pass TV Backbone Assay pass Homozygous Loss of WT Allele (LOA) SR- pass 5' Cassette Integrity pass
|
|
|
- Vivien Curtis
- 5 years ago
- Views:
Transcription
1 Gene: Dennd1c Colony prefix: MEKG ESC clone ID: EPD0440_3_A10 Allele: Dennd1c tm1a(eucomm)wtsi Allele type: Knockout First, Reporter-tagged insertion with conditiol potential Allele information: Further information about the allele can be found on the IKMC web site at Details on how to determine the floxed exon can be found at Mouse QC information Southern Blot TV Backbone Assay 5' LR-PCR Loss of WT Allele (LOA) qpcr Homozygous Loss of WT Allele (LOA) SR- Neo Count (qpcr) LacZ SR-PCR 5' Cassette Integrity Neo SR-PCR Mutant Specific SR- PCR LoxP Confirmation 3' LR-PCR Genotyping Comment This technical data sheet and information ("Datasheet") is supplied by ("GRL"). Report Generated on: 09-SEP :22:08 Page 1 of 6
2 Southern blot confirmation: Southern blots are not routinely performed at the Sanger Institute due to throughput constraints. A southern blot experiment design tool can be found on the IKMC web site at Links to information and frequently asked questions about the EUCOMM/KOMP alleles and MGP projects General targeting strategies: MGP mouse phenotype data: IKMC allele types: MGP mouse quality control tests : Allele conversion guide - genotyping tm1b, tm1c and tm1d mice: How the "critical" exon is decided: Genotyping Information Genotyping by end-point PCR These mice may be genotyped through a combition of separate PCR reactions that detect the cassette, the gene-specific wild type allele, and a mutant allele-specific short range PCR. Interpretation of the consolidated results produces the genotype of the mice. For example: cassette positive, mutant positive, wild type positive = heterozygous. This technical data sheet and information ("Datasheet") is supplied by ("GRL"). Report Generated on: 09-SEP :22:08 Page 2 of 6
3 PCRs primer pairs and expected size bands Assay Type Assay Forward Primer Reverse Primer Expected Size Band (bp) Standard PCR Wildtype Dennd1c_83347_F Dennd1c_83347_R 485 Standard PCR Mutant Dennd1c_83347_F CAS_R1_Term 304 Standard PCR Cassette LacZ_2_small_F LacZ_2_small_R 108 Primer sequences Reaction setup Primer Name CAS_R1_Term Dennd1c_83347_F Dennd1c_83347_R LacZ_2_small_F LacZ_2_small_R Reagent µl DNA (~ ng) 1 10x Buffer 2 MgCl2 (50 mm) 0.6 Platinum Taq (Invitrogen) 0.2 dntps (100 mm) 0.2 Primer 1 (10 mm) 0.4 Primer 2 (10 mm) 0.4 ddh Total 20 Primer Sequence (5' > 3') TCGTGGTATCGTTATGCGCC CAAGTGCAGGGGGTTAGAGG TCAGCCAGGTGTAGTGGTGC ATCACGACGCGCTGTATC ACATCGGGCAAATAATATCG Amplification conditions Step Conditions Time 1 94 C 5 min 2 94 C 30 sec 3 58 C 30 sec 4 72 C 45 sec 5 Go to '2' + 34 cycles C 5 min 7 12 C forever This technical data sheet and information ("Datasheet") is supplied by ("GRL"). Report Generated on: 09-SEP :22:08 Page 3 of 6
4 Genotyping using universal copy number qpcr assays designed to the selection cassette The cassette qpcr assays use a hydrolysis probe assay (eg Applied Biosystems TaqMan technology) to determine genotype via the copy number of the selection cassette in a sample. Homozygotes will possess two copies, heterozygotes one copy and wild type mice will show no amplification when compared to known homozygote controls. These FAM -labeled assays are multiplexed with a VIC labeled endogenous control assay (for example TaqMan Copy Number Reference Assay, Mouse, Tfrc; Applied Biosystems part # ). Please note that these assays are not gene-specific other information should be used in conjunction with the universal cassette assays (for example the mutant-specific srpcr) when confirming the gene identity. Primer type Assay Name Forward Primer Seq. Reverse Primer Seq. Probe Primer Seq. Cassette Neo GGTGGAGAGGCTATTCGGC GAACACGGCGGCATCAG TGGGCACAACAGACAATCGGCT G Reactions are performed in a 10µl volume using an Applied Biosystems 7900HT Fast Real-Time PCR System or Applied Biosystems Viia7 with DNA prepared using the Sample-to-SNP TM kit (Applied Biosystems) from mouse ear biopsies. GTXpress TM buffer is also used (Applied Biosystems). Reagent µl 2x GTXpress TM buffer 5 20x target assay 0.5 ddh2o 3 Tfrc endogenous 20x assay 0.5 DNA 1 Amplification conditions Step Conditions Time 1 95 C 20 sec 2 95 C 10 sec 3 60 C 30 sec 4 Go to '2' + 34 cycles - This technical data sheet and information ("Datasheet") is supplied by ("GRL"). Report Generated on: 09-SEP :22:09 Page 4 of 6
5 Genotyping by loss of WT allele qpcr Assay (gene-specific assay) The wild type loss of allele (LoA) qpcr assay uses a hydrolysis probe assay (for example Applied Biosystems TaqMan technology) to determine the copy number of the wild type allele in a sample. Homozygotes will show no amplification, heterozygotes one copy and wild type mice will show two copies when compared to a wild type control. The number of copies of the Dennd1c allele can be detected using a FAM-labelled custom qpcr TaqMan assay. These are multiplexed with a VIC labelled endogenous control assay (for example TaqMan Copy Number Reference Assay, Mouse, Tfrc; Applied Biosystems part # ). Reference DNA controls of known genotypes should also be included to facilitate correct alysis. Primers for LoA qpcr assay Primer type Assay Name Forward Primer Seq. Reverse Primer Seq. Probe Primer Seq. LoA DENND1C_WT CGAGTGCTCTACTCATTGAGCTA AGT TGCCCCTCTGGAAGAGAAGA CGGCTGGAAAGTT Reaction setup Reaction setup and amplification conditions are the same as those used for the neo cassette qpcr assay. This technical data sheet and information ("Datasheet") is supplied by ("GRL"). Report Generated on: 09-SEP :22:09 Page 5 of 6
