SUPPLEMENTARY INFORMATION
|
|
|
- Arthur Henry
- 5 years ago
- Views:
Transcription
1 Substantial and robust changes in microrna transcriptome support postnatal development of the hypothalamus in rat Soraya Doubi-Kadmiri 1#, Charlotte Benoit 1,2#, Xavier Benigni 1,3, Guillaume Beaumont 1,4, Claire-Marie Vacher 1, Mohammed Taouis 1, Anne Baroin- Tourancheau 1 and Laurence Amar 1* 1 Université Paris-Saclay, Université Paris-Sud, CNRS, UMR 9197-Institut des Neurosciences Paris-Saclay, Orsay F-91405, France 2 Present address : Nodea Medical, 1 mail du Professeur Georges Mathé, Villejuif F-94800, France 3 Present address : Institute of Plant Sciences - Paris-Saclay, 2 Rue Gaston Crémieux, Evry F-91000, France 4 Present address : Laboratoire de Bioinformatique, Centre National de Génotypage, Commissariat à l Énergie Atomique, 2 rue Gaston Crémieux, CP5721, Evry F-91057, France # These authors contributed equally to this work SUPPLEMENTARY INFORMATION * Corresponding author : laurence.amar@u-psud.fr 1
2 A Rela(ve Ra(o 1,E+01' 1,E+00' 1,E$01' 1,E$02' 1,E$03' 1,E$04' P4 P8 P14 P21 P28 PVN B Rela(ve Ra(o 1,E+01' 1,E+00' 1,E$01' 1,E$02' 1,E$03' 1,E$04' P4 P8 P14 P21 P28 HF- PROGENY Figure S1. Assessment of ARC/MEs. (A) POMC transcripts were quantified relatively to GAPDH transcripts in ARC/MEs at stages P4, P8, P14, P21 or P28 by using RT-qPCRs. PVNs of 3 adults were used as POMC non-expressing controls. The relative ratio of POMC to GAPDH mrnas of one ARC/ME at stage P28 (identified by a blue hatched bar) was taken as the reference ratio of 1. Note that the Y-axis is drawn using a log(10) scale. PVNs displayed relative ratios lower than The relative ratio of (10 fold higher than the highest PVN relative ratio) was therefore taken as the threshold value for assessing ARC/ME. This threshold is drawn as a dashed red line. Three ARC/MEs of stage P4 and 1 of stage P8 were discarded. One ARC/ME of P28 which ratio was much lower than the other values of the P28 sample (lower than mean minus 1.7 standard deviation) was also discarded. (B) The same analysis was performed on ARC/MEs of HF-progeny at stages P4, P8, P14, P21 orp28. Two ARC/MEs of stage P4 and 3 of stage P14 were discarded. 2
3 A P4 P8 P14 P21 Fold- Change P28 B P14.4 P21.4 Fold- Change Figure S2. Quantification of mirna expression homogeneity across biological replicates of ARC/ME. Levels of homogeneity were quantified by computing the coefficients of variation of expression (ratios of the standard deviation to the mean) either over all mirnas (global CVs) (see Figure 1) or over sliding windows of 50 mirnas of decreasing abundance (sliding CVs). Sliding CVs were plotted against mean expression of corresponding mirnas. (A) Sliding CVs at stages P4, P8, P14, P21 or P28. Bulge at stage P8 is the consequence of the high CV of mir-328-3p (see Supplemental Table S2). (B) Sliding CVs at sub-stages P14.4 or P
4 Plasma Lep(n in Dams (ng/ml) 20" 15" 10" 5" 0" P4" P8" P14" P21" P28" Postnatal Stage Figure S3. Dams of C- and HF-progeny display similar levels of plasma leptin during lactation. Data are shown as mean +- SEM. C- and HF-dams are identified by blue and red colors, respectively. 4
5 P4 P8 4HF1 4HF2 4HF3 8HF1 8HF2 8HF3 8HF4 8HF5 8HF6 8HF7 P14 P21 14HF1 14HF2 14HF3 14HF4 21HF1 21HF2 21HF3 21HF4 21HF5 21HF6 21HF7 P28 Fold-Change 28HF1 28HF2 28HF3 28HF4 28HF5 28HF6 28HF7 HF#PROGENY Stage n mirnas5>5105reads Global55CV5(mean5+#5SEM) P (+*(0.01 P (+*(0.02 P (+*(0.01 P (+*(0.01 P (+*(0.01 Figure S4. Analysis of mirna expression homogeneity across ARC/ME biological replicates in HF-progeny. Plots depict mirna relative expressions of ARC/MEs at P4, P8, P14, P21 orp28. In each plot, the X-axis denotes mean read counts between biological replicates, the Y-axis, fold changes of expression relatively to the mean expression. X- and Y-axes are drawn using a log(10) scale. Each dot represents the relative expression of an individual mirna expression and each color, a biological replicate. Dots of positive or negative values on the Y-axis denote up- or down-regulated mirna expressions relatively to mean expressions, respectively. Values on the Y-axis close to 1 denote mirnas of highly homogeneous expression. Many Y-values were distant from 1 for mirnas of low mean expression. This accounted for the fact that the lower the mean expression, the less precise its quantification. Coefficients of variations (standard deviation/ mean; CV) were calculated for each mirna (see Supplemental Table S5), and global CV (mean +/ SEM), for each stage. Table in bottom right corner summarizes data for each stage. 5
6 P4 P8 P14 P21 Fold- Change P28 Figure S5. Quantification of mirna expression homogeneity across biological replicates of ARC/ME in HF-progeny. Levels of homogeneity were quantified by computing the coefficients of variation of expression (ratios of the standard deviation to the mean) either over all mirnas (global CVs) (see Supplemental Figure S4) or over sliding windows of 50 mirnas of decreasing abundance (sliding CVs) at stages P4, P8, P14, P21 or P28. Sliding CVs were plotted against mean expression of corresponding mirnas. 6
7 Supplemental Tables S1 S7 are provided as individual.xls files. Table S1: Sequencing data. Total read counts, counts and percentages of reads >16 bases and <40 bases, counts and percentages of mir-related reads among reads >16 bases and <40 bases are indicated for each cdna library. Tables S2-S6. In each Table, mirnas are ranked by mean expression. Note that the denomination of mir-3p/mir-5p is currently replacing the denomination of mirna/mirna*. Denomination changes are in process so that one or the other denomination were used depending on the mirna. Table S2. mirna expression profiles in developing ARC/MEs. Table S3. Comparisons of mirna expressions at P4, P8, P14.4, or P21.4 versus P28. Table S4. Comparisons of mirna expressions between side-stages. Table S5. mirna expression profiles in developing ARC/MEs of HFprogeny. Table S6. Comparisons of mirna expressions at P4, P8, P14 P21 and P28 between ARC/MEs of C- and HF-progeny. Note that comparisons at stages P14 and P21 involved C-progeny of sub-stages P14.4 and P21.4. Table S7. Oligonucleotide Information. Oligonucleotides used for cdna library construction and specific RT-qPCR analyses are indicated. 7
reverse transcription! RT 1! RT 2! RT 3!
 Supplementary Figure! Entire workflow repeated 3 times! mirna! stock! -fold dilution series! to yield - copies! per µl in PCR reaction! reverse transcription! RT! pre-pcr! mastermix!!!! 3!!! 3! ddpcr!
