ABI3 Controls Embryo De-greening Through Mendel's I locus
|
|
|
- Darleen McCoy
- 5 years ago
- Views:
Transcription
1 Supporting Online Material for ABI3 Controls Embryo De-greening Through Mendel's I locus Frédéric Delmas a,b,c, Subramanian Sankaranarayanan d, Srijani Deb d, Ellen Widdup d, Céline Bournonville b,c,norbert Bollier b,c, Julian G. B. Northey a, Peter McCourt a,1, and Marcus A. Samuel d,1 1
2 Supplemental Table 1. Genes with altered expression (log 2 ) in abi3-6 embryos relative to expression in Col-0 embryos at 13 DAF AGI ID Gene Description Log 2 ratio Plant development-related At4g34520 FAE1 (FATTY ACID ELONGATION1) very long chain fatty acids content of seed oil At5g51210 OLEOSIN3 involved in seed lipid accumulation At4g S seed storage protein At4g S seed storage protein At4g S seed storage protein At3g15670 Late embryogenesis abundant protein, putative / LEA protein At1g32560 Late embryogenesis abundant group 1 / LEA group At4g21020 Late embryogenesis abundant domain-containing protein / LEA Light reaction-related At2g40100 LHCB4.3 (LIGHT HARVESTING COMPLEX PSII) 2.31 At1g45474 LHCA5 (Photosystem I light harvesting complex gene 5) 1.15 At3g50820 PSBO2 (PHOTOSYSTEM II SUBUNIT O-2); oxygen evolving 1.86 At4g21280 PSBQ oxygen evolving complex of Photosystem II 1.94 At2g46820 TMP14 encodes the P- subunit of Photosystem I 2.34 At1g76450 Oxygen evolving complex-related, chloroplast precursor 1.80 At1g76100 Plastocyanin minor isoform, chloroplast precursor 2.69 Tetrapyrrole synthesis-related At4g18480 CHLI1 (CHLORINA 42) Encodes the CHLI subunit of magnesium chelatase 2.69 At3g59400 GUN4 (Genomes uncoupled 4) enhances the activity of Mg-chelatase 1.88 At1g44446 CH1 (CHLORINA 1) chlorophyll a oxygenase 1.62 At3g51820 ATG4 / CHLG / G4 (CHLOROPHYLL SYNTHASE) chlorophyll synthase 1.44 activity At3g14110 FLU (FLUORESCENT IN BLUE LIGHT) involved in chlorophyll biosynthesis 1.21 Senescence-related At4g02380 SAG21 (SENESCENCE-ASSOCIATED GENE 21) encodes AtLEA At1g17020 SRG1 (SENESCENCE-RELATED GENE 1) oxidoreductase 2.95 At1g21460 Nodulin MtN3 family protein similar to senescence-associated protein-like 2.05 At1g79970 Similar to senescence-associated protein-related 1.72 At2g44670 Senescence-associated protein-related 1.58 At1g74940 Senescence-associated protein-related 1.35 At5g47060 Senescence-associated protein-related 1.23 At4g22920 Similar to tomato senescence-inducible chloroplast stay-green protein
3 Figure S1. A. Protein domains of ABI3 with the locations of the published mutations (modified from Nambara et al., 2002) B. Transient expression of GFP- ABI3, GFP-ABI3-6 and GFP-ABI3-8 in suspension-cultured tobacco cells. C. EFP-browser data showing ABI3 and SGR1 expression during Arabidopsis embryo maturation stages. 3
4 Figure S2. Gene expression profiles of abi3-6 embryos at 13 DAF. Fold changes (log 2 ) of expression values of abi3-6 embryos relative to expression in Col-0 with P <0.01 are represented as blue (upregulated), red (downregulated) and white (unchanged) squares for each gene based on the pathway analysis program MapMan ( 4
5 Figure S3. Vegetative phenotypes of abi3-6/35s::sgr1 and abi3-6/35s::sgr2. A. SGR1 overexpression in abi3-6 background resulted in leaf yellowing phenotypes, while SGR2 overexpression did not cause any shoot phenotypes (B). 5
6 Figure S4. Protein profiles were analyzed from mature seeds of Col-0, abi3-6 and abi3-6/35s::sgr overexpressors A. Freshly harvested mature seeds from the transgenic lines were tested for ABA sensitivity by germinating them on 10 and 25 μm ABA without stratification. Radicle emergence was observed at either 24h (B) or 72h (C) following plating. Values are means ± SEM of three biological replicates. Germination index was measured as the ratio of % germination on treatment plates / % germination of abi3-6 on half strength MS plates 6
7 Figure S5. Protein profiles were analyzed from mature seeds of Col-0, abi3-6, sgr1-1, sgr2-2 and sgr1-1/sgr2-2 7
8 Figure S6. Induced senescence assay indicates a seed specific de-greening phenotype in abi3-6. Rosette phenotypes of Col-0, abi3-6, sgr1-1 and sgr2-2 homozygous T-DNA insertional lines after 7 days of dark-induced senescence 8
9 Figure S7. RT-qPCR analysis of RAB18 expression in seeds (11-13 DAF) that were either left untreated or exposed to freezing as in Figure 4A and allowed to recover for either 1 day or 2 days at ambient temperature (n=4). Asterisk indicates p<0.05 compared to untreated Col-0. 9
10 Supplemental methods Identification of stable reference genes for RT-qPCR analysis during seed maturation and cold treatment The reference genes tubulin 4 (TUB4), actin 2 (ACT2), ubiquitin 10 (UBQ10) and glyceraldehyde 3-phosphate dehydrogenase A (GAPDA) were analyzed for their transcript stability during seed maturation (13 DAF vs 16 DAF) in both Col-0 and abi3-6 genotypes. Their stability following cold treatment in Col-0 and ABI3-OX seeds was also analyzed to identify the reference gene with the least variation following cold treatment. Reference gene specific qpcr reactions were performed with cdna synthesized from seed RNA isolated from various developmental stages or cold treatment conditions. The Ct (cycle threshold) values obtained for the four reference genes were processed using BestKeeper software to identify the reference genes that displayed the highest transcript stability. The variations in expression of the reference genes were displayed as the standard deviation (SD) of the Ct values and the co-efficient of variance (CV) expressed as a percentage of the Ct level. Reference genes with SD values greater than 1 were considered as unstable under the conditions tested. Based on both these values, reference genes with the least variation were identified as the most stable ones. Stability of reference genes during seed maturation: RNA extracted from Col- 0 and abi3-6 seeds at 13 DAF and 16 DAF were used for RT-qPCR and BestKeeper analyses to identify the most stable reference genes. Expression of 10