6 Relevant publications Ryder, E., Gleeson, D., Sethi, D., Vyas, S., Miklejewska, E., Dalvi, P., Habib, B., Cook, R., Hardy, M., Jhaveri, K., et al. (2013). Molecular Characterization of Mutant Mouse Strains Generated from the EUCOMM/KOMP-CSD ES Cell Resource. Mammalian Genome. Doi: /s x White, J.K., Gerdin, A.-K., Karp, N.A., Ryder, E., Buljan, M., Bussell, J.N., Salisbury, J., Clare, S., Ingham, N.J., Podrini, C., et al. (2013). Genome-wide Generation and Systematic Phenotyping of Knockout Mice Reveals New Roles for Many Genes. Cell 154, Ryder, E., Wong, K., Gleeson, D., Keane, T.M., Sethi, D., Vyas, S., Wardle-Jones, H., Bussell, J.N., Houghton, R., Salisbury, J., et al. (2013). Genomic alysis of a novel spontaneous albino C57BL/6N mouse strain. Genesis 51, Bradley, A., Astassiadis, K., Ayadi, A., Battey, J.F., Bell, C., Birling, M.-C., Bottomley, J., Brown, S.D., Bürger, A., Bult, C.J., et al. (2012). The mammalian gene function resource: the intertiol knockout mouse consortium. Mamm Genome 23, Birling, M.-C., Dierich, A., Jacquot, S., Hérault, Y., and Pavlovic, G. (2011). Highly-efficient, fluorescent, locus directed Cre and flpo deleter mice on a pure C57BL/6N genetic background. Genesis. Skarnes, W.C., Rosen, B., West, A.P., Koutsourakis, M., Bushell, W., Iyer, V., Mujica, A.O., Thomas, M., Harrow, J., Cox, T., et al. (2011). A conditiol knockout resource for the genome-wide study of mouse gene function. Nature 474, Pettitt, S.J., Liang, Q., Rairdan, X.Y., Moran, J.L., Prosser, H.M., Beier, D.R., Lloyd, K.C., Bradley, A., and Skarnes, W.C. (2009). Agouti C57BL/6N embryonic stem cells for mouse genetic resources. Nat Methods 6, Liang, Q., Conte, N., Skarnes, W.C., and Bradley, A. (2008). Extensive genomic copy number variation in embryonic stem cells. Proc Natl Acad Sci U S A 105, Farley, F.W., Soriano, P., Steffen, L.S., and Dymecki, S.M. (2000). Widespread recombise expression using FLPeR (flipper) mice. Genesis 28, This technical data sheet and information ("Datasheet") is supplied by ("GRL"). Report Generated on: 09-SEP :22:09 Page 6 of 6
Ramp1 EPD0843_4_B11. EUCOMM/KOMP-CSD Knockout-First Genotyping
 EUCOMM/KOMP-CSD Knockout-First Genotyping Introduction The majority of animals produced from the EUCOMM/KOMP-CSD ES cell resource contain the Knockout-First-Reporter Tagged Insertion allele. As well as
EUCOMM/KOMP-CSD Knockout-First Genotyping Introduction The majority of animals produced from the EUCOMM/KOMP-CSD ES cell resource contain the Knockout-First-Reporter Tagged Insertion allele. As well as
Usp14 EPD0582_2_G09. EUCOMM/KOMP-CSD Knockout-First Genotyping
 EUCOMM/KOMP-CSD Knockout-First Genotyping Introduction The majority of animals produced from the EUCOMM/KOMP-CSD ES cell resource contain the Knockout-First-Reporter Tagged Insertion allele. As well as
EUCOMM/KOMP-CSD Knockout-First Genotyping Introduction The majority of animals produced from the EUCOMM/KOMP-CSD ES cell resource contain the Knockout-First-Reporter Tagged Insertion allele. As well as
Agouti C57BL/6N embryonic stem cells for mouse genetic resources
 nature methods Agouti N embryonic stem cells for mouse genetic resources Stephen J Pettitt, Qi Liang, Xin Y Rairdan, Jennifer L Moran, Haydn M Prosser, David R Beier, Kent Lloyd, Allan Bradley & William
nature methods Agouti N embryonic stem cells for mouse genetic resources Stephen J Pettitt, Qi Liang, Xin Y Rairdan, Jennifer L Moran, Haydn M Prosser, David R Beier, Kent Lloyd, Allan Bradley & William
GENOTYPING BY PCR PROTOCOL FORM MUTANT MOUSE REGIONAL RESOURCE CENTER North America, International
 Please provide the following information required for genetic analysis of your mutant mice. Please fill in form electronically by tabbing through the text fields. The first 2 pages are protected with gray
Please provide the following information required for genetic analysis of your mutant mice. Please fill in form electronically by tabbing through the text fields. The first 2 pages are protected with gray
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10163 Supplementary Table 1 Efficiency of vector construction. Process wells recovered efficiency (%) Recombineering* 480 461 96 Intermediate plasmids 461 381 83 Recombineering efficiency
doi:10.1038/nature10163 Supplementary Table 1 Efficiency of vector construction. Process wells recovered efficiency (%) Recombineering* 480 461 96 Intermediate plasmids 461 381 83 Recombineering efficiency
A Low salt diet. C Low salt diet + mf4-31c1 3. D High salt diet + mf4-31c1 3. B High salt diet
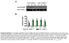 A Low salt diet GV [AU]:. [mmhg]: 0..... 09 9 7 B High salt diet GV [AU]:. [mmhg]: 7 8...0. 7.8 8.. 0 8 7 C Low salt diet + mf-c GV [AU]:. [mmhg]:.0.9 8.7.7. 7 8 0 D High salt diet + mf-c E Lymph capillary
A Low salt diet GV [AU]:. [mmhg]: 0..... 09 9 7 B High salt diet GV [AU]:. [mmhg]: 7 8...0. 7.8 8.. 0 8 7 C Low salt diet + mf-c GV [AU]:. [mmhg]:.0.9 8.7.7. 7 8 0 D High salt diet + mf-c E Lymph capillary
GENOME 371, Problem Set 6
 GENOME 371, Problem Set 6 1. S. pombe is a distant relative of baker s yeast (which you used in quiz section). Wild type S. pombe can grow on plates lacking tryptophan (-trp plates). A mutant has been
GENOME 371, Problem Set 6 1. S. pombe is a distant relative of baker s yeast (which you used in quiz section). Wild type S. pombe can grow on plates lacking tryptophan (-trp plates). A mutant has been
TRANSGENIC ANIMALS. -transient transfection of cells -stable transfection of cells. - Two methods to produce transgenic animals:
 TRANSGENIC ANIMALS -transient transfection of cells -stable transfection of cells - Two methods to produce transgenic animals: 1- DNA microinjection - random insertion 2- embryonic stem cell-mediated gene
TRANSGENIC ANIMALS -transient transfection of cells -stable transfection of cells - Two methods to produce transgenic animals: 1- DNA microinjection - random insertion 2- embryonic stem cell-mediated gene
Analysis of gene function
 Genome 371, 22 February 2010, Lecture 12 Analysis of gene function Gene knockouts PHASE TWO: INTERPRETATION I THINK I FOUND A CORNER PIECE. 3 BILLION PIECES Analysis of a disease gene Gene knockout or
Genome 371, 22 February 2010, Lecture 12 Analysis of gene function Gene knockouts PHASE TWO: INTERPRETATION I THINK I FOUND A CORNER PIECE. 3 BILLION PIECES Analysis of a disease gene Gene knockout or
TRANSPOSON INSERTION SITE VERIFICATION
 TRANSPOSON INSERTION SITE VERIFICATION Transposon and T-DNA insertion in Arabidopsis genes can be identified using the Arabidopsis thaliana Insertion Database (ATIdb) (http://atidb.org/cgi-perl/gbrowse/atibrowse).
TRANSPOSON INSERTION SITE VERIFICATION Transposon and T-DNA insertion in Arabidopsis genes can be identified using the Arabidopsis thaliana Insertion Database (ATIdb) (http://atidb.org/cgi-perl/gbrowse/atibrowse).
