i, j max S i 1, j S i, j 1 max i k j S i, k S k 1, j Independent frequency of seeing the symbols (A, C, G, T) in locations i or j depending on symbol. Frequencies obtained by aligning model to training data consists of sample sequences. Reflect values which optimize alignment of sequences to model. 76
77 Covariance Model (CM) Training Algorithm S(i, j) = Score at indices i and j in RNA when aligned to the Covariance Model. S i 1, j 1 M i, j S i, j max S i 1, j S i, j 1 max i k j S i, k S k 1, j Independent frequency of seeing the symbols (A, C, G, U) in locations i or j depending on symbol. Frequencies obtained by aligning model to training data consists of sample sequences. Reflect values which optimize alignment of sequences to model. 77
78 Alignment to CM Algorithm Calculate the probability score of aligning RNA to CM. Three dimensional matrix O(n³) Align sequence to given subtrees in CM. For each subsequence, calculate all possible states. Subtrees evolve from bifurcations For simplicity, left singlet is default. Images Eddy et al. 78
79 Alignment to CM Algorithm For each calculation, take into account: Transition (T) to next state. Emission probability (P) in the state as determined by training data. Images Eddy et al. 79
80 Alignment to CM Algorithm For each calculation, take into account: Transition (T) to next state. Emission probability (P) in the state as determined by training data. Images Eddy et al. 80
81 Alignment to CM Algorithm For each calculation, take into account: Transition (T) to next state. Emission probability (P) in the state as determined by training data. Images Eddy et al. 81
82 Alignment to CM Algorithm For each calculation, take into account: Transition (T) to next state. Emission probability (P) in the state as determined by training data. Deletion does not have emission probability associated with it. Images Eddy et al. 82
83 Alignment to CM Algorithm For each calculation, take into account: Transition (T) to next state. Emission probability (P) in the state as determined by training data. Deletion does not have emission probability associated with it. Bifurcation does not have state probability associated with it. Images Eddy et al. 83
84 Covariance Model Drawbacks Needs to be well trained. Not suitable for searches of large RNA. Structural complexity of large RNA cannot be modeled Runtime Memory requirements 84
RNA Secondary Structure Prediction Computational Genomics Seyoung Kim
 RNA Secondary Structure Prediction 02-710 Computational Genomics Seyoung Kim Outline RNA folding Dynamic programming for RNA secondary structure prediction Covariance model for RNA structure prediction
RNA Secondary Structure Prediction 02-710 Computational Genomics Seyoung Kim Outline RNA folding Dynamic programming for RNA secondary structure prediction Covariance model for RNA structure prediction
RNA Secondary Structure Prediction
 RNA Secondary Structure Prediction Outline 1) Introduction: RNA structure basics 2) Dynamic programming for RNA secondary structure prediction The Central Dogma of Molecular Biology DNA CCTGAGCCAACTATTGATGAA
RNA Secondary Structure Prediction Outline 1) Introduction: RNA structure basics 2) Dynamic programming for RNA secondary structure prediction The Central Dogma of Molecular Biology DNA CCTGAGCCAACTATTGATGAA
RNA folding & ncrna discovery
 I519 Introduction to Bioinformatics RNA folding & ncrna discovery Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Non-coding RNAs and their functions RNA structures RNA folding
I519 Introduction to Bioinformatics RNA folding & ncrna discovery Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Contents Non-coding RNAs and their functions RNA structures RNA folding
90 Algorithms in Bioinformatics I, WS 06, ZBIT, D. Huson, December 4, 2006
 90 Algorithms in Bioinformatics I, WS 06, ZBIT, D. Huson, December 4, 2006 8 RNA Secondary Structure Sources for this lecture: R. Durbin, S. Eddy, A. Krogh und G. Mitchison. Biological sequence analysis,
90 Algorithms in Bioinformatics I, WS 06, ZBIT, D. Huson, December 4, 2006 8 RNA Secondary Structure Sources for this lecture: R. Durbin, S. Eddy, A. Krogh und G. Mitchison. Biological sequence analysis,
RNA Structure Prediction. Algorithms in Bioinformatics. SIGCSE 2009 RNA Secondary Structure Prediction. Transfer RNA. RNA Structure Prediction
 Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri RNA Structure Prediction Secondary
Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri RNA Structure Prediction Secondary
RNA is a single strand molecule composed of subunits called nucleotides joined by phosphodiester bonds.
 The Versatility of RNA Primary structure of RNA RNA is a single strand molecule composed of subunits called nucleotides joined by phosphodiester bonds. Each nucleotide subunit is composed of a ribose sugar,
The Versatility of RNA Primary structure of RNA RNA is a single strand molecule composed of subunits called nucleotides joined by phosphodiester bonds. Each nucleotide subunit is composed of a ribose sugar,
Genes and How They Work. Chapter 15
 Genes and How They Work Chapter 15 The Nature of Genes They proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes The central
Genes and How They Work Chapter 15 The Nature of Genes They proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes The central
Computational analysis of non-coding RNA. Andrew Uzilov BME110 Tue, Nov 16, 2010
 Computational analysis of non-coding RNA Andrew Uzilov auzilov@ucsc.edu BME110 Tue, Nov 16, 2010 1 Corrected/updated talk slides are here: http://tinyurl.com/uzilovrna redirects to: http://users.soe.ucsc.edu/~auzilov/bme110/fall2010/
Computational analysis of non-coding RNA Andrew Uzilov auzilov@ucsc.edu BME110 Tue, Nov 16, 2010 1 Corrected/updated talk slides are here: http://tinyurl.com/uzilovrna redirects to: http://users.soe.ucsc.edu/~auzilov/bme110/fall2010/
There are four major types of introns. Group I introns, found in some rrna genes, are self-splicing: they can catalyze their own removal.
