Box 7905, Engineering Building I, North Carolina State University, Raleigh, NC 27695, Phone: Fax:
|
|
|
- Cameron Buck Powers
- 5 years ago
- Views:
Transcription
1 Inefficient ribosomal skipping enables simultaneous secretion and display of proteins in Saccharomyces cerevisiae Carlos A. Cruz-Teran 1, Karthik Tiruthani 1, Adam Mischler, and Balaji M. Rao * Department of Chemical and Biomolecular Engineering, North Carolina State University, Raleigh, NC 1 These authors contributed equally * Address Correspondence to: Box 7905, Engineering Building I, North Carolina State University, Raleigh, NC 27695, Phone: Fax: bmrao@ncsu.edu SUPPORTING INFORMATION Contents Figure S1: List of constructs generated for simultaneous secretion and yeast cell surface display. Figure S2: Confirmation of the presence of immobilized IgG on streptavidin beads. Figure S3: Yeast surface expression of Sso7dstrep and Sso7dhFc in pctcon, pct-ct-f2a, and pct-ct-f2a. Figure S4: Enrichment of high affinity binders from a yeast display library of Sso7d mutants incorporating the F2A peptide. Figure S5: Activity of GOx expressed as yeast cell surface fusions. Figure S6: Yeast surface expression and target binding of Sso7dstrep and Sso7dhFc in pctcon, pct-ct-f2a, and pct-ct-f2a-sed1sp. Figure S7: Secretion and display of Sso7dhFc in pct-ct-f2a-prepro and pct-ct-f2a-sed1sp. Table S1: List of lysozyme binders isolated from the combinatorial library. Table S2: List of primers. Table S3: List of gene fragments.
2 Figure S1: List of plasmid constructs generated for N-terminal and C-terminal secretion and display. A) Protein of interest (shown as Sso7d here) is expressed as an N-terminal fusion to Aga2. B) Protein of interest is expressed as a C-terminal fusion to Aga2 and lacks a secretory signal upstream of the Sso7d gene. C) Protein of interest is expressed as a C-terminal fusion to Aga2 and contains the SED1SP secretory signal upstream of the Sso7d gene. D) Protein of interest is expressed as a C-terminal fusion to Aga2 and contains the prepro secretory signal upstream of the Sso7d gene. Vector maps were generated with SnapGene Software (GSL Biotech, Chicago, IL; available at snapgene.com).
3 Figure S2: Confirmation of the presence of immobilized IgG on streptavidin beads. Streptavidin beads (control; black) or beads coated with human IgG (red) were labeled with an anti-human IgG antibody.
4 Figure S3: Yeast surface expression of Sso7dstrep (A) and Sso7dhFc (B) in pctcon (red), pct-nt-f2a (green), and pct-ct-f2a (blue).
5 Figure S4: Enrichment of high affinity binders from a yeast display library of Sso7d mutants incorporating the F2A peptide. Fluorescence ratio was computed and cells were placed in one of 6 bins. The fraction of cells in each bin relative to total cells analyzed is plotted for lysozyme labeling at 250 nm. The bin corresponding to all cells with fluorescence ratio is denoted by 0.25, by 0.5, and so on. All error bars indicate standard error of the mean from triplicate experiments. *p<0.05 for comparison with pre-facs population in the same bin.
6 Figure S5: Activity of GOx expressed as yeast cell surface fusions. GOx activity on the yeast surface after 24, 48, and 72 hours of protein induction in SGCAA medium at 20 C for GOx in pctcon (grey), and GOx (green) and Sso7dhFc-GOx (orange) in pct-nt-f2a, and GOx in pct-ct-f2a (blue). The absorbance of the quenched assay solution at 450 nm after 5 minutes of reaction is plotted as a measure of GOx activity. Error bars correspond to standard error of three independent repeats.
7 Figure S6: Yeast surface expression and target binding of Sso7dstrep and Sso7dhFc in pctcon (A and D), pct-ct-f2a (B and E) and pct-ct-f2a-sed1sp (C and F). Sso7dstrep (A-C) was labeled with 50 nm streptavidin-pe and an anti-c-myc antibody. Sso7dhFc (D-F) was labeled with 500nM human IgG-biotin and an anti-c-myc antibody.
8 Figure S7: Secretion and yeast surface display of Sso7dhFc in pct-ct-f2a-prepro and pct- CT-F2A-SED1SP. A) Yeast cells expressing cell surface Sso7dhFc were labeled with 500 nm IgG and an anti-c-myc antibody. B) Protein in supernatant containing a c-myc tag was captured using His-Tag isolation beads and detected by cytometry using an anti-c-myc antibody. C) Immunoblotting analysis of Sso7dhFc in cell culture supernatant from cells with pct-ct-f2aprepro (lane 1), pct-ct-f2a (lane 2), pct-ct-f2a-sed1sp (lane 3). A molecular weight ladder is included in lane 4.
9 Position WT I K K W V M S T E T R A E 1 I V D D K K A V L W S V G 2 I V D D K K A V L W N V E 3 I T Y F W H V D L S N E E 4 I F F I D D L K L A L E E 5 T V Y F H G F D L F I E E 6 I T Y L F E Y D L Q N G E 7 T V Y F H G F D L F I E E 8 T V Y F H G F D L W L E E 9 I C F F W C C D L Q S C E 10 T N N D F Q F N L I T F E Table S1: List of lysozyme binders isolated by combinatorial library screening. The library was generated by random mutagenesis of low affinity binders isolated from a library of Sso7d mutants. Mutations outside the 10 positions mutagenized in the original Sso7d library are in bold font. Wild type Sso7d (WT) is shown as a reference.
10 Primer Sequence (5 3 ) Pf1.1 GAGTCCGGATCCGAGAACCTGTACTTCCAAGGG Pf1.2 Pr1.1 Pf2 Pr2 Pf3.1 Pf3.2 Pf3.3 Pr3.1 Pr3.2 Pf4 Pr4 Pf5 Pr5 Pf6 Pr6 Pr7 GAGTACCCTAGGTACCCGTACGACGTTCCAGACTACGCTCAGGAACTGACAACTATAT GCG GAGTCACTCGAGTCATTAAAAAACATACTGT GAGTCAGCTAGCATGGCGACCGTGAAATTTAAATAT GAGTCAGGATCCTTTTTTCTGTTTTTCCAGCATCTG GCATTCGAATTCATGAAGGTTTTGATTGTCTT CAGGAGCCTAGGTACCCGTACGACGTTCCAGAC GGCTGGCATATGAAGGTTTTGATTGTCTTGTTG GCATTTCTCGAGTCATTACAAGTCTTCTTC GAGTATGCTAGCCCCTTGGAAGTACAGGTTCTC CAGAAGGCTCTTTGGACAAGAGACACCACCACCACCACCACGCTAGCATGGCGACCG TGAAATTTAAATAT GGAGATAAGCTTTTGTTCCCCTTGGAAGTACAGGTTCTCGGATCCTTTTTTCTGTTTTT CCAGCATCTG CCTTTCCTCTCTCATTCCCTCA AATGCCCTTGTTTGGTAGTAAT GATTACCCTAGGATGCAGACTCTCCTTGTGAGCTCGCTT GAGCACGGATCCCTGCATGGAAGCATAATCTTCCAAGATAG GAGCACCTCGAGCTGCATGGAAGCATAATCTTCCAAGATAG Table S2: List of oligonucleotide primers.