Supplementary Figure! Entire workflow repeated 3 times! mirna! stock! -fold dilution series! to yield - copies! per µl in PCR reaction! reverse transcription! RT! pre-pcr! mastermix!!!! 3!!! 3! ddpcr!
QPCR ASSAYS FOR MIRNA EXPRESSION PROFILING
 TECH NOTE 4320 Forest Park Ave Suite 303 Saint Louis, MO 63108 +1 (314) 833-9764 mirna qpcr ASSAYS - powered by NAWGEN Our mirna qpcr Assays were developed by mirna experts at Nawgen to improve upon previously
TECH NOTE 4320 Forest Park Ave Suite 303 Saint Louis, MO 63108 +1 (314) 833-9764 mirna qpcr ASSAYS - powered by NAWGEN Our mirna qpcr Assays were developed by mirna experts at Nawgen to improve upon previously
Technical Review. Real time PCR
 Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Supplementary Information for Single-cell sequencing of the small-rna transcriptome
 Supplementary Information for Single-cell sequencing of the small-rna transcriptome Omid R. Faridani 1,6,*, Ilgar Abdullayev 1,2,6, Michael Hagemann-Jensen 1,3, John P. Schell 4, Fredrik Lanner 4,5 and
Supplementary Information for Single-cell sequencing of the small-rna transcriptome Omid R. Faridani 1,6,*, Ilgar Abdullayev 1,2,6, Michael Hagemann-Jensen 1,3, John P. Schell 4, Fredrik Lanner 4,5 and
Agilent GeneSpring GX 10: Beyond. Pam Tangvoranuntakul Product Manager, GeneSpring October 1, 2008
 Agilent GeneSpring GX 10: Gene Expression and Beyond Pam Tangvoranuntakul Product Manager, GeneSpring October 1, 2008 GeneSpring GX 10 in the News Our Goals for GeneSpring GX 10 Goal 1: Bring back GeneSpring
Agilent GeneSpring GX 10: Gene Expression and Beyond Pam Tangvoranuntakul Product Manager, GeneSpring October 1, 2008 GeneSpring GX 10 in the News Our Goals for GeneSpring GX 10 Goal 1: Bring back GeneSpring
MeCP2. MeCP2/α-tubulin. GFP mir1-1 mir132
 Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
Conservation Figure S1. Schematic showing 3 UTR (top; thick black line), mir132 MRE (arrow) and nucleotide sequence conservation (vertical black lines; http://genome.ucsc.edu). a GFP mir1-1 mir132 b GFP
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Supplementary Figure 1 Endogenous gene tagging to study subcellular localization and chromatin binding. a, b, Schematic of experimental set-up to endogenously tag RNAi factors using the CRISPR Cas9 technology,
Supplementary Figure 1. Details of the component modelling and steps in generating and validating a PTGS. The steps (boxes) and the flow of
 Supplementary Figure 1. Details of the component modelling and steps in generating and validating a PTGS. The steps (boxes) and the flow of information (arrows) through the modelling process, including
Supplementary Figure 1. Details of the component modelling and steps in generating and validating a PTGS. The steps (boxes) and the flow of information (arrows) through the modelling process, including
SUPPLEMENTARY INFORMATION. Massive release of extracellular vesicles from cancer cells after
 SUPPLEMENTARY INFORMATION Massive release of extracellular vesicles from cancer cells after photodynamic treatment or chemotherapy Kelly Aubertin a, Amanda K. A. Silva a*, Nathalie Luciani a, Ana Espinosa
SUPPLEMENTARY INFORMATION Massive release of extracellular vesicles from cancer cells after photodynamic treatment or chemotherapy Kelly Aubertin a, Amanda K. A. Silva a*, Nathalie Luciani a, Ana Espinosa
TaqMan Advanced mirna Assays
 PRODUCT BULLETIN TaqMan Advanced mirna Assays TaqMan Advanced mirna Assays Key features Universal reverse transcription (RT) one RT step for all Applied Biosystems TaqMan Advanced mirna Assays Sensitive
PRODUCT BULLETIN TaqMan Advanced mirna Assays TaqMan Advanced mirna Assays Key features Universal reverse transcription (RT) one RT step for all Applied Biosystems TaqMan Advanced mirna Assays Sensitive
Nature Biotechnology: doi: /nbt.4199
 Supplementary Figure 1 Base editing activities of engineered A3A-BE3 variants with mutations designed to disrupt non-specific interactions with substrate ssdna. Graphs illustrating the frequencies of C
Supplementary Figure 1 Base editing activities of engineered A3A-BE3 variants with mutations designed to disrupt non-specific interactions with substrate ssdna. Graphs illustrating the frequencies of C
Next-Generation Sequencing Gene Expression Analysis Using Agilent GeneSpring GX
 Next-Generation Sequencing Gene Expression Analysis Using Agilent GeneSpring GX Technical Overview Introduction RNA Sequencing (RNA-Seq) is one of the most commonly used next-generation sequencing (NGS)
Next-Generation Sequencing Gene Expression Analysis Using Agilent GeneSpring GX Technical Overview Introduction RNA Sequencing (RNA-Seq) is one of the most commonly used next-generation sequencing (NGS)
Supplementary Figures and Tables
 Genome assembly using Nanopore-guided Long and Error-free DNA Reads Mohammed-Amin Madoui 1 *, Stefan Engelen 1 *, Corinne Cruaud 1, Caroline Belser 1, Laurie Bertrand 1, Adriana Alberti 1, Arnaud Lemainque
Genome assembly using Nanopore-guided Long and Error-free DNA Reads Mohammed-Amin Madoui 1 *, Stefan Engelen 1 *, Corinne Cruaud 1, Caroline Belser 1, Laurie Bertrand 1, Adriana Alberti 1, Arnaud Lemainque
Simultaneous profiling of transcriptome and DNA methylome from a single cell
 Additional file 1: Supplementary materials Simultaneous profiling of transcriptome and DNA methylome from a single cell Youjin Hu 1, 2, Kevin Huang 1, 3, Qin An 1, Guizhen Du 1, Ganlu Hu 2, Jinfeng Xue
Additional file 1: Supplementary materials Simultaneous profiling of transcriptome and DNA methylome from a single cell Youjin Hu 1, 2, Kevin Huang 1, 3, Qin An 1, Guizhen Du 1, Ganlu Hu 2, Jinfeng Xue
APPLICATION NOTE. MagMAX mirvana Total RNA Isolation Kit TaqMan Advanced mirna Assays
 APPLICATION NOTE MagMAX mirvana Total RNA Isolation Kit TaqMan Advanced mirna Assays A complete workflow for high-throughput isolation of serum mirnas and downstream analysis by qrt-pcr: application for
APPLICATION NOTE MagMAX mirvana Total RNA Isolation Kit TaqMan Advanced mirna Assays A complete workflow for high-throughput isolation of serum mirnas and downstream analysis by qrt-pcr: application for
Supplemental Information. Boundary Formation through a Direct. Threshold-Based Readout. of Mobile Small RNA Gradients
 Developmental Cell, Volume 43 Supplemental Information Boundary Formation through a Direct Threshold-Based Readout of Mobile Small RNA Gradients Damianos S. Skopelitis, Anna H. Benkovics, Aman Y. Husbands,
Developmental Cell, Volume 43 Supplemental Information Boundary Formation through a Direct Threshold-Based Readout of Mobile Small RNA Gradients Damianos S. Skopelitis, Anna H. Benkovics, Aman Y. Husbands,
Supplementary material
 Supplementary material Manuscript Title Multiple effects of a commercial Roundup formulation on the soil filamentous fungus Aspergillus nidulans at low doses: evidence of an unexpected impact on energetic
Supplementary material Manuscript Title Multiple effects of a commercial Roundup formulation on the soil filamentous fungus Aspergillus nidulans at low doses: evidence of an unexpected impact on energetic
MicroRNA sequencing (mirnaseq)
 , Robust experimental design Data analysis using the CAP-miRSeq: A comprehensive analysis pipeline for deep microrna sequencing E. Starr Hazard Over view of this first lecture 1) Review of very basic mirna
, Robust experimental design Data analysis using the CAP-miRSeq: A comprehensive analysis pipeline for deep microrna sequencing E. Starr Hazard Over view of this first lecture 1) Review of very basic mirna
Fig. S1. Phenotypic analysis and assessment of meiotic silencing in Dgcr8 and Dicer cko males.