 1 2 Continuous genes - Intron: Many eukaryotic genes contain coding regions called exons and noncoding regions called intervening sequences or introns. The average human gene contains from eight to nine
1 2 Continuous genes - Intron: Many eukaryotic genes contain coding regions called exons and noncoding regions called intervening sequences or introns. The average human gene contains from eight to nine
RNA Genomics. BME 110: CompBio Tools Todd Lowe May 14, 2010
 RNA Genomics BME 110: CompBio Tools Todd Lowe May 14, 2010 Admin WebCT quiz on Tuesday cover reading, using Jalview & Pfam Homework #3 assigned today due next Friday (8 days) In Genomes, Two Types of Genes
RNA Genomics BME 110: CompBio Tools Todd Lowe May 14, 2010 Admin WebCT quiz on Tuesday cover reading, using Jalview & Pfam Homework #3 assigned today due next Friday (8 days) In Genomes, Two Types of Genes
Molecular biology WID Masters of Science in Tropical and Infectious Diseases-Transcription Lecture Series RNA I. Introduction and Background:
 Molecular biology WID 602 - Masters of Science in Tropical and Infectious Diseases-Transcription Lecture Series RNA I. Introduction and Background: DNA and RNA each consists of only four different nucleotides.
Molecular biology WID 602 - Masters of Science in Tropical and Infectious Diseases-Transcription Lecture Series RNA I. Introduction and Background: DNA and RNA each consists of only four different nucleotides.
RNA Genomics II. BME 110: CompBio Tools Todd Lowe & Andrew Uzilov May 17, 2011
 RNA Genomics II BME 110: CompBio Tools Todd Lowe & Andrew Uzilov May 17, 2011 1 TIME Why RNA? An evolutionary perspective The RNA World hypotheses: life arose as self-replicating non-coding RNA (ncrna)
RNA Genomics II BME 110: CompBio Tools Todd Lowe & Andrew Uzilov May 17, 2011 1 TIME Why RNA? An evolutionary perspective The RNA World hypotheses: life arose as self-replicating non-coding RNA (ncrna)
An Overview of Probabilistic Methods for RNA Secondary Structure Analysis. David W Richardson CSE527 Project Presentation 12/15/2004
 An Overview of Probabilistic Methods for RNA Secondary Structure Analysis David W Richardson CSE527 Project Presentation 12/15/2004 RNA - a quick review RNA s primary structure is sequence of nucleotides
An Overview of Probabilistic Methods for RNA Secondary Structure Analysis David W Richardson CSE527 Project Presentation 12/15/2004 RNA - a quick review RNA s primary structure is sequence of nucleotides
Themes: RNA and RNA Processing. Messenger RNA (mrna) What is a gene? RNA is very versatile! RNA-RNA interactions are very important!
 Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Themes: RNA is very versatile! RNA and RNA Processing Chapter 14 RNA-RNA interactions are very important! Prokaryotes and Eukaryotes have many important differences. Messenger RNA (mrna) Carries genetic
Feedback D. Incorrect! No, although this is a correct characteristic of RNA, this is not the best response to the questions.
 Biochemistry - Problem Drill 23: RNA No. 1 of 10 1. Which of the following statements best describes the structural highlights of RNA? (A) RNA can be single or double stranded. (B) G-C pairs have 3 hydrogen
Biochemistry - Problem Drill 23: RNA No. 1 of 10 1. Which of the following statements best describes the structural highlights of RNA? (A) RNA can be single or double stranded. (B) G-C pairs have 3 hydrogen
COMP598: Advanced Computational Biology Methods & Research
 COMP598: Advanced Computational Biology Methods & Research Introduction to RNA secondary structure prediction Jérôme Waldispühl School of Computer Science, McGill RNA world In prebiotic world, RNA thought
COMP598: Advanced Computational Biology Methods & Research Introduction to RNA secondary structure prediction Jérôme Waldispühl School of Computer Science, McGill RNA world In prebiotic world, RNA thought
CH 17 :From Gene to Protein
 CH 17 :From Gene to Protein Defining a gene gene gene Defining a gene is problematic because one gene can code for several protein products, some genes code only for RNA, two genes can overlap, and there
CH 17 :From Gene to Protein Defining a gene gene gene Defining a gene is problematic because one gene can code for several protein products, some genes code only for RNA, two genes can overlap, and there
8/21/2014. From Gene to Protein
 From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
From Gene to Protein Chapter 17 Objectives Describe the contributions made by Garrod, Beadle, and Tatum to our understanding of the relationship between genes and enzymes Briefly explain how information
Figure A summary of spontaneous alterations likely to require DNA repair.
 DNA Damage Figure 5-46. A summary of spontaneous alterations likely to require DNA repair. The sites on each nucleotide that are known to be modified by spontaneous oxidative damage (red arrows), hydrolytic
DNA Damage Figure 5-46. A summary of spontaneous alterations likely to require DNA repair. The sites on each nucleotide that are known to be modified by spontaneous oxidative damage (red arrows), hydrolytic
BIOCHEMISTRY REVIEW. Overview of Biomolecules. Chapter 13 Protein Synthesis
 BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 13 Protein Synthesis 2 3 4 5 6 7 8 9 10 Are You Getting It?? Which properties are characteristic of the normal genetic code? (multiple answers) a) A
BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 13 Protein Synthesis 2 3 4 5 6 7 8 9 10 Are You Getting It?? Which properties are characteristic of the normal genetic code? (multiple answers) a) A
Prediction of noncoding RNAs with RNAz
 Prediction of noncoding RNAs with RNAz John Dzmil, III Steve Griesmer Philip Murillo April 4, 2007 What is non-coding RNA (ncrna)? RNA molecules that are not translated into proteins Size range from 20
Prediction of noncoding RNAs with RNAz John Dzmil, III Steve Griesmer Philip Murillo April 4, 2007 What is non-coding RNA (ncrna)? RNA molecules that are not translated into proteins Size range from 20
The Nature of Genes. The Nature of Genes. The Nature of Genes. The Nature of Genes. The Nature of Genes. The Genetic Code. Genes and How They Work
 Genes and How They Work Chapter 15 Early ideas to explain how genes work came from studying human diseases. Archibald Garrod studied alkaptonuria, 1902 Garrod recognized that the disease is inherited via
Genes and How They Work Chapter 15 Early ideas to explain how genes work came from studying human diseases. Archibald Garrod studied alkaptonuria, 1902 Garrod recognized that the disease is inherited via
BIO 311C Spring Lecture 36 Wednesday 28 Apr.