11 Gene block Sequence (5 3 ) Gene block 1 Gene block 2 GAATTCATGAAGGTTTTGATTGTCTTGTTGGCTATCTTCGCTGCTTTGCCATT GGCCTTAGCTCAACCGGTTATTTCTACTACCGTCGGTTCCGCTGCAGAAGGC TCTTTGGACAAGAGACACCACCACCACCACCACGCTAGCATGGCGACCGTG AAATTTAAATATAAAGGCGAAGAAAAACAGGTGGATATTAGCAAAATTTGGAT TGTGGCCCGCGATGGCAAATGGATTGACTTTTCTTATGATCTGGGCGGCGG CAAATCAGGCATAGGCTACGTGAGCGAAAAAGATGCGCCGAAAGAACTGCT GCAGATGCTGGAAAAACAGAAAAAAGGATCCGAGAACCTGTACTTCCAAGG GGAACAAAAGCTTATCTCCGAAGAAGACTTGGGAAGCGGAGTGAAACAGAC TTTGAATTTTGACCTTCTCAAGTTGGCGGGAGACGTGGAGTCCAACCCTGGA CCTCAGGAACTGACAACTATATGCGAGCAAATCCCCTCACCAACTTTAGAAT CGACGCCGTACTCTTTGTCAACGACTACTATTTTGGCCAACGGGAAGGCAAT GCAAGGAGTTTTTGAATATTACAAATCAGTAACGTTTGTCAGTAATTGCGGTT CTCACCCCTCAACAACTAGCAAAGGCAGCCCCATAAACACACAGTATGTTTT TTAATGACTCGAG GAGAACCTGTACTTCCAAGGGGAACAAAAGCTTATCTCCGAAGAAGACTTGG GAAGCGGAGTGAAACAGACTTTGAATTTTGACCTTCTCAAGTTGGCGGGAGA CGTGGAGTCCAACCCTGGACCTCCCGGGGGTGGTGGTGGTTCTGGAGGAG GCTCTGGTGGTGGTGGTTCTGGTGGTGGTGGTTCTCCTAGGCAGGAACTGA CAACTATATGCGAGCAAATCCCCTCACCAACTTTAGAATCGACGCCGTACTC TTTGTCAACGACTACTATTTTGGCCAACGGGAAGGCAATGCAAGGAGTTTTT GAATATTACAAATCAGTAACGTTTGTCAGTAATTGCGGTTCTCACCCCTCAAC AACTAGCAAAGGCAGCCCCATAAACACACAGTATGTTTTTTAATGA Gene block 3 GAATTCATGAAGGTTTTGATTGTCTTGTTGGCTATCTTCGCTGCTTTGCCATT GGCCTTAGCTCAACCGGTTATTTCTACTACCGTCGGTTCCGCTGCAGAAGGC TCTTTGGACAAGAGACAGGAACTGACAACTATATGCGAGCAAATCCCCTCAC CAACTTTAGAATCGACGCCGTACTCTTTGTCAACGACTACTATTTTGGCCAAC GGGAAGGCAATGCAAGGAGTTTTTGAATATTACAAATCAGTAACGTTTGTCA GTAATTGCGGTTCTCACCCCTCAACAACTAGCAAAGGCAGCCCCATAAACAC ACAGTATGTTTTTCCCGGGGGTGGTGGTGGTTCTGGAGGAGGCTCTGGTGG TGGTGGTTCTGGTGGTGGTGGTTCTCCTAGGTATCCGTACGACGTTCCAGA CTACGCTGGAAGCGGAGTGAAACAGACTTTGAATTTTGACCTTCTCAAGTTG GCGGGAGACGTGGAGTCCAACCCTGGACCTCATCACCACCATCACCACGAG AACCTGTACTTCCAAGGGGCTAGCATGGCGACCGTGAAATTTAAATATAAAG GCGAAGAAAAACAGGTGGATATTAGCAAAATTTGGATTGTGGCCCGCGATG GCAAATGGATTGACTTTTCTTATGATCTGGGCGGCGGCAAATCAGGCATAGG CTACGTGAGCGAAAAAGATGCGCCGAAAGAACTGCTGCAGATGCTGGAAAA ACAGAAAAAAGGATCCGAACAAAAGCTTATCTCCGAAGAAGACTTGTAATGA CTCGAG Gene 4 TACCCGTACGACGTTCCAGACTACGCTGGAAGCGGAGTGAAACAGACTTTG AATTTTGACCTTCTCAAGTTGGCGGGAGACGTGGAGTCCAACCCTGGACCT CATATGAAGTTATCCACAGTCCTACTGAGTGCCGGACTTGCGAGCACGACC CTAGCCCAGCACCATCACCATCACCACCCGACTCCTACGCCTACACCGACG CCCACAGAGAACCTGTACTTCCAAGGGGCTAGCATGGCGACCGTGAAATTT AAATATAAAGGCGAAGAAAAACAGGTGGATATTAGCAAAATTTGGATTGTGG CCCGCGATGGCAAATGGATTGACTTTTCTTATGATCTGGGCGGCGGCAAATC AGGCATAGGCTACGTGAGCGAAAAAGATGCGCCGAAAGAACTGCTGCAGAT
12 GCTGGAAAAACAGAAAAAAGGATCCGAACAAAAGCTTATCTCCGAAGAAGAC TTG Gene 5 CATATGAAGGTTTTGATTGTCTTGTTGGCTATCTTCGCTGCTTTGCCATTGGC CTTAGCTCAACCGGTTATTTCTACTACCGTCGGTTCCGCTGCAGAAGGCTCT TTGGACAAGAGACACCATCACCATCACCACGAGAACCTGTACTTCCAAGGG GCTAGCATGGCGACCGTGAAATTTAAATATAAAGGCGAAGAAAAACAGGTGG ATATTAGCAAAATTTACCTTGTGTCGCGCATTGGCAAACGTATTCTTTTTATG TATGATCTGGGCGGCGGCAAATATGGCATTGGCCGCGTGAGCGAAAAAGAT GCGCCGAAAGAACTGCTGCAGATGCTGGAAAAACAGAAAAAAGGATCC Table S3: List of gene fragments
Supplemental Data. Farmer et al. (2010) Plant Cell /tpc
 Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
SUPPLEMENTARY INFORMATION. Tolerance of a knotted near infrared fluorescent protein to random circular permutation
 SUPPLEMENTARY INFORMATION Tolerance of a knotted near infrared fluorescent protein to random circular permutation Naresh Pandey 1,3, Brianna E. Kuypers 2,4, Barbara Nassif 1, Emily E. Thomas 1,3, Razan
SUPPLEMENTARY INFORMATION Tolerance of a knotted near infrared fluorescent protein to random circular permutation Naresh Pandey 1,3, Brianna E. Kuypers 2,4, Barbara Nassif 1, Emily E. Thomas 1,3, Razan
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the
 Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Supplementary Fig. 1. Schematic structure of TRAIP and RAP80. The prey line below TRAIP indicates bait and the two lines above RAP80 highlight the prey clones identified in the yeast two hybrid screen.