11 TUB4 and ACT2 was stable during seed maturation (Figure S8A and B). UBQ10 expression was not stable as it was upregulated at 16 DAF in both genotypes (Figure S8A and B). Supplemental Table 2. BestKeeper analysis of the expression of reference genes in maturing seeds (13 DAF and 16 DAF) from Col-0 and abi3-6 TUB4 ACT2 UBQ10 GAPDA N Geo Mean [Ct] Ar Mean [Ct] Min [Ct] Max [Ct] Std dev [± Ct] CV [% Ct] Ranking Abbreviations- N: number of samples; Geo Mean: Geometric mean of Ct; Ar Mean: Arithmetic mean of Ct Min and Max [Ct]: extreme values of Ct; std dev [± Ct]: standard deviation of the Ct; CV [% Ct]: the coefficient of variance expressed as a percentage on the Ct level, Stability ranking indicates ranking based on std dev [± Ct] and CV [% Ct]. 11
12 Figure S8. Cycle threshold (Ct) values for the expression of various reference genes in Col-0 and abi3-6 seeds at 13 DAF and 16DAF (A). Biological replicates are plotted together. Standard deviations of the Cts as identified by BestKeeper analysis of the Ct values of the same samples (B). Genes with the least variations are the most stable under the conditions tested. 12
13 Stability of reference genes under cold treatment: Seed RNA extracted from both untreated and cold-treated (frozen and allowed to recover for either one or two days) plants were subjected to RT-qPCR and BestKeeper analyses to identify the most stable reference gene. Although UBQ10 expression was not stable between 13 DAF and 16 DAF, it was the most stable reference gene when similar stages of seeds from Col-0 and ABI3-OX were analyzed following cold treatment (Figure S9A and B). Supplemental Table 3. BestKeeper analysis of the expression of reference genes in control and cold-treated Col-0 and ABI3-OX seeds TUB4 ACT2 UBQ10 GAPDA N Geo Mean [Ct] Ar Mean [Ct] Min [Ct] Max [Ct] Std dev [± Ct] CV [% Ct] Ranking
14 Figure S9. Cycle threshold (Ct) values for the expression of various reference genes in control and cold-treated seeds from Col-0 and ABI3-OX lines (A). Biological replicates are plotted together. UT: untreated; 1D: one day after treatment; 2D: two days after treatment. Standard deviations of the Cts as identified by BestKeeper analysis of the Ct values of the same samples (B). Genes with the least variations are the most stable under the conditions tested. 14
15 Supplemental Table 4. Sequences of primers used RT-qPCR Gene AGI ID Primer Seq. Product size (bp) F Primer SGR1 AT4G22920 AGCAGCAGCAGCTCACTCTTCTT 165 R Primer SGR1 AT4G22920 CCTAGGGAGCGTTGAAGGAT 165 F Primer SGR2 AT4G11910 ATTAGCGGAGGCCACTTCTT 183 R Primer SGR2 AT4G11910 GTTGTACTCCGGGATGTTGG 183 F Primer TUB4 AT5G44340 AACGCTGACGAGTGTATGGTTTT 86 R Primer TUB4 AT5G44340 CCAAAGGTAGGATTAGCGAGCTT 86 F Primer UBQ10 AT4G05320 GGCCTTGTATAATCCCTGATGAATAAG 63 R Primer UBQ10 AT4G05320 AAAGAGATAACAGGAACGGAAACATAGT 63 F Primer ACT2 AT3G18780 ACTGTGCCAATCTACGAGGGTTTC 150 R Primer ACT2 AT3G18780 GTCTCTTACAATTTCCCGCTCTGC 150 F Primer GAPDA AT3G26650 TAACCGAAACCCGTCTCTTC 107 R Primer GAPDA AT3G26650 CTTCAATGTGTTTCCCTGCAC 107 F Primer ABI3 AT3G24650 GCTGCTGTGTTTTTGGAGTG 110 R Primer ABI3 AT3G24650 AGTCTTCTTGCCGCTGATTC 110 F Primer RAB18 AT5G66440 GGTCATCATGATCAGTCTGG 112 R Primer RAB18 AT5G66440 TCTTGTCCATCATCCCCTTC 112 RT-PCR F Primer SGR1 AT4G22920 CATCCTTCAACGCTCCCTAG 430 R Primer SGR1 AT4G22920 CAGTCTTGTGACCATCAGGC 430 F Primer SGR2 AT4G11910 ACACGCAAGAATAACGCGAG 477 R Primer SGR2 AT4G11910 CACTCGCCCACTACTTCGTC
Supplemental Data. Benstein et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. Purification of the heterologously expressed PGDH1, PGDH2 and PGDH3 enzymes by Ni-NTA affinity chromatography. Protein extracts (2 µl) of different fractions (lane 1 = total extract,
Supplemental Figure 1. Purification of the heterologously expressed PGDH1, PGDH2 and PGDH3 enzymes by Ni-NTA affinity chromatography. Protein extracts (2 µl) of different fractions (lane 1 = total extract,
Supplemental Data. Na Xu et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Supplemental Figure 1. The weak fluorescence phenotype is not caused by the mutation in At3g60240. (A) A mutation mapped to the gene At3g60240. Map-based cloning strategy was used to map the mutated site
Supplemental Data. Tang et al. Plant Cell. (2012) /tpc
 Supplemental Figure 1. Relative Pchlide Fluorescence of Various Mutants and Wild Type. Seedlings were grown in darkness for 5 d. Experiments were repeated 3 times with same results. Supplemental Figure
Supplemental Figure 1. Relative Pchlide Fluorescence of Various Mutants and Wild Type. Seedlings were grown in darkness for 5 d. Experiments were repeated 3 times with same results. Supplemental Figure
Supplemental Data. Cui et al. (2012). Plant Cell /tpc a b c d. Stem UBC32 ACTIN
 A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
A Root Stem Leaf Flower Silique Senescence leaf B a b c d UBC32 ACTIN C * Supplemental Figure 1. Expression Pattern and Protein Sequence of UBC32 Homologues in Yeast, Human, and Arabidopsis. (A) Expression
Supplemental Data. Wu et al. (2). Plant Cell..5/tpc RGLG Hormonal treatment H2O B RGLG µm ABA µm ACC µm GA Time (hours) µm µm MJ µm IA
 Supplemental Data. Wu et al. (2). Plant Cell..5/tpc..4. A B Supplemental Figure. Immunoblot analysis verifies the expression of the AD-PP2C and BD-RGLG proteins in the Y2H assay. Total proteins were extracted
Supplemental Data. Wu et al. (2). Plant Cell..5/tpc..4. A B Supplemental Figure. Immunoblot analysis verifies the expression of the AD-PP2C and BD-RGLG proteins in the Y2H assay. Total proteins were extracted
Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination.
 Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination. Seeds of Col-0 were harvested from plants grown at 16 C, stored for 2 months, imbibed for indicated
Supplementary Figure 1 qrt-pcr expression analysis of NLP8 with and without KNO 3 during germination. Seeds of Col-0 were harvested from plants grown at 16 C, stored for 2 months, imbibed for indicated
Supplementary Information. c d e
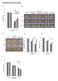 Supplementary Information a b c d e f Supplementary Figure 1. atabcg30, atabcg31, and atabcg40 mutant seeds germinate faster than the wild type on ½ MS medium supplemented with ABA (a and d-f) Germination
Supplementary Information a b c d e f Supplementary Figure 1. atabcg30, atabcg31, and atabcg40 mutant seeds germinate faster than the wild type on ½ MS medium supplemented with ABA (a and d-f) Germination
Supplementary Fig 1. The responses of ERF109 to different hormones and stresses. (a to k) The induced expression of ERF109 in 7-day-old Arabidopsis
 Supplementary Fig 1. The responses of ERF109 to different hormones and stresses. (a to k) The induced expression of ERF109 in 7-day-old Arabidopsis seedlings expressing ERF109pro-GUS. The GUS staining
Supplementary Fig 1. The responses of ERF109 to different hormones and stresses. (a to k) The induced expression of ERF109 in 7-day-old Arabidopsis seedlings expressing ERF109pro-GUS. The GUS staining
Supplementary Information
 Supplementary Information Supplementary Fig. 1. Seed dormancy and germination responses of RVE1 and PIF1. (a) Diagram of RVE1 and the T-DNA insertion of the rve1-2 mutant (SAIL_326_A01). Black boxes represent
Supplementary Information Supplementary Fig. 1. Seed dormancy and germination responses of RVE1 and PIF1. (a) Diagram of RVE1 and the T-DNA insertion of the rve1-2 mutant (SAIL_326_A01). Black boxes represent
Supplemental Data. Guo et al. (2015). Plant Cell /tpc
 Supplemental Figure 1. The Mutant exb1-d Displayed Pleiotropic Phenotypes and Produced Branches in the Axils of Cotyledons. (A) Branches were developed in exb1-d but not in wild-type plants. (B) and (C)
Supplemental Figure 1. The Mutant exb1-d Displayed Pleiotropic Phenotypes and Produced Branches in the Axils of Cotyledons. (A) Branches were developed in exb1-d but not in wild-type plants. (B) and (C)
SUPPLEMENTARY INFORMATION
 AS-NMD modulates FLM-dependent thermosensory flowering response in Arabidopsis NATURE PLANTS www.nature.com/natureplants 1 Supplementary Figure 1. Genomic sequence of FLM along with the splice sites. Sequencing
AS-NMD modulates FLM-dependent thermosensory flowering response in Arabidopsis NATURE PLANTS www.nature.com/natureplants 1 Supplementary Figure 1. Genomic sequence of FLM along with the splice sites. Sequencing
Supplementary Information
 Supplementary Information MED18 interaction with distinct transcription factors regulates plant immunity, flowering time and responses to hormones Supplementary Figure 1. Diagram showing T-DNA insertion
Supplementary Information MED18 interaction with distinct transcription factors regulates plant immunity, flowering time and responses to hormones Supplementary Figure 1. Diagram showing T-DNA insertion
Supplemental materials
 Supplemental materials Materials and methods for supplemental figures Yeast two-hybrid assays TAP46-PP2Ac interactions I. The TAP46 was used as the bait and the full-length cdnas of the five C subunits
Supplemental materials Materials and methods for supplemental figures Yeast two-hybrid assays TAP46-PP2Ac interactions I. The TAP46 was used as the bait and the full-length cdnas of the five C subunits
Supplemental Data. Wang et al. Plant Cell. (2013) /tpc
 SNL1 SNL Supplemental Figure 1. The Expression Patterns of SNL1 and SNL in Different Tissues of Arabidopsis from Genevestigator Web Site (https://www.genevestigator.com/gv/index.jsp). 1 A 1. 1..8.6.4..
SNL1 SNL Supplemental Figure 1. The Expression Patterns of SNL1 and SNL in Different Tissues of Arabidopsis from Genevestigator Web Site (https://www.genevestigator.com/gv/index.jsp). 1 A 1. 1..8.6.4..
Supporting Information
 Supporting Information Yuan et al. 10.1073/pnas.0906869106 Fig. S1. Heat map showing that Populus ICS is coregulated with orthologs of Arabidopsis genes involved in PhQ biosynthesis and PSI function, but
Supporting Information Yuan et al. 10.1073/pnas.0906869106 Fig. S1. Heat map showing that Populus ICS is coregulated with orthologs of Arabidopsis genes involved in PhQ biosynthesis and PSI function, but
Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified
 Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified by primers used for mrna expression analysis. Gray
Supplementary Figure 1. jmj30-2 and jmj32-1 produce null mutants. (a) Schematic drawing of JMJ30 and JMJ32 genome structure showing regions amplified by primers used for mrna expression analysis. Gray
Supplemental Data. Fan et al. (2014). Plant Cell /tpc
 Supplemental Data. Fan et al. (). Plant Cell./tpc.. Cell wall EXP EXP EXP9 HLH/bHLH AIF HFR HBI PAR ABAR Atg778 At3g39 Atg38 Atg3 EXP EXP6 EXP8 Photosynthesis PSAD- PSAD- PSBY PSBO- PSAN LHCB6 PSBS LHCB
Supplemental Data. Fan et al. (). Plant Cell./tpc.. Cell wall EXP EXP EXP9 HLH/bHLH AIF HFR HBI PAR ABAR Atg778 At3g39 Atg38 Atg3 EXP EXP6 EXP8 Photosynthesis PSAD- PSAD- PSBY PSBO- PSAN LHCB6 PSBS LHCB
Supplemental Figure S1. Validation of auxin-related gene expression from Fig. 1A. Relative gene expression in 2x and 3x seeds 6 days after
 Supplemental Figure S1. Validation of auxin-related gene expression from Fig. 1A. Relative gene expression in 2x and 3x seeds 6 days after pollination (6 DAP), as determined by RT-qPCR for YUC10 (A), YUC11
Supplemental Figure S1. Validation of auxin-related gene expression from Fig. 1A. Relative gene expression in 2x and 3x seeds 6 days after pollination (6 DAP), as determined by RT-qPCR for YUC10 (A), YUC11
Nature Genetics: doi: /ng Supplementary Figure 1. ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets.
 Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
Supplementary Figure 1 ChIP-seq genome browser views of BRM occupancy at previously identified BRM targets. Gene structures are shown underneath each panel. Supplementary Figure 2 pref6::ref6-gfp complements
- 1 - Supplemental Data
 - 1-1 Supplemental Data 2 3 4 5 6 7 8 9 Supplemental Figure S1. Differential expression of AtPIP Genes in DC3000-inoculated plants. Gene expression in leaves was analyzed by real-time RT-PCR and expression
- 1-1 Supplemental Data 2 3 4 5 6 7 8 9 Supplemental Figure S1. Differential expression of AtPIP Genes in DC3000-inoculated plants. Gene expression in leaves was analyzed by real-time RT-PCR and expression
A Repressor Complex Governs the Integration of
 Developmental Cell 15 Supplemental Data A Repressor Complex Governs the Integration of Flowering Signals in Arabidopsis Dan Li, Chang Liu, Lisha Shen, Yang Wu, Hongyan Chen, Masumi Robertson, Chris A.
Developmental Cell 15 Supplemental Data A Repressor Complex Governs the Integration of Flowering Signals in Arabidopsis Dan Li, Chang Liu, Lisha Shen, Yang Wu, Hongyan Chen, Masumi Robertson, Chris A.
Supplemental Data. Huo et al. (2013). Plant Cell /tpc
 Supplemental Data. Huo et al. (2013). Plant Cell 10.1105/tpc.112.108902 1 Supplemental Figure 1. Alignment of NCED4 Amino Acid Sequences from Different Lettuce Varieties. NCED4 amino acid sequences are
Supplemental Data. Huo et al. (2013). Plant Cell 10.1105/tpc.112.108902 1 Supplemental Figure 1. Alignment of NCED4 Amino Acid Sequences from Different Lettuce Varieties. NCED4 amino acid sequences are
Supplemental Materials
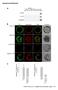 Supplemental Materials Flores-Pérez et al., Supplemental Materials, page 1 of 5 Supplemental Figure S1. Pull-down and BiFC controls, and quantitative analyses associated with the BiFC studies. (A) Controls
Supplemental Materials Flores-Pérez et al., Supplemental Materials, page 1 of 5 Supplemental Figure S1. Pull-down and BiFC controls, and quantitative analyses associated with the BiFC studies. (A) Controls
PIE1 ARP6 SWC6 KU70 ARP6 PIE1. HSA SNF2_N HELICc SANT. pie1-3 A1,A2 K1,K2 K1,K3 K3,LB2 A3, A4 A3,LB1 A1,A2 K1,K2 K1,K3. swc6-1 A3,A4.