 BIO 311C Spring 2010 1 Lecture 36 Wednesday 28 Apr. Synthesis of a Polypeptide Chain 5 direction of ribosome movement along the mrna 3 ribosome mrna NH 2 polypeptide chain direction of mrna movement through
BIO 311C Spring 2010 1 Lecture 36 Wednesday 28 Apr. Synthesis of a Polypeptide Chain 5 direction of ribosome movement along the mrna 3 ribosome mrna NH 2 polypeptide chain direction of mrna movement through
Gene Expression: Transcription, Translation, RNAs and the Genetic Code
 Lecture 28-29 Gene Expression: Transcription, Translation, RNAs and the Genetic Code Central dogma of molecular biology During transcription, the information in a DNA sequence (a gene) is copied into a
Lecture 28-29 Gene Expression: Transcription, Translation, RNAs and the Genetic Code Central dogma of molecular biology During transcription, the information in a DNA sequence (a gene) is copied into a
The Nature of Genes. The Nature of Genes. Genes and How They Work. Chapter 15/16
 Genes and How They Work Chapter 15/16 The Nature of Genes Beadle and Tatum proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes
Genes and How They Work Chapter 15/16 The Nature of Genes Beadle and Tatum proposed the one gene one enzyme hypothesis. Today we know this as the one gene one polypeptide hypothesis. 2 The Nature of Genes
RNA Structure and the Versatility of RNA. Mitesh Shrestha
 RNA Structure and the Versatility of RNA Mitesh Shrestha Ribonucleic Acid (RNA) Nitrogenous Bases (Adenine, Uracil, Guanine, Cytosine) Ribose Sugar Ribonucleic Acid (RNA) Phosphate Group RNA world Hypothesis
RNA Structure and the Versatility of RNA Mitesh Shrestha Ribonucleic Acid (RNA) Nitrogenous Bases (Adenine, Uracil, Guanine, Cytosine) Ribose Sugar Ribonucleic Acid (RNA) Phosphate Group RNA world Hypothesis
1/4/18 NUCLEIC ACIDS. Nucleic Acids. Nucleic Acids. ECS129 Instructor: Patrice Koehl
 NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
NUCLEIC ACIDS. ECS129 Instructor: Patrice Koehl
 NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
NUCLEIC ACIDS ECS129 Instructor: Patrice Koehl Nucleic Acids Nucleotides DNA Structure RNA Synthesis Function Secondary structure Tertiary interactions Wobble hypothesis DNA RNA Replication Transcription
BASIC MOLECULAR GENETIC MECHANISMS Introduction:
 BASIC MOLECULAR GENETIC MECHANISMS Introduction: nucleic acids. (1) contain the information for determining the amino acid sequence & the structure and function of proteins (1) part of the cellular structures:
BASIC MOLECULAR GENETIC MECHANISMS Introduction: nucleic acids. (1) contain the information for determining the amino acid sequence & the structure and function of proteins (1) part of the cellular structures:
Bio11 Announcements. Ch 21: DNA Biology and Technology. DNA Functions. DNA and RNA Structure. How do DNA and RNA differ? What are genes?
 Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
Bio11 Announcements TODAY Genetics (review) and quiz (CP #4) Structure and function of DNA Extra credit due today Next week in lab: Case study presentations Following week: Lab Quiz 2 Ch 21: DNA Biology
Chapter 17. From Gene to Protein
 Chapter 17 From Gene to Protein Overview: The Flow of Genetic Information The information content of DNA is in the form of specific sequences of nucleotides The DNA inherited by an organism leads to specific
Chapter 17 From Gene to Protein Overview: The Flow of Genetic Information The information content of DNA is in the form of specific sequences of nucleotides The DNA inherited by an organism leads to specific
BIOLOGY - CLUTCH CH.17 - GENE EXPRESSION.