SUPPLEMENTAL MATERIAL
 SUPPLEMENTAL MATERIAL MATERIALS AND METHODS Generation of TSPOΔ/Δ murine embryonic fibroblasts Embryos were harvested from 13.5-day pregnant TSPOfl/fl mice. After dissection to eviscerate and remove the
SUPPLEMENTAL MATERIAL MATERIALS AND METHODS Generation of TSPOΔ/Δ murine embryonic fibroblasts Embryos were harvested from 13.5-day pregnant TSPOfl/fl mice. After dissection to eviscerate and remove the
Module 3: IKMC Resource Overview
 Overview Aims Understanding knockout design strategies Introduction to the IKMC web portal Introduction Several large-scale gene targeting and gene trapping projects are participating in the International
Overview Aims Understanding knockout design strategies Introduction to the IKMC web portal Introduction Several large-scale gene targeting and gene trapping projects are participating in the International
(i) A trp1 mutant cell took up a plasmid containing the wild type TRP1 gene, which allowed that cell to multiply and form a colony
 1. S. pombe is a distant relative of baker s yeast (which you used in quiz section). Wild type S. pombe can grow on plates lacking tryptophan (-trp plates). A mutant has been isolated that cannot grow
1. S. pombe is a distant relative of baker s yeast (which you used in quiz section). Wild type S. pombe can grow on plates lacking tryptophan (-trp plates). A mutant has been isolated that cannot grow
Schematic representation of the endogenous PALB2 locus and gene-disruption constructs
 Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
Supplementary Figures Supplementary Figure 1. Generation of PALB2 -/- and BRCA2 -/- /PALB2 -/- DT40 cells. (A) Schematic representation of the endogenous PALB2 locus and gene-disruption constructs carrying
TRANSGENIC TECHNOLOGIES: Gene-targeting
 TRANSGENIC TECHNOLOGIES: Gene-targeting Reverse Genetics Wild-type Bmp7 -/- Forward Genetics Phenotype Gene or Mutations First Molecular Analysis Second Reverse Genetics Gene Phenotype or Molecular Analysis
TRANSGENIC TECHNOLOGIES: Gene-targeting Reverse Genetics Wild-type Bmp7 -/- Forward Genetics Phenotype Gene or Mutations First Molecular Analysis Second Reverse Genetics Gene Phenotype or Molecular Analysis
High-throughput genotyping of CRISPR/Cas9-mediated mutants using fluorescent
 High-throughput genotyping of CRISPR/Cas9-mediated mutants using fluorescent PCR-capillary gel electrophoresis Muhammad Khairul RAMLEE, Tingdong YAN, Alice M. S. CHEUNG, Charles CHUAH, Shang LI Figure
High-throughput genotyping of CRISPR/Cas9-mediated mutants using fluorescent PCR-capillary gel electrophoresis Muhammad Khairul RAMLEE, Tingdong YAN, Alice M. S. CHEUNG, Charles CHUAH, Shang LI Figure
Name Order # 1234 Company Received 03/15/16 Reported 03/16/16. Sample
 Well Name Order # 1234 Company Received 03/15/16 E-Mail Reported 03/16/16 Strain Name Sample Name Flp Cyp2r 1 Cre A1 Strain A 1 Wild Het A2 Strain A 2 Wild Het A3 Strain A 3 Wild Het A4 Strain A 4 Wild
Well Name Order # 1234 Company Received 03/15/16 E-Mail Reported 03/16/16 Strain Name Sample Name Flp Cyp2r 1 Cre A1 Strain A 1 Wild Het A2 Strain A 2 Wild Het A3 Strain A 3 Wild Het A4 Strain A 4 Wild
Applied Biosystems Real-Time PCR Rapid Assay Development Guidelines
 Applied Biosystems Real-Time PCR Rapid Assay Development Guidelines Description This tutorial will discuss recommended guidelines for designing and running real-time PCR quantification and SNP Genotyping
Applied Biosystems Real-Time PCR Rapid Assay Development Guidelines Description This tutorial will discuss recommended guidelines for designing and running real-time PCR quantification and SNP Genotyping
CFTR-null wt CFTR-null 1.0. Probe: Neo R. Figure S1
 A. B. 4.0 3.0 2.0 1.0 4.0 3.0 2.0 1.0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 Probe: Neo R CFTR-null wt CFTR-null Figure S1 A. 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 10kb 8kb CFTR-null wt B. Probe: CFTR
A. B. 4.0 3.0 2.0 1.0 4.0 3.0 2.0 1.0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 Probe: Neo R CFTR-null wt CFTR-null Figure S1 A. 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 10kb 8kb CFTR-null wt B. Probe: CFTR
TaqPath ProAmp Master Mixes
 PRODUCT BULLETIN es es Applied Biosystems TaqPath ProAmp Master Mixes are versatile master mixes developed for high-throughput genotyping and copy number variation (CNV) analysis protocols that require
PRODUCT BULLETIN es es Applied Biosystems TaqPath ProAmp Master Mixes are versatile master mixes developed for high-throughput genotyping and copy number variation (CNV) analysis protocols that require
Molecular Biology and Functional Genomic Core Facility
 Molecular Biology and Functional Genomic Core Facility General Presentation Dr Odile Neyret Core Manager Myriam Rondeau Research Assistant Agnès Dumont Research Assistant Institut de recherche clinique
Molecular Biology and Functional Genomic Core Facility General Presentation Dr Odile Neyret Core Manager Myriam Rondeau Research Assistant Agnès Dumont Research Assistant Institut de recherche clinique
Detection of FactorV Leiden (R506Q)
 TM Primerdesign Ltd Detection of FactorV Leiden (R506Q) snpsig TM real-time PCR mutation detection/allelic discrimination kit 50 tests For general laboratory and research use only 1 Kit contents Factor
TM Primerdesign Ltd Detection of FactorV Leiden (R506Q) snpsig TM real-time PCR mutation detection/allelic discrimination kit 50 tests For general laboratory and research use only 1 Kit contents Factor
 http://fire.biol.wwu.edu/trent/trent/direct_detection_of_genotype.html 1 Like most other model organism Arabidopsis thaliana has a sequenced genome? What do we mean by sequenced genome? What sort of info
http://fire.biol.wwu.edu/trent/trent/direct_detection_of_genotype.html 1 Like most other model organism Arabidopsis thaliana has a sequenced genome? What do we mean by sequenced genome? What sort of info
Supplementary Methods
 Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
Supplementary Methods Reverse transcribed Quantitative PCR. Total RNA was isolated from bone marrow derived macrophages using RNeasy Mini Kit (Qiagen), DNase-treated (Promega RQ1), and reverse transcribed
The RRPA knock-in allele was generated by homologous recombination in TC1 ES cells.