 A B N-terminal SWC2 H2A.Z SWC6 ARP6 PIE1 HSA SNF2_N HELICc SANT C pie1-3 D PIE1 ARP6 5 Kb A1 200 bp A3 A2 LB1 arp6-3 A4 E A1,A2 A3, A4 A3,LB1 K1,K2 K1,K3 K3,LB2 SWC6 swc6-1 A1,A2 A3,A4 K1,K2 K1,K3 100
A B N-terminal SWC2 H2A.Z SWC6 ARP6 PIE1 HSA SNF2_N HELICc SANT C pie1-3 D PIE1 ARP6 5 Kb A1 200 bp A3 A2 LB1 arp6-3 A4 E A1,A2 A3, A4 A3,LB1 K1,K2 K1,K3 K3,LB2 SWC6 swc6-1 A1,A2 A3,A4 K1,K2 K1,K3 100
Supplementary Figures 1-12
 Supplementary Figures 1-12 Supplementary Figure 1. The specificity of anti-abi1 antibody. Total Proteins extracted from the wild type seedlings or abi1-3 null mutant seedlings were used for immunoblotting
Supplementary Figures 1-12 Supplementary Figure 1. The specificity of anti-abi1 antibody. Total Proteins extracted from the wild type seedlings or abi1-3 null mutant seedlings were used for immunoblotting
Supplemental Materials
 Supplemental Materials Supplemental Figure S. Phenotypic assessment of alb4 mutant plants under different stress conditions. (A) High-light stress and drought stress. Wild-type (WT) and alb4 mutant plants
Supplemental Materials Supplemental Figure S. Phenotypic assessment of alb4 mutant plants under different stress conditions. (A) High-light stress and drought stress. Wild-type (WT) and alb4 mutant plants
Alternative Cleavage and Polyadenylation of RNA
 Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
Developmental Cell 18 Supplemental Information The Spen Family Protein FPA Controls Alternative Cleavage and Polyadenylation of RNA Csaba Hornyik, Lionel C. Terzi, and Gordon G. Simpson Figure S1, related
Supplemental Data. Yang et al. (2014). Plant Cell /tpc
 Supplemental Figure 1. Activities of the Cm-BBX24 promoter in response to hormone treatments and abiotic stresses in transgenic Arabidopsis plants. (A) Cis-element analysis of the Cm-BBX24 promoter. (B)
Supplemental Figure 1. Activities of the Cm-BBX24 promoter in response to hormone treatments and abiotic stresses in transgenic Arabidopsis plants. (A) Cis-element analysis of the Cm-BBX24 promoter. (B)
Supplemental Figure 1. VLN5 retains conserved residues at both type 1 and type 2 Ca 2+ -binding
 Supplemental Figure 1. VLN5 retains conserved residues at both type 1 and type 2 Ca 2+ -binding sites in the G1 domain. Multiple sequence alignment was performed with DNAMAN6.0.40. Secondary structural
Supplemental Figure 1. VLN5 retains conserved residues at both type 1 and type 2 Ca 2+ -binding sites in the G1 domain. Multiple sequence alignment was performed with DNAMAN6.0.40. Secondary structural
Supplemental Data. Lee et al. Plant Cell. (2010) /tpc Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2.
 Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Supplemental Figure 1. Protein and Gene Structures of DWA1 and DWA2. (A) Protein structures of DWA1 and DWA2. WD40 region was determined based on the NCBI conserved domain databases (B, C) Schematic representation
Construction of plant complementation vector and generation of transgenic plants
 MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
MATERIAL S AND METHODS Plant materials and growth conditions Arabidopsis ecotype Columbia (Col0) was used for this study. SALK_072009, SALK_076309, and SALK_027645 were obtained from the Arabidopsis Biological
Supplemental Data. Zhang et al. Plant Cell (2014) /tpc
 Supplemental Data. Zhang et al. Plant Cell (214) 1.115/tpc.114.134163 55 - T C N SDIRIP1-GFP 35-25 - Psb 18 - Histone H3 Supplemental Figure 1. Detection of SDIRIP1-GFP in the nuclear fraction by Western
Supplemental Data. Zhang et al. Plant Cell (214) 1.115/tpc.114.134163 55 - T C N SDIRIP1-GFP 35-25 - Psb 18 - Histone H3 Supplemental Figure 1. Detection of SDIRIP1-GFP in the nuclear fraction by Western
Supplemental Data. Farmer et al. (2010) Plant Cell /tpc
 Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
Supplemental Figure 1. Amino acid sequence comparison of RAD23 proteins. Identical and similar residues are shown in the black and gray boxes, respectively. Dots denote gaps. The sequence of plant Ub is
Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon Song, Richard M. Amasino, Bosl Noh, and Yoo-Sun Noh
 Developmental Cell, Volume 22 Supplemental Information Control of Seed Germination by Light-Induced Histone Arginine Demethylation Activity Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon
Developmental Cell, Volume 22 Supplemental Information Control of Seed Germination by Light-Induced Histone Arginine Demethylation Activity Jung-Nam Cho, Jee-Youn Ryu, Young-Min Jeong, Jihye Park, Ji-Joon
A Nucleus-Encoded Chloroplast Protein YL1 Is Involved in Chloroplast. Development and Efficient Biogenesis of Chloroplast ATP Synthase in Rice
 A Nucleus-Encoded Chloroplast Protein YL1 Is Involved in Chloroplast Development and Efficient Biogenesis of Chloroplast ATP Synthase in Rice Fei Chen 1,*, Guojun Dong 2,*, Limin Wu 1, Fang Wang 3, Xingzheng
A Nucleus-Encoded Chloroplast Protein YL1 Is Involved in Chloroplast Development and Efficient Biogenesis of Chloroplast ATP Synthase in Rice Fei Chen 1,*, Guojun Dong 2,*, Limin Wu 1, Fang Wang 3, Xingzheng
Supplemental Data. Jing et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. Characterization of epp1 Mutants. (A) Cotyledon angles of 5-d-old Col wild-type (gray bars) and epp1-1 (black bars) seedlings under red (R), far-red (FR) and blue (BL) light conditions,
Supplemental Figure 1. Characterization of epp1 Mutants. (A) Cotyledon angles of 5-d-old Col wild-type (gray bars) and epp1-1 (black bars) seedlings under red (R), far-red (FR) and blue (BL) light conditions,
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature10928 Materials and Methods 1. Plant material and growth conditions. All plant lines used were in Col-0 background unless otherwise specified. pif4-101 mutant
SUPPLEMENTARY INFORMATION doi:10.1038/nature10928 Materials and Methods 1. Plant material and growth conditions. All plant lines used were in Col-0 background unless otherwise specified. pif4-101 mutant
Supplementary Materials
 Supplementary Materials Table S1. Oligonucleotide sequences and PCR conditions used to amplify the indicated genes. TA = annealing temperature; gdna = genomic DNA; cdna = complementary DNA; c = concentration.
Supplementary Materials Table S1. Oligonucleotide sequences and PCR conditions used to amplify the indicated genes. TA = annealing temperature; gdna = genomic DNA; cdna = complementary DNA; c = concentration.
Supplemental Data. Seo et al. (2014). Plant Cell /tpc
 Supplemental Figure 1. Protein alignment of ABD1 from other model organisms. The alignment was performed with H. sapiens DCAF8, M. musculus DCAF8 and O. sativa Os10g0544500. The WD40 domains are underlined.
Supplemental Figure 1. Protein alignment of ABD1 from other model organisms. The alignment was performed with H. sapiens DCAF8, M. musculus DCAF8 and O. sativa Os10g0544500. The WD40 domains are underlined.