 !! www.clutchprep.com CONCEPT: GENES Beadle and Tatum develop the one gene one enzyme hypothesis through their work with Neurospora (bread mold). This idea was later revised as the one gene one polypeptide
!! www.clutchprep.com CONCEPT: GENES Beadle and Tatum develop the one gene one enzyme hypothesis through their work with Neurospora (bread mold). This idea was later revised as the one gene one polypeptide
RNA World Hypothesis and RNA folding. By Lixin Dai
 RNA World Hypothesis and RNA folding By Lixin Dai Outline: RNA World Hypothesis RNA structure primary secondary tertiary Folding of RNA structure Problems with the folding Solutions RNA World Hypothesis:
RNA World Hypothesis and RNA folding By Lixin Dai Outline: RNA World Hypothesis RNA structure primary secondary tertiary Folding of RNA structure Problems with the folding Solutions RNA World Hypothesis:
Analyzed Fungi Neurospora crassa mutants. Mutants were UNABLE to grow without Arginine (an amino acid) Other biochemical experiments indicated:
 From Gene to Protein Beadle and Tatum Analyzed Fungi Neurospora crassa mutants Mutants were UNABLE to grow without Arginine (an amino acid) Other biochemical experiments indicated: Precursor Ornithine
From Gene to Protein Beadle and Tatum Analyzed Fungi Neurospora crassa mutants Mutants were UNABLE to grow without Arginine (an amino acid) Other biochemical experiments indicated: Precursor Ornithine
The Central Dogma. DNA makes RNA makes Proteins
 The Central Dogma DNA makes RNA makes Proteins TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION OF
The Central Dogma DNA makes RNA makes Proteins TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION OF
Ch Molecular Biology of the Gene
 Ch. 12 - Molecular Biology of the Gene AP BIOLOGY CHAPTER GUIDE 1. In the middle of the unraveling the mysteries of DNA, researchers knew that genetic material must be able to. It must be stable so it
Ch. 12 - Molecular Biology of the Gene AP BIOLOGY CHAPTER GUIDE 1. In the middle of the unraveling the mysteries of DNA, researchers knew that genetic material must be able to. It must be stable so it
Regulation of bacterial gene expression
 Regulation of bacterial gene expression Gene Expression Gene Expression: RNA and protein synthesis DNA ----------> RNA ----------> Protein transcription translation! DNA replication only occurs in cells
Regulation of bacterial gene expression Gene Expression Gene Expression: RNA and protein synthesis DNA ----------> RNA ----------> Protein transcription translation! DNA replication only occurs in cells
From DNA to Protein: Genotype to Phenotype
 12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
From RNA To Protein
 From RNA To Protein 22-11-2016 Introduction mrna Processing heterogeneous nuclear RNA (hnrna) RNA that comprises transcripts of nuclear genes made by RNA polymerase II; it has a wide size distribution
From RNA To Protein 22-11-2016 Introduction mrna Processing heterogeneous nuclear RNA (hnrna) RNA that comprises transcripts of nuclear genes made by RNA polymerase II; it has a wide size distribution
Chapter 12: Molecular Biology of the Gene
 Biology Textbook Notes Chapter 12: Molecular Biology of the Gene p. 214-219 The Genetic Material (12.1) - Genetic Material must: 1. Be able to store information that pertains to the development, structure,
Biology Textbook Notes Chapter 12: Molecular Biology of the Gene p. 214-219 The Genetic Material (12.1) - Genetic Material must: 1. Be able to store information that pertains to the development, structure,
A. Incorrect! This feature does help with it suitability as genetic material.
 College Biology - Problem Drill 08: Gene Structures and Functions No. 1 of 10 1. Which of the statements below is NOT true in explaining why DNA is a suitable genetic material? #01 (A) Its double helix
College Biology - Problem Drill 08: Gene Structures and Functions No. 1 of 10 1. Which of the statements below is NOT true in explaining why DNA is a suitable genetic material? #01 (A) Its double helix
RNA secondary structure
 Sequence Analysis '17 -- Lecture 17 RNA secondary structure Functions Representations Predictions Many slides courtesy of M. Zuker, RP Math When RNA secondary structure matters mrna --> protein protein
Sequence Analysis '17 -- Lecture 17 RNA secondary structure Functions Representations Predictions Many slides courtesy of M. Zuker, RP Math When RNA secondary structure matters mrna --> protein protein
DNA. translation. base pairing rules for DNA Replication. thymine. cytosine. amino acids. The building blocks of proteins are?
 2 strands, has the 5-carbon sugar deoxyribose, and has the nitrogen base Thymine. The actual process of assembling the proteins on the ribosome is called? DNA translation Adenine pairs with Thymine, Thymine
2 strands, has the 5-carbon sugar deoxyribose, and has the nitrogen base Thymine. The actual process of assembling the proteins on the ribosome is called? DNA translation Adenine pairs with Thymine, Thymine
Gene Expression: Transcription
 Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
Gene Expression: Transcription The majority of genes are expressed as the proteins they encode. The process occurs in two steps: Transcription = DNA RNA Translation = RNA protein Taken together, they make
Bioinformatics: Sequence Analysis. COMP 571 Luay Nakhleh, Rice University
 Bioinformatics: Sequence Analysis COMP 571 Luay Nakhleh, Rice University Course Information Instructor: Luay Nakhleh (nakhleh@rice.edu); office hours by appointment (office: DH 3119) TA: Leo Elworth (DH
Bioinformatics: Sequence Analysis COMP 571 Luay Nakhleh, Rice University Course Information Instructor: Luay Nakhleh (nakhleh@rice.edu); office hours by appointment (office: DH 3119) TA: Leo Elworth (DH
Transcription in Eukaryotes
 Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Transcription in Eukaryotes Biology I Hayder A Giha Transcription Transcription is a DNA-directed synthesis of RNA, which is the first step in gene expression. Gene expression, is transformation of the
Chapter 13. From DNA to Protein
 Chapter 13 From DNA to Protein Proteins All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequenceof a gene The Path From Genes to
Chapter 13 From DNA to Protein Proteins All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequenceof a gene The Path From Genes to
Transcription and Post Transcript Modification
 Transcription and Post Transcript Modification You Should Be Able To 1. Describe transcription. 2. Compare and contrast eukaryotic + prokaryotic transcription. 3. Explain mrna processing in eukaryotes.
Transcription and Post Transcript Modification You Should Be Able To 1. Describe transcription. 2. Compare and contrast eukaryotic + prokaryotic transcription. 3. Explain mrna processing in eukaryotes.