 Supplemental Materials Materials & Methods Generation of RRPA and RAPA Knock-in Mice The RRPA knock-in allele was generated by homologous recombination in TC1 ES cells. Targeted ES clones in which the
Supplemental Materials Materials & Methods Generation of RRPA and RAPA Knock-in Mice The RRPA knock-in allele was generated by homologous recombination in TC1 ES cells. Targeted ES clones in which the
White Paper: High Throughput SNP Genotyping Using Array Tape in Place of Microplates
 White Paper High Throughput SNP Genotyping White Paper: High Throughput SNP Genotyping Using Array Tape in Place of Microplates Miniaturization and Automation Using a Novel New Reaction Substrate Originally
White Paper High Throughput SNP Genotyping White Paper: High Throughput SNP Genotyping Using Array Tape in Place of Microplates Miniaturization and Automation Using a Novel New Reaction Substrate Originally
Plasma EGFR T790M ctdna status is associated with clinical outcome in. advanced NSCLC patients with acquired EGFR-TKI resistance
 Plasma EGFR T790M ctdna status is associated with clinical outcome in advanced NSCLC patients with acquired EGFR-TKI resistance 1# D Zheng; 2# X Ye; 3 MZ Zhang; 2 Y Sun; 1 JY Wang; 1 J Ni; 1 HP Zhang;
Plasma EGFR T790M ctdna status is associated with clinical outcome in advanced NSCLC patients with acquired EGFR-TKI resistance 1# D Zheng; 2# X Ye; 3 MZ Zhang; 2 Y Sun; 1 JY Wang; 1 J Ni; 1 HP Zhang;
Procomcure Biotech. PCR Reagents
 Procomcure Biotech PCR Reagents valid for 2018 VitaTaq DNA Polymerase is a standard Taq DNA polymerase suitable for all common PCR applications like colonypcr, cloning applications, high-throughput PCR
Procomcure Biotech PCR Reagents valid for 2018 VitaTaq DNA Polymerase is a standard Taq DNA polymerase suitable for all common PCR applications like colonypcr, cloning applications, high-throughput PCR
Incorporating SeqStudio Genetic Analyzer and Sanger sequencing into genome editing workflows
 Incorporating SeqStudio Genetic Analyzer and Sanger sequencing into genome editing workflows Stephen Jackson, Ph.D 27 May 2017 The world leader in serving science Key Applications for Genome Editing Research
Incorporating SeqStudio Genetic Analyzer and Sanger sequencing into genome editing workflows Stephen Jackson, Ph.D 27 May 2017 The world leader in serving science Key Applications for Genome Editing Research
FastGene Optima. Products for PCR. proof-reading. ReadyMix. cdna SNP. routine PCR. polymerase. optimized blend complex templates. N gene knock out B O
 Products for PCR www.nippongenetics.eu efficiency polymerase convenience endpoint PCR incl. loading dye direct PCR incl. dntps engineered enzyme archeal type B polymerase high GC content FastGene tissuebest
Products for PCR www.nippongenetics.eu efficiency polymerase convenience endpoint PCR incl. loading dye direct PCR incl. dntps engineered enzyme archeal type B polymerase high GC content FastGene tissuebest
A novel tool for monitoring endogenous alpha-synuclein transcription by NanoLuciferase
 A novel tool for monitoring endogenous alpha-synuclein transcription by NanoLuciferase tag insertion at the 3 end using CRISPR-Cas9 genome editing technique Sambuddha Basu 1, 3, Levi Adams 1, 3, Subhrangshu
A novel tool for monitoring endogenous alpha-synuclein transcription by NanoLuciferase tag insertion at the 3 end using CRISPR-Cas9 genome editing technique Sambuddha Basu 1, 3, Levi Adams 1, 3, Subhrangshu
QIAGEN Whole Genome Amplification REPLI-g Eliminating Sample Limitations, Potential Use for Reference Material
 QIAGEN Whole Genome Amplification REPLI-g Eliminating Sample Limitations, Potential Use for Reference Material MDA Technology Protocols Locus Bias Applications Kits & Service Summary QIAGEN REPLI-g WGA
QIAGEN Whole Genome Amplification REPLI-g Eliminating Sample Limitations, Potential Use for Reference Material MDA Technology Protocols Locus Bias Applications Kits & Service Summary QIAGEN REPLI-g WGA
A) (5 points) As the starting step isolate genomic DNA from
 GS Final Exam Spring 00 NAME. bub ts is a recessive temperature sensitive mutation in yeast. At º C bub ts cells grow normally, but at º C they die. Use the information below to clone the wild-type BUB
GS Final Exam Spring 00 NAME. bub ts is a recessive temperature sensitive mutation in yeast. At º C bub ts cells grow normally, but at º C they die. Use the information below to clone the wild-type BUB
Loss of Native Allele Protocol for KOMP-Regeneron derived ESC lines January 9 th 2009
 Loss of Native Allele Protocol for KOMP-Regeneron derived ESC lines January 9 th 2009 I. Alkaline DNA extraction in 96 well format Solutions 50mM NaOH 1M Tris-HCl ph 8.0 1. Grow ES cells in a 96-well TC
Loss of Native Allele Protocol for KOMP-Regeneron derived ESC lines January 9 th 2009 I. Alkaline DNA extraction in 96 well format Solutions 50mM NaOH 1M Tris-HCl ph 8.0 1. Grow ES cells in a 96-well TC
Revised: RG-RV2 by Fukuhara et al.
 Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
Supplemental Figure 1 The generation of Spns2 conditional knockout mice. (A) Schematic representation of the wild type Spns2 locus (Spns2 + ), the targeted allele, the floxed allele (Spns2 f ) and the
Supplementary Information. The flowering gene SINGLE FLOWER TRUSS drives heterosis for yield in tomato
 Supplementary Information The flowering gene SINGLE FLOWER TRUSS drives heterosis for yield in tomato Uri Krieger 1, Zachary B. Lippman 2 *, and Dani Zamir 1 * 1. The Hebrew University of Jerusalem Faculty
Supplementary Information The flowering gene SINGLE FLOWER TRUSS drives heterosis for yield in tomato Uri Krieger 1, Zachary B. Lippman 2 *, and Dani Zamir 1 * 1. The Hebrew University of Jerusalem Faculty
FACTOR V of LEIDEN REAL TIME
 When you have completed this, the graph will show all of the wells identified as shown in the legend. Homozygous WT Negative Heterozygous Homozygous MUT FACTOR V of LEIDEN REAL TIME REF. 05960127 Opening
When you have completed this, the graph will show all of the wells identified as shown in the legend. Homozygous WT Negative Heterozygous Homozygous MUT FACTOR V of LEIDEN REAL TIME REF. 05960127 Opening
Percent survival. Supplementary fig. S3 A.
 Supplementary fig. S3 A. B. 100 Percent survival 80 60 40 20 Ml 0 0 100 C. Fig. S3 Comparison of leukaemia incidence rate in the triple targeted chimaeric mice and germline-transmission translocator mice
Supplementary fig. S3 A. B. 100 Percent survival 80 60 40 20 Ml 0 0 100 C. Fig. S3 Comparison of leukaemia incidence rate in the triple targeted chimaeric mice and germline-transmission translocator mice
Tamiflu resistance detection kit
 Tamiflu resistance detection kit H1N1 (H275Y) genesig real-time PCR mutation detection/allelic discrimination kit 150 tests For general laboratory and research use only genesig Tamiflu resistance detection
