Supplemental Figure 1
|
|
|
- Roger Gardner
- 5 years ago
- Views:
Transcription
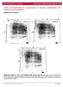
1 Supplemental Figure 1 Supplemental Fig. 1. (A) The binding avidity between Fz4-FcH6 and MBP-Norrin dimers measured by biolayer interferometry. 5 µg/ml Fz4-Fc protein was loaded onto anti human Fc capture sensors, incubated with kinetic buffer, and then incubated with MBP- Norrin protein (80 nm and 32 nm) in kinetic buffer for the association phase, followed with kinetic buffer for the dissociation phase. A K d value of 4.6±1.1 nm was calculated using the associated kinetic analysis program (version 6.4). (B) Stereo view of the Norrin monomeric structure. Shown are the main chain atoms in stick models with carbon atoms in green, oxygen atoms in red, and nitrogen atoms in blue. The 2F o -F c map contoured at 1.0 σ is also shown.
2 Supplemental Figure 2
3 Supplemental Fig. 2. Crystal structure of MBP-Norrin. (A) MBP-Norrin crystal packing. Four MBP-Norrin dimers are shown in yellow, magenta, blue and grey, respectively. The unit cell box is shown in green. The majority of the interactions for crystal packing are due to MBP MBP interactions (site 1) and MBP Norrin interactions (sites 2 and 3). The figures were generated using the Pymol program. (B) Structure of the MBP-Norrin dimer. On top are two MBP molecules (green and yellow) and on the bottom are two Norrin monomers (cyan and orange). One MBP-Norrin molecule (green+cyan) is related to another molecule (yellow+orange) by a crystallographic twofold symmetry.
4 Supplemental Figure 3
5 Supplemental Fig. 3. Structural comparison of Norrin with TGF-β3. (A) The monomeric structures of Norrin and TGF-β3 (PDB code: 1TGJ) are shown in cartoon representation with a rainbow color scheme from N-terminus (blue) to C-terminus (red). The disulfide bonds including the conserved cystine knot structure (a cluster of three intramolecular disulfide bonds) are shown as stick models. The significant structural differences between them are indicated in four boxes (Boxes 1-4). (B) Comparison of the dimeric structures of Norrin with TGF-β3. Two monomers in cartoon representation are colored in green and yellow. The intermolecular disulfide bonds are shown as stick models. (C) A display of the missense Norrie disease mutations onto the Norrin dimeric structure. A Norrin dimer is shown as a Cα backbone ribbon diagram with two monomer colored in green and magenta, respectively. The residues whose mutations cause Norrie disease or related diseases are shown as spheres on the green monomer. The green spheres are cysteine residues; yellow spheres are residues involved in forming the dimer interface; blue spheres are residues involved in Fz4 binding; red spheres are residues involved in Lrp5/6 binding; grey spheres are residues involved in hydrophobic packing. The disulfide bonds are shown as stick models.
6 Supplemental Figure 4
7 Supplemental Fig. 4. The dimer structure is important for Norrin function. (A) Norrin-mediated activation of the downstream TCF luciferase reporter activity is dependent on Fz4 expression. 293STF cells were transfected with Lrp5, Lrp6, Fz4 alone or Fz4+Lrp5, Fz4+Lrp6 in the presence or absence of Norrin wild type (WT) expression vector along with TKRlu (an internal control) on day 1. Cell culture media were changed on day 2. Cells were harvested and luciferase activities were measured on day 3 using the Dual luciferase kit (Promega). TCF mediated firefly luciferase activity was normalized to renilla luciferase activity. The value for Norrin+Fz4+Lrp5 transfected cells is set to 100% and the values for the other groups are adjusted accordingly. Error bars indicate SD (n=3). (B, C) Effects of Norrin cysteine mutations and hydrophobic residue mutations at the dimer interface on Norrin function. (B) 293STF cells were transfected with Fz4 or Fz4+Lrp5 in the presence or absence of Norrin WT or mutants along with TKRlu and luciferase activities were measured similarly as described in A. Error bars indicate SD (n=3). (C) 293STF cells were transfected with Fz4+ Lrp6 in the presence or absence of Norrin WT or mutants along with TKRlu and luciferase activities were measured similarly as described in A. Error bars indicate SD (n=3).
8 Supplemental Figure 5
9 Supplemental Fig. 5. Control experiments to examine Fz4 cell surface expression, Fz4 protein expression and Fz4-YFP fluorescence. (A) Fz4 surface expression for HEK293 cells transfected with Fz4 receptor with or without different tags. HEK293 cells were transfected with Fz4 receptor with or without a v5, YFP, YFP-N, YFP-C or Rlu tag at the C-terminus for 2 d. Cells were harvested by Cell Dissociation Solution (Sigma), and stained with a biotinylated Fz4 antibody (R&D systems) on ice for 30 min. Then cells were incubated with Streptavidin-Allophycocyanin (APC) (BD Biosciences) on ice for 30 min. Cells were washed with FACS buffer (1xPBS, 2%FBS, 1% Azide) between different steps and analyzed by flow cytometry using MoFlo Astrios (Beckman Coulter, Inc.). (B) Coexpression with Norrin or treatment with MBP-Norrin protein did not change overall protein expression for Fz4 receptor. HEK293 cells were transfected with Fz4±Norrin on day 1. HEK293 cells transfected with Fz4 were treated with or without 5 µg/ml MBP-Norrin on day 2 and harvested on day 3 for Western blot analysis. The blot was first probed with an anti-v5 antibody (Thermo Scientific), and then stripped and reprobed for β-actin as a loading control. (C) The cell surface YFP fluorescence for HEK293 cells transfected with Fz4YFP±Norrin DNAs. The surface YFP fluorescence is used as an indication for Fz4 surface expression. (D) The cell surface YFP fluorescence for cells transfected with Fz4-YFP, Fz4-YFP-N, Fz4-YFP-C or Fz4-YFP-N+Fz4-YFP-C. COS-1 cells were transfected with either intact YFP construct or the complementary YFP-N and YFP-C constructs.
10 Supplemental Figure 6
11 Supplemental Fig. 6. Both BP1 and BP2 domains of Lrp6 can bind to Norrin. (A) MBP-Norrin competes the interaction between biotinylated DKK1 peptide and His8- tagged BP1-2 protein. The AlphaScreen binding data were plotted with increasing concentrations of MBP-Norrin protein. The first two columns are negative controls. The binding signal of biotinylated DKK1 peptide and His8-tagged Lrp6 BP1-2 protein in the absence of MBP-Norrin protein was set to 100%. Error bars indicate SD (n=3). (B, C) Lrp6 BP2-Fc fusion protein binds to Norrin. (B) Binding of Lrp6 BP2-Fc protein to MBP-Norrin was determined by biolayer interferometry. Protein A sensors were loaded with 5 µg/ml Fz4-Fc, Lrp6 BP2-Fc, or human IgG and then incubated with buffer. The loaded sensors were incubated sequentially with 20 µg/ml MBP-Norrin in the association step and with buffer in the dissociation step. The interaction between Fz4-Fc and MBP- Norrin was used as a positive control. (C) The binding of MBP-Norrin to Lrp6 BP2-Fc measured by AlphaScreen assay. 50 nm biotinylated MBP-Norrin was mixed with 50 nm His6 tagged Lrp6-BP2-Fc protein at room temperature for 4 h. The reaction mixtures also contained 10 µg/ml of the streptavidin-coated donor beads and Ni-chelate-coated acceptor beads. The data represent the average of triplicate samples.