Transcription is the first stage of gene expression
 Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
Transcription is the first stage of gene expression RNA synthesis is catalyzed by RNA polymerase, which pries the DNA strands apart and hooks together the RNA nucleotides The RNA is complementary to the
PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein
 PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein This is also known as: The central dogma of molecular biology Protein Proteins are made
PROTEIN SYNTHESIS Flow of Genetic Information The flow of genetic information can be symbolized as: DNA RNA Protein This is also known as: The central dogma of molecular biology Protein Proteins are made
Lecture Overview. Overview of the Genetic Information. Marieb s Human Anatomy and Physiology. Chapter 3 DNA & RNA Protein Synthesis Lecture 6
 Marieb s Human Anatomy and Physiology Marieb Hoehn Chapter 3 DNA & RNA Protein Synthesis Lecture 6 Lecture Overview The Genetic Information Structure of DNA/RNA DNA Replication Overview of protein synthesis
Marieb s Human Anatomy and Physiology Marieb Hoehn Chapter 3 DNA & RNA Protein Synthesis Lecture 6 Lecture Overview The Genetic Information Structure of DNA/RNA DNA Replication Overview of protein synthesis
GENETICS - CLUTCH CH.10 TRANSCRIPTION.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF TRANSCRIPTION Transcription is the process of using DNA as a template to RNA RNA polymerase is the enzyme that transcribes DNA - There are many different types
!! www.clutchprep.com CONCEPT: OVERVIEW OF TRANSCRIPTION Transcription is the process of using DNA as a template to RNA RNA polymerase is the enzyme that transcribes DNA - There are many different types
Chapter 14: From DNA to Protein
 Chapter 14: From DNA to Protein Steps from DNA to Proteins Same two steps produce all proteins: 1) DNA is transcribed to form RNA Occurs in the nucleus RNA moves into cytoplasm 2) RNA is translated in
Chapter 14: From DNA to Protein Steps from DNA to Proteins Same two steps produce all proteins: 1) DNA is transcribed to form RNA Occurs in the nucleus RNA moves into cytoplasm 2) RNA is translated in
RNA Structure Prediction and Comparison. RNA Biology Background
 RN Structure Prediction and omparison Session 1 RN Biology Background Robert iegerich Faculty of Technology robert@techfak.ni-bielefeld.de October 13, 2013 Robert iegerich Overview of lecture topics The
RN Structure Prediction and omparison Session 1 RN Biology Background Robert iegerich Faculty of Technology robert@techfak.ni-bielefeld.de October 13, 2013 Robert iegerich Overview of lecture topics The
Control of Eukaryotic Genes. AP Biology
 Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
Control of Eukaryotic Genes The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? Evolution
From DNA to Protein: Genotype to Phenotype
 12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
12 From DNA to Protein: Genotype to Phenotype 12.1 What Is the Evidence that Genes Code for Proteins? The gene-enzyme relationship is one-gene, one-polypeptide relationship. Example: In hemoglobin, each
RNA Structure Prediction and Comparison. RNA Biology Background
 RN Structure Prediction and omparison Session 1 RN Biology Background édric Saule Faculty of Technology cedric.saule@ni-bielefeld.de pril 13, 2015 édric Saule Overview of lecture topics The lecture plan
RN Structure Prediction and omparison Session 1 RN Biology Background édric Saule Faculty of Technology cedric.saule@ni-bielefeld.de pril 13, 2015 édric Saule Overview of lecture topics The lecture plan
Lecture Summary: Regulation of transcription. General mechanisms-what are the major regulatory points?
 BCH 401G Lecture 37 Andres Lecture Summary: Regulation of transcription. General mechanisms-what are the major regulatory points? RNA processing: Capping, polyadenylation, splicing. Why process mammalian
BCH 401G Lecture 37 Andres Lecture Summary: Regulation of transcription. General mechanisms-what are the major regulatory points? RNA processing: Capping, polyadenylation, splicing. Why process mammalian
Fermentation. Lesson Overview. Lesson Overview 13.1 RNA
 13.1 RNA THINK ABOUT IT DNA is the genetic material of cells. The sequence of nucleotide bases in the strands of DNA carries some sort of code. In order for that code to work, the cell must be able to
13.1 RNA THINK ABOUT IT DNA is the genetic material of cells. The sequence of nucleotide bases in the strands of DNA carries some sort of code. In order for that code to work, the cell must be able to
Algorithmic Approaches to Modelling and Predicting RNA Structure
 Algorithmic Approaches to Modelling and Predicting RNA Structure Literature Review Carlos Gonzalez Oliver Supervisor: Jérôme Waldispühl August 27, 2017 Abstract Ribonucleic acid (RNA) is a chain-like molecule
Algorithmic Approaches to Modelling and Predicting RNA Structure Literature Review Carlos Gonzalez Oliver Supervisor: Jérôme Waldispühl August 27, 2017 Abstract Ribonucleic acid (RNA) is a chain-like molecule
COMP364. Introduction to RNA secondary structure prediction. Jérôme Waldispühl School of Computer Science, McGill
 COMP364 Introduction to RNA secondary structure prediction Jérôme Waldispühl School of Computer Science, McGill RNA world In prebiotic world, RNA thought to have filled two distinct roles: 1. an information
COMP364 Introduction to RNA secondary structure prediction Jérôme Waldispühl School of Computer Science, McGill RNA world In prebiotic world, RNA thought to have filled two distinct roles: 1. an information