Supplemental Data. Zhou et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. Confirmation of mutant mapping results. (A) Complementation assay with stably transformed genomic fragments (ComN-N) (2 kb upstream of TSS and 1.5 kb downstream of TES) and CaMV
Supplemental Figure 1. Confirmation of mutant mapping results. (A) Complementation assay with stably transformed genomic fragments (ComN-N) (2 kb upstream of TSS and 1.5 kb downstream of TES) and CaMV
A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 Journal of Plant Research A subclass of HSP70s regulate development and abiotic stress responses in Arabidopsis thaliana Linna Leng 1 Qianqian Liang
Supplemental Data. Hu et al. Plant Cell (2017) /tpc
 1 2 3 4 Supplemental Figure 1. DNA gel blot analysis of homozygous transgenic plants. (Supports Figure 1.) 5 6 7 8 Rice genomic DNA was digested with the restriction enzymes EcoRⅠ and BamHⅠ. Lanes in the
1 2 3 4 Supplemental Figure 1. DNA gel blot analysis of homozygous transgenic plants. (Supports Figure 1.) 5 6 7 8 Rice genomic DNA was digested with the restriction enzymes EcoRⅠ and BamHⅠ. Lanes in the
Supplementary information
 Supplementary information Supplementary figures Figure S1 Level of mycdet1 protein in DET1 OE-1, OE-2 and OE-3 transgenic lines. Total protein extract from wild type Col0, det1-1 mutant and DET1 OE lines
Supplementary information Supplementary figures Figure S1 Level of mycdet1 protein in DET1 OE-1, OE-2 and OE-3 transgenic lines. Total protein extract from wild type Col0, det1-1 mutant and DET1 OE lines
Intron distance to transcription start (nt)*
 Schwab et al. SOM: Enhanced microrna accumulation through stemloop-adjacent introns 1 Supplemental Table 1. Characteristics of introns used in this study Intron name Length intron (nt) Length cloned fragment
Schwab et al. SOM: Enhanced microrna accumulation through stemloop-adjacent introns 1 Supplemental Table 1. Characteristics of introns used in this study Intron name Length intron (nt) Length cloned fragment
Supplementary Materials: 1. Supplementary Figures S1-S9. 2. Supplementary Tables S1-S2
 Supplementary Materials: 1. Supplementary Figures S1-S9 2. Supplementary Tables S1-S2 S1 1. Supplementary Figures Fig. S1. Genotypes of tcp20 mutants. Fig. S2. Root phenotypes of tcp20 mutants. Fig. S3.
Supplementary Materials: 1. Supplementary Figures S1-S9 2. Supplementary Tables S1-S2 S1 1. Supplementary Figures Fig. S1. Genotypes of tcp20 mutants. Fig. S2. Root phenotypes of tcp20 mutants. Fig. S3.
Supplemental Table S1. Gene co-expression analysis using HHT/ASFT (At5g41040) as the bait*
 1 2 Supplemental Table S1. Gene co-expression analysis using HHT/ASFT (At5g41040) as the bait* 3 4 5 AGI ID R value GeneChip ID Annotation At5g41040 1.000 249289_at HXXXD-type acyl-transferase family protein
1 2 Supplemental Table S1. Gene co-expression analysis using HHT/ASFT (At5g41040) as the bait* 3 4 5 AGI ID R value GeneChip ID Annotation At5g41040 1.000 249289_at HXXXD-type acyl-transferase family protein
WiscDsLox485 ATG < > //----- E1 E2 E3 E4 E bp. Col-0 arr7 ARR7 ACTIN7. s of mrna/ng total RNA (x10 3 ) ARR7.
 A WiscDsLox8 ATG < > -9 +0 > ---------//----- E E E E E UTR +9 UTR +0 > 00bp B S D ol-0 arr7 ol-0 arr7 ARR7 ATIN7 D s of mrna/ng total RNA opie 0. ol-0 (x0 ) arr7 ARR7 Supplemental Fig.. Genotyping and
A WiscDsLox8 ATG < > -9 +0 > ---------//----- E E E E E UTR +9 UTR +0 > 00bp B S D ol-0 arr7 ol-0 arr7 ARR7 ATIN7 D s of mrna/ng total RNA opie 0. ol-0 (x0 ) arr7 ARR7 Supplemental Fig.. Genotyping and
Supplemental Data. Wu and Xue (2010). Plant Cell /tpc
 Supplemental Data. Wu and Xue (21). Plant Cell 1.115/tpc.11.75564 A P1-S P1-A P2-S P2-A P3-S P3-A P4-S P4-A B Relative expression Relative expression C 5. 4.5 4. 3.5 3. 2.5 2. 1.5 1..5 5. 4.5 4. 3.5 3.
Supplemental Data. Wu and Xue (21). Plant Cell 1.115/tpc.11.75564 A P1-S P1-A P2-S P2-A P3-S P3-A P4-S P4-A B Relative expression Relative expression C 5. 4.5 4. 3.5 3. 2.5 2. 1.5 1..5 5. 4.5 4. 3.5 3.
Supplemental Data. Liu et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. The GFP Tag Does Not Disturb the Physiological Functions of WDL3. (A) RT-PCR analysis of WDL3 expression in wild-type, WDL3-GFP, and WDL3 (without the GFP tag) transgenic seedlings.
Supplemental Figure 1. The GFP Tag Does Not Disturb the Physiological Functions of WDL3. (A) RT-PCR analysis of WDL3 expression in wild-type, WDL3-GFP, and WDL3 (without the GFP tag) transgenic seedlings.
Nature Genetics: doi: /ng.3556 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE. Supplementary Figure 1
 INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
INTEGRATED SUPPLEMENTARY FIGURE TEMPLATE Supplementary Figure 1 REF6 expression in transgenic lines. (a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT qpcr (a) and immunoblot
S156AT168AY175A (AAA) were purified as GST-fusion proteins and incubated with GSTfused
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 Supplemental Materials Supplemental Figure S1 (a) Phenotype of the wild type and grik1-2 grik2-1 plants after 8 days in darkness.
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 Supplemental Materials Supplemental Figure S1 (a) Phenotype of the wild type and grik1-2 grik2-1 plants after 8 days in darkness.
Chemical hijacking of auxin signaling with an engineered auxin-tir1
 1 SUPPLEMENTARY INFORMATION Chemical hijacking of auxin signaling with an engineered auxin-tir1 pair Naoyuki Uchida 1,2*, Koji Takahashi 2*, Rie Iwasaki 1, Ryotaro Yamada 2, Masahiko Yoshimura 2, Takaho
1 SUPPLEMENTARY INFORMATION Chemical hijacking of auxin signaling with an engineered auxin-tir1 pair Naoyuki Uchida 1,2*, Koji Takahashi 2*, Rie Iwasaki 1, Ryotaro Yamada 2, Masahiko Yoshimura 2, Takaho
Supplemental Figure 1. Transcript profiles of Arabidopsis IPMS1 and IPMS2 in different tissues and developmental stages.