Module 2 overview SPRING BREAK
 1 Module 2 overview lecture lab 1. Introduction to the module 1. Start-up protein eng. 2. Rational protein design 2. Site-directed mutagenesis 3. Fluorescence and sensors 3. DNA amplification 4. Protein
1 Module 2 overview lecture lab 1. Introduction to the module 1. Start-up protein eng. 2. Rational protein design 2. Site-directed mutagenesis 3. Fluorescence and sensors 3. DNA amplification 4. Protein
Supplemental Figure 1. Immunoblot analysis of wild-type and mutant forms of JAZ3
 Supplemental Data. Chung and Howe (2009). A Critical Role for the TIFY Motif in Repression of Jasmonate Signaling by a Stabilized Splice Variant of the JASMONATE ZIM-domain protein JAZ10 in Arabidopsis.
Supplemental Data. Chung and Howe (2009). A Critical Role for the TIFY Motif in Repression of Jasmonate Signaling by a Stabilized Splice Variant of the JASMONATE ZIM-domain protein JAZ10 in Arabidopsis.
Enhancers mutations that make the original mutant phenotype more extreme. Suppressors mutations that make the original mutant phenotype less extreme
 Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Interactomics and Proteomics 1. Interactomics The field of interactomics is concerned with interactions between genes or proteins. They can be genetic interactions, in which two genes are involved in the
Module 2 overview SPRING BREAK
 1 Module 2 overview lecture lab 1. Introduction to the module 1. Start-up protein eng. 2. Rational protein design 2. Site-directed mutagenesis 3. Fluorescence and sensors 3. DNA amplification 4. Protein
1 Module 2 overview lecture lab 1. Introduction to the module 1. Start-up protein eng. 2. Rational protein design 2. Site-directed mutagenesis 3. Fluorescence and sensors 3. DNA amplification 4. Protein
Phage Antibody Selection With Reichert SPR System
 Phage Antibody Selection With Reichert SPR System Reichert Technologies Webinar April 4, 2016 Mark A. Sullivan, Ph.D. Department of Microbiology and Immunology University of Rochester Presentation Outline
Phage Antibody Selection With Reichert SPR System Reichert Technologies Webinar April 4, 2016 Mark A. Sullivan, Ph.D. Department of Microbiology and Immunology University of Rochester Presentation Outline
Supplementary Methods
 Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
SUPPLEMENTARY FIGURES and LEGENDS
 DARPins recognizing the tumor-associated antigen EpCAM selected by phage and ribosome display and engineered for multivalency Nikolas Stefan 1, 3, Patricia Martin-Killias 1, 3, Sascha Wyss-Stoeckle 1,
DARPins recognizing the tumor-associated antigen EpCAM selected by phage and ribosome display and engineered for multivalency Nikolas Stefan 1, 3, Patricia Martin-Killias 1, 3, Sascha Wyss-Stoeckle 1,
transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected with the wwp-luc reporter, and FLAG-tagged FHL1,
 Supplementary Data Supplementary Figure Legends Supplementary Figure 1 FHL-mediated TGFβ-responsive reporter transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected
Supplementary Data Supplementary Figure Legends Supplementary Figure 1 FHL-mediated TGFβ-responsive reporter transcription and the promoter occupancy of Smad proteins. (A) HepG2 cells were co-transfected
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Dissecting binding of a β-barrel outer membrane
Engineering splicing factors with designed specificities
 nature methods Engineering splicing factors with designed specificities Yang Wang, Cheom-Gil Cheong, Traci M Tanaka Hall & Zefeng Wang Supplementary figures and text: Supplementary Figure 1 Supplementary
nature methods Engineering splicing factors with designed specificities Yang Wang, Cheom-Gil Cheong, Traci M Tanaka Hall & Zefeng Wang Supplementary figures and text: Supplementary Figure 1 Supplementary
Supplementary Figure 1 - Characterization of rbag3 binding on macrophages cell surface.
 Supplementary Figure 1 - Characterization of rbag3 binding on macrophages cell surface. (a) Human PDAC cell lines were treated as indicated in Figure 1 panel F. Cells were analyzed for FITC-rBAG3 binding
Supplementary Figure 1 - Characterization of rbag3 binding on macrophages cell surface. (a) Human PDAC cell lines were treated as indicated in Figure 1 panel F. Cells were analyzed for FITC-rBAG3 binding
Module 2 overview! SPRING BREAK!
 1! Module 2 overview! lecture!!!!!lab! 1. Introduction to the module!!1. Start-up protein eng.! 2. Rational protein design!!2. Site-directed mutagenesis! 3. Fluorescence and sensors!!3. DNA amplification!
1! Module 2 overview! lecture!!!!!lab! 1. Introduction to the module!!1. Start-up protein eng.! 2. Rational protein design!!2. Site-directed mutagenesis! 3. Fluorescence and sensors!!3. DNA amplification!
Supplementary methods
 Supplementary methods Cell culture, infection, transfection, and RNA interference HEK293 cells and its derivatives were grown in DMEM supplemented with 10% FBS. Various constructs were introduced into
Supplementary methods Cell culture, infection, transfection, and RNA interference HEK293 cells and its derivatives were grown in DMEM supplemented with 10% FBS. Various constructs were introduced into
Polyclonal ARHGAP25 antibody was prepared from rabbit serum after intracutaneous
 Preparation and purification of polyclonal antibodies Polyclonal ARHGAP25 antibody was prepared from rabbit serum after intracutaneous injections of glutathione S-transferase-ARHGAP25-(509-619) (GST-coiled
Preparation and purification of polyclonal antibodies Polyclonal ARHGAP25 antibody was prepared from rabbit serum after intracutaneous injections of glutathione S-transferase-ARHGAP25-(509-619) (GST-coiled
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
DOI: 10.1038/ncb3240 Supplementary Figure 1 GBM cell lines display similar levels of p100 to p52 processing but respond differentially to TWEAK-induced TERT expression according to TERT promoter mutation
Supplementary Information
 Supplementary Information Supplementary Figure 1: Over-expression of CD300f in NIH3T3 cells enhances their capacity to phagocytize AC. (a) NIH3T3 cells were stably transduced by EV, CD300f WT or CD300f
Supplementary Information Supplementary Figure 1: Over-expression of CD300f in NIH3T3 cells enhances their capacity to phagocytize AC. (a) NIH3T3 cells were stably transduced by EV, CD300f WT or CD300f
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table.
 Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Supplementary Table 1. The Q-PCR primer sequence is summarized in the following table. Name Sequence (5-3 ) Application Flag-u ggactacaaggacgacgatgac Shared upstream primer for all the amplifications of
Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan).
 1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
1 2 3 4 5 6 7 8 Supplemental Materials and Methods Cell proliferation assay Cell proliferation was measured with Cell Counting Kit-8 (Dojindo Laboratories, Kumamoto, Japan). GCs were plated at 96-well
Supplementary Results Supplementary Table 1. P1 and P2 enrichment scores for wild-type subtiligase.
 Supplementary Results Supplementary Table 1. P1 and P2 enrichment scores for wild-type subtiligase. Supplementary Table 2. Masses of subtiligase mutants measured by LC-MS. Supplementary Table 2 (cont d).
Supplementary Results Supplementary Table 1. P1 and P2 enrichment scores for wild-type subtiligase. Supplementary Table 2. Masses of subtiligase mutants measured by LC-MS. Supplementary Table 2 (cont d).
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Map of pct-vhvl-k1 native V H :V L display vector.