 Fig. S1. Phenotypic analysis and assessment of meiotic silencing in Dgcr8 and Dicer cko males. A-B Photographic representation of whole testes removed from Dgcr8 (A) and Dicer (B) litters (wild type (WT):
Fig. S1. Phenotypic analysis and assessment of meiotic silencing in Dgcr8 and Dicer cko males. A-B Photographic representation of whole testes removed from Dgcr8 (A) and Dicer (B) litters (wild type (WT):
RNA-Sequencing analysis
 RNA-Sequencing analysis Markus Kreuz 25. 04. 2012 Institut für Medizinische Informatik, Statistik und Epidemiologie Content: Biological background Overview transcriptomics RNA-Seq RNA-Seq technology Challenges
RNA-Sequencing analysis Markus Kreuz 25. 04. 2012 Institut für Medizinische Informatik, Statistik und Epidemiologie Content: Biological background Overview transcriptomics RNA-Seq RNA-Seq technology Challenges
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Distribution of mirnas between lncrna and protein-coding genes. Pie chart showing distribution of human mirna between protein coding and lncrna genes. To the right, lncrna mirna
Supplementary Figure 1 Distribution of mirnas between lncrna and protein-coding genes. Pie chart showing distribution of human mirna between protein coding and lncrna genes. To the right, lncrna mirna
TECH NOTE Pushing the Limit: A Complete Solution for Generating Stranded RNA Seq Libraries from Picogram Inputs of Total Mammalian RNA
 TECH NOTE Pushing the Limit: A Complete Solution for Generating Stranded RNA Seq Libraries from Picogram Inputs of Total Mammalian RNA Stranded, Illumina ready library construction in
TECH NOTE Pushing the Limit: A Complete Solution for Generating Stranded RNA Seq Libraries from Picogram Inputs of Total Mammalian RNA Stranded, Illumina ready library construction in
Supplementary Material for
 www.sciencemag.org/cgi/content/full/science.aaa6090/dc1 Supplementary Material for Spatially resolved, highly multiplexed RNA profiling in single cells Kok Hao Chen, Alistair N. Boettiger, Jeffrey R. Moffitt,
www.sciencemag.org/cgi/content/full/science.aaa6090/dc1 Supplementary Material for Spatially resolved, highly multiplexed RNA profiling in single cells Kok Hao Chen, Alistair N. Boettiger, Jeffrey R. Moffitt,
Better appreciation of true biological mirna expression differences using an improved version of the global mean normalization strategy
 Better appreciation of true biological mirna expression differences using an improved version of the global mean normalization strategy Jo Vandesompele professor, Ghent University co-founder and CEO, Biogazelle
Better appreciation of true biological mirna expression differences using an improved version of the global mean normalization strategy Jo Vandesompele professor, Ghent University co-founder and CEO, Biogazelle
TaqMan Advanced mirna Assays
 PRODUCT BULLETIN Key features Universal reverse transcription (RT) one RT step for all TaqMan Advanced mirna Assays Sensitive detect as few as 60 copies of input microrna (mirna) Specific detect only mature
PRODUCT BULLETIN Key features Universal reverse transcription (RT) one RT step for all TaqMan Advanced mirna Assays Sensitive detect as few as 60 copies of input microrna (mirna) Specific detect only mature
Measuring and Understanding Gene Expression
 Measuring and Understanding Gene Expression Dr. Lars Eijssen Dept. Of Bioinformatics BiGCaT Sciences programme 2014 Why are genes interesting? TRANSCRIPTION Genome Genomics Transcriptome Transcriptomics
Measuring and Understanding Gene Expression Dr. Lars Eijssen Dept. Of Bioinformatics BiGCaT Sciences programme 2014 Why are genes interesting? TRANSCRIPTION Genome Genomics Transcriptome Transcriptomics
Application Note Selective transcript depletion
 Application Note Selective transcript depletion Sample Authors Laura de Jager RED Scientist Michael Berry Bioinformatics Scientist Luke Esau RED Senior Scientist Ross Wadsworth RED Team Lead Roche Sequencing
Application Note Selective transcript depletion Sample Authors Laura de Jager RED Scientist Michael Berry Bioinformatics Scientist Luke Esau RED Senior Scientist Ross Wadsworth RED Team Lead Roche Sequencing
A comprehensive assessment of RNA-seq accuracy, reproducibility and information content by the Sequencing Quality Control Consortium
 A comprehensive assessment of RNA-seq accuracy, reproducibility and information content by the Sequencing Quality Control Consortium SEQC/MAQC-III Consortium* 214 Nature America, Inc. All rights reserved.
A comprehensive assessment of RNA-seq accuracy, reproducibility and information content by the Sequencing Quality Control Consortium SEQC/MAQC-III Consortium* 214 Nature America, Inc. All rights reserved.
Novel methods for RNA and DNA- Seq analysis using SMART Technology. Andrew Farmer, D. Phil. Vice President, R&D Clontech Laboratories, Inc.
 Novel methods for RNA and DNA- Seq analysis using SMART Technology Andrew Farmer, D. Phil. Vice President, R&D Clontech Laboratories, Inc. Agenda Enabling Single Cell RNA-Seq using SMART Technology SMART
Novel methods for RNA and DNA- Seq analysis using SMART Technology Andrew Farmer, D. Phil. Vice President, R&D Clontech Laboratories, Inc. Agenda Enabling Single Cell RNA-Seq using SMART Technology SMART
Evaluation of extraction kits and RT-qPCR systems adapted to high-throughput platform for
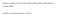 Evaluation of extraction kits and RT-qPCR systems adapted to high-throughput platform for circulating mirnas. Geok Wee Tan, Alan Soo Beng Khoo, Lu Ping Tan Supplementary Table S1. ICC (two-way mixed model
Evaluation of extraction kits and RT-qPCR systems adapted to high-throughput platform for circulating mirnas. Geok Wee Tan, Alan Soo Beng Khoo, Lu Ping Tan Supplementary Table S1. ICC (two-way mixed model
Q: Using at least 3 biological replicates in an experiment is recommended to do. What do you suggest: At which step of calculation of the relative
 The questions below have been asked by attendees of the qpcr webinar series, Part 2: Analyzing Your Data. All the questions, including the questions that could not be answered during the webinar have been
The questions below have been asked by attendees of the qpcr webinar series, Part 2: Analyzing Your Data. All the questions, including the questions that could not be answered during the webinar have been
Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency.
 Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency. (a) Transfection of different concentration of GAS5-overexpressing
Supplementary Figure 1, related to Figure 1. GAS5 is highly expressed in the cytoplasm of hescs, and positively correlates with pluripotency. (a) Transfection of different concentration of GAS5-overexpressing
Nature Methods: doi: /nmeth Supplementary Figure 1. Pilot CrY2H-seq experiments to confirm strain and plasmid functionality.
 Supplementary Figure 1 Pilot CrY2H-seq experiments to confirm strain and plasmid functionality. (a) RT-PCR on HIS3 positive diploid cell lysate containing known interaction partners AT3G62420 (bzip53)
Supplementary Figure 1 Pilot CrY2H-seq experiments to confirm strain and plasmid functionality. (a) RT-PCR on HIS3 positive diploid cell lysate containing known interaction partners AT3G62420 (bzip53)
OPTICAL CONTROL OF TUMOR INDUCTION IN THE ZEBRAFISH
 OPTICAL CONTROL OF TUMOR INDUCTION IN THE ZEBRAFISH Zhiping Feng, 1,12 Suzy Nam, 2 Fatima Hamouri, 3,4 Isabelle Aujard, 5,6 Bertrand Ducos, 3,4 Sophie Vriz, 7,8 Michel Volovitch, 7,9 Ludovic Jullien, 5,6
OPTICAL CONTROL OF TUMOR INDUCTION IN THE ZEBRAFISH Zhiping Feng, 1,12 Suzy Nam, 2 Fatima Hamouri, 3,4 Isabelle Aujard, 5,6 Bertrand Ducos, 3,4 Sophie Vriz, 7,8 Michel Volovitch, 7,9 Ludovic Jullien, 5,6
Development of quantitative targeted RNA-seq methodology for use in differential gene expression
 Development of quantitative targeted RNA-seq methodology for use in differential gene expression Dr. Jens Winter, Market Development Group Biological Biological Research Content EMEA QIAGEN Universal Workflows
Development of quantitative targeted RNA-seq methodology for use in differential gene expression Dr. Jens Winter, Market Development Group Biological Biological Research Content EMEA QIAGEN Universal Workflows
Competition for XPO5 binding between Dicer mrna, pre-mirna and viral RNA regulates human Dicer levels
 CORRECTION NOTICE Nat. Struct. Mol. Biol. 18, 323 327 (211) Competition for binding between mrna, pre-mirna and viral RNA regulates human levels Yamina Bennasser, Christine Chable-Bessia, Robinson Triboulet,
CORRECTION NOTICE Nat. Struct. Mol. Biol. 18, 323 327 (211) Competition for binding between mrna, pre-mirna and viral RNA regulates human levels Yamina Bennasser, Christine Chable-Bessia, Robinson Triboulet,
Applications and Uses. (adapted from Roche RealTime PCR Application Manual)
 What Can You Do With qpcr? Applications and Uses (adapted from Roche RealTime PCR Application Manual) What is qpcr? Real time PCR also known as quantitative PCR (qpcr) measures PCR amplification as it
What Can You Do With qpcr? Applications and Uses (adapted from Roche RealTime PCR Application Manual) What is qpcr? Real time PCR also known as quantitative PCR (qpcr) measures PCR amplification as it
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Multiplet rate and sensitivity of the GemCode single cell platform from scrna-seq of 50:50 mixing of 293Ts and 3T3s. (a) Inferred multiplet rate as a function
Supplementary Figures Supplementary Figure 1. Multiplet rate and sensitivity of the GemCode single cell platform from scrna-seq of 50:50 mixing of 293Ts and 3T3s. (a) Inferred multiplet rate as a function
Supplementary Figure 1. Limit of quantitation (LOQ) and limit of detection (LOD) were
 Supplementary Figure 1. Limit of quantitation (LOQ) and limit of detection (LOD) were determined for SIL peptides in buffer solution. Presented in this figure are the regression analysis to demonstrate
Supplementary Figure 1. Limit of quantitation (LOQ) and limit of detection (LOD) were determined for SIL peptides in buffer solution. Presented in this figure are the regression analysis to demonstrate
Gene Expression on the Fluidigm BioMark HD
 Gene Expression on the Fluidigm BioMark HD Overview Introduction to Fluidigm James Miller Advantages of the technology Running a Fluidigm gene expression project Paul Lacaze Assay design, chemistry, experimental
Gene Expression on the Fluidigm BioMark HD Overview Introduction to Fluidigm James Miller Advantages of the technology Running a Fluidigm gene expression project Paul Lacaze Assay design, chemistry, experimental
Direct RNA vs cdna sequencing of C. elegans transcriptome
 Direct RNA vs cdna sequencing of C. elegans transcriptome Rachael Workman Timp Lab Johns Hopkins University Why compare direct RNA vs cdna sequencing? Many advantages to direct RNA: Poly-A profiling Modification
Direct RNA vs cdna sequencing of C. elegans transcriptome Rachael Workman Timp Lab Johns Hopkins University Why compare direct RNA vs cdna sequencing? Many advantages to direct RNA: Poly-A profiling Modification
Experimental design of RNA-Seq Data
 Experimental design of RNA-Seq Data RNA-seq course: The Power of RNA-seq Thursday June 6 th 2013, Marco Bink Biometris Overview Acknowledgements Introduction Experimental designs Randomization, Replication,
Experimental design of RNA-Seq Data RNA-seq course: The Power of RNA-seq Thursday June 6 th 2013, Marco Bink Biometris Overview Acknowledgements Introduction Experimental designs Randomization, Replication,
Supplementary Table S1
 Primers used in RT-qPCR, ChIP and Bisulphite-Sequencing. Quantitative real-time RT-PCR primers Supplementary Table S1 gene Forward primer sequence Reverse primer sequence Product TRAIL CAACTCCGTCAGCTCGTTAGAAAG
Primers used in RT-qPCR, ChIP and Bisulphite-Sequencing. Quantitative real-time RT-PCR primers Supplementary Table S1 gene Forward primer sequence Reverse primer sequence Product TRAIL CAACTCCGTCAGCTCGTTAGAAAG
Gene Expression Analysis with Pathway-Centric DNA Microarrays
 Gene Expression Analysis with Pathway-Centric DNA Microarrays SuperArray Bioscience Corporation George J. Quellhorst, Jr. Ph.D. Manager, Customer Education Topics to be Covered Introduction to DNA Microarrays
Gene Expression Analysis with Pathway-Centric DNA Microarrays SuperArray Bioscience Corporation George J. Quellhorst, Jr. Ph.D. Manager, Customer Education Topics to be Covered Introduction to DNA Microarrays
Miniaturized Quantitative PCR in a 1536-well Format Using the Echo Liquid Handler
 TM Application Note G101 Echo Liquid Handler Miniaturized Quantitative PCR in a 1536-well Format Using the Echo Liquid Handler Celeste Glazer, Sammy Datwani Labcyte Inc. Abstract Miniaturizing quantitative
TM Application Note G101 Echo Liquid Handler Miniaturized Quantitative PCR in a 1536-well Format Using the Echo Liquid Handler Celeste Glazer, Sammy Datwani Labcyte Inc. Abstract Miniaturizing quantitative
Regulation of Alternative Splicing at the Single-Cell Level
 Appendix Regulation of Alternative Splicing at the Single-Cell Level Lior Faigenbloom 1,, Nimrod D. Rubinstein 2,4,, Yoel Kloog 1, Itay Mayrose 3, Tal Pupko 2,*, Reuven Stein 1,* 1 The Department of Neurobiology,