Structural Bioinformatics GENOME 541 Spring 2018
 Molecular composition of a rapidly dividing Escherichia coli cell Structural Bioinformatics GENOME 541 Spring 2018 Lecture 4: Nucleic Acids Frank DiMaio (dimaio@uw.edu) The major biopolymers DNA structure
Molecular composition of a rapidly dividing Escherichia coli cell Structural Bioinformatics GENOME 541 Spring 2018 Lecture 4: Nucleic Acids Frank DiMaio (dimaio@uw.edu) The major biopolymers DNA structure
RNA : functional role
 RNA : functional role Hamad Yaseen, PhD MLS Department, FAHS Hamad.ali@hsc.edu.kw RNA mrna rrna trna 1 From DNA to Protein -Outline- From DNA to RNA From RNA to Protein From DNA to RNA Transcription: Copying
RNA : functional role Hamad Yaseen, PhD MLS Department, FAHS Hamad.ali@hsc.edu.kw RNA mrna rrna trna 1 From DNA to Protein -Outline- From DNA to RNA From RNA to Protein From DNA to RNA Transcription: Copying
RNA does not adopt the classic B-DNA helix conformation when it forms a self-complementary double helix
 Reason: RNA has ribose sugar ring, with a hydroxyl group (OH) If RNA in B-from conformation there would be unfavorable steric contact between the hydroxyl group, base, and phosphate backbone. RNA structure
Reason: RNA has ribose sugar ring, with a hydroxyl group (OH) If RNA in B-from conformation there would be unfavorable steric contact between the hydroxyl group, base, and phosphate backbone. RNA structure
Lesson 8. DNA: The Molecule of Heredity. Gene Expression and Regulation. Introduction to Life Processes - SCI 102 1
 Lesson 8 DNA: The Molecule of Heredity Gene Expression and Regulation Introduction to Life Processes - SCI 102 1 Genes and DNA Hereditary information is found in discrete units called genes Genes are segments
Lesson 8 DNA: The Molecule of Heredity Gene Expression and Regulation Introduction to Life Processes - SCI 102 1 Genes and DNA Hereditary information is found in discrete units called genes Genes are segments
Lecture for Wednesday. Dr. Prince BIOL 1408
 Lecture for Wednesday Dr. Prince BIOL 1408 THE FLOW OF GENETIC INFORMATION FROM DNA TO RNA TO PROTEIN Copyright 2009 Pearson Education, Inc. Genes are expressed as proteins A gene is a segment of DNA that
Lecture for Wednesday Dr. Prince BIOL 1408 THE FLOW OF GENETIC INFORMATION FROM DNA TO RNA TO PROTEIN Copyright 2009 Pearson Education, Inc. Genes are expressed as proteins A gene is a segment of DNA that
Chapter 8 From DNA to Proteins. Chapter 8 From DNA to Proteins
 KEY CONCEPT Section 1 DNA was identified as the genetic material through a series of experiments. Griffith finds a transforming principle. Griffith experimented with the bacteria that cause pneumonia.
KEY CONCEPT Section 1 DNA was identified as the genetic material through a series of experiments. Griffith finds a transforming principle. Griffith experimented with the bacteria that cause pneumonia.
DNA. Is a molecule that encodes the genetic instructions used in the development and functioning of all known living organisms and many viruses.
 Is a molecule that encodes the genetic instructions used in the development and functioning of all known living organisms and many viruses. Genetic information is encoded as a sequence of nucleotides (guanine,
Is a molecule that encodes the genetic instructions used in the development and functioning of all known living organisms and many viruses. Genetic information is encoded as a sequence of nucleotides (guanine,
produces an RNA copy of the coding region of a gene
 1. Transcription Gene Expression The expression of a gene into a protein occurs by: 1) Transcription of a gene into RNA produces an RNA copy of the coding region of a gene the RNA transcript may be the
1. Transcription Gene Expression The expression of a gene into a protein occurs by: 1) Transcription of a gene into RNA produces an RNA copy of the coding region of a gene the RNA transcript may be the
DNA Transcription. Dr Aliwaini
 DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
DNA Transcription 1 DNA Transcription-Introduction The synthesis of an RNA molecule from DNA is called Transcription. All eukaryotic cells have five major classes of RNA: ribosomal RNA (rrna), messenger
Chapter 31. Transcription and RNA processing
 Chapter 31 Transcription and RNA processing RNA polymerase (RNAP) E. coli promoters Components of E. coli RNA Polymerase Holoenzyme (α 2 ββ'ωσ) Structure of prokaryotic RNAP The closed and open state of
Chapter 31 Transcription and RNA processing RNA polymerase (RNAP) E. coli promoters Components of E. coli RNA Polymerase Holoenzyme (α 2 ββ'ωσ) Structure of prokaryotic RNAP The closed and open state of
Nucleic Acids: DNA and RNA
 Nucleic Acids: DNA and RNA Living organisms are complex systems. Hundreds of thousands of proteins exist inside each one of us to help carry out our daily functions. These proteins are produced locally,
Nucleic Acids: DNA and RNA Living organisms are complex systems. Hundreds of thousands of proteins exist inside each one of us to help carry out our daily functions. These proteins are produced locally,
TERTIARY MOTIF INTERACTIONS ON RNA STRUCTURE
 1 TERTIARY MOTIF INTERACTIONS ON RNA STRUCTURE Bioinformatics Senior Project Wasay Hussain Spring 2009 Overview of RNA 2 The central Dogma of Molecular biology is DNA RNA Proteins The RNA (Ribonucleic
1 TERTIARY MOTIF INTERACTIONS ON RNA STRUCTURE Bioinformatics Senior Project Wasay Hussain Spring 2009 Overview of RNA 2 The central Dogma of Molecular biology is DNA RNA Proteins The RNA (Ribonucleic
Exam 2 Key - Spring 2008 A#: Please see us if you have any questions!