 Supplementary Figure 1 Map of pct-vhvl-k1 native V H :V L display vector. Natively-paired V H :V L sequences are cloned en masse into this vector for human antibody repertoire mining, and their corresponding
Supplementary Figure 1 Map of pct-vhvl-k1 native V H :V L display vector. Natively-paired V H :V L sequences are cloned en masse into this vector for human antibody repertoire mining, and their corresponding
COMPAS for the Analysis of SELEX Experiments
 COMPAS for the Analysis of SELEX Experiments COMPAS (COMmon PAtternS) is a software tool that was especially developed to harness the technology of next generation sequencing (NGS) to bring light into
COMPAS for the Analysis of SELEX Experiments COMPAS (COMmon PAtternS) is a software tool that was especially developed to harness the technology of next generation sequencing (NGS) to bring light into
Designed BH3 peptides with high affinity and specificity for
 Designed BH3 peptides with high affinity and specificity for targeting Mcl-1 in cells Glenna Wink Foight a, Jeremy A. Ryan b, Stefano Gullá a,c, Anthony Letai b,d and Amy E. Keating a * Author affiliations:
Designed BH3 peptides with high affinity and specificity for targeting Mcl-1 in cells Glenna Wink Foight a, Jeremy A. Ryan b, Stefano Gullá a,c, Anthony Letai b,d and Amy E. Keating a * Author affiliations:
Table S1. Antibodies and recombinant proteins used in this study
 Table S1. Antibodies and recombinant proteins used in this study Labeled Antibody Clone Cat. no. Streptavidin-PerCP BD Biosciences 554064 Biotin anti-mouse CD25 7D4 BD Biosciences 553070 PE anti-mouse
Table S1. Antibodies and recombinant proteins used in this study Labeled Antibody Clone Cat. no. Streptavidin-PerCP BD Biosciences 554064 Biotin anti-mouse CD25 7D4 BD Biosciences 553070 PE anti-mouse
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3562 In the format provided by the authors and unedited. Supplementary Figure 1 Glucose deficiency induced FH-ATF2 interaction. In b-m, immunoblotting or immunoprecipitation analyses were
DOI: 10.1038/ncb3562 In the format provided by the authors and unedited. Supplementary Figure 1 Glucose deficiency induced FH-ATF2 interaction. In b-m, immunoblotting or immunoprecipitation analyses were
Supplemental Data. Sethi et al. (2014). Plant Cell /tpc
 Supplemental Data Supplemental Figure 1. MYC2 Binds to the E-box but not the E1-box of the MPK6 Promoter. (A) E1-box and E-box (wild type) containing MPK6 promoter fragment. The region shown in red denotes
Supplemental Data Supplemental Figure 1. MYC2 Binds to the E-box but not the E1-box of the MPK6 Promoter. (A) E1-box and E-box (wild type) containing MPK6 promoter fragment. The region shown in red denotes
Supplementary Information
 Supplementary Information Supplementary Figure 1 Supplementary Figure 1 Scheme of isolating broadly neutralizing Abs against influenza viruses from human memory B cell repertoire. Representative fluorescence-labeled
Supplementary Information Supplementary Figure 1 Supplementary Figure 1 Scheme of isolating broadly neutralizing Abs against influenza viruses from human memory B cell repertoire. Representative fluorescence-labeled
Fig. S1. CrgA intracellular levels in M. tuberculosis. Ten and twenty micrograms of
 Supplementary data Fig. S1. CrgA intracellular levels in M. tuberculosis. Ten and twenty micrograms of cell free protein lysates from WT M. tuberculosis (Rv) together with various known concentrations
Supplementary data Fig. S1. CrgA intracellular levels in M. tuberculosis. Ten and twenty micrograms of cell free protein lysates from WT M. tuberculosis (Rv) together with various known concentrations
Supplementary Table 1: Antigenic regions/sites on Ebola-GP identified using GFPDL*
 Supplementary Table 1: Antigenic regions/sites on Ebola-GP identified using GFPDL* Site AA Sequence 3x10 6 3x10 6 20x10 6 20x10 6 100x10 6 100x10 6 Post-1 st Post-2 nd Post-1 st Post-2 nd Post-1 st Post-2
Supplementary Table 1: Antigenic regions/sites on Ebola-GP identified using GFPDL* Site AA Sequence 3x10 6 3x10 6 20x10 6 20x10 6 100x10 6 100x10 6 Post-1 st Post-2 nd Post-1 st Post-2 nd Post-1 st Post-2
Supplemental Information
 Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
Supplemental Information ATP-dependent unwinding of U4/U6 snrnas by the Brr2 helicase requires the C-terminus of Prp8 Corina Maeder 1,3, Alan K. Kutach 1,2,3, and Christine Guthrie 1 1 Department of Biochemistry
Lecture 25 (11/15/17)
 Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Lecture 25 (11/15/17) Reading: Ch9; 328-332 Ch25; 990-995, 1005-1012 Problems: Ch9 (study-guide: applying); 1,2 Ch9 (study-guide: facts); 7,8 Ch25 (text); 1-3,5-7,9,10,13-15 Ch25 (study-guide: applying);
Supplementary Information
 Supplementary Information Supplementary Figures Figure S1. Study of mgtl translation in vitro. (A) Detection of 5 LR RNA using wild-type and anti-sd (91-95) substituted templates in a transcription-translation
Supplementary Information Supplementary Figures Figure S1. Study of mgtl translation in vitro. (A) Detection of 5 LR RNA using wild-type and anti-sd (91-95) substituted templates in a transcription-translation
Supplementary Figure 1. GST pull-down analysis of the interaction of GST-cIAP1 (A, B), GSTcIAP1
 Legends Supplementary Figure 1. GST pull-down analysis of the interaction of GST- (A, B), GST mutants (B) or GST- (C) with indicated proteins. A, B, Cell lysate from untransfected HeLa cells were loaded
Legends Supplementary Figure 1. GST pull-down analysis of the interaction of GST- (A, B), GST mutants (B) or GST- (C) with indicated proteins. A, B, Cell lysate from untransfected HeLa cells were loaded
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3363 Supplementary Figure 1 Several WNTs bind to the extracellular domains of PKD1. (a) HEK293T cells were co-transfected with indicated plasmids. Flag-tagged proteins were immunoprecipiated
DOI: 10.1038/ncb3363 Supplementary Figure 1 Several WNTs bind to the extracellular domains of PKD1. (a) HEK293T cells were co-transfected with indicated plasmids. Flag-tagged proteins were immunoprecipiated
Supporting Information: A GPCR dimerization interface in human cone opsins
 Supporting Information: A GPCR dimerization interface in human cone opsins Beata Jastrzebska *, William D. Comar *, Megan J. Kaliszewski, Kevin C. Skinner, Morgan H. Torcasio, Anthony S. Esway, Hui Jin,
Supporting Information: A GPCR dimerization interface in human cone opsins Beata Jastrzebska *, William D. Comar *, Megan J. Kaliszewski, Kevin C. Skinner, Morgan H. Torcasio, Anthony S. Esway, Hui Jin,
Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA.
 Summary of Supplemental Information Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA. Figure S2: rrna removal procedure is effective for clearing out
Summary of Supplemental Information Figure S1: NUN preparation yields nascent, unadenylated RNA with a different profile from Total RNA. Figure S2: rrna removal procedure is effective for clearing out
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Supplementary Figure 1 Origin use and efficiency are similar among WT, rrm3, pif1-m2, and pif1-m2; rrm3 strains. A. Analysis of fork progression around confirmed and likely origins (from cerevisiae.oridb.org).