 Transcript level (intensity) Supplemental Data. Xing et al. Plant Cell (2017) 10.1105/tpc.17.00186 1600 1400 1200 1000 800 600 400 200 0 IPMS1 IPMS2 Supplemental Figure 1. Transcript profiles of Arabidopsis
Transcript level (intensity) Supplemental Data. Xing et al. Plant Cell (2017) 10.1105/tpc.17.00186 1600 1400 1200 1000 800 600 400 200 0 IPMS1 IPMS2 Supplemental Figure 1. Transcript profiles of Arabidopsis
PrimePCR Assay Validation Report
 Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID Glyceraldehyde-3-phosphate dehydrogenase Gapdh Rat This gene encodes a member of
Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID Glyceraldehyde-3-phosphate dehydrogenase Gapdh Rat This gene encodes a member of
Technical Review. Real time PCR
 Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Technical Review Real time PCR Normal PCR: Analyze with agarose gel Normal PCR vs Real time PCR Real-time PCR, also known as quantitative PCR (qpcr) or kinetic PCR Key feature: Used to amplify and simultaneously
Supplementary Information
 Supplementary Information ER-localized auxin transporter PIN8 regulates auxin homeostasis and male gametophyte development in Arabidopsis Supplemental Figures a 8 6 4 2 Absolute Expression b Absolute Expression
Supplementary Information ER-localized auxin transporter PIN8 regulates auxin homeostasis and male gametophyte development in Arabidopsis Supplemental Figures a 8 6 4 2 Absolute Expression b Absolute Expression
M Col/1 Col/2 C24/1 C24/2 rpf2-1/1 rpf2-1/2
 M Col/1 Col/2 C24/1 C24/2 rpf2-1/1 rpf2-1/2 250 bp 260 bp -243 220 bp -202 Supplemental Figure S1: Detection of the -202 5 end of the nad9 mrna in Col, C24 and rpf2-1. CR-RT-PCR was performed with primers
M Col/1 Col/2 C24/1 C24/2 rpf2-1/1 rpf2-1/2 250 bp 260 bp -243 220 bp -202 Supplemental Figure S1: Detection of the -202 5 end of the nad9 mrna in Col, C24 and rpf2-1. CR-RT-PCR was performed with primers
Supplemental Figure 1. Alignment of the NbGAPC amino acid sequences with their Arabidopsis homologues.
 Supplemental Figure 1. Alignment of the NbGAPC amino acid sequences with their Arabidopsis homologues. Homologs from N. benthamiana (NbGAPC1, NbGAPC2, NbGAPC3), Arabidopsis (AtGAPC1, AT3G04120; AtGAPC2,
Supplemental Figure 1. Alignment of the NbGAPC amino acid sequences with their Arabidopsis homologues. Homologs from N. benthamiana (NbGAPC1, NbGAPC2, NbGAPC3), Arabidopsis (AtGAPC1, AT3G04120; AtGAPC2,
Supplemental Information
 Supplemental Information Supplemental Figure 1. The Heterologous Yeast System for Screening of Arabidopsis JA Transporters. (A) Exogenous JA inhibited yeast cell growth. Yeast cells were diluted to various
Supplemental Information Supplemental Figure 1. The Heterologous Yeast System for Screening of Arabidopsis JA Transporters. (A) Exogenous JA inhibited yeast cell growth. Yeast cells were diluted to various
Supplemental Data. Bai et al. Plant Cell. (2012) /tpc A
 A B Unconserved Conserved AIF4 AIF2 AIF3 AIF1 UPB1 IBH1 Supplemental Figure 1. IBH1, UPB1 and AIFs belong to the same HLH family. (A), Part of a phylogenetic tree constructed using conserved domains shows
A B Unconserved Conserved AIF4 AIF2 AIF3 AIF1 UPB1 IBH1 Supplemental Figure 1. IBH1, UPB1 and AIFs belong to the same HLH family. (A), Part of a phylogenetic tree constructed using conserved domains shows
sides of the aleurone (Al) but it is excluded from the basal endosperm transfer layer
 Supplemental Data. Gómez et al. (2009). The maize transcription factor MRP-1 (Myb-Related-Protein-1) is a key regulator of the differentiation of transfer cells. Supplemental Figure 1. Expression analyses
Supplemental Data. Gómez et al. (2009). The maize transcription factor MRP-1 (Myb-Related-Protein-1) is a key regulator of the differentiation of transfer cells. Supplemental Figure 1. Expression analyses
PrimePCR Assay Validation Report
 Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID glyceraldehyde-3-phosphate dehydrogenase GAPDH Human The product of this gene catalyzes
Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID glyceraldehyde-3-phosphate dehydrogenase GAPDH Human The product of this gene catalyzes
Experimental Tools and Resources Available in Arabidopsis. Manish Raizada, University of Guelph, Canada
 Experimental Tools and Resources Available in Arabidopsis Manish Raizada, University of Guelph, Canada Community website: The Arabidopsis Information Resource (TAIR) at http://www.arabidopsis.org Can order
Experimental Tools and Resources Available in Arabidopsis Manish Raizada, University of Guelph, Canada Community website: The Arabidopsis Information Resource (TAIR) at http://www.arabidopsis.org Can order
Supplemental Data. Sethi et al. (2014). Plant Cell /tpc
 Supplemental Data Supplemental Figure 1. MYC2 Binds to the E-box but not the E1-box of the MPK6 Promoter. (A) E1-box and E-box (wild type) containing MPK6 promoter fragment. The region shown in red denotes
Supplemental Data Supplemental Figure 1. MYC2 Binds to the E-box but not the E1-box of the MPK6 Promoter. (A) E1-box and E-box (wild type) containing MPK6 promoter fragment. The region shown in red denotes
Supplemental Figure legends Figure S1. (A) (B) (C) (D) Figure S2. Figure S3. (A-E) Figure S4. Figure S5. (A, C, E, G, I) (B, D, F, H, Figure S6.
 Supplemental Figure legends Figure S1. Map-based cloning and complementation testing for ZOP1. (A) ZOP1 was mapped to a ~273-kb interval on Chromosome 1. In the interval, a single-nucleotide G to A substitution
Supplemental Figure legends Figure S1. Map-based cloning and complementation testing for ZOP1. (A) ZOP1 was mapped to a ~273-kb interval on Chromosome 1. In the interval, a single-nucleotide G to A substitution
SUPPLEMENTARY INFORMATION
 Local auxin metabolism regulates environment-induced hypocotyl elongation Zuyu Zheng 1,2, Yongxia Guo 3, Ondřej Novák 4,5, William Chen 2, Karin Ljung 4, Joseph P. Noel 1,3, *, and Joanne Chory 1,2, *
Local auxin metabolism regulates environment-induced hypocotyl elongation Zuyu Zheng 1,2, Yongxia Guo 3, Ondřej Novák 4,5, William Chen 2, Karin Ljung 4, Joseph P. Noel 1,3, *, and Joanne Chory 1,2, *
Supplemental Materials. Zhu et al. RNAi suppression of DDM1 in poplar
 %mc Supplemental Materials Zhu et al. RNAi suppression of in poplar Supplemental Fig. S1 Total cellular cytosine methylation for selected events determined by HPLC. The association between methylcytosine
%mc Supplemental Materials Zhu et al. RNAi suppression of in poplar Supplemental Fig. S1 Total cellular cytosine methylation for selected events determined by HPLC. The association between methylcytosine
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description:
 File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. dcas9-mq1 fusion protein induces de novo
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. dcas9-mq1 fusion protein induces de novo
Supplemental Data. Álvarez et al. (2012). Plant Cell /tpc oas-a1.1. Ponceau!