12 Supplemental Figure 7
13 Supplemental Fig. 7. Norrin binds to Fz4 and Lrp6 extracellular domains to promote their interaction. (A) Norrin forms a ternary complex with Fz4-CRD and Lrp6 BP1-2 domains measured by biolayer interferometry. Protein A sensors were incubated sequentially with 5 µg/ml Fz4-FcH6 (step1, loading), kinetic buffer (step2, baseline), 10 µg/ml MBP-Norrin (step3, association 1). The bound sensors were further incubated sequentially in kinetic buffer (step4, baseline), different Lrp6 ECD proteins (40 µg/ml Lrp6BP1-2, 40 µg/ml Lrp6BP3-4 and 80 µg/ml Lrp6BP1-4) (step5, association 2), and kinetic buffer (step6, dissociation). All the curves are aligned to the beginning point of Step 5 using the OctetRed analysis program (version 6.4). (B) Fz4-FcH6 protein does not directly bind Lrp6 β-propeller domains. Protein A sensors were loaded with 5 µg/ml Fz4-FcH6 protein and then incubated with kinetic buffer. The loaded sensors were then incubated sequentially with the indicated proteins (20 µg/ml MBP-Norrin, 70 µg/ml Lrp6 BP1-2, 70 µg/ml BP3-4, or 140 µg/ml BP1-4) in the association step and with kinetic buffer in the dissociation step. MBP-Norrin protein was used as a positive control.
14 Supplemental Figure 8
15 Supplemental Fig. 8. The sequence alignment of Norrin proteins from different species and with other growth factors. (A) Norrin sequences from different species were aligned by using the ClustalW program. All the DNA sequences were obtained from the NCBI protein database. The identical residues are highlighted in yellow whereas the conserved residues are highlighted in light yellow. The secondary structures based on human Norrin structure are noted above the sequence alignment. The black stars denote the cysteines forming intramolecular disulfide bonds whereas the red stars denote the cysteines forming intermolecular disulfide bonds. The filled triangles denote the residues for Fz4 binding whereas the filled circles denote residues for Lrp5/6 binding. (B) Sequence alignment of human Norrin, von Willebrand factor (vwf), mucin-2, TGF-β and pig submaxillary mucin proteins. The initial alignment was made by using ClustalW program and improved by small manual adjustments. All the DNA sequences were obtained from the NCBI protein database. The highly conserved residues are highlighted in yellow whereas the less conserved residues are highlighted in light yellow. The black stars denote these cysteines forming the cystine knot structure, whereas the red stars denote the cysteines forming intermolecular disulfide bonds. The green stars denote the two cysteines forming the TGF-β specific intramolecular disulfide bond, whereas the blue stars denote the two cysteines forming the Norrin specific intramolecular disulfide bond, which are also conserved in vwf and mucin proteins. The intramolecular disulfide bond pattern for Norrin is shown with black lines. The alignment figures were prepared by using the Aline program.
16 Supplemental Table 1. IC 50 values obtained by using purified MBP-Norrin wild type and mutant proteins to compete the Fz4-Fc-H6 and biotin-mbp-norrin interaction or H8- Lrp6 BP1-2 and biotin DKK1 peptide interaction. IC 50 (nm) Competition with Fz4-CRD interaction Lrp6 BP1-2 interaction Wild type R41E NC 288 K54E/R109E 18 NC C55A/C110A NC stands for no competition.
17 Supplemental Table 2. Functional categorization of Norrin missense mutations, related to Figure 2 and Figure 5. Mutations were categorized into six groups based on our structural and functional studies of Norrin (Cysteine mutations, Dimer interface mutations, Hydrophobic packing and protein stability mutations, Fz4 binding site mutations, Lrp5/6 binding site mutations, and other mutations). The full names of the disease acronyms are shown below the table. The information for Norrin mutations is obtained from the website ( Cysteine mutations Fz4 binding site Lp5/6 binding site Mutation Disease Mutation Disease C39R ND Y44C ND C55R, C55F ND L62P ND C65Y,C65W ND A63D,A63S ND C69S ND E66K PFV C95R,C95S ND G67R,G67E ND (severe) C96Y ND,EVR R74C ND,FEVR C96W CD S75P,S75C ND C110R,C110S ND F89L ND Dimer C110G FEVR R97P ND Interface C126S ND P98L ND C128R ND L108P ROP R41K EVR A118D ND R41T ND Y120C EVR R41S PFV R121G ND,PRDX H43R, H43Q ND R121W ND,FEVR,ROP V45M, V45E ND R121Q ND,FEVR L61F ND, VI R121L FEVR L61I FEVR I123N ND L61P ND Hydrophobic V60E ND L124F FEVR packing and protein R90C, ND stability R90P K54N ND, L103V FEVR FEVR R115L FEVR L108P ROP K104Q ND Y120C EVR (mild) K104N ND I123N ND S92P ND S101F ND, PHPV A105T ND ND = Norrie Disease (F)EVR = (Familial) Exudative Vitreoretinopathy PFV = Persistent Fetal Vasculature ROP = Retinopathy Of Prematurity CD = Coats Disease PRDX = Primary Retinal Dysplasia VI = Venous Insufficiency PHPV = Primary Hyperplastic Persistent Vitreous
HEK293T. Fig. 1 in the
 Supplementary Information Supplementary Figure 1 Zinc uptake assay of hzip4 and hzip4-δecd transiently expressed in HEK293T cells. The results of one representative e experiment are shown in Fig. 1 in
Supplementary Information Supplementary Figure 1 Zinc uptake assay of hzip4 and hzip4-δecd transiently expressed in HEK293T cells. The results of one representative e experiment are shown in Fig. 1 in
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than. SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with
 Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Supplementary Fig. S1. SAMHD1c has a more potent dntpase activity than SAMHD1c. Purified recombinant SAMHD1c and SAMHD1c proteins (with concentration of 800nM) were incubated with 1mM dgtp for the indicated
Structural basis of a novel PD-L1 nanobody for immune checkpoint blockade
 Structural basis of a novel PD-L1 nanobody for immune checkpoint blockade Supplementary Materials Supplementary methods Table S1-S Figure S1-S 1 1 1 1 1 1 1 1 1 0 1 0 Supplementary Methods Competitive
Structural basis of a novel PD-L1 nanobody for immune checkpoint blockade Supplementary Materials Supplementary methods Table S1-S Figure S1-S 1 1 1 1 1 1 1 1 1 0 1 0 Supplementary Methods Competitive
SUPPLEMENTARY INFORMATION. Supplementary Figures 1-8
 SUPPLEMENTARY INFORMATION Supplementary Figures 1-8 Supplementary Figure 1. TFAM residues contacting the DNA minor groove (A) TFAM contacts on nonspecific DNA. Leu58, Ile81, Asn163, Pro178, and Leu182
SUPPLEMENTARY INFORMATION Supplementary Figures 1-8 Supplementary Figure 1. TFAM residues contacting the DNA minor groove (A) TFAM contacts on nonspecific DNA. Leu58, Ile81, Asn163, Pro178, and Leu182
SUPPLEMENTARY INFORMATION
 Results Construct purification and coupling. Two A1-GP1bα ReaLiSM constructs, with and without cysteine residues near the N and C-termini (Fig. S2a), were expressed and purified by Ni affinity chromatography
Results Construct purification and coupling. Two A1-GP1bα ReaLiSM constructs, with and without cysteine residues near the N and C-termini (Fig. S2a), were expressed and purified by Ni affinity chromatography
Supplementary methods
 Supplementary methods Cell culture, infection, transfection, and RNA interference HEK293 cells and its derivatives were grown in DMEM supplemented with 10% FBS. Various constructs were introduced into
Supplementary methods Cell culture, infection, transfection, and RNA interference HEK293 cells and its derivatives were grown in DMEM supplemented with 10% FBS. Various constructs were introduced into
Supplementary Information
 Supplementary Information Supplementary Figure 1 Supplementary Figure 1 Scheme of isolating broadly neutralizing Abs against influenza viruses from human memory B cell repertoire. Representative fluorescence-labeled
Supplementary Information Supplementary Figure 1 Supplementary Figure 1 Scheme of isolating broadly neutralizing Abs against influenza viruses from human memory B cell repertoire. Representative fluorescence-labeled
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
doi: 10.1038/nature06147 SUPPLEMENTARY INFORMATION Figure S1 The genomic and domain structure of Dscam. The Dscam gene comprises 24 exons, encoding a signal peptide (SP), 10 IgSF domains, 6 fibronectin
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10016 Supplementary discussion on binding site density for protein complexes on the surface: The density of biotin sites on the chip is ~10 3 biotin-peg per µm 2. The biotin sites are
doi:10.1038/nature10016 Supplementary discussion on binding site density for protein complexes on the surface: The density of biotin sites on the chip is ~10 3 biotin-peg per µm 2. The biotin sites are
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Multiple sequence alignments of four Swi2/Snf2 subfamily proteins, ScChd1, SsoRad54 and the RNA helicase Vasa. The sequence alignments of the Swi2/Snf2 subfamily proteins, ScChd1
Supplementary Figure 1 Multiple sequence alignments of four Swi2/Snf2 subfamily proteins, ScChd1, SsoRad54 and the RNA helicase Vasa. The sequence alignments of the Swi2/Snf2 subfamily proteins, ScChd1
Coleman et al., Supplementary Figure 1
 Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Supplemental Figure 1. HepG2 cells were transfected with GLI luciferase reporter construct
 Supplemental Figure 1. HepG2 cells were transfected with GLI luciferase reporter construct (pgl38xgli), EWS-FLI1 luciferase reporter construct (NROB1-Luc) with or without GLI1, EWS- FLI1 and cdnas respectively.