Identification and characterization of DNA aptamers specific for
 SUPPLEMENTARY INFORMATION Identification and characterization of DNA aptamers specific for phosphorylation epitopes of tau protein I-Ting Teng 1,, Xiaowei Li 1,, Hamad Ahmad Yadikar #, Zhihui Yang #, Long
SUPPLEMENTARY INFORMATION Identification and characterization of DNA aptamers specific for phosphorylation epitopes of tau protein I-Ting Teng 1,, Xiaowei Li 1,, Hamad Ahmad Yadikar #, Zhihui Yang #, Long
Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells.
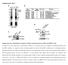 Supplementary Fig. 1 Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells. (a) FRTL-5 cells were treated with 1 mm dibutyryl camp for 24 h, and the lysates
Supplementary Fig. 1 Supplementary Fig. 1 Identification of Nedd4 as an IRS-2-associated protein in camp-treated FRTL-5 cells. (a) FRTL-5 cells were treated with 1 mm dibutyryl camp for 24 h, and the lysates
Contents... vii. List of Figures... xii. List of Tables... xiv. Abbreviatons... xv. Summary... xvii. 1. Introduction In vitro evolution...
 vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
vii Contents Contents... vii List of Figures... xii List of Tables... xiv Abbreviatons... xv Summary... xvii 1. Introduction...1 1.1 In vitro evolution... 1 1.2 Phage Display Technology... 3 1.3 Cell surface
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or
 Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Figure 1: TDP-43 is subject to lysine acetylation within the RNA-binding domain a) QBI-293 cells were transfected with TDP-43 in the presence or absence of the acetyltransferase CBP and acetylated TDP-43
Coleman et al., Supplementary Figure 1
 Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Coleman et al., Supplementary Figure 1 BrdU Merge G1 Early S Mid S Supplementary Figure 1. Sequential destruction of CRL4 Cdt2 targets during the G1/S transition. HCT116 cells were synchronized by sequential
Supplemental Figure 1 Kam et al., 2017
 nti-zikv IgG response (OD 450 nm) % Infectivity (Relative to virus only) 3 2 1 2 0 1 0 0 8 0 6 0 1-H1L1 2F-H1L3 SIgN-3C SIgN-3C-LL 1 4 0 2 0 0 1-H1L1 2F-H1L3 SIgN-3C SIgN-3C -LL 0 0.1 1 1 0 1 0 0 [b],
nti-zikv IgG response (OD 450 nm) % Infectivity (Relative to virus only) 3 2 1 2 0 1 0 0 8 0 6 0 1-H1L1 2F-H1L3 SIgN-3C SIgN-3C-LL 1 4 0 2 0 0 1-H1L1 2F-H1L3 SIgN-3C SIgN-3C -LL 0 0.1 1 1 0 1 0 0 [b],
Practice Problems 5. Location of LSA-GFP fluorescence
 Life Sciences 1a Practice Problems 5 1. Soluble proteins that are normally secreted from the cell into the extracellular environment must go through a series of steps referred to as the secretory pathway.
Life Sciences 1a Practice Problems 5 1. Soluble proteins that are normally secreted from the cell into the extracellular environment must go through a series of steps referred to as the secretory pathway.
Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified
 Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified by primers used for mrna expression analysis. Gray
Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified by primers used for mrna expression analysis. Gray
Supporting Information
 Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany RNA ligands that distinguish metabolite-induced conformations in the TPP riboswitch Günter Mayer, Marie-Sophie L. Raddatz, Julia D. Grunwald,
Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany RNA ligands that distinguish metabolite-induced conformations in the TPP riboswitch Günter Mayer, Marie-Sophie L. Raddatz, Julia D. Grunwald,
Supplementary Fig. 1
 a FL (1-2266) NL (1-1190) CL (1191-2266) HA-ICE1: - HA-ICE1: - - - FLAG-ICE2: + + + + FLAG-ELL: + + + + + + IP: anti-ha FLAG-ICE2 HA-ICE1-FL HA-ICE1-NL HA-ICE1-CL FLAG-ICE2 b IP: anti-ha FL (1-2266) NL
a FL (1-2266) NL (1-1190) CL (1191-2266) HA-ICE1: - HA-ICE1: - - - FLAG-ICE2: + + + + FLAG-ELL: + + + + + + IP: anti-ha FLAG-ICE2 HA-ICE1-FL HA-ICE1-NL HA-ICE1-CL FLAG-ICE2 b IP: anti-ha FL (1-2266) NL
This is the author's accepted version of the manuscript.
 This is the author's accepted version of the manuscript. The definitive version is published in Nature Communications Online Edition: 2015/4/16 (Japan time), doi:10.1038/ncomms7780. The final version published
This is the author's accepted version of the manuscript. The definitive version is published in Nature Communications Online Edition: 2015/4/16 (Japan time), doi:10.1038/ncomms7780. The final version published
B. Transgenic plants with strong phenotype (%)
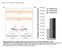 A. TCTAGTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT AAATATGGTCTAAAGAAGAAGAATATGGTCTAAAGAAGAAGAATATGG 2XP35S STTM165 5 GGGGGATGAAGctaCCTGGTCCGA3 3 CCCCCUACUUC---GGACCAGGCU5 mir165 HindIII mir165 96 nt GTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT
A. TCTAGTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT AAATATGGTCTAAAGAAGAAGAATATGGTCTAAAGAAGAAGAATATGG 2XP35S STTM165 5 GGGGGATGAAGctaCCTGGTCCGA3 3 CCCCCUACUUC---GGACCAGGCU5 mir165 HindIII mir165 96 nt GTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT
Deep sequencing reveals global patterns of mrna recruitment
 Supplementary information for: Deep sequencing reveals global patterns of mrna recruitment during translation initiation Rong Gao 1#*, Kai Yu 1#, Ju-Kui Nie 1,Teng-Fei Lian 1, Jian-Shi Jin 1, Anders Liljas
Supplementary information for: Deep sequencing reveals global patterns of mrna recruitment during translation initiation Rong Gao 1#*, Kai Yu 1#, Ju-Kui Nie 1,Teng-Fei Lian 1, Jian-Shi Jin 1, Anders Liljas
Nature Methods: doi: /nmeth Supplementary Figure 1. Construction of a sensitive TetR mediated auxotrophic off-switch.
 Supplementary Figure 1 Construction of a sensitive TetR mediated auxotrophic off-switch. A Production of the Tet repressor in yeast when conjugated to either the LexA4 or LexA8 promoter DNA binding sequences.