Appendix Regulation of Alternative Splicing at the Single-Cell Level Lior Faigenbloom 1,, Nimrod D. Rubinstein 2,4,, Yoel Kloog 1, Itay Mayrose 3, Tal Pupko 2,*, Reuven Stein 1,* 1 The Department of Neurobiology,
Data Analysis on the ABI PRISM 7700 Sequence Detection System: Setting Baselines and Thresholds. Overview. Data Analysis Tutorial
 Data Analysis on the ABI PRISM 7700 Sequence Detection System: Setting Baselines and Thresholds Overview In order for accuracy and precision to be optimal, the assay must be properly evaluated and a few
Data Analysis on the ABI PRISM 7700 Sequence Detection System: Setting Baselines and Thresholds Overview In order for accuracy and precision to be optimal, the assay must be properly evaluated and a few
Quantitative Real time PCR. Only for teaching purposes - not for reproduction or sale
 Quantitative Real time PCR PCR reaction conventional versus real time PCR real time PCR principles threshold cycle C T efficiency relative quantification reference genes primers detection chemistry GLP
Quantitative Real time PCR PCR reaction conventional versus real time PCR real time PCR principles threshold cycle C T efficiency relative quantification reference genes primers detection chemistry GLP
Rapid Method for the Purification of Total RNA from Formalin- Fixed Paraffin-Embedded (FFPE) Tissue Samples
 Application Note 17 RNA Sample Preparation Rapid Method for the Purification of Total RNA from Formalin- Fixed Paraffin-Embedded (FFPE) Tissue Samples M. Melmogy 1, V. Misic 1, B. Lam, PhD 1, C. Dobbin,
Application Note 17 RNA Sample Preparation Rapid Method for the Purification of Total RNA from Formalin- Fixed Paraffin-Embedded (FFPE) Tissue Samples M. Melmogy 1, V. Misic 1, B. Lam, PhD 1, C. Dobbin,
Gene Regulation Solutions. Microarrays and Next-Generation Sequencing
 Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
Gene Regulation Solutions Microarrays and Next-Generation Sequencing Gene Regulation Solutions The Microarrays Advantage Microarrays Lead the Industry in: Comprehensive Content SurePrint G3 Human Gene
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Development of conditions to convert 4-thiouridine (s 4 U) into a convertible nucleoside. Development of conditions to convert 4-thiouridine (s 4 U) into a convertible nucleoside.
Supplementary Figure 1 Development of conditions to convert 4-thiouridine (s 4 U) into a convertible nucleoside. Development of conditions to convert 4-thiouridine (s 4 U) into a convertible nucleoside.
Supplementary Figure 1
 Supplementary Figure 1 CITE-seq library preparation. (a) Illustration of the DNA-barcoded antibodies used in CITE-seq. (b) Antibody-oligonucleotide complexes appear as a high-molecular-weight smear when
Supplementary Figure 1 CITE-seq library preparation. (a) Illustration of the DNA-barcoded antibodies used in CITE-seq. (b) Antibody-oligonucleotide complexes appear as a high-molecular-weight smear when
less sensitive than RNA-seq but more robust analysis pipelines expensive but quantitiatve standard but typically not high throughput
 Chapter 11: Gene Expression The availability of an annotated genome sequence enables massively parallel analysis of gene expression. The expression of all genes in an organism can be measured in one experiment.
Chapter 11: Gene Expression The availability of an annotated genome sequence enables massively parallel analysis of gene expression. The expression of all genes in an organism can be measured in one experiment.
Spermatazoa. Sertoli cells. Morc1 WT adult. Morc1 KO adult. Morc1 WT P14.5. Morc1 KO P14.5. Germ cells. Germ cells. Spermatids.
 a. Spermatazoa Sertoli cells Morc1 WT adult c. Morc1 KO adult Morc1 WT P1.5 Morc1 KO P1.5 Morc1 WT adult Morc1 KO adult Germ cells Germ cells Spermatids Spermatozoa Supplementary Figure 1. Confirmation
a. Spermatazoa Sertoli cells Morc1 WT adult c. Morc1 KO adult Morc1 WT P1.5 Morc1 KO P1.5 Morc1 WT adult Morc1 KO adult Germ cells Germ cells Spermatids Spermatozoa Supplementary Figure 1. Confirmation
initial single-cell analysis, with a pragmatic focus on surface markers with the highest potential for
 Supplementary Figure 1: Summary of the exclusionary approach to surface marker selection for initial single-cell analysis, with a pragmatic focus on surface markers with the highest potential for protein
Supplementary Figure 1: Summary of the exclusionary approach to surface marker selection for initial single-cell analysis, with a pragmatic focus on surface markers with the highest potential for protein
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/496/eaam6291/dc1 Supplementary Materials for Regulation of autophagy, NF-κB signaling, and cell viability by mir-124 in KRAS mutant mesenchymal-like NSCLC cells
www.sciencesignaling.org/cgi/content/full/10/496/eaam6291/dc1 Supplementary Materials for Regulation of autophagy, NF-κB signaling, and cell viability by mir-124 in KRAS mutant mesenchymal-like NSCLC cells
i-stop codon positions in the mcherry gene
 Supplementary Figure 1 i-stop codon positions in the mcherry gene The grnas (green) that can potentially generate stop codons from Trp (63 th and 98 th aa, upper panel) and Gln (47 th and 114 th aa, bottom
Supplementary Figure 1 i-stop codon positions in the mcherry gene The grnas (green) that can potentially generate stop codons from Trp (63 th and 98 th aa, upper panel) and Gln (47 th and 114 th aa, bottom
of NOBOX. Supplemental Figure S1F shows the staining profile of LHX8 antibody (Abcam, ab41519), which is a
 SUPPLEMENTAL MATERIALS AND METHODS LHX8 Staining During the course of this work, it came to our attention that the NOBOX antibody used in this study had been discontinued. There are a number of other nuclear
SUPPLEMENTAL MATERIALS AND METHODS LHX8 Staining During the course of this work, it came to our attention that the NOBOX antibody used in this study had been discontinued. There are a number of other nuclear
Assessing a novel approach for predicting local 3D protein structures from sequence
 Assessing a novel approach for predicting local 3D protein structures from sequence Cristina Benros*, Alexandre G. de Brevern, Catherine Etchebest and Serge Hazout Equipe de Bioinformatique Génomique et
Assessing a novel approach for predicting local 3D protein structures from sequence Cristina Benros*, Alexandre G. de Brevern, Catherine Etchebest and Serge Hazout Equipe de Bioinformatique Génomique et
Virus-induced gene complementation reveals a transcription factor network in modulation of tomato fruit ripening
 Supplementary Information Virus-induced gene complementation reveals a transcription factor network in modulation of tomato fruit ripening Tao Zhou 2,3, Hang Zhang 2,4, Tongfei Lai 1, Cheng Qin 1, Nongnong
Supplementary Information Virus-induced gene complementation reveals a transcription factor network in modulation of tomato fruit ripening Tao Zhou 2,3, Hang Zhang 2,4, Tongfei Lai 1, Cheng Qin 1, Nongnong