Supplemental Figure 1. HepG2 cells were transfected with GLI luciferase reporter construct (pgl38xgli), EWS-FLI1 luciferase reporter construct (NROB1-Luc) with or without GLI1, EWS- FLI1 and cdnas respectively.
Nature Structural & Molecular Biology: doi: /nsmb.2548
 Supplementary Figure 1. Structure of GltPhout. (a) Stereo view of a slice through a single GltPhout protomer shown in stick representation along with 2Fo-Fc and anomalous difference electron maps. The
Supplementary Figure 1. Structure of GltPhout. (a) Stereo view of a slice through a single GltPhout protomer shown in stick representation along with 2Fo-Fc and anomalous difference electron maps. The
Supplementary Online Material. Structural mimicry in transcription regulation of human RNA polymerase II by the. DNA helicase RECQL5
 Supplementary Online Material Structural mimicry in transcription regulation of human RNA polymerase II by the DNA helicase RECQL5 Susanne A. Kassube, Martin Jinek, Jie Fang, Susan Tsutakawa and Eva Nogales
Supplementary Online Material Structural mimicry in transcription regulation of human RNA polymerase II by the DNA helicase RECQL5 Susanne A. Kassube, Martin Jinek, Jie Fang, Susan Tsutakawa and Eva Nogales
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.
 Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Fig. S1. Effect of p120-catenin overexpression on the interaction of SCUBE2 with E-cadherin. The expression plasmid encoding FLAG.SCUBE2, E-cadherin.Myc, or HA.p120-catenin was transfected in a combination
Box 7905, Engineering Building I, North Carolina State University, Raleigh, NC 27695, Phone: Fax:
 Inefficient ribosomal skipping enables simultaneous secretion and display of proteins in Saccharomyces cerevisiae Carlos A. Cruz-Teran 1, Karthik Tiruthani 1, Adam Mischler, and Balaji M. Rao * Department
Inefficient ribosomal skipping enables simultaneous secretion and display of proteins in Saccharomyces cerevisiae Carlos A. Cruz-Teran 1, Karthik Tiruthani 1, Adam Mischler, and Balaji M. Rao * Department
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description:
 File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. dcas9-mq1 fusion protein induces de novo
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. dcas9-mq1 fusion protein induces de novo
Supplementary Figure S1. Binding of HSA mutants to hfcrn. (a) The levels of titrated amounts of HSA
 Supplementary Figure S1. Binding of HSA mutants to hfcrn. (a) The levels of titrated amounts of HSA variants (5.0-0.002 μg/ml) directly coated in the wells at ph 6.0 were controlled using a horseradish
Supplementary Figure S1. Binding of HSA mutants to hfcrn. (a) The levels of titrated amounts of HSA variants (5.0-0.002 μg/ml) directly coated in the wells at ph 6.0 were controlled using a horseradish
CD93 and dystroglycan cooperation in human endothelial cell adhesion and migration
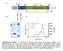
 /, Supplementary Advance Publications Materials 2016 CD93 and dystroglycan cooperation in human endothelial cell adhesion and migration Supplementary Materials Supplementary Figure S1: In ECs CD93 silencing
/, Supplementary Advance Publications Materials 2016 CD93 and dystroglycan cooperation in human endothelial cell adhesion and migration Supplementary Materials Supplementary Figure S1: In ECs CD93 silencing
Supporting Information: A GPCR dimerization interface in human cone opsins
 Supporting Information: A GPCR dimerization interface in human cone opsins Beata Jastrzebska *, William D. Comar *, Megan J. Kaliszewski, Kevin C. Skinner, Morgan H. Torcasio, Anthony S. Esway, Hui Jin,
Supporting Information: A GPCR dimerization interface in human cone opsins Beata Jastrzebska *, William D. Comar *, Megan J. Kaliszewski, Kevin C. Skinner, Morgan H. Torcasio, Anthony S. Esway, Hui Jin,
SUPPLEMENTAL MATERIAL. Supplemental Methods:
 SUPPLEMENTAL MATERIAL Supplemental Methods: Immunoprecipitation- As we described but with some modifications [22]. As part of another ongoing project, lysate from human umbilical vein endothelial cells
SUPPLEMENTAL MATERIAL Supplemental Methods: Immunoprecipitation- As we described but with some modifications [22]. As part of another ongoing project, lysate from human umbilical vein endothelial cells
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3363 Supplementary Figure 1 Several WNTs bind to the extracellular domains of PKD1. (a) HEK293T cells were co-transfected with indicated plasmids. Flag-tagged proteins were immunoprecipiated
DOI: 10.1038/ncb3363 Supplementary Figure 1 Several WNTs bind to the extracellular domains of PKD1. (a) HEK293T cells were co-transfected with indicated plasmids. Flag-tagged proteins were immunoprecipiated
Conformational changes in IgE contribute to its. uniquely slow dissociation rate from receptor FcεRI
 Conformational changes in IgE contribute to its uniquely slow dissociation rate from receptor FcεRI M.D. Holdom, A.M. Davies, J.E. Nettleship, S.C. Bagby, B. Dhaliwal, E. Girardi, J. Hunt, H.J. Gould,
Conformational changes in IgE contribute to its uniquely slow dissociation rate from receptor FcεRI M.D. Holdom, A.M. Davies, J.E. Nettleship, S.C. Bagby, B. Dhaliwal, E. Girardi, J. Hunt, H.J. Gould,
Data Sheet GITR / NF-ĸB-Luciferase Reporter (Luc) - Jurkat Cell Line Catalog #60546
 Data Sheet GITR / NF-ĸB-Luciferase Reporter (Luc) - Jurkat Cell Line Catalog #60546 Description This cell line expresses a surface human GITR (glucocorticoid-induced TNFR family related gene; TNFRSF18;
Data Sheet GITR / NF-ĸB-Luciferase Reporter (Luc) - Jurkat Cell Line Catalog #60546 Description This cell line expresses a surface human GITR (glucocorticoid-induced TNFR family related gene; TNFRSF18;
Minicollagen cysteine-rich domains encode distinct modes of. Anja Tursch, Davide Mercadante, Jutta Tennigkeit, Frauke Gräter and Suat
 Supplementary Figure S1. Conformational dynamics of partially reduced CRDs. (A) Root mean square deviation (RMSD) and (B) radius of gyration (R G ) distributions for the N-CRD (green) and C-CRD (red) domains,
Supplementary Figure S1. Conformational dynamics of partially reduced CRDs. (A) Root mean square deviation (RMSD) and (B) radius of gyration (R G ) distributions for the N-CRD (green) and C-CRD (red) domains,