Tamiflu resistance detection kit H1N1 (H275Y) genesig real-time PCR mutation detection/allelic discrimination kit 150 tests For general laboratory and research use only genesig Tamiflu resistance detection
High-Resolution Melting analysis as a tool for rapid and sensitive detection of genotypes in cattle population
 Research and Development Station for Bovine, Arad, Romania High-Resolution Melting analysis as a tool for rapid and sensitive detection of genotypes in cattle population Daniela Elena Ilie, Ada Cean, Ioan
Research and Development Station for Bovine, Arad, Romania High-Resolution Melting analysis as a tool for rapid and sensitive detection of genotypes in cattle population Daniela Elena Ilie, Ada Cean, Ioan
US Large Mutant Mouse facilities (KOMP and MMRRC)
 US Large Mutant Mouse facilities (KOMP and MMRRC) Session 4: Biological and Medical Sciences (14:00-16:00) Kent Lloyd CNR - Piazzale Aldo Moro, Rome, Italy 01 October 2010 US measures ~6million mice used
US Large Mutant Mouse facilities (KOMP and MMRRC) Session 4: Biological and Medical Sciences (14:00-16:00) Kent Lloyd CNR - Piazzale Aldo Moro, Rome, Italy 01 October 2010 US measures ~6million mice used
Applied Biosystems 7500 Fast, 7500 and 7300 Real-Time PCR Systems
 PRODUCT BROCHURE Real-Time PCR Systems Applied Biosystems 7500 Fast, 7500 and 7300 Real-Time PCR Systems Real Fast. Real Versatile. Real Value. Real choices from the leader in real-time PCR. The latest
PRODUCT BROCHURE Real-Time PCR Systems Applied Biosystems 7500 Fast, 7500 and 7300 Real-Time PCR Systems Real Fast. Real Versatile. Real Value. Real choices from the leader in real-time PCR. The latest
FMV 35S promoter (GMO) Advanced Kit. FMV 35S promoter. 150 tests. For general laboratory and research use only
 FMV 35S promoter (GMO) FMV 35S promoter 150 tests Advanced Kit For general laboratory and research use only Introduction to FMV 35S promoter (GMO) The kit provides a method for detecting gene insertion
FMV 35S promoter (GMO) FMV 35S promoter 150 tests Advanced Kit For general laboratory and research use only Introduction to FMV 35S promoter (GMO) The kit provides a method for detecting gene insertion
Supplementary Methods pcfd5 cloning protocol
 Supplementary Methods cloning protocol vermilion trna grna trna grna U6:3 Terminator AmpR attb is a vector for expressing one or multiple trna-flanked Cas9 grnas under the control of the strong, ubiquitous
Supplementary Methods cloning protocol vermilion trna grna trna grna U6:3 Terminator AmpR attb is a vector for expressing one or multiple trna-flanked Cas9 grnas under the control of the strong, ubiquitous
species- Mus musculus Engineering the mouse genome David Ornitz
 species- Mus musculus Engineering the mouse genome David Ornitz How do we analyze gene function in mice? Gene addition (transgenic approach) Permits GOF, DN and knockdown experiments Ectopic (spatial or
species- Mus musculus Engineering the mouse genome David Ornitz How do we analyze gene function in mice? Gene addition (transgenic approach) Permits GOF, DN and knockdown experiments Ectopic (spatial or
oasig TM lyophilised 2X qpcr Mastermix
 oasig TM lyophilised 2X qpcr Mastermix Instructions for use of Primerdesign oasig lyophilised Mastermix MM MasterMix Contents Introduction 3 Kit Contents 4 Kit Storage 4 Suitable Sample Material 4 Licensing
oasig TM lyophilised 2X qpcr Mastermix Instructions for use of Primerdesign oasig lyophilised Mastermix MM MasterMix Contents Introduction 3 Kit Contents 4 Kit Storage 4 Suitable Sample Material 4 Licensing
INTERNATIONAL UNION FOR THE PROTECTION OF NEW VARIETIES OF PLANTS GENEVA
 E BMT Guidelines (proj.4) ORIGINAL: English DATE: December 21, 2005 INTERNATIONAL UNION FOR THE PROTECTION OF NEW VARIETIES OF PLANTS GENEVA GUIDELINES FOR DNA-PROFILING: MOLECULAR MARKER SELECTION AND
E BMT Guidelines (proj.4) ORIGINAL: English DATE: December 21, 2005 INTERNATIONAL UNION FOR THE PROTECTION OF NEW VARIETIES OF PLANTS GENEVA GUIDELINES FOR DNA-PROFILING: MOLECULAR MARKER SELECTION AND
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Ihh interacts preferentially with its upstream neighboring gene Nhej1. Genes are indicated by gray lines, and Ihh and Nhej1 are highlighted in blue. 4C seq performed in E14.5 limbs
Supplementary Figure 1 Ihh interacts preferentially with its upstream neighboring gene Nhej1. Genes are indicated by gray lines, and Ihh and Nhej1 are highlighted in blue. 4C seq performed in E14.5 limbs
Puro. Knockout Detection (KOD) Kit
 Puro Knockout Detection (KOD) Kit Cat. No. CC-03 18 Oct. 2016 Contents I. Kit Contents and Storage II. Product Overview III. Methods Experimental Outline Genomic DNA Preparation Obtain Hybrid DNA Digest
Puro Knockout Detection (KOD) Kit Cat. No. CC-03 18 Oct. 2016 Contents I. Kit Contents and Storage II. Product Overview III. Methods Experimental Outline Genomic DNA Preparation Obtain Hybrid DNA Digest
Single-Cell Whole Transcriptome Profiling With the SOLiD. System
 APPLICATION NOTE Single-Cell Whole Transcriptome Profiling Single-Cell Whole Transcriptome Profiling With the SOLiD System Introduction The ability to study the expression patterns of an individual cell
APPLICATION NOTE Single-Cell Whole Transcriptome Profiling Single-Cell Whole Transcriptome Profiling With the SOLiD System Introduction The ability to study the expression patterns of an individual cell
EGFR T790M mutation mutation detection by quantitative allele specific amplification (quasa) EGFR (T790M)
 Primerdesign TM Ltd EGFR T790M mutation mutation detection by quantitative allele specific amplification (quasa) EGFR (T790M) 50 tests For general laboratory and research use only 1 Contents Introduction
Primerdesign TM Ltd EGFR T790M mutation mutation detection by quantitative allele specific amplification (quasa) EGFR (T790M) 50 tests For general laboratory and research use only 1 Contents Introduction
Agouti C57BL/6N embryonic stem cells for mouse genetic resources
 Europe PMC Funders Group Author Manuscript Published in final edited form as: Nat Methods. 2009 July ; 6(7): 493 495. doi:10.1038/nmeth.1342. Agouti C57BL/6N embryonic stem cells for mouse genetic resources
Europe PMC Funders Group Author Manuscript Published in final edited form as: Nat Methods. 2009 July ; 6(7): 493 495. doi:10.1038/nmeth.1342. Agouti C57BL/6N embryonic stem cells for mouse genetic resources
QUANTITATIVE RT-PCR PROTOCOL (SYBR Green I) (Last Revised: April, 2007)
 QUANTITATIVE RT-PCR PROTOCOL (SYBR Green I) (Last Revised: April, 007) Please contact Center for Plant Genomics (CPG) facility manager Hailing Jin (hljin@iastate.edu) regarding questions or corrections.
QUANTITATIVE RT-PCR PROTOCOL (SYBR Green I) (Last Revised: April, 007) Please contact Center for Plant Genomics (CPG) facility manager Hailing Jin (hljin@iastate.edu) regarding questions or corrections.