 A! oas-a1.1 B! oas-a1.1 Col-0 des1-1! Free ATG8 proteins! ATG8-PE adducts! α-atg8! Ponceau! Supplemental Figure 1 online. Detec'on of SAVs (A) and (B) immunoblot analysis of ATG8 accumula'on in oas- a1.1
A! oas-a1.1 B! oas-a1.1 Col-0 des1-1! Free ATG8 proteins! ATG8-PE adducts! α-atg8! Ponceau! Supplemental Figure 1 online. Detec'on of SAVs (A) and (B) immunoblot analysis of ATG8 accumula'on in oas- a1.1
1. Introduction Drought stress and climate change Three strategies of plants in response to water stress 3
 Contents 1. Introduction 1 1.1 Drought stress and climate change 3 1.2 Three strategies of plants in response to water stress 3 1.3 Three closely related species of Linderniaceae family are experimental
Contents 1. Introduction 1 1.1 Drought stress and climate change 3 1.2 Three strategies of plants in response to water stress 3 1.3 Three closely related species of Linderniaceae family are experimental
6/256 1/256 0/256 1/256 2/256 7/256 10/256. At3g06290 (SAC3B)
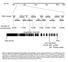 Chr.III M 5M 1M 15M 2M 23M BAC clones F22F7 F1A16 F24F17 F24P17 T8E24 F17A9 F21O3 F17A17 F18C1 F2O1 F28L1 F5E6 F3E22 T1B9 MLP3 Number of recombinants 6/256 1/256 /256 1/256 2/256 7/256 1/256 At3g629 (SAC3B)
Chr.III M 5M 1M 15M 2M 23M BAC clones F22F7 F1A16 F24F17 F24P17 T8E24 F17A9 F21O3 F17A17 F18C1 F2O1 F28L1 F5E6 F3E22 T1B9 MLP3 Number of recombinants 6/256 1/256 /256 1/256 2/256 7/256 1/256 At3g629 (SAC3B)
Gene expression analysis. Biosciences 741: Genomics Fall, 2013 Week 5. Gene expression analysis
 Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Gene expression analysis Biosciences 741: Genomics Fall, 2013 Week 5 Gene expression analysis From EST clusters to spotted cdna microarrays Long vs. short oligonucleotide microarrays vs. RT-PCR Methods
Functional Genomics Research Stream. Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update
 Functional Genomics Research Stream Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update Updates Alternate Lab Meeting Fridays 11:30-1:00 WEL 4.224 Welcome to attend either one Lab Log thanks
Functional Genomics Research Stream Research Meeting: June 19, 2012 SYBR Green qpcr, Research Update Updates Alternate Lab Meeting Fridays 11:30-1:00 WEL 4.224 Welcome to attend either one Lab Log thanks
Nature Genetics: doi: /ng Supplementary Figure 1. High-confidence PRC2 targets and candidate PREs.
 Supplementary Figure 1 High-confidence PRC2 targets and candidate PREs. (a) Flowchart for identification of candidate Arabidopsis PREs. We identified 1504 genomic regions marked by at least 3 of the following:
Supplementary Figure 1 High-confidence PRC2 targets and candidate PREs. (a) Flowchart for identification of candidate Arabidopsis PREs. We identified 1504 genomic regions marked by at least 3 of the following:
Design. Construction. Characterization
 Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Design Construction Characterization DNA mrna (messenger) A C C transcription translation C A C protein His A T G C T A C G Plasmids replicon copy number incompatibility selection marker origin of replication
Supplementary Information. The flowering gene SINGLE FLOWER TRUSS drives heterosis for yield in tomato
 Supplementary Information The flowering gene SINGLE FLOWER TRUSS drives heterosis for yield in tomato Uri Krieger 1, Zachary B. Lippman 2 *, and Dani Zamir 1 * 1. The Hebrew University of Jerusalem Faculty
Supplementary Information The flowering gene SINGLE FLOWER TRUSS drives heterosis for yield in tomato Uri Krieger 1, Zachary B. Lippman 2 *, and Dani Zamir 1 * 1. The Hebrew University of Jerusalem Faculty
(phosphatase tensin) domain is shown in dark gray, the FH1 domain in black, and the
 Supplemental Figure 1. Predicted Domain Organization of the AFH14 Protein. (A) Schematic representation of the predicted domain organization of AFH14. The PTEN (phosphatase tensin) domain is shown in dark
Supplemental Figure 1. Predicted Domain Organization of the AFH14 Protein. (A) Schematic representation of the predicted domain organization of AFH14. The PTEN (phosphatase tensin) domain is shown in dark
To investigate the heredity of the WFP gene, we selected plants that were homozygous
 Supplementary information Supplementary Note ST-12 WFP allele is semi-dominant To investigate the heredity of the WFP gene, we selected plants that were homozygous for chromosome 1 of Nipponbare and heterozygous
Supplementary information Supplementary Note ST-12 WFP allele is semi-dominant To investigate the heredity of the WFP gene, we selected plants that were homozygous for chromosome 1 of Nipponbare and heterozygous
SUPPLEMENTARY INFORMATION
 Ca 2+ /calmodulin Regulates Salicylic Acid-mediated Plant Immunity Liqun Du, Gul S. Ali, Kayla A. Simons, Jingguo Hou, Tianbao Yang, A.S.N. Reddy and B. W. Poovaiah * *To whom correspondence should be
Ca 2+ /calmodulin Regulates Salicylic Acid-mediated Plant Immunity Liqun Du, Gul S. Ali, Kayla A. Simons, Jingguo Hou, Tianbao Yang, A.S.N. Reddy and B. W. Poovaiah * *To whom correspondence should be
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity.
 Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
Supplemental Figure 1 HDA18 has an HDAC domain and therefore has concentration dependent and TSA inhibited histone deacetylase activity. (A) Amino acid alignment of HDA5, HDA15 and HDA18. The blue line
Supplemental Figure 1. Conserved regions of the kinase domain of PEPR1, PEPR2, CLV1 and BRI1.
 Supplemental Figure 1. Conserved regions of the kinase domain of PEPR1, PEPR2, CLV1 and BRI1. The asterisks and colons indicate the important residues for ATP-binding pocket and substrate binding pocket,
Supplemental Figure 1. Conserved regions of the kinase domain of PEPR1, PEPR2, CLV1 and BRI1. The asterisks and colons indicate the important residues for ATP-binding pocket and substrate binding pocket,
David M. Rocke Division of Biostatistics and Department of Biomedical Engineering University of California, Davis
 David M. Rocke Division of Biostatistics and Department of Biomedical Engineering University of California, Davis Outline RNA-Seq for differential expression analysis Statistical methods for RNA-Seq: Structure
David M. Rocke Division of Biostatistics and Department of Biomedical Engineering University of California, Davis Outline RNA-Seq for differential expression analysis Statistical methods for RNA-Seq: Structure
Table S1. List of primers used in this study.
 Gene/Construct/ mutant Primer name Sequence (5 ->3 ) Confimation of T-DNA insertion clh1-1 LBb1 5 GCGTGGACCGCTTGCTGCAACT3 1LP1 5 CCGAAAATGATAAATGCATGG3 1RP1 5 ATGTCCAGCTCGAAAGATTCC3 clh2-1 Lb4 5 CTACAAATTGCCTTTTCTTATCGAC3
Gene/Construct/ mutant Primer name Sequence (5 ->3 ) Confimation of T-DNA insertion clh1-1 LBb1 5 GCGTGGACCGCTTGCTGCAACT3 1LP1 5 CCGAAAATGATAAATGCATGG3 1RP1 5 ATGTCCAGCTCGAAAGATTCC3 clh2-1 Lb4 5 CTACAAATTGCCTTTTCTTATCGAC3
PrimePCR Assay Validation Report
 Gene Information Gene Name minichromosome maintenance complex component 8 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID MCM8 Human The protein encoded by
Gene Information Gene Name minichromosome maintenance complex component 8 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID MCM8 Human The protein encoded by
Supplemental Figure 1. Identification of the sid2 ald1 (sid2-1 ald1) double mutant.