Supplementary Figure 1 Construction of a sensitive TetR mediated auxotrophic off-switch. A Production of the Tet repressor in yeast when conjugated to either the LexA4 or LexA8 promoter DNA binding sequences.
evaluated with UAS CLB eliciting UAS CIT -N Libraries increase in the
 Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Supplementary Figures Supplementary Figure 1: Promoter scaffold library assemblies. Many ensembless of libraries were evaluated in this work. As a legend, the box outline color in top half of the figure
Figure S6. Detection of anti-gfp antibodies in anti-dna and normal plasma without competition DNA--9
 Supplementary Information Ultrasensitive antibody detection by agglutination-pcr (ADAP) Cheng-ting Tsai 1 *, Peter V. Robinson 1 *, Carole A. Spencer 2 and Carolyn R. Bertozzi 3,4ǂ Department of 1 Chemistry,
Supplementary Information Ultrasensitive antibody detection by agglutination-pcr (ADAP) Cheng-ting Tsai 1 *, Peter V. Robinson 1 *, Carole A. Spencer 2 and Carolyn R. Bertozzi 3,4ǂ Department of 1 Chemistry,
Sarker et al. Supplementary Material. Subcellular Fractionation
 Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Supplementary Material Subcellular Fractionation Transfected 293T cells were harvested with phosphate buffered saline (PBS) and centrifuged at 2000 rpm (500g) for 3 min. The pellet was washed, re-centrifuged
Multiplex Assay Design
 Multiplex Assay Design Geeta Bhat, Luminex Molecular Diagnostics; Toronto. APHL/CDC Newborn Screening Molecular Workshop, CDC, Atlanta, GA June 28-30, 2011 Luminex Multiplexed Solutions. For Life. Luminex
Multiplex Assay Design Geeta Bhat, Luminex Molecular Diagnostics; Toronto. APHL/CDC Newborn Screening Molecular Workshop, CDC, Atlanta, GA June 28-30, 2011 Luminex Multiplexed Solutions. For Life. Luminex
Site-Directed Mutagenesis. Mutations in four Smad4 sites of mouse Gat1 promoter
 Supplement Supporting Materials and Methods Site-Directed Mutagenesis. Mutations in four Smad4 sites of mouse Gat1 promoter were independently generated using a two-step PCR method. The Smad4 binding site
Supplement Supporting Materials and Methods Site-Directed Mutagenesis. Mutations in four Smad4 sites of mouse Gat1 promoter were independently generated using a two-step PCR method. The Smad4 binding site
Supporting information
 Supporting information Construction of strains and plasmids To create ptc67, a PCR product obtained with primers cc2570-162f (gcatgggcaagcttgaggacggcgtcatgt) and cc2570+512f (gaggccgtggtaccatagaggcgggcg),
Supporting information Construction of strains and plasmids To create ptc67, a PCR product obtained with primers cc2570-162f (gcatgggcaagcttgaggacggcgtcatgt) and cc2570+512f (gaggccgtggtaccatagaggcgggcg),
Supplementary Figure 1 Telomerase RNA fragments used in single-molecule FRET experiments. A pseudoknot fragment (nts ) labeled at position U42
 Supplementary Figure 1 Telomerase RNA fragments used in single-molecule FRET experiments. A pseudoknot fragment (nts 32-195) labeled at position U42 with Cy3 (green circle) was constructed by a two piece
Supplementary Figure 1 Telomerase RNA fragments used in single-molecule FRET experiments. A pseudoknot fragment (nts 32-195) labeled at position U42 with Cy3 (green circle) was constructed by a two piece
Zhang et al., RepID facilitates replication Initiation. Supplemental Information:
 Supplemental Information: a b 1 Supplementary Figure 1 (a) DNA sequence of all the oligonucleotides used in this study. Only one strand is shown. The unshaded nucleotide sequences show changes from the
Supplemental Information: a b 1 Supplementary Figure 1 (a) DNA sequence of all the oligonucleotides used in this study. Only one strand is shown. The unshaded nucleotide sequences show changes from the
Advanced Therapeutic Antibody Discovery with Multiplexed Screening
 Advanced Therapeutic Antibody Discovery with Multiplexed Screening White Paper Scientists need powerful tools that can deliver results to fully understand the ability of candidate antibodies to interrupt
Advanced Therapeutic Antibody Discovery with Multiplexed Screening White Paper Scientists need powerful tools that can deliver results to fully understand the ability of candidate antibodies to interrupt
Combinatorial Evolution of Enzymes and Synthetic pathways Using
 Supplementary data Combinatorial Evolution of Enzymes and Synthetic pathways Using One-Step PCR Peng Jin,, Zhen Kang,,, *, Junli Zhang,, Linpei Zhang,, Guocheng Du,, and Jian Chen, The Key Laboratory of
Supplementary data Combinatorial Evolution of Enzymes and Synthetic pathways Using One-Step PCR Peng Jin,, Zhen Kang,,, *, Junli Zhang,, Linpei Zhang,, Guocheng Du,, and Jian Chen, The Key Laboratory of
Supplementary Figure 1. α-synuclein is truncated in PD and LBD brains. Nature Structural & Molecular Biology: doi: /nsmb.
 Supplementary Figure 1 α-synuclein is truncated in PD and LBD brains. (a) Specificity of anti-n103 antibody. Anti-N103 antibody was coated on an ELISA plate and different concentrations of full-length
Supplementary Figure 1 α-synuclein is truncated in PD and LBD brains. (a) Specificity of anti-n103 antibody. Anti-N103 antibody was coated on an ELISA plate and different concentrations of full-length
supplementary information
 DOI: 10.1038/ncb2172 Figure S1 p53 regulates cellular NADPH and lipid levels via inhibition of G6PD. (a) U2OS cells stably expressing p53 shrna or a control shrna were transfected with control sirna or
DOI: 10.1038/ncb2172 Figure S1 p53 regulates cellular NADPH and lipid levels via inhibition of G6PD. (a) U2OS cells stably expressing p53 shrna or a control shrna were transfected with control sirna or
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling
 Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
Supplementary information to accompany: A novel role for the DNA repair gene Rad51 in Netrin-1 signalling Glendining KA 1, Markie D 2, Gardner RJM 4, Franz EA 3, Robertson SP 4, Jasoni CL 1 Supplementary
CHAPTER 9 DNA Technologies
 CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
CHAPTER 9 DNA Technologies Recombinant DNA Artificially created DNA that combines sequences that do not occur together in the nature Basis of much of the modern molecular biology Molecular cloning of genes
Supplementatry Fig 1. Domain structure, biophysical characterisation and electron microscopy of a TD. (a) XTACC3/Maskin and XMAP215/chTOG domain
 Supplementatry Fig 1. Domain structure, biophysical characterisation and electron microscopy of a TD. (a) XTACC3/Maskin and XMAP215/chTOG domain architecture. Various C-terminal fragments were cloned and
Supplementatry Fig 1. Domain structure, biophysical characterisation and electron microscopy of a TD. (a) XTACC3/Maskin and XMAP215/chTOG domain architecture. Various C-terminal fragments were cloned and
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/9/eaat5401/dc1 Supplementary Materials for GLK-IKKβ signaling induces dimerization and translocation of the AhR-RORγt complex in IL-17A induction and autoimmune
advances.sciencemag.org/cgi/content/full/4/9/eaat5401/dc1 Supplementary Materials for GLK-IKKβ signaling induces dimerization and translocation of the AhR-RORγt complex in IL-17A induction and autoimmune