human Cdc45 Figure 1c. (c)
 1 Details of the refined crystallographic model of human Cdc45 and comparison of its active-site region with that of bacterial RecJ. (a) Stereo view of a representative example of the final 2F o -F c electron
1 Details of the refined crystallographic model of human Cdc45 and comparison of its active-site region with that of bacterial RecJ. (a) Stereo view of a representative example of the final 2F o -F c electron
Supplementary Figure 1 PZA inhibits root hair formation as well as cell elongation in the maturation zone of eto1-2 roots. (A) The PI staining of the
 Supplementary Figure 1 PZA inhibits root hair formation as well as cell elongation in the maturation zone of eto1-2 roots. (A) The PI staining of the roots of three-day-old etiolated seedlings of Col-0
Supplementary Figure 1 PZA inhibits root hair formation as well as cell elongation in the maturation zone of eto1-2 roots. (A) The PI staining of the roots of three-day-old etiolated seedlings of Col-0
pt7ht vector and over-expressed in E. coli as inclusion bodies. Cells were lysed in 6 M
 Supplementary Methods MIG6 production, purification, inhibition, and kinase assays MIG6 segment 1 (30mer, residues 334 364) peptide was synthesized using standard solid-phase peptide synthesis as described
Supplementary Methods MIG6 production, purification, inhibition, and kinase assays MIG6 segment 1 (30mer, residues 334 364) peptide was synthesized using standard solid-phase peptide synthesis as described
Supplementary Table 1. DNA sequence synthesized to express the Zika virus NS5 protein.
 Supplementary Table 1. DNA sequence synthesized to express the Zika virus NS5 protein. MR766 NS5 sequence 1 ACAGAGAACAGATTGGTGGTGGAGGTGGGACGGGAGAGACTCTGGGAGAGAAGTGGAAAG 61 CTCGTCTGAATCAGATGTCGGCCCTGGAGTTCTACTCTTATAAAAAGTCAGGTATCACTG
Supplementary Table 1. DNA sequence synthesized to express the Zika virus NS5 protein. MR766 NS5 sequence 1 ACAGAGAACAGATTGGTGGTGGAGGTGGGACGGGAGAGACTCTGGGAGAGAAGTGGAAAG 61 CTCGTCTGAATCAGATGTCGGCCCTGGAGTTCTACTCTTATAAAAAGTCAGGTATCACTG
Supplementary Figure 1. Supporting data for Figure 1
 Supplementary Figure 1 Supporting data for Figure 1 (A) Schematic of BLI assay used to measure dissociation. 5 monobiotinylated substrate DNA (identical to λ1, Figure 2) is associated with streptavidin-coated
Supplementary Figure 1 Supporting data for Figure 1 (A) Schematic of BLI assay used to measure dissociation. 5 monobiotinylated substrate DNA (identical to λ1, Figure 2) is associated with streptavidin-coated
Supplementary Information
 Supplementary Information Supplementary Figure 1: Over-expression of CD300f in NIH3T3 cells enhances their capacity to phagocytize AC. (a) NIH3T3 cells were stably transduced by EV, CD300f WT or CD300f
Supplementary Information Supplementary Figure 1: Over-expression of CD300f in NIH3T3 cells enhances their capacity to phagocytize AC. (a) NIH3T3 cells were stably transduced by EV, CD300f WT or CD300f
Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and
 Supplementary Tables Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and their side chain contacting residues in the second chain (B) a Interface Res. in Contacting
Supplementary Tables Supplementary Table 1: List of CH3 domain interface residues in the first chain (A) and their side chain contacting residues in the second chain (B) a Interface Res. in Contacting
Supplementary Information
 Supplementary Information Peroxiredoxin-2 and STAT3 form a redox relay for H 2 O 2 signaling Mirko C. Sobotta 1, Willy Liou 1, Sarah Stöcker 1, Deepti Talwar 1, Michael Oehler 1, Thomas Ruppert 2, Annette
Supplementary Information Peroxiredoxin-2 and STAT3 form a redox relay for H 2 O 2 signaling Mirko C. Sobotta 1, Willy Liou 1, Sarah Stöcker 1, Deepti Talwar 1, Michael Oehler 1, Thomas Ruppert 2, Annette
Data Sheet CD137/NF-κB Reporter - HEK293 Recombinant Cell Line Catalog # 79289
 Data Sheet CD137/NF-κB Reporter - HEK293 Recombinant Cell Line Catalog # 79289 Background Human CD137 (4-1BB; TNFRS9) is an inducible co-stimulatory molecule that activates T cells. CD137:CD137L-mediated
Data Sheet CD137/NF-κB Reporter - HEK293 Recombinant Cell Line Catalog # 79289 Background Human CD137 (4-1BB; TNFRS9) is an inducible co-stimulatory molecule that activates T cells. CD137:CD137L-mediated
Nature Structural & Molecular Biology: doi: /nsmb.3018
 Supplementary Figure 1 Validation of genetic complementation assay in Bmal1 / Per2 Luc fibroblasts. (a) Only Bmal1, not Bmal2, rescues circadian rhythms from cells. Cells expressing various Bmal constructs
Supplementary Figure 1 Validation of genetic complementation assay in Bmal1 / Per2 Luc fibroblasts. (a) Only Bmal1, not Bmal2, rescues circadian rhythms from cells. Cells expressing various Bmal constructs
Supplementary Table, Figures and Videos
 Supplementary Table, Figures and Videos Table S1. Oligonucleotides used for different approaches. (A) RT-qPCR study. (B) qpcr study after ChIP assay. (C) Probes used for EMSA. Figure S1. Notch activation
Supplementary Table, Figures and Videos Table S1. Oligonucleotides used for different approaches. (A) RT-qPCR study. (B) qpcr study after ChIP assay. (C) Probes used for EMSA. Figure S1. Notch activation
Sarker et al. Supplementary Material. Subcellular Fractionation
 Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
ASPP1 Fw GGTTGGGAATCCACGTGTTG ASPP1 Rv GCCATATCTTGGAGCTCTGAGAG
 Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplemental Materials and Methods Plasmids: the following plasmids were used in the supplementary data: pwzl-myc- Lats2 (Aylon et al, 2006), pretrosuper-vector and pretrosuper-shp53 (generous gift of
Supplementatry Fig 1. Domain structure, biophysical characterisation and electron microscopy of a TD. (a) XTACC3/Maskin and XMAP215/chTOG domain
 Supplementatry Fig 1. Domain structure, biophysical characterisation and electron microscopy of a TD. (a) XTACC3/Maskin and XMAP215/chTOG domain architecture. Various C-terminal fragments were cloned and
Supplementatry Fig 1. Domain structure, biophysical characterisation and electron microscopy of a TD. (a) XTACC3/Maskin and XMAP215/chTOG domain architecture. Various C-terminal fragments were cloned and
Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid
 Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
Supplementary Figures Suppl. Figure 1: RCC1 sequence and sequence alignments. (a) Amino acid sequence of Drosophila RCC1. Same colors are for Figure 1 with sequence of β-wedge that interacts with Ran in
supplementary information
 DOI: 10.1038/ncb2172 Figure S1 p53 regulates cellular NADPH and lipid levels via inhibition of G6PD. (a) U2OS cells stably expressing p53 shrna or a control shrna were transfected with control sirna or
DOI: 10.1038/ncb2172 Figure S1 p53 regulates cellular NADPH and lipid levels via inhibition of G6PD. (a) U2OS cells stably expressing p53 shrna or a control shrna were transfected with control sirna or
Molecular design principles underlying β-strand swapping. in the adhesive dimerization of cadherins
 Supplementary information for: Molecular design principles underlying β-strand swapping in the adhesive dimerization of cadherins Jeremie Vendome 1,2,3,5, Shoshana Posy 1,2,3,5,6, Xiangshu Jin, 1,3 Fabiana
Supplementary information for: Molecular design principles underlying β-strand swapping in the adhesive dimerization of cadherins Jeremie Vendome 1,2,3,5, Shoshana Posy 1,2,3,5,6, Xiangshu Jin, 1,3 Fabiana
Masayoshi Honda, Jeehae Park, Robert A. Pugh, Taekjip Ha, and Maria Spies
 Molecular Cell, Volume 35 Supplemental Data Single-Molecule Analysis Reveals Differential Effect of ssdna-binding Proteins on DNA Translocation by XPD Helicase Masayoshi Honda, Jeehae Park, Robert A. Pugh,
Molecular Cell, Volume 35 Supplemental Data Single-Molecule Analysis Reveals Differential Effect of ssdna-binding Proteins on DNA Translocation by XPD Helicase Masayoshi Honda, Jeehae Park, Robert A. Pugh,
Colchicine. Colchicine. a b c d e
 α1-tub Colchicine Laulimalide RB3 β1-tub Vinblastine α2-tub Taxol Colchicine β2-tub Maytansine a b c d e Supplementary Figure 1 Structures of microtubule and tubulin (a)the cartoon of head to tail arrangement
α1-tub Colchicine Laulimalide RB3 β1-tub Vinblastine α2-tub Taxol Colchicine β2-tub Maytansine a b c d e Supplementary Figure 1 Structures of microtubule and tubulin (a)the cartoon of head to tail arrangement
Recruitment of Grb2 to surface IgG and IgE provides antigen receptor-intrinsic costimulation to class-switched B cells
 SUPPLEMENTARY FIGURES Recruitment of Grb2 to surface IgG and IgE provides antigen receptor-intrinsic costimulation to class-switched B cells Niklas Engels, Lars Morten König, Christina Heemann, Johannes
SUPPLEMENTARY FIGURES Recruitment of Grb2 to surface IgG and IgE provides antigen receptor-intrinsic costimulation to class-switched B cells Niklas Engels, Lars Morten König, Christina Heemann, Johannes
Supplementary information Bi-specific Aptamers mediating Tumour Cell Lysis
 Supplementary information Bi-specific Aptamers mediating Tumour Cell Lysis Figure S1 SELEX scheme of different CD16 DNA selection approaches. Nine rounds of filter SELEX were conducted in parallel, only
Supplementary information Bi-specific Aptamers mediating Tumour Cell Lysis Figure S1 SELEX scheme of different CD16 DNA selection approaches. Nine rounds of filter SELEX were conducted in parallel, only
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb327 a b Sequence coverage (%) 4 3 2 IP: -GFP isoform IP: GFP IP: -GFP IP: GFP Sequence coverage (%) 4 3 2 IP: -GFP IP: GFP 33 52 58 isoform 2 33 49 47 IP: Control IP: Peptide Sequence Start
DOI:.38/ncb327 a b Sequence coverage (%) 4 3 2 IP: -GFP isoform IP: GFP IP: -GFP IP: GFP Sequence coverage (%) 4 3 2 IP: -GFP IP: GFP 33 52 58 isoform 2 33 49 47 IP: Control IP: Peptide Sequence Start
Supplementary data. sienigma. F-Enigma F-EnigmaSM. a-p53
 Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
Supplementary data Supplemental Figure 1 A sienigma #2 sienigma sicontrol a-enigma - + ++ - - - - - - + ++ - - - - - - ++ B sienigma F-Enigma F-EnigmaSM a-flag HLK3 cells - - - + ++ + ++ - + - + + - -
1 24 C63 C β- β-
 M40 Signal leaved RS1 Domain 59 110 142 Discoidin Domain 223 1 24 63 219 224 + - β- β- M M e e O O H H His 6 -Tag 250 150 100 75 50 37 * 25 20 Supplementary Figure 1 Purification of wild-type retinoschisin.
M40 Signal leaved RS1 Domain 59 110 142 Discoidin Domain 223 1 24 63 219 224 + - β- β- M M e e O O H H His 6 -Tag 250 150 100 75 50 37 * 25 20 Supplementary Figure 1 Purification of wild-type retinoschisin.
Data Sheet PD-1 / NFAT - Reporter - Jurkat Recombinant Cell Line Catalog #: 60535
 Data Sheet PD-1 / NFAT - Reporter - Jurkat Recombinant Cell Line Catalog #: 60535 Product Description Recombinant Jurkat T cell expressing firefly luciferase gene under the control of NFAT response elements
Data Sheet PD-1 / NFAT - Reporter - Jurkat Recombinant Cell Line Catalog #: 60535 Product Description Recombinant Jurkat T cell expressing firefly luciferase gene under the control of NFAT response elements
SUPPLEMENTAL MATERIAL BIOCHEMICAL AND STRUCTURAL STUDIES ON THE M. TUBERCULOSIS O 6 -METHYLGUANINE METHYLTRANSFERASE AND MUTATED VARIANTS
 SUPPLEMENTAL MATERIAL BIOCHEMICAL AND STRUCTURAL STUDIES ON THE M. TUBERCULOSIS O 6 -METHYLGUANINE METHYLTRANSFERASE AND MUTATED VARIANTS Riccardo Miggiano 1, Valentina Casazza 1, Silvia Garavaglia 1,
SUPPLEMENTAL MATERIAL BIOCHEMICAL AND STRUCTURAL STUDIES ON THE M. TUBERCULOSIS O 6 -METHYLGUANINE METHYLTRANSFERASE AND MUTATED VARIANTS Riccardo Miggiano 1, Valentina Casazza 1, Silvia Garavaglia 1,
Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells.
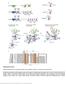
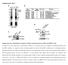 Supplementary Fig. 1 Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells. (a) FRTL-5 cells were treated with 1 mm dibutyryl camp for 24 h, and the lysates
Supplementary Fig. 1 Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells. (a) FRTL-5 cells were treated with 1 mm dibutyryl camp for 24 h, and the lysates
Expanded View Figures
 EMO reports rystal structure of Mis18 Yippee-like domain Lakxmi Subramanian et al Expanded View Figures Figure EV1. Structural characterization of the N-terminal Yippee-like globular domain of spmis18.