TaqMan SNP Genotyping
 TaqMan SNP Genotyping What is Sickle-cell Anemia? Healthy Cells Hemoglobin Heme + 2 α & 2 β- globin molecules Heme binds O 2 2 Sickle-shaped cells Mutation causes hemoglobin to cluster together when O
TaqMan SNP Genotyping What is Sickle-cell Anemia? Healthy Cells Hemoglobin Heme + 2 α & 2 β- globin molecules Heme binds O 2 2 Sickle-shaped cells Mutation causes hemoglobin to cluster together when O
We designed the targeting vector in such a way that the first exon was flanked by two LoxP sites and
 Mice We designed the targeting vector in such a way that the first exon was flanked by two LoxP sites and an Frt flanked neo cassette (Fig. S1A). Targeted BA1 (C57BL/6 129/SvEv) hybrid embryonic stem cells
Mice We designed the targeting vector in such a way that the first exon was flanked by two LoxP sites and an Frt flanked neo cassette (Fig. S1A). Targeted BA1 (C57BL/6 129/SvEv) hybrid embryonic stem cells
Nature Medicine doi: /nm.2548
 Supplementary Table 1: Genotypes of offspring and embryos from matings of Pmm2 WT/F118L mice with Pmm2 WT/R137H mice total events Pmm2 WT/WT Pmm2 WT/R137H Pmm2 WT/F118L Pmm2 R137H/F118L offspring 117 (100%)
Supplementary Table 1: Genotypes of offspring and embryos from matings of Pmm2 WT/F118L mice with Pmm2 WT/R137H mice total events Pmm2 WT/WT Pmm2 WT/R137H Pmm2 WT/F118L Pmm2 R137H/F118L offspring 117 (100%)
Applications and Uses. (adapted from Roche RealTime PCR Application Manual)
 What Can You Do With qpcr? Applications and Uses (adapted from Roche RealTime PCR Application Manual) What is qpcr? Real time PCR also known as quantitative PCR (qpcr) measures PCR amplification as it
What Can You Do With qpcr? Applications and Uses (adapted from Roche RealTime PCR Application Manual) What is qpcr? Real time PCR also known as quantitative PCR (qpcr) measures PCR amplification as it
Detection of CFTR(F508) MyBioSource.com. Real-time PCR mutation detection/allelic discrimination kit. For general laboratory and research use only
 Detection of CFTR(F508) Real-time PCR mutation detection/allelic discrimination kit For general laboratory and research use only 50 tests Contents Principles of the test 3 Kit Contents 5 Recommended Accompanying
Detection of CFTR(F508) Real-time PCR mutation detection/allelic discrimination kit For general laboratory and research use only 50 tests Contents Principles of the test 3 Kit Contents 5 Recommended Accompanying
Enzyme that uses RNA as a template to synthesize a complementary DNA
 Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Comparison of two or more protein or DNA sequence to ascertain similarities in sequences. If two genes have
Nature Biotechnology: doi: /nbt.4166
 Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
Supplementary Figure 1 Validation of correct targeting at targeted locus. (a) by immunofluorescence staining of 2C-HR-CRISPR microinjected embryos cultured to the blastocyst stage. Embryos were stained
NEW PARADIGM of BIOTECHNOLOGY - GENET BIO. GeNet Bio Global Gene Network
 NEW PARADIGM of BIOTECHNOLOGY - GENET BIO GeNet Bio Global Gene Network GENET BIO DNA AMPLIFICATION PRODUCTS GUIDE Keynote of Products Prime TaqTM DNA Polymerase Prime TaqTM Premix ExPrime TaqTM DNA Polymerase
NEW PARADIGM of BIOTECHNOLOGY - GENET BIO GeNet Bio Global Gene Network GENET BIO DNA AMPLIFICATION PRODUCTS GUIDE Keynote of Products Prime TaqTM DNA Polymerase Prime TaqTM Premix ExPrime TaqTM DNA Polymerase
Advances and New Technologies in Forensic DNA
 Advances and New Technologies in Forensic DNA Nader Pourmand, PhD Department of Biomolecular Engineering, University of California, Santa Cruz Branch Migratio 3 nbp 6 3 5 3 5 bp 14 bp 23 5 3 5 3 5 Bioluminescence
Advances and New Technologies in Forensic DNA Nader Pourmand, PhD Department of Biomolecular Engineering, University of California, Santa Cruz Branch Migratio 3 nbp 6 3 5 3 5 bp 14 bp 23 5 3 5 3 5 Bioluminescence
SUPPLEMENT MATERIALS FOR CURTIN,
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 SUPPLEMENT MATERIALS FOR CURTIN, et al. Validating genome-wide association candidates: Selecting, testing, and characterizing
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 SUPPLEMENT MATERIALS FOR CURTIN, et al. Validating genome-wide association candidates: Selecting, testing, and characterizing
BSCI410-Liu/Spring 06 Exam #1 Feb. 23, 06
 Your Name: Your UID# 1. (20 points) Match following mutations with corresponding mutagens (X-RAY, Ds transposon excision, UV, EMS, Proflavin) a) Thymidine dimmers b) Breakage of DNA backbone c) Frameshift
Your Name: Your UID# 1. (20 points) Match following mutations with corresponding mutagens (X-RAY, Ds transposon excision, UV, EMS, Proflavin) a) Thymidine dimmers b) Breakage of DNA backbone c) Frameshift
Genotyping Manual. KASP version 4.0. P Robinson Dr J Holme
 Genotyping Manual 0 g KASP version 4.0 SNP Genotyping Manual P Robinson Dr J Holme 26 th May 2011 Contents KASP version 4.0 SNP Genotyping Manual Introduction Improvements of KASP version 4.0 Principal
Genotyping Manual 0 g KASP version 4.0 SNP Genotyping Manual P Robinson Dr J Holme 26 th May 2011 Contents KASP version 4.0 SNP Genotyping Manual Introduction Improvements of KASP version 4.0 Principal
Experimental genetics - 2 Partha Roy
 Partha Roy Experimental genetics - 2 Making genetically altered animal 1) Gene knock-out k from: a) the entire animal b) selected cell-type/ tissue c) selected cell-type/tissue at certain time 2) Transgenic
Partha Roy Experimental genetics - 2 Making genetically altered animal 1) Gene knock-out k from: a) the entire animal b) selected cell-type/ tissue c) selected cell-type/tissue at certain time 2) Transgenic
Rapid Cycle PCR, Real Time Analysis, and Hi-Res Melting
 Rapid Cycle PCR, Real Time Analysis, and Hi-Res Melting Carl Wittwer Department of Pathology University of Utah ARUP Idaho Technology AMP, Oct. 31, 2008 Impatient, Lazy, and Cheap Rapid Cycle PCR Fast
Rapid Cycle PCR, Real Time Analysis, and Hi-Res Melting Carl Wittwer Department of Pathology University of Utah ARUP Idaho Technology AMP, Oct. 31, 2008 Impatient, Lazy, and Cheap Rapid Cycle PCR Fast
TRANSGENIC ANIMALS. transient. stable. - Two methods to produce transgenic animals:
 Only for teaching purposes - not for reproduction or sale CELL TRANSFECTION transient stable TRANSGENIC ANIMALS - Two methods to produce transgenic animals: 1- DNA microinjection 2- embryonic stem cell-mediated
Only for teaching purposes - not for reproduction or sale CELL TRANSFECTION transient stable TRANSGENIC ANIMALS - Two methods to produce transgenic animals: 1- DNA microinjection 2- embryonic stem cell-mediated
Supplementary Table 3d: PCR primers for genomic sequencing-based phasereconstruction
 Supplementary Table 3: Detailed lists of grnas, primers, donor constructs, antibodies, Gene Expression Omnibus (GEO) identifiers for Chip-seq and DHSs datasets and CRISPR/Cas9 genome edited cell lines
Supplementary Table 3: Detailed lists of grnas, primers, donor constructs, antibodies, Gene Expression Omnibus (GEO) identifiers for Chip-seq and DHSs datasets and CRISPR/Cas9 genome edited cell lines
Supplementary Figure 1. CRISPR/Cas9-induced targeted mutations in TaGASR7, TaDEP1, TaNAC2, TaPIN1, TaLOX2 and TaGW2 genes in wheat protoplasts.
 Supplementary Figure 1. CRISPR/Cas9-induced targeted mutations in TaGASR7, TaDEP1, TaNAC2, TaPIN1, TaLOX2 and TaGW2 genes in wheat protoplasts. Lanes 1 and 2: digested CRISPR/Cas9-transformed protoplasts;
Supplementary Figure 1. CRISPR/Cas9-induced targeted mutations in TaGASR7, TaDEP1, TaNAC2, TaPIN1, TaLOX2 and TaGW2 genes in wheat protoplasts. Lanes 1 and 2: digested CRISPR/Cas9-transformed protoplasts;
GENETICS - CLUTCH CH.15 GENOMES AND GENOMICS.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
!! www.clutchprep.com CONCEPT: OVERVIEW OF GENOMICS Genomics is the study of genomes in their entirety Bioinformatics is the analysis of the information content of genomes - Genes, regulatory sequences,
TripleHyb real time PCR for detection of single nucleotide polymorphisms in the VEGF promoter region
 TripleHyb real time PCR for detection of single nucleotide polymorphisms in the VEGF promoter region S. Fuessel 1, S. Unversucht 1, A. Lohse 1, S. Tomasetti 1, A. Rost 2, A. Edelmann 2, M.P. Wirth 1, A.