measuring gene expression December 11, 2018
 measuring gene expression December 11, 2018 Intervening Sequences (introns): how does the cell get rid of them? Splicing!!! Highly conserved ribonucleoprotein complex recognizes intron/exon junctions and
measuring gene expression December 11, 2018 Intervening Sequences (introns): how does the cell get rid of them? Splicing!!! Highly conserved ribonucleoprotein complex recognizes intron/exon junctions and
ABI3 Controls Embryo De-greening Through Mendel's I locus
 Supporting Online Material for ABI3 Controls Embryo De-greening Through Mendel's I locus Frédéric Delmas a,b,c, Subramanian Sankaranarayanan d, Srijani Deb d, Ellen Widdup d, Céline Bournonville b,c,norbert
Supporting Online Material for ABI3 Controls Embryo De-greening Through Mendel's I locus Frédéric Delmas a,b,c, Subramanian Sankaranarayanan d, Srijani Deb d, Ellen Widdup d, Céline Bournonville b,c,norbert
B. Transgenic plants with strong phenotype (%)
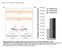 A. TCTAGTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT AAATATGGTCTAAAGAAGAAGAATATGGTCTAAAGAAGAAGAATATGG 2XP35S STTM165 5 GGGGGATGAAGctaCCTGGTCCGA3 3 CCCCCUACUUC---GGACCAGGCU5 mir165 HindIII mir165 96 nt GTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT
A. TCTAGTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT AAATATGGTCTAAAGAAGAAGAATATGGTCTAAAGAAGAAGAATATGG 2XP35S STTM165 5 GGGGGATGAAGctaCCTGGTCCGA3 3 CCCCCUACUUC---GGACCAGGCU5 mir165 HindIII mir165 96 nt GTTGTTGTTGTTATGGTCTAGTTGTTGTTGTTATGGTCTAATTT
Benchmarking of RNA-seq data processing pipelines using whole transcriptome qpcr expression data
 Benchmarking of RNA-seq data processing pipelines using whole transcriptome qpcr expression data Jan Hellemans 7th international qpcr & NGS Event - Freising March 24 th, 2015 Therapeutics lncrna oncology
Benchmarking of RNA-seq data processing pipelines using whole transcriptome qpcr expression data Jan Hellemans 7th international qpcr & NGS Event - Freising March 24 th, 2015 Therapeutics lncrna oncology
Supplemental Figure S1. Validation of auxin-related gene expression from Fig. 1A. Relative gene expression in 2x and 3x seeds 6 days after
 Supplemental Figure S1. Validation of auxin-related gene expression from Fig. 1A. Relative gene expression in 2x and 3x seeds 6 days after pollination (6 DAP), as determined by RT-qPCR for YUC10 (A), YUC11
Supplemental Figure S1. Validation of auxin-related gene expression from Fig. 1A. Relative gene expression in 2x and 3x seeds 6 days after pollination (6 DAP), as determined by RT-qPCR for YUC10 (A), YUC11
(A-B) P2ry14 expression was assessed by (A) genotyping (upper arrow: WT; lower
 Supplementary Figures S1. (A-B) P2ry14 expression was assessed by (A) genotyping (upper arrow: ; lower arrow: KO) and (B) q-pcr analysis with Lin- cells, The white vertical line in panel A indicates that
Supplementary Figures S1. (A-B) P2ry14 expression was assessed by (A) genotyping (upper arrow: ; lower arrow: KO) and (B) q-pcr analysis with Lin- cells, The white vertical line in panel A indicates that
A normalization method based on variance and median adjustment for massive mrna polyadenylation data
 ISSN : 0974-7435 Volume 8 Issue 4 A normalization method based on variance and median adjustment for massive mrna polyadenylation data Guoli Ji, Ying Wang, Mingchen Wu, Yangzi Zhang, Xiaohui Wu* Department
ISSN : 0974-7435 Volume 8 Issue 4 A normalization method based on variance and median adjustment for massive mrna polyadenylation data Guoli Ji, Ying Wang, Mingchen Wu, Yangzi Zhang, Xiaohui Wu* Department
Measuring gene expression (Microarrays) Ulf Leser
 Measuring gene expression (Microarrays) Ulf Leser This Lecture Gene expression Microarrays Idea Technologies Problems Quality control Normalization Analysis next week! 2 http://learn.genetics.utah.edu/content/molecules/transcribe/
Measuring gene expression (Microarrays) Ulf Leser This Lecture Gene expression Microarrays Idea Technologies Problems Quality control Normalization Analysis next week! 2 http://learn.genetics.utah.edu/content/molecules/transcribe/
Gene Expression Analysis Superior Solutions for any Project
 Gene Expression Analysis Superior Solutions for any Project Find Your Perfect Match ArrayXS Global Array-to-Go Focussed Comprehensive: detect the whole transcriptome reliably Certified: discover exceptional
Gene Expression Analysis Superior Solutions for any Project Find Your Perfect Match ArrayXS Global Array-to-Go Focussed Comprehensive: detect the whole transcriptome reliably Certified: discover exceptional
Regulation of Synaptic Structure and Function by FMRP- Associated MicroRNAs mir-125b and mir-132
 Neuron, Volume 65 Regulation of Synaptic Structure and Function by FMRP- Associated MicroRNAs mir-125b and mir-132 Dieter Edbauer, Joel R. Neilson, Kelly A. Foster, Chi-Fong Wang, Daniel P. Seeburg, Matthew
Neuron, Volume 65 Regulation of Synaptic Structure and Function by FMRP- Associated MicroRNAs mir-125b and mir-132 Dieter Edbauer, Joel R. Neilson, Kelly A. Foster, Chi-Fong Wang, Daniel P. Seeburg, Matthew
Rapid Method for the Purification of Total RNA from Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Samples
 Application Note 17 RNA Sample Preparation Rapid Method for the Purification of Total RNA from Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Samples M. Melmogy 1, V. Misic 1, B. Lam, PhD 1, C. Dobbin,
Application Note 17 RNA Sample Preparation Rapid Method for the Purification of Total RNA from Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Samples M. Melmogy 1, V. Misic 1, B. Lam, PhD 1, C. Dobbin,
Supplementary Materials and Methods:
 Supplementary Materials and Methods: Preclinical chemoprevention experimental design Mice were weighed and each mammary tumor was manually palpated and measured with digital calipers once a week from 4
Supplementary Materials and Methods: Preclinical chemoprevention experimental design Mice were weighed and each mammary tumor was manually palpated and measured with digital calipers once a week from 4
Alain Sewer 1*, Sylvain Gubian 1, Ulrike Kogel 2, Emilija Veljkovic 1, Wanjiang Han 1, Arnd Hengstermann 2, Manuel C Peitsch 1 and Julia Hoeng 1
 Sewer et al. BMC Research Notes 214, 7:32 RESEARCH ARTICLE Open Access Assessment of a novel multi-array normalization method based on spike-in control probes suitable for microrna datasets with global
Sewer et al. BMC Research Notes 214, 7:32 RESEARCH ARTICLE Open Access Assessment of a novel multi-array normalization method based on spike-in control probes suitable for microrna datasets with global
supplementary information
 Figure S1 ZEB1 full length mrna. (a) Analysis of the ZEB1 mrna using the UCSC genome browser (http://genome.ucsc.edu) revealed truncation of the annotated Refseq sequence (NM_030751). The probable terminus