EMO reports rystal structure of Mis18 Yippee-like domain Lakxmi Subramanian et al Expanded View Figures Figure EV1. Structural characterization of the N-terminal Yippee-like globular domain of spmis18.
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL Materials and Methods Circular dichroism (CD) spectroscopy. Far ultraviolet (UV) CD spectra of apo- and holo- CaM and the CaM mutants were recorded on a Jasco J-715 spectropolarimeter
SUPPLEMENTARY MATERIAL Materials and Methods Circular dichroism (CD) spectroscopy. Far ultraviolet (UV) CD spectra of apo- and holo- CaM and the CaM mutants were recorded on a Jasco J-715 spectropolarimeter
HPV E6 oncoprotein targets histone methyltransferases for modulating specific. Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu,
 1 HPV E oncoprotein targets histone methyltransferases for modulating specific gene transcription 3 5 Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu, Cheng-Ming Chiang, Sheng-Chung
1 HPV E oncoprotein targets histone methyltransferases for modulating specific gene transcription 3 5 Chih-Hung Hsu, Kai-Lin Peng, Hua-Ci Jhang, Chia-Hui Lin, Shwu-Yuan Wu, Cheng-Ming Chiang, Sheng-Chung
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Conserved arginines on the rim of Hfq catalyze base pair formation and exchange Subrata Panja and Sarah A. Woodson T.C. Jenkins Department of Biophysics, Johns Hopkins University,
SUPPLEMENTARY INFORMATION Conserved arginines on the rim of Hfq catalyze base pair formation and exchange Subrata Panja and Sarah A. Woodson T.C. Jenkins Department of Biophysics, Johns Hopkins University,
T H E J O U R N A L O F C E L L B I O L O G Y
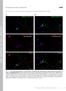 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Kanaani et al., http://www.jcb.org/cgi/content/full/jcb.200912101/dc1 Figure S1. The K2 rabbit polyclonal antibody is specific for GAD67,
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Kanaani et al., http://www.jcb.org/cgi/content/full/jcb.200912101/dc1 Figure S1. The K2 rabbit polyclonal antibody is specific for GAD67,
Supplementary Figure 1. Electron microscopy of gb-698glyco/1g2 Fab complex. a)
 Supplementary Figure 1. Electron microscopy of gb-698glyco/1g2 Fab complex. a) Representative images of 2D class averages of gb-698glyc bound to 1G2 Fab. Top views of the complex were underrepresented
Supplementary Figure 1. Electron microscopy of gb-698glyco/1g2 Fab complex. a) Representative images of 2D class averages of gb-698glyc bound to 1G2 Fab. Top views of the complex were underrepresented
Supporting Information
 Supporting Information DNA Crystals as Vehicles for Biocatalysis. Chun Geng and Paul J. Paukstelis* *Email: paukstel@umd.edu Recombinant MBP-tagged RNase inhibitor expression and purification. The porcine
Supporting Information DNA Crystals as Vehicles for Biocatalysis. Chun Geng and Paul J. Paukstelis* *Email: paukstel@umd.edu Recombinant MBP-tagged RNase inhibitor expression and purification. The porcine
Supplementary Figure 1 a
 3 min PMA 45 min PMA AnnexinV-FITC Supplementary Figure 1 5 min PMA 15 min PMA a 9 min PMA 12 min PMA 5 min FGF7 15 min FGF7 3 min FGF7 6 min FGF7 9 min FGF7 12 min FGF7 5 min control 3 min control 6 min
3 min PMA 45 min PMA AnnexinV-FITC Supplementary Figure 1 5 min PMA 15 min PMA a 9 min PMA 12 min PMA 5 min FGF7 15 min FGF7 3 min FGF7 6 min FGF7 9 min FGF7 12 min FGF7 5 min control 3 min control 6 min
No wash 2 Washes 2 Days ** ** IgG-bead phagocytosis (%)
 Supplementary Figures Supplementary Figure 1. No wash 2 Washes 2 Days Tat Control ** ** 2 4 6 8 IgG-bead phagocytosis (%) Supplementary Figure 1. Reversibility of phagocytosis inhibition by Tat. Human
Supplementary Figures Supplementary Figure 1. No wash 2 Washes 2 Days Tat Control ** ** 2 4 6 8 IgG-bead phagocytosis (%) Supplementary Figure 1. Reversibility of phagocytosis inhibition by Tat. Human
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Domain architecture and conformational states of the decapping complex, as revealed by structural studies. (a) Domain organization of Schizosaccharomyces pombe (Sp) and Saccharomyces
Supplementary Figure 1 Domain architecture and conformational states of the decapping complex, as revealed by structural studies. (a) Domain organization of Schizosaccharomyces pombe (Sp) and Saccharomyces
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature15713 Supplementary Table 1. The DNAs used in this study D1 D2 D3 D4 D5 D6 D7 D8 D9 D10 12-bp FAM-18-bp biotinylated 26-bp s Biotin-free 26-bp s 58-bpDNA 58-bpDNA AT-rich 58-bpDNA CG-rich
doi:10.1038/nature15713 Supplementary Table 1. The DNAs used in this study D1 D2 D3 D4 D5 D6 D7 D8 D9 D10 12-bp FAM-18-bp biotinylated 26-bp s Biotin-free 26-bp s 58-bpDNA 58-bpDNA AT-rich 58-bpDNA CG-rich
Supplementary Figure 1. Botrocetin induces binding of human VWF to human
 Supplementary Figure 1: Supplementary Figure 1. Botrocetin induces binding of human VWF to human platelets in the absence of elevated shear and induces platelet agglutination as detected by flow cytometry.
Supplementary Figure 1: Supplementary Figure 1. Botrocetin induces binding of human VWF to human platelets in the absence of elevated shear and induces platelet agglutination as detected by flow cytometry.
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Map of pct-vhvl-k1 native V H :V L display vector.
 Supplementary Figure 1 Map of pct-vhvl-k1 native V H :V L display vector. Natively-paired V H :V L sequences are cloned en masse into this vector for human antibody repertoire mining, and their corresponding
Supplementary Figure 1 Map of pct-vhvl-k1 native V H :V L display vector. Natively-paired V H :V L sequences are cloned en masse into this vector for human antibody repertoire mining, and their corresponding
Figure S1. Sequence alignments of ATRIP and ATR TopBP1 interacting regions.
 A H. sapiens 204 TKLQTS--ERANKLAAPSVSH VSPRKNPSVVIKPEACS-PQFGKTSFPTKESFSANMS LP 259 B. taurus 201 TKLQSS--ERANKLAVPTVSH VSPRKSPSVVIKPEACS-PQFGKPSFPTKESFSANKS LP 257 M. musculus 204 TKSQSN--GRTNKPAAPSVSH
A H. sapiens 204 TKLQTS--ERANKLAAPSVSH VSPRKNPSVVIKPEACS-PQFGKTSFPTKESFSANMS LP 259 B. taurus 201 TKLQSS--ERANKLAVPTVSH VSPRKSPSVVIKPEACS-PQFGKPSFPTKESFSANKS LP 257 M. musculus 204 TKSQSN--GRTNKPAAPSVSH
Data Sheet. TCR activator / PD-L1 - CHO Recombinant Cell line Cat. #: 60536
 Data Sheet TCR activator / PD-L1 - CHO Recombinant Cell line Cat. #: 60536 Product Description Recombinant CHO-K1 cells constitutively expressing human PD-L1 (Programmed Cell Death 1 Ligand 1, CD274, B7
Data Sheet TCR activator / PD-L1 - CHO Recombinant Cell line Cat. #: 60536 Product Description Recombinant CHO-K1 cells constitutively expressing human PD-L1 (Programmed Cell Death 1 Ligand 1, CD274, B7
Document S1. Supplemental Experimental Procedures and Three Figures (see next page)
 Supplemental Data Document S1. Supplemental Experimental Procedures and Three Figures (see next page) Table S1. List of Candidate Genes Identified from the Screen. Candidate genes, corresponding dsrnas
Supplemental Data Document S1. Supplemental Experimental Procedures and Three Figures (see next page) Table S1. List of Candidate Genes Identified from the Screen. Candidate genes, corresponding dsrnas
This is the author's accepted version of the manuscript.