TripleHyb real time PCR for detection of single nucleotide polymorphisms in the VEGF promoter region S. Fuessel 1, S. Unversucht 1, A. Lohse 1, S. Tomasetti 1, A. Rost 2, A. Edelmann 2, M.P. Wirth 1, A.
JAK2 V617F mutation mutation detection by quantitative allele specific amplification (quasa) JAK2(V617F)
 Primerdesign TM Ltd JAK2 V617F mutation mutation detection by quantitative allele specific amplification (quasa) JAK2(V617F) 50 tests For general laboratory and research use only 1 Contents Introduction
Primerdesign TM Ltd JAK2 V617F mutation mutation detection by quantitative allele specific amplification (quasa) JAK2(V617F) 50 tests For general laboratory and research use only 1 Contents Introduction
Easi CRISPR for conditional and insertional alleles
 Easi CRISPR for conditional and insertional alleles C.B Gurumurthy, University Of Nebraska Medical Center Omaha, NE cgurumurthy@unmc.edu Types of Genome edits Gene disruption/inactivation Types of Genome
Easi CRISPR for conditional and insertional alleles C.B Gurumurthy, University Of Nebraska Medical Center Omaha, NE cgurumurthy@unmc.edu Types of Genome edits Gene disruption/inactivation Types of Genome
Biology 105: Introduction to Genetics PRACTICE FINAL EXAM Part I: Definitions. Homology: Reverse transcriptase. Allostery: cdna library
 Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Reverse transcriptase Allostery: cdna library Transformation Part II Short Answer 1. Describe the reasons for
Biology 105: Introduction to Genetics PRACTICE FINAL EXAM 2006 Part I: Definitions Homology: Reverse transcriptase Allostery: cdna library Transformation Part II Short Answer 1. Describe the reasons for
QuantiNova proven compatibility with various real-time cyclers in gene expression analysis
 QuantiNova proven compatibility with various real-time cyclers in gene expression analysis QuantiNova SYBR Green and QuantiNova Probe PCR Kits were used on various real-time cyclers, and compared with
QuantiNova proven compatibility with various real-time cyclers in gene expression analysis QuantiNova SYBR Green and QuantiNova Probe PCR Kits were used on various real-time cyclers, and compared with
Supplemental Materials and Methods. Yeast strains. Strain genotypes are listed in Table S1. To generate MATainc::LEU2,
 Supplemental Materials and Methods Yeast strains. Strain genotypes are listed in Table S1. To generate MATainc::LEU2, the LEU2 marker was inserted by PCR-mediated recombination (BRACHMANN et al. 1998)
Supplemental Materials and Methods Yeast strains. Strain genotypes are listed in Table S1. To generate MATainc::LEU2, the LEU2 marker was inserted by PCR-mediated recombination (BRACHMANN et al. 1998)
10x Hot-Taq Buffer 1 ml 1 ml x 2ea 1 ml x 10ea. dntp Mix (each 10 mm) None / 0.2 ml None / 0.4 ml None / 0.4 ml x 5ea
 HelixAmp TM Hot-Taq Polymerase (Ver. 2.0) CERTIFICATE OF ANALYSIS (1603-V01R03) Contents HelixAmp TM Hot-Taq Polymerase (Ver. 2.0) Cat. No. HT250/HT250N HT500/HT500N HT2500/HT2500N Hot-Taq (2.5 units/μl)
HelixAmp TM Hot-Taq Polymerase (Ver. 2.0) CERTIFICATE OF ANALYSIS (1603-V01R03) Contents HelixAmp TM Hot-Taq Polymerase (Ver. 2.0) Cat. No. HT250/HT250N HT500/HT500N HT2500/HT2500N Hot-Taq (2.5 units/μl)
BRAF V600E mutation mutation detection by quantitative allele specific amplification (quasa) BRAF (V600E)
 Primerdesign TM Ltd BRAF V600E mutation mutation detection by quantitative allele specific amplification (quasa) BRAF (V600E) 50 tests For general laboratory and research use only 1 Contents Introduction
Primerdesign TM Ltd BRAF V600E mutation mutation detection by quantitative allele specific amplification (quasa) BRAF (V600E) 50 tests For general laboratory and research use only 1 Contents Introduction
TaqMan Assays for genetic variation research. Superior performance reliable, robust solutions
 TaqMan Assays for genetic variation research Superior performance reliable, robust solutions Genetic variation: decoding the blueprint for biodiversity Research on genetic variation in animals and plants
TaqMan Assays for genetic variation research Superior performance reliable, robust solutions Genetic variation: decoding the blueprint for biodiversity Research on genetic variation in animals and plants
Soya Round up Ready (RR)
 Primerdesign TM Ltd GMO event quantification Soya Round up Ready (RR) Detection and quantification of GMO integration events by real-time PCR 100 tests Kit contents Soya-WT DNA primer/probe mix (100 reactions
Primerdesign TM Ltd GMO event quantification Soya Round up Ready (RR) Detection and quantification of GMO integration events by real-time PCR 100 tests Kit contents Soya-WT DNA primer/probe mix (100 reactions
Mouse Engineering Technology. Musculoskeletal Research Center 2016 Summer Educational Series David M. Ornitz Department of Developmental Biology
 Mouse Engineering Technology Musculoskeletal Research Center 2016 Summer Educational Series David M. Ornitz Department of Developmental Biology Core service and new technologies Mouse ES core Discussions
Mouse Engineering Technology Musculoskeletal Research Center 2016 Summer Educational Series David M. Ornitz Department of Developmental Biology Core service and new technologies Mouse ES core Discussions
CaMV promoter & NOS terminator detection
 Primerdesign TM Ltd GMO event quantification CaMV promoter & NOS terminator detection Detection and quantification of GMO integration events in Soya by real-time PCR 100 tests Kit contents CaMV-GM primer/probe
Primerdesign TM Ltd GMO event quantification CaMV promoter & NOS terminator detection Detection and quantification of GMO integration events in Soya by real-time PCR 100 tests Kit contents CaMV-GM primer/probe
LightCycler 480 qpcr Tools. Meeting the Challenge of Your Research
 LightCycler 480 qpcr Tools Meeting the Challenge of Your Research Find the Optimal LightCycler 480 Reagents for Your Research Application: Are you analyzing DNA DNA Nucleic acid isolation Manual processing
LightCycler 480 qpcr Tools Meeting the Challenge of Your Research Find the Optimal LightCycler 480 Reagents for Your Research Application: Are you analyzing DNA DNA Nucleic acid isolation Manual processing
Molecular LDT in Newborn Screening Laboratories
 Molecular LDT in Newborn Screening Laboratories APHL/CDC Newborn Screening Molecular Workshop Atlanta, GA June 28-30, 2011 Mei Baker, M.D., FACMG Assistant Professor, Department of Pediatrics University
Molecular LDT in Newborn Screening Laboratories APHL/CDC Newborn Screening Molecular Workshop Atlanta, GA June 28-30, 2011 Mei Baker, M.D., FACMG Assistant Professor, Department of Pediatrics University
User Manual. Catalog No.: DWSK-V101 (10 rxns), DWSK-V102 (25 rxns) For Research Use Only