Figure S1 ZEB1 full length mrna. (a) Analysis of the ZEB1 mrna using the UCSC genome browser (http://genome.ucsc.edu) revealed truncation of the annotated Refseq sequence (NM_030751). The probable terminus
ChIP-seq and RNA-seq. Farhat Habib
 ChIP-seq and RNA-seq Farhat Habib fhabib@iiserpune.ac.in Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions
ChIP-seq and RNA-seq Farhat Habib fhabib@iiserpune.ac.in Biological Goals Learn how genomes encode the diverse patterns of gene expression that define each cell type and state. Protein-DNA interactions
PrimerArray Analysis Tool for Embryonic Stem Cells
 For Research Use Manual Table of Contents I. Calculating and exporting Ct values... 3 II. Relative quantification... 4 III. Troubleshooting... 8 2 URL:http://www.takara-bio.com The PrimerArray Analysis
For Research Use Manual Table of Contents I. Calculating and exporting Ct values... 3 II. Relative quantification... 4 III. Troubleshooting... 8 2 URL:http://www.takara-bio.com The PrimerArray Analysis
Introduction to RNA-Seq. David Wood Winter School in Mathematics and Computational Biology July 1, 2013
 Introduction to RNA-Seq David Wood Winter School in Mathematics and Computational Biology July 1, 2013 Abundance RNA is... Diverse Dynamic Central DNA rrna Epigenetics trna RNA mrna Time Protein Abundance
Introduction to RNA-Seq David Wood Winter School in Mathematics and Computational Biology July 1, 2013 Abundance RNA is... Diverse Dynamic Central DNA rrna Epigenetics trna RNA mrna Time Protein Abundance
Result Tables The Result Table, which indicates chromosomal positions and annotated gene names, promoter regions and CpG islands, is the best way for
 Result Tables The Result Table, which indicates chromosomal positions and annotated gene names, promoter regions and CpG islands, is the best way for you to discover methylation changes at specific genomic
Result Tables The Result Table, which indicates chromosomal positions and annotated gene names, promoter regions and CpG islands, is the best way for you to discover methylation changes at specific genomic
Identification of cytokine-induced modulation of microrna expression and secretion as measured by a novel microrna specific qpcr assay
 Supplementary Figure Legends Supplementary Figure 1: Detection, analysis and comparison of mirnas expression in mouse liver and heart by using miqpcr and TaqMan mirna assays. A) Microarray analysis of
Supplementary Figure Legends Supplementary Figure 1: Detection, analysis and comparison of mirnas expression in mouse liver and heart by using miqpcr and TaqMan mirna assays. A) Microarray analysis of
A synthetic recombinase-based feedback loop results in robust expression Supporting Information
 A synthetic recombinase-based feedback loop results in robust expression Supporting Information 1 Fig. S1: Control experiments performed to demonstrate the expected function of our closed-loop feedback
A synthetic recombinase-based feedback loop results in robust expression Supporting Information 1 Fig. S1: Control experiments performed to demonstrate the expected function of our closed-loop feedback
Nanocrystals of the Fe(phen) 2 (NCS) 2 prototypical compound and the size-dependent spin-crossover characteristics
 Electronic Supplementary Material (ESI) for Dalton Transactions. This journal is The Royal Society of Chemistry 2015 Nanocrystals of the Fe(phen) 2 (NCS) 2 prototypical compound and the size-dependent
Electronic Supplementary Material (ESI) for Dalton Transactions. This journal is The Royal Society of Chemistry 2015 Nanocrystals of the Fe(phen) 2 (NCS) 2 prototypical compound and the size-dependent
PrimerArray Analysis Tool Ver. 2.2
 For Research Use PrimerArray Analysis Tool Ver. 2.2 Manual Table of Contents I. Calculating and exporting Ct values... 3 II. Relative quantification... 4 III. Troubleshooting...10 2 URL:http://www.takara-bio.com
For Research Use PrimerArray Analysis Tool Ver. 2.2 Manual Table of Contents I. Calculating and exporting Ct values... 3 II. Relative quantification... 4 III. Troubleshooting...10 2 URL:http://www.takara-bio.com
Nature Immunology: doi: /ni Supplementary Figure 1. Data-processing pipeline.
 Supplementary Figure 1 Data-processing pipeline. Steps for processing data from multiple sorted B cell populations derived from a single individual at a single time point are shown. Parameters used are
Supplementary Figure 1 Data-processing pipeline. Steps for processing data from multiple sorted B cell populations derived from a single individual at a single time point are shown. Parameters used are
Supplement to: Deconvoluting Stress-Responsive Proteostasis Signaling Pathways for Pharmacologic Activation using Targeted RNA-sequencing
 Supplement to: Deconvoluting Stress-Responsive Proteostasis Signaling Pathways for Pharmacologic Activation using Targeted RNA-sequencing Julia M.D. Grandjean 1, Lars Plate 1,2, Richard I. Morimoto 3,
Supplement to: Deconvoluting Stress-Responsive Proteostasis Signaling Pathways for Pharmacologic Activation using Targeted RNA-sequencing Julia M.D. Grandjean 1, Lars Plate 1,2, Richard I. Morimoto 3,
mrna Sequencing Quality Control (V6)
 mrna Sequencing Quality Control (V6) Notes: the following analyses are based on 8 adult brains sequenced in USC and Yale 1. Error Rates The error rates of each sequencing cycle are reported for 120 tiles
mrna Sequencing Quality Control (V6) Notes: the following analyses are based on 8 adult brains sequenced in USC and Yale 1. Error Rates The error rates of each sequencing cycle are reported for 120 tiles
Going further with 3-color Crystal Digital PCR: characterizing CNV and indels. 4th qpcr & Digital PCR Congress 09/13/2018
 Going further with 3-color Crystal Digital PCR: characterizing CNV and indels 4th qpcr & Digital PCR Congress 09/13/2018 Overview of presentation I. Introduction to digital PCR and Naica System II. Reliable
Going further with 3-color Crystal Digital PCR: characterizing CNV and indels 4th qpcr & Digital PCR Congress 09/13/2018 Overview of presentation I. Introduction to digital PCR and Naica System II. Reliable
Technical Note. GeneChip 3 IVT PLUS Reagent Kit vs. GeneChip 3 IVT Express Reagent Kit Comparison. Introduction:
 Technical Note GeneChip 3 IVT PLUS Reagent Kit vs. GeneChip 3 IVT Express Reagent Kit Comparison Introduction: Affymetrix has launched a new 3 IVT PLUS Reagent Kit which creates hybridization ready target
Technical Note GeneChip 3 IVT PLUS Reagent Kit vs. GeneChip 3 IVT Express Reagent Kit Comparison Introduction: Affymetrix has launched a new 3 IVT PLUS Reagent Kit which creates hybridization ready target
Standardized Next Generation Sequencing Abundance Measurements (StarSeq) using Competitive Template Mixtures. AccuGenomics Inc.