 This is the author's accepted version of the manuscript. The definitive version is published in Nature Communications Online Edition: 2015/4/16 (Japan time), doi:10.1038/ncomms7780. The final version published
This is the author's accepted version of the manuscript. The definitive version is published in Nature Communications Online Edition: 2015/4/16 (Japan time), doi:10.1038/ncomms7780. The final version published
SUPPLEMENTARY INFORMATION. Chemical modulation of Chaperone-mediated autophagy by novel
 SUPPLEMENTARY INFORMATION Chemical modulation of Chaperone-mediated autophagy by novel retinoic acid derivatives Jaime Anguiano 1, Thomas P Garner 2, Murugesan Mahalingam 1, Bhaskar C. Das 1, Evripidis
SUPPLEMENTARY INFORMATION Chemical modulation of Chaperone-mediated autophagy by novel retinoic acid derivatives Jaime Anguiano 1, Thomas P Garner 2, Murugesan Mahalingam 1, Bhaskar C. Das 1, Evripidis
Comparative Analysis of Argonaute-Dependent Small RNA Pathways in Drosophila
 Molecular Cell, Volume 32 Supplemental Data Comparative Analysis of Argonaute-Dependent Small RNA Pathways in Drosophila Rui Zhou, Ikuko Hotta, Ahmet M. Denli, Pengyu Hong, Norbert Perrimon, and Gregory
Molecular Cell, Volume 32 Supplemental Data Comparative Analysis of Argonaute-Dependent Small RNA Pathways in Drosophila Rui Zhou, Ikuko Hotta, Ahmet M. Denli, Pengyu Hong, Norbert Perrimon, and Gregory
SUPPLEMENTAL FIGURE LEGENDS. Figure S1: Homology alignment of DDR2 amino acid sequence. Shown are
 SUPPLEMENTAL FIGURE LEGENDS Figure S1: Homology alignment of DDR2 amino acid sequence. Shown are the amino acid sequences of human DDR2, mouse DDR2 and the closest homologs in zebrafish and C. Elegans.
SUPPLEMENTAL FIGURE LEGENDS Figure S1: Homology alignment of DDR2 amino acid sequence. Shown are the amino acid sequences of human DDR2, mouse DDR2 and the closest homologs in zebrafish and C. Elegans.
Supplementary Figure 1.
 Supplementary Figure 1. Assessment of quaternary structure of soluble RSV F proteins. Soluble variants of F proteins from A2 and B1 RSV strains were expressed in HEK293 cells. The cell culture supernatants
Supplementary Figure 1. Assessment of quaternary structure of soluble RSV F proteins. Soluble variants of F proteins from A2 and B1 RSV strains were expressed in HEK293 cells. The cell culture supernatants
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity.
 Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
Supplemental Figure 1 Human REEP family of proteins can be divided into two distinct subfamilies. Residues (single letter amino acid code) identical
 Supplemental Figure Human REEP family of proteins can be divided into two distinct subfamilies. Residues (single letter amino acid code) identical in all six REEPs are highlighted in green. Additional
Supplemental Figure Human REEP family of proteins can be divided into two distinct subfamilies. Residues (single letter amino acid code) identical in all six REEPs are highlighted in green. Additional
Supplemental Information for:
 Supplemental Information for: Antibody-induced dimerization of FGFR1 promotes receptor endocytosis independently of its kinase activity Łukasz Opaliński*, Aleksandra Sokołowska-Wędzina, Martyna Szczepara,
Supplemental Information for: Antibody-induced dimerization of FGFR1 promotes receptor endocytosis independently of its kinase activity Łukasz Opaliński*, Aleksandra Sokołowska-Wędzina, Martyna Szczepara,
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
SUPPLEMENTARY INFORMATION. Reengineering Protein Interfaces Yields Copper-Inducible Ferritin Cage Assembly
 SUPPLEMENTARY INFORMATION Reengineering Protein Interfaces Yields Copper-Inducible Ferritin Cage Assembly Dustin J. E. Huard, Kathleen M. Kane and F. Akif Tezcan* Department of Chemistry and Biochemistry,
SUPPLEMENTARY INFORMATION Reengineering Protein Interfaces Yields Copper-Inducible Ferritin Cage Assembly Dustin J. E. Huard, Kathleen M. Kane and F. Akif Tezcan* Department of Chemistry and Biochemistry,
Supplemental Information Molecular Cell, Volume 41
 Supplemental Information Molecular Cell, Volume 41 Molecular Mechanisms for the RNA-Dependent ATPase Activity of Upf1 and Its Regulation by Upf2 Sutapa Chakrabarti, Uma Jayachandran, Fabien Bonneau, Francesca
Supplemental Information Molecular Cell, Volume 41 Molecular Mechanisms for the RNA-Dependent ATPase Activity of Upf1 and Its Regulation by Upf2 Sutapa Chakrabarti, Uma Jayachandran, Fabien Bonneau, Francesca
Supplemental Material Igreja and Izaurralde 1. CUP promotes deadenylation and inhibits decapping of mrna targets. Catia Igreja and Elisa Izaurralde
 Supplemental Material Igreja and Izaurralde 1 CUP promotes deadenylation and inhibits decapping of mrna targets Catia Igreja and Elisa Izaurralde Supplemental Materials and methods Functional assays and
Supplemental Material Igreja and Izaurralde 1 CUP promotes deadenylation and inhibits decapping of mrna targets Catia Igreja and Elisa Izaurralde Supplemental Materials and methods Functional assays and
PHF20 is an effector protein of p53 double lysine methylation
 SUPPLEMENTARY INFORMATION PHF20 is an effector protein of p53 double lysine methylation that stabilizes and activates p53 Gaofeng Cui 1, Sungman Park 2, Aimee I Badeaux 3, Donghwa Kim 2, Joseph Lee 1,
SUPPLEMENTARY INFORMATION PHF20 is an effector protein of p53 double lysine methylation that stabilizes and activates p53 Gaofeng Cui 1, Sungman Park 2, Aimee I Badeaux 3, Donghwa Kim 2, Joseph Lee 1,
Supporting Information
 Supporting Information Powell and Xu 10.1073/pnas.0807274105 SI Methods Design of Estrogen Receptor (ER) Fusion Constructs for Use in Bioluminescence Resonance Energy Transfer (BRET) Assay. Because the
Supporting Information Powell and Xu 10.1073/pnas.0807274105 SI Methods Design of Estrogen Receptor (ER) Fusion Constructs for Use in Bioluminescence Resonance Energy Transfer (BRET) Assay. Because the
SUPPLEMENTARY INFORMATION. Design and Characterization of Bivalent BET Inhibitors
 SUPPLEMENTARY INFORMATION Design and Characterization of Bivalent BET Inhibitors Minoru Tanaka 1,2,#, Justin M. Roberts 1,#, Hyuk-Soo Seo 3, Amanda Souza 1, Joshiawa Paulk 1, Thomas G. Scott 1, Stephen
SUPPLEMENTARY INFORMATION Design and Characterization of Bivalent BET Inhibitors Minoru Tanaka 1,2,#, Justin M. Roberts 1,#, Hyuk-Soo Seo 3, Amanda Souza 1, Joshiawa Paulk 1, Thomas G. Scott 1, Stephen
Supplemental Materials and Methods
 Supplemental Materials and Methods Co-immunoprecipitation (Co-IP) assay Cells were lysed with NETN buffer (20 mm Tris-HCl, ph 8.0, 0 mm NaCl, 1 mm EDT, 0.5% Nonidet P-40) containing 50 mm β-glycerophosphate,
Supplemental Materials and Methods Co-immunoprecipitation (Co-IP) assay Cells were lysed with NETN buffer (20 mm Tris-HCl, ph 8.0, 0 mm NaCl, 1 mm EDT, 0.5% Nonidet P-40) containing 50 mm β-glycerophosphate,
Synthesis of the pyridinyl analogues of dibenzylideneacetone. (pyr-dba) via an improved Claisen-Schmidt condensation,
 Synthesis of the pyridinyl analogues of dibenzylideneacetone (pyr-dba) via an improved Claisen-Schmidt condensation, displaying diverse biological activities as curcumin analogues Bin Cao, Yong Wang, Kan
Synthesis of the pyridinyl analogues of dibenzylideneacetone (pyr-dba) via an improved Claisen-Schmidt condensation, displaying diverse biological activities as curcumin analogues Bin Cao, Yong Wang, Kan
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Figure S2. Verification of ihog and boi
 Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
Supplementary Figure S1 Purification of deubiquitinases HEK293 cells were transfected with the indicated DUB-expressing plasmids.