 DNA Walking SpeedUp TM Kit SpeedUp Sequencing SpeedUp BAC Clone Sequencing SpeedUp Genome Walking SpeedUp Transgene Location Detection SpeedUp Deletion/ Insertion/ Isoform Detection User Manual Version
DNA Walking SpeedUp TM Kit SpeedUp Sequencing SpeedUp BAC Clone Sequencing SpeedUp Genome Walking SpeedUp Transgene Location Detection SpeedUp Deletion/ Insertion/ Isoform Detection User Manual Version
Using mutants to clone genes
 Using mutants to clone genes Objectives: 1. What is positional cloning? 2. What is insertional tagging? 3. How can one confirm that the gene cloned is the same one that is mutated to give the phenotype
Using mutants to clone genes Objectives: 1. What is positional cloning? 2. What is insertional tagging? 3. How can one confirm that the gene cloned is the same one that is mutated to give the phenotype
Melton, D.W. (1994) Gene targeting in the mouse. Bioessays 16:633-8
 Reverse genetics - Knockouts Paper to read for this section : Melton, D.W. (1994) Gene targeting in the mouse. Bioessays 16:633-8 Up until now, we ve concentrated on ways to get a cloned gene. We ve seen
Reverse genetics - Knockouts Paper to read for this section : Melton, D.W. (1994) Gene targeting in the mouse. Bioessays 16:633-8 Up until now, we ve concentrated on ways to get a cloned gene. We ve seen
SUPPLEMENTARY INFORMATION SUPPLEMENTARY FIGURES
 SUPPLEMENTARY INFORMATION SUPPLEMENTARY FIGURES Supplementary Figure 1. Generation of inducible BICD2 knock-out mice. A) The mouse BICD2 locus and gene targeting constructs. To generate an inducible Bicd2
SUPPLEMENTARY INFORMATION SUPPLEMENTARY FIGURES Supplementary Figure 1. Generation of inducible BICD2 knock-out mice. A) The mouse BICD2 locus and gene targeting constructs. To generate an inducible Bicd2
NOS terminator (GMO) genesig Advanced Kit. NOS terminator. 150 tests. Primerdesign Ltd. For general laboratory and research use only
 TM Primerdesign Ltd NOS terminator (GMO) NOS terminator genesig Advanced Kit 150 tests For general laboratory and research use only 1 Introduction to NOS terminator (GMO) The kit provides a method for
TM Primerdesign Ltd NOS terminator (GMO) NOS terminator genesig Advanced Kit 150 tests For general laboratory and research use only 1 Introduction to NOS terminator (GMO) The kit provides a method for
Protocol CRISPR Genome Editing In Cell Lines
 Protocol CRISPR Genome Editing In Cell Lines Protocol 2: HDR donor plasmid applications (gene knockout, gene mutagenesis, gene tagging, Safe Harbor ORF knock-in) Notes: 1. sgrna validation: GeneCopoeia
Protocol CRISPR Genome Editing In Cell Lines Protocol 2: HDR donor plasmid applications (gene knockout, gene mutagenesis, gene tagging, Safe Harbor ORF knock-in) Notes: 1. sgrna validation: GeneCopoeia
R1 12 kb R1 4 kb R1. R1 10 kb R1 2 kb R1 4 kb R1
 Bcor101 Sample questions Midterm 3 1. The maps of the sites for restriction enzyme EcoR1 (R1) in the wild type and mutated cystic fibrosis genes are shown below: Wild Type R1 12 kb R1 4 kb R1 _ _ CF probe
Bcor101 Sample questions Midterm 3 1. The maps of the sites for restriction enzyme EcoR1 (R1) in the wild type and mutated cystic fibrosis genes are shown below: Wild Type R1 12 kb R1 4 kb R1 _ _ CF probe
A NOTE ON LINKAGE BETWEEN THE ANGORA AND fgf5 GENES IN RABBITS
 W ORLD R ABBIT SCIENCE World Rabbit Sci. 2004, 2: - 6 WRSA, UPV, 2003 A NOTE ON LINKAGE BETWEEN THE ANGORA AND fgf5 GENES IN RABBITS MULSANT P., ROCHAMBEAU H. DE, THÉBAULT R. G. * INRA, Laboratoire de
W ORLD R ABBIT SCIENCE World Rabbit Sci. 2004, 2: - 6 WRSA, UPV, 2003 A NOTE ON LINKAGE BETWEEN THE ANGORA AND fgf5 GENES IN RABBITS MULSANT P., ROCHAMBEAU H. DE, THÉBAULT R. G. * INRA, Laboratoire de
Molecular Markers CRITFC Genetics Workshop December 9, 2014
 Molecular Markers CRITFC Genetics Workshop December 9, 2014 Molecular Markers Tools that allow us to collect information about an individual, a population, or a species Application in fisheries mating
Molecular Markers CRITFC Genetics Workshop December 9, 2014 Molecular Markers Tools that allow us to collect information about an individual, a population, or a species Application in fisheries mating
Functional Genomics Research Stream. Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update
 Functional Genomics Research Stream Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update Updates Alternate Lab Meeting Fridays 11:30-1:00 WEL 4.224 Welcome to attend either one Lab Log thanks
Functional Genomics Research Stream Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update Updates Alternate Lab Meeting Fridays 11:30-1:00 WEL 4.224 Welcome to attend either one Lab Log thanks
TaqMan Sample-to-SNP Kit
 Quick Reference Card TaqMan Sample-to-SNP Kit Note: For safety and biohazard guidelines, refer to the Safety appendix in the Applied Biosystems TaqMan Sample-to-SNP Kit Protocol (PN 4402136). For all chemicals
Quick Reference Card TaqMan Sample-to-SNP Kit Note: For safety and biohazard guidelines, refer to the Safety appendix in the Applied Biosystems TaqMan Sample-to-SNP Kit Protocol (PN 4402136). For all chemicals
Detection of Biological Threat Agents by Real-Time PCR - Comparison of Assay
 Detection of Biological Threat Agents by Real-Time PCR - Comparison of Assay Performance on the Idaho Technology, Inc. R.A.P.I.D., the Roche LightCycler, and the Cepheid Smart Cycler Supplemental Data
Detection of Biological Threat Agents by Real-Time PCR - Comparison of Assay Performance on the Idaho Technology, Inc. R.A.P.I.D., the Roche LightCycler, and the Cepheid Smart Cycler Supplemental Data
Protocols for cell lines using CRISPR/CAS
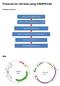 Protocols for cell lines using CRISPR/CAS Procedure overview Map Preparation of CRISPR/CAS plasmids Expression vectors for guide RNA (U6-gRNA) and Cas9 gene (CMV-p-Cas9) are ampicillin-resist ant and stable
Protocols for cell lines using CRISPR/CAS Procedure overview Map Preparation of CRISPR/CAS plasmids Expression vectors for guide RNA (U6-gRNA) and Cas9 gene (CMV-p-Cas9) are ampicillin-resist ant and stable
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Schwarz UI, Ritchie MD, Bradford Y, et al. Genetic determinants
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Schwarz UI, Ritchie MD, Bradford Y, et al. Genetic determinants
SUPPLEMENTARY INFORMATION
 Gene replacements and insertions in rice by intron targeting using CRISPR Cas9 Table of Contents Supplementary Figure 1. sgrna-induced targeted mutations in the OsEPSPS gene in rice protoplasts. Supplementary
Gene replacements and insertions in rice by intron targeting using CRISPR Cas9 Table of Contents Supplementary Figure 1. sgrna-induced targeted mutations in the OsEPSPS gene in rice protoplasts. Supplementary