 Standardized Next Generation Sequencing Abundance Measurements (StarSeq) using Competitive Template Mixtures AccuGenomics Inc. Need for NGS Standardization * Nature Biotechnology November 2012 Targeted
Standardized Next Generation Sequencing Abundance Measurements (StarSeq) using Competitive Template Mixtures AccuGenomics Inc. Need for NGS Standardization * Nature Biotechnology November 2012 Targeted
% Viability. isw2 ino isw2 ino isw2 ino isw2 ino mM HU 4-NQO CPT
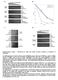 a Drug concentration b 1.3% MMS nhp1 nhp1 8 nhp1 mag1.5% MMS.3% MMS nhp1 nhp1 ino8 9 ino8 9 % Viability 4.5% MMS ino8 9 ino8 9 2.5.1.15 % MMS c d nhp1 nhp1 nhp1 nhp1 nhp1 nhp1 Control (YPD) γ IR (1 gy)
a Drug concentration b 1.3% MMS nhp1 nhp1 8 nhp1 mag1.5% MMS.3% MMS nhp1 nhp1 ino8 9 ino8 9 % Viability 4.5% MMS ino8 9 ino8 9 2.5.1.15 % MMS c d nhp1 nhp1 nhp1 nhp1 nhp1 nhp1 Control (YPD) γ IR (1 gy)
Interplay Between Intracellular Ca 2+ Oscillations and Ca 2+ -stimulated Mitochondrial Metabolism
 Interplay Between Intracellular Ca 2+ Oscillations and Ca 2+ -stimulated Mitochondrial Metabolism Benjamin Wacquier 1, Laurent Combettes 2,3, Guy Tran Van Nhieu 4,5,6,7, and Geneviève Dupont 1,* 1 Unité
Interplay Between Intracellular Ca 2+ Oscillations and Ca 2+ -stimulated Mitochondrial Metabolism Benjamin Wacquier 1, Laurent Combettes 2,3, Guy Tran Van Nhieu 4,5,6,7, and Geneviève Dupont 1,* 1 Unité
Supplementary Figure 1: sgrna library generation and the length of sgrnas for the functional screen. (a) A diagram of the retroviral vector for sgrna
 Supplementary Figure 1: sgrna library generation and the length of sgrnas for the functional screen. (a) A diagram of the retroviral vector for sgrna expression. It contains a U6-promoter-driven sgrna
Supplementary Figure 1: sgrna library generation and the length of sgrnas for the functional screen. (a) A diagram of the retroviral vector for sgrna expression. It contains a U6-promoter-driven sgrna
Principals of Real-Time PCR. Amira A. T. AL-Hosary Lecturer of Infectious Diseases, Faculty of Veterinary Medicine, Assiut University, Egypt
 Principals of Real-Time PCR Amira A. T. AL-Hosary Lecturer of Infectious Diseases, Faculty of Veterinary Medicine, Assiut University, Egypt What Is Real-Time PCR? Nucleic acid (DNA) amplification and detection
Principals of Real-Time PCR Amira A. T. AL-Hosary Lecturer of Infectious Diseases, Faculty of Veterinary Medicine, Assiut University, Egypt What Is Real-Time PCR? Nucleic acid (DNA) amplification and detection
Nature Methods: doi: /nmeth Supplementary Figure 1. DMS-MaPseq data are highly reproducible at elevated DMS concentrations.
 Supplementary Figure 1 DMS-MaPseq data are highly reproducible at elevated DMS concentrations. a, Correlation of Gini index for 202 yeast mrna regions with 15x coverage at 2.5% or 5% v/v DMS concentrations
Supplementary Figure 1 DMS-MaPseq data are highly reproducible at elevated DMS concentrations. a, Correlation of Gini index for 202 yeast mrna regions with 15x coverage at 2.5% or 5% v/v DMS concentrations
Quantitative Real time PCR. Only for teaching purposes - not for reproduction or sale
 Quantitative Real time PCR PCR reaction conventional versus real time PCR real time PCR principles threshold cycle C T efficiency relative quantification reference genes primers detection chemistry GLP
Quantitative Real time PCR PCR reaction conventional versus real time PCR real time PCR principles threshold cycle C T efficiency relative quantification reference genes primers detection chemistry GLP
mir-24-mediated down-regulation of H2AX suppresses DNA repair
 Supplemental Online Material mir-24-mediated down-regulation of H2AX suppresses DNA repair in terminally differentiated blood cells Ashish Lal 1,4, Yunfeng Pan 2,4, Francisco Navarro 1,4, Derek M. Dykxhoorn
Supplemental Online Material mir-24-mediated down-regulation of H2AX suppresses DNA repair in terminally differentiated blood cells Ashish Lal 1,4, Yunfeng Pan 2,4, Francisco Navarro 1,4, Derek M. Dykxhoorn
Gene Expression Profiling and Validation Using Agilent SurePrint G3 Gene Expression Arrays
 Gene Expression Profiling and Validation Using Agilent SurePrint G3 Gene Expression Arrays Application Note Authors Bahram Arezi, Nilanjan Guha and Anne Bergstrom Lucas Agilent Technologies Inc. Santa
Gene Expression Profiling and Validation Using Agilent SurePrint G3 Gene Expression Arrays Application Note Authors Bahram Arezi, Nilanjan Guha and Anne Bergstrom Lucas Agilent Technologies Inc. Santa
digital PCR Maido Remm BIOINFO Journal Club
 digital PCR Maido Remm BIOINFO Journal Club 04.10.2013 real-.me PCR (qpcr) = analog PCR h=p://origin- ars.els- cdn.com/content/image/1- s2.0- S073497501000090X- gr3.jpg real-.me PCR (qpcr) increase in
digital PCR Maido Remm BIOINFO Journal Club 04.10.2013 real-.me PCR (qpcr) = analog PCR h=p://origin- ars.els- cdn.com/content/image/1- s2.0- S073497501000090X- gr3.jpg real-.me PCR (qpcr) increase in
Supporting Information
 Supporting Information Table S1. Overview of samples used for sequencing, and the number of sequences obtained from each sample. Visit 1 is day 0, Visit 2 is day 7, Visit 3 is day 28, and Visit 4 is day
Supporting Information Table S1. Overview of samples used for sequencing, and the number of sequences obtained from each sample. Visit 1 is day 0, Visit 2 is day 7, Visit 3 is day 28, and Visit 4 is day
Supplementary Figure 1 Validate the expression of mir-302b after bacterial infection by northern
 Supplementary Figure 1 Validate the expression of mir-302b after bacterial infection by northern blot. Northern blot analysis of mir-302b expression following infection with PAO1, PAK and Kp in (A) lung
Supplementary Figure 1 Validate the expression of mir-302b after bacterial infection by northern blot. Northern blot analysis of mir-302b expression following infection with PAO1, PAK and Kp in (A) lung
Acclaro Protein Contaminant ID
 TECHNICAL NOTE Acclaro Protein Contaminant ID Detection of Protein in Nucleic Acid Samples Using the NanoDrop One Spectrophotometer No. 52853 Sean Loughrey and Brian Matlock, Thermo Fisher Scientific,
TECHNICAL NOTE Acclaro Protein Contaminant ID Detection of Protein in Nucleic Acid Samples Using the NanoDrop One Spectrophotometer No. 52853 Sean Loughrey and Brian Matlock, Thermo Fisher Scientific,