New Product: X-Aptamer Selection Kit. Updated April 2015
 New Product: X-Aptamer Selection Kit Updated April 2015 Technology Basics X-Aptamers are synthetic affinity molecules Similar in function to monoclonal antibodies X-Aptamers combine a variety of chemical
New Product: X-Aptamer Selection Kit Updated April 2015 Technology Basics X-Aptamers are synthetic affinity molecules Similar in function to monoclonal antibodies X-Aptamers combine a variety of chemical
SUPPLEMENTARY INFORMATION. Supplementary Figures 1-8
 SUPPLEMENTARY INFORMATION Supplementary Figures 1-8 Supplementary Figure 1. TFAM residues contacting the DNA minor groove (A) TFAM contacts on nonspecific DNA. Leu58, Ile81, Asn163, Pro178, and Leu182
SUPPLEMENTARY INFORMATION Supplementary Figures 1-8 Supplementary Figure 1. TFAM residues contacting the DNA minor groove (A) TFAM contacts on nonspecific DNA. Leu58, Ile81, Asn163, Pro178, and Leu182
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Figure S2. Verification of ihog and boi
 Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
Figure S1. Verification of ihog Mutation by Protein Immunoblotting Extracts from S2R+ cells, embryos, and adults were analyzed by immunoprecipitation and immunoblotting with anti-ihog antibody. The Ihog
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb54 sensitivity (+/-) 3 det- WS bri-5 BL (nm) bzr-dbri- bzr-d/ bri- g h i a b c d e f Hypcocotyl length(cm) 5 5 PAC Hypocotyl length (cm)..8..4. 5 5 PPZ - - + + + + - + - + - + PPZ - - + + +
DOI:.38/ncb54 sensitivity (+/-) 3 det- WS bri-5 BL (nm) bzr-dbri- bzr-d/ bri- g h i a b c d e f Hypcocotyl length(cm) 5 5 PAC Hypocotyl length (cm)..8..4. 5 5 PPZ - - + + + + - + - + - + PPZ - - + + +
Supplementary Information. A novel human endogenous retroviral protein inhibits cell-cell fusion. Supplementary Figures:
 Supplementary Information A novel human endogenous retroviral protein inhibits cell-cell fusion Jun Sugimoto, Makiko Sugimoto, Helene Bernstein, Yoshihiro Jinno and Danny J. Schust Supplementary Figures:
Supplementary Information A novel human endogenous retroviral protein inhibits cell-cell fusion Jun Sugimoto, Makiko Sugimoto, Helene Bernstein, Yoshihiro Jinno and Danny J. Schust Supplementary Figures:
Supplemental Data. Hu et al. Plant Cell (2017) /tpc
 1 2 3 4 Supplemental Figure 1. DNA gel blot analysis of homozygous transgenic plants. (Supports Figure 1.) 5 6 7 8 Rice genomic DNA was digested with the restriction enzymes EcoRⅠ and BamHⅠ. Lanes in the
1 2 3 4 Supplemental Figure 1. DNA gel blot analysis of homozygous transgenic plants. (Supports Figure 1.) 5 6 7 8 Rice genomic DNA was digested with the restriction enzymes EcoRⅠ and BamHⅠ. Lanes in the
Supplementary Figure 1 Collision-induced dissociation (CID) mass spectra of peptides from PPK1, PPK2, PPK3 and PPK4 respectively.
 Supplementary Figure 1 lision-induced dissociation (CID) mass spectra of peptides from PPK1, PPK, PPK3 and PPK respectively. % of nuclei with signal / field a 5 c ppif3:gus pppk1:gus 0 35 30 5 0 15 10
Supplementary Figure 1 lision-induced dissociation (CID) mass spectra of peptides from PPK1, PPK, PPK3 and PPK respectively. % of nuclei with signal / field a 5 c ppif3:gus pppk1:gus 0 35 30 5 0 15 10
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
doi:10.1038/nature12119 SUPPLEMENTARY FIGURES AND LEGENDS pre-let-7a- 1 +14U pre-let-7a- 1 Ddx3x Dhx30 Dis3l2 Elavl1 Ggt5 Hnrnph 2 Osbpl5 Puf60 Rnpc3 Rpl7 Sf3b3 Sf3b4 Tia1 Triobp U2af1 U2af2 1 6 2 4 3
Supplementary Figure 1. Intracellular distribution of the EPE peptide. HeLa cells were serum-starved (16 h, 0.1%), and treated with EPE peptide,
 Supplementary Figure 1. Intracellular distribution of the EPE peptide. HeLa cells were serum-starved (16 h, 0.1%), and treated with EPE peptide, conjugated with either TAT or Myristic acid and biotin for
Supplementary Figure 1. Intracellular distribution of the EPE peptide. HeLa cells were serum-starved (16 h, 0.1%), and treated with EPE peptide, conjugated with either TAT or Myristic acid and biotin for
TARGETS Anti-Biotin Anti-Streptavidin Anti-FITC Anti-Phycoerythrin Anti-Mouse IgG Anti-Rabbit IgG (H+L) Anti-Human IgG (H+L)
 Signal Amplifiers Genisphere s UltraAmp reagents are 3DNA dendrimers customized with a variety of labels and targeting moieties, to increase the sensitivity in any immunoassay or nucleic acid detection
Signal Amplifiers Genisphere s UltraAmp reagents are 3DNA dendrimers customized with a variety of labels and targeting moieties, to increase the sensitivity in any immunoassay or nucleic acid detection
Supplementary Infomation
 Supplementary Infomation Mycobacterium avium MAV2054 protein induces macrophage apoptosis through targeting to mitochondria and reduces intracellular growth of the bacteria. Kang-In Lee 1,2, Jake Whang
Supplementary Infomation Mycobacterium avium MAV2054 protein induces macrophage apoptosis through targeting to mitochondria and reduces intracellular growth of the bacteria. Kang-In Lee 1,2, Jake Whang
pgbkt7 Anti- Myc AH109 strain (KDa) 50
 pgbkt7 (KDa) 50 37 Anti- Myc AH109 strain Supplementary Figure 1. Protein expression of CRN and TDR in yeast. To analyse the protein expression of CRNKD and TDRKD, total proteins extracted from yeast culture
pgbkt7 (KDa) 50 37 Anti- Myc AH109 strain Supplementary Figure 1. Protein expression of CRN and TDR in yeast. To analyse the protein expression of CRNKD and TDRKD, total proteins extracted from yeast culture
SUPPLEMENTARY INFORMATION
 doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
doi:.38/nature899 Supplementary Figure Suzuki et al. a c p7 -/- / WT ratio (+)/(-) p7 -/- / WT ratio Log X 3. Fold change by treatment ( (+)/(-)) Log X.5 3-3. -. b Fold change by treatment ( (+)/(-)) 8
Supplementary Figure 1. Nature Structural & Molecular Biology: doi: /nsmb.3494
 Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
Supplementary Figure 1 Pol structure-function analysis (a) Inactivating polymerase and helicase mutations do not alter the stability of Pol. Flag epitopes were introduced using CRISPR/Cas9 gene targeting
Chapter 14 Regulation of Transcription
 Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
Chapter 14 Regulation of Transcription Cis-acting sequences Distance-independent cis-acting elements Dissecting regulatory elements Transcription factors Overview transcriptional regulation Transcription
AD BD TOC1. Supplementary Figure 1: Yeast two-hybrid assays showing the interaction between
 AD X BD TOC1 AD BD X PIFΔAD PIF TOC1 TOC1 PIFΔAD PIF N TOC1 TOC1 C1 PIFΔAD PIF C1 TOC1 TOC1 C PIFΔAD PIF C TOC1 Supplementary Figure 1: Yeast two-hybrid assays showing the interaction between PIF and TOC1
AD X BD TOC1 AD BD X PIFΔAD PIF TOC1 TOC1 PIFΔAD PIF N TOC1 TOC1 C1 PIFΔAD PIF C1 TOC1 TOC1 C PIFΔAD PIF C TOC1 Supplementary Figure 1: Yeast two-hybrid assays showing the interaction between PIF and TOC1
Supplementary Figure 1. Botrocetin induces binding of human VWF to human
 Supplementary Figure 1: Supplementary Figure 1. Botrocetin induces binding of human VWF to human platelets in the absence of elevated shear and induces platelet agglutination as detected by flow cytometry.