 Supplemental Data. Bernsdorff et al. (216). Plant Cell 1.115/tpc.15.496-1 A B C Supplemental Figure 1. Identification of the sid2 ald1 (sid2-1 ald1) double mutant. (A) PCR analysis of genomic DNA isolated
Supplemental Data. Bernsdorff et al. (216). Plant Cell 1.115/tpc.15.496-1 A B C Supplemental Figure 1. Identification of the sid2 ald1 (sid2-1 ald1) double mutant. (A) PCR analysis of genomic DNA isolated
Supplemental Data. Challa et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. DEX-induced TCP4 activity rescues jaw-d phenotypes. (A) and (B) Rosettess of 32-day-old Col-0;ProTCP4:mTCP4:GR plants (A) and jaw-d;protcp4:mtcp4:gr (B) plants grown in mock or in
Supplemental Figure 1. DEX-induced TCP4 activity rescues jaw-d phenotypes. (A) and (B) Rosettess of 32-day-old Col-0;ProTCP4:mTCP4:GR plants (A) and jaw-d;protcp4:mtcp4:gr (B) plants grown in mock or in
Supplemental Data. Borg et al. Plant Cell (2014) /tpc
 Supplementary Figure 1 - Alignment of selected angiosperm DAZ1 and DAZ2 homologs Multiple sequence alignment of selected DAZ1 and DAZ2 homologs. A consensus sequence built using default parameters is shown
Supplementary Figure 1 - Alignment of selected angiosperm DAZ1 and DAZ2 homologs Multiple sequence alignment of selected DAZ1 and DAZ2 homologs. A consensus sequence built using default parameters is shown
Supplemental Data. Wang et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. Time-Course Analysis of Storage Proteins during Endosperm Development of the Wild-type 9311 and the gpa4-1 Mutant. (A) SDS-PAGE analyses of seed storage proteins during wild-type
Supplemental Figure 1. Time-Course Analysis of Storage Proteins during Endosperm Development of the Wild-type 9311 and the gpa4-1 Mutant. (A) SDS-PAGE analyses of seed storage proteins during wild-type
mirna Purification from Plant Tissues: Corn, Soybean and Arabidopsis
 mirna Purification from Plant Tissues: Corn, Soybean and Arabidopsis A Maxwell RSC mirna Tissue Kit Application Note Materials Required: Maxwell RSC mirna Tissue Kit (Cat.# AS1480) QuantiFluor RNA System
mirna Purification from Plant Tissues: Corn, Soybean and Arabidopsis A Maxwell RSC mirna Tissue Kit Application Note Materials Required: Maxwell RSC mirna Tissue Kit (Cat.# AS1480) QuantiFluor RNA System
Supplemental Data. mir156-regulated SPL Transcription. Factors Define an Endogenous Flowering. Pathway in Arabidopsis thaliana
 Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Cell, Volume 138 Supplemental Data mir156-regulated SPL Transcription Factors Define an Endogenous Flowering Pathway in Arabidopsis thaliana Jia-Wei Wang, Benjamin Czech, and Detlef Weigel Table S1. Interaction
Supplementary Data 1.
 Supplementary Data 1. Evaluation of the effects of number of F2 progeny to be bulked (n) and average sequencing coverage (depth) of the genome (G) on the levels of false positive SNPs (SNP index = 1).
Supplementary Data 1. Evaluation of the effects of number of F2 progeny to be bulked (n) and average sequencing coverage (depth) of the genome (G) on the levels of false positive SNPs (SNP index = 1).
Supplemental Figure 1. Mutation in NLA Causes Increased Pi Uptake Activity and
 Supplemental Figure 1. Mutation in NLA Causes Increased Pi Uptake Activity and PHT1 Protein Amounts. (A) Shoot morphology of 19-day-old nla mutants under Pi-sufficient conditions. (B) [ 33 P]Pi uptake
Supplemental Figure 1. Mutation in NLA Causes Increased Pi Uptake Activity and PHT1 Protein Amounts. (A) Shoot morphology of 19-day-old nla mutants under Pi-sufficient conditions. (B) [ 33 P]Pi uptake
PrimePCR Assay Validation Report
 Gene Information Gene Name integrin, alpha 1 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID ITGA1 Human This gene encodes the alpha 1 subunit of integrin
Gene Information Gene Name integrin, alpha 1 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID ITGA1 Human This gene encodes the alpha 1 subunit of integrin
PrimePCR Assay Validation Report
 Gene Information Gene Name 3-hydroxybutyrate dehydrogenase, type 1 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID BDH1 Human This gene encodes a member of
Gene Information Gene Name 3-hydroxybutyrate dehydrogenase, type 1 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID BDH1 Human This gene encodes a member of
Supplemental Data. Pillitteri et al. (2011) /tpc
 A B C D Supplemental Figure 1. Images of representative 5-dag seedlings used in this study.(a) WT; (B) spch; (C) scrm-d; (D) scrm-d. mute. Images were taken under the same magnification. Scale bar, 1 mm.
A B C D Supplemental Figure 1. Images of representative 5-dag seedlings used in this study.(a) WT; (B) spch; (C) scrm-d; (D) scrm-d. mute. Images were taken under the same magnification. Scale bar, 1 mm.
Fig. S1. Molecular phylogenetic analysis of AtHD-ZIP IV family. A phylogenetic tree was constructed using Bayesian analysis with Markov Chain Monte
 Fig. S1. Molecular phylogenetic analysis of AtHD-ZIP IV family. A phylogenetic tree was constructed using Bayesian analysis with Markov Chain Monte Carlo algorithm for one million generations to obtain
Fig. S1. Molecular phylogenetic analysis of AtHD-ZIP IV family. A phylogenetic tree was constructed using Bayesian analysis with Markov Chain Monte Carlo algorithm for one million generations to obtain
Aromatase Activity and CYP19 Gene Expression in male adult Xenopus laevis Exposed to Atrazine
 Aromatase Activity and CYP19 Gene Expression in male adult Xenopus laevis Exposed to Atrazine June-Woo Park*, Markus Hecker*, Margaret Murphy*, Paul Jones*, Glen van der Kraak**, Keith Solomon**, Ron Kendall***,
Aromatase Activity and CYP19 Gene Expression in male adult Xenopus laevis Exposed to Atrazine June-Woo Park*, Markus Hecker*, Margaret Murphy*, Paul Jones*, Glen van der Kraak**, Keith Solomon**, Ron Kendall***,
SUPPLEMENTAL DATA. Supplementary data
 1 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 SUPPLEMENTAL DATA Supplementary data Figure S1: Verification of pla-iα1 knockdown lines a: Semiquantitative
1 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 SUPPLEMENTAL DATA Supplementary data Figure S1: Verification of pla-iα1 knockdown lines a: Semiquantitative
Relative Quantification: Data Management & Analysis Settings
 Relative Quantification: Data Management & Analysis Settings Ray Meng, Ph.D. International Field Applications Specialist Gene Expression Division Bio-Rad Laboratories Gene Expression AMPLIFICATION Assay
Relative Quantification: Data Management & Analysis Settings Ray Meng, Ph.D. International Field Applications Specialist Gene Expression Division Bio-Rad Laboratories Gene Expression AMPLIFICATION Assay
Quantitative Real Time PCR USING SYBR GREEN
 Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
Quantitative Real Time PCR USING SYBR GREEN SYBR Green SYBR Green is a cyanine dye that binds to double stranded DNA. When it is bound to D.S. DNA it has a much greater fluorescence than when bound to