 Supplementary Figure S1 Purification of deubiquitinases HEK293 cells were transfected with the indicated DUB-expressing plasmids. The cells were harvested 72 h after transfection. FLAG-tagged deubiquitinases
Supplementary Figure S1 Purification of deubiquitinases HEK293 cells were transfected with the indicated DUB-expressing plasmids. The cells were harvested 72 h after transfection. FLAG-tagged deubiquitinases
Drug Discovery Research Clinical Screening. Comparison of ELISA and AlphaScreen Assay Technologies for Measurement of Protein Expression Levels
 Drug Discovery Research Clinical Screening FLAG Expression Measurement Application Note Comparison of ELISA and AlphaScreen Assay Technologies for Measurement of Protein Expression Levels ALPHASCREEN APPLICATION
Drug Discovery Research Clinical Screening FLAG Expression Measurement Application Note Comparison of ELISA and AlphaScreen Assay Technologies for Measurement of Protein Expression Levels ALPHASCREEN APPLICATION
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/11/e1601625/dc1 Supplementary Materials for A molecular mechanism of chaperone-client recognition This PDF file includes: Lichun He, Timothy Sharpe, Adam Mazur,
advances.sciencemag.org/cgi/content/full/2/11/e1601625/dc1 Supplementary Materials for A molecular mechanism of chaperone-client recognition This PDF file includes: Lichun He, Timothy Sharpe, Adam Mazur,
Supporting Information
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 215 Supporting Information Quantitative Description of Thermodynamic and Kinetic Properties of the
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 215 Supporting Information Quantitative Description of Thermodynamic and Kinetic Properties of the
SUPPLEMENTARY INFORMATION
 doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan).
 1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
How to run Alpha assay: How to setup an Alpha assay Make your own assay!
 How to run Alpha assay: How to setup an Alpha assay Make your own assay! 1 2009 PerkinElmer AlphaLISA kits - recommendations before starting the assay Samples: Phenol red and hemoglobin: choose AlphaLISA
How to run Alpha assay: How to setup an Alpha assay Make your own assay! 1 2009 PerkinElmer AlphaLISA kits - recommendations before starting the assay Samples: Phenol red and hemoglobin: choose AlphaLISA
X2-C/X1-Y X2-C/VCAM-Y. FRET efficiency. Ratio YFP/CFP
 FRET efficiency.7.6..4.3.2 X2-C/X1-Y X2-C/VCAM-Y.1 1 2 3 Ratio YFP/CFP Supplemental Data 1. Analysis of / heterodimers in live cells using FRET. FRET saturation curves were obtained using cells transiently
FRET efficiency.7.6..4.3.2 X2-C/X1-Y X2-C/VCAM-Y.1 1 2 3 Ratio YFP/CFP Supplemental Data 1. Analysis of / heterodimers in live cells using FRET. FRET saturation curves were obtained using cells transiently
p53 increases MHC class I expression by upregulating the endoplasmic reticulum aminopeptidase ERAP1
 Supplementary Information p53 increases MHC class I expression by upregulating the endoplasmic reticulum aminopeptidase ERAP1 Bei Wang 1, Dandan Niu 1, Liyun Lai 1, & Ee Chee Ren 1,2 1 Singapore Immunology
Supplementary Information p53 increases MHC class I expression by upregulating the endoplasmic reticulum aminopeptidase ERAP1 Bei Wang 1, Dandan Niu 1, Liyun Lai 1, & Ee Chee Ren 1,2 1 Singapore Immunology
Breast cell lines with Combination Index (CI) and p53 mutation status
 Supplementary Table 1 Breast cell lines with Combination Index (CI) and p53 mutation status Cell Line CI p53 status a BT20 0.44 K132Q BT549 0.53 R249S CAL120 0.61 c.672+2t>g CAL51 0.73 wild-type CAMA1
Supplementary Table 1 Breast cell lines with Combination Index (CI) and p53 mutation status Cell Line CI p53 status a BT20 0.44 K132Q BT549 0.53 R249S CAL120 0.61 c.672+2t>g CAL51 0.73 wild-type CAMA1
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the
 Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
J. Cell Sci. 128: doi: /jcs : Supplementary Material. Supplemental Figures. Journal of Cell Science Supplementary Material
 Supplemental Figures Figure S1. Trio controls endothelial barrier function. (A) TagRFP-shTrio constructs were expressed in ECs. Western blot shows efficient Trio knockdown in TagRFP-expressing ECs. (B)
Supplemental Figures Figure S1. Trio controls endothelial barrier function. (A) TagRFP-shTrio constructs were expressed in ECs. Western blot shows efficient Trio knockdown in TagRFP-expressing ECs. (B)
Supplementary Figure 1. FRET probe labeling locations in the Cas9-RNA-DNA complex.
 Supplementary Figure 1. FRET probe labeling locations in the Cas9-RNA-DNA complex. (a) Cy3 and Cy5 labeling locations shown in the crystal structure of Cas9-RNA bound to a cognate DNA target (PDB ID: 4UN3)
Supplementary Figure 1. FRET probe labeling locations in the Cas9-RNA-DNA complex. (a) Cy3 and Cy5 labeling locations shown in the crystal structure of Cas9-RNA bound to a cognate DNA target (PDB ID: 4UN3)
Supplementary Information. Arrays of Individual DNA Molecules on Nanopatterned Substrates
 Supplementary Information Arrays of Individual DNA Molecules on Nanopatterned Substrates Roland Hager, Alma Halilovic, Jonathan R. Burns, Friedrich Schäffler, Stefan Howorka S1 Figure S-1. Characterization
Supplementary Information Arrays of Individual DNA Molecules on Nanopatterned Substrates Roland Hager, Alma Halilovic, Jonathan R. Burns, Friedrich Schäffler, Stefan Howorka S1 Figure S-1. Characterization