Supplementary Figure 1: Supplementary Figure 1. Botrocetin induces binding of human VWF to human platelets in the absence of elevated shear and induces platelet agglutination as detected by flow cytometry.
Surrogate reporter-based enrichment of cells containing RNA-guided Cas9 nucleaseinduced
 Supplementary Data Surrogate reporter-based enrichment of cells containing RNA-guided Cas9 nucleaseinduced mutations Suresh Ramakrishna 1, Seung Woo Cho 2, Sojung Kim 2, Myungjae Song 1, Ramu Gopalappa
Supplementary Data Surrogate reporter-based enrichment of cells containing RNA-guided Cas9 nucleaseinduced mutations Suresh Ramakrishna 1, Seung Woo Cho 2, Sojung Kim 2, Myungjae Song 1, Ramu Gopalappa
Mechanism of Transcription Termination by RNA Polymerase III Utilizes a Non-template Strand Sequence-Specific Signal Element
 Molecular Cell Supplemental Information Mechanism of ranscription ermination by RNA Polymerase III Utilizes a Non-template Strand Sequence-Specific Signal Element Aneeshkumar G. Arimbasseri and Richard
Molecular Cell Supplemental Information Mechanism of ranscription ermination by RNA Polymerase III Utilizes a Non-template Strand Sequence-Specific Signal Element Aneeshkumar G. Arimbasseri and Richard
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description:
 File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. dcas9-mq1 fusion protein induces de novo
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. dcas9-mq1 fusion protein induces de novo
TITLE: Engineering Anti-EGFR Antibodies for Treatment of Breast Cancers with Poor Prognosis
 AD Award Number: W81XWH-07-1-0499 TITLE: Engineering Anti-EGFR Antibodies for Treatment of Breast Cancers with Poor Prognosis PRINCIPAL INVESTIGATOR: James D. Marks, M.D., Ph.D. Yu Zhou, Ph.D. CONTRACTING
AD Award Number: W81XWH-07-1-0499 TITLE: Engineering Anti-EGFR Antibodies for Treatment of Breast Cancers with Poor Prognosis PRINCIPAL INVESTIGATOR: James D. Marks, M.D., Ph.D. Yu Zhou, Ph.D. CONTRACTING
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11070 Supplementary Figure 1 Purification of FLAG-tagged proteins. a, Purification of FLAG-RNF12 by FLAG-affinity from nuclear extracts of wild-type (WT) and two FLAG- RNF12 transgenic
doi:10.1038/nature11070 Supplementary Figure 1 Purification of FLAG-tagged proteins. a, Purification of FLAG-RNF12 by FLAG-affinity from nuclear extracts of wild-type (WT) and two FLAG- RNF12 transgenic
A synthetic recombinase-based feedback loop results in robust expression Supporting Information
 A synthetic recombinase-based feedback loop results in robust expression Supporting Information 1 Fig. S1: Control experiments performed to demonstrate the expected function of our closed-loop feedback
A synthetic recombinase-based feedback loop results in robust expression Supporting Information 1 Fig. S1: Control experiments performed to demonstrate the expected function of our closed-loop feedback
Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana.
 SUPPLEMENTARY FIGURES Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana. YFP:, :CFP, HA: and (A) and HA, CFP and YFPtagged and AVR1-CO39 (B) were expressed
SUPPLEMENTARY FIGURES Figure S1. Immunoblotting showing proper expression of tagged R and AVR proteins in N. benthamiana. YFP:, :CFP, HA: and (A) and HA, CFP and YFPtagged and AVR1-CO39 (B) were expressed
Supplementary Figure 1. Supporting data for Figure 1
 Supplementary Figure 1 Supporting data for Figure 1 (A) Schematic of BLI assay used to measure dissociation. 5 monobiotinylated substrate DNA (identical to λ1, Figure 2) is associated with streptavidin-coated
Supplementary Figure 1 Supporting data for Figure 1 (A) Schematic of BLI assay used to measure dissociation. 5 monobiotinylated substrate DNA (identical to λ1, Figure 2) is associated with streptavidin-coated
Structural basis of a novel PD-L1 nanobody for immune checkpoint blockade
 Structural basis of a novel PD-L1 nanobody for immune checkpoint blockade Supplementary Materials Supplementary methods Table S1-S Figure S1-S 1 1 1 1 1 1 1 1 1 0 1 0 Supplementary Methods Competitive
Structural basis of a novel PD-L1 nanobody for immune checkpoint blockade Supplementary Materials Supplementary methods Table S1-S Figure S1-S 1 1 1 1 1 1 1 1 1 0 1 0 Supplementary Methods Competitive
Table S1. List of DNA constructs and primers, part 1 Construct
 SUPPLEMENTARY TABLES: Table S1. List of DNA constructs and primers, part 1 Construct Comment (Expressed protein) Vector, PCR primers and template or source of Restriction Sites name Resistance construct
SUPPLEMENTARY TABLES: Table S1. List of DNA constructs and primers, part 1 Construct Comment (Expressed protein) Vector, PCR primers and template or source of Restriction Sites name Resistance construct
Orthogonal Ribosome Biofirewall
 1 2 3 4 5 Supporting Information Orthogonal Ribosome Biofirewall Bin Jia,, Hao Qi,, Bing-Zhi Li,, Shuo pan,, Duo Liu,, Hong Liu,, Yizhi Cai, and Ying-Jin Yuan*, 6 7 8 9 10 11 12 Key Laboratory of Systems
1 2 3 4 5 Supporting Information Orthogonal Ribosome Biofirewall Bin Jia,, Hao Qi,, Bing-Zhi Li,, Shuo pan,, Duo Liu,, Hong Liu,, Yizhi Cai, and Ying-Jin Yuan*, 6 7 8 9 10 11 12 Key Laboratory of Systems
Technical tips Session 4
 Technical tips Session 4 Biotinylation assay: Streptavidin is a small bacterial protein that binds with high affinity to the vitamin biotin. This streptavidin-biotin combination can be used to link molecules
Technical tips Session 4 Biotinylation assay: Streptavidin is a small bacterial protein that binds with high affinity to the vitamin biotin. This streptavidin-biotin combination can be used to link molecules
Supplemental Figure 1
 Supplemental Figure 1 Supplemental Fig. 1. (A) The binding avidity between Fz4-FcH6 and MBP-Norrin dimers measured by biolayer interferometry. 5 µg/ml Fz4-Fc protein was loaded onto anti human Fc capture
Supplemental Figure 1 Supplemental Fig. 1. (A) The binding avidity between Fz4-FcH6 and MBP-Norrin dimers measured by biolayer interferometry. 5 µg/ml Fz4-Fc protein was loaded onto anti human Fc capture
p53 increases MHC class I expression by upregulating the endoplasmic reticulum aminopeptidase ERAP1
 Supplementary Information p53 increases MHC class I expression by upregulating the endoplasmic reticulum aminopeptidase ERAP1 Bei Wang 1, Dandan Niu 1, Liyun Lai 1, & Ee Chee Ren 1,2 1 Singapore Immunology
Supplementary Information p53 increases MHC class I expression by upregulating the endoplasmic reticulum aminopeptidase ERAP1 Bei Wang 1, Dandan Niu 1, Liyun Lai 1, & Ee Chee Ren 1,2 1 Singapore Immunology