 Page 1 of 5 Exam 2 Key - Spring 2008 A#: Please see us if you have any questions! 1. A mutation in which parts of two nonhomologous chromosomes change places is called a(n) A. translocation. B. transition.
Page 1 of 5 Exam 2 Key - Spring 2008 A#: Please see us if you have any questions! 1. A mutation in which parts of two nonhomologous chromosomes change places is called a(n) A. translocation. B. transition.
RNA GENE DISCOVERY THROUGH THE CLASSIFICATION OF RNA SECONDARY STRUCTURE ELEMENTS
 RNA GENE DISCOVERY THROUGH THE CLASSIFICATION OF RNA SECONDARY STRUCTURE ELEMENTS by Nicholas Erho B. Sc., Computing Science, Trinity Western University, 2007 a Thesis submitted in partial fulfillment
RNA GENE DISCOVERY THROUGH THE CLASSIFICATION OF RNA SECONDARY STRUCTURE ELEMENTS by Nicholas Erho B. Sc., Computing Science, Trinity Western University, 2007 a Thesis submitted in partial fulfillment
Control of Eukaryotic Gene Expression (Learning Objectives)
 Control of Eukaryotic Gene Expression (Learning Objectives) 1. Compare and contrast chromatin and chromosome: composition, proteins involved and level of packing. Explain the structure and function of
Control of Eukaryotic Gene Expression (Learning Objectives) 1. Compare and contrast chromatin and chromosome: composition, proteins involved and level of packing. Explain the structure and function of
Sequence Analysis. II: Sequence Patterns and Matrices. George Bell, Ph.D. WIBR Bioinformatics and Research Computing
 Sequence Analysis II: Sequence Patterns and Matrices George Bell, Ph.D. WIBR Bioinformatics and Research Computing Sequence Patterns and Matrices Multiple sequence alignments Sequence patterns Sequence
Sequence Analysis II: Sequence Patterns and Matrices George Bell, Ph.D. WIBR Bioinformatics and Research Computing Sequence Patterns and Matrices Multiple sequence alignments Sequence patterns Sequence
Protein Synthesis ~Biology AP~
 Protein Synthesis ~Biology AP~ A Meridian Study Guide by David Guan, Jennifer Zheng [Edited by Lei Gong] Introduction: - DNA and RNA are essential for life because they code for enzymes, which regulate
Protein Synthesis ~Biology AP~ A Meridian Study Guide by David Guan, Jennifer Zheng [Edited by Lei Gong] Introduction: - DNA and RNA are essential for life because they code for enzymes, which regulate
Chapters 31-32: Ribonucleic Acid (RNA)
 Chapters 31-32: Ribonucleic Acid (RNA) Short segments from the transcription, processing and translation sections of each chapter Slide 1 RNA In comparison with DNA RNA utilizes uracil in place of thymine
Chapters 31-32: Ribonucleic Acid (RNA) Short segments from the transcription, processing and translation sections of each chapter Slide 1 RNA In comparison with DNA RNA utilizes uracil in place of thymine
Key Area 1.3: Gene Expression
 Key Area 1.3: Gene Expression RNA There is a second type of nucleic acid in the cell, called RNA. RNA plays a vital role in the production of protein from the code in the DNA. What is gene expression?
Key Area 1.3: Gene Expression RNA There is a second type of nucleic acid in the cell, called RNA. RNA plays a vital role in the production of protein from the code in the DNA. What is gene expression?
From Gene to Protein. Chapter 17
 From Gene to Protein Chapter 17 What you need to know: The key terms: gene expression, transcription, and translation. The major events of transcription. How eukaryotic cells modify RNA after transcription.
From Gene to Protein Chapter 17 What you need to know: The key terms: gene expression, transcription, and translation. The major events of transcription. How eukaryotic cells modify RNA after transcription.
Chapter 17. From Gene to Protein. Slide 1. Slide 2. Slide 3. Gene Expression. Which of the following is the best example of gene expression? Why?
 Slide 1 Chapter 17 From Gene to Protein PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Slide 1 Chapter 17 From Gene to Protein PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Lecture Overview. Overview of the Genetic Information. Chapter 3 DNA & RNA Lecture 6
 Visual Anatomy & Physiology First Edition Martini & Ober Chapter 3 DNA & RNA Lecture 6 Lecture Overview What is the cell s genetic information? How/where is the genetic information stored in eukaryotic
Visual Anatomy & Physiology First Edition Martini & Ober Chapter 3 DNA & RNA Lecture 6 Lecture Overview What is the cell s genetic information? How/where is the genetic information stored in eukaryotic
BIOCHEMISTRY REVIEW. Overview of Biomolecules. Chapter 12 Transcription
 BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 12 Transcription 2 3 4 5 Are You Getting It?? Which are general characteristics of transcription? (multiple answers) a) An entire DNA molecule is transcribed
BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 12 Transcription 2 3 4 5 Are You Getting It?? Which are general characteristics of transcription? (multiple answers) a) An entire DNA molecule is transcribed
Molecular Cell Biology - Problem Drill 08: Transcription, Translation and the Genetic Code
 Molecular Cell Biology - Problem Drill 08: Transcription, Translation and the Genetic Code Question No. 1 of 10 1. Which of the following statements about how genes function is correct? Question #1 (A)
Molecular Cell Biology - Problem Drill 08: Transcription, Translation and the Genetic Code Question No. 1 of 10 1. Which of the following statements about how genes function is correct? Question #1 (A)
Protein Synthesis: From Gene RNA Protein Trait
 Protein Synthesis: From Gene RNA Protein Trait Human Genome The human genome contains about genes. Each gene is a of DNA (sequence of nitrogen bases) contained within each chromosome. Each chromosome contains
Protein Synthesis: From Gene RNA Protein Trait Human Genome The human genome contains about genes. Each gene is a of DNA (sequence of nitrogen bases) contained within each chromosome. Each chromosome contains
Chapter 17 From Gene to Protein
 Chapter 17 From Gene to Protein The Flow of Genetic Information The information content of DNA is in the form of specific sequences of nucleotides The DNA inherited by an organism leads to specific traits
Chapter 17 From Gene to Protein The Flow of Genetic Information The information content of DNA is in the form of specific sequences of nucleotides The DNA inherited by an organism leads to specific traits
Chapter 8 Lecture Outline. Transcription, Translation, and Bioinformatics
 Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
Chapter 8 Lecture Outline Transcription, Translation, and Bioinformatics Replication, Transcription, Translation n Repetitive processes Build polymers of nucleotides or amino acids n All have 3 major steps
I. Gene Expression Figure 1: Central Dogma of Molecular Biology
 I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
I. Gene Expression Figure 1: Central Dogma of Molecular Biology Central Dogma: Gene Expression: RNA Structure RNA nucleotides contain the pentose sugar Ribose instead of deoxyribose. Contain the bases
BIOLOGICAL SCIENCE. Lecture Presentation by Cindy S. Malone, PhD, California State University Northridge. FIFTH EDITION Freeman Quillin Allison
 BIOLOGICAL SCIENCE FIFTH EDITION Freeman Quillin Allison 4 Lecture Presentation by Cindy S. Malone, PhD, California State University Northridge In this chapter you will learn that Nucleic acids store the
BIOLOGICAL SCIENCE FIFTH EDITION Freeman Quillin Allison 4 Lecture Presentation by Cindy S. Malone, PhD, California State University Northridge In this chapter you will learn that Nucleic acids store the
CSE : Computational Issues in Molecular Biology. Lecture 19. Spring 2004
 CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
CSE 397-497: Computational Issues in Molecular Biology Lecture 19 Spring 2004-1- Protein structure Primary structure of protein is determined by number and order of amino acids within polypeptide chain.
EUKARYOTIC GENE CONTROL
 EUKARYOTIC GENE CONTROL THE BIG QUESTIONS How are genes turned on and off? How do cells with the same DNA/ genes differentiate to perform completely different and specialized functions? GENE EXPRESSION
EUKARYOTIC GENE CONTROL THE BIG QUESTIONS How are genes turned on and off? How do cells with the same DNA/ genes differentiate to perform completely different and specialized functions? GENE EXPRESSION
Nucleic Acids. OpenStax College. 1 DNA and RNA
 OpenStax-CNX module: m44403 1 Nucleic Acids OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 4.0 By the end of this section, you will be
OpenStax-CNX module: m44403 1 Nucleic Acids OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 4.0 By the end of this section, you will be
From Gene to Protein
 Chapter 17 From Gene to Protein PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp
Chapter 17 From Gene to Protein PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp
Transcription steps. Transcription steps. Eukaryote RNA processing
 Transcription steps Initiation at 5 end of gene binding of RNA polymerase to promoter unwinding of DNA Elongation addition of nucleotides to 3 end rules of base pairing requires Mg 2+ energy from NTP substrates
Transcription steps Initiation at 5 end of gene binding of RNA polymerase to promoter unwinding of DNA Elongation addition of nucleotides to 3 end rules of base pairing requires Mg 2+ energy from NTP substrates
DNA REPLICATION. DNA structure. Semiconservative replication. DNA structure. Origin of replication. Replication bubbles and forks.
 DNA REPLICATION 5 4 Phosphate 3 DNA structure Nitrogenous base 1 Deoxyribose 2 Nucleotide DNA strand = DNA polynucleotide 2004 Biology Olympiad Preparation Program 2 2004 Biology Olympiad Preparation Program
DNA REPLICATION 5 4 Phosphate 3 DNA structure Nitrogenous base 1 Deoxyribose 2 Nucleotide DNA strand = DNA polynucleotide 2004 Biology Olympiad Preparation Program 2 2004 Biology Olympiad Preparation Program
DNA and RNA Structure Guided Notes
 Nucleic acids, especially DNA, are considered as the key biomolecules that guarantee the continuity of life. DNA is the prime genetic molecule which carry all the hereditary information that's passed from
Nucleic acids, especially DNA, are considered as the key biomolecules that guarantee the continuity of life. DNA is the prime genetic molecule which carry all the hereditary information that's passed from
DNA makes RNA makes Proteins. The Central Dogma
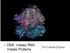 DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
DNA makes RNA makes Proteins The Central Dogma TRANSCRIPTION DNA RNA transcript RNA polymerase RNA PROCESSING Exon RNA transcript (pre-mrna) Intron Aminoacyl-tRNA synthetase NUCLEUS CYTOPLASM FORMATION
DNA Structure and Replication, and Virus Structure and Replication Test Review
 DNA Structure and Replication, and Virus Structure and Replication Test Review What does DNA stand for? Deoxyribonucleic Acid DNA is what type of macromolecule? DNA is a nucleic acid The building blocks
DNA Structure and Replication, and Virus Structure and Replication Test Review What does DNA stand for? Deoxyribonucleic Acid DNA is what type of macromolecule? DNA is a nucleic acid The building blocks
CLEP Biology - Problem Drill 11: Transcription, Translation and The Genetic Code
 CLEP Biology - Problem Drill 11: Transcription, Translation and The Genetic Code No. 1 of 10 1. Three types of RNA comprise the structural and functional core for protein synthesis, serving as a template
CLEP Biology - Problem Drill 11: Transcription, Translation and The Genetic Code No. 1 of 10 1. Three types of RNA comprise the structural and functional core for protein synthesis, serving as a template
